Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
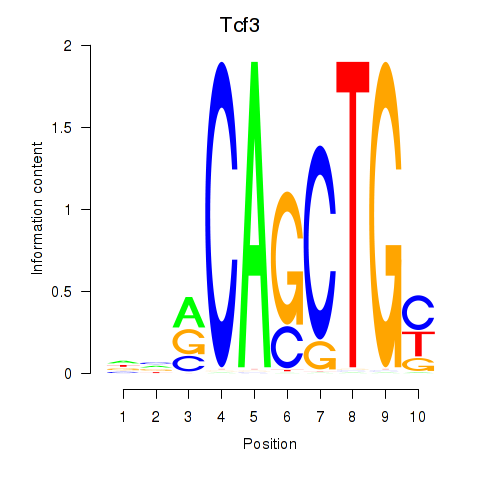
Results for Tcf3
Z-value: 1.29
Transcription factors associated with Tcf3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Tcf3
|
ENSMUSG00000020167.15 | Tcf3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Tcf3 | mm39_v1_chr10_-_80269436_80269488 | 0.32 | 6.1e-03 | Click! |
Activity profile of Tcf3 motif
Sorted Z-values of Tcf3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Tcf3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_43523595 | 17.03 |
ENSMUST00000107914.10
|
Tpm2
|
tropomyosin 2, beta |
| chr4_-_43523745 | 16.68 |
ENSMUST00000150592.2
|
Tpm2
|
tropomyosin 2, beta |
| chr4_-_43523388 | 15.72 |
ENSMUST00000107913.10
ENSMUST00000030184.12 |
Tpm2
|
tropomyosin 2, beta |
| chr11_-_100036792 | 8.60 |
ENSMUST00000007317.8
|
Krt19
|
keratin 19 |
| chr8_+_46338498 | 7.56 |
ENSMUST00000034053.7
|
Pdlim3
|
PDZ and LIM domain 3 |
| chr1_+_75336965 | 7.43 |
ENSMUST00000027409.10
|
Des
|
desmin |
| chr2_+_131028861 | 7.19 |
ENSMUST00000028804.15
ENSMUST00000079857.9 |
Cdc25b
|
cell division cycle 25B |
| chr8_+_23629080 | 6.91 |
ENSMUST00000033947.15
|
Ank1
|
ankyrin 1, erythroid |
| chr8_+_46338557 | 6.88 |
ENSMUST00000210422.2
|
Pdlim3
|
PDZ and LIM domain 3 |
| chr8_-_112417633 | 6.87 |
ENSMUST00000034435.7
|
Ctrb1
|
chymotrypsinogen B1 |
| chr16_+_57173456 | 6.75 |
ENSMUST00000159816.8
|
Filip1l
|
filamin A interacting protein 1-like |
| chr15_-_101268036 | 6.69 |
ENSMUST00000077196.6
|
Krt80
|
keratin 80 |
| chr2_-_28453374 | 6.47 |
ENSMUST00000028161.6
|
Cel
|
carboxyl ester lipase |
| chr7_+_142025575 | 6.29 |
ENSMUST00000038946.9
|
Lsp1
|
lymphocyte specific 1 |
| chr7_+_142025817 | 6.27 |
ENSMUST00000105966.2
|
Lsp1
|
lymphocyte specific 1 |
| chr6_-_41291634 | 5.91 |
ENSMUST00000064324.12
|
Try5
|
trypsin 5 |
| chr6_+_30639217 | 5.79 |
ENSMUST00000031806.10
|
Cpa1
|
carboxypeptidase A1, pancreatic |
| chr6_+_17307639 | 5.67 |
ENSMUST00000115453.2
|
Cav1
|
caveolin 1, caveolae protein |
| chr7_+_140659672 | 5.62 |
ENSMUST00000066873.5
ENSMUST00000163041.2 |
Pkp3
|
plakophilin 3 |
| chr8_+_23629046 | 5.41 |
ENSMUST00000121075.8
|
Ank1
|
ankyrin 1, erythroid |
| chr7_+_141995545 | 5.29 |
ENSMUST00000105971.8
ENSMUST00000145287.8 |
Tnni2
|
troponin I, skeletal, fast 2 |
| chr6_+_17307272 | 5.25 |
ENSMUST00000115454.2
|
Cav1
|
caveolin 1, caveolae protein |
| chr17_+_75485791 | 5.19 |
ENSMUST00000135447.8
ENSMUST00000112516.8 |
Ltbp1
|
latent transforming growth factor beta binding protein 1 |
| chr11_+_98555167 | 5.18 |
ENSMUST00000017348.3
|
Gsdma
|
gasdermin A |
| chr8_-_106660470 | 5.16 |
ENSMUST00000034368.8
|
Ctrl
|
chymotrypsin-like |
| chr16_-_56706494 | 4.68 |
ENSMUST00000023435.6
|
Tmem45a
|
transmembrane protein 45a |
| chr7_-_105131407 | 4.55 |
ENSMUST00000047040.4
|
Cavin3
|
caveolae associated 3 |
| chr3_+_106393348 | 4.43 |
ENSMUST00000183271.2
|
Dennd2d
|
DENN/MADD domain containing 2D |
| chr1_+_135252508 | 4.43 |
ENSMUST00000059352.3
|
Lmod1
|
leiomodin 1 (smooth muscle) |
| chr14_-_70415117 | 4.42 |
ENSMUST00000022681.11
|
Pdlim2
|
PDZ and LIM domain 2 |
| chr17_+_75485906 | 4.34 |
ENSMUST00000112514.2
|
Ltbp1
|
latent transforming growth factor beta binding protein 1 |
| chr7_+_141996067 | 4.32 |
ENSMUST00000149529.8
|
Tnni2
|
troponin I, skeletal, fast 2 |
| chr3_-_20329823 | 4.25 |
ENSMUST00000011607.6
|
Cpb1
|
carboxypeptidase B1 (tissue) |
| chr19_+_5118103 | 4.25 |
ENSMUST00000070630.8
|
Cd248
|
CD248 antigen, endosialin |
| chr9_+_21249118 | 4.22 |
ENSMUST00000034697.8
|
Slc44a2
|
solute carrier family 44, member 2 |
| chr15_-_101759212 | 4.14 |
ENSMUST00000023790.5
|
Krt1
|
keratin 1 |
| chr13_-_111945499 | 4.12 |
ENSMUST00000109267.9
|
Map3k1
|
mitogen-activated protein kinase kinase kinase 1 |
| chr14_+_30601157 | 4.11 |
ENSMUST00000040715.8
|
Mustn1
|
musculoskeletal, embryonic nuclear protein 1 |
| chr13_-_55676334 | 4.04 |
ENSMUST00000047877.5
|
Dok3
|
docking protein 3 |
| chr4_+_42255693 | 4.01 |
ENSMUST00000178864.3
|
Ccl21b
|
chemokine (C-C motif) ligand 21B (leucine) |
| chr2_-_125348305 | 4.00 |
ENSMUST00000028633.13
|
Fbn1
|
fibrillin 1 |
| chrX_-_111608339 | 3.99 |
ENSMUST00000039887.4
|
Pof1b
|
premature ovarian failure 1B |
| chr4_+_41903610 | 3.91 |
ENSMUST00000098128.4
|
Ccl21d
|
chemokine (C-C motif) ligand 21D |
| chr15_-_101588714 | 3.88 |
ENSMUST00000023786.7
|
Krt6b
|
keratin 6B |
| chr4_+_42114817 | 3.87 |
ENSMUST00000098123.4
|
Gm13304
|
predicted gene 13304 |
| chr6_-_68609426 | 3.86 |
ENSMUST00000103328.3
|
Igkv10-96
|
immunoglobulin kappa variable 10-96 |
| chr11_+_69856222 | 3.85 |
ENSMUST00000018713.13
|
Cldn7
|
claudin 7 |
| chr15_-_101621332 | 3.81 |
ENSMUST00000023709.7
|
Krt5
|
keratin 5 |
| chr12_+_51640097 | 3.72 |
ENSMUST00000164782.10
ENSMUST00000085412.7 |
Coch
|
cochlin |
| chr6_-_69792108 | 3.71 |
ENSMUST00000103367.3
|
Igkv12-44
|
immunoglobulin kappa variable 12-44 |
| chr2_-_91854844 | 3.69 |
ENSMUST00000028663.5
|
Creb3l1
|
cAMP responsive element binding protein 3-like 1 |
| chr1_+_177270101 | 3.69 |
ENSMUST00000194319.2
|
Zbtb18
|
zinc finger and BTB domain containing 18 |
| chr12_-_114672701 | 3.62 |
ENSMUST00000103505.3
ENSMUST00000193855.2 |
Ighv1-19
|
immunoglobulin heavy variable V1-19 |
| chr1_-_79649683 | 3.61 |
ENSMUST00000162342.8
|
Ap1s3
|
adaptor-related protein complex AP-1, sigma 3 |
| chr4_+_97665992 | 3.59 |
ENSMUST00000107062.9
ENSMUST00000052018.12 ENSMUST00000107057.8 |
Nfia
|
nuclear factor I/A |
| chr17_-_36014892 | 3.56 |
ENSMUST00000097333.10
ENSMUST00000003628.13 |
Ddr1
|
discoidin domain receptor family, member 1 |
| chr12_-_114901026 | 3.53 |
ENSMUST00000103516.2
ENSMUST00000191868.2 |
Ighv1-42
|
immunoglobulin heavy variable V1-42 |
| chr19_-_57185988 | 3.50 |
ENSMUST00000099294.9
|
Ablim1
|
actin-binding LIM protein 1 |
| chr3_-_100396635 | 3.47 |
ENSMUST00000061455.9
|
Tent5c
|
terminal nucleotidyltransferase 5C |
| chr11_-_3477916 | 3.45 |
ENSMUST00000020718.10
|
Smtn
|
smoothelin |
| chr5_+_21748523 | 3.44 |
ENSMUST00000035651.6
|
Lrrc17
|
leucine rich repeat containing 17 |
| chr7_-_126224848 | 3.41 |
ENSMUST00000032961.4
|
Nupr1
|
nuclear protein transcription regulator 1 |
| chr6_-_68713748 | 3.40 |
ENSMUST00000183936.2
ENSMUST00000196863.2 |
Igkv19-93
|
immunoglobulin kappa chain variable 19-93 |
| chr2_-_35322581 | 3.40 |
ENSMUST00000079424.11
|
Ggta1
|
glycoprotein galactosyltransferase alpha 1, 3 |
| chr9_-_31824758 | 3.34 |
ENSMUST00000116615.5
|
Barx2
|
BarH-like homeobox 2 |
| chr16_-_19019100 | 3.33 |
ENSMUST00000103749.3
|
Iglc2
|
immunoglobulin lambda constant 2 |
| chr11_+_115790768 | 3.32 |
ENSMUST00000152171.8
|
Smim5
|
small integral membrane protein 5 |
| chr4_+_140737955 | 3.31 |
ENSMUST00000071977.9
|
Mfap2
|
microfibrillar-associated protein 2 |
| chr1_+_78286946 | 3.30 |
ENSMUST00000036172.10
|
Sgpp2
|
sphingosine-1-phosphate phosphatase 2 |
| chrX_-_47551990 | 3.25 |
ENSMUST00000033429.9
ENSMUST00000140486.2 |
Elf4
|
E74-like factor 4 (ets domain transcription factor) |
| chr7_-_44180700 | 3.25 |
ENSMUST00000205506.2
|
Spib
|
Spi-B transcription factor (Spi-1/PU.1 related) |
| chr3_+_89344006 | 3.25 |
ENSMUST00000038942.10
ENSMUST00000130858.8 ENSMUST00000146630.8 ENSMUST00000145753.2 |
Pbxip1
|
pre B cell leukemia transcription factor interacting protein 1 |
| chr7_+_44866635 | 3.14 |
ENSMUST00000097216.5
ENSMUST00000209343.2 ENSMUST00000209678.2 |
Tead2
|
TEA domain family member 2 |
| chr10_+_128745214 | 3.11 |
ENSMUST00000220308.2
|
Cd63
|
CD63 antigen |
| chr8_-_106198112 | 3.09 |
ENSMUST00000014990.13
|
Tppp3
|
tubulin polymerization-promoting protein family member 3 |
| chr15_+_78810919 | 3.09 |
ENSMUST00000089377.6
|
Lgals1
|
lectin, galactose binding, soluble 1 |
| chrX_+_100492684 | 3.08 |
ENSMUST00000033674.6
|
Itgb1bp2
|
integrin beta 1 binding protein 2 |
| chr13_-_60325170 | 3.06 |
ENSMUST00000065086.6
|
Gas1
|
growth arrest specific 1 |
| chr5_-_98178811 | 3.02 |
ENSMUST00000031281.14
|
Antxr2
|
anthrax toxin receptor 2 |
| chr17_-_48739874 | 2.99 |
ENSMUST00000046549.5
|
Apobec2
|
apolipoprotein B mRNA editing enzyme, catalytic polypeptide 2 |
| chr5_+_17779721 | 2.96 |
ENSMUST00000169603.2
|
Sema3c
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3C |
| chr6_-_68812291 | 2.96 |
ENSMUST00000199143.5
ENSMUST00000103335.3 |
Igkv12-89
|
immunoglobulin kappa chain variable 12-89 |
| chr2_-_118593087 | 2.87 |
ENSMUST00000059997.9
|
Ccdc9b
|
coiled-coil domain containing 9B |
| chr7_-_89176294 | 2.86 |
ENSMUST00000207932.2
|
Prss23
|
protease, serine 23 |
| chr2_-_152775122 | 2.86 |
ENSMUST00000099200.3
|
Foxs1
|
forkhead box S1 |
| chr8_+_3715747 | 2.86 |
ENSMUST00000014118.4
|
Mcemp1
|
mast cell expressed membrane protein 1 |
| chr5_-_113968483 | 2.84 |
ENSMUST00000100874.6
|
Selplg
|
selectin, platelet (p-selectin) ligand |
| chr2_+_49677688 | 2.81 |
ENSMUST00000028103.13
|
Lypd6b
|
LY6/PLAUR domain containing 6B |
| chrX_-_56384089 | 2.80 |
ENSMUST00000033468.11
|
Arhgef6
|
Rac/Cdc42 guanine nucleotide exchange factor (GEF) 6 |
| chr4_+_129181407 | 2.80 |
ENSMUST00000102599.4
|
Sync
|
syncoilin |
| chr19_-_11243530 | 2.79 |
ENSMUST00000169159.3
|
Ms4a1
|
membrane-spanning 4-domains, subfamily A, member 1 |
| chr6_-_134609350 | 2.79 |
ENSMUST00000047443.5
|
Mansc1
|
MANSC domain containing 1 |
| chr12_-_115299134 | 2.78 |
ENSMUST00000195359.6
ENSMUST00000103530.3 |
Ighv1-59
|
immunoglobulin heavy variable V1-59 |
| chr7_-_109322993 | 2.78 |
ENSMUST00000106735.9
ENSMUST00000033334.5 |
BC051019
|
cDNA sequence BC051019 |
| chr7_+_19024994 | 2.74 |
