Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
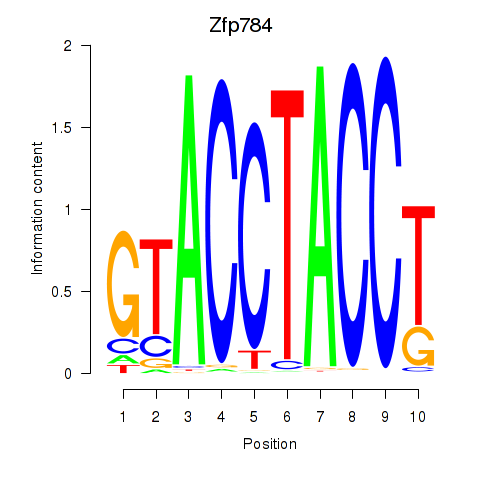
Results for Zfp784
Z-value: 1.02
Transcription factors associated with Zfp784
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Zfp784
|
ENSMUSG00000043290.7 | Zfp784 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Zfp784 | mm39_v1_chr7_-_5041427_5041450 | 0.33 | 4.4e-03 | Click! |
Activity profile of Zfp784 motif
Sorted Z-values of Zfp784 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Zfp784
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_+_8574420 | 7.45 |
ENSMUST00000029002.9
|
Stmn2
|
stathmin-like 2 |
| chr9_-_121324744 | 6.97 |
ENSMUST00000035120.6
|
Cck
|
cholecystokinin |
| chr16_-_31133622 | 6.96 |
ENSMUST00000115230.2
ENSMUST00000130560.8 |
Apod
|
apolipoprotein D |
| chr9_-_121324705 | 6.08 |
ENSMUST00000216138.2
|
Cck
|
cholecystokinin |
| chr1_+_106862171 | 5.41 |
ENSMUST00000081277.9
|
Serpinb12
|
serine (or cysteine) peptidase inhibitor, clade B (ovalbumin), member 12 |
| chr14_-_61275340 | 4.94 |
ENSMUST00000225730.2
ENSMUST00000111236.4 |
Tnfrsf19
|
tumor necrosis factor receptor superfamily, member 19 |
| chr1_-_133681419 | 4.77 |
ENSMUST00000125659.8
ENSMUST00000048953.14 ENSMUST00000165602.9 |
Atp2b4
|
ATPase, Ca++ transporting, plasma membrane 4 |
| chr7_-_27242170 | 4.66 |
ENSMUST00000150964.2
|
Pld3
|
phospholipase D family, member 3 |
| chr2_+_70392351 | 4.63 |
ENSMUST00000094934.11
|
Gad1
|
glutamate decarboxylase 1 |
| chr3_+_92325386 | 4.59 |
ENSMUST00000029533.3
|
Sprr2j-ps
|
small proline-rich protein 2J, pseudogene |
| chr6_+_86605146 | 4.50 |
ENSMUST00000043400.9
|
Asprv1
|
aspartic peptidase, retroviral-like 1 |
| chr7_-_105230807 | 4.40 |
ENSMUST00000191011.7
|
Apbb1
|
amyloid beta (A4) precursor protein-binding, family B, member 1 |
| chr7_-_24705320 | 4.29 |
ENSMUST00000102858.10
ENSMUST00000196684.2 ENSMUST00000080882.11 |
Atp1a3
|
ATPase, Na+/K+ transporting, alpha 3 polypeptide |
| chr7_-_105230395 | 4.22 |
ENSMUST00000188726.2
ENSMUST00000188440.7 |
Apbb1
|
amyloid beta (A4) precursor protein-binding, family B, member 1 |
| chr7_-_105230698 | 4.00 |
ENSMUST00000189378.7
|
Apbb1
|
amyloid beta (A4) precursor protein-binding, family B, member 1 |
| chr10_-_127031578 | 3.88 |
ENSMUST00000038217.14
ENSMUST00000130855.8 ENSMUST00000116229.2 ENSMUST00000144322.8 |
Dtx3
|
deltex 3, E3 ubiquitin ligase |
| chr7_-_105230479 | 3.84 |
ENSMUST00000191601.7
|
Apbb1
|
amyloid beta (A4) precursor protein-binding, family B, member 1 |
| chrX_+_73372664 | 3.79 |
ENSMUST00000004326.4
|
Plxna3
|
plexin A3 |
| chr4_+_42917228 | 3.70 |
ENSMUST00000107976.9
ENSMUST00000069184.9 |
Phf24
|
PHD finger protein 24 |
| chr13_+_73615316 | 3.69 |
ENSMUST00000022099.15
|
Lpcat1
|
lysophosphatidylcholine acyltransferase 1 |
| chr13_+_43019718 | 3.62 |
ENSMUST00000131942.2
|
Phactr1
|
phosphatase and actin regulator 1 |
| chr17_-_82045800 | 3.60 |
ENSMUST00000235015.2
ENSMUST00000163123.3 |
Slc8a1
|
solute carrier family 8 (sodium/calcium exchanger), member 1 |
| chr15_+_76544763 | 3.56 |
ENSMUST00000004294.12
|
Kifc2
|
kinesin family member C2 |
| chr4_+_41903610 | 3.53 |
ENSMUST00000098128.4
|
Ccl21d
|
chemokine (C-C motif) ligand 21D |
| chr7_+_19251763 | 3.49 |
ENSMUST00000207576.2
|
Nkpd1
|
NTPase, KAP family P-loop domain containing 1 |
| chr3_+_88536749 | 3.43 |
ENSMUST00000176316.8
ENSMUST00000176879.8 |
Arhgef2
|
rho/rac guanine nucleotide exchange factor (GEF) 2 |
| chr4_+_42612121 | 3.41 |
ENSMUST00000178168.3
|
Gm10591
|
predicted gene 10591 |
| chr9_-_119897328 | 3.41 |
ENSMUST00000177637.2
|
Cx3cr1
|
chemokine (C-X3-C motif) receptor 1 |
| chr9_+_122980006 | 3.39 |
ENSMUST00000026890.6
|
Clec3b
|
C-type lectin domain family 3, member b |
| chr3_+_84859453 | 3.30 |
ENSMUST00000029727.8
|
Fbxw7
|
F-box and WD-40 domain protein 7 |
| chr11_+_78215026 | 3.30 |
ENSMUST00000102478.4
|
Aldoc
|
aldolase C, fructose-bisphosphate |
| chr6_-_28831746 | 3.28 |
ENSMUST00000062304.7
|
Lrrc4
|
leucine rich repeat containing 4 |
| chr9_-_119897358 | 3.26 |
ENSMUST00000064165.5
|
Cx3cr1
|
chemokine (C-X3-C motif) receptor 1 |
| chr11_-_101979297 | 3.26 |
ENSMUST00000017458.11
|
Mpp2
|
membrane protein, palmitoylated 2 (MAGUK p55 subfamily member 2) |
| chr9_+_110075133 | 3.20 |
ENSMUST00000199736.2
|
Cspg5
|
chondroitin sulfate proteoglycan 5 |
| chr8_+_122457302 | 3.19 |
ENSMUST00000026357.12
|
Jph3
|
junctophilin 3 |
| chr2_+_103242027 | 3.08 |
ENSMUST00000239273.2
ENSMUST00000164172.8 |
Elf5
|
E74-like factor 5 |
| chr4_+_48049080 | 3.07 |
ENSMUST00000153369.2
|
Nr4a3
|
nuclear receptor subfamily 4, group A, member 3 |
| chr4_+_42114817 | 3.07 |
ENSMUST00000098123.4
|
Gm13304
|
predicted gene 13304 |
| chr1_+_149975782 | 3.04 |
ENSMUST00000035065.9
|
Ptgs2
|
prostaglandin-endoperoxide synthase 2 |
| chr11_-_42072990 | 2.99 |
ENSMUST00000205546.2
|
Gabra1
|
gamma-aminobutyric acid (GABA) A receptor, subunit alpha 1 |
| chr4_-_152561896 | 2.95 |
ENSMUST00000238738.2
ENSMUST00000162017.3 ENSMUST00000030768.10 |
Kcnab2
|
potassium voltage-gated channel, shaker-related subfamily, beta member 2 |
| chr5_+_66903174 | 2.91 |
ENSMUST00000101164.11
ENSMUST00000238785.2 |
Limch1
|
LIM and calponin homology domains 1 |
| chr4_+_42255693 | 2.90 |
ENSMUST00000178864.3
|
Ccl21b
|
chemokine (C-C motif) ligand 21B (leucine) |
| chr4_-_141966662 | 2.90 |
ENSMUST00000036476.10
|
Kazn
|
kazrin, periplakin interacting protein |
| chr18_+_56533389 | 2.86 |
ENSMUST00000237355.2
ENSMUST00000237422.2 |
Gramd3
|
GRAM domain containing 3 |
| chr6_+_86055018 | 2.81 |
ENSMUST00000205034.3
ENSMUST00000203724.3 |
Add2
|
adducin 2 (beta) |
| chr4_-_4138432 | 2.78 |
ENSMUST00000070375.8
|
Penk
|
preproenkephalin |
| chr4_-_42773987 | 2.78 |
ENSMUST00000095114.5
|
Ccl21a
|
chemokine (C-C motif) ligand 21A (serine) |
| chr6_-_67512768 | 2.74 |
ENSMUST00000058178.6
|
Tacstd2
|
tumor-associated calcium signal transducer 2 |
| chr15_+_92059224 | 2.71 |
ENSMUST00000068378.6
|
Cntn1
|
contactin 1 |
| chr15_-_97629209 | 2.68 |
ENSMUST00000100249.10
|
Endou
|
endonuclease, polyU-specific |
| chr7_+_139474612 | 2.67 |
ENSMUST00000053445.18
ENSMUST00000121839.2 |
Kndc1
|
kinase non-catalytic C-lobe domain (KIND) containing 1 |
| chr17_+_6047112 | 2.63 |
ENSMUST00000115786.8
|
Synj2
|
synaptojanin 2 |
| chr1_+_106908709 | 2.61 |
ENSMUST00000027564.8
|
Serpinb13
|
serine (or cysteine) peptidase inhibitor, clade B (ovalbumin), member 13 |
| chr7_+_4925781 | 2.61 |
ENSMUST00000207527.2
ENSMUST00000207687.2 ENSMUST00000208754.2 |
Nat14
|
N-acetyltransferase 14 |
| chr3_-_151960948 | 2.59 |
ENSMUST00000199423.5
ENSMUST00000198460.5 |
Nexn
|
nexilin |
| chr9_+_110074574 | 2.55 |
ENSMUST00000197850.5
|
Cspg5
|
chondroitin sulfate proteoglycan 5 |
| chr11_-_42072920 | 2.55 |
ENSMUST00000207274.2
|
Gabra1
|
gamma-aminobutyric acid (GABA) A receptor, subunit alpha 1 |
| chr7_-_45315762 | 2.49 |
ENSMUST00000210620.2
ENSMUST00000069800.6 |
Fut2
|
fucosyltransferase 2 |
| chr9_-_107109108 | 2.47 |
ENSMUST00000044532.11
|
Dock3
|
dedicator of cyto-kinesis 3 |
| chr17_+_12338161 | 2.43 |
ENSMUST00000024594.9
|
Agpat4
|
1-acylglycerol-3-phosphate O-acyltransferase 4 (lysophosphatidic acid acyltransferase, delta) |
| chr3_+_90561560 | 2.43 |
ENSMUST00000079286.4
|
S100a7a
|
S100 calcium binding protein A7A |
| chr2_+_24852409 | 2.39 |
ENSMUST00000028351.9
|
Dph7
|
diphthamine biosynethesis 7 |
| chr19_+_17114108 | 2.37 |
ENSMUST00000223920.2
ENSMUST00000225351.2 |
Prune2
|
prune homolog 2 |
| chr4_+_152160713 | 2.35 |
ENSMUST00000239025.2
|
Plekhg5
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 5 |
| chr8_+_59364789 | 2.27 |
ENSMUST00000062978.7
|
BC030500
|
cDNA sequence BC030500 |
| chr10_-_80834010 | 2.24 |
ENSMUST00000118465.8
|
Gng7
|
guanine nucleotide binding protein (G protein), gamma 7 |
| chr15_-_103123711 | 2.22 |
ENSMUST00000122182.2
ENSMUST00000108813.10 ENSMUST00000127191.2 |
Cbx5
|
chromobox 5 |
| chr9_+_108883907 | 2.22 |
ENSMUST00000154184.5
|
Shisa5
|
shisa family member 5 |
| chr6_+_8949669 | 2.21 |
ENSMUST00000060369.4
|
Nxph1
|
neurexophilin 1 |
| chr19_+_23881821 | 2.20 |
ENSMUST00000237688.2
|
Apba1
|
amyloid beta (A4) precursor protein binding, family A, member 1 |
| chr11_-_100026754 | 2.17 |
ENSMUST00000107411.3
|
Krt15
|
keratin 15 |
| chr4_+_101944740 | 2.17 |
ENSMUST00000106911.8
|
Pde4b
|
phosphodiesterase 4B, cAMP specific |
| chr9_-_97915227 | 2.14 |
ENSMUST00000035027.13
|
Clstn2
|
calsyntenin 2 |
| chr3_-_151960992 | 2.14 |
ENSMUST00000198750.5
|
Nexn
|
nexilin |
| chr6_+_86055048 | 2.14 |
ENSMUST00000032069.8
|
Add2
|
adducin 2 (beta) |
| chr2_+_14393127 | 2.12 |
ENSMUST00000114731.8
ENSMUST00000082290.8 |
Slc39a12
|
solute carrier family 39 (zinc transporter), member 12 |
| chr5_+_66903195 | 2.10 |
ENSMUST00000118242.8
|
Limch1
|
LIM and calponin homology domains 1 |
| chr11_+_6511133 | 2.10 |
ENSMUST00000160633.8
ENSMUST00000109721.3 |
Ccm2
|
cerebral cavernous malformation 2 |
| chr11_-_95405368 | 2.06 |
ENSMUST00000058866.8
|
Nxph3
|
neurexophilin 3 |
| chr6_-_99412306 | 2.05 |
ENSMUST00000113322.9
ENSMUST00000176850.8 ENSMUST00000176632.8 |
Foxp1
|
forkhead box P1 |
| chr12_-_79054050 | 2.03 |
ENSMUST00000056660.13
ENSMUST00000174721.8 |
Tmem229b
|
transmembrane protein 229B |
| chr16_-_74208180 | 2.03 |
ENSMUST00000117200.8
|
Robo2
|
roundabout guidance receptor 2 |
| chr1_+_40363701 | 1.99 |
ENSMUST00000095020.9
ENSMUST00000194296.6 |
Il1rl2
|
interleukin 1 receptor-like 2 |
| chr7_+_66393252 | 1.98 |
ENSMUST00000066475.11
|
Cers3
|
ceramide synthase 3 |
| chr5_+_37185673 | 1.96 |
ENSMUST00000173836.8
|
Jakmip1
|
janus kinase and microtubule interacting protein 1 |
| chr1_+_146373352 | 1.96 |
ENSMUST00000132847.8
ENSMUST00000166814.8 |
Brinp3
|
bone morphogenetic protein/retinoic acid inducible neural specific 3 |
| chr11_-_97934368 | 1.96 |
ENSMUST00000131519.2
|
Stac2
|
SH3 and cysteine rich domain 2 |
| chrX_-_132799041 | 1.94 |
ENSMUST00000176718.8
ENSMUST00000176641.2 |
Tspan6
|
tetraspanin 6 |
| chr10_-_20600442 | 1.93 |
ENSMUST00000170265.8
|
Pde7b
|
phosphodiesterase 7B |
| chr4_+_57821050 | 1.88 |
ENSMUST00000238994.2
|
Pakap
|
paralemmin A kinase anchor protein |
| chr12_-_76842263 | 1.87 |
ENSMUST00000082431.6
|
Gpx2
|
glutathione peroxidase 2 |
| chr7_+_142025817 | 1.87 |
ENSMUST00000105966.2
|
Lsp1
|
lymphocyte specific 1 |
| chr10_-_20600797 | 1.86 |
ENSMUST00000020165.14
|
Pde7b
|
phosphodiesterase 7B |
| chr15_-_72418189 | 1.86 |
ENSMUST00000044624.8
|
Kcnk9
|
potassium channel, subfamily K, member 9 |
| chr7_-_12537533 | 1.79 |
ENSMUST00000210650.2
|
Zscan18
|
zinc finger and SCAN domain containing 18 |
| chr5_-_24534554 | 1.79 |
ENSMUST00000115098.7
|
Kcnh2
|
potassium voltage-gated channel, subfamily H (eag-related), member 2 |
| chr7_+_83234118 | 1.79 |
ENSMUST00000039317.14
ENSMUST00000164944.2 |
Tmc3
|
transmembrane channel-like gene family 3 |
| chr11_+_59197746 | 1.79 |
ENSMUST00000000128.10
ENSMUST00000108783.4 |
Wnt9a
|
wingless-type MMTV integration site family, member 9A |
| chr4_-_59783780 | 1.75 |
ENSMUST00000107526.8
ENSMUST00000095063.11 |
Inip
|
INTS3 and NABP interacting protein |
| chr4_-_110144676 | 1.75 |
ENSMUST00000106598.8
ENSMUST00000102723.11 ENSMUST00000153906.2 |
Elavl4
|
ELAV like RNA binding protein 4 |
| chr11_+_92989229 | 1.66 |
ENSMUST00000107859.8
ENSMUST00000107861.8 ENSMUST00000042943.13 ENSMUST00000107858.9 |
Car10
|
carbonic anhydrase 10 |
| chrX_+_106132840 | 1.65 |
ENSMUST00000118666.8
ENSMUST00000053375.4 |
P2ry10
|
purinergic receptor P2Y, G-protein coupled 10 |
| chr2_+_70392491 | 1.63 |
ENSMUST00000148210.8
|
Gad1
|
glutamate decarboxylase 1 |
| chr12_+_102095260 | 1.56 |
ENSMUST00000079020.12
|
Slc24a4
|
solute carrier family 24 (sodium/potassium/calcium exchanger), member 4 |
| chr16_-_84970617 | 1.55 |
ENSMUST00000226232.2
ENSMUST00000227021.2 ENSMUST00000005406.12 ENSMUST00000227723.2 |
App
|
amyloid beta (A4) precursor protein |
| chr12_+_55171254 | 1.55 |
ENSMUST00000177768.3
|
Fam177a
|
family with sequence similarity 177, member A |
| chr15_+_80861966 | 1.55 |
ENSMUST00000139517.9
ENSMUST00000137255.3 ENSMUST00000137004.2 |
Sgsm3
|
small G protein signaling modulator 3 |
| chrX_+_150961960 | 1.53 |
ENSMUST00000096275.5
|
Iqsec2
|
IQ motif and Sec7 domain 2 |
| chr19_-_40371016 | 1.52 |
ENSMUST00000225766.3
|
Sorbs1
|
sorbin and SH3 domain containing 1 |
| chr2_+_130119077 | 1.52 |
ENSMUST00000028890.15
ENSMUST00000159373.2 |
Nop56
|
NOP56 ribonucleoprotein |
| chr7_-_108774367 | 1.51 |
ENSMUST00000207178.2
|
Lmo1
|
LIM domain only 1 |
| chr2_+_174126103 | 1.51 |
ENSMUST00000109095.8
ENSMUST00000180362.8 ENSMUST00000109096.9 |
Gnas
|
GNAS (guanine nucleotide binding protein, alpha stimulating) complex locus |
| chr9_-_60048503 | 1.50 |
ENSMUST00000034829.6
|
Thsd4
|
thrombospondin, type I, domain containing 4 |
| chr1_-_132470672 | 1.50 |
ENSMUST00000086521.11
|
Cntn2
|
contactin 2 |
| chr1_-_86598286 | 1.49 |
ENSMUST00000027449.6
|
Nppc
|
natriuretic peptide type C |
| chr2_+_110551927 | 1.48 |
ENSMUST00000111017.9
|
Muc15
|
mucin 15 |
| chr2_+_156317416 | 1.47 |
ENSMUST00000029155.16
|
Epb41l1
|
erythrocyte membrane protein band 4.1 like 1 |
| chr16_-_84970580 | 1.47 |
ENSMUST00000227737.2
ENSMUST00000226801.2 |
App
|
amyloid beta (A4) precursor protein |
| chr2_+_118219096 | 1.40 |
ENSMUST00000110877.8
ENSMUST00000005233.12 |
Eif2ak4
|
eukaryotic translation initiation factor 2 alpha kinase 4 |
| chr1_+_133965228 | 1.39 |
ENSMUST00000162779.2
|
Fmod
|
fibromodulin |
| chr2_+_110551685 | 1.39 |
ENSMUST00000111016.9
|
Muc15
|
mucin 15 |
| chr14_-_119336809 | 1.38 |
ENSMUST00000156203.8
|
Uggt2
|
UDP-glucose glycoprotein glucosyltransferase 2 |
| chr14_-_29443792 | 1.38 |
ENSMUST00000022567.9
|
Cacna2d3
|
calcium channel, voltage-dependent, alpha2/delta subunit 3 |
| chr9_-_97915036 | 1.36 |
ENSMUST00000162295.2
|
Clstn2
|
calsyntenin 2 |
| chr19_+_8966641 | 1.34 |
ENSMUST00000092956.4
ENSMUST00000092955.11 |
Ahnak
|
AHNAK nucleoprotein (desmoyokin) |
| chr17_-_23959334 | 1.33 |
ENSMUST00000024702.5
|
Paqr4
|
progestin and adipoQ receptor family member IV |
| chr7_+_120442048 | 1.32 |
ENSMUST00000047875.16
|
Eef2k
|
eukaryotic elongation factor-2 kinase |
| chr4_-_154181562 | 1.31 |
ENSMUST00000105643.8
ENSMUST00000133533.8 ENSMUST00000097762.11 |
Trp73
|
transformation related protein 73 |
| chr13_-_67569483 | 1.30 |
ENSMUST00000224113.2
ENSMUST00000223644.2 ENSMUST00000056470.10 |
Zfp459
|
zinc finger protein 459 |
| chr2_-_23462052 | 1.29 |
ENSMUST00000132484.7
|
Spopl
|
speckle-type BTB/POZ protein-like |
| chr12_-_40298072 | 1.27 |
ENSMUST00000169926.8
|
Ifrd1
|
interferon-related developmental regulator 1 |
| chr5_-_99876671 | 1.25 |
ENSMUST00000209346.2
|
Rasgef1b
|
RasGEF domain family, member 1B |
| chr19_-_29344694 | 1.21 |
ENSMUST00000138051.2
|
Plgrkt
|
plasminogen receptor, C-terminal lysine transmembrane protein |
| chr6_+_29348068 | 1.21 |
ENSMUST00000173216.8
ENSMUST00000173694.5 ENSMUST00000172974.8 ENSMUST00000031779.17 ENSMUST00000090481.14 |
Calu
|
calumenin |
| chr7_+_142025575 | 1.19 |
ENSMUST00000038946.9
|
Lsp1
|
lymphocyte specific 1 |
| chr12_-_46863726 | 1.17 |
ENSMUST00000219330.2
|
Nova1
|
NOVA alternative splicing regulator 1 |
| chr7_+_120442773 | 1.17 |
ENSMUST00000143279.3
ENSMUST00000106489.9 |
Eef2k
|
eukaryotic elongation factor-2 kinase |
| chr19_-_6964988 | 1.15 |
ENSMUST00000130048.8
ENSMUST00000025914.7 |
Vegfb
|
vascular endothelial growth factor B |
| chr7_+_81824544 | 1.15 |
ENSMUST00000032874.14
|
Sh3gl3
|
SH3-domain GRB2-like 3 |
| chr14_-_70864448 | 1.14 |
ENSMUST00000110984.4
|
Dmtn
|
dematin actin binding protein |
| chr5_-_123887434 | 1.14 |
ENSMUST00000182955.8
ENSMUST00000182489.8 ENSMUST00000183147.9 ENSMUST00000050827.14 ENSMUST00000057795.12 ENSMUST00000111515.8 ENSMUST00000182309.8 |
Rsrc2
|
arginine/serine-rich coiled-coil 2 |
| chr13_+_48816466 | 1.14 |
ENSMUST00000021813.5
|
Barx1
|
BarH-like homeobox 1 |
| chr9_-_20864096 | 1.14 |
ENSMUST00000004202.17
|
Dnmt1
|
DNA methyltransferase (cytosine-5) 1 |
| chrX_-_135644424 | 1.13 |
ENSMUST00000166478.8
ENSMUST00000113097.8 |
Morf4l2
|
mortality factor 4 like 2 |
| chr6_+_21215472 | 1.13 |
ENSMUST00000081542.6
|
Kcnd2
|
potassium voltage-gated channel, Shal-related family, member 2 |
| chr11_+_115865535 | 1.12 |
ENSMUST00000021107.14
ENSMUST00000169928.8 ENSMUST00000106461.8 |
Itgb4
|
integrin beta 4 |
| chr2_+_14393245 | 1.12 |
ENSMUST00000133258.2
|
Slc39a12
|
solute carrier family 39 (zinc transporter), member 12 |
| chr5_+_30745447 | 1.09 |
ENSMUST00000066295.5
|
Kcnk3
|
potassium channel, subfamily K, member 3 |
| chr2_+_124910037 | 1.08 |
ENSMUST00000070353.4
|
Slc24a5
|
solute carrier family 24, member 5 |
| chr11_-_101917745 | 1.08 |
ENSMUST00000107167.2
ENSMUST00000062801.11 |
Mpp3
|
membrane protein, palmitoylated 3 (MAGUK p55 subfamily member 3) |
| chr9_+_98868418 | 1.07 |
ENSMUST00000035038.8
ENSMUST00000112911.9 |
Faim
|
Fas apoptotic inhibitory molecule |
| chr8_-_34237752 | 1.06 |
ENSMUST00000179364.3
|
Smim18
|
small integral membrane protein 18 |
| chr14_-_65662609 | 1.04 |
ENSMUST00000224594.2
|
Pnoc
|
prepronociceptin |
| chr14_-_70864666 | 1.03 |
ENSMUST00000022694.17
|
Dmtn
|
dematin actin binding protein |
| chr7_+_43418321 | 1.03 |
ENSMUST00000107970.8
|
Klk12
|
kallikrein related-peptidase 12 |
| chr5_-_53864595 | 1.03 |
ENSMUST00000200691.4
|
Cckar
|
cholecystokinin A receptor |
| chr4_-_110143777 | 1.02 |
ENSMUST00000138972.8
|
Elavl4
|
ELAV like RNA binding protein 4 |
| chr7_+_123582021 | 1.02 |
ENSMUST00000106437.2
|
Hs3st4
|
heparan sulfate (glucosamine) 3-O-sulfotransferase 4 |
| chr3_+_88536805 | 1.01 |
ENSMUST00000175745.2
|
Arhgef2
|
rho/rac guanine nucleotide exchange factor (GEF) 2 |
| chr7_+_101583283 | 1.01 |
ENSMUST00000209639.2
ENSMUST00000210679.2 |
Numa1
|
nuclear mitotic apparatus protein 1 |
| chr6_-_118539187 | 0.99 |
ENSMUST00000112830.3
|
Ankrd26
|
ankyrin repeat domain 26 |
| chr5_-_92190859 | 0.99 |
ENSMUST00000069937.11
ENSMUST00000086978.12 |
Cdkl2
|
cyclin-dependent kinase-like 2 (CDC2-related kinase) |
| chr13_+_52750883 | 0.99 |
ENSMUST00000055087.7
|
Syk
|
spleen tyrosine kinase |
| chr11_-_34674677 | 0.99 |
ENSMUST00000093193.12
ENSMUST00000101365.9 |
Dock2
|
dedicator of cyto-kinesis 2 |
| chr2_+_32465730 | 0.98 |
ENSMUST00000055304.14
|
Pip5kl1
|
phosphatidylinositol-4-phosphate 5-kinase-like 1 |
| chr14_-_47655621 | 0.98 |
ENSMUST00000180299.8
|
Dlgap5
|
DLG associated protein 5 |
| chr7_-_24997291 | 0.98 |
ENSMUST00000148150.8
ENSMUST00000155118.2 |
Pafah1b3
|
platelet-activating factor acetylhydrolase, isoform 1b, subunit 3 |
| chr7_+_107166925 | 0.98 |
ENSMUST00000239087.2
|
Olfml1
|
olfactomedin-like 1 |
| chr11_+_28803188 | 0.97 |
ENSMUST00000020759.12
|
Efemp1
|
epidermal growth factor-containing fibulin-like extracellular matrix protein 1 |
| chr16_+_4756963 | 0.97 |
ENSMUST00000229961.2
ENSMUST00000050881.10 |
Nudt16l1
|
nudix (nucleoside diphosphate linked moiety X)-type motif 16-like 1 |
| chr11_+_50267808 | 0.97 |
ENSMUST00000109142.8
|
Hnrnph1
|
heterogeneous nuclear ribonucleoprotein H1 |
| chr5_+_122344854 | 0.96 |
ENSMUST00000145854.8
|
Hvcn1
|
hydrogen voltage-gated channel 1 |
| chr16_-_18107046 | 0.96 |
ENSMUST00000232424.2
ENSMUST00000009321.11 |
Dgcr8
|
DGCR8, microprocessor complex subunit |
| chr8_-_91544021 | 0.95 |
ENSMUST00000209208.2
|
Gm19935
|
predicted gene, 19935 |
| chr10_-_30679071 | 0.95 |
ENSMUST00000068567.5
|
Ncoa7
|
nuclear receptor coactivator 7 |
| chr4_-_41774097 | 0.94 |
ENSMUST00000108036.8
ENSMUST00000108037.9 ENSMUST00000108032.3 ENSMUST00000173865.9 ENSMUST00000155240.2 |
Ccl27a
|
chemokine (C-C motif) ligand 27A |
| chr19_-_40371242 | 0.93 |
ENSMUST00000224583.2
|
Sorbs1
|
sorbin and SH3 domain containing 1 |
| chr7_-_24997393 | 0.93 |
ENSMUST00000005583.12
|
Pafah1b3
|
platelet-activating factor acetylhydrolase, isoform 1b, subunit 3 |
| chr5_-_53864874 | 0.92 |
ENSMUST00000031093.5
|
Cckar
|
cholecystokinin A receptor |
| chr13_+_107083613 | 0.92 |
ENSMUST00000022203.10
|
Dimt1
|
DIM1 dimethyladenosine transferase 1-like (S. cerevisiae) |
| chr11_+_73051228 | 0.91 |
ENSMUST00000006104.10
ENSMUST00000135202.8 ENSMUST00000136894.3 |
P2rx5
|
purinergic receptor P2X, ligand-gated ion channel, 5 |
| chr19_-_12093187 | 0.91 |
ENSMUST00000208391.3
ENSMUST00000214103.2 |
Olfr1428
|
olfactory receptor 1428 |
| chr1_+_151447232 | 0.90 |
ENSMUST00000111875.2
|
Niban1
|
niban apoptosis regulator 1 |
| chr14_+_27057980 | 0.89 |
ENSMUST00000224981.2
|
Arhgef3
|
Rho guanine nucleotide exchange factor (GEF) 3 |
| chr18_+_42186713 | 0.89 |
ENSMUST00000072008.11
|
Sh3rf2
|
SH3 domain containing ring finger 2 |
| chr4_-_87951565 | 0.86 |
ENSMUST00000078090.12
|
Mllt3
|
myeloid/lymphoid or mixed-lineage leukemia; translocated to, 3 |
| chr16_-_16345197 | 0.84 |
ENSMUST00000069284.14
|
Fgd4
|
FYVE, RhoGEF and PH domain containing 4 |
| chr15_+_32244947 | 0.83 |
ENSMUST00000067458.7
|
Sema5a
|
sema domain, seven thrombospondin repeats (type 1 and type 1-like), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 5A |
| chr11_-_30218167 | 0.83 |
ENSMUST00000006629.14
|
Sptbn1
|
spectrin beta, non-erythrocytic 1 |
| chr9_+_44151962 | 0.81 |
ENSMUST00000092426.5
ENSMUST00000217221.2 ENSMUST00000213891.2 |
Ccdc153
|
coiled-coil domain containing 153 |
| chr15_+_80862074 | 0.79 |
ENSMUST00000229727.2
|
Sgsm3
|
small G protein signaling modulator 3 |
| chr17_+_35844091 | 0.79 |
ENSMUST00000025273.9
|
Psors1c2
|
psoriasis susceptibility 1 candidate 2 (human) |
| chr3_-_136031799 | 0.79 |
ENSMUST00000196159.5
ENSMUST00000041577.13 |
Bank1
|
B cell scaffold protein with ankyrin repeats 1 |
| chrX_+_10583629 | 0.78 |
ENSMUST00000115524.8
ENSMUST00000008179.7 ENSMUST00000156321.2 |
Mid1ip1
|
Mid1 interacting protein 1 (gastrulation specific G12-like (zebrafish)) |
| chr3_+_125474628 | 0.77 |
ENSMUST00000174648.6
|
Ndst4
|
N-deacetylase/N-sulfotransferase (heparin glucosaminyl) 4 |
| chr2_-_45002902 | 0.77 |
ENSMUST00000076836.13
ENSMUST00000176732.8 ENSMUST00000200844.4 |
Zeb2
|
zinc finger E-box binding homeobox 2 |
| chr6_+_31375670 | 0.75 |
ENSMUST00000026699.15
|
Mkln1
|
muskelin 1, intracellular mediator containing kelch motifs |
| chr12_+_112772530 | 0.74 |
ENSMUST00000037014.11
ENSMUST00000177808.3 |
Clba1
|
clathrin binding box of aftiphilin containing 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.7 | 16.5 | GO:0050760 | thymidylate synthase biosynthetic process(GO:0050757) regulation of thymidylate synthase biosynthetic process(GO:0050758) negative regulation of thymidylate synthase biosynthetic process(GO:0050760) |
| 2.3 | 7.0 | GO:2000097 | negative regulation of lipoprotein oxidation(GO:0034443) regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 2.2 | 6.7 | GO:0002545 | chronic inflammatory response to non-antigenic stimulus(GO:0002545) regulation of chronic inflammatory response to non-antigenic stimulus(GO:0002880) |
| 1.9 | 7.4 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 1.6 | 4.8 | GO:0045763 | negative regulation of cellular amino acid metabolic process(GO:0045763) |
| 1.0 | 3.0 | GO:0090362 | positive regulation of platelet-derived growth factor production(GO:0090362) |
| 1.0 | 3.0 | GO:0051563 | smooth endoplasmic reticulum calcium ion homeostasis(GO:0051563) cellular response to norepinephrine stimulus(GO:0071874) |
| 0.9 | 2.8 | GO:2000547 | regulation of dendritic cell dendrite assembly(GO:2000547) |
| 0.9 | 3.7 | GO:2001245 | regulation of phosphatidylcholine biosynthetic process(GO:2001245) |
| 0.9 | 6.3 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 0.9 | 13.0 | GO:0051901 | positive regulation of mitochondrial depolarization(GO:0051901) |
| 0.8 | 3.1 | GO:1900625 | positive regulation of mast cell cytokine production(GO:0032765) regulation of monocyte aggregation(GO:1900623) positive regulation of monocyte aggregation(GO:1900625) |
| 0.7 | 2.0 | GO:0050925 | negative regulation of negative chemotaxis(GO:0050925) |
| 0.7 | 3.3 | GO:1903378 | positive regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903378) |
| 0.6 | 4.4 | GO:0071802 | negative regulation of podosome assembly(GO:0071802) |
| 0.5 | 2.7 | GO:0090191 | negative regulation of branching involved in ureteric bud morphogenesis(GO:0090191) |
| 0.5 | 3.8 | GO:0021637 | trigeminal nerve morphogenesis(GO:0021636) trigeminal nerve structural organization(GO:0021637) semaphorin-plexin signaling pathway involved in axon guidance(GO:1902287) |
| 0.5 | 1.9 | GO:0009609 | response to symbiont(GO:0009608) response to symbiotic bacterium(GO:0009609) |
| 0.5 | 7.9 | GO:0071436 | sodium ion export(GO:0071436) |
| 0.4 | 1.8 | GO:1902303 | negative regulation of potassium ion export(GO:1902303) |
| 0.4 | 3.1 | GO:0001712 | ectodermal cell fate commitment(GO:0001712) |
| 0.4 | 2.0 | GO:0017182 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.4 | 1.5 | GO:0060168 | positive regulation of adenosine receptor signaling pathway(GO:0060168) |
| 0.4 | 3.7 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.4 | 2.2 | GO:0035585 | calcium-mediated signaling using extracellular calcium source(GO:0035585) |
| 0.3 | 1.4 | GO:0071262 | regulation of eIF2 alpha phosphorylation by amino acid starvation(GO:0060733) regulation of translational initiation in response to starvation(GO:0071262) positive regulation of translational initiation in response to starvation(GO:0071264) |
| 0.3 | 1.0 | GO:0002277 | myeloid dendritic cell activation involved in immune response(GO:0002277) |
| 0.3 | 1.9 | GO:0002023 | reduction of food intake in response to dietary excess(GO:0002023) positive regulation of somatostatin secretion(GO:0090274) |
| 0.3 | 2.3 | GO:0048227 | plasma membrane to endosome transport(GO:0048227) |
| 0.3 | 2.6 | GO:0097283 | keratinocyte apoptotic process(GO:0097283) regulation of keratinocyte apoptotic process(GO:1902172) |
| 0.3 | 1.1 | GO:0090116 | C-5 methylation of cytosine(GO:0090116) positive regulation of methylation-dependent chromatin silencing(GO:0090309) |
| 0.3 | 1.1 | GO:1900377 | negative regulation of melanin biosynthetic process(GO:0048022) negative regulation of secondary metabolite biosynthetic process(GO:1900377) |
| 0.3 | 1.3 | GO:1902167 | positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator(GO:1902167) |
| 0.3 | 1.0 | GO:1902365 | regulation of spindle elongation(GO:0032887) regulation of mitotic spindle elongation(GO:0032888) anastral spindle assembly(GO:0055048) protein localization to spindle pole body(GO:0071988) regulation of protein localization to spindle pole body(GO:1902363) positive regulation of protein localization to spindle pole body(GO:1902365) positive regulation of mitotic spindle elongation(GO:1902846) |
| 0.2 | 1.0 | GO:0032752 | serotonin secretion by platelet(GO:0002554) positive regulation of interleukin-3 production(GO:0032752) interleukin-3 biosynthetic process(GO:0042223) regulation of interleukin-3 biosynthetic process(GO:0045399) positive regulation of interleukin-3 biosynthetic process(GO:0045401) positive regulation of granulocyte macrophage colony-stimulating factor biosynthetic process(GO:0045425) cellular response to molecule of fungal origin(GO:0071226) |
| 0.2 | 0.7 | GO:0006428 | isoleucyl-tRNA aminoacylation(GO:0006428) |
| 0.2 | 5.5 | GO:0071420 | cellular response to histamine(GO:0071420) |
| 0.2 | 2.6 | GO:1901250 | regulation of lung goblet cell differentiation(GO:1901249) negative regulation of lung goblet cell differentiation(GO:1901250) |
| 0.2 | 1.2 | GO:0035470 | positive regulation of vascular wound healing(GO:0035470) regulation of vascular wound healing(GO:0061043) |
| 0.2 | 4.7 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 0.2 | 1.5 | GO:0046013 | regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.2 | 3.3 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.2 | 1.3 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.2 | 0.5 | GO:0060734 | regulation of endoplasmic reticulum stress-induced eIF2 alpha phosphorylation(GO:0060734) |
| 0.2 | 2.7 | GO:0021684 | cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.2 | 0.7 | GO:0072362 | regulation of glycolytic process by negative regulation of transcription from RNA polymerase II promoter(GO:0072362) |
| 0.2 | 3.2 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.2 | 1.5 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) |
| 0.2 | 0.7 | GO:0015786 | UDP-glucose transport(GO:0015786) |
| 0.2 | 2.2 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.2 | 0.8 | GO:0048842 | positive regulation of axon extension involved in axon guidance(GO:0048842) |
| 0.2 | 3.3 | GO:0097119 | postsynaptic density protein 95 clustering(GO:0097119) |
| 0.2 | 2.8 | GO:0002118 | aggressive behavior(GO:0002118) |
| 0.2 | 0.9 | GO:0071677 | positive regulation of mononuclear cell migration(GO:0071677) |
| 0.2 | 3.0 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.2 | 1.5 | GO:1900454 | positive regulation of long term synaptic depression(GO:1900454) |
| 0.2 | 0.5 | GO:1904431 | response to intra-S DNA damage checkpoint signaling(GO:0072429) positive regulation of t-circle formation(GO:1904431) |
| 0.1 | 2.1 | GO:0060837 | blood vessel endothelial cell differentiation(GO:0060837) |
| 0.1 | 2.9 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.1 | 0.4 | GO:1901535 | cell-cell signaling involved in kidney development(GO:0060995) Wnt signaling pathway involved in kidney development(GO:0061289) canonical Wnt signaling pathway involved in metanephric kidney development(GO:0061290) cell-cell signaling involved in metanephros development(GO:0072204) regulation of DNA demethylation(GO:1901535) negative regulation of DNA demethylation(GO:1901536) |
| 0.1 | 0.7 | GO:0000727 | double-strand break repair via break-induced replication(GO:0000727) |
| 0.1 | 1.5 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.1 | 1.1 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.1 | 0.9 | GO:0035735 | intraciliary transport involved in cilium morphogenesis(GO:0035735) |
| 0.1 | 0.5 | GO:0021564 | vagus nerve development(GO:0021564) |
| 0.1 | 3.2 | GO:0060314 | regulation of ryanodine-sensitive calcium-release channel activity(GO:0060314) |
| 0.1 | 2.6 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.1 | 5.0 | GO:0050901 | leukocyte tethering or rolling(GO:0050901) |
| 0.1 | 1.9 | GO:0039532 | negative regulation of viral-induced cytoplasmic pattern recognition receptor signaling pathway(GO:0039532) |
| 0.1 | 1.6 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.1 | 0.3 | GO:0034476 | U1 snRNA 3'-end processing(GO:0034473) U5 snRNA 3'-end processing(GO:0034476) |
| 0.1 | 1.0 | GO:0007182 | common-partner SMAD protein phosphorylation(GO:0007182) |
| 0.1 | 3.7 | GO:0050966 | detection of mechanical stimulus involved in sensory perception of pain(GO:0050966) |
| 0.1 | 1.0 | GO:2001032 | regulation of double-strand break repair via nonhomologous end joining(GO:2001032) |
| 0.1 | 1.5 | GO:1900165 | negative regulation of interleukin-6 secretion(GO:1900165) |
| 0.1 | 1.0 | GO:0031053 | primary miRNA processing(GO:0031053) |
| 0.1 | 1.1 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.1 | 0.8 | GO:0030035 | microspike assembly(GO:0030035) |
| 0.1 | 2.2 | GO:0014051 | gamma-aminobutyric acid secretion(GO:0014051) |
| 0.1 | 2.6 | GO:0097186 | amelogenesis(GO:0097186) |
| 0.1 | 0.4 | GO:2000211 | regulation of glutamate metabolic process(GO:2000211) |
| 0.1 | 1.4 | GO:0071712 | ER-associated misfolded protein catabolic process(GO:0071712) |
| 0.1 | 0.9 | GO:0032074 | negative regulation of nuclease activity(GO:0032074) |
| 0.1 | 1.0 | GO:1903975 | regulation of glial cell migration(GO:1903975) |
| 0.1 | 1.0 | GO:0071294 | cellular response to zinc ion(GO:0071294) |
| 0.1 | 0.5 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.1 | 2.9 | GO:0031424 | keratinization(GO:0031424) |
| 0.1 | 0.6 | GO:0010756 | positive regulation of plasminogen activation(GO:0010756) |
| 0.1 | 0.3 | GO:0046341 | CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.1 | 0.9 | GO:0043416 | regulation of skeletal muscle tissue regeneration(GO:0043416) |
| 0.1 | 0.6 | GO:2000270 | negative regulation of fibroblast apoptotic process(GO:2000270) |
| 0.1 | 0.5 | GO:2001181 | positive regulation of interleukin-10 secretion(GO:2001181) |
| 0.1 | 2.5 | GO:0061003 | positive regulation of dendritic spine morphogenesis(GO:0061003) |
| 0.1 | 2.4 | GO:0000154 | rRNA modification(GO:0000154) |
| 0.1 | 2.3 | GO:0035767 | endothelial cell chemotaxis(GO:0035767) |
| 0.1 | 2.9 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.1 | 0.2 | GO:0060447 | bud outgrowth involved in lung branching(GO:0060447) |
| 0.1 | 1.3 | GO:0048671 | negative regulation of collateral sprouting(GO:0048671) |
| 0.1 | 0.2 | GO:0030311 | poly-N-acetyllactosamine metabolic process(GO:0030309) poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.1 | 0.9 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.1 | 2.5 | GO:0045725 | positive regulation of glycogen biosynthetic process(GO:0045725) |
| 0.1 | 0.6 | GO:0030854 | positive regulation of granulocyte differentiation(GO:0030854) |
| 0.0 | 0.4 | GO:0042590 | antigen processing and presentation of exogenous peptide antigen via MHC class I(GO:0042590) |
| 0.0 | 0.5 | GO:0071493 | cellular response to UV-B(GO:0071493) |
| 0.0 | 0.1 | GO:0016078 | tRNA catabolic process(GO:0016078) |
| 0.0 | 0.7 | GO:0060134 | prepulse inhibition(GO:0060134) |
| 0.0 | 0.6 | GO:0006027 | glycosaminoglycan catabolic process(GO:0006027) |
| 0.0 | 1.6 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.0 | 0.4 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.0 | 0.4 | GO:0060346 | bone trabecula formation(GO:0060346) |
| 0.0 | 0.5 | GO:0090050 | positive regulation of cell migration involved in sprouting angiogenesis(GO:0090050) |
| 0.0 | 0.4 | GO:0071073 | positive regulation of phospholipid biosynthetic process(GO:0071073) |
| 0.0 | 0.2 | GO:1903070 | negative regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903070) |
| 0.0 | 1.1 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 1.4 | GO:0060259 | regulation of feeding behavior(GO:0060259) |
| 0.0 | 0.4 | GO:0045759 | negative regulation of action potential(GO:0045759) |
| 0.0 | 4.4 | GO:0060078 | regulation of postsynaptic membrane potential(GO:0060078) |
| 0.0 | 0.3 | GO:0036371 | protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) |
| 0.0 | 3.5 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.0 | 1.8 | GO:0035115 | embryonic forelimb morphogenesis(GO:0035115) |
| 0.0 | 8.0 | GO:0007219 | Notch signaling pathway(GO:0007219) |
| 0.0 | 2.2 | GO:0042771 | intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator(GO:0042771) |
| 0.0 | 0.6 | GO:0046498 | S-adenosylhomocysteine metabolic process(GO:0046498) |
| 0.0 | 4.1 | GO:0001942 | hair follicle development(GO:0001942) |
| 0.0 | 2.2 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.0 | 1.3 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.0 | 0.3 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.0 | 2.0 | GO:0046513 | ceramide biosynthetic process(GO:0046513) |
| 0.0 | 0.2 | GO:1900028 | negative regulation of ruffle assembly(GO:1900028) |
| 0.0 | 3.1 | GO:1900006 | positive regulation of dendrite development(GO:1900006) |
| 0.0 | 0.3 | GO:0002024 | diet induced thermogenesis(GO:0002024) |
| 0.0 | 8.2 | GO:0031032 | actomyosin structure organization(GO:0031032) |
| 0.0 | 0.8 | GO:0031114 | negative regulation of microtubule depolymerization(GO:0007026) regulation of microtubule depolymerization(GO:0031114) |
| 0.0 | 1.1 | GO:1902042 | negative regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902042) |
| 0.0 | 0.5 | GO:0051382 | kinetochore assembly(GO:0051382) |
| 0.0 | 4.8 | GO:0002244 | hematopoietic progenitor cell differentiation(GO:0002244) |
| 0.0 | 2.1 | GO:0045582 | positive regulation of T cell differentiation(GO:0045582) |
| 0.0 | 2.4 | GO:0042107 | cytokine metabolic process(GO:0042107) |
| 0.0 | 0.3 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.0 | 1.6 | GO:0031397 | negative regulation of protein ubiquitination(GO:0031397) |
| 0.0 | 5.1 | GO:0050804 | modulation of synaptic transmission(GO:0050804) |
| 0.0 | 0.6 | GO:0030835 | negative regulation of actin filament depolymerization(GO:0030835) |
| 0.0 | 0.3 | GO:0009191 | ribonucleoside diphosphate catabolic process(GO:0009191) |
| 0.0 | 6.3 | GO:0016042 | lipid catabolic process(GO:0016042) |
| 0.0 | 0.3 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 0.0 | 0.6 | GO:0009250 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.0 | 1.6 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.0 | 1.0 | GO:0046488 | phosphatidylinositol metabolic process(GO:0046488) |
| 0.0 | 0.5 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.0 | 1.9 | GO:0010212 | response to ionizing radiation(GO:0010212) |
| 0.0 | 2.3 | GO:0008654 | phospholipid biosynthetic process(GO:0008654) |
| 0.0 | 0.2 | GO:1990001 | inhibition of cysteine-type endopeptidase activity involved in apoptotic process(GO:1990001) |
| 0.0 | 0.1 | GO:2000860 | positive regulation of mineralocorticoid secretion(GO:2000857) positive regulation of aldosterone secretion(GO:2000860) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 19.5 | GO:1990761 | growth cone lamellipodium(GO:1990761) growth cone filopodium(GO:1990812) |
| 1.0 | 3.0 | GO:1990031 | pinceau fiber(GO:1990031) |
| 0.9 | 4.3 | GO:0044326 | dendritic spine neck(GO:0044326) |
| 0.8 | 3.3 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.8 | 13.0 | GO:0043203 | axon hillock(GO:0043203) |
| 0.7 | 2.8 | GO:0032280 | symmetric synapse(GO:0032280) |
| 0.4 | 3.4 | GO:0001652 | granular component(GO:0001652) |
| 0.4 | 2.5 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.4 | 1.8 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.3 | 1.0 | GO:0070877 | microprocessor complex(GO:0070877) ribonuclease III complex(GO:1903095) |
| 0.3 | 2.2 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 0.3 | 5.0 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.3 | 1.7 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.3 | 3.2 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.2 | 0.7 | GO:0000811 | GINS complex(GO:0000811) |
| 0.2 | 1.5 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.2 | 2.6 | GO:0031315 | extrinsic component of mitochondrial outer membrane(GO:0031315) |
| 0.2 | 1.0 | GO:0061673 | mitotic spindle astral microtubule(GO:0061673) |
| 0.2 | 5.5 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.2 | 2.8 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.2 | 6.3 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.1 | 7.8 | GO:0032809 | neuronal cell body membrane(GO:0032809) |
| 0.1 | 0.8 | GO:0008091 | spectrin(GO:0008091) |
| 0.1 | 2.7 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.1 | 0.6 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.1 | 5.8 | GO:0030660 | Golgi-associated vesicle membrane(GO:0030660) |
| 0.1 | 1.0 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.1 | 3.3 | GO:0032590 | dendrite membrane(GO:0032590) |
| 0.1 | 0.5 | GO:0048476 | Holliday junction resolvase complex(GO:0048476) |
| 0.1 | 1.1 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.1 | 1.5 | GO:0001527 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.1 | 2.9 | GO:0030057 | desmosome(GO:0030057) |
| 0.1 | 3.4 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.1 | 1.5 | GO:0033268 | node of Ranvier(GO:0033268) |
| 0.1 | 4.4 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 10.1 | GO:0030315 | T-tubule(GO:0030315) |
| 0.1 | 2.2 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.1 | 1.8 | GO:0035267 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.1 | 8.5 | GO:0022626 | cytosolic ribosome(GO:0022626) |
| 0.1 | 0.4 | GO:0034750 | Scrib-APC-beta-catenin complex(GO:0034750) |
| 0.1 | 0.6 | GO:0042587 | glycogen granule(GO:0042587) |
| 0.1 | 3.8 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.1 | 2.3 | GO:0005921 | gap junction(GO:0005921) |
| 0.1 | 0.7 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.0 | 0.5 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 3.7 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 1.0 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 0.9 | GO:0005639 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.0 | 0.1 | GO:0005879 | axonemal microtubule(GO:0005879) |
| 0.0 | 2.0 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 0.3 | GO:0036396 | MIS complex(GO:0036396) |
| 0.0 | 10.9 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 0.7 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.0 | 2.9 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 0.2 | GO:0032444 | activin responsive factor complex(GO:0032444) SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.0 | 0.4 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.0 | 4.7 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 1.4 | GO:0016529 | sarcoplasmic reticulum(GO:0016529) |
| 0.0 | 0.9 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 2.2 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 12.3 | GO:0060076 | excitatory synapse(GO:0060076) |
| 0.0 | 3.0 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 0.1 | GO:0097651 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) phosphatidylinositol 3-kinase complex, class I(GO:0097651) |
| 0.0 | 0.3 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.0 | 0.4 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.0 | 2.2 | GO:0043197 | dendritic spine(GO:0043197) |
| 0.0 | 0.1 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.0 | 3.3 | GO:0043679 | axon terminus(GO:0043679) |
| 0.0 | 2.2 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 1.6 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 0.5 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.0 | 0.3 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.0 | 2.6 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 0.7 | GO:0005762 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 1.5 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.0 | 0.6 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 3.7 | GO:0047192 | 1-alkylglycerophosphocholine O-acetyltransferase activity(GO:0047192) |
| 1.2 | 3.6 | GO:0099580 | ion antiporter activity involved in regulation of postsynaptic membrane potential(GO:0099580) |
| 0.9 | 6.3 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 0.8 | 2.5 | GO:0008107 | galactoside 2-alpha-L-fucosyltransferase activity(GO:0008107) alpha-(1,2)-fucosyltransferase activity(GO:0031127) |
| 0.8 | 2.5 | GO:0004686 | elongation factor-2 kinase activity(GO:0004686) |
| 0.7 | 6.7 | GO:0019960 | C-X3-C chemokine binding(GO:0019960) |
| 0.7 | 5.5 | GO:1904315 | transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.6 | 4.7 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.6 | 19.3 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.5 | 3.3 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.5 | 16.5 | GO:0048156 | tau protein binding(GO:0048156) |
| 0.5 | 2.0 | GO:0004909 | interleukin-1, Type I, activating receptor activity(GO:0004909) |
| 0.4 | 2.6 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 0.4 | 1.2 | GO:0043183 | vascular endothelial growth factor receptor 1 binding(GO:0043183) |
| 0.4 | 3.0 | GO:0051425 | PTB domain binding(GO:0051425) |
| 0.4 | 4.8 | GO:0030235 | nitric-oxide synthase regulator activity(GO:0030235) |
| 0.4 | 4.3 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.3 | 1.4 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.3 | 3.3 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.3 | 1.0 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.3 | 0.9 | GO:0008988 | rRNA (adenine-N6,N6-)-dimethyltransferase activity(GO:0000179) rRNA (adenine-N6-)-methyltransferase activity(GO:0008988) |
| 0.3 | 1.8 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.3 | 3.8 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.3 | 2.9 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.3 | 1.9 | GO:0047179 | platelet-activating factor acetyltransferase activity(GO:0047179) |
| 0.2 | 0.7 | GO:0004822 | isoleucine-tRNA ligase activity(GO:0004822) |
| 0.2 | 1.1 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.2 | 1.3 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.2 | 2.6 | GO:0004439 | phosphatidylinositol-4,5-bisphosphate 5-phosphatase activity(GO:0004439) |
| 0.2 | 1.0 | GO:0004716 | receptor signaling protein tyrosine kinase activity(GO:0004716) |
| 0.2 | 2.0 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.2 | 1.0 | GO:0004525 | ribonuclease III activity(GO:0004525) double-stranded RNA-specific ribonuclease activity(GO:0032296) |
| 0.2 | 1.3 | GO:0061629 | RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 0.2 | 1.1 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.2 | 1.0 | GO:0030171 | voltage-gated proton channel activity(GO:0030171) |
| 0.2 | 0.6 | GO:0031720 | haptoglobin binding(GO:0031720) hemoglobin alpha binding(GO:0031721) |
| 0.1 | 5.4 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.1 | 5.7 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.1 | 2.9 | GO:0022841 | potassium ion leak channel activity(GO:0022841) |
| 0.1 | 6.0 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.1 | 3.2 | GO:0099604 | calcium-release channel activity(GO:0015278) ligand-gated calcium channel activity(GO:0099604) |
| 0.1 | 4.5 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.1 | 5.5 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.1 | 1.4 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 0.1 | 1.5 | GO:0051430 | corticotropin-releasing hormone receptor 1 binding(GO:0051430) |
| 0.1 | 0.6 | GO:0004719 | protein-L-isoaspartate (D-aspartate) O-methyltransferase activity(GO:0004719) |
| 0.1 | 0.3 | GO:0003881 | CDP-diacylglycerol-inositol 3-phosphatidyltransferase activity(GO:0003881) |
| 0.1 | 3.0 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.1 | 1.0 | GO:0016308 | 1-phosphatidylinositol-4-phosphate 5-kinase activity(GO:0016308) |
| 0.1 | 0.2 | GO:0003986 | acetyl-CoA hydrolase activity(GO:0003986) |
| 0.1 | 2.4 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.1 | 0.9 | GO:0035381 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.1 | 7.4 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.1 | 1.1 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.1 | 2.9 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.1 | 0.4 | GO:0004949 | cannabinoid receptor activity(GO:0004949) |
| 0.1 | 0.5 | GO:0017108 | 5'-flap endonuclease activity(GO:0017108) |
| 0.1 | 3.1 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.1 | 0.7 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.1 | 3.0 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen(GO:0016701) oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.1 | 0.8 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.1 | 1.5 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.1 | 0.3 | GO:0000702 | oxidized base lesion DNA N-glycosylase activity(GO:0000702) |
| 0.1 | 7.5 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.1 | 1.1 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 1.6 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.1 | 2.0 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.1 | 3.2 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.1 | 0.2 | GO:0016262 | protein N-acetylglucosaminyltransferase activity(GO:0016262) |
| 0.1 | 4.8 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.1 | 3.1 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.1 | 0.5 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.1 | 0.5 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.1 | 0.4 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.1 | 1.4 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 2.4 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.1 | 7.4 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 0.2 | GO:0030618 | transforming growth factor beta receptor, pathway-specific cytoplasmic mediator activity(GO:0030618) |
| 0.1 | 1.3 | GO:0097371 | MDM2/MDM4 family protein binding(GO:0097371) |
| 0.1 | 0.7 | GO:0030976 | thiamine pyrophosphate binding(GO:0030976) |
| 0.1 | 1.9 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.1 | 0.3 | GO:0004382 | guanosine-diphosphatase activity(GO:0004382) |
| 0.1 | 1.0 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
| 0.0 | 0.1 | GO:0000171 | ribonuclease MRP activity(GO:0000171) |
| 0.0 | 1.3 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 0.7 | GO:0043138 | 3'-5' DNA helicase activity(GO:0043138) |
| 0.0 | 0.6 | GO:0003906 | DNA-(apurinic or apyrimidinic site) lyase activity(GO:0003906) |
| 0.0 | 2.2 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 2.5 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 1.5 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.0 | 2.2 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 0.2 | GO:1904288 | BAT3 complex binding(GO:1904288) |
| 0.0 | 0.4 | GO:0016595 | glutamate binding(GO:0016595) |
| 0.0 | 0.8 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.0 | 0.4 | GO:0001055 | RNA polymerase I activity(GO:0001054) RNA polymerase II activity(GO:0001055) |
| 0.0 | 0.5 | GO:0008301 | AT DNA binding(GO:0003680) DNA binding, bending(GO:0008301) |
| 0.0 | 3.7 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.0 | 0.2 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.0 | 1.4 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 2.2 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.0 | 2.6 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.0 | 3.8 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 1.0 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 2.2 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.0 | 2.2 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.0 | 1.0 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.0 | 2.3 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.0 | 2.8 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
| 0.0 | 1.4 | GO:0043621 | protein self-association(GO:0043621) |
| 0.0 | 1.4 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 0.1 | GO:0008169 | C-methyltransferase activity(GO:0008169) |
| 0.0 | 0.1 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 2.3 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 2.2 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 0.3 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.0 | 0.4 | GO:0015036 | disulfide oxidoreductase activity(GO:0015036) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 7.7 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.2 | 2.7 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.1 | 18.4 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.1 | 2.5 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.1 | 1.2 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.1 | 3.0 | PID S1P S1P1 PATHWAY | S1P1 pathway |
| 0.1 | 3.3 | PID MYC PATHWAY | C-MYC pathway |
| 0.1 | 3.1 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 3.2 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.1 | 4.4 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.1 | 14.5 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 2.1 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.1 | 1.9 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.1 | 0.7 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.1 | 1.0 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.1 | 1.5 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.0 | 0.4 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 2.2 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 2.5 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 1.1 | PID ERBB1 RECEPTOR PROXIMAL PATHWAY | EGF receptor (ErbB1) signaling pathway |
| 0.0 | 1.0 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 1.0 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 2.4 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 1.0 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.0 | 1.1 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 1.5 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 0.4 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.0 | 0.9 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.9 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 1.3 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 1.5 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 1.1 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.0 | 1.3 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.0 | 0.6 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 6.3 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.2 | 5.5 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.2 | 3.8 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.2 | 3.0 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.2 | 2.9 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.2 | 3.8 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.2 | 3.6 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.1 | 3.5 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.1 | 2.5 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.1 | 1.2 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.1 | 6.7 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 17.4 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.1 | 4.8 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 5.9 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 1.6 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.1 | 1.4 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.1 | 4.3 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.1 | 3.8 | REACTOME PROSTACYCLIN SIGNALLING THROUGH PROSTACYCLIN RECEPTOR | Genes involved in Prostacyclin signalling through prostacyclin receptor |
| 0.1 | 2.4 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.1 | 1.6 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 0.1 | 6.2 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.1 | 2.7 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.1 | 1.0 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.1 | 2.8 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.1 | 1.0 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.0 | 0.6 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.0 | 2.2 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 2.5 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 2.0 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.5 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.0 | 1.8 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 2.0 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 0.7 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.0 | 1.8 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 1.0 | REACTOME THE ROLE OF NEF IN HIV1 REPLICATION AND DISEASE PATHOGENESIS | Genes involved in The role of Nef in HIV-1 replication and disease pathogenesis |
| 0.0 | 0.8 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.8 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 1.5 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.0 | 0.3 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.0 | 1.9 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.0 | 3.1 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 3.5 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.0 | 0.3 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.0 | 0.7 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 0.7 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.0 | 0.4 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.0 | 0.5 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.0 | 0.7 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.0 | 0.3 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 0.4 | REACTOME VIRAL MESSENGER RNA SYNTHESIS | Genes involved in Viral Messenger RNA Synthesis |
| 0.0 | 2.6 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 0.5 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 1.5 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 1.9 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 0.4 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.3 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 0.4 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |