Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
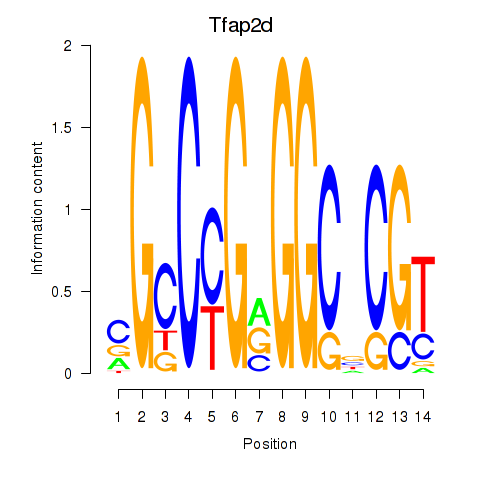
Results for Tfap2d
Z-value: 0.94
Transcription factors associated with Tfap2d
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Tfap2d
|
ENSMUSG00000042596.8 | Tfap2d |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Tfap2d | mm39_v1_chr1_+_19173246_19173267 | 0.04 | 7.1e-01 | Click! |
Activity profile of Tfap2d motif
Sorted Z-values of Tfap2d motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Tfap2d
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_163760603 | 6.04 |
ENSMUST00000044734.3
|
Rims4
|
regulating synaptic membrane exocytosis 4 |
| chrX_+_93278526 | 4.09 |
ENSMUST00000113908.8
ENSMUST00000113916.10 |
Klhl15
|
kelch-like 15 |
| chr5_+_34445944 | 4.04 |
ENSMUST00000207754.2
|
Cfap99
|
cilia and flagella associated protein 99 |
| chr3_+_37117676 | 3.94 |
ENSMUST00000144629.8
|
Adad1
|
adenosine deaminase domain containing 1 (testis specific) |
| chr2_+_92430043 | 3.88 |
ENSMUST00000065797.7
|
Chst1
|
carbohydrate sulfotransferase 1 |
| chrX_+_93278588 | 3.85 |
ENSMUST00000096369.10
ENSMUST00000113911.9 |
Klhl15
|
kelch-like 15 |
| chr3_+_37117779 | 3.84 |
ENSMUST00000029274.14
|
Adad1
|
adenosine deaminase domain containing 1 (testis specific) |
| chr2_+_155907658 | 3.70 |
ENSMUST00000137966.2
|
Spag4
|
sperm associated antigen 4 |
| chr16_-_38533597 | 3.67 |
ENSMUST00000023487.5
|
Arhgap31
|
Rho GTPase activating protein 31 |
| chr2_+_9887427 | 3.67 |
ENSMUST00000114919.2
|
4930412O13Rik
|
RIKEN cDNA 4930412O13 gene |
| chr11_-_109613040 | 3.64 |
ENSMUST00000020938.8
|
Fam20a
|
FAM20A, golgi associated secretory pathway pseudokinase |
| chr5_+_114706077 | 3.36 |
ENSMUST00000043650.8
|
Fam222a
|
family with sequence similarity 222, member A |
| chr8_+_94537460 | 3.32 |
ENSMUST00000034198.15
ENSMUST00000125716.8 |
Gnao1
|
guanine nucleotide binding protein, alpha O |
| chrX_+_93278203 | 3.22 |
ENSMUST00000153900.8
|
Klhl15
|
kelch-like 15 |
| chr8_+_86219191 | 3.19 |
ENSMUST00000034136.12
|
Gpt2
|
glutamic pyruvate transaminase (alanine aminotransferase) 2 |
| chr7_+_106969950 | 3.18 |
ENSMUST00000073459.12
|
Syt9
|
synaptotagmin IX |
| chr3_-_107366868 | 3.16 |
ENSMUST00000009617.10
ENSMUST00000238670.2 |
Kcnc4
|
potassium voltage gated channel, Shaw-related subfamily, member 4 |
| chr15_+_39061612 | 2.94 |
ENSMUST00000082054.12
ENSMUST00000227243.2 ENSMUST00000042917.10 |
Rims2
|
regulating synaptic membrane exocytosis 2 |
| chr11_-_97391166 | 2.94 |
ENSMUST00000056955.2
|
4933428G20Rik
|
RIKEN cDNA 4933428G20 gene |
| chr11_+_98277276 | 2.90 |
ENSMUST00000041301.8
|
Pnmt
|
phenylethanolamine-N-methyltransferase |
| chr14_+_73790105 | 2.89 |
ENSMUST00000160507.8
ENSMUST00000022706.7 |
Sucla2
|
succinate-Coenzyme A ligase, ADP-forming, beta subunit |
| chr12_+_103498542 | 2.81 |
ENSMUST00000021631.12
|
Ppp4r4
|
protein phosphatase 4, regulatory subunit 4 |
| chr3_+_40904253 | 2.75 |
ENSMUST00000048490.13
|
Larp1b
|
La ribonucleoprotein domain family, member 1B |
| chr3_-_54962458 | 2.70 |
ENSMUST00000199352.2
ENSMUST00000198320.5 ENSMUST00000029368.7 |
Ccna1
|
cyclin A1 |
| chr11_+_103664976 | 2.64 |
ENSMUST00000000127.6
|
Wnt3
|
wingless-type MMTV integration site family, member 3 |
| chr2_-_35994819 | 2.60 |
ENSMUST00000148852.4
|
Lhx6
|
LIM homeobox protein 6 |
| chr16_-_18405709 | 2.60 |
ENSMUST00000232335.2
|
Tbx1
|
T-box 1 |
| chr5_+_16139760 | 2.57 |
ENSMUST00000101581.10
ENSMUST00000039370.14 ENSMUST00000199704.5 ENSMUST00000180204.8 ENSMUST00000078272.13 ENSMUST00000115281.7 |
Cacna2d1
|
calcium channel, voltage-dependent, alpha2/delta subunit 1 |
| chr2_+_91087668 | 2.56 |
ENSMUST00000111349.9
ENSMUST00000131711.8 |
Pacsin3
|
protein kinase C and casein kinase substrate in neurons 3 |
| chr19_-_10582672 | 2.52 |
ENSMUST00000236478.2
ENSMUST00000236950.2 |
Tkfc
|
triokinase, FMN cyclase |
| chr6_-_72876269 | 2.51 |
ENSMUST00000204598.3
|
Kcmf1
|
potassium channel modulatory factor 1 |
| chr12_+_71158192 | 2.47 |
ENSMUST00000021482.6
|
Tomm20l
|
translocase of outer mitochondrial membrane 20-like |
| chr9_-_29323032 | 2.46 |
ENSMUST00000115236.2
|
Ntm
|
neurotrimin |
| chr2_+_155907100 | 2.45 |
ENSMUST00000038860.12
|
Spag4
|
sperm associated antigen 4 |
| chr4_-_108637700 | 2.40 |
ENSMUST00000106658.8
|
Zfyve9
|
zinc finger, FYVE domain containing 9 |
| chr8_-_65146079 | 2.29 |
ENSMUST00000048967.9
|
Cpe
|
carboxypeptidase E |
| chr16_+_17464428 | 2.27 |
ENSMUST00000182117.8
ENSMUST00000182671.8 ENSMUST00000182344.8 |
Ccdc74a
|
coiled-coil domain containing 74A |
| chr4_-_108637979 | 2.24 |
ENSMUST00000106657.8
|
Zfyve9
|
zinc finger, FYVE domain containing 9 |
| chr6_-_88851579 | 2.24 |
ENSMUST00000061262.11
ENSMUST00000140455.8 ENSMUST00000145780.2 |
Podxl2
|
podocalyxin-like 2 |
| chr19_-_4748696 | 2.19 |
ENSMUST00000225896.2
ENSMUST00000177696.8 |
Gm960
|
predicted gene 960 |
| chr4_-_46991842 | 2.17 |
ENSMUST00000107749.4
|
Gabbr2
|
gamma-aminobutyric acid (GABA) B receptor, 2 |
| chr5_+_119808890 | 2.15 |
ENSMUST00000121021.8
|
Tbx3
|
T-box 3 |
| chr5_+_119808722 | 2.13 |
ENSMUST00000079719.11
|
Tbx3
|
T-box 3 |
| chr16_+_17464328 | 2.12 |
ENSMUST00000056962.11
|
Ccdc74a
|
coiled-coil domain containing 74A |
| chr14_-_19420488 | 2.10 |
ENSMUST00000166494.2
|
Gm2897
|
predicted gene 2897 |
| chr13_-_58421910 | 2.08 |
ENSMUST00000224505.2
|
Gkap1
|
G kinase anchoring protein 1 |
| chr7_+_118454957 | 2.06 |
ENSMUST00000208658.2
ENSMUST00000098087.9 ENSMUST00000106547.2 |
Iqck
|
IQ motif containing K |
| chr11_+_69826603 | 2.01 |
ENSMUST00000018698.12
|
Ybx2
|
Y box protein 2 |
| chr10_-_31485180 | 2.00 |
ENSMUST00000081989.8
|
Rnf217
|
ring finger protein 217 |
| chr2_-_104324745 | 2.00 |
ENSMUST00000028600.14
|
Hipk3
|
homeodomain interacting protein kinase 3 |
| chr1_-_157240138 | 1.99 |
ENSMUST00000078308.13
|
Rasal2
|
RAS protein activator like 2 |
| chr14_+_14901127 | 1.95 |
ENSMUST00000163790.2
|
Gm3558
|
predicted gene 3558 |
| chr5_-_24806960 | 1.94 |
ENSMUST00000030791.12
|
Smarcd3
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3 |
| chr2_-_35995283 | 1.92 |
ENSMUST00000112960.8
ENSMUST00000112967.12 ENSMUST00000112963.8 |
Lhx6
|
LIM homeobox protein 6 |
| chr2_-_32872019 | 1.91 |
ENSMUST00000153484.7
|
Slc2a8
|
solute carrier family 2, (facilitated glucose transporter), member 8 |
| chr9_+_51959534 | 1.89 |
ENSMUST00000061352.11
|
Rdx
|
radixin |
| chr8_-_121361097 | 1.87 |
ENSMUST00000123927.9
|
1190005I06Rik
|
RIKEN cDNA 1190005I06 gene |
| chr5_+_34817704 | 1.86 |