ENSMUST00000108468.5
|
Rtn2
|
reticulon 2 (Z-band associated protein) |
| chr6_-_67919524 | 2.74 |
ENSMUST00000196768.2
|
Igkv9-124
|
immunoglobulin kappa chain variable 9-124 |
| chr11_-_46057224 | 2.73 |
ENSMUST00000020679.3
|
Nipal4
|
NIPA-like domain containing 4 |
| chr3_+_146558114 | 2.71 |
ENSMUST00000170055.8
ENSMUST00000037942.11 |
Ttll7
|
tubulin tyrosine ligase-like family, member 7 |
| chr3_+_95496239 | 2.71 |
ENSMUST00000177390.8
ENSMUST00000060323.12 ENSMUST00000098861.11 |
Golph3l
|
golgi phosphoprotein 3-like |
| chr17_+_35413415 | 2.69 |
ENSMUST00000025262.6
ENSMUST00000173600.2 |
Ltb
|
lymphotoxin B |
| chr3_+_95496270 | 2.69 |
ENSMUST00000176674.8
ENSMUST00000177389.8 ENSMUST00000176755.8 ENSMUST00000177399.2 |
Golph3l
|
golgi phosphoprotein 3-like |
| chr12_-_114646685 | 2.68 |
ENSMUST00000194350.6
ENSMUST00000103504.3 |
Ighv1-18
|
immunoglobulin heavy variable V1-18 |
| chr12_-_115172211 | 2.68 |
ENSMUST00000103526.3
|
Ighv1-55
|
immunoglobulin heavy variable 1-55 |
| chr15_+_99489018 | 2.67 |
ENSMUST00000229728.2
ENSMUST00000231163.2 |
Aqp5
|
aquaporin 5 |
| chr3_-_151971391 | 2.65 |
ENSMUST00000199470.5
ENSMUST00000200589.5 |
Nexn
|
nexilin |
| chr19_-_57185928 | 2.63 |
ENSMUST00000111544.8
|
Ablim1
|
actin-binding LIM protein 1 |
| chr4_-_141325517 | 2.63 |
ENSMUST00000131317.8
ENSMUST00000006381.11 ENSMUST00000129602.8 |
Fblim1
|
filamin binding LIM protein 1 |
| chr8_-_123425805 | 2.63 |
ENSMUST00000127984.9
|
Cbfa2t3
|
CBFA2/RUNX1 translocation partner 3 |
| chr12_+_9624437 | 2.61 |
ENSMUST00000057021.9
|
Osr1
|
odd-skipped related transcription factor 1 |
| chr11_-_102255999 | 2.60 |
ENSMUST00000006749.10
|
Slc4a1
|
solute carrier family 4 (anion exchanger), member 1 |
| chr7_-_131756447 | 2.58 |
ENSMUST00000033149.5
|
Cpxm2
|
carboxypeptidase X 2 (M14 family) |
| chr17_+_75772475 | 2.57 |
ENSMUST00000095204.6
|
Rasgrp3
|
RAS, guanyl releasing protein 3 |
| chr19_-_45548942 | 2.56 |
ENSMUST00000026239.7
|
Poll
|
polymerase (DNA directed), lambda |
| chr19_-_57185861 | 2.55 |
ENSMUST00000111550.8
|
Ablim1
|
actin-binding LIM protein 1 |
| chr2_+_174602412 | 2.54 |
ENSMUST00000029030.9
|
Edn3
|
endothelin 3 |
| chr9_-_58065800 | 2.53 |
ENSMUST00000168864.4
|
Islr
|
immunoglobulin superfamily containing leucine-rich repeat |
| chr4_+_141148068 | 2.52 |
ENSMUST00000102486.5
|
Hspb7
|
heat shock protein family, member 7 (cardiovascular) |
| chr12_-_114878652 | 2.52 |
ENSMUST00000103515.2
|
Ighv1-39
|
immunoglobulin heavy variable 1-39 |
| chr19_-_57185808 | 2.52 |
ENSMUST00000111546.8
|
Ablim1
|
actin-binding LIM protein 1 |
| chr8_+_23629173 | 2.49 |
ENSMUST00000174435.2
|
Ank1
|
ankyrin 1, erythroid |
| chrX_-_72478938 | 2.47 |
ENSMUST00000033738.8
|
Trex2
|
three prime repair exonuclease 2 |
| chr19_+_6952580 | 2.45 |
ENSMUST00000237084.2
ENSMUST00000236218.2 ENSMUST00000237235.2 |
Ppp1r14b
|
protein phosphatase 1, regulatory inhibitor subunit 14B |
| chr4_-_130068902 | 2.44 |
ENSMUST00000105998.8
|
Tinagl1
|
tubulointerstitial nephritis antigen-like 1 |
| chr19_-_6065181 | 2.44 |
ENSMUST00000236537.2
ENSMUST00000025891.11 |
Capn1
|
calpain 1 |
| chr4_-_133600308 | 2.42 |
ENSMUST00000137486.3
|
Rps6ka1
|
ribosomal protein S6 kinase polypeptide 1 |
| chr10_-_78554104 | 2.42 |
ENSMUST00000005488.9
|
Casp14
|
caspase 14 |
| chr6_-_69741999 | 2.42 |
ENSMUST00000103365.3
|
Igkv12-46
|
immunoglobulin kappa variable 12-46 |
| chr11_+_70548622 | 2.40 |
ENSMUST00000170716.8
|
Eno3
|
enolase 3, beta muscle |
| chr6_+_70648743 | 2.39 |
ENSMUST00000103401.3
|
Igkv3-4
|
immunoglobulin kappa variable 3-4 |
| chr6_+_41331039 | 2.36 |
ENSMUST00000072103.7
|
Try10
|
trypsin 10 |
| chr2_+_103800459 | 2.31 |
ENSMUST00000111143.8
ENSMUST00000138815.2 |
Lmo2
|
LIM domain only 2 |
| chr2_+_103254465 | 2.31 |
ENSMUST00000171693.8
|
Elf5
|
E74-like factor 5 |
| chr3_-_75177378 | 2.30 |
ENSMUST00000039047.5
|
Serpini2
|
serine (or cysteine) peptidase inhibitor, clade I, member 2 |
| chr5_-_98178834 | 2.27 |
ENSMUST00000199088.2
|
Antxr2
|
anthrax toxin receptor 2 |
| chr11_+_96820091 | 2.25 |
ENSMUST00000054311.6
ENSMUST00000107636.4 |
Prr15l
|
proline rich 15-like |
| chr5_+_123214332 | 2.24 |
ENSMUST00000067505.15
ENSMUST00000111619.10 ENSMUST00000160344.2 |
Tmem120b
|
transmembrane protein 120B |
| chr17_+_75772567 | 2.24 |
ENSMUST00000234660.2
|
Rasgrp3
|
RAS, guanyl releasing protein 3 |
| chr8_+_72050292 | 2.22 |
ENSMUST00000143662.8
|
Niban3
|
niban apoptosis regulator 3 |
| chr15_-_78657640 | 2.22 |
ENSMUST00000018313.6
|
Mfng
|
MFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr7_+_19144950 | 2.21 |
ENSMUST00000208710.2
ENSMUST00000003643.3 |
Ckm
|
creatine kinase, muscle |
| chr11_+_63019799 | 2.21 |
ENSMUST00000108702.8
|
Pmp22
|
peripheral myelin protein 22 |
| chr1_-_171061838 | 2.21 |
ENSMUST00000193973.2
|
Fcer1g
|
Fc receptor, IgE, high affinity I, gamma polypeptide |
| chr6_-_69835868 | 2.19 |
ENSMUST00000103369.2
|
Igkv12-41
|
immunoglobulin kappa chain variable 12-41 |
| chr10_-_120037464 | 2.17 |
ENSMUST00000020448.11
|
Irak3
|
interleukin-1 receptor-associated kinase 3 |
| chr15_-_77527470 | 2.17 |
ENSMUST00000181154.2
ENSMUST00000180949.8 ENSMUST00000166623.10 |
Apol11b
|
apolipoprotein L 11b |
| chr15_-_36598263 | 2.16 |
ENSMUST00000155116.2
|
Pabpc1
|
poly(A) binding protein, cytoplasmic 1 |
| chr14_+_26760898 | 2.15 |
ENSMUST00000035336.5
|
Il17rd
|
interleukin 17 receptor D |
| chr1_-_192880260 | 2.15 |
ENSMUST00000161367.2
|
Traf3ip3
|
TRAF3 interacting protein 3 |
| chr17_-_71309012 | 2.14 |
ENSMUST00000128179.2
ENSMUST00000150456.2 ENSMUST00000233357.2 ENSMUST00000233417.2 |
Myl12a
Gm49909
|
myosin, light chain 12A, regulatory, non-sarcomeric predicted gene, 49909 |
| chr7_-_80037153 | 2.13 |
ENSMUST00000206728.2
|
Fes
|
feline sarcoma oncogene |
| chr2_+_103800553 | 2.12 |
ENSMUST00000111140.3
ENSMUST00000111139.3 |
Lmo2
|
LIM domain only 2 |
| chr19_+_6952319 | 2.11 |
ENSMUST00000070850.8
|
Ppp1r14b
|
protein phosphatase 1, regulatory inhibitor subunit 14B |
| chr7_+_3339077 | 2.11 |
ENSMUST00000203566.3
|
Myadm
|
myeloid-associated differentiation marker |
| chr7_+_3339059 | 2.11 |
ENSMUST00000096744.8
|
Myadm
|
myeloid-associated differentiation marker |
| chr10_-_128579879 | 2.10 |
ENSMUST00000026414.9
|
Dgka
|
diacylglycerol kinase, alpha |
| chr9_-_105837842 | 2.09 |
ENSMUST00000190193.8
ENSMUST00000165165.3 |
Col6a5
|
collagen, type VI, alpha 5 |
| chr19_+_8975249 | 2.08 |
ENSMUST00000236390.2
|
Ahnak
|
AHNAK nucleoprotein (desmoyokin) |
| chr1_-_153061758 | 2.08 |
ENSMUST00000185356.7
|
Lamc2
|
laminin, gamma 2 |
| chr7_+_27770655 | 2.07 |
ENSMUST00000138392.8
ENSMUST00000076648.8 |
Fcgbp
|
Fc fragment of IgG binding protein |
| chr3_-_151871867 | 2.06 |
ENSMUST00000046614.10
|
Gipc2
|
GIPC PDZ domain containing family, member 2 |
| chr15_-_77653979 | 2.04 |
ENSMUST00000229259.2
|
Myh9
|
myosin, heavy polypeptide 9, non-muscle |
| chr11_-_101357046 | 2.03 |
ENSMUST00000040430.8
|
Vat1
|
vesicle amine transport 1 |
| chr2_-_113883285 | 2.03 |
ENSMUST00000090269.7
|
Actc1
|
actin, alpha, cardiac muscle 1 |
| chr12_-_114451189 | 2.03 |
ENSMUST00000103493.3
|
Ighv1-4
|
immunoglobulin heavy variable 1-4 |
| chr15_+_61857226 | 2.02 |
ENSMUST00000161976.8
ENSMUST00000022971.8 |
Myc
|
myelocytomatosis oncogene |
| chr2_+_125876566 | 2.02 |
ENSMUST00000064794.14
|
Fgf7
|
fibroblast growth factor 7 |
| chr1_+_135060431 | 2.02 |
ENSMUST00000187985.7
ENSMUST00000049449.11 |
Ptpn7
|
protein tyrosine phosphatase, non-receptor type 7 |
| chr3_-_151541603 | 2.01 |
ENSMUST00000106126.2
|
Ptgfr
|
prostaglandin F receptor |
| chr9_-_107863062 | 2.01 |
ENSMUST00000048568.6
|
Inka1
|
inka box actin regulator 1 |
| chr6_-_124715618 | 2.01 |
ENSMUST00000171549.9
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr2_-_93164812 | 1.99 |
ENSMUST00000111265.9
|
Tspan18
|
tetraspanin 18 |
| chr5_+_99002293 | 1.98 |
ENSMUST00000031278.6
ENSMUST00000200388.2 |
Bmp3
|
bone morphogenetic protein 3 |
| chr6_-_124715560 | 1.98 |
ENSMUST00000004377.15
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr6_-_52222776 | 1.97 |
ENSMUST00000048026.10
|
Hoxa11
|
homeobox A11 |
| chr15_+_102927366 | 1.96 |
ENSMUST00000165375.3
|
Hoxc4
|
homeobox C4 |
| chr11_-_7163897 | 1.96 |
ENSMUST00000020702.11
ENSMUST00000135887.3 |
Igfbp3
|
insulin-like growth factor binding protein 3 |
| chr8_+_83891972 | 1.96 |
ENSMUST00000034145.11
|
Tbc1d9
|
TBC1 domain family, member 9 |
| chr6_-_148345834 | 1.94 |
ENSMUST00000060095.15
ENSMUST00000100772.10 |
Tmtc1
|
transmembrane and tetratricopeptide repeat containing 1 |
| chr6_+_67873135 | 1.94 |
ENSMUST00000103310.3
|
Igkv14-126
|
immunoglobulin kappa variable 14-126 |
| chr3_+_89090437 | 1.94 |
ENSMUST00000140473.2
ENSMUST00000041913.13 |
Fam189b
|
family with sequence similarity 189, member B |
| chr13_+_37529184 | 1.94 |
ENSMUST00000021860.7
|
Ly86
|
lymphocyte antigen 86 |
| chr8_+_36561982 | 1.93 |
ENSMUST00000110492.2
|
Prag1
|
PEAK1 related kinase activating pseudokinase 1 |
| chr6_+_112436466 | 1.93 |
ENSMUST00000075477.8
|
Cav3
|
caveolin 3 |
| chr11_+_11636213 | 1.92 |
ENSMUST00000076700.11
ENSMUST00000048122.13 |
Ikzf1
|
IKAROS family zinc finger 1 |
| chr11_+_115790951 | 1.92 |
ENSMUST00000142089.2
ENSMUST00000131566.2 |
Smim5
|
small integral membrane protein 5 |
| chrX_-_144131890 | 1.91 |
ENSMUST00000040084.10
ENSMUST00000123443.2 |
Lhfpl1
|
lipoma HMGIC fusion partner-like 1 |
| chr12_-_115884332 | 1.91 |
ENSMUST00000103548.3
|
Ighv1-81
|
immunoglobulin heavy variable 1-81 |
| chr4_-_133694607 | 1.91 |
ENSMUST00000105893.8
|
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr10_+_75729237 | 1.90 |
ENSMUST00000009236.6
ENSMUST00000217811.2 |
Derl3
|
Der1-like domain family, member 3 |
| chr10_+_126899468 | 1.90 |
ENSMUST00000120226.8
ENSMUST00000133115.8 |
Cdk4
|
cyclin-dependent kinase 4 |
| chr17_-_73706284 | 1.89 |
ENSMUST00000095208.4
|
Capn13
|
calpain 13 |
| chr5_-_134643805 | 1.89 |
ENSMUST00000202085.2
ENSMUST00000036362.13 ENSMUST00000077636.8 |
Lat2
|
linker for activation of T cells family, member 2 |
| chr9_+_37439367 | 1.88 |
ENSMUST00000002011.14
|
Esam
|
endothelial cell-specific adhesion molecule |
| chr6_-_3494587 | 1.88 |
ENSMUST00000049985.15
|
Hepacam2
|
HEPACAM family member 2 |
| chr6_-_41354538 | 1.88 |
ENSMUST00000096003.7
|
Prss3
|
protease, serine 3 |
| chr19_+_53128901 | 1.88 |
ENSMUST00000235754.2
ENSMUST00000237301.2 ENSMUST00000238130.2 |
Add3
|
adducin 3 (gamma) |
| chr6_-_87473260 | 1.88 |
ENSMUST00000101197.9
|
Arhgap25
|
Rho GTPase activating protein 25 |
| chr11_-_100288566 | 1.87 |
ENSMUST00000001592.15
ENSMUST00000107403.2 |
Jup
|
junction plakoglobin |
| chr13_-_56326511 | 1.86 |
ENSMUST00000169652.3
|
Tifab
|
TRAF-interacting protein with forkhead-associated domain, family member B |
| chr7_+_126811831 | 1.86 |
ENSMUST00000127710.3
|
Mylpf
|
myosin light chain, phosphorylatable, fast skeletal muscle |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 12.6 | GO:1900085 | negative regulation of peptidyl-tyrosine autophosphorylation(GO:1900085) negative regulation of inward rectifier potassium channel activity(GO:1903609) |
| 1.2 | 7.5 | GO:0002159 | desmosome assembly(GO:0002159) |
| 1.2 | 3.6 | GO:0061033 | secretion by lung epithelial cell involved in lung growth(GO:0061033) |
| 1.2 | 3.5 | GO:0098583 | mastication(GO:0071626) learned vocalization behavior(GO:0098583) |
| 1.1 | 4.5 | GO:1901003 | negative regulation of fermentation(GO:1901003) |
| 1.1 | 5.5 | GO:0072186 | metanephric cap development(GO:0072185) metanephric cap morphogenesis(GO:0072186) metanephric cap mesenchymal cell proliferation involved in metanephros development(GO:0090094) |
| 1.0 | 3.1 | GO:0034117 | erythrocyte aggregation(GO:0034117) regulation of erythrocyte aggregation(GO:0034118) |
| 1.0 | 4.0 | GO:0035582 | sequestering of BMP in extracellular matrix(GO:0035582) |
| 1.0 | 3.0 | GO:0003350 | pulmonary myocardium development(GO:0003350) |
| 0.9 | 2.8 | GO:0033082 | regulation of extrathymic T cell differentiation(GO:0033082) |
| 0.9 | 3.4 | GO:0042335 | cuticle development(GO:0042335) |
| 0.8 | 3.4 | GO:0042125 | protein glycosylation at cell surface(GO:0033575) protein galactosylation at cell surface(GO:0033580) protein galactosylation(GO:0042125) |
| 0.8 | 7.2 | GO:0007144 | female meiosis I(GO:0007144) |
| 0.7 | 3.7 | GO:0070278 | extracellular matrix constituent secretion(GO:0070278) |
| 0.7 | 2.9 | GO:0032765 | positive regulation of mast cell cytokine production(GO:0032765) |
| 0.7 | 5.8 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.7 | 1.4 | GO:0003032 | detection of oxygen(GO:0003032) |
| 0.7 | 2.0 | GO:1903465 | motogenic signaling involved in postnatal olfactory bulb interneuron migration(GO:0021837) positive regulation of mitotic cell cycle DNA replication(GO:1903465) |
| 0.6 | 1.9 | GO:0045660 | positive regulation of neutrophil differentiation(GO:0045660) |
| 0.6 | 0.6 | GO:0060715 | syncytiotrophoblast cell differentiation involved in labyrinthine layer development(GO:0060715) |
| 0.6 | 4.1 | GO:0001712 | ectodermal cell fate commitment(GO:0001712) |
| 0.6 | 1.8 | GO:0002017 | regulation of blood volume by renal aldosterone(GO:0002017) |
| 0.6 | 1.7 | GO:0060032 | floor plate formation(GO:0021508) notochord regression(GO:0060032) |
| 0.6 | 2.8 | GO:0050902 | leukocyte adhesive activation(GO:0050902) |
| 0.6 | 1.7 | GO:0002378 | immunoglobulin biosynthetic process(GO:0002378) |
| 0.6 | 1.7 | GO:0072268 | pattern specification involved in metanephros development(GO:0072268) |
| 0.6 | 1.7 | GO:0021682 | nerve maturation(GO:0021682) |
| 0.5 | 1.6 | GO:0098903 | membrane repolarization during action potential(GO:0086011) regulation of membrane repolarization during action potential(GO:0098903) |
| 0.5 | 1.6 | GO:2000402 | negative regulation of lymphocyte migration(GO:2000402) |
| 0.5 | 4.2 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.5 | 1.6 | GO:1903904 | negative regulation of establishment of T cell polarity(GO:1903904) |
| 0.5 | 3.1 | GO:2000680 | regulation of rubidium ion transport(GO:2000680) |
| 0.5 | 4.1 | GO:1902732 | positive regulation of chondrocyte proliferation(GO:1902732) |
| 0.5 | 3.1 | GO:0003100 | regulation of systemic arterial blood pressure by endothelin(GO:0003100) vein smooth muscle contraction(GO:0014826) |
| 0.5 | 2.0 | GO:1903921 | protein processing in phagocytic vesicle(GO:1900756) regulation of protein processing in phagocytic vesicle(GO:1903921) positive regulation of protein processing in phagocytic vesicle(GO:1903923) |
| 0.5 | 1.5 | GO:2000832 | negative regulation of steroid hormone secretion(GO:2000832) |
| 0.5 | 2.0 | GO:0071798 | response to prostaglandin D(GO:0071798) cellular response to prostaglandin D stimulus(GO:0071799) |
| 0.5 | 4.4 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.5 | 6.3 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.5 | 1.4 | GO:0097374 | sensory neuron axon guidance(GO:0097374) |
| 0.5 | 1.4 | GO:0061193 | taste bud development(GO:0061193) |
| 0.4 | 3.6 | GO:0042713 | sperm ejaculation(GO:0042713) |
| 0.4 | 1.8 | GO:0060988 | lipid tube assembly(GO:0060988) |
| 0.4 | 5.7 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.4 | 3.9 | GO:2000491 | positive regulation of hepatic stellate cell activation(GO:2000491) |
| 0.4 | 3.4 | GO:0048539 | bone marrow development(GO:0048539) |
| 0.4 | 2.6 | GO:0006287 | base-excision repair, gap-filling(GO:0006287) |
| 0.4 | 3.8 | GO:0045084 | positive regulation of interleukin-12 biosynthetic process(GO:0045084) |
| 0.4 | 1.3 | GO:0002874 | regulation of chronic inflammatory response to antigenic stimulus(GO:0002874) |
| 0.4 | 1.2 | GO:0051385 | response to mineralocorticoid(GO:0051385) |
| 0.4 | 1.2 | GO:0010615 | positive regulation of cardiac muscle adaptation(GO:0010615) positive regulation of cardiac muscle hypertrophy in response to stress(GO:1903244) |
| 0.4 | 0.8 | GO:0002572 | pro-T cell differentiation(GO:0002572) |
| 0.4 | 13.2 | GO:0045109 | intermediate filament organization(GO:0045109) |
| 0.4 | 2.3 | GO:0031444 | slow-twitch skeletal muscle fiber contraction(GO:0031444) |
| 0.4 | 1.5 | GO:0000379 | tRNA-type intron splice site recognition and cleavage(GO:0000379) |
| 0.4 | 1.5 | GO:0033088 | negative regulation of immature T cell proliferation in thymus(GO:0033088) |
| 0.4 | 1.8 | GO:0070268 | cornification(GO:0070268) |
| 0.4 | 1.5 | GO:0032972 | regulation of muscle filament sliding speed(GO:0032972) |
| 0.4 | 2.2 | GO:0010936 | negative regulation of macrophage cytokine production(GO:0010936) |
| 0.4 | 7.8 | GO:0046514 | ceramide catabolic process(GO:0046514) |
| 0.3 | 2.4 | GO:0010288 | response to lead ion(GO:0010288) |
| 0.3 | 2.0 | GO:0010637 | negative regulation of mitochondrial fusion(GO:0010637) |
| 0.3 | 5.4 | GO:0048194 | Golgi vesicle budding(GO:0048194) |
| 0.3 | 1.0 | GO:1900108 | negative regulation of nodal signaling pathway(GO:1900108) |
| 0.3 | 1.3 | GO:2001137 | positive regulation of endocytic recycling(GO:2001137) |
| 0.3 | 3.3 | GO:0001866 | NK T cell proliferation(GO:0001866) |
| 0.3 | 14.8 | GO:0006779 | porphyrin-containing compound biosynthetic process(GO:0006779) |
| 0.3 | 1.9 | GO:1904180 | negative regulation of membrane depolarization(GO:1904180) |
| 0.3 | 4.1 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.3 | 9.5 | GO:0032967 | positive regulation of collagen biosynthetic process(GO:0032967) |
| 0.3 | 1.3 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 0.3 | 3.1 | GO:0032224 | positive regulation of synaptic transmission, cholinergic(GO:0032224) |
| 0.3 | 1.6 | GO:0061056 | sclerotome development(GO:0061056) |
| 0.3 | 2.2 | GO:2000622 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.3 | 0.9 | GO:0002426 | immunoglobulin production in mucosal tissue(GO:0002426) |
| 0.3 | 5.5 | GO:0072189 | ureter development(GO:0072189) |
| 0.3 | 1.2 | GO:0002036 | regulation of L-glutamate transport(GO:0002036) |
| 0.3 | 4.6 | GO:0060033 | anatomical structure regression(GO:0060033) |
| 0.3 | 0.9 | GO:0030327 | prenylated protein catabolic process(GO:0030327) |
| 0.3 | 0.8 | GO:0072054 | renal outer medulla development(GO:0072054) regulation of chromatin-mediated maintenance of transcription(GO:1904499) positive regulation of chromatin-mediated maintenance of transcription(GO:1904501) regulation of euchromatin binding(GO:1904793) |
| 0.3 | 0.8 | GO:2000451 | positive regulation of CD8-positive, alpha-beta T cell extravasation(GO:2000451) |
| 0.3 | 2.0 | GO:0009786 | regulation of asymmetric cell division(GO:0009786) |
| 0.3 | 0.8 | GO:1990426 | homologous recombination-dependent replication fork processing(GO:1990426) |
| 0.3 | 1.1 | GO:0000415 | negative regulation of histone H3-K36 methylation(GO:0000415) |
| 0.3 | 4.4 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 0.3 | 1.0 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.3 | 1.3 | GO:0048069 | eye pigmentation(GO:0048069) |
| 0.3 | 2.6 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.2 | 0.7 | GO:0002270 | plasmacytoid dendritic cell activation(GO:0002270) |
| 0.2 | 0.7 | GO:0097065 | anterior head development(GO:0097065) regulation of anterior head development(GO:2000742) positive regulation of anterior head development(GO:2000744) |
| 0.2 | 2.7 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 0.2 | 2.6 | GO:1903715 | regulation of aerobic respiration(GO:1903715) |
| 0.2 | 1.9 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.2 | 0.7 | GO:0014735 | regulation of muscle atrophy(GO:0014735) death-inducing signaling complex assembly(GO:0071550) negative regulation of mitochondrial membrane permeability involved in apoptotic process(GO:1902109) |
| 0.2 | 3.7 | GO:0048368 | lateral mesoderm development(GO:0048368) |
| 0.2 | 2.1 | GO:0042637 | catagen(GO:0042637) |
| 0.2 | 34.5 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.2 | 1.1 | GO:0038032 | termination of G-protein coupled receptor signaling pathway(GO:0038032) |
| 0.2 | 1.8 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.2 | 0.7 | GO:1903334 | positive regulation of protein folding(GO:1903334) |
| 0.2 | 0.7 | GO:0035638 | patched ligand maturation(GO:0007225) signal maturation(GO:0035638) |
| 0.2 | 1.3 | GO:0051533 | positive regulation of NFAT protein import into nucleus(GO:0051533) |
| 0.2 | 1.5 | GO:0044336 | canonical Wnt signaling pathway involved in negative regulation of apoptotic process(GO:0044336) |
| 0.2 | 0.9 | GO:0090472 | dibasic protein processing(GO:0090472) |
| 0.2 | 1.7 | GO:0010897 | negative regulation of triglyceride catabolic process(GO:0010897) |
| 0.2 | 10.1 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.2 | 1.5 | GO:0014816 | skeletal muscle satellite cell differentiation(GO:0014816) |
| 0.2 | 2.7 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.2 | 0.6 | GO:0051305 | chromosome movement towards spindle pole(GO:0051305) |
| 0.2 | 0.6 | GO:1903347 | negative regulation of bicellular tight junction assembly(GO:1903347) |
| 0.2 | 1.8 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) |
| 0.2 | 2.0 | GO:0030240 | skeletal muscle thin filament assembly(GO:0030240) |
| 0.2 | 3.9 | GO:0032463 | negative regulation of protein homooligomerization(GO:0032463) |
| 0.2 | 3.6 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.2 | 0.8 | GO:0098886 | modification of dendritic spine(GO:0098886) |
| 0.2 | 2.1 | GO:0035878 | nail development(GO:0035878) |
| 0.2 | 3.4 | GO:2000194 | regulation of female gonad development(GO:2000194) |
| 0.2 | 0.9 | GO:2000584 | regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000583) negative regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000584) |
| 0.2 | 0.5 | GO:0001698 | gastrin-induced gastric acid secretion(GO:0001698) |
| 0.2 | 2.0 | GO:0060272 | embryonic skeletal joint morphogenesis(GO:0060272) |
| 0.2 | 2.7 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 0.2 | 0.5 | GO:0070103 | regulation of interleukin-6-mediated signaling pathway(GO:0070103) |
| 0.2 | 8.1 | GO:0060706 | cell differentiation involved in embryonic placenta development(GO:0060706) |
| 0.2 | 1.4 | GO:0006564 | L-serine biosynthetic process(GO:0006564) |
| 0.2 | 3.3 | GO:0006670 | sphingosine metabolic process(GO:0006670) |
| 0.2 | 0.7 | GO:1902202 | regulation of hepatocyte growth factor receptor signaling pathway(GO:1902202) |
| 0.2 | 0.5 | GO:0060676 | ureteric bud formation(GO:0060676) |
| 0.2 | 3.1 | GO:0060628 | regulation of ER to Golgi vesicle-mediated transport(GO:0060628) |
| 0.2 | 3.6 | GO:0045103 | intermediate filament-based process(GO:0045103) |
| 0.2 | 2.6 | GO:0033623 | regulation of integrin activation(GO:0033623) |
| 0.2 | 1.3 | GO:0030035 | microspike assembly(GO:0030035) |
| 0.2 | 2.7 | GO:0090308 | regulation of methylation-dependent chromatin silencing(GO:0090308) |
| 0.2 | 1.7 | GO:0033631 | cell-cell adhesion mediated by integrin(GO:0033631) |
| 0.2 | 0.9 | GO:0014043 | negative regulation of neuron maturation(GO:0014043) |
| 0.2 | 0.8 | GO:0071921 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 0.2 | 3.2 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 0.2 | 1.1 | GO:0006868 | glutamine transport(GO:0006868) |
| 0.2 | 0.6 | GO:1901675 | negative regulation of histone H3-K27 acetylation(GO:1901675) |
| 0.2 | 0.3 | GO:0060450 | positive regulation of hindgut contraction(GO:0060450) |
| 0.2 | 2.7 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.2 | 2.1 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.2 | 1.4 | GO:1900262 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.2 | 0.5 | GO:0014908 | myotube differentiation involved in skeletal muscle regeneration(GO:0014908) |
| 0.2 | 0.8 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.1 | 0.9 | GO:1903566 | positive regulation of protein localization to cilium(GO:1903566) |
| 0.1 | 1.6 | GO:0003344 | pericardium morphogenesis(GO:0003344) |
| 0.1 | 1.5 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.1 | 1.3 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.1 | 3.6 | GO:0040034 | regulation of development, heterochronic(GO:0040034) |
| 0.1 | 2.1 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.1 | 0.3 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.1 | 1.5 | GO:0045059 | positive thymic T cell selection(GO:0045059) |
| 0.1 | 2.5 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.1 | 1.6 | GO:0045618 | positive regulation of keratinocyte differentiation(GO:0045618) |
| 0.1 | 2.3 | GO:0043586 | tongue development(GO:0043586) |
| 0.1 | 0.9 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.1 | 1.3 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 0.1 | 2.4 | GO:0097205 | renal filtration(GO:0097205) |
| 0.1 | 0.5 | GO:1901187 | regulation of ephrin receptor signaling pathway(GO:1901187) |
| 0.1 | 0.7 | GO:0044727 | DNA demethylation of male pronucleus(GO:0044727) |
| 0.1 | 13.4 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.1 | 3.2 | GO:0061029 | eyelid development in camera-type eye(GO:0061029) |
| 0.1 | 2.3 | GO:1904776 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.1 | 3.3 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.1 | 45.5 | GO:0006936 | muscle contraction(GO:0006936) |
| 0.1 | 2.1 | GO:0046339 | phosphatidic acid biosynthetic process(GO:0006654) diacylglycerol metabolic process(GO:0046339) |
| 0.1 | 0.2 | GO:0021524 | visceral motor neuron differentiation(GO:0021524) peripheral nervous system neuron axonogenesis(GO:0048936) |
| 0.1 | 14.1 | GO:0007586 | digestion(GO:0007586) |
| 0.1 | 1.6 | GO:0006301 | postreplication repair(GO:0006301) |
| 0.1 | 0.2 | GO:0001705 | ectoderm formation(GO:0001705) |
| 0.1 | 2.2 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.1 | 0.9 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) |
| 0.1 | 0.1 | GO:2000137 | negative regulation of cell proliferation involved in heart morphogenesis(GO:2000137) |
| 0.1 | 1.1 | GO:0043922 | negative regulation by host of viral transcription(GO:0043922) |
| 0.1 | 2.2 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.1 | 0.3 | GO:0021664 | rhombomere formation(GO:0021594) rhombomere 3 formation(GO:0021660) rhombomere 5 morphogenesis(GO:0021664) rhombomere 5 formation(GO:0021666) |
| 0.1 | 1.6 | GO:0045586 | regulation of gamma-delta T cell differentiation(GO:0045586) regulation of gamma-delta T cell activation(GO:0046643) |
| 0.1 | 0.7 | GO:0010669 | epithelial structure maintenance(GO:0010669) |
| 0.1 | 1.5 | GO:0071803 | positive regulation of podosome assembly(GO:0071803) |
| 0.1 | 0.8 | GO:0060482 | lobar bronchus epithelium development(GO:0060481) lobar bronchus development(GO:0060482) |
| 0.1 | 4.7 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.1 | 1.3 | GO:0070296 | sarcoplasmic reticulum calcium ion transport(GO:0070296) |
| 0.1 | 1.5 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.1 | 3.6 | GO:0031116 | positive regulation of microtubule polymerization(GO:0031116) |
| 0.1 | 1.6 | GO:0015732 | prostaglandin transport(GO:0015732) |
| 0.1 | 3.3 | GO:0030220 | platelet formation(GO:0030220) |
| 0.1 | 4.1 | GO:0070830 | bicellular tight junction assembly(GO:0070830) |
| 0.1 | 1.3 | GO:1900113 | negative regulation of histone H3-K9 trimethylation(GO:1900113) |
| 0.1 | 1.1 | GO:0051418 | interphase microtubule nucleation by interphase microtubule organizing center(GO:0051415) microtubule nucleation by microtubule organizing center(GO:0051418) |
| 0.1 | 0.3 | GO:0044849 | estrous cycle(GO:0044849) regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.1 | 1.7 | GO:0007263 | nitric oxide mediated signal transduction(GO:0007263) |
| 0.1 | 1.8 | GO:0010831 | positive regulation of myotube differentiation(GO:0010831) |
| 0.1 | 1.2 | GO:0032494 | response to peptidoglycan(GO:0032494) |
| 0.1 | 3.3 | GO:0030225 | macrophage differentiation(GO:0030225) |
| 0.1 | 0.5 | GO:0045075 | interleukin-12 biosynthetic process(GO:0042090) regulation of interleukin-12 biosynthetic process(GO:0045075) |
| 0.1 | 0.4 | GO:0021564 | vagus nerve development(GO:0021564) |
| 0.1 | 0.7 | GO:1902730 | regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010908) positive regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010909) positive regulation of proteoglycan biosynthetic process(GO:1902730) |
| 0.1 | 0.5 | GO:2000210 | positive regulation of anoikis(GO:2000210) |
| 0.1 | 2.8 | GO:0030835 | negative regulation of actin filament depolymerization(GO:0030835) |
| 0.1 | 3.5 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.1 | 0.5 | GO:0098719 | sodium ion import(GO:0097369) sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.1 | 0.5 | GO:0032202 | telomere assembly(GO:0032202) |
| 0.1 | 0.5 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.1 | 0.9 | GO:0061299 | retina vasculature morphogenesis in camera-type eye(GO:0061299) |
| 0.1 | 0.4 | GO:0045657 | positive regulation of monocyte differentiation(GO:0045657) |
| 0.1 | 0.4 | GO:0010766 | negative regulation of sodium ion transport(GO:0010766) |
| 0.1 | 2.4 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.1 | 0.2 | GO:0021571 | rhombomere 5 development(GO:0021571) |
| 0.1 | 0.2 | GO:0071733 | transcriptional activation by promoter-enhancer looping(GO:0071733) gene looping(GO:0090202) dsDNA loop formation(GO:0090579) |
| 0.1 | 2.0 | GO:0000731 | DNA synthesis involved in DNA repair(GO:0000731) |
| 0.1 | 0.3 | GO:0002143 | tRNA wobble position uridine thiolation(GO:0002143) |
| 0.1 | 0.6 | GO:0005513 | detection of calcium ion(GO:0005513) activation of store-operated calcium channel activity(GO:0032237) positive regulation of store-operated calcium channel activity(GO:1901341) |
| 0.1 | 0.7 | GO:0021527 | spinal cord association neuron differentiation(GO:0021527) |
| 0.1 | 0.5 | GO:0031179 | peptide modification(GO:0031179) |
| 0.1 | 0.5 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.1 | 1.1 | GO:0038092 | nodal signaling pathway(GO:0038092) |
| 0.1 | 13.6 | GO:0007519 | skeletal muscle tissue development(GO:0007519) |
| 0.1 | 1.2 | GO:0032897 | negative regulation of viral transcription(GO:0032897) |
| 0.1 | 2.4 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.1 | 2.6 | GO:0031648 | protein destabilization(GO:0031648) |
| 0.1 | 0.1 | GO:0033686 | positive regulation of gonadotropin secretion(GO:0032278) regulation of luteinizing hormone secretion(GO:0033684) positive regulation of luteinizing hormone secretion(GO:0033686) |
| 0.1 | 0.5 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) strand invasion(GO:0042148) |
| 0.1 | 0.4 | GO:0043569 | negative regulation of insulin-like growth factor receptor signaling pathway(GO:0043569) |
| 0.1 | 0.2 | GO:0001830 | trophectodermal cell fate commitment(GO:0001830) |
| 0.1 | 0.9 | GO:1903392 | negative regulation of focal adhesion assembly(GO:0051895) negative regulation of adherens junction organization(GO:1903392) |
| 0.1 | 1.0 | GO:0002693 | positive regulation of cellular extravasation(GO:0002693) |
| 0.1 | 0.6 | GO:0021516 | dorsal spinal cord development(GO:0021516) |
| 0.1 | 0.6 | GO:0061032 | visceral serous pericardium development(GO:0061032) |
| 0.1 | 0.4 | GO:0034134 | toll-like receptor 2 signaling pathway(GO:0034134) |
| 0.1 | 1.4 | GO:0045214 | sarcomere organization(GO:0045214) |
| 0.1 | 1.2 | GO:0071548 | response to dexamethasone(GO:0071548) cellular response to dexamethasone stimulus(GO:0071549) |
| 0.1 | 13.3 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.1 | 1.2 | GO:0030574 | collagen catabolic process(GO:0030574) multicellular organism catabolic process(GO:0044243) |
| 0.1 | 0.2 | GO:0006436 | tryptophanyl-tRNA aminoacylation(GO:0006436) |
| 0.1 | 0.3 | GO:0060923 | cardiac muscle cell fate commitment(GO:0060923) |
| 0.1 | 0.3 | GO:2000399 | negative regulation of T cell differentiation in thymus(GO:0033085) negative regulation of thymocyte aggregation(GO:2000399) |
| 0.1 | 0.6 | GO:0014850 | response to muscle activity(GO:0014850) |
| 0.1 | 0.4 | GO:0043619 | regulation of transcription from RNA polymerase II promoter in response to oxidative stress(GO:0043619) |
| 0.1 | 0.3 | GO:1902962 | regulation of metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902962) negative regulation of metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902963) |
| 0.1 | 1.9 | GO:0040018 | positive regulation of multicellular organism growth(GO:0040018) |
| 0.0 | 0.2 | GO:0046601 | positive regulation of centriole replication(GO:0046601) |
| 0.0 | 0.2 | GO:0045077 | negative regulation of interferon-gamma biosynthetic process(GO:0045077) |
| 0.0 | 0.4 | GO:0008228 | opsonization(GO:0008228) |
| 0.0 | 0.5 | GO:0035372 | protein localization to microtubule(GO:0035372) |
| 0.0 | 1.5 | GO:0010862 | positive regulation of pathway-restricted SMAD protein phosphorylation(GO:0010862) |
| 0.0 | 1.8 | GO:0046785 | microtubule polymerization(GO:0046785) |
| 0.0 | 0.2 | GO:0032298 | positive regulation of DNA-dependent DNA replication initiation(GO:0032298) |
| 0.0 | 1.4 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 0.3 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.0 | 1.0 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.0 | 0.2 | GO:0001550 | ovarian cumulus expansion(GO:0001550) fused antrum stage(GO:0048165) |
| 0.0 | 0.6 | GO:0035414 | negative regulation of catenin import into nucleus(GO:0035414) |
| 0.0 | 0.3 | GO:0010457 | centriole-centriole cohesion(GO:0010457) |
| 0.0 | 0.1 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 0.0 | 0.9 | GO:0051497 | negative regulation of stress fiber assembly(GO:0051497) |
| 0.0 | 0.5 | GO:0048747 | muscle fiber development(GO:0048747) |
| 0.0 | 1.0 | GO:0033599 | regulation of mammary gland epithelial cell proliferation(GO:0033599) |
| 0.0 | 0.4 | GO:0033234 | negative regulation of protein sumoylation(GO:0033234) |
| 0.0 | 1.8 | GO:1901998 | toxin transport(GO:1901998) |
| 0.0 | 0.6 | GO:0022011 | myelination in peripheral nervous system(GO:0022011) peripheral nervous system axon ensheathment(GO:0032292) |
| 0.0 | 2.1 | GO:0007569 | cell aging(GO:0007569) |
| 0.0 | 0.3 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.0 | 2.3 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.0 | 0.1 | GO:0035020 | regulation of Rac protein signal transduction(GO:0035020) |
| 0.0 | 0.1 | GO:0003199 | heart valve formation(GO:0003188) endocardial cushion to mesenchymal transition involved in heart valve formation(GO:0003199) |
| 0.0 | 0.2 | GO:0072386 | plus-end-directed vesicle transport along microtubule(GO:0072383) plus-end-directed organelle transport along microtubule(GO:0072386) |
| 0.0 | 1.1 | GO:0002279 | mast cell activation involved in immune response(GO:0002279) mast cell mediated immunity(GO:0002448) mast cell degranulation(GO:0043303) |
| 0.0 | 0.6 | GO:0034142 | toll-like receptor 4 signaling pathway(GO:0034142) |
| 0.0 | 1.0 | GO:0000154 | rRNA modification(GO:0000154) |
| 0.0 | 0.1 | GO:0006713 | glucocorticoid catabolic process(GO:0006713) |
| 0.0 | 1.4 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.0 | 0.2 | GO:1902187 | negative regulation of viral release from host cell(GO:1902187) |
| 0.0 | 3.7 | GO:0045089 | positive regulation of innate immune response(GO:0045089) |
| 0.0 | 0.6 | GO:0001958 | endochondral ossification(GO:0001958) replacement ossification(GO:0036075) |
| 0.0 | 0.1 | GO:1904690 | regulation of cap-independent translational initiation(GO:1903677) positive regulation of cap-independent translational initiation(GO:1903679) regulation of cytoplasmic translational initiation(GO:1904688) positive regulation of cytoplasmic translational initiation(GO:1904690) |
| 0.0 | 0.7 | GO:0045022 | early endosome to late endosome transport(GO:0045022) |
| 0.0 | 0.1 | GO:0008295 | spermidine biosynthetic process(GO:0008295) |
| 0.0 | 0.3 | GO:0043496 | regulation of protein homodimerization activity(GO:0043496) |
| 0.0 | 0.3 | GO:0007379 | segment specification(GO:0007379) |
| 0.0 | 1.2 | GO:0071806 | intracellular protein transmembrane transport(GO:0065002) protein transmembrane transport(GO:0071806) |
| 0.0 | 0.2 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.0 | 0.2 | GO:1902236 | negative regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signaling pathway(GO:1902236) |
| 0.0 | 0.3 | GO:0035162 | embryonic hemopoiesis(GO:0035162) |
| 0.0 | 0.1 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.0 | 0.2 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.0 | 0.1 | GO:0019372 | lipoxygenase pathway(GO:0019372) |
| 0.0 | 0.3 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 0.0 | 1.7 | GO:0019827 | stem cell population maintenance(GO:0019827) |
| 0.0 | 1.7 | GO:0060606 | neural tube closure(GO:0001843) tube closure(GO:0060606) |
| 0.0 | 0.1 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.3 | 49.4 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 3.2 | 9.5 | GO:0038045 | large latent transforming growth factor-beta complex(GO:0038045) |
| 1.3 | 1.3 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 1.2 | 3.6 | GO:0032998 | Fc receptor complex(GO:0032997) Fc-epsilon receptor I complex(GO:0032998) |
| 1.1 | 5.6 | GO:0005914 | spot adherens junction(GO:0005914) |
| 1.0 | 3.1 | GO:0031904 | endosome lumen(GO:0031904) |
| 1.0 | 8.6 | GO:1990357 | terminal web(GO:1990357) |
| 0.9 | 3.7 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 0.8 | 14.9 | GO:0005861 | troponin complex(GO:0005861) |
| 0.7 | 12.6 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.7 | 7.8 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.7 | 2.1 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 0.7 | 2.7 | GO:0014802 | terminal cisterna(GO:0014802) |
| 0.7 | 2.0 | GO:0042643 | actomyosin, actin portion(GO:0042643) |
| 0.6 | 7.3 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.6 | 2.4 | GO:0034667 | integrin alpha3-beta1 complex(GO:0034667) |
| 0.5 | 3.2 | GO:0097149 | centralspindlin complex(GO:0097149) |
| 0.5 | 7.3 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.5 | 13.3 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.4 | 2.0 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.4 | 1.6 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.4 | 16.2 | GO:0045095 | keratin filament(GO:0045095) |
| 0.4 | 1.4 | GO:0036284 | tubulobulbar complex(GO:0036284) |
| 0.3 | 1.7 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 0.3 | 7.5 | GO:0005865 | striated muscle thin filament(GO:0005865) |
| 0.3 | 2.4 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 0.3 | 2.0 | GO:0097513 | myosin II filament(GO:0097513) |
| 0.3 | 34.9 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.3 | 1.5 | GO:0000214 | tRNA-intron endonuclease complex(GO:0000214) |
| 0.2 | 3.0 | GO:0016011 | dystroglycan complex(GO:0016011) |
| 0.2 | 2.7 | GO:0071664 | catenin-TCF7L2 complex(GO:0071664) |
| 0.2 | 4.8 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.2 | 3.1 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.2 | 1.4 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 0.2 | 2.5 | GO:0097451 | glial limiting end-foot(GO:0097451) |
| 0.2 | 1.8 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.2 | 3.5 | GO:0043034 | costamere(GO:0043034) |
| 0.2 | 2.8 | GO:0001931 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.2 | 1.3 | GO:0071458 | integral component of cytoplasmic side of endoplasmic reticulum membrane(GO:0071458) integral component of lumenal side of endoplasmic reticulum membrane(GO:0071556) lumenal side of endoplasmic reticulum membrane(GO:0098553) |
| 0.2 | 24.8 | GO:0097517 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.2 | 2.4 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.2 | 6.5 | GO:0042588 | zymogen granule(GO:0042588) |
| 0.2 | 1.1 | GO:0008274 | gamma-tubulin large complex(GO:0000931) gamma-tubulin ring complex(GO:0008274) |
| 0.2 | 27.6 | GO:0030018 | Z disc(GO:0030018) |
| 0.1 | 2.7 | GO:0031674 | I band(GO:0031674) |
| 0.1 | 1.7 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.1 | 2.1 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.1 | 4.0 | GO:0030057 | desmosome(GO:0030057) |
| 0.1 | 5.3 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.1 | 4.0 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.1 | 11.1 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.1 | 1.6 | GO:0016589 | NURF complex(GO:0016589) |
| 0.1 | 1.0 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.1 | 0.6 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.1 | 1.0 | GO:0030478 | actin cap(GO:0030478) |
| 0.1 | 0.7 | GO:0070695 | FHF complex(GO:0070695) |
| 0.1 | 1.1 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.1 | 1.9 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.1 | 0.8 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 0.1 | 6.0 | GO:0005720 | nuclear heterochromatin(GO:0005720) |
| 0.1 | 2.1 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.1 | 0.7 | GO:0001940 | male pronucleus(GO:0001940) |
| 0.1 | 1.6 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 0.1 | 1.6 | GO:0044232 | organelle membrane contact site(GO:0044232) |
| 0.1 | 5.4 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.1 | 1.3 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.1 | 0.9 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 0.1 | 0.5 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.1 | 1.5 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.1 | 3.1 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.1 | 1.3 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.1 | 2.8 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 1.5 | GO:0043218 | compact myelin(GO:0043218) |
| 0.1 | 4.4 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.1 | 0.8 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.1 | 1.1 | GO:0031672 | A band(GO:0031672) |
| 0.1 | 2.3 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.1 | 0.4 | GO:0005785 | signal recognition particle receptor complex(GO:0005785) |
| 0.1 | 0.3 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.1 | 2.1 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 0.5 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.1 | 0.6 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.0 | 6.9 | GO:0000922 | spindle pole(GO:0000922) |
| 0.0 | 2.5 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 0.6 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.0 | 5.3 | GO:0030863 | cortical cytoskeleton(GO:0030863) |
| 0.0 | 2.2 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 0.2 | GO:0000839 | Hrd1p ubiquitin ligase ERAD-L complex(GO:0000839) |
| 0.0 | 1.8 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.0 | 4.9 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 0.4 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.0 | 0.7 | GO:0036038 | MKS complex(GO:0036038) |
| 0.0 | 1.0 | GO:0005605 | basal lamina(GO:0005605) |
| 0.0 | 1.0 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.0 | 0.3 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 0.0 | 0.4 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.0 | 0.4 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.0 | 5.0 | GO:0005923 | bicellular tight junction(GO:0005923) |
| 0.0 | 4.7 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.0 | 1.3 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.0 | 0.4 | GO:0034709 | methylosome(GO:0034709) |
| 0.0 | 0.8 | GO:0043296 | apical junction complex(GO:0043296) |
| 0.0 | 3.3 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 0.1 | GO:0000938 | GARP complex(GO:0000938) |
| 0.0 | 0.2 | GO:0033268 | node of Ranvier(GO:0033268) |
| 0.0 | 0.1 | GO:0045180 | basal cortex(GO:0045180) |
| 0.0 | 4.0 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 47.3 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 0.5 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
| 0.0 | 1.3 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 1.6 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 3.5 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 5.0 | GO:0098852 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.0 | 1.2 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 1.8 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 1.1 | GO:0032420 | stereocilium(GO:0032420) |
| 0.0 | 8.4 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 2.2 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 0.3 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.0 | 4.2 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 2.1 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 0.3 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.0 | 2.4 | GO:0000794 | condensed nuclear chromosome(GO:0000794) |
| 0.0 | 0.2 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.0 | 0.4 | GO:0000152 | nuclear ubiquitin ligase complex(GO:0000152) |
| 0.0 | 1.1 | GO:0005819 | spindle(GO:0005819) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.6 | 10.2 | GO:0050436 | microfibril binding(GO:0050436) |
| 2.2 | 6.5 | GO:0050253 | sterol esterase activity(GO:0004771) retinyl-palmitate esterase activity(GO:0050253) |
| 2.1 | 12.6 | GO:0070320 | inward rectifier potassium channel inhibitor activity(GO:0070320) |
| 1.5 | 14.8 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 1.2 | 11.9 | GO:0031014 | troponin T binding(GO:0031014) |
| 1.1 | 79.1 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.8 | 3.3 | GO:0042392 | sphingosine-1-phosphate phosphatase activity(GO:0042392) |
| 0.8 | 3.1 | GO:0048030 | disaccharide binding(GO:0048030) |
| 0.8 | 3.1 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.7 | 3.0 | GO:0004699 | calcium-independent protein kinase C activity(GO:0004699) |
| 0.7 | 2.2 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.7 | 3.6 | GO:0019767 | IgE receptor activity(GO:0019767) |
| 0.7 | 4.1 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.7 | 4.1 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.7 | 2.7 | GO:0070740 | tubulin-glutamic acid ligase activity(GO:0070740) |
| 0.7 | 2.0 | GO:0004958 | prostaglandin F receptor activity(GO:0004958) |
| 0.7 | 3.3 | GO:0070051 | fibrinogen binding(GO:0070051) |
| 0.6 | 6.5 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.6 | 1.8 | GO:0003845 | 11-beta-hydroxysteroid dehydrogenase [NAD(P)] activity(GO:0003845) |
| 0.5 | 2.2 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 0.5 | 8.3 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.5 | 1.5 | GO:0070773 | protein-N-terminal glutamine amidohydrolase activity(GO:0070773) |
| 0.5 | 3.6 | GO:0005111 | type 2 fibroblast growth factor receptor binding(GO:0005111) |
| 0.5 | 2.5 | GO:0008853 | exodeoxyribonuclease III activity(GO:0008853) |
| 0.5 | 2.0 | GO:0008160 | protein tyrosine phosphatase activator activity(GO:0008160) |
| 0.5 | 3.4 | GO:0047276 | N-acetyllactosaminide 3-alpha-galactosyltransferase activity(GO:0047276) |
| 0.5 | 1.4 | GO:0005330 | dopamine:sodium symporter activity(GO:0005330) |
| 0.5 | 0.5 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.4 | 5.8 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.4 | 2.7 | GO:0044729 | hemi-methylated DNA-binding(GO:0044729) |
| 0.4 | 1.5 | GO:0030172 | troponin C binding(GO:0030172) |
| 0.3 | 2.1 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.3 | 5.7 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.3 | 1.0 | GO:0038100 | nodal binding(GO:0038100) |
| 0.3 | 3.0 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.3 | 1.6 | GO:0004668 | protein-arginine deiminase activity(GO:0004668) |
| 0.3 | 2.2 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.3 | 9.5 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.3 | 4.4 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.3 | 38.5 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.3 | 5.4 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.3 | 2.6 | GO:0051575 | 5'-deoxyribose-5-phosphate lyase activity(GO:0051575) |
| 0.3 | 3.4 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.3 | 2.4 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.3 | 1.5 | GO:0035877 | death effector domain binding(GO:0035877) |
| 0.3 | 1.5 | GO:0000213 | tRNA-intron endonuclease activity(GO:0000213) |
| 0.2 | 0.7 | GO:0000402 | open form four-way junction DNA binding(GO:0000401) crossed form four-way junction DNA binding(GO:0000402) |
| 0.2 | 2.4 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.2 | 2.1 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.2 | 1.4 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 0.2 | 0.7 | GO:0005171 | hepatocyte growth factor receptor binding(GO:0005171) |
| 0.2 | 3.0 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 0.2 | 1.3 | GO:0046624 | sphingolipid transporter activity(GO:0046624) |
| 0.2 | 1.5 | GO:0015315 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.2 | 1.4 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.2 | 5.5 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.2 | 3.7 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.2 | 2.0 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.2 | 0.5 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.2 | 1.2 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.2 | 2.4 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.2 | 1.4 | GO:0005166 | neurotrophin p75 receptor binding(GO:0005166) |
| 0.2 | 4.4 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.2 | 0.8 | GO:0005275 | amine transmembrane transporter activity(GO:0005275) |
| 0.2 | 0.6 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.2 | 1.6 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.2 | 1.4 | GO:0042285 | xylosyltransferase activity(GO:0042285) |
| 0.2 | 1.7 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.2 | 3.7 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 0.2 | 1.8 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 0.2 | 1.1 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.1 | 1.6 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.1 | 1.9 | GO:0071253 | connexin binding(GO:0071253) |
| 0.1 | 1.3 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.1 | 5.4 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.1 | 2.4 | GO:0043495 | protein anchor(GO:0043495) |
| 0.1 | 4.8 | GO:0043236 | laminin binding(GO:0043236) |
| 0.1 | 2.7 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 0.5 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.1 | 2.0 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.1 | 1.7 | GO:0046934 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol bisphosphate kinase activity(GO:0052813) |
| 0.1 | 1.3 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.1 | 1.4 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.1 | 1.2 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.1 | 1.5 | GO:0003997 | acyl-CoA oxidase activity(GO:0003997) |
| 0.1 | 2.1 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.1 | 1.4 | GO:0033170 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.1 | 2.8 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.1 | 0.9 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 0.1 | 4.0 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.1 | 2.4 | GO:0031005 | filamin binding(GO:0031005) |
| 0.1 | 4.9 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.1 | 4.7 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 2.7 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.1 | 4.7 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 27.8 | GO:0008236 | serine-type peptidase activity(GO:0008236) |
| 0.1 | 1.7 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 7.1 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 1.6 | GO:0015245 | fatty acid transporter activity(GO:0015245) |
| 0.1 | 0.5 | GO:0042134 | rRNA primary transcript binding(GO:0042134) |
| 0.1 | 3.3 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.1 | 1.6 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.1 | 1.8 | GO:0030548 | acetylcholine receptor regulator activity(GO:0030548) neurotransmitter receptor regulator activity(GO:0099602) |
| 0.1 | 2.7 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.1 | 0.4 | GO:1902379 | chemoattractant activity involved in axon guidance(GO:1902379) |
| 0.1 | 1.1 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.1 | 3.9 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.1 | 2.7 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.1 | 0.9 | GO:0016670 | oxidoreductase activity, acting on a sulfur group of donors, oxygen as acceptor(GO:0016670) |
| 0.1 | 0.9 | GO:0016004 | phospholipase activator activity(GO:0016004) |
| 0.1 | 2.6 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.1 | 0.7 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 0.1 | 0.5 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.1 | 1.7 | GO:0031702 | type 1 angiotensin receptor binding(GO:0031702) |
| 0.1 | 3.7 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.1 | 1.0 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.1 | 0.4 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.1 | 0.7 | GO:0070579 | methylcytosine dioxygenase activity(GO:0070579) |
| 0.1 | 1.9 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.1 | 2.8 | GO:0000980 | RNA polymerase II distal enhancer sequence-specific DNA binding(GO:0000980) |
| 0.1 | 0.6 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.1 | 1.3 | GO:0030296 | protein tyrosine kinase activator activity(GO:0030296) |
| 0.1 | 1.8 | GO:0048156 | tau protein binding(GO:0048156) |
| 0.1 | 0.3 | GO:0031800 | type 3 metabotropic glutamate receptor binding(GO:0031800) |
| 0.1 | 3.9 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.1 | 1.1 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.1 | 2.0 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 1.3 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.1 | 3.5 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.1 | 2.6 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.1 | 1.0 | GO:0015250 | water channel activity(GO:0015250) |
| 0.1 | 1.6 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.1 | 3.2 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.1 | 0.2 | GO:0004830 | tryptophan-tRNA ligase activity(GO:0004830) |
| 0.1 | 2.6 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.1 | 0.5 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.1 | 2.4 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 31.5 | GO:0003779 | actin binding(GO:0003779) |
| 0.1 | 2.8 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 2.0 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 0.7 | GO:0015197 | peptide transporter activity(GO:0015197) |
| 0.0 | 1.3 | GO:0015026 | coreceptor activity(GO:0015026) |
| 0.0 | 0.6 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 3.6 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.0 | 1.5 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.0 | 0.6 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 6.4 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 1.3 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 2.3 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.0 | 0.7 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.0 | 0.7 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.0 | 0.5 | GO:0001158 | enhancer sequence-specific DNA binding(GO:0001158) |
| 0.0 | 0.4 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 0.0 | 1.9 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 0.3 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.0 | 0.3 | GO:0043560 | insulin receptor substrate binding(GO:0043560) |
| 0.0 | 0.9 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.0 | 1.3 | GO:0008378 | galactosyltransferase activity(GO:0008378) |
| 0.0 | 0.6 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.0 | 1.2 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.0 | 1.0 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.0 | 0.4 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.0 | 1.8 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.4 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 4.1 | GO:0008565 | protein transporter activity(GO:0008565) |
| 0.0 | 0.4 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.0 | 11.3 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 0.6 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.0 | 1.1 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.1 | GO:0004052 | arachidonate 12-lipoxygenase activity(GO:0004052) |
| 0.0 | 0.2 | GO:0043141 | ATP-dependent 5'-3' DNA helicase activity(GO:0043141) |
| 0.0 | 0.3 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.0 | 0.5 | GO:0070034 | telomerase RNA binding(GO:0070034) |
| 0.0 | 0.2 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.0 | 14.6 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 0.1 | GO:0098821 | BMP receptor activity(GO:0098821) |
| 0.0 | 0.6 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.0 | 4.2 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 1.4 | GO:0003823 | antigen binding(GO:0003823) |
| 0.0 | 0.3 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
| 0.0 | 0.3 | GO:0004180 | carboxypeptidase activity(GO:0004180) |
| 0.0 | 0.5 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.4 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.0 | 0.3 | GO:0098748 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.0 | 0.8 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.2 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.0 | 0.1 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.0 | 0.1 | GO:0016422 | mRNA (2'-O-methyladenosine-N6-)-methyltransferase activity(GO:0016422) |
| 0.0 | 0.7 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.2 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 0.0 | 0.1 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.0 | 1.3 | GO:0015108 | chloride transmembrane transporter activity(GO:0015108) |
| 0.0 | 1.3 | GO:0001076 | transcription factor activity, RNA polymerase II transcription factor binding(GO:0001076) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 7.2 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.7 | 3.4 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.4 | 14.1 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.4 | 6.5 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.3 | 11.2 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.2 | 6.4 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.2 | 1.7 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.2 | 5.3 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.2 | 1.4 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.2 | 4.0 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.2 | 6.2 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.2 | 10.1 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.1 | 2.4 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.1 | 3.3 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.1 | 7.6 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.1 | 3.5 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.1 | 4.5 | PID MYC PATHWAY | C-MYC pathway |
| 0.1 | 2.2 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 11.0 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.1 | 7.3 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.1 | 9.7 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.1 | 3.7 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.1 | 5.3 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.1 | 2.6 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.1 | 5.5 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.1 | 1.4 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.1 | 1.4 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.1 | 2.8 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.1 | 4.9 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 1.7 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
| 0.1 | 3.3 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.1 | 2.2 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.1 | 3.6 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.1 | 3.1 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.1 | 3.6 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 4.1 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.1 | 4.2 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.1 | 0.8 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.1 | 12.5 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.1 | 2.1 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.1 | 0.9 | PID EPO PATHWAY | EPO signaling pathway |
| 0.1 | 1.6 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.1 | 3.2 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.1 | 0.9 | ST P38 MAPK PATHWAY | p38 MAPK Pathway |
| 0.1 | 0.4 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.1 | 2.3 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.1 | 2.2 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 0.9 | PID TCPTP PATHWAY | Signaling events mediated by TCPTP |
| 0.0 | 1.5 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 1.7 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.0 | 2.1 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.0 | 0.8 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.0 | 1.4 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 2.2 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.0 | 3.2 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.0 | 2.7 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 0.5 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.0 | 1.1 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 1.4 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 0.5 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.0 | 0.6 | PID CD8 TCR PATHWAY | TCR signaling in naïve CD8+ T cells |
| 0.0 | 0.6 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 0.4 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.0 | 0.4 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.0 | 2.6 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.0 | 5.4 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.2 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 1.7 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 0.9 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 0.9 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.0 | 0.6 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 0.7 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.3 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.0 | 0.5 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 74.0 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.7 | 2.0 | REACTOME FGFR1 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR1 ligand binding and activation |
| 0.4 | 10.8 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.4 | 11.5 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.4 | 7.2 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.3 | 7.3 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 0.3 | 15.6 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.3 | 9.1 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.3 | 3.6 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.2 | 3.6 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.2 | 2.0 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.2 | 5.4 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.2 | 3.4 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 0.2 | 0.6 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.2 | 5.3 | REACTOME TRAF6 MEDIATED NFKB ACTIVATION | Genes involved in TRAF6 mediated NF-kB activation |
| 0.2 | 5.9 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.2 | 5.7 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.2 | 1.4 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.1 | 0.8 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.1 | 4.2 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.1 | 1.8 | REACTOME PROSTACYCLIN SIGNALLING THROUGH PROSTACYCLIN RECEPTOR | Genes involved in Prostacyclin signalling through prostacyclin receptor |
| 0.1 | 3.6 | REACTOME ACTIVATED POINT MUTANTS OF FGFR2 | Genes involved in Activated point mutants of FGFR2 |
| 0.1 | 1.6 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.1 | 3.0 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.1 | 6.7 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 2.0 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.1 | 9.4 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.1 | 3.6 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.1 | 4.1 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.1 | 3.9 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.1 | 2.2 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 3.2 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.1 | 0.6 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.1 | 2.4 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 1.0 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.1 | 3.3 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.1 | 0.9 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.1 | 0.8 | REACTOME EXTENSION OF TELOMERES | Genes involved in Extension of Telomeres |
| 0.1 | 3.0 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.1 | 1.7 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 1.8 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.0 | 2.4 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 1.4 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.0 | 1.1 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.0 | 2.3 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.0 | 2.2 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 2.2 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.0 | 0.9 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.0 | 0.5 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.0 | 0.5 | REACTOME GLUCAGON TYPE LIGAND RECEPTORS | Genes involved in Glucagon-type ligand receptors |
| 0.0 | 0.9 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 5.4 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.3 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.0 | 0.5 | REACTOME REGULATION OF HYPOXIA INDUCIBLE FACTOR HIF BY OXYGEN | Genes involved in Regulation of Hypoxia-inducible Factor (HIF) by Oxygen |
| 0.0 | 0.3 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.0 | 0.1 | REACTOME DEFENSINS | Genes involved in Defensins |
| 0.0 | 1.9 | REACTOME TOLL RECEPTOR CASCADES | Genes involved in Toll Receptor Cascades |
| 0.0 | 0.4 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 5.2 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 1.2 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.3 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.0 | 1.1 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.3 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.3 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.0 | 1.0 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.3 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 0.2 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |