Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
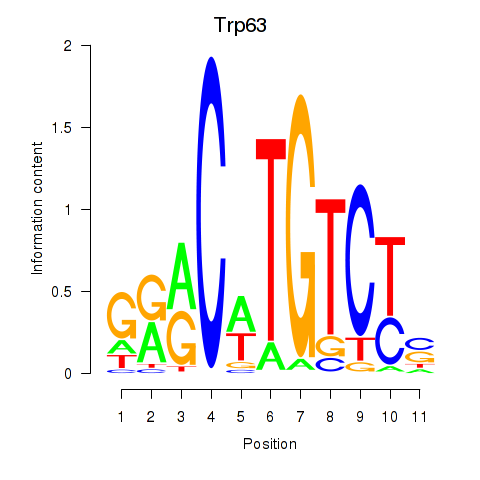
Results for Trp63
Z-value: 0.81
Transcription factors associated with Trp63
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Trp63
|
ENSMUSG00000022510.15 | Trp63 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Trp63 | mm39_v1_chr16_+_25620652_25620678 | 0.31 | 8.0e-03 | Click! |
Activity profile of Trp63 motif
Sorted Z-values of Trp63 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Trp63
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_+_29309942 | 6.47 |
ENSMUST00000119901.9
|
Cdkn1a
|
cyclin-dependent kinase inhibitor 1A (P21) |
| chr2_+_119068012 | 6.34 |
ENSMUST00000110817.3
|
Spint1
|
serine protease inhibitor, Kunitz type 1 |
| chr2_+_119067832 | 6.32 |
ENSMUST00000028783.14
|
Spint1
|
serine protease inhibitor, Kunitz type 1 |
| chr2_+_119067929 | 6.09 |
ENSMUST00000110816.8
|
Spint1
|
serine protease inhibitor, Kunitz type 1 |
| chr12_-_114646685 | 5.74 |
ENSMUST00000194350.6
ENSMUST00000103504.3 |
Ighv1-18
|
immunoglobulin heavy variable V1-18 |
| chr2_-_13496624 | 4.96 |
ENSMUST00000091436.7
|
Cubn
|
cubilin (intrinsic factor-cobalamin receptor) |
| chr2_-_170269748 | 4.69 |
ENSMUST00000013667.3
ENSMUST00000109152.9 ENSMUST00000068137.11 |
Bcas1
|
breast carcinoma amplified sequence 1 |
| chr3_-_92777521 | 4.02 |
ENSMUST00000163439.3
|
2310050C09Rik
|
RIKEN cDNA 2310050C09 gene |
| chr12_-_107969853 | 3.31 |
ENSMUST00000066060.11
|
Bcl11b
|
B cell leukemia/lymphoma 11B |
| chr11_-_69838971 | 3.20 |
ENSMUST00000179298.3
ENSMUST00000018710.13 ENSMUST00000135437.3 ENSMUST00000141837.9 ENSMUST00000142500.8 |
Slc2a4
|
solute carrier family 2 (facilitated glucose transporter), member 4 |
| chr6_+_4505493 | 2.89 |
ENSMUST00000031668.10
|
Col1a2
|
collagen, type I, alpha 2 |
| chr4_-_127247864 | 2.76 |
ENSMUST00000106090.8
ENSMUST00000060419.2 |
Gjb4
|
gap junction protein, beta 4 |
| chr8_+_45388466 | 2.74 |
ENSMUST00000191428.7
|
Fat1
|
FAT atypical cadherin 1 |
| chr12_-_107969673 | 2.69 |
ENSMUST00000109887.8
ENSMUST00000109891.3 |
Bcl11b
|
B cell leukemia/lymphoma 11B |
| chr6_+_34686543 | 2.65 |
ENSMUST00000031775.13
|
Cald1
|
caldesmon 1 |
| chr19_+_53128901 | 2.65 |
ENSMUST00000235754.2
ENSMUST00000237301.2 ENSMUST00000238130.2 |
Add3
|
adducin 3 (gamma) |
| chr19_+_53128861 | 2.26 |
ENSMUST00000111741.10
|
Add3
|
adducin 3 (gamma) |
| chr10_-_90918566 | 2.24 |
ENSMUST00000162618.8
ENSMUST00000020157.13 ENSMUST00000160788.2 |
Apaf1
|
apoptotic peptidase activating factor 1 |
| chr19_-_5925239 | 2.20 |
ENSMUST00000155227.2
|
Frmd8
|
FERM domain containing 8 |
| chrX_+_158038778 | 1.90 |
ENSMUST00000126686.8
ENSMUST00000033671.13 |
Rps6ka3
|
ribosomal protein S6 kinase polypeptide 3 |
| chr6_+_34686373 | 1.89 |
ENSMUST00000115021.8
|
Cald1
|
caldesmon 1 |
| chrX_+_158039107 | 1.83 |
ENSMUST00000148570.8
|
Rps6ka3
|
ribosomal protein S6 kinase polypeptide 3 |
| chrX_+_158038915 | 1.63 |
ENSMUST00000112492.8
|
Rps6ka3
|
ribosomal protein S6 kinase polypeptide 3 |
| chr19_-_5925296 | 1.55 |
ENSMUST00000025728.13
|
Frmd8
|
FERM domain containing 8 |
| chr8_+_27575611 | 1.53 |
ENSMUST00000178514.8
ENSMUST00000033876.14 |
Adgra2
|
adhesion G protein-coupled receptor A2 |
| chr11_-_101357046 | 1.50 |
ENSMUST00000040430.8
|
Vat1
|
vesicle amine transport 1 |
| chr11_+_98754434 | 1.47 |
ENSMUST00000142414.8