ENSMUST00000074651.11
ENSMUST00000001112.14 |
Grk4
|
G protein-coupled receptor kinase 4 |
| chr9_-_29323500 | 1.85 |
ENSMUST00000115237.8
|
Ntm
|
neurotrimin |
| chr7_+_106969919 | 1.81 |
ENSMUST00000137663.2
|
Syt9
|
synaptotagmin IX |
| chrX_+_85235370 | 1.80 |
ENSMUST00000026036.5
|
Nr0b1
|
nuclear receptor subfamily 0, group B, member 1 |
| chr3_+_9315662 | 1.79 |
ENSMUST00000155203.2
|
Zbtb10
|
zinc finger and BTB domain containing 10 |
| chr2_-_32872032 | 1.79 |
ENSMUST00000195863.2
ENSMUST00000028129.13 |
Slc2a8
|
solute carrier family 2, (facilitated glucose transporter), member 8 |
| chr14_-_18817743 | 1.78 |
ENSMUST00000167430.8
|
Gm3020
|
predicted gene 3020 |
| chr14_-_24054927 | 1.76 |
ENSMUST00000145596.3
|
Kcnma1
|
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr5_+_28370687 | 1.71 |
ENSMUST00000036177.9
|
En2
|
engrailed 2 |
| chr1_-_60082246 | 1.71 |
ENSMUST00000027172.13
ENSMUST00000191251.7 |
Ica1l
|
islet cell autoantigen 1-like |
| chr10_+_11219117 | 1.68 |
ENSMUST00000069106.5
|
Epm2a
|
epilepsy, progressive myoclonic epilepsy, type 2 gene alpha |
| chr8_-_124242194 | 1.67 |
ENSMUST00000176286.8
ENSMUST00000176155.2 |
Dbndd1
|
dysbindin (dystrobrevin binding protein 1) domain containing 1 |
| chr4_-_41640321 | 1.65 |
ENSMUST00000127306.2
|
Enho
|
energy homeostasis associated |
| chr14_+_16508028 | 1.65 |
ENSMUST00000163885.2
|
Gm3248
|
predicted gene 3248 |
| chr6_+_72332423 | 1.63 |
ENSMUST00000069695.9
ENSMUST00000132243.3 |
Tmem150a
|
transmembrane protein 150A |
| chr6_-_88852017 | 1.62 |
ENSMUST00000145944.3
|
Podxl2
|
podocalyxin-like 2 |
| chr14_-_19261196 | 1.61 |
ENSMUST00000112797.12
ENSMUST00000225885.2 |
D830030K20Rik
|
RIKEN cDNA D830030K20 gene |
| chr2_+_119572770 | 1.59 |
ENSMUST00000028758.8
|
Itpka
|
inositol 1,4,5-trisphosphate 3-kinase A |
| chr13_+_47276132 | 1.58 |
ENSMUST00000068891.12
|
Rnf144b
|
ring finger protein 144B |
| chr5_-_122187884 | 1.57 |
ENSMUST00000111752.10
|
Cux2
|
cut-like homeobox 2 |
| chr8_-_124242072 | 1.55 |
ENSMUST00000177240.8
|
Dbndd1
|
dysbindin (dystrobrevin binding protein 1) domain containing 1 |
| chr6_-_56339251 | 1.53 |
ENSMUST00000166102.8
|
Pde1c
|
phosphodiesterase 1C |
| chr5_+_108842294 | 1.50 |
ENSMUST00000013633.12
|
Fgfrl1
|
fibroblast growth factor receptor-like 1 |
| chr2_+_84489890 | 1.50 |
ENSMUST00000133437.3
|
Btbd18
|
BTB (POZ) domain containing 18 |
| chr5_-_22549688 | 1.49 |
ENSMUST00000062372.14
ENSMUST00000161356.8 |
Reln
|
reelin |
| chr2_-_9888804 | 1.48 |
ENSMUST00000114915.3
|
9230102O04Rik
|
RIKEN cDNA 9230102O04 gene |
| chr7_+_65511777 | 1.48 |
ENSMUST00000055576.12
ENSMUST00000098391.11 |
Pcsk6
|
proprotein convertase subtilisin/kexin type 6 |
| chr19_-_10582773 | 1.47 |
ENSMUST00000237788.2
|
Tkfc
|
triokinase, FMN cyclase |
| chr14_+_16589391 | 1.47 |
ENSMUST00000164484.8
|
Gm8237
|
predicted gene 8237 |
| chr6_+_50087167 | 1.47 |
ENSMUST00000166318.8
ENSMUST00000036236.15 ENSMUST00000204545.3 ENSMUST00000036225.15 |
Mpp6
|
membrane protein, palmitoylated 6 (MAGUK p55 subfamily member 6) |
| chr18_+_82928959 | 1.43 |
ENSMUST00000171238.8
|
Zfp516
|
zinc finger protein 516 |
| chr13_-_69147639 | 1.41 |
ENSMUST00000022013.8
|
Adcy2
|
adenylate cyclase 2 |
| chr2_-_44817173 | 1.40 |
ENSMUST00000130991.8
|
Gtdc1
|
glycosyltransferase-like domain containing 1 |
| chr2_-_146927365 | 1.39 |
ENSMUST00000067020.3
|
Nkx2-4
|
NK2 homeobox 4 |
| chr9_+_54606832 | 1.38 |
ENSMUST00000070070.8
|
Dnaja4
|
DnaJ heat shock protein family (Hsp40) member A4 |
| chr7_+_128858730 | 1.38 |
ENSMUST00000094018.6
ENSMUST00000205896.2 |
Plpp4
|
phospholipid phosphatase 4 |
| chr8_+_70354828 | 1.38 |
ENSMUST00000050373.7
|
Tssk6
|
testis-specific serine kinase 6 |
| chr9_+_54606798 | 1.37 |
ENSMUST00000154690.8
|
Dnaja4
|
DnaJ heat shock protein family (Hsp40) member A4 |
| chr5_+_16139909 | 1.36 |
ENSMUST00000196750.2
|
Cacna2d1
|
calcium channel, voltage-dependent, alpha2/delta subunit 1 |
| chr9_-_58943670 | 1.36 |
ENSMUST00000214547.2
ENSMUST00000068664.7 |
Neo1
|
neogenin |
| chr6_+_72332449 | 1.33 |
ENSMUST00000206064.2
|
Tmem150a
|
transmembrane protein 150A |
| chr2_+_24839758 | 1.31 |
ENSMUST00000028350.9
|
Zmynd19
|
zinc finger, MYND domain containing 19 |
| chr2_+_27567213 | 1.29 |
ENSMUST00000077257.12
|
Rxra
|
retinoid X receptor alpha |
| chr2_+_167922924 | 1.26 |
ENSMUST00000052125.7
|
Pard6b
|
par-6 family cell polarity regulator beta |
| chr7_+_65294630 | 1.24 |
ENSMUST00000032728.9
|
Tarsl2
|
threonyl-tRNA synthetase-like 2 |
| chr10_-_114638202 | 1.22 |
ENSMUST00000239411.2
|
Trhde
|
TRH-degrading enzyme |
| chr11_+_77821626 | 1.20 |
ENSMUST00000093995.10
ENSMUST00000000646.14 |
Sez6
|
seizure related gene 6 |
| chr19_-_5148506 | 1.19 |
ENSMUST00000025805.8
|
Cnih2
|
cornichon family AMPA receptor auxiliary protein 2 |
| chr2_-_162502994 | 1.19 |
ENSMUST00000109442.8
ENSMUST00000109445.9 ENSMUST00000109443.8 ENSMUST00000109441.2 |
Ptprt
|
protein tyrosine phosphatase, receptor type, T |
| chr2_-_73605684 | 1.19 |
ENSMUST00000112024.10
ENSMUST00000180045.8 |
Chn1
|
chimerin 1 |
| chr8_+_93084253 | 1.18 |
ENSMUST00000210246.2
ENSMUST00000034184.12 |
Irx5
|
Iroquois homeobox 5 |
| chr11_+_117545618 | 1.18 |
ENSMUST00000106344.8
|
Tnrc6c
|
trinucleotide repeat containing 6C |
| chr6_-_52168675 | 1.15 |
ENSMUST00000101395.3
|
Hoxa4
|
homeobox A4 |
| chr19_-_5474787 | 1.14 |
ENSMUST00000148219.9
|
Drap1
|
Dr1 associated protein 1 (negative cofactor 2 alpha) |
| chr9_-_43117052 | 1.14 |
ENSMUST00000176636.5
|
Pou2f3
|
POU domain, class 2, transcription factor 3 |
| chr5_+_88712840 | 1.14 |
ENSMUST00000196894.5
ENSMUST00000198965.5 |
Rufy3
|
RUN and FYVE domain containing 3 |
| chr4_+_13743424 | 1.13 |
ENSMUST00000006761.10
|
Runx1t1
|
RUNX1 translocation partner 1 |
| chr13_+_119565669 | 1.13 |
ENSMUST00000173627.8
ENSMUST00000126957.9 ENSMUST00000174691.8 |
Paip1
|
polyadenylate binding protein-interacting protein 1 |
| chr2_-_26096547 | 1.11 |
ENSMUST00000028302.8
|
Lhx3
|
LIM homeobox protein 3 |
| chr14_-_19057159 | 1.10 |
ENSMUST00000170123.2
|
Gm10409
|
predicted gene 10409 |
| chr2_+_90677499 | 1.09 |
ENSMUST00000136872.8
ENSMUST00000150232.8 ENSMUST00000111467.4 |
Mtch2
|
mitochondrial carrier 2 |
| chr5_+_32768515 | 1.09 |
ENSMUST00000202543.4
ENSMUST00000072311.13 |
Yes1
|
YES proto-oncogene 1, Src family tyrosine kinase |
| chr17_-_13271183 | 1.09 |
ENSMUST00000091648.4
|
Gpr31b
|
G protein-coupled receptor 31, D17Leh66b region |
| chr10_-_120815232 | 1.08 |
ENSMUST00000119944.8
ENSMUST00000119093.2 |
Lemd3
|
LEM domain containing 3 |
| chr16_+_65612394 | 1.07 |
ENSMUST00000227997.2
|
Vgll3
|
vestigial like family member 3 |
| chr4_+_152093260 | 1.07 |
ENSMUST00000097773.4
|
Klhl21
|
kelch-like 21 |
| chr2_-_151474391 | 1.05 |
ENSMUST00000137936.2
ENSMUST00000146172.8 ENSMUST00000094456.10 ENSMUST00000148755.8 ENSMUST00000109875.8 ENSMUST00000028951.14 ENSMUST00000109877.10 |
Snph
|
syntaphilin |
| chr4_+_84802650 | 1.04 |
ENSMUST00000169371.9
|
Cntln
|
centlein, centrosomal protein |
| chr5_+_141227245 | 1.04 |
ENSMUST00000085774.11
|
Sdk1
|
sidekick cell adhesion molecule 1 |
| chr14_-_16968099 | 1.03 |
ENSMUST00000181562.8
|
Gm3488
|
predicted gene, 3488 |
| chr9_+_106354463 | 1.03 |
ENSMUST00000047721.10
|
Rrp9
|
ribosomal RNA processing 9, U3 small nucleolar RNA binding protein |
| chr19_-_37340010 | 1.03 |
ENSMUST00000131070.3
|
Ide
|
insulin degrading enzyme |
| chr19_-_50666923 | 1.02 |
ENSMUST00000211687.2
ENSMUST00000211008.2 ENSMUST00000111756.4 ENSMUST00000209783.2 |
Sorcs1
|
sortilin-related VPS10 domain containing receptor 1 |
| chr17_-_29456750 | 1.02 |
ENSMUST00000137727.3
|
Cpne5
|
copine V |
| chr7_+_65511482 | 1.01 |
ENSMUST00000176199.8
|
Pcsk6
|
proprotein convertase subtilisin/kexin type 6 |
| chr13_+_112937320 | 1.00 |
ENSMUST00000016144.12
|
Plpp1
|
phospholipid phosphatase 1 |
| chr3_-_105594865 | 0.99 |
ENSMUST00000090680.11
|
Ddx20
|
DEAD box helicase 20 |
| chr8_+_84699580 | 0.99 |
ENSMUST00000005606.8
|
Prkaca
|
protein kinase, cAMP dependent, catalytic, alpha |
| chr18_+_82929037 | 0.99 |
ENSMUST00000236858.2
|
Zfp516
|
zinc finger protein 516 |
| chr12_+_112978051 | 0.99 |
ENSMUST00000223502.2
ENSMUST00000084891.5 ENSMUST00000220541.2 |
Pacs2
|
phosphofurin acidic cluster sorting protein 2 |
| chr8_+_80366247 | 0.99 |
ENSMUST00000173078.8
ENSMUST00000173286.8 |
Otud4
|
OTU domain containing 4 |
| chr14_+_15579811 | 0.98 |
ENSMUST00000171906.2
|
Gm3667
|
predicted gene 3667 |
| chr16_-_4238280 | 0.98 |
ENSMUST00000120080.8
|
Adcy9
|
adenylate cyclase 9 |
| chr6_-_30936013 | 0.97 |
ENSMUST00000101589.5
|
Klf14
|
Kruppel-like factor 14 |
| chr5_-_36641456 | 0.97 |
ENSMUST00000119916.2
ENSMUST00000031097.8 |
Tada2b
|
transcriptional adaptor 2B |
| chr14_-_31552608 | 0.97 |
ENSMUST00000014640.9
|
Ankrd28
|
ankyrin repeat domain 28 |
| chr14_-_24053994 | 0.96 |
ENSMUST00000225431.2
ENSMUST00000188210.8 ENSMUST00000224787.2 ENSMUST00000225315.2 ENSMUST00000225556.2 ENSMUST00000223727.2 ENSMUST00000223655.2 ENSMUST00000224077.2 ENSMUST00000224812.2 ENSMUST00000224285.2 ENSMUST00000225471.2 ENSMUST00000224232.2 ENSMUST00000223749.2 ENSMUST00000224025.2 |
Kcnma1
|
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr12_+_53294940 | 0.95 |
ENSMUST00000223358.3
|
Npas3
|
neuronal PAS domain protein 3 |
| chr14_-_24054273 | 0.95 |
ENSMUST00000188285.7
ENSMUST00000190044.7 |
Kcnma1
|
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr19_-_5474509 | 0.94 |
ENSMUST00000136579.3
ENSMUST00000113673.9 |
Drap1
|
Dr1 associated protein 1 (negative cofactor 2 alpha) |
| chr18_-_25886750 | 0.94 |
ENSMUST00000224553.2
ENSMUST00000025117.14 |
Celf4
|
CUGBP, Elav-like family member 4 |
| chr15_+_87428483 | 0.91 |
ENSMUST00000230414.2
|
Tafa5
|
TAFA chemokine like family member 5 |
| chr15_-_31367872 | 0.91 |
ENSMUST00000123325.9
|
Ankrd33b
|
ankyrin repeat domain 33B |
| chr14_-_24054186 | 0.91 |
ENSMUST00000188991.7
ENSMUST00000224468.2 |
Kcnma1
|
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr9_+_107277242 | 0.90 |
ENSMUST00000166799.8
ENSMUST00000170737.3 |
Cacna2d2
|
calcium channel, voltage-dependent, alpha 2/delta subunit 2 |
| chr17_+_87415049 | 0.90 |
ENSMUST00000041369.8
ENSMUST00000234803.2 |
Socs5
|
suppressor of cytokine signaling 5 |
| chr12_+_95658987 | 0.90 |
ENSMUST00000057324.4
|
Flrt2
|
fibronectin leucine rich transmembrane protein 2 |
| chr15_+_34837501 | 0.90 |
ENSMUST00000072868.5
|
Kcns2
|
K+ voltage-gated channel, subfamily S, 2 |
| chr17_-_66826661 | 0.89 |
ENSMUST00000167962.2
ENSMUST00000070538.12 |
Rab12
|
RAB12, member RAS oncogene family |
| chr5_+_100994230 | 0.88 |
ENSMUST00000092990.4
ENSMUST00000145612.2 |
Gpat3
|
glycerol-3-phosphate acyltransferase 3 |
| chr8_+_36924702 | 0.87 |
ENSMUST00000135373.8
ENSMUST00000147525.9 |
Trmt9b
|
tRNA methyltransferase 9B |
| chr2_+_30061469 | 0.87 |
ENSMUST00000015481.6
|
Endog
|
endonuclease G |
| chr12_+_13319110 | 0.87 |
ENSMUST00000042953.10
|
Nbas
|
neuroblastoma amplified sequence |
| chr10_+_95350975 | 0.86 |
ENSMUST00000099329.5
|
Ube2n
|
ubiquitin-conjugating enzyme E2N |
| chr5_-_76452365 | 0.86 |
ENSMUST00000075159.5
|
Clock
|
circadian locomotor output cycles kaput |
| chr11_-_72686853 | 0.86 |
ENSMUST00000156294.8
|
Cyb5d2
|
cytochrome b5 domain containing 2 |
| chr5_+_37399284 | 0.85 |
ENSMUST00000202434.4
ENSMUST00000114158.9 |
Crmp1
|
collapsin response mediator protein 1 |
| chr18_-_81029986 | 0.85 |
ENSMUST00000057950.9
|
Sall3
|
spalt like transcription factor 3 |
| chr7_-_37472290 | 0.84 |
ENSMUST00000176205.8
|
Zfp536
|
zinc finger protein 536 |
| chr4_+_59626185 | 0.84 |
ENSMUST00000070150.11
ENSMUST00000052420.7 |
E130308A19Rik
|
RIKEN cDNA E130308A19 gene |
| chr19_-_10434837 | 0.83 |
ENSMUST00000171400.4
|
Lrrc10b
|
leucine rich repeat containing 10B |
| chr14_+_15295240 | 0.82 |
ENSMUST00000172431.8
|
Gm3512
|
predicted gene 3512 |
| chr7_-_64806164 | 0.81 |
ENSMUST00000148459.3
ENSMUST00000119118.8 |
Fam189a1
|
family with sequence similarity 189, member A1 |
| chr19_-_34856853 | 0.80 |
ENSMUST00000036584.13
|
Pank1
|
pantothenate kinase 1 |
| chr6_+_72333209 | 0.80 |
ENSMUST00000206531.2
|
Tmem150a
|
transmembrane protein 150A |
| chr12_+_103354915 | 0.79 |
ENSMUST00000101094.9
ENSMUST00000021620.13 |
Otub2
|
OTU domain, ubiquitin aldehyde binding 2 |
| chr11_+_49500090 | 0.79 |
ENSMUST00000020617.3
|
Flt4
|
FMS-like tyrosine kinase 4 |
| chr2_+_27567246 | 0.79 |
ENSMUST00000166775.8
|
Rxra
|
retinoid X receptor alpha |
| chr13_-_30170031 | 0.78 |
ENSMUST00000102948.11
|
E2f3
|
E2F transcription factor 3 |
| chr5_-_72716942 | 0.78 |
ENSMUST00000074948.5
ENSMUST00000087216.12 |
Nfxl1
|
nuclear transcription factor, X-box binding-like 1 |
| chr4_+_84802592 | 0.78 |
ENSMUST00000102819.10
|
Cntln
|
centlein, centrosomal protein |
| chr12_+_84161095 | 0.78 |
ENSMUST00000123491.8
ENSMUST00000046340.9 ENSMUST00000136159.2 |
Dnal1
|
dynein, axonemal, light chain 1 |
| chr7_-_34914675 | 0.77 |
ENSMUST00000118444.3
ENSMUST00000122409.8 |
Lrp3
|
low density lipoprotein receptor-related protein 3 |
| chr6_-_113172340 | 0.77 |
ENSMUST00000162280.2
|
Lhfpl4
|
lipoma HMGIC fusion partner-like protein 4 |
| chr2_-_80411578 | 0.76 |
ENSMUST00000028386.12
|
Nckap1
|
NCK-associated protein 1 |
| chr11_+_11065782 | 0.76 |
ENSMUST00000056344.5
|
Vwc2
|
von Willebrand factor C domain containing 2 |
| chr14_-_18135010 | 0.75 |
ENSMUST00000165466.8
|
2610042L04Rik
|
RIKEN cDNA 2610042L04 gene |
| chr4_+_84802513 | 0.75 |
ENSMUST00000047023.13
|
Cntln
|
centlein, centrosomal protein |
| chr10_-_32765671 | 0.75 |
ENSMUST00000218645.2
|
Nkain2
|
Na+/K+ transporting ATPase interacting 2 |
| chr4_+_115642147 | 0.75 |
ENSMUST00000239030.2
|
Atpaf1
|
ATP synthase mitochondrial F1 complex assembly factor 1 |
| chr11_-_107685383 | 0.75 |
ENSMUST00000021066.4
|
Cacng4
|
calcium channel, voltage-dependent, gamma subunit 4 |
| chr5_-_100563974 | 0.74 |
ENSMUST00000182886.8
ENSMUST00000094578.11 |
Sec31a
|
Sec31 homolog A (S. cerevisiae) |
| chr7_+_40547608 | 0.74 |
ENSMUST00000044705.12
|
Vstm2b
|
V-set and transmembrane domain containing 2B |
| chr11_-_35689711 | 0.74 |
ENSMUST00000160726.4
|
Fbll1
|
fibrillarin-like 1 |
| chr19_-_50667079 | 0.74 |
ENSMUST00000209413.2
ENSMUST00000072685.13 ENSMUST00000164039.9 |
Sorcs1
|
sortilin-related VPS10 domain containing receptor 1 |
| chr5_-_123038329 | 0.74 |
ENSMUST00000031435.14
|
Kdm2b
|
lysine (K)-specific demethylase 2B |
| chr14_-_20844074 | 0.72 |
ENSMUST00000080440.14
ENSMUST00000100837.11 ENSMUST00000071816.7 |
Camk2g
|
calcium/calmodulin-dependent protein kinase II gamma |
| chr14_-_18897750 | 0.72 |
ENSMUST00000178728.2
|
Gm3005
|
predicted gene 3005 |
| chr11_+_59738866 | 0.72 |
ENSMUST00000102695.4
|
Nt5m
|
5',3'-nucleotidase, mitochondrial |
| chr6_+_107506678 | 0.70 |
ENSMUST00000049285.10
|
Lrrn1
|
leucine rich repeat protein 1, neuronal |
| chr4_+_141095415 | 0.70 |
ENSMUST00000006380.5
|
Fam131c
|
family with sequence similarity 131, member C |
| chr7_-_141055044 | 0.69 |
ENSMUST00000043870.9
|
Polr2l
|
polymerase (RNA) II (DNA directed) polypeptide L |
| chr7_+_141055135 | 0.69 |
ENSMUST00000026585.14
|
Tspan4
|
tetraspanin 4 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 11.2 | GO:0071630 | nucleus-associated proteasomal ubiquitin-dependent protein catabolic process(GO:0071630) |
| 1.4 | 4.3 | GO:0003167 | atrioventricular bundle cell differentiation(GO:0003167) |
| 1.1 | 3.2 | GO:1904456 | negative regulation of neuronal action potential(GO:1904456) |
| 1.0 | 3.9 | GO:0042339 | keratan sulfate metabolic process(GO:0042339) |
| 0.9 | 3.7 | GO:0015755 | fructose transport(GO:0015755) |
| 0.9 | 2.6 | GO:1904954 | canonical Wnt signaling pathway involved in midbrain dopaminergic neuron differentiation(GO:1904954) |
| 0.9 | 2.6 | GO:0021644 | vagus nerve morphogenesis(GO:0021644) |
| 0.8 | 3.2 | GO:0042851 | L-alanine metabolic process(GO:0042851) |
| 0.8 | 3.9 | GO:0086048 | membrane depolarization during bundle of His cell action potential(GO:0086048) |
| 0.8 | 4.7 | GO:0060082 | response to carbon monoxide(GO:0034465) eye blink reflex(GO:0060082) |
| 0.8 | 2.3 | GO:0030070 | insulin processing(GO:0030070) |
| 0.7 | 2.8 | GO:0080163 | regulation of protein serine/threonine phosphatase activity(GO:0080163) |
| 0.7 | 4.0 | GO:0039534 | negative regulation of MDA-5 signaling pathway(GO:0039534) |
| 0.6 | 2.9 | GO:0042414 | epinephrine metabolic process(GO:0042414) |
| 0.6 | 4.5 | GO:0021853 | cerebral cortex GABAergic interneuron migration(GO:0021853) interneuron migration(GO:1904936) |
| 0.5 | 2.2 | GO:0042138 | meiotic DNA double-strand break formation(GO:0042138) |
| 0.5 | 2.5 | GO:0035927 | RNA import into mitochondrion(GO:0035927) |
| 0.4 | 2.1 | GO:0045994 | regulation of translational initiation by iron(GO:0006447) positive regulation of translational initiation by iron(GO:0045994) |
| 0.4 | 2.5 | GO:0032902 | nerve growth factor production(GO:0032902) |
| 0.4 | 1.2 | GO:0006435 | threonyl-tRNA aminoacylation(GO:0006435) |
| 0.4 | 6.1 | GO:0090286 | cytoskeletal anchoring at nuclear membrane(GO:0090286) |
| 0.4 | 1.5 | GO:0097477 | lateral motor column neuron migration(GO:0097477) |
| 0.3 | 2.0 | GO:0046726 | positive regulation by virus of viral protein levels in host cell(GO:0046726) |
| 0.3 | 1.2 | GO:1902683 | regulation of receptor localization to synapse(GO:1902683) |
| 0.3 | 1.2 | GO:0033615 | mitochondrial proton-transporting ATP synthase complex assembly(GO:0033615) |
| 0.3 | 2.9 | GO:0006104 | succinyl-CoA metabolic process(GO:0006104) |
| 0.3 | 1.7 | GO:0006663 | platelet activating factor biosynthetic process(GO:0006663) |
| 0.3 | 1.9 | GO:0003219 | cardiac right ventricle formation(GO:0003219) |
| 0.3 | 1.1 | GO:0021526 | medial motor column neuron differentiation(GO:0021526) |
| 0.3 | 14.0 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.3 | 2.1 | GO:0007199 | G-protein coupled receptor signaling pathway coupled to cGMP nucleotide second messenger(GO:0007199) |
| 0.3 | 1.5 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.2 | 0.7 | GO:0034963 | box C/D snoRNA 3'-end processing(GO:0000494) box C/D snoRNA metabolic process(GO:0033967) box C/D snoRNA processing(GO:0034963) histone glutamine methylation(GO:1990258) |
| 0.2 | 1.9 | GO:1902966 | regulation of protein localization to early endosome(GO:1902965) positive regulation of protein localization to early endosome(GO:1902966) |
| 0.2 | 3.6 | GO:0070166 | enamel mineralization(GO:0070166) |
| 0.2 | 0.8 | GO:1904616 | regulation of actin filament binding(GO:1904529) regulation of actin binding(GO:1904616) |
| 0.2 | 1.3 | GO:0016185 | synaptic vesicle budding from presynaptic endocytic zone membrane(GO:0016185) |
| 0.2 | 1.8 | GO:0035902 | response to immobilization stress(GO:0035902) |
| 0.2 | 1.7 | GO:0046959 | habituation(GO:0046959) |
| 0.2 | 0.7 | GO:0021993 | initiation of neural tube closure(GO:0021993) |
| 0.2 | 0.5 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.2 | 0.7 | GO:0006244 | pyrimidine nucleotide catabolic process(GO:0006244) pyrimidine deoxyribonucleotide catabolic process(GO:0009223) |
| 0.2 | 0.2 | GO:0060423 | foregut regionalization(GO:0060423) lung field specification(GO:0060424) lung induction(GO:0060492) |
| 0.2 | 0.5 | GO:0016561 | protein import into peroxisome matrix, translocation(GO:0016561) |
| 0.2 | 0.8 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.2 | 1.6 | GO:0034398 | telomere tethering at nuclear periphery(GO:0034398) |
| 0.2 | 0.7 | GO:0015766 | disaccharide transport(GO:0015766) sucrose transport(GO:0015770) oligosaccharide transport(GO:0015772) |
| 0.2 | 0.7 | GO:0060849 | regulation of transcription involved in lymphatic endothelial cell fate commitment(GO:0060849) |
| 0.2 | 0.5 | GO:1901994 | negative regulation of meiotic cell cycle phase transition(GO:1901994) |
| 0.2 | 0.8 | GO:0070602 | regulation of centromeric sister chromatid cohesion(GO:0070602) |
| 0.1 | 0.4 | GO:0002654 | positive regulation of tolerance induction dependent upon immune response(GO:0002654) regulation of peripheral tolerance induction(GO:0002658) positive regulation of peripheral tolerance induction(GO:0002660) regulation of peripheral T cell tolerance induction(GO:0002849) positive regulation of peripheral T cell tolerance induction(GO:0002851) |
| 0.1 | 0.9 | GO:1902512 | positive regulation of apoptotic DNA fragmentation(GO:1902512) |
| 0.1 | 2.6 | GO:0010457 | centriole-centriole cohesion(GO:0010457) |
| 0.1 | 1.0 | GO:0071880 | adenylate cyclase-activating adrenergic receptor signaling pathway(GO:0071880) |
| 0.1 | 0.5 | GO:2000384 | regulation of ectoderm development(GO:2000383) negative regulation of ectoderm development(GO:2000384) |
| 0.1 | 0.5 | GO:0097168 | condensed mesenchymal cell proliferation(GO:0072137) mesenchymal stem cell proliferation(GO:0097168) |
| 0.1 | 0.9 | GO:0060024 | rhythmic synaptic transmission(GO:0060024) |
| 0.1 | 0.9 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 0.1 | 0.9 | GO:2000622 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.1 | 1.6 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.1 | 2.3 | GO:0002031 | G-protein coupled receptor internalization(GO:0002031) |
| 0.1 | 4.1 | GO:0046854 | phosphatidylinositol phosphorylation(GO:0046854) |
| 0.1 | 1.0 | GO:0010815 | bradykinin catabolic process(GO:0010815) |
| 0.1 | 1.7 | GO:0090084 | negative regulation of inclusion body assembly(GO:0090084) |
| 0.1 | 1.7 | GO:1990403 | embryonic brain development(GO:1990403) |
| 0.1 | 0.6 | GO:0035795 | negative regulation of mitochondrial membrane permeability(GO:0035795) |
| 0.1 | 1.1 | GO:0035701 | hematopoietic stem cell migration(GO:0035701) hematopoietic stem cell homeostasis(GO:0061484) |
| 0.1 | 0.7 | GO:2000819 | regulation of nucleotide-excision repair(GO:2000819) |
| 0.1 | 0.8 | GO:0030950 | establishment or maintenance of actin cytoskeleton polarity(GO:0030950) |
| 0.1 | 1.0 | GO:0090520 | sphingolipid mediated signaling pathway(GO:0090520) |
| 0.1 | 0.4 | GO:0098582 | innate vocalization behavior(GO:0098582) |
| 0.1 | 0.9 | GO:2000323 | negative regulation of glucocorticoid receptor signaling pathway(GO:2000323) |
| 0.1 | 3.9 | GO:0050901 | leukocyte tethering or rolling(GO:0050901) |
| 0.1 | 1.0 | GO:0090170 | regulation of Golgi inheritance(GO:0090170) |
| 0.1 | 1.7 | GO:0035278 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.1 | 1.1 | GO:0043922 | negative regulation by host of viral transcription(GO:0043922) |
| 0.1 | 0.8 | GO:1904378 | maintenance of unfolded protein(GO:0036506) maintenance of unfolded protein involved in ERAD pathway(GO:1904378) |
| 0.1 | 0.5 | GO:0014870 | response to inactivity(GO:0014854) response to muscle inactivity(GO:0014870) response to muscle inactivity involved in regulation of muscle adaptation(GO:0014877) response to denervation involved in regulation of muscle adaptation(GO:0014894) |
| 0.1 | 0.3 | GO:0072194 | kidney smooth muscle tissue development(GO:0072194) |
| 0.1 | 1.2 | GO:0060040 | retinal bipolar neuron differentiation(GO:0060040) |
| 0.1 | 2.6 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.1 | 0.2 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 0.1 | 0.4 | GO:1902303 | negative regulation of potassium ion export(GO:1902303) |
| 0.1 | 0.3 | GO:0016240 | autophagosome docking(GO:0016240) |
| 0.1 | 0.3 | GO:0035938 | estradiol secretion(GO:0035938) regulation of estradiol secretion(GO:2000864) |
| 0.1 | 1.6 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.1 | 1.4 | GO:0035092 | sperm chromatin condensation(GO:0035092) |
| 0.1 | 0.4 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.1 | 0.9 | GO:1902866 | regulation of retina development in camera-type eye(GO:1902866) |
| 0.1 | 0.6 | GO:0002432 | granuloma formation(GO:0002432) |
| 0.1 | 1.0 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.1 | 0.1 | GO:2001055 | positive regulation of mesenchymal cell apoptotic process(GO:2001055) |
| 0.1 | 0.1 | GO:0060137 | maternal process involved in parturition(GO:0060137) |
| 0.1 | 0.4 | GO:0021615 | glossopharyngeal nerve morphogenesis(GO:0021615) |
| 0.1 | 1.4 | GO:0036120 | cellular response to platelet-derived growth factor stimulus(GO:0036120) |
| 0.1 | 1.7 | GO:0007614 | short-term memory(GO:0007614) |
| 0.1 | 0.8 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.1 | 0.8 | GO:0021891 | olfactory bulb interneuron development(GO:0021891) |
| 0.1 | 0.9 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.1 | 1.5 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.1 | 0.6 | GO:0060449 | bud elongation involved in lung branching(GO:0060449) |
| 0.1 | 0.8 | GO:0035871 | protein K11-linked deubiquitination(GO:0035871) |
| 0.1 | 0.8 | GO:0090037 | positive regulation of protein kinase C signaling(GO:0090037) |
| 0.1 | 0.5 | GO:0019348 | dolichol metabolic process(GO:0019348) |
| 0.1 | 3.8 | GO:0009409 | response to cold(GO:0009409) |
| 0.1 | 1.2 | GO:0042711 | maternal behavior(GO:0042711) |
| 0.1 | 1.0 | GO:0034497 | protein localization to pre-autophagosomal structure(GO:0034497) |
| 0.1 | 0.8 | GO:0060025 | regulation of synaptic activity(GO:0060025) |
| 0.1 | 1.5 | GO:0040037 | negative regulation of fibroblast growth factor receptor signaling pathway(GO:0040037) diaphragm development(GO:0060539) |
| 0.1 | 0.4 | GO:0045200 | establishment or maintenance of neuroblast polarity(GO:0045196) establishment of neuroblast polarity(GO:0045200) |
| 0.1 | 1.2 | GO:0090036 | regulation of protein kinase C signaling(GO:0090036) |
| 0.0 | 9.8 | GO:0007286 | spermatid development(GO:0007286) |
| 0.0 | 0.5 | GO:0097623 | potassium ion export across plasma membrane(GO:0097623) |
| 0.0 | 0.5 | GO:0002002 | regulation of angiotensin levels in blood(GO:0002002) |
| 0.0 | 0.5 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.0 | 0.2 | GO:0021564 | vagus nerve development(GO:0021564) |
| 0.0 | 0.8 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.0 | 1.4 | GO:1902259 | regulation of delayed rectifier potassium channel activity(GO:1902259) |
| 0.0 | 0.7 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.0 | 1.0 | GO:0048148 | behavioral response to cocaine(GO:0048148) |
| 0.0 | 0.9 | GO:0000729 | DNA double-strand break processing(GO:0000729) |
| 0.0 | 0.2 | GO:0045039 | protein import into mitochondrial inner membrane(GO:0045039) |
| 0.0 | 1.1 | GO:2000114 | regulation of establishment of cell polarity(GO:2000114) |
| 0.0 | 0.1 | GO:0046462 | monoacylglycerol metabolic process(GO:0046462) |
| 0.0 | 1.2 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
| 0.0 | 1.2 | GO:0043171 | peptide catabolic process(GO:0043171) |
| 0.0 | 0.8 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.0 | 1.0 | GO:1903861 | positive regulation of dendrite extension(GO:1903861) |
| 0.0 | 0.2 | GO:1902498 | regulation of protein autoubiquitination(GO:1902498) |
| 0.0 | 1.9 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.0 | 0.3 | GO:0045003 | DNA recombinase assembly(GO:0000730) double-strand break repair via synthesis-dependent strand annealing(GO:0045003) |
| 0.0 | 0.4 | GO:2000465 | regulation of glycogen (starch) synthase activity(GO:2000465) |
| 0.0 | 0.7 | GO:1901897 | regulation of relaxation of cardiac muscle(GO:1901897) |
| 0.0 | 1.1 | GO:0034629 | cellular protein complex localization(GO:0034629) |
| 0.0 | 1.2 | GO:0046460 | neutral lipid biosynthetic process(GO:0046460) acylglycerol biosynthetic process(GO:0046463) |
| 0.0 | 0.0 | GO:0018307 | enzyme active site formation(GO:0018307) |
| 0.0 | 0.3 | GO:0090166 | Golgi disassembly(GO:0090166) |
| 0.0 | 0.3 | GO:0090336 | positive regulation of brown fat cell differentiation(GO:0090336) |
| 0.0 | 1.5 | GO:0007212 | dopamine receptor signaling pathway(GO:0007212) |
| 0.0 | 0.1 | GO:0023016 | signal transduction by trans-phosphorylation(GO:0023016) |
| 0.0 | 1.1 | GO:0045599 | negative regulation of fat cell differentiation(GO:0045599) |
| 0.0 | 0.7 | GO:0045723 | positive regulation of fatty acid biosynthetic process(GO:0045723) |
| 0.0 | 0.3 | GO:0035879 | lactate transport(GO:0015727) lactate transmembrane transport(GO:0035873) plasma membrane lactate transport(GO:0035879) |
| 0.0 | 1.0 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 0.3 | GO:0042249 | establishment of planar polarity of embryonic epithelium(GO:0042249) |
| 0.0 | 0.7 | GO:0071711 | basement membrane organization(GO:0071711) |
| 0.0 | 0.7 | GO:0035115 | embryonic forelimb morphogenesis(GO:0035115) |
| 0.0 | 0.3 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.0 | 0.5 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.0 | 0.0 | GO:0061762 | CAMKK-AMPK signaling cascade(GO:0061762) |
| 0.0 | 3.9 | GO:0042787 | protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0042787) |
| 0.0 | 0.9 | GO:0019933 | cAMP-mediated signaling(GO:0019933) |
| 0.0 | 0.9 | GO:0008333 | endosome to lysosome transport(GO:0008333) |
| 0.0 | 0.5 | GO:0070534 | protein K63-linked ubiquitination(GO:0070534) |
| 0.0 | 0.2 | GO:1900194 | negative regulation of oocyte maturation(GO:1900194) |
| 0.0 | 0.8 | GO:0030514 | negative regulation of BMP signaling pathway(GO:0030514) |
| 0.0 | 0.5 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.0 | 0.4 | GO:1902230 | negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902230) |
| 0.0 | 0.1 | GO:0015820 | branched-chain amino acid transport(GO:0015803) leucine transport(GO:0015820) |
| 0.0 | 0.1 | GO:0035106 | operant conditioning(GO:0035106) |
| 0.0 | 0.4 | GO:0007184 | SMAD protein import into nucleus(GO:0007184) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.7 | GO:0097123 | cyclin A1-CDK2 complex(GO:0097123) |
| 0.4 | 2.2 | GO:0038039 | G-protein coupled receptor heterodimeric complex(GO:0038039) |
| 0.4 | 1.3 | GO:0098830 | presynaptic endosome(GO:0098830) |
| 0.4 | 7.3 | GO:0031045 | dense core granule(GO:0031045) |
| 0.3 | 0.9 | GO:0070939 | Dsl1p complex(GO:0070939) |
| 0.3 | 0.9 | GO:0031372 | UBC13-MMS2 complex(GO:0031372) |
| 0.3 | 4.7 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.3 | 3.9 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.2 | 0.6 | GO:0036488 | CHOP-C/EBP complex(GO:0036488) |
| 0.2 | 2.5 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.2 | 1.8 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.2 | 9.0 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.2 | 1.5 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.2 | 1.6 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.1 | 0.6 | GO:0071149 | TEAD-1-YAP complex(GO:0071148) TEAD-2-YAP complex(GO:0071149) |
| 0.1 | 13.0 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 0.9 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.1 | 0.4 | GO:0097632 | extrinsic component of pre-autophagosomal structure membrane(GO:0097632) |
| 0.1 | 1.8 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.1 | 0.5 | GO:0033185 | dolichol-phosphate-mannose synthase complex(GO:0033185) |
| 0.1 | 0.6 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.1 | 0.4 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.1 | 0.8 | GO:0072379 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.1 | 1.0 | GO:0090571 | RNA polymerase II transcription repressor complex(GO:0090571) |
| 0.1 | 2.3 | GO:0043196 | varicosity(GO:0043196) |
| 0.1 | 1.0 | GO:0031597 | cytosolic proteasome complex(GO:0031597) |
| 0.1 | 0.8 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.1 | 0.5 | GO:0030689 | Noc complex(GO:0030689) |
| 0.1 | 1.0 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.1 | 1.1 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.1 | 0.8 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.0 | 0.7 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 0.7 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.0 | 1.1 | GO:0071437 | invadopodium(GO:0071437) |
| 0.0 | 1.1 | GO:0051286 | cell tip(GO:0051286) |
| 0.0 | 1.0 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.0 | 3.2 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 2.0 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 2.7 | GO:0008287 | protein serine/threonine phosphatase complex(GO:0008287) phosphatase complex(GO:1903293) |
| 0.0 | 3.7 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 0.2 | GO:0072487 | MSL complex(GO:0072487) |
| 0.0 | 0.3 | GO:0033061 | DNA recombinase mediator complex(GO:0033061) |
| 0.0 | 2.0 | GO:0005844 | polysome(GO:0005844) |
| 0.0 | 0.7 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.0 | 0.4 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.0 | 0.4 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.0 | 5.9 | GO:0031514 | motile cilium(GO:0031514) |
| 0.0 | 0.9 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 1.0 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 0.5 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 0.3 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.0 | 0.6 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.0 | 0.9 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.0 | 0.5 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 2.3 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 3.8 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.7 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 4.5 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 0.4 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 0.2 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.0 | 2.4 | GO:0043204 | perikaryon(GO:0043204) |
| 0.0 | 2.3 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.0 | 0.5 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.0 | 0.7 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.0 | 0.8 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.0 | 0.5 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.0 | 1.0 | GO:0031903 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 3.2 | GO:0000151 | ubiquitin ligase complex(GO:0000151) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 4.0 | GO:0034012 | glycerone kinase activity(GO:0004371) FAD-AMP lyase (cyclizing) activity(GO:0034012) triokinase activity(GO:0050354) |
| 1.3 | 3.9 | GO:0086057 | voltage-gated calcium channel activity involved in bundle of His cell action potential(GO:0086057) |
| 1.2 | 3.7 | GO:0005353 | fructose transmembrane transporter activity(GO:0005353) uniporter activity(GO:0015292) |
| 1.0 | 2.9 | GO:0004774 | succinate-CoA ligase activity(GO:0004774) |
| 0.8 | 2.5 | GO:0030943 | mitochondrion targeting sequence binding(GO:0030943) |
| 0.6 | 3.2 | GO:0047635 | L-alanine:2-oxoglutarate aminotransferase activity(GO:0004021) alanine-oxo-acid transaminase activity(GO:0047635) |
| 0.6 | 4.7 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.6 | 7.8 | GO:0004000 | adenosine deaminase activity(GO:0004000) |
| 0.6 | 3.9 | GO:0001517 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) |
| 0.5 | 2.2 | GO:0003918 | DNA topoisomerase type II (ATP-hydrolyzing) activity(GO:0003918) DNA topoisomerase II activity(GO:0061505) |
| 0.5 | 1.5 | GO:0001571 | non-tyrosine kinase fibroblast growth factor receptor activity(GO:0001571) |
| 0.4 | 1.2 | GO:0004829 | threonine-tRNA ligase activity(GO:0004829) |
| 0.4 | 2.2 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.3 | 2.8 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.3 | 1.7 | GO:2001070 | starch binding(GO:2001070) |
| 0.3 | 1.7 | GO:0004142 | diacylglycerol cholinephosphotransferase activity(GO:0004142) |
| 0.3 | 4.8 | GO:0043495 | protein anchor(GO:0043495) |
| 0.3 | 1.9 | GO:0004703 | G-protein coupled receptor kinase activity(GO:0004703) |
| 0.3 | 2.1 | GO:0070644 | vitamin D response element binding(GO:0070644) |
| 0.2 | 0.7 | GO:0036009 | protein-glutamine N-methyltransferase activity(GO:0036009) histone-glutamine methyltransferase activity(GO:1990259) |
| 0.2 | 1.5 | GO:0070326 | very-low-density lipoprotein particle receptor binding(GO:0070326) |
| 0.2 | 2.3 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.2 | 0.8 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
| 0.2 | 0.7 | GO:0015157 | sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.2 | 0.5 | GO:0016419 | [acyl-carrier-protein] S-malonyltransferase activity(GO:0004314) S-malonyltransferase activity(GO:0016419) malonyltransferase activity(GO:0016420) |
| 0.2 | 2.6 | GO:0019855 | calcium channel inhibitor activity(GO:0019855) |
| 0.1 | 1.2 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.1 | 1.7 | GO:0001162 | RNA polymerase II intronic transcription regulatory region sequence-specific DNA binding(GO:0001162) |
| 0.1 | 1.5 | GO:0048101 | calmodulin-dependent cyclic-nucleotide phosphodiesterase activity(GO:0004117) calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.1 | 2.5 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.1 | 1.0 | GO:0019784 | NEDD8-specific protease activity(GO:0019784) |
| 0.1 | 1.2 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.1 | 1.4 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.1 | 2.9 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.1 | 2.8 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.1 | 0.5 | GO:0004582 | dolichyl-phosphate beta-D-mannosyltransferase activity(GO:0004582) |
| 0.1 | 1.0 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.1 | 1.4 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.1 | 0.8 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.1 | 0.4 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.1 | 0.4 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.1 | 1.0 | GO:0043559 | insulin binding(GO:0043559) |
| 0.1 | 0.5 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.1 | 1.0 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.1 | 0.5 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.1 | 1.2 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.1 | 0.3 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.1 | 5.0 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.1 | 0.8 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.1 | 0.4 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.1 | 0.6 | GO:0019158 | fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.1 | 0.8 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.1 | 3.6 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.1 | 1.1 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.1 | 0.5 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.1 | 1.5 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.1 | 1.8 | GO:0000217 | DNA secondary structure binding(GO:0000217) |
| 0.1 | 1.4 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.1 | 0.5 | GO:0001025 | RNA polymerase III transcription factor binding(GO:0001025) |
| 0.1 | 1.8 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 0.7 | GO:0001055 | RNA polymerase I activity(GO:0001054) RNA polymerase II activity(GO:0001055) |
| 0.1 | 0.5 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 0.1 | 2.0 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.1 | 0.5 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.0 | 0.4 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.0 | 2.0 | GO:0045182 | translation regulator activity(GO:0045182) |
| 0.0 | 1.2 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.0 | 0.4 | GO:1902282 | voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization(GO:1902282) |
| 0.0 | 1.6 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.8 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.0 | 0.8 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.0 | 3.6 | GO:0005249 | voltage-gated potassium channel activity(GO:0005249) |
| 0.0 | 1.0 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 10.4 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 2.9 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.0 | 1.0 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 0.3 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.0 | 0.9 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.0 | 2.4 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.0 | 0.3 | GO:0061676 | importin-alpha family protein binding(GO:0061676) |
| 0.0 | 0.1 | GO:0019870 | potassium channel inhibitor activity(GO:0019870) |
| 0.0 | 0.3 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 3.0 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.0 | 0.7 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.0 | 1.8 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.0 | 1.4 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 0.6 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.0 | 0.4 | GO:0016500 | protein-hormone receptor activity(GO:0016500) |
| 0.0 | 11.7 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 1.2 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 3.7 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.0 | 0.1 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.0 | 0.6 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.8 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 2.0 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.0 | 0.5 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.0 | 1.9 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.0 | 0.9 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.0 | 0.3 | GO:0003756 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.0 | 0.4 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 2.0 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 2.0 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.3 | GO:0031005 | filamin binding(GO:0031005) |
| 0.0 | 0.5 | GO:0001190 | transcriptional activator activity, RNA polymerase II transcription factor binding(GO:0001190) transcriptional repressor activity, RNA polymerase II activating transcription factor binding(GO:0098811) |
| 0.0 | 0.8 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 2.7 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 0.9 | GO:0004536 | deoxyribonuclease activity(GO:0004536) |
| 0.0 | 1.0 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.6 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.1 | 0.8 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.1 | 1.6 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.1 | 2.1 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.9 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 0.9 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 0.0 | 1.0 | SA TRKA RECEPTOR | The TrkA receptor binds nerve growth factor to activate MAP kinase pathways and promote cell growth. |
| 0.0 | 2.4 | ST GA13 PATHWAY | G alpha 13 Pathway |
| 0.0 | 0.9 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.0 | 1.4 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.0 | 0.7 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.0 | 1.5 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 1.9 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.0 | 1.9 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 2.2 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.5 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.0 | 0.8 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.0 | 0.4 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.0 | 0.6 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.0 | 0.3 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.0 | 0.6 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 4.0 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.2 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.0 | 0.4 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.7 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.1 | 2.7 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.1 | 2.9 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.1 | 2.9 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.1 | 3.2 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.1 | 3.9 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.1 | 2.9 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.1 | 2.2 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.1 | 2.3 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.1 | 0.8 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.1 | 4.4 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 2.1 | REACTOME REGULATORY RNA PATHWAYS | Genes involved in Regulatory RNA pathways |
| 0.1 | 1.1 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.1 | 1.8 | REACTOME PKA MEDIATED PHOSPHORYLATION OF CREB | Genes involved in PKA-mediated phosphorylation of CREB |
| 0.1 | 2.0 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.1 | 0.7 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.1 | 1.4 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.0 | 1.7 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 0.8 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.0 | 1.4 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 1.8 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.0 | 0.7 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.0 | 0.7 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.0 | 0.6 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 0.7 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.0 | 0.6 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.4 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.0 | 0.9 | REACTOME DOWNSTREAM TCR SIGNALING | Genes involved in Downstream TCR signaling |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.0 | 1.4 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.5 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.0 | 1.8 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.3 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.0 | 1.0 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.0 | 0.6 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.5 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |