Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
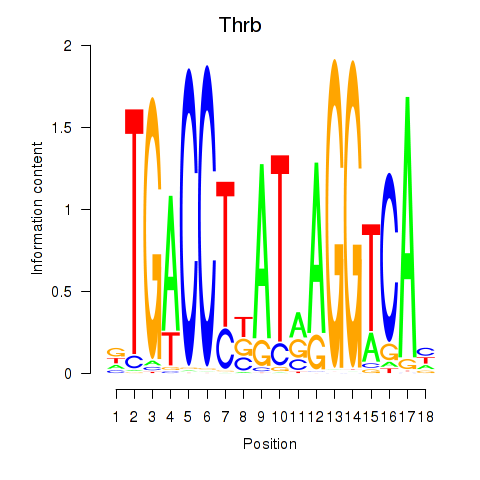
Results for Thrb
Z-value: 1.24
Transcription factors associated with Thrb
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Thrb
|
ENSMUSG00000021779.20 | Thrb |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Thrb | mm39_v1_chr14_-_4506874_4506894 | 0.14 | 2.5e-01 | Click! |
Activity profile of Thrb motif
Sorted Z-values of Thrb motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Thrb
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_+_105775224 | 21.08 |
ENSMUST00000093222.13
ENSMUST00000093223.5 |
Ces3a
|
carboxylesterase 3A |
| chr7_-_105249308 | 20.62 |
ENSMUST00000210531.2
ENSMUST00000033185.10 |
Hpx
|
hemopexin |
| chr2_+_58457370 | 17.51 |
ENSMUST00000071543.12
|
Upp2
|
uridine phosphorylase 2 |
| chr10_+_87357782 | 17.02 |
ENSMUST00000219813.2
|
Pah
|
phenylalanine hydroxylase |
| chr10_+_87357657 | 14.51 |
ENSMUST00000020241.17
|
Pah
|
phenylalanine hydroxylase |
| chr3_+_14928561 | 14.19 |
ENSMUST00000029076.6
|
Car3
|
carbonic anhydrase 3 |
| chr2_+_118998235 | 12.20 |
ENSMUST00000057454.4
|
Gchfr
|
GTP cyclohydrolase I feedback regulator |
| chr9_-_107546166 | 11.17 |
ENSMUST00000177567.8
|
Slc38a3
|
solute carrier family 38, member 3 |
| chr2_-_25359752 | 11.09 |
ENSMUST00000114259.3
ENSMUST00000015234.13 |
Ptgds
|
prostaglandin D2 synthase (brain) |
| chr9_-_107546195 | 10.55 |
ENSMUST00000192990.6
|
Slc38a3
|
solute carrier family 38, member 3 |
| chr2_-_25360043 | 9.33 |
ENSMUST00000114251.8
|
Ptgds
|
prostaglandin D2 synthase (brain) |
| chr12_+_108300599 | 7.99 |
ENSMUST00000021684.6
|
Cyp46a1
|
cytochrome P450, family 46, subfamily a, polypeptide 1 |
| chr2_+_92213256 | 6.94 |
ENSMUST00000054316.9
ENSMUST00000111280.3 |
1700029I15Rik
|
RIKEN cDNA 1700029I15 gene |
| chr16_-_28571820 | 6.70 |
ENSMUST00000232352.2
|
Fgf12
|
fibroblast growth factor 12 |
| chr1_-_83385911 | 6.16 |
ENSMUST00000160953.8
|
Sphkap
|
SPHK1 interactor, AKAP domain containing |
| chr17_-_28908757 | 6.12 |
ENSMUST00000233398.2
|
Slc26a8
|
solute carrier family 26, member 8 |
| chr13_-_59892731 | 6.03 |
ENSMUST00000180139.3
|
1700014D04Rik
|
RIKEN cDNA 1700014D04 gene |
| chr14_-_61597843 | 4.19 |
ENSMUST00000022494.10
|
Ebpl
|
emopamil binding protein-like |
| chr14_+_14475188 | 4.14 |
ENSMUST00000026315.8
|
Dnase1l3
|
deoxyribonuclease 1-like 3 |
| chr17_-_14223966 | 4.08 |
ENSMUST00000189454.3
|
Gm7356
|
predicted gene 7356 |
| chr16_+_8498618 | 3.93 |
ENSMUST00000201722.2
|
Gm5767
|
predicted gene 5767 |
| chr13_+_34923589 | 3.90 |
ENSMUST00000221037.2
|
Fam50b
|
family with sequence similarity 50, member B |
| chr11_+_46295547 | 3.90 |
ENSMUST00000063166.6
|
Fam71b
|
family with sequence similarity 71, member B |
| chr15_-_76193955 | 3.89 |
ENSMUST00000210024.2
|
Oplah
|
5-oxoprolinase (ATP-hydrolysing) |
| chr7_+_27222678 | 3.52 |
ENSMUST00000108353.9
|
Hipk4
|
homeodomain interacting protein kinase 4 |
| chr7_-_24705320 | 3.44 |
ENSMUST00000102858.10
ENSMUST00000196684.2 ENSMUST00000080882.11 |
Atp1a3
|
ATPase, Na+/K+ transporting, alpha 3 polypeptide |
| chr11_-_113600346 | 3.33 |
ENSMUST00000173655.8
ENSMUST00000100248.6 |
Cpsf4l
|
cleavage and polyadenylation specific factor 4-like |
| chr2_+_152804405 | 3.06 |
ENSMUST00000099197.9
ENSMUST00000103155.10 |
Ttll9
|
tubulin tyrosine ligase-like family, member 9 |
| chr11_-_113600838 | 2.56 |
ENSMUST00000018871.8
|
Cpsf4l
|
cleavage and polyadenylation specific factor 4-like |
| chr11_+_109376432 | 2.54 |
ENSMUST00000106697.8
|
Arsg
|
arylsulfatase G |
| chr7_-_33065747 | 2.51 |
ENSMUST00000179248.2
|
Scgb2b20
|
secretoglobin, family 2B, member 20 |
| chr10_-_18662526 | 2.17 |
ENSMUST00000216654.2
|
Gm4922
|
predicted gene 4922 |
| chr5_+_130248547 | 2.15 |
ENSMUST00000202305.4
ENSMUST00000065329.13 ENSMUST00000200802.4 |
Tmem248
|
transmembrane protein 248 |
| chr19_-_46314945 | 2.10 |
ENSMUST00000225781.2
ENSMUST00000223903.2 |
Psd
|
pleckstrin and Sec7 domain containing |
| chr11_-_83412952 | 2.09 |
ENSMUST00000019069.4
|
Heatr9
|
HEAT repeat containing 9 |
| chr6_+_114435480 | 2.02 |
ENSMUST00000160780.2
|
Hrh1
|
histamine receptor H1 |
| chr8_+_112262729 | 1.94 |
ENSMUST00000172856.8
|
Znrf1
|
zinc and ring finger 1 |
| chr4_+_130519788 | 1.86 |
ENSMUST00000070478.4
|
Sdc3
|
syndecan 3 |
| chr5_+_35971697 | 1.85 |
ENSMUST00000130233.8
|
Ablim2
|
actin-binding LIM protein 2 |
| chr5_-_30619246 | 1.84 |
ENSMUST00000114747.9
ENSMUST00000074171.10 |
Otof
|
otoferlin |
| chr3_-_79749949 | 1.80 |
ENSMUST00000029568.7
|
Tmem144
|
transmembrane protein 144 |
| chr16_-_16962279 | 1.58 |
ENSMUST00000232033.2
ENSMUST00000231597.2 ENSMUST00000232540.2 |
Ccdc116
|
coiled-coil domain containing 116 |
| chr6_-_121450547 | 1.56 |
ENSMUST00000046373.8
ENSMUST00000151397.3 |
Iqsec3
|
IQ motif and Sec7 domain 3 |
| chr9_-_102503180 | 1.51 |
ENSMUST00000216281.2
ENSMUST00000093791.10 |
Cep63
|
centrosomal protein 63 |
| chr16_-_16962256 | 1.49 |
ENSMUST00000115711.10
|
Ccdc116
|
coiled-coil domain containing 116 |
| chr11_+_70350436 | 1.44 |
ENSMUST00000039093.10
|
Zmynd15
|
zinc finger, MYND-type containing 15 |
| chr14_+_54713703 | 1.28 |
ENSMUST00000164697.8
|
Rem2
|
rad and gem related GTP binding protein 2 |
| chr14_+_53315909 | 1.11 |
ENSMUST00000103608.4
|
Trav14d-3-dv8
|
T cell receptor alpha variable 14D-3-DV8 |
| chr9_-_117843228 | 1.11 |
ENSMUST00000187803.3
|
Zcwpw2
|
zinc finger, CW type with PWWP domain 2 |
| chr11_+_70350252 | 1.07 |
ENSMUST00000108563.9
|
Zmynd15
|
zinc finger, MYND-type containing 15 |
| chr3_+_67281424 | 1.03 |
ENSMUST00000077916.12
|
Mlf1
|
myeloid leukemia factor 1 |
| chr14_+_54713557 | 0.99 |
ENSMUST00000164766.8
|
Rem2
|
rad and gem related GTP binding protein 2 |
| chr9_-_102503708 | 0.88 |
ENSMUST00000161645.8
ENSMUST00000162297.2 ENSMUST00000213636.2 ENSMUST00000162655.9 |
Cep63
|
centrosomal protein 63 |
| chr16_-_58930996 | 0.85 |
ENSMUST00000076262.4
|
Olfr193
|
olfactory receptor 193 |
| chr3_+_67281449 | 0.84 |
ENSMUST00000061322.10
|
Mlf1
|
myeloid leukemia factor 1 |
| chr5_+_95021656 | 0.82 |
ENSMUST00000180076.2
|
Gm3183
|
predicted gene 3183 |
| chr9_+_95739650 | 0.79 |
ENSMUST00000034980.9
|
Atr
|
ataxia telangiectasia and Rad3 related |
| chr11_+_24030663 | 0.76 |
ENSMUST00000118955.2
|
Bcl11a
|
B cell CLL/lymphoma 11A (zinc finger protein) |
| chr11_+_49094292 | 0.73 |
ENSMUST00000150284.8
ENSMUST00000109197.8 ENSMUST00000151228.2 |
Zfp62
|
zinc finger protein 62 |
| chr1_-_24626492 | 0.58 |
ENSMUST00000051344.6
ENSMUST00000115244.9 |
Col19a1
|
collagen, type XIX, alpha 1 |
| chr15_-_103160082 | 0.56 |
ENSMUST00000149111.8
ENSMUST00000132836.8 |
Nfe2
|
nuclear factor, erythroid derived 2 |
| chr4_+_116078830 | 0.54 |
ENSMUST00000030464.14
|
Pik3r3
|
phosphoinositide-3-kinase regulatory subunit 3 |
| chr14_+_53607470 | 0.52 |
ENSMUST00000103652.5
|
Trav14n-3
|
T cell receptor alpha variable 14N-3 |
| chr2_-_58457168 | 0.49 |
ENSMUST00000056376.12
|
Acvr1
|
activin A receptor, type 1 |
| chr5_+_94365864 | 0.47 |
ENSMUST00000179743.3
|
Pramel38
|
PRAME like 38 |
| chr17_+_57297280 | 0.37 |
ENSMUST00000002735.9
|
Clpp
|
caseinolytic mitochondrial matrix peptidase proteolytic subunit |
| chr9_+_102503815 | 0.35 |
ENSMUST00000038673.14
ENSMUST00000186693.2 |
Anapc13
|
anaphase promoting complex subunit 13 |
| chr9_+_102503476 | 0.35 |
ENSMUST00000190279.7
ENSMUST00000188398.7 |
Anapc13
|
anaphase promoting complex subunit 13 |
| chr15_-_103159892 | 0.20 |
ENSMUST00000133600.8
ENSMUST00000134554.2 ENSMUST00000156927.8 |
Nfe2
|
nuclear factor, erythroid derived 2 |
| chr7_+_101546059 | 0.14 |
ENSMUST00000143835.8
|
Anapc15
|
anaphase promoting complex C subunit 15 |
| chr18_+_77369654 | 0.13 |
ENSMUST00000096547.11
ENSMUST00000148341.9 ENSMUST00000123410.9 |
Loxhd1
|
lipoxygenase homology domains 1 |
| chr6_-_38814159 | 0.09 |
ENSMUST00000160962.8
|
Hipk2
|
homeodomain interacting protein kinase 2 |
| chr8_-_120228434 | 0.08 |
ENSMUST00000133821.2
ENSMUST00000036748.15 |
Slc38a8
|
solute carrier family 38, member 8 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.5 | 31.5 | GO:0006571 | tyrosine biosynthetic process(GO:0006571) aromatic amino acid family biosynthetic process(GO:0009073) aromatic amino acid family biosynthetic process, prephenate pathway(GO:0009095) |
| 7.2 | 21.7 | GO:2000487 | asparagine transport(GO:0006867) positive regulation of glutamine transport(GO:2000487) |
| 3.4 | 20.4 | GO:2000255 | negative regulation of male germ cell proliferation(GO:2000255) |
| 2.9 | 20.6 | GO:0060332 | positive regulation of humoral immune response mediated by circulating immunoglobulin(GO:0002925) heme transport(GO:0015886) positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) |
| 1.3 | 6.7 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.7 | 6.1 | GO:0019532 | oxalate transport(GO:0019532) |
| 0.5 | 5.9 | GO:0098789 | pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.5 | 8.0 | GO:0006707 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.4 | 2.3 | GO:1901842 | negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.3 | 2.0 | GO:0071421 | manganese ion transmembrane transport(GO:0071421) |
| 0.3 | 6.2 | GO:0035404 | histone-serine phosphorylation(GO:0035404) |
| 0.3 | 2.4 | GO:0098535 | de novo centriole assembly(GO:0098535) |
| 0.3 | 14.2 | GO:0006730 | one-carbon metabolic process(GO:0006730) |
| 0.2 | 3.4 | GO:0036376 | sodium ion export from cell(GO:0036376) |
| 0.2 | 0.8 | GO:1904884 | telomerase catalytic core complex assembly(GO:1904868) regulation of telomerase catalytic core complex assembly(GO:1904882) positive regulation of telomerase catalytic core complex assembly(GO:1904884) |
| 0.2 | 0.8 | GO:1904800 | regulation of neuron remodeling(GO:1904799) negative regulation of neuron remodeling(GO:1904800) |
| 0.2 | 4.1 | GO:0006309 | apoptotic DNA fragmentation(GO:0006309) |
| 0.1 | 6.2 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.1 | 1.9 | GO:0002318 | myeloid progenitor cell differentiation(GO:0002318) |
| 0.1 | 17.5 | GO:0009166 | nucleotide catabolic process(GO:0009166) |
| 0.1 | 3.7 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 0.5 | GO:0003274 | endocardial cushion fusion(GO:0003274) positive regulation of determination of dorsal identity(GO:2000017) |
| 0.1 | 3.5 | GO:0016572 | histone phosphorylation(GO:0016572) |
| 0.1 | 1.8 | GO:0034219 | carbohydrate transmembrane transport(GO:0034219) |
| 0.1 | 1.8 | GO:0016082 | synaptic vesicle priming(GO:0016082) |
| 0.1 | 12.2 | GO:0051291 | protein heterooligomerization(GO:0051291) |
| 0.1 | 0.8 | GO:0034242 | negative regulation of syncytium formation by plasma membrane fusion(GO:0034242) |
| 0.0 | 3.9 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.0 | 0.7 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.0 | 4.2 | GO:0016125 | sterol metabolic process(GO:0016125) |
| 0.0 | 1.9 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.0 | 2.5 | GO:0007286 | spermatid development(GO:0007286) |
| 0.0 | 0.4 | GO:0006515 | misfolded or incompletely synthesized protein catabolic process(GO:0006515) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 17.5 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.3 | 3.4 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.3 | 12.2 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.3 | 5.9 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.2 | 6.2 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.2 | 0.5 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.2 | 20.4 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.1 | 1.8 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.1 | 17.5 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.1 | 3.9 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.1 | 21.7 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 1.9 | GO:0097546 | ciliary base(GO:0097546) |
| 0.0 | 1.6 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.0 | 2.1 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.8 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 2.4 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 0.5 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 37.0 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.0 | 2.4 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 1.8 | GO:0030016 | myofibril(GO:0030016) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.4 | 21.7 | GO:0015182 | L-asparagine transmembrane transporter activity(GO:0015182) |
| 5.3 | 31.5 | GO:0004505 | phenylalanine 4-monooxygenase activity(GO:0004505) |
| 5.1 | 20.4 | GO:0004667 | prostaglandin-D synthase activity(GO:0004667) |
| 3.4 | 20.6 | GO:0015232 | heme transporter activity(GO:0015232) |
| 2.9 | 17.5 | GO:0004850 | uridine phosphorylase activity(GO:0004850) |
| 1.1 | 14.2 | GO:0016151 | nickel cation binding(GO:0016151) |
| 0.5 | 2.0 | GO:0051381 | histamine binding(GO:0051381) |
| 0.4 | 3.9 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 0.3 | 6.1 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.3 | 12.2 | GO:0030742 | GTP-dependent protein binding(GO:0030742) |
| 0.3 | 3.4 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.3 | 6.2 | GO:0035174 | histone serine kinase activity(GO:0035174) |
| 0.2 | 4.2 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 0.2 | 8.0 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.2 | 6.7 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.1 | 2.5 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.1 | 0.8 | GO:0032405 | MutLalpha complex binding(GO:0032405) |
| 0.1 | 21.1 | GO:0052689 | carboxylic ester hydrolase activity(GO:0052689) |
| 0.1 | 5.9 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.1 | 1.8 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.1 | 3.7 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.1 | 6.2 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.1 | 0.5 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.1 | 4.1 | GO:0004520 | endodeoxyribonuclease activity(GO:0004520) |
| 0.0 | 0.5 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 1.8 | GO:1901476 | carbohydrate transmembrane transporter activity(GO:0015144) carbohydrate transporter activity(GO:1901476) |
| 0.0 | 0.8 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 2.5 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 3.1 | GO:0016874 | ligase activity(GO:0016874) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 19.3 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.5 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 0.8 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.0 | 6.7 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 17.5 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.6 | 12.2 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.5 | 8.0 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.3 | 21.7 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.3 | 2.5 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.1 | 31.5 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.1 | 3.9 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 1.9 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.1 | 0.8 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.1 | 3.4 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 2.0 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.0 | 2.4 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 0.5 | REACTOME CD28 DEPENDENT PI3K AKT SIGNALING | Genes involved in CD28 dependent PI3K/Akt signaling |