ENSMUST00000037480.9 |
Wipf2
|
WAS/WASL interacting protein family, member 2 |
| chr19_-_40365318 | 1.28 |
ENSMUST00000239304.2
|
Sorbs1
|
sorbin and SH3 domain containing 1 |
| chr4_-_63414188 | 1.09 |
ENSMUST00000063650.10
ENSMUST00000102867.8 ENSMUST00000107393.8 ENSMUST00000084510.8 ENSMUST00000095038.8 ENSMUST00000119294.8 ENSMUST00000095037.2 ENSMUST00000063672.10 |
Whrn
|
whirlin |
| chr6_+_11907808 | 1.09 |
ENSMUST00000155037.4
|
Phf14
|
PHD finger protein 14 |
| chr1_-_160134873 | 1.05 |
ENSMUST00000193185.6
|
Rabgap1l
|
RAB GTPase activating protein 1-like |
| chr7_+_24584197 | 0.93 |
ENSMUST00000156372.8
ENSMUST00000124035.2 |
Rps19
|
ribosomal protein S19 |
| chr6_-_94260806 | 0.91 |
ENSMUST00000203519.3
ENSMUST00000204347.3 |
Magi1
|
membrane associated guanylate kinase, WW and PDZ domain containing 1 |
| chr17_+_47983587 | 0.87 |
ENSMUST00000152724.2
|
Usp49
|
ubiquitin specific peptidase 49 |
| chr17_-_84495364 | 0.85 |
ENSMUST00000060366.7
|
Zfp36l2
|
zinc finger protein 36, C3H type-like 2 |
| chr7_-_65020955 | 0.83 |
ENSMUST00000102592.10
|
Tjp1
|
tight junction protein 1 |
| chr9_+_121245036 | 0.82 |
ENSMUST00000211187.2
|
Trak1
|
trafficking protein, kinesin binding 1 |
| chr2_+_119803230 | 0.78 |
ENSMUST00000229024.2
|
Mapkbp1
|
mitogen-activated protein kinase binding protein 1 |
| chr7_+_24584076 | 0.70 |
ENSMUST00000153451.9
ENSMUST00000108429.8 |
Rps19
|
ribosomal protein S19 |
| chr10_+_3784877 | 0.64 |
ENSMUST00000239159.2
|
Plekhg1
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 1 |
| chr1_+_180762587 | 0.63 |
ENSMUST00000037361.9
|
Lefty1
|
left right determination factor 1 |
| chr2_+_59442378 | 0.62 |
ENSMUST00000112568.8
ENSMUST00000037526.11 |
Tanc1
|
tetratricopeptide repeat, ankyrin repeat and coiled-coil containing 1 |
| chr2_+_119803180 | 0.61 |
ENSMUST00000066058.8
|
Mapkbp1
|
mitogen-activated protein kinase binding protein 1 |
| chr13_-_103470937 | 0.55 |
ENSMUST00000167058.8
ENSMUST00000164111.2 |
Mast4
|
microtubule associated serine/threonine kinase family member 4 |
| chr7_-_113853894 | 0.51 |
ENSMUST00000033012.9
|
Copb1
|
coatomer protein complex, subunit beta 1 |
| chr7_+_24583994 | 0.49 |
ENSMUST00000108428.8
|
Rps19
|
ribosomal protein S19 |
| chr7_-_102264194 | 0.43 |
ENSMUST00000098222.2
|
Olfr553
|
olfactory receptor 553 |
| chr14_+_64331130 | 0.38 |
ENSMUST00000224112.2
ENSMUST00000165710.2 ENSMUST00000170709.2 |
Prss51
|
protease, serine 51 |
| chr9_+_108437485 | 0.37 |
ENSMUST00000081111.14
ENSMUST00000193421.2 |
Impdh2
|
inosine monophosphate dehydrogenase 2 |
| chr2_-_91795910 | 0.30 |
ENSMUST00000239257.2
|
Dgkz
|
diacylglycerol kinase zeta |
| chr2_-_87524291 | 0.30 |
ENSMUST00000077471.5
|
Olfr1136
|
olfactory receptor 1136 |
| chr17_-_68311073 | 0.29 |
ENSMUST00000024840.12
|
Arhgap28
|
Rho GTPase activating protein 28 |
| chr7_-_65020655 | 0.19 |
ENSMUST00000032729.8
|
Tjp1
|
tight junction protein 1 |
| chr18_-_15851103 | 0.12 |
ENSMUST00000053017.13
|
Chst9
|
carbohydrate (N-acetylgalactosamine 4-0) sulfotransferase 9 |
| chr8_-_32408380 | 0.07 |
ENSMUST00000208497.3
ENSMUST00000207584.3 |
Nrg1
|
neuregulin 1 |
| chr3_+_75655504 | 0.06 |
ENSMUST00000189155.4
|
Gm29133
|
predicted gene 29133 |
| chr14_-_31299275 | 0.03 |
ENSMUST00000112027.9
|
Colq
|
collagen-like tail subunit (single strand of homotrimer) of asymmetric acetylcholinesterase |
| chr5_-_5315968 | 0.03 |
ENSMUST00000115451.8
ENSMUST00000115452.8 ENSMUST00000131392.8 |
Cdk14
|
cyclin-dependent kinase 14 |
| chr14_-_55114989 | 0.02 |
ENSMUST00000168622.2
ENSMUST00000177403.2 |
Ppp1r3e
|
protein phosphatase 1, regulatory subunit 3E |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 6.0 | GO:0097535 | lymphoid lineage cell migration(GO:0097534) lymphoid lineage cell migration into thymus(GO:0097535) |
| 0.9 | 18.7 | GO:0060670 | branching involved in labyrinthine layer morphogenesis(GO:0060670) |
| 0.8 | 5.0 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.5 | 2.1 | GO:0060265 | positive regulation of respiratory burst involved in inflammatory response(GO:0060265) |
| 0.4 | 6.5 | GO:0071493 | cellular response to UV-B(GO:0071493) |
| 0.3 | 1.5 | GO:0010637 | negative regulation of mitochondrial fusion(GO:0010637) |
| 0.2 | 1.5 | GO:0090210 | regulation of establishment of blood-brain barrier(GO:0090210) |
| 0.2 | 1.1 | GO:2000583 | regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000583) negative regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000584) |
| 0.2 | 1.4 | GO:0007256 | activation of JNKK activity(GO:0007256) negative regulation of defense response to bacterium(GO:1900425) |
| 0.2 | 2.2 | GO:0008635 | activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c(GO:0008635) |
| 0.2 | 2.9 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 0.2 | 0.9 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.2 | 2.8 | GO:0042048 | olfactory behavior(GO:0042048) |
| 0.2 | 0.8 | GO:0098957 | anterograde axonal transport of mitochondrion(GO:0098957) |
| 0.1 | 0.6 | GO:1900108 | negative regulation of nodal signaling pathway(GO:1900108) |
| 0.1 | 3.2 | GO:0044381 | glucose import in response to insulin stimulus(GO:0044381) |
| 0.1 | 0.9 | GO:1904628 | response to phorbol 13-acetate 12-myristate(GO:1904627) cellular response to phorbol 13-acetate 12-myristate(GO:1904628) |
| 0.1 | 0.3 | GO:1904425 | negative regulation of GTP binding(GO:1904425) |
| 0.1 | 1.1 | GO:0045217 | cell-cell junction maintenance(GO:0045217) |
| 0.1 | 4.0 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.1 | 1.0 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.1 | 0.4 | GO:0006177 | GMP biosynthetic process(GO:0006177) |
| 0.0 | 2.7 | GO:0045197 | establishment or maintenance of epithelial cell apical/basal polarity(GO:0045197) |
| 0.0 | 4.4 | GO:0006940 | regulation of smooth muscle contraction(GO:0006940) |
| 0.0 | 5.7 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 5.4 | GO:0002224 | toll-like receptor signaling pathway(GO:0002224) |
| 0.0 | 1.3 | GO:0070875 | positive regulation of glycogen biosynthetic process(GO:0045725) positive regulation of glycogen metabolic process(GO:0070875) |
| 0.0 | 1.0 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.0 | 0.6 | GO:0097062 | dendritic spine maintenance(GO:0097062) |
| 0.0 | 4.5 | GO:0042493 | response to drug(GO:0042493) |
| 0.0 | 0.3 | GO:0046339 | diacylglycerol metabolic process(GO:0046339) |
| 0.0 | 0.5 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.2 | 6.5 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 1.0 | 5.0 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 1.0 | 2.9 | GO:0005584 | collagen type I trimer(GO:0005584) |
| 0.5 | 4.5 | GO:0030478 | actin cap(GO:0030478) |
| 0.4 | 2.2 | GO:0043293 | apoptosome(GO:0043293) |
| 0.2 | 1.1 | GO:1990696 | stereocilia ankle link complex(GO:0002142) USH2 complex(GO:1990696) |
| 0.2 | 1.3 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.2 | 3.2 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.1 | 2.8 | GO:0005922 | connexon complex(GO:0005922) |
| 0.1 | 4.0 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 1.0 | GO:0005921 | gap junction(GO:0005921) |
| 0.0 | 1.4 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.0 | 5.7 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 0.8 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.0 | 0.5 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.0 | 2.1 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 4.9 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 2.7 | GO:0030175 | filopodium(GO:0030175) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.2 | 6.5 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 0.7 | 5.0 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.6 | 3.2 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
| 0.2 | 2.9 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.2 | 0.6 | GO:0038100 | nodal binding(GO:0038100) |
| 0.1 | 2.2 | GO:0008656 | cysteine-type endopeptidase activator activity involved in apoptotic process(GO:0008656) |
| 0.1 | 18.7 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 5.4 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.1 | 1.0 | GO:0071253 | connexin binding(GO:0071253) |
| 0.1 | 0.4 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.1 | 0.9 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.1 | 2.1 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.0 | 4.9 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 5.7 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 0.8 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 0.1 | GO:0001537 | N-acetylgalactosamine 4-O-sulfotransferase activity(GO:0001537) |
| 0.0 | 1.3 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 0.3 | GO:0004143 | lipid kinase activity(GO:0001727) diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 0.9 | GO:0051393 | alpha-actinin binding(GO:0051393) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 6.5 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.1 | 5.4 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.1 | 2.2 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.1 | 3.2 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 2.9 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.0 | 1.3 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 1.0 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 5.0 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.2 | 6.5 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.2 | 3.2 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.2 | 2.9 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.1 | 2.8 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 5.8 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 5.4 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.0 | 1.0 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 0.5 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.0 | 2.1 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 1.4 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.0 | 0.6 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 0.4 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |