Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
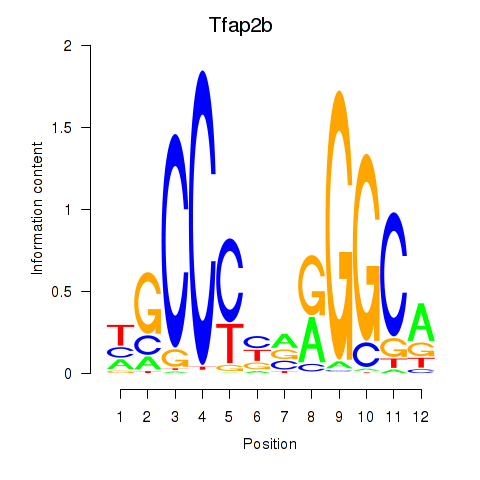
Results for Tfap2b
Z-value: 1.00
Transcription factors associated with Tfap2b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Tfap2b
|
ENSMUSG00000025927.14 | Tfap2b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Tfap2b | mm39_v1_chr1_+_19279138_19279176 | -0.19 | 1.2e-01 | Click! |
Activity profile of Tfap2b motif
Sorted Z-values of Tfap2b motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Tfap2b
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_131671751 | 8.89 |
ENSMUST00000049027.10
|
Slc26a9
|
solute carrier family 26, member 9 |
| chr11_-_99383938 | 7.83 |
ENSMUST00000006969.8
|
Krt23
|
keratin 23 |
| chr15_-_101862711 | 5.90 |
ENSMUST00000164932.3
|
Krt78
|
keratin 78 |
| chr7_+_24335969 | 5.60 |
ENSMUST00000080718.6
|
Lypd3
|
Ly6/Plaur domain containing 3 |
| chr4_-_140501507 | 4.95 |
ENSMUST00000026381.7
|
Padi4
|
peptidyl arginine deiminase, type IV |
| chr13_+_3887757 | 4.70 |
ENSMUST00000042219.6
|
Calm4
|
calmodulin 4 |
| chr13_+_48816466 | 4.20 |
ENSMUST00000021813.5
|
Barx1
|
BarH-like homeobox 1 |
| chr11_-_70146156 | 4.15 |
ENSMUST00000108574.3
ENSMUST00000000329.9 |
Alox12
|
arachidonate 12-lipoxygenase |
| chr1_+_131678223 | 3.94 |
ENSMUST00000147800.2
|
Slc26a9
|
solute carrier family 26, member 9 |
| chr15_-_101833160 | 3.67 |
ENSMUST00000023797.8
|
Krt4
|
keratin 4 |
| chr18_-_20247666 | 3.29 |
ENSMUST00000224432.2
|
Dsc1
|
desmocollin 1 |
| chr18_-_20247824 | 3.03 |
ENSMUST00000038710.6
|
Dsc1
|
desmocollin 1 |
| chr19_+_6952319 | 2.93 |
ENSMUST00000070850.8
|
Ppp1r14b
|
protein phosphatase 1, regulatory inhibitor subunit 14B |
| chr3_-_84387700 | 2.86 |
ENSMUST00000194027.2
ENSMUST00000107689.7 |
Fhdc1
|
FH2 domain containing 1 |
| chr6_+_113435716 | 2.85 |
ENSMUST00000203661.3
ENSMUST00000204774.3 ENSMUST00000053569.7 ENSMUST00000101065.8 |
Il17re
|
interleukin 17 receptor E |
| chr19_+_6952580 | 2.79 |
ENSMUST00000237084.2
ENSMUST00000236218.2 ENSMUST00000237235.2 |
Ppp1r14b
|
protein phosphatase 1, regulatory inhibitor subunit 14B |
| chr7_+_43441315 | 2.72 |
ENSMUST00000005891.7
|
Klk9
|
kallikrein related-peptidase 9 |
| chr11_-_102255999 | 2.57 |
ENSMUST00000006749.10
|
Slc4a1
|
solute carrier family 4 (anion exchanger), member 1 |
| chr9_+_118881838 | 2.45 |
ENSMUST00000051386.13
ENSMUST00000074734.13 |
Vill
|
villin-like |
| chr6_-_124710084 | 2.29 |
ENSMUST00000112484.10
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr1_+_34840785 | 2.26 |
ENSMUST00000047664.16
ENSMUST00000211073.2 |
Arhgef4
SMIM39
|
Rho guanine nucleotide exchange factor (GEF) 4 novel protein |
| chr18_-_20247928 | 2.24 |
ENSMUST00000226115.2
|
Dsc1
|
desmocollin 1 |
| chr12_+_85733415 | 2.04 |
ENSMUST00000040536.6
|
Batf
|
basic leucine zipper transcription factor, ATF-like |
| chr5_-_103777145 | 2.02 |
ENSMUST00000031263.2
|
Slc10a6
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 6 |
| chr2_+_25179903 | 2.00 |
ENSMUST00000028337.7
|
Lrrc26
|
leucine rich repeat containing 26 |
| chr7_+_25386418 | 1.97 |
ENSMUST00000002678.10
|
Tgfb1
|
transforming growth factor, beta 1 |
| chr1_+_86454431 | 1.96 |
ENSMUST00000045897.15
ENSMUST00000186255.7 ENSMUST00000188699.7 |
Ptma
|
prothymosin alpha |
| chr15_-_102165884 | 1.95 |
ENSMUST00000043172.15
|
Rarg
|
retinoic acid receptor, gamma |
| chr11_+_69546140 | 1.95 |
ENSMUST00000047373.6
|
Sox15
|
SRY (sex determining region Y)-box 15 |
| chr9_+_118881926 | 1.90 |
ENSMUST00000131647.2
|
Vill
|
villin-like |
| chr3_-_137687284 | 1.87 |
ENSMUST00000136613.4
ENSMUST00000029806.13 |
Dapp1
|
dual adaptor for phosphotyrosine and 3-phosphoinositides 1 |
| chr19_+_38043446 | 1.86 |
ENSMUST00000236044.2
ENSMUST00000116506.8 ENSMUST00000096096.12 ENSMUST00000169673.3 |
Cep55
|
centrosomal protein 55 |
| chr13_-_40891849 | 1.80 |
ENSMUST00000223869.2
|
Tfap2a
|
transcription factor AP-2, alpha |
| chr4_-_133360749 | 1.79 |
ENSMUST00000084238.5
|
Zdhhc18
|
zinc finger, DHHC domain containing 18 |
| chr10_-_81201642 | 1.74 |
ENSMUST00000020456.5
|
4930404N11Rik
|
RIKEN cDNA 4930404N11 gene |
| chr15_-_102165740 | 1.73 |
ENSMUST00000135466.2
|
Rarg
|
retinoic acid receptor, gamma |
| chr6_+_7844759 | 1.71 |
ENSMUST00000040159.6
|
C1galt1
|
core 1 synthase, glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase, 1 |
| chr7_-_45175570 | 1.67 |
ENSMUST00000167273.2
ENSMUST00000042105.11 |
Ppp1r15a
|
protein phosphatase 1, regulatory subunit 15A |
| chr17_+_34809132 | 1.66 |
ENSMUST00000173772.2
|
Gpsm3
|
G-protein signalling modulator 3 (AGS3-like, C. elegans) |
| chr15_-_75620060 | 1.65 |
ENSMUST00000062002.6
|
Mafa
|
v-maf musculoaponeurotic fibrosarcoma oncogene family, protein A (avian) |
| chr17_+_33774681 | 1.63 |
ENSMUST00000087605.13
ENSMUST00000174695.2 |
Myo1f
|
myosin IF |
| chr15_+_61859255 | 1.61 |
ENSMUST00000160009.2
|
Myc
|
myelocytomatosis oncogene |
| chr12_+_70872266 | 1.60 |
ENSMUST00000057859.9
|
Frmd6
|
FERM domain containing 6 |
| chr18_+_61044830 | 1.59 |
ENSMUST00000040359.6
|
Arsi
|
arylsulfatase i |
| chr7_-_97811525 | 1.58 |
ENSMUST00000107112.2
|
Capn5
|
calpain 5 |
| chr19_-_6065872 | 1.54 |
ENSMUST00000164843.10
|
Capn1
|
calpain 1 |
| chr12_-_111638722 | 1.52 |
ENSMUST00000001304.9
|
Ckb
|
creatine kinase, brain |
| chr17_+_34808772 | 1.51 |
ENSMUST00000038244.15
|
Gpsm3
|
G-protein signalling modulator 3 (AGS3-like, C. elegans) |
| chr1_+_86454511 | 1.51 |
ENSMUST00000188533.2
|
Ptma
|
prothymosin alpha |
| chr4_-_133694607 | 1.50 |
ENSMUST00000105893.8
|
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr7_+_28834276 | 1.43 |
ENSMUST00000161522.8
ENSMUST00000204845.3 ENSMUST00000205027.3 ENSMUST00000204194.3 ENSMUST00000203070.3 ENSMUST00000203380.3 |
Rasgrp4
|
RAS guanyl releasing protein 4 |
| chr8_+_72050292 | 1.42 |
ENSMUST00000143662.8
|
Niban3
|
niban apoptosis regulator 3 |
| chr6_+_97783975 | 1.42 |
ENSMUST00000203884.3
ENSMUST00000043637.14 |
Mitf
|
melanogenesis associated transcription factor |
| chr8_+_85449632 | 1.40 |
ENSMUST00000098571.5
|
G430095P16Rik
|
RIKEN cDNA G430095P16 gene |
| chr15_-_103242697 | 1.39 |
ENSMUST00000229373.2
|
Zfp385a
|
zinc finger protein 385A |
| chr7_-_126046814 | 1.36 |
ENSMUST00000146973.2
|
Atp2a1
|
ATPase, Ca++ transporting, cardiac muscle, fast twitch 1 |
| chr16_-_92155762 | 1.34 |
ENSMUST00000166707.3
|
Kcne1
|
potassium voltage-gated channel, Isk-related subfamily, member 1 |
| chr2_+_158610003 | 1.34 |
ENSMUST00000029183.3
|
Fam83d
|
family with sequence similarity 83, member D |
| chr16_-_92156312 | 1.30 |
ENSMUST00000051705.7
|
Kcne1
|
potassium voltage-gated channel, Isk-related subfamily, member 1 |
| chr1_+_74830675 | 1.29 |
ENSMUST00000006718.15
|
Wnt10a
|
wingless-type MMTV integration site family, member 10A |
| chr2_-_131001916 | 1.28 |
ENSMUST00000103188.10
ENSMUST00000133602.8 ENSMUST00000028800.12 |
1700037H04Rik
|
RIKEN cDNA 1700037H04 gene |
| chr16_-_20245138 | 1.28 |
ENSMUST00000079158.13
|
Abcc5
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 5 |
| chr3_-_132655804 | 1.25 |
ENSMUST00000117164.8
ENSMUST00000093971.5 ENSMUST00000042729.16 |
Npnt
|
nephronectin |
| chr9_-_50256268 | 1.23 |
ENSMUST00000076364.6
|
Rpl10-ps3
|
ribosomal protein L10, pseudogene 3 |
| chr3_-_132655954 | 1.23 |
ENSMUST00000042744.16
ENSMUST00000117811.8 |
Npnt
|
nephronectin |
| chrX_+_158410229 | 1.22 |
ENSMUST00000112456.9
|
Sh3kbp1
|
SH3-domain kinase binding protein 1 |
| chr7_+_141027557 | 1.22 |
ENSMUST00000106004.8
|
Rplp2
|
ribosomal protein, large P2 |
| chr15_-_97729341 | 1.21 |
ENSMUST00000079838.14
ENSMUST00000118294.8 |
Hdac7
|
histone deacetylase 7 |
| chr7_+_141027763 | 1.20 |
ENSMUST00000106003.2
|
Rplp2
|
ribosomal protein, large P2 |
| chr18_+_60907698 | 1.20 |
ENSMUST00000118551.8
|
Rps14
|
ribosomal protein S14 |
| chr2_-_11506893 | 1.19 |
ENSMUST00000114845.10
ENSMUST00000171188.9 ENSMUST00000179584.8 ENSMUST00000028114.13 ENSMUST00000114846.9 ENSMUST00000170196.9 ENSMUST00000191668.6 ENSMUST00000049849.12 ENSMUST00000114844.8 ENSMUST00000100411.4 |
Pfkfb3
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3 |
| chr2_-_30720345 | 1.19 |
ENSMUST00000041726.4
|
Asb6
|
ankyrin repeat and SOCS box-containing 6 |
| chr12_-_15866763 | 1.19 |
ENSMUST00000020922.8
ENSMUST00000221215.2 ENSMUST00000221518.2 |
Trib2
|
tribbles pseudokinase 2 |
| chr14_+_54431947 | 1.19 |
ENSMUST00000103715.2
|
Traj26
|
T cell receptor alpha joining 26 |
| chr8_+_105897542 | 1.17 |
ENSMUST00000109394.3
|
Cbfb
|
core binding factor beta |
| chr4_-_133694543 | 1.15 |
ENSMUST00000123234.8
|
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr5_-_115439016 | 1.15 |
ENSMUST00000009157.4
|
Dynll1
|
dynein light chain LC8-type 1 |
| chr18_+_60907668 | 1.15 |
ENSMUST00000025511.11
|
Rps14
|
ribosomal protein S14 |
| chr7_+_24990596 | 1.14 |
ENSMUST00000164820.2
|
Cic
|
capicua transcriptional repressor |
| chr16_-_20245071 | 1.14 |
ENSMUST00000115547.9
ENSMUST00000096199.5 |
Abcc5
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 5 |
| chr4_+_59035088 | 1.14 |
ENSMUST00000041160.13
|
Gng10
|
guanine nucleotide binding protein (G protein), gamma 10 |
| chr12_-_76756772 | 1.13 |
ENSMUST00000166101.2
|
Sptb
|
spectrin beta, erythrocytic |
| chr19_+_38043506 | 1.12 |
ENSMUST00000237408.2
|
Cep55
|
centrosomal protein 55 |
| chr8_-_91074971 | 1.09 |
ENSMUST00000109621.10
|
Tox3
|
TOX high mobility group box family member 3 |
| chrX_-_9335525 | 1.09 |
ENSMUST00000015484.10
|
Cybb
|
cytochrome b-245, beta polypeptide |
| chr13_-_53440087 | 1.09 |
ENSMUST00000021918.10
|
Ror2
|
receptor tyrosine kinase-like orphan receptor 2 |
| chr7_+_58307930 | 1.08 |
ENSMUST00000168747.3
|
Atp10a
|
ATPase, class V, type 10A |
| chr8_+_105897300 | 1.06 |
ENSMUST00000052209.9
ENSMUST00000109392.9 ENSMUST00000109395.8 |
Cbfb
|
core binding factor beta |
| chr5_-_115438971 | 1.04 |
ENSMUST00000112090.2
|
Dynll1
|
dynein light chain LC8-type 1 |
| chr3_+_130904000 | 1.02 |
ENSMUST00000029611.14
ENSMUST00000106341.9 ENSMUST00000066849.13 |
Lef1
|
lymphoid enhancer binding factor 1 |
| chr1_-_132294807 | 1.01 |
ENSMUST00000136828.3
|
Tmcc2
|
transmembrane and coiled-coil domains 2 |
| chr4_+_48045143 | 0.98 |
ENSMUST00000030025.10
|
Nr4a3
|
nuclear receptor subfamily 4, group A, member 3 |
| chr8_+_93084253 | 0.97 |
ENSMUST00000210246.2
ENSMUST00000034184.12 |
Irx5
|
Iroquois homeobox 5 |
| chr9_+_62249341 | 0.96 |
ENSMUST00000135395.8
|
Anp32a
|
acidic (leucine-rich) nuclear phosphoprotein 32 family, member A |
| chr2_-_147887810 | 0.96 |
ENSMUST00000109964.8
|
Foxa2
|
forkhead box A2 |
| chr19_+_6451667 | 0.96 |
ENSMUST00000113471.3
ENSMUST00000113469.3 |
Rasgrp2
|
RAS, guanyl releasing protein 2 |
| chr3_-_108322868 | 0.95 |
ENSMUST00000090558.10
|
Celsr2
|
cadherin, EGF LAG seven-pass G-type receptor 2 |
| chr4_-_133695264 | 0.94 |
ENSMUST00000102553.11
|
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr5_-_139798992 | 0.93 |
ENSMUST00000110832.8
|
Tmem184a
|
transmembrane protein 184a |
| chr5_-_24534554 | 0.92 |
ENSMUST00000115098.7
|
Kcnh2
|
potassium voltage-gated channel, subfamily H (eag-related), member 2 |
| chr2_+_3714515 | 0.90 |
ENSMUST00000115053.9
|
Fam107b
|
family with sequence similarity 107, member B |
| chr17_-_71156639 | 0.89 |
ENSMUST00000134654.2
ENSMUST00000172229.8 ENSMUST00000127719.2 |
Tgif1
|
TGFB-induced factor homeobox 1 |
| chr17_+_43327412 | 0.89 |
ENSMUST00000024708.6
|
Tnfrsf21
|
tumor necrosis factor receptor superfamily, member 21 |
| chr10_-_84938350 | 0.88 |
ENSMUST00000059383.8
ENSMUST00000216889.2 |
Fhl4
|
four and a half LIM domains 4 |
| chr19_-_6065415 | 0.87 |
ENSMUST00000237519.2
|
Capn1
|
calpain 1 |
| chr8_+_106024294 | 0.87 |
ENSMUST00000015003.10
|
E2f4
|
E2F transcription factor 4 |
| chr18_+_82929451 | 0.85 |
ENSMUST00000235902.2
|
Zfp516
|
zinc finger protein 516 |
| chr17_-_48189815 | 0.80 |
ENSMUST00000154108.2
|
Foxp4
|
forkhead box P4 |
| chr17_+_29709723 | 0.80 |
ENSMUST00000024811.9
|
Pim1
|
proviral integration site 1 |
| chr1_-_135211813 | 0.79 |
ENSMUST00000077340.14
ENSMUST00000074357.8 |
Rnpep
|
arginyl aminopeptidase (aminopeptidase B) |
| chr5_+_149335214 | 0.79 |
ENSMUST00000093110.12
|
Medag
|
mesenteric estrogen dependent adipogenesis |
| chr7_+_127078371 | 0.78 |
ENSMUST00000205432.3
|
Fbrs
|
fibrosin |
| chr11_-_94133527 | 0.78 |
ENSMUST00000061469.4
|
Wfikkn2
|
WAP, follistatin/kazal, immunoglobulin, kunitz and netrin domain containing 2 |
| chr14_-_86986541 | 0.78 |
ENSMUST00000226254.2
|
Diaph3
|
diaphanous related formin 3 |
| chr15_+_102927366 | 0.77 |
ENSMUST00000165375.3
|
Hoxc4
|
homeobox C4 |
| chr11_-_100160697 | 0.77 |
ENSMUST00000017270.8
|
Krt42
|
keratin 42 |
| chr4_-_133695204 | 0.77 |
ENSMUST00000100472.10
ENSMUST00000136327.2 |
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr8_+_106412905 | 0.76 |
ENSMUST00000213019.2
|
Carmil2
|
capping protein regulator and myosin 1 linker 2 |
| chr1_-_96799832 | 0.76 |
ENSMUST00000071985.6
|
Slco4c1
|
solute carrier organic anion transporter family, member 4C1 |
| chr14_-_30329765 | 0.75 |
ENSMUST00000112207.8
ENSMUST00000112206.8 ENSMUST00000112202.8 ENSMUST00000112203.2 |
Prkcd
|
protein kinase C, delta |
| chr7_-_66339319 | 0.74 |
ENSMUST00000124899.8
|
Asb7
|
ankyrin repeat and SOCS box-containing 7 |
| chr3_+_106943472 | 0.73 |
ENSMUST00000052718.5
|
Kcna3
|
potassium voltage-gated channel, shaker-related subfamily, member 3 |
| chr2_+_60040231 | 0.73 |
ENSMUST00000102748.11
|
Marchf7
|
membrane associated ring-CH-type finger 7 |
| chr9_+_62249730 | 0.73 |
ENSMUST00000156461.2
|
Anp32a
|
acidic (leucine-rich) nuclear phosphoprotein 32 family, member A |
| chr2_-_154411765 | 0.72 |
ENSMUST00000103145.11
|
E2f1
|
E2F transcription factor 1 |
| chr12_+_108376801 | 0.71 |
ENSMUST00000054955.14
|
Eml1
|
echinoderm microtubule associated protein like 1 |
| chr10_-_116417333 | 0.70 |
ENSMUST00000218744.2
ENSMUST00000105267.8 ENSMUST00000105265.8 ENSMUST00000167706.8 ENSMUST00000168036.8 ENSMUST00000169921.8 ENSMUST00000020374.6 |
Cnot2
|
CCR4-NOT transcription complex, subunit 2 |
| chr6_-_124710030 | 0.70 |
ENSMUST00000173647.2
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr4_+_41762309 | 0.70 |
ENSMUST00000108042.3
|
Il11ra1
|
interleukin 11 receptor, alpha chain 1 |
| chr5_+_91175323 | 0.69 |
ENSMUST00000202724.4
ENSMUST00000041516.9 |
Epgn
|
epithelial mitogen |
| chr9_-_54554483 | 0.67 |
ENSMUST00000128163.8
|
Acsbg1
|
acyl-CoA synthetase bubblegum family member 1 |
| chr7_+_24981604 | 0.67 |
ENSMUST00000163320.8
ENSMUST00000005578.13 |
Cic
|
capicua transcriptional repressor |
| chr5_+_31070739 | 0.67 |
ENSMUST00000031055.8
|
Emilin1
|
elastin microfibril interfacer 1 |
| chr15_+_103411461 | 0.66 |
ENSMUST00000023132.5
|
Pde1b
|
phosphodiesterase 1B, Ca2+-calmodulin dependent |
| chr7_+_87927293 | 0.65 |
ENSMUST00000032779.12
ENSMUST00000131108.9 ENSMUST00000128791.2 |
Ctsc
|
cathepsin C |
| chr11_-_50183129 | 0.64 |
ENSMUST00000059458.5
|
Maml1
|
mastermind like transcriptional coactivator 1 |
| chr1_+_39940043 | 0.63 |
ENSMUST00000168431.7
ENSMUST00000163854.9 |
Map4k4
|
mitogen-activated protein kinase kinase kinase kinase 4 |
| chr13_+_13612136 | 0.63 |
ENSMUST00000005532.9
|
Nid1
|
nidogen 1 |
| chr10_-_121312212 | 0.63 |
ENSMUST00000026902.9
|
Rassf3
|
Ras association (RalGDS/AF-6) domain family member 3 |
| chr4_-_133965683 | 0.63 |
ENSMUST00000030645.9
|
Cnksr1
|
connector enhancer of kinase suppressor of Ras 1 |
| chr7_+_140659038 | 0.62 |
ENSMUST00000159375.8
|
Pkp3
|
plakophilin 3 |
| chr11_+_60590498 | 0.61 |
ENSMUST00000052346.10
|
Llgl1
|
LLGL1 scribble cell polarity complex component |
| chr6_-_39702381 | 0.60 |
ENSMUST00000002487.15
|
Braf
|
Braf transforming gene |
| chr11_+_60590584 | 0.60 |
ENSMUST00000108719.4
|
Llgl1
|
LLGL1 scribble cell polarity complex component |
| chr12_+_108376884 | 0.60 |
ENSMUST00000109857.8
|
Eml1
|
echinoderm microtubule associated protein like 1 |
| chr10_+_67371295 | 0.59 |
ENSMUST00000145754.8
ENSMUST00000145936.2 |
Egr2
|
early growth response 2 |
| chr16_-_20244631 | 0.58 |
ENSMUST00000077867.10
|
Abcc5
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 5 |
| chr16_+_24266829 | 0.58 |
ENSMUST00000078988.10
|
Lpp
|
LIM domain containing preferred translocation partner in lipoma |
| chr8_-_91074754 | 0.57 |
ENSMUST00000176034.2
ENSMUST00000176616.8 |
Tox3
|
TOX high mobility group box family member 3 |
| chr6_-_115652831 | 0.57 |
ENSMUST00000112949.8
|
Raf1
|
v-raf-leukemia viral oncogene 1 |
| chr7_+_89053562 | 0.56 |
ENSMUST00000058755.5
|
Fzd4
|
frizzled class receptor 4 |
| chr8_+_40876827 | 0.56 |
ENSMUST00000049389.11
ENSMUST00000128166.8 ENSMUST00000167766.2 |
Zdhhc2
|
zinc finger, DHHC domain containing 2 |
| chr7_-_97958679 | 0.56 |
ENSMUST00000033020.14
|
Acer3
|
alkaline ceramidase 3 |
| chr6_-_115653583 | 0.54 |
ENSMUST00000000451.14
|
Raf1
|
v-raf-leukemia viral oncogene 1 |
| chr17_+_69690018 | 0.54 |
ENSMUST00000112674.8
|
Zbtb14
|
zinc finger and BTB domain containing 14 |
| chr9_+_37400577 | 0.53 |
ENSMUST00000211060.2
ENSMUST00000239463.2 ENSMUST00000048604.8 |
Msantd2
|
Myb/SANT-like DNA-binding domain containing 2 |
| chr11_+_35011953 | 0.53 |
ENSMUST00000069837.4
|
Slit3
|
slit guidance ligand 3 |
| chr10_-_59787646 | 0.53 |
ENSMUST00000020308.5
|
Ddit4
|
DNA-damage-inducible transcript 4 |
| chr2_-_181335518 | 0.52 |
ENSMUST00000108776.8
ENSMUST00000108771.2 |
Rgs19
|
regulator of G-protein signaling 19 |
| chr7_-_34353767 | 0.52 |
ENSMUST00000206501.2
ENSMUST00000108069.8 |
Kctd15
|
potassium channel tetramerisation domain containing 15 |
| chr6_-_83654789 | 0.51 |
ENSMUST00000037882.8
|
Cd207
|
CD207 antigen |
| chr17_+_87590308 | 0.51 |
ENSMUST00000041110.12
ENSMUST00000125875.8 |
Ttc7
|
tetratricopeptide repeat domain 7 |
| chr8_+_106331866 | 0.50 |
ENSMUST00000043531.10
|
Ripor1
|
RHO family interacting cell polarization regulator 1 |
| chr19_-_6065799 | 0.50 |
ENSMUST00000235138.2
|
Capn1
|
calpain 1 |
| chr19_+_53665719 | 0.49 |
ENSMUST00000164202.9
|
Rbm20
|
RNA binding motif protein 20 |
| chr9_+_61280890 | 0.49 |
ENSMUST00000161689.8
|
Tle3
|
transducin-like enhancer of split 3 |
| chr7_+_79882609 | 0.47 |
ENSMUST00000062915.9
|
Gdpgp1
|
GDP-D-glucose phosphorylase 1 |
| chr13_-_10410857 | 0.46 |
ENSMUST00000187510.7
|
Chrm3
|
cholinergic receptor, muscarinic 3, cardiac |
| chr4_+_155924181 | 0.46 |
ENSMUST00000030947.4
|
Mxra8
|
matrix-remodelling associated 8 |
| chr11_+_121150798 | 0.45 |
ENSMUST00000106113.2
|
Foxk2
|
forkhead box K2 |
| chr10_+_19232281 | 0.45 |
ENSMUST00000053225.7
|
Olig3
|
oligodendrocyte transcription factor 3 |
| chr1_+_55445033 | 0.44 |
ENSMUST00000042986.10
|
Plcl1
|
phospholipase C-like 1 |
| chr17_-_13211374 | 0.43 |
ENSMUST00000159551.8
ENSMUST00000160781.8 |
Wtap
|
Wilms tumour 1-associating protein |
| chr13_+_24685508 | 0.43 |
ENSMUST00000238974.2
|
Ripor2
|
RHO family interacting cell polarization regulator 2 |
| chr2_+_152873772 | 0.41 |
ENSMUST00000037235.7
|
Xkr7
|
X-linked Kx blood group related 7 |
| chr1_-_177085736 | 0.41 |
ENSMUST00000111159.2
|
Akt3
|
thymoma viral proto-oncogene 3 |
| chr1_-_22031718 | 0.40 |
ENSMUST00000029667.13
ENSMUST00000173058.8 ENSMUST00000173404.2 |
Kcnq5
|
potassium voltage-gated channel, subfamily Q, member 5 |
| chr10_-_119075910 | 0.40 |
ENSMUST00000020315.13
|
Cand1
|
cullin associated and neddylation disassociated 1 |
| chr2_-_155199300 | 0.40 |
ENSMUST00000165234.2
ENSMUST00000077626.13 |
Pigu
|
phosphatidylinositol glycan anchor biosynthesis, class U |
| chr10_-_58511476 | 0.40 |
ENSMUST00000003312.5
|
Edar
|
ectodysplasin-A receptor |
| chr11_-_97635484 | 0.37 |
ENSMUST00000018691.9
|
Pip4k2b
|
phosphatidylinositol-5-phosphate 4-kinase, type II, beta |
| chr16_-_21982049 | 0.37 |
ENSMUST00000100052.11
|
Igf2bp2
|
insulin-like growth factor 2 mRNA binding protein 2 |
| chr14_+_56122404 | 0.37 |
ENSMUST00000022831.5
|
Khnyn
|
KH and NYN domain containing |
| chr9_-_106762818 | 0.37 |
ENSMUST00000185707.2
|
Rbm15b
|
RNA binding motif protein 15B |
| chr19_+_5790918 | 0.37 |
ENSMUST00000081496.6
|
Ltbp3
|
latent transforming growth factor beta binding protein 3 |
| chr15_-_60696790 | 0.36 |
ENSMUST00000100635.5
|
Lratd2
|
LRAT domain containing 1 |
| chr6_-_71299184 | 0.36 |
ENSMUST00000173297.2
ENSMUST00000114188.3 |
Smyd1
|
SET and MYND domain containing 1 |
| chr7_+_28834391 | 0.36 |
ENSMUST00000160194.8
|
Rasgrp4
|
RAS guanyl releasing protein 4 |
| chr6_+_91492910 | 0.36 |
ENSMUST00000040607.6
|
Lsm3
|
LSM3 homolog, U6 small nuclear RNA and mRNA degradation associated |
| chr10_-_116984386 | 0.35 |
ENSMUST00000020381.5
|
Frs2
|
fibroblast growth factor receptor substrate 2 |
| chr10_+_84938452 | 0.35 |
ENSMUST00000095383.6
|
Tmem263
|
transmembrane protein 263 |
| chr17_+_87270504 | 0.35 |
ENSMUST00000024956.15
|
Rhoq
|
ras homolog family member Q |
| chr3_+_145823205 | 0.35 |
ENSMUST00000140214.3
|
Mcoln3
|
mucolipin 3 |
| chr2_-_11506511 | 0.34 |
ENSMUST00000183869.8
|
Pfkfb3
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3 |
| chrX_+_7439839 | 0.34 |
ENSMUST00000144719.9
ENSMUST00000234896.2 |
Flicr
Foxp3
|
Foxp3 regulating long intergenic noncoding RNA forkhead box P3 |
| chr2_-_91854844 | 0.34 |
ENSMUST00000028663.5
|
Creb3l1
|
cAMP responsive element binding protein 3-like 1 |
| chr2_+_30176395 | 0.33 |
ENSMUST00000064447.12
|
Nup188
|
nucleoporin 188 |
| chr7_+_46025890 | 0.33 |
ENSMUST00000072514.3
|
Myod1
|
myogenic differentiation 1 |
| chr2_-_93787441 | 0.33 |
ENSMUST00000099689.5
|
Gm13889
|
predicted gene 13889 |
| chr10_-_61288437 | 0.33 |
ENSMUST00000167087.2
ENSMUST00000020288.15 |
Eif4ebp2
|
eukaryotic translation initiation factor 4E binding protein 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 5.0 | GO:0036414 | protein citrullination(GO:0018101) histone citrullination(GO:0036414) |
| 0.8 | 4.2 | GO:0051122 | hepoxilin metabolic process(GO:0051121) hepoxilin biosynthetic process(GO:0051122) |
| 0.7 | 3.7 | GO:0003430 | growth plate cartilage chondrocyte growth(GO:0003430) |
| 0.7 | 2.0 | GO:0019049 | evasion or tolerance of host defenses by virus(GO:0019049) avoidance of host defenses(GO:0044413) evasion or tolerance of host defenses(GO:0044415) avoidance of defenses of other organism involved in symbiotic interaction(GO:0051832) evasion or tolerance of defenses of other organism involved in symbiotic interaction(GO:0051834) negative regulation of hyaluronan biosynthetic process(GO:1900126) |
| 0.6 | 1.9 | GO:0048627 | myoblast development(GO:0048627) positive regulation of G0 to G1 transition(GO:0070318) |
| 0.6 | 12.8 | GO:0008272 | sulfate transport(GO:0008272) |
| 0.6 | 1.7 | GO:0060734 | regulation of endoplasmic reticulum stress-induced eIF2 alpha phosphorylation(GO:0060734) |
| 0.5 | 2.7 | GO:2000721 | positive regulation of transcription from RNA polymerase II promoter involved in smooth muscle cell differentiation(GO:2000721) |
| 0.5 | 1.9 | GO:0032696 | negative regulation of interleukin-13 production(GO:0032696) |
| 0.5 | 1.4 | GO:0051659 | maintenance of mitochondrion location(GO:0051659) relaxation of skeletal muscle(GO:0090076) |
| 0.4 | 1.6 | GO:0090095 | lactic acid secretion(GO:0046722) regulation of metanephric cap mesenchymal cell proliferation(GO:0090095) positive regulation of metanephric cap mesenchymal cell proliferation(GO:0090096) |
| 0.4 | 1.2 | GO:0045074 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.4 | 3.0 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.4 | 1.1 | GO:0071550 | death-inducing signaling complex assembly(GO:0071550) |
| 0.4 | 1.8 | GO:0003409 | optic cup structural organization(GO:0003409) |
| 0.3 | 1.4 | GO:0010609 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902164) |
| 0.3 | 1.6 | GO:0007223 | Wnt signaling pathway, calcium modulating pathway(GO:0007223) |
| 0.3 | 1.0 | GO:0045014 | carbon catabolite repression of transcription(GO:0045013) negative regulation of transcription by glucose(GO:0045014) |
| 0.3 | 0.8 | GO:0032976 | release of matrix enzymes from mitochondria(GO:0032976) B cell receptor apoptotic signaling pathway(GO:1990117) |
| 0.3 | 2.6 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.3 | 0.8 | GO:2000813 | negative regulation of barbed-end actin filament capping(GO:2000813) |
| 0.2 | 1.7 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.2 | 1.0 | GO:0032765 | positive regulation of mast cell cytokine production(GO:0032765) regulation of monocyte aggregation(GO:1900623) positive regulation of monocyte aggregation(GO:1900625) |
| 0.2 | 0.9 | GO:2001107 | negative regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001107) |
| 0.2 | 0.9 | GO:1902303 | negative regulation of potassium ion export(GO:1902303) |
| 0.2 | 1.6 | GO:0003383 | apical constriction(GO:0003383) |
| 0.2 | 0.6 | GO:0021666 | rhombomere formation(GO:0021594) rhombomere 3 formation(GO:0021660) rhombomere 5 morphogenesis(GO:0021664) rhombomere 5 formation(GO:0021666) |
| 0.2 | 0.8 | GO:0009786 | regulation of asymmetric cell division(GO:0009786) |
| 0.2 | 0.8 | GO:0060697 | positive regulation of phospholipid catabolic process(GO:0060697) |
| 0.2 | 0.7 | GO:0071930 | negative regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0071930) |
| 0.2 | 3.4 | GO:1903818 | positive regulation of voltage-gated potassium channel activity(GO:1903818) |
| 0.2 | 0.5 | GO:0070100 | negative regulation of chemokine-mediated signaling pathway(GO:0070100) |
| 0.2 | 0.6 | GO:0003162 | atrioventricular node development(GO:0003162) |
| 0.2 | 0.9 | GO:0033326 | cerebrospinal fluid secretion(GO:0033326) |
| 0.2 | 0.9 | GO:0018992 | germ-line sex determination(GO:0018992) |
| 0.2 | 2.9 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.2 | 1.5 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.1 | 2.2 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.1 | 4.4 | GO:0040034 | regulation of development, heterochronic(GO:0040034) |
| 0.1 | 3.2 | GO:1900017 | positive regulation of cytokine production involved in inflammatory response(GO:1900017) |
| 0.1 | 1.1 | GO:0097411 | hypoxia-inducible factor-1alpha signaling pathway(GO:0097411) |
| 0.1 | 2.0 | GO:0072540 | T-helper 17 cell lineage commitment(GO:0072540) |
| 0.1 | 3.0 | GO:0000920 | cell separation after cytokinesis(GO:0000920) |
| 0.1 | 0.8 | GO:2000138 | positive regulation of cell proliferation involved in heart morphogenesis(GO:2000138) |
| 0.1 | 0.4 | GO:0036363 | transforming growth factor beta activation(GO:0036363) regulation of mesenchymal stem cell proliferation(GO:1902460) positive regulation of mesenchymal stem cell proliferation(GO:1902462) |
| 0.1 | 3.5 | GO:0043486 | histone exchange(GO:0043486) |
| 0.1 | 0.4 | GO:0046619 | optic placode formation involved in camera-type eye formation(GO:0046619) |
| 0.1 | 0.3 | GO:0002362 | CD4-positive, CD25-positive, alpha-beta regulatory T cell lineage commitment(GO:0002362) |
| 0.1 | 0.3 | GO:0014908 | myotube differentiation involved in skeletal muscle regeneration(GO:0014908) |
| 0.1 | 0.2 | GO:0030997 | regulation of centriole-centriole cohesion(GO:0030997) |
| 0.1 | 1.4 | GO:0044336 | canonical Wnt signaling pathway involved in negative regulation of apoptotic process(GO:0044336) |
| 0.1 | 0.6 | GO:0002159 | desmosome assembly(GO:0002159) |
| 0.1 | 0.3 | GO:2000597 | optic nerve formation(GO:0021634) optic chiasma development(GO:0061360) regulation of optic nerve formation(GO:2000595) positive regulation of optic nerve formation(GO:2000597) |
| 0.1 | 0.9 | GO:2000301 | negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.1 | 0.2 | GO:0061153 | trachea submucosa development(GO:0061152) trachea gland development(GO:0061153) |
| 0.1 | 0.5 | GO:0060857 | establishment of glial blood-brain barrier(GO:0060857) |
| 0.1 | 0.4 | GO:0097476 | spinal cord motor neuron migration(GO:0097476) |
| 0.1 | 0.3 | GO:1904924 | negative regulation of mitophagy in response to mitochondrial depolarization(GO:1904924) |
| 0.1 | 1.6 | GO:0043312 | neutrophil degranulation(GO:0043312) |
| 0.1 | 2.3 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.1 | 1.0 | GO:0060040 | retinal bipolar neuron differentiation(GO:0060040) |
| 0.1 | 0.2 | GO:0008588 | release of cytoplasmic sequestered NF-kappaB(GO:0008588) |
| 0.1 | 2.3 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.1 | 1.3 | GO:0048733 | sebaceous gland development(GO:0048733) |
| 0.1 | 0.8 | GO:1901250 | regulation of lung goblet cell differentiation(GO:1901249) negative regulation of lung goblet cell differentiation(GO:1901250) |
| 0.1 | 1.2 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 0.1 | 5.5 | GO:0051693 | actin filament capping(GO:0051693) |
| 0.1 | 0.3 | GO:0070278 | extracellular matrix constituent secretion(GO:0070278) |
| 0.1 | 0.4 | GO:0097473 | cellular response to light intensity(GO:0071484) cellular response to high light intensity(GO:0071486) retinal rod cell apoptotic process(GO:0097473) |
| 0.1 | 0.2 | GO:0000715 | nucleotide-excision repair, DNA damage recognition(GO:0000715) |
| 0.1 | 0.3 | GO:0046501 | protoporphyrinogen IX metabolic process(GO:0046501) |
| 0.1 | 1.6 | GO:0007263 | nitric oxide mediated signal transduction(GO:0007263) |
| 0.1 | 0.9 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.1 | 4.2 | GO:0048536 | spleen development(GO:0048536) |
| 0.1 | 0.7 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.1 | 0.2 | GO:1900110 | negative regulation of histone H3-K9 dimethylation(GO:1900110) |
| 0.1 | 0.3 | GO:0035092 | sperm chromatin condensation(GO:0035092) |
| 0.1 | 0.4 | GO:0090521 | glomerular visceral epithelial cell migration(GO:0090521) |
| 0.1 | 0.4 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.1 | 0.8 | GO:0036006 | response to macrophage colony-stimulating factor(GO:0036005) cellular response to macrophage colony-stimulating factor stimulus(GO:0036006) |
| 0.1 | 2.7 | GO:0060216 | definitive hemopoiesis(GO:0060216) |
| 0.1 | 0.3 | GO:0031133 | regulation of axon diameter(GO:0031133) |
| 0.1 | 0.3 | GO:0034465 | response to carbon monoxide(GO:0034465) eye blink reflex(GO:0060082) |
| 0.1 | 0.8 | GO:0045820 | negative regulation of glycolytic process(GO:0045820) |
| 0.1 | 0.3 | GO:0072531 | pyrimidine-containing compound transmembrane transport(GO:0072531) |
| 0.0 | 0.4 | GO:0060662 | tube lumen cavitation(GO:0060605) salivary gland cavitation(GO:0060662) |
| 0.0 | 0.4 | GO:1900122 | positive regulation of receptor binding(GO:1900122) |
| 0.0 | 7.1 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.0 | 0.9 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.0 | 0.6 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.0 | 0.1 | GO:0008292 | acetylcholine biosynthetic process(GO:0008292) acetate ester biosynthetic process(GO:1900620) |
| 0.0 | 0.3 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.0 | 0.1 | GO:0090327 | negative regulation of locomotion involved in locomotory behavior(GO:0090327) |
| 0.0 | 2.4 | GO:0006414 | translational elongation(GO:0006414) |
| 0.0 | 0.1 | GO:0072049 | comma-shaped body morphogenesis(GO:0072049) |
| 0.0 | 0.9 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.0 | 0.7 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) |
| 0.0 | 0.6 | GO:0030644 | cellular chloride ion homeostasis(GO:0030644) |
| 0.0 | 0.7 | GO:0046697 | decidualization(GO:0046697) |
| 0.0 | 0.6 | GO:0046512 | sphingosine biosynthetic process(GO:0046512) |
| 0.0 | 0.6 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.0 | 1.6 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 | 0.4 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.0 | 0.6 | GO:0033962 | cytoplasmic mRNA processing body assembly(GO:0033962) |
| 0.0 | 0.3 | GO:0000414 | regulation of histone H3-K36 methylation(GO:0000414) |
| 0.0 | 0.3 | GO:0072513 | positive regulation of secondary heart field cardioblast proliferation(GO:0072513) |
| 0.0 | 0.4 | GO:0045793 | positive regulation of cell size(GO:0045793) |
| 0.0 | 1.3 | GO:0051310 | metaphase plate congression(GO:0051310) |
| 0.0 | 0.1 | GO:0019509 | L-methionine biosynthetic process from methylthioadenosine(GO:0019509) |
| 0.0 | 0.1 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.0 | 0.4 | GO:0045899 | positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.0 | 0.3 | GO:0034472 | snRNA 3'-end processing(GO:0034472) |
| 0.0 | 1.8 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 0.2 | GO:2000786 | positive regulation of autophagosome assembly(GO:2000786) |
| 0.0 | 0.3 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.0 | 4.0 | GO:1990830 | response to leukemia inhibitory factor(GO:1990823) cellular response to leukemia inhibitory factor(GO:1990830) |
| 0.0 | 5.1 | GO:0007160 | cell-matrix adhesion(GO:0007160) |
| 0.0 | 0.7 | GO:0001913 | T cell mediated cytotoxicity(GO:0001913) |
| 0.0 | 1.1 | GO:0060612 | adipose tissue development(GO:0060612) |
| 0.0 | 2.7 | GO:0050680 | negative regulation of epithelial cell proliferation(GO:0050680) |
| 0.0 | 1.1 | GO:0048146 | positive regulation of fibroblast proliferation(GO:0048146) |
| 0.0 | 0.5 | GO:0045987 | positive regulation of smooth muscle contraction(GO:0045987) |
| 0.0 | 0.3 | GO:0021520 | spinal cord motor neuron cell fate specification(GO:0021520) |
| 0.0 | 0.3 | GO:0045947 | negative regulation of translational initiation(GO:0045947) |
| 0.0 | 0.8 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 1.3 | GO:0046579 | positive regulation of Ras protein signal transduction(GO:0046579) |
| 0.0 | 1.1 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.0 | 1.0 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.0 | 0.3 | GO:0033169 | histone H3-K9 demethylation(GO:0033169) |
| 0.0 | 0.2 | GO:0033148 | positive regulation of intracellular estrogen receptor signaling pathway(GO:0033148) |
| 0.0 | 0.3 | GO:0021854 | hypothalamus development(GO:0021854) |
| 0.0 | 1.4 | GO:0007052 | mitotic spindle organization(GO:0007052) |
| 0.0 | 0.4 | GO:0045663 | positive regulation of myoblast differentiation(GO:0045663) |
| 0.0 | 0.8 | GO:0048747 | muscle fiber development(GO:0048747) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.5 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 0.3 | 0.8 | GO:0097144 | BAX complex(GO:0097144) |
| 0.3 | 9.2 | GO:0030057 | desmosome(GO:0030057) |
| 0.2 | 3.0 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.2 | 0.9 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.2 | 3.0 | GO:0090543 | Flemming body(GO:0090543) |
| 0.1 | 0.7 | GO:0035189 | Rb-E2F complex(GO:0035189) |
| 0.1 | 1.1 | GO:0008091 | spectrin(GO:0008091) |
| 0.1 | 0.9 | GO:0031673 | H zone(GO:0031673) |
| 0.1 | 1.7 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.1 | 0.4 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.1 | 2.2 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.1 | 12.5 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.1 | 2.6 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.1 | 1.0 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.1 | 1.2 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.1 | 0.6 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.1 | 0.8 | GO:0044354 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 0.1 | 0.6 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.1 | 0.3 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.1 | 1.1 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.1 | 0.7 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.1 | 0.4 | GO:0036396 | MIS complex(GO:0036396) |
| 0.1 | 0.4 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.1 | 0.4 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.1 | 1.1 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.0 | 0.2 | GO:0071942 | XPC complex(GO:0071942) |
| 0.0 | 0.5 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.0 | 2.4 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 1.6 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.0 | 2.4 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 4.2 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.0 | 0.3 | GO:0019907 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) |
| 0.0 | 0.1 | GO:1902737 | dendritic filopodium(GO:1902737) |
| 0.0 | 1.6 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.0 | 0.4 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.0 | 0.3 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.0 | 2.6 | GO:0014704 | intercalated disc(GO:0014704) |
| 0.0 | 0.6 | GO:0005605 | basal lamina(GO:0005605) |
| 0.0 | 4.4 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 8.1 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.0 | 0.4 | GO:0071004 | U2-type prespliceosome(GO:0071004) |
| 0.0 | 1.1 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 1.5 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 0.7 | GO:0008305 | integrin complex(GO:0008305) |
| 0.0 | 0.3 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.0 | 1.9 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 0.6 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 0.6 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 0.2 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 0.1 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.0 | 0.6 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 0.2 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 0.1 | GO:0001739 | sex chromatin(GO:0001739) |
| 0.0 | 0.5 | GO:0016235 | aggresome(GO:0016235) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 4.2 | GO:0004052 | arachidonate 12-lipoxygenase activity(GO:0004052) |
| 1.0 | 5.0 | GO:0004668 | protein-arginine deiminase activity(GO:0004668) |
| 0.7 | 12.8 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.3 | 2.0 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 0.3 | 2.8 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.3 | 2.2 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.3 | 1.0 | GO:0030284 | estrogen receptor activity(GO:0030284) |
| 0.2 | 1.7 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.2 | 3.0 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.2 | 0.7 | GO:0098640 | integrin binding involved in cell-matrix adhesion(GO:0098640) |
| 0.2 | 2.3 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.2 | 0.8 | GO:0070976 | calcium-independent protein kinase C activity(GO:0004699) TIR domain binding(GO:0070976) |
| 0.2 | 3.7 | GO:0003708 | retinoic acid receptor activity(GO:0003708) |
| 0.2 | 0.7 | GO:0019970 | interleukin-11 receptor activity(GO:0004921) interleukin-11 binding(GO:0019970) |
| 0.2 | 1.5 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.2 | 0.6 | GO:0030629 | U6 snRNA 3'-end binding(GO:0030629) |
| 0.2 | 1.7 | GO:0048531 | beta-1,3-galactosyltransferase activity(GO:0048531) |
| 0.2 | 0.9 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.1 | 3.2 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.1 | 6.2 | GO:0043236 | laminin binding(GO:0043236) |
| 0.1 | 4.5 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 2.2 | GO:0030235 | nitric-oxide synthase regulator activity(GO:0030235) |
| 0.1 | 0.7 | GO:0001226 | RNA polymerase II transcription corepressor binding(GO:0001226) |
| 0.1 | 0.5 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.1 | 1.4 | GO:0005338 | nucleotide-sugar transmembrane transporter activity(GO:0005338) |
| 0.1 | 5.7 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.1 | 1.4 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.1 | 3.2 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.1 | 1.6 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.1 | 0.8 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.1 | 2.0 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.1 | 0.5 | GO:0004645 | phosphorylase activity(GO:0004645) |
| 0.1 | 1.7 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.1 | 2.3 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.1 | 0.6 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.1 | 0.2 | GO:0016309 | 1-phosphatidylinositol-5-phosphate 4-kinase activity(GO:0016309) |
| 0.1 | 0.8 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.1 | 2.7 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.1 | 2.3 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.1 | 0.7 | GO:0004117 | calmodulin-dependent cyclic-nucleotide phosphodiesterase activity(GO:0004117) calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.1 | 0.4 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.1 | 0.7 | GO:0016505 | peptidase activator activity involved in apoptotic process(GO:0016505) |
| 0.0 | 1.9 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.0 | 0.8 | GO:0050897 | cobalt ion binding(GO:0050897) |
| 0.0 | 2.2 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.0 | 1.6 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.0 | 0.6 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.0 | 0.3 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.0 | 0.1 | GO:0097677 | STAT family protein binding(GO:0097677) |
| 0.0 | 0.1 | GO:0035939 | microsatellite binding(GO:0035939) |
| 0.0 | 0.3 | GO:0042799 | histone methyltransferase activity (H4-K20 specific)(GO:0042799) |
| 0.0 | 0.8 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 2.7 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.7 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 0.3 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.0 | 1.4 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 0.9 | GO:0070410 | JUN kinase binding(GO:0008432) co-SMAD binding(GO:0070410) |
| 0.0 | 1.6 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.0 | 1.1 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 0.3 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 0.4 | GO:0070679 | inositol 1,4,5 trisphosphate binding(GO:0070679) |
| 0.0 | 0.5 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 0.4 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.0 | 0.9 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.0 | 0.3 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.0 | 0.3 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.0 | 0.5 | GO:0042166 | acetylcholine binding(GO:0042166) |
| 0.0 | 2.4 | GO:0005249 | voltage-gated potassium channel activity(GO:0005249) |
| 0.0 | 0.2 | GO:0001164 | RNA polymerase I regulatory region DNA binding(GO:0001013) RNA polymerase I regulatory region sequence-specific DNA binding(GO:0001163) RNA polymerase I CORE element sequence-specific DNA binding(GO:0001164) |
| 0.0 | 0.4 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.0 | 0.3 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.0 | 5.3 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 3.0 | GO:0043492 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) ATPase activity, coupled to movement of substances(GO:0043492) |
| 0.0 | 1.3 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 0.9 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 0.1 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.0 | 0.3 | GO:0001104 | RNA polymerase II transcription cofactor activity(GO:0001104) |
| 0.0 | 0.8 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.0 | 0.9 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.0 | 0.9 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.0 | 1.0 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.0 | 0.6 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.1 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.0 | 0.3 | GO:0015215 | nucleotide transmembrane transporter activity(GO:0015215) |
| 0.0 | 0.4 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.3 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.0 | 1.4 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 0.7 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 10.3 | GO:0005198 | structural molecule activity(GO:0005198) |
| 0.0 | 0.4 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.0 | 0.4 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.0 | 0.1 | GO:0030619 | U1 snRNA binding(GO:0030619) |
| 0.0 | 0.3 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.0 | 6.8 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 2.6 | GO:0003713 | transcription coactivator activity(GO:0003713) |
| 0.0 | 0.8 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 7.7 | GO:0005509 | calcium ion binding(GO:0005509) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.0 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.1 | 2.0 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.1 | 3.7 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.1 | 0.2 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 3.1 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.1 | 0.8 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.1 | 4.4 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.1 | 3.6 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
| 0.1 | 0.9 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.1 | 2.2 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 2.9 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.0 | 3.8 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.0 | 4.5 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 1.6 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 3.6 | PID MET PATHWAY | Signaling events mediated by Hepatocyte Growth Factor Receptor (c-Met) |
| 0.0 | 2.2 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.0 | 2.6 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 0.6 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.0 | 3.5 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 1.8 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 2.6 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 0.9 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.0 | 1.0 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 1.5 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 1.0 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 0.3 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.0 | 0.3 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 0.9 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 1.3 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 0.8 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.0 | 0.5 | PID MTOR 4PATHWAY | mTOR signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.6 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.2 | 3.0 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.1 | 3.0 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 0.1 | 2.0 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.1 | 3.4 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 3.6 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.1 | 3.4 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.1 | 2.0 | REACTOME SIGNALLING TO P38 VIA RIT AND RIN | Genes involved in Signalling to p38 via RIT and RIN |
| 0.1 | 1.4 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.1 | 11.8 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.1 | 1.3 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.1 | 0.8 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.0 | 1.5 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 4.7 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 4.8 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 0.4 | REACTOME PROLONGED ERK ACTIVATION EVENTS | Genes involved in Prolonged ERK activation events |
| 0.0 | 0.5 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.0 | 2.7 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 1.5 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 1.1 | REACTOME G BETA GAMMA SIGNALLING THROUGH PLC BETA | Genes involved in G beta:gamma signalling through PLC beta |
| 0.0 | 1.2 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.0 | 0.6 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.0 | 0.8 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.0 | 0.1 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.0 | 1.1 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 1.1 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.0 | 1.1 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 1.8 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 0.5 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.0 | 0.6 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.7 | REACTOME DAG AND IP3 SIGNALING | Genes involved in DAG and IP3 signaling |
| 0.0 | 0.5 | REACTOME REGULATION OF MRNA STABILITY BY PROTEINS THAT BIND AU RICH ELEMENTS | Genes involved in Regulation of mRNA Stability by Proteins that Bind AU-rich Elements |
| 0.0 | 0.3 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.0 | 0.6 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 1.0 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.7 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.0 | 0.4 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |