Project
PRJNA375882: Comprehensive Mouse Transcriptomic BodyMap
Navigation
Downloads
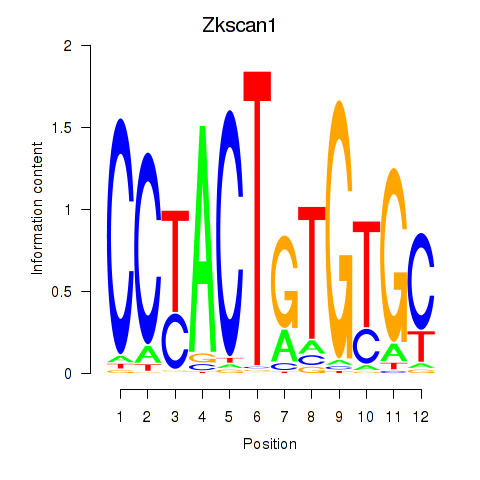
Results for Zkscan1
Z-value: 0.84
Transcription factors associated with Zkscan1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Zkscan1
|
ENSMUSG00000029729.13 | Zkscan1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Zkscan1 | mm39_v1_chr5_+_138083345_138083410 | 0.07 | 5.4e-01 | Click! |
Activity profile of Zkscan1 motif
Sorted Z-values of Zkscan1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Zkscan1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_+_75027089 | 5.63 |
ENSMUST00000190262.2
|
Ly6g
|
lymphocyte antigen 6 complex, locus G |
| chr17_-_33937565 | 5.15 |
ENSMUST00000174040.2
ENSMUST00000173015.8 ENSMUST00000066121.13 ENSMUST00000186022.7 ENSMUST00000173329.8 ENSMUST00000172767.9 |
Marchf2
|
membrane associated ring-CH-type finger 2 |
| chr19_-_61216834 | 4.90 |
ENSMUST00000076046.7
|
Csf2ra
|
colony stimulating factor 2 receptor, alpha, low-affinity (granulocyte-macrophage) |
| chr10_+_79716876 | 4.58 |
ENSMUST00000166201.2
|
Prtn3
|
proteinase 3 |
| chr3_-_88291740 | 4.57 |
ENSMUST00000076048.5
|
Bglap
|
bone gamma carboxyglutamate protein |
| chr8_-_72178340 | 4.40 |
ENSMUST00000153800.8
ENSMUST00000146100.8 |
Fcho1
|
FCH domain only 1 |
| chr3_-_88286006 | 4.40 |
ENSMUST00000098956.3
|
Bglap2
|
bone gamma-carboxyglutamate protein 2 |
| chr15_-_74983786 | 3.79 |
ENSMUST00000191451.2
ENSMUST00000100542.10 |
Ly6c2
|
lymphocyte antigen 6 complex, locus C2 |
| chr2_+_31135813 | 3.77 |
ENSMUST00000000199.8
|
Ncs1
|
neuronal calcium sensor 1 |
| chr17_+_44114894 | 3.66 |
ENSMUST00000044895.13
|
Rcan2
|
regulator of calcineurin 2 |
| chr15_-_82505132 | 3.30 |
ENSMUST00000109515.3
|
Cyp2d34
|
cytochrome P450, family 2, subfamily d, polypeptide 34 |
| chr2_+_84564394 | 3.14 |
ENSMUST00000238573.2
ENSMUST00000090729.9 |
Ypel4
|
yippee like 4 |
| chr15_+_83664196 | 2.95 |
ENSMUST00000046168.12
|
Mpped1
|
metallophosphoesterase domain containing 1 |
| chr12_-_76865148 | 2.94 |
ENSMUST00000118604.8
|
Rab15
|
RAB15, member RAS oncogene family |
| chr12_+_112586465 | 2.89 |
ENSMUST00000021726.8
|
Adssl1
|
adenylosuccinate synthetase like 1 |
| chr6_-_126717590 | 2.89 |
ENSMUST00000185333.2
|
Kcna6
|
potassium voltage-gated channel, shaker-related, subfamily, member 6 |
| chr6_-_126717114 | 2.83 |
ENSMUST00000112242.2
|
Kcna6
|
potassium voltage-gated channel, shaker-related, subfamily, member 6 |
| chr8_+_73373304 | 2.83 |
ENSMUST00000093427.11
|
Nwd1
|
NACHT and WD repeat domain containing 1 |
| chr8_+_73373356 | 2.80 |
ENSMUST00000161254.9
|
Nwd1
|
NACHT and WD repeat domain containing 1 |
| chr11_-_61344818 | 2.74 |
ENSMUST00000060255.14
ENSMUST00000054927.14 ENSMUST00000102661.4 |
Rnf112
|
ring finger protein 112 |
| chr10_+_126836578 | 2.60 |
ENSMUST00000026500.12
ENSMUST00000142698.8 |
Avil
|
advillin |
| chr12_+_112586501 | 2.58 |
ENSMUST00000180015.9
|
Adssl1
|
adenylosuccinate synthetase like 1 |
| chr15_-_74855264 | 2.48 |
ENSMUST00000023250.11
ENSMUST00000166694.2 |
Ly6i
|
lymphocyte antigen 6 complex, locus I |
| chr5_-_8417982 | 2.41 |
ENSMUST00000088761.11
ENSMUST00000115386.8 ENSMUST00000050166.14 ENSMUST00000046838.14 ENSMUST00000115388.9 ENSMUST00000088744.12 ENSMUST00000115385.2 |
Adam22
|
a disintegrin and metallopeptidase domain 22 |
| chr15_-_79048674 | 2.39 |
ENSMUST00000230261.2
ENSMUST00000040019.5 |
Sox10
|
SRY (sex determining region Y)-box 10 |
| chr15_+_83664175 | 2.36 |
ENSMUST00000163723.8
|
Mpped1
|
metallophosphoesterase domain containing 1 |
| chr4_+_42922253 | 2.33 |
ENSMUST00000139100.2
|
Phf24
|
PHD finger protein 24 |
| chr11_+_42310557 | 2.31 |
ENSMUST00000007797.10
|
Gabrb2
|
gamma-aminobutyric acid (GABA) A receptor, subunit beta 2 |
| chr9_-_103357564 | 2.07 |
ENSMUST00000124310.5
|
Bfsp2
|
beaded filament structural protein 2, phakinin |
| chr12_-_79054050 | 2.02 |
ENSMUST00000056660.13
ENSMUST00000174721.8 |
Tmem229b
|
transmembrane protein 229B |
| chr15_-_74920518 | 2.00 |
ENSMUST00000185372.2
ENSMUST00000187347.7 ENSMUST00000188845.7 ENSMUST00000185200.7 ENSMUST00000179762.8 ENSMUST00000191216.7 ENSMUST00000065408.16 |
Ly6c1
|
lymphocyte antigen 6 complex, locus C1 |
| chr7_-_97811525 | 1.98 |
ENSMUST00000107112.2
|
Capn5
|
calpain 5 |
| chr10_-_81360059 | 1.97 |
ENSMUST00000043709.8
|
Gna15
|
guanine nucleotide binding protein, alpha 15 |
| chr7_+_43959637 | 1.86 |
ENSMUST00000107938.8
|
Shank1
|
SH3 and multiple ankyrin repeat domains 1 |
| chr9_+_106245792 | 1.86 |
ENSMUST00000172306.3
|
Dusp7
|
dual specificity phosphatase 7 |
| chr11_+_104468107 | 1.84 |
ENSMUST00000106956.10
|
Myl4
|
myosin, light polypeptide 4 |
| chr17_+_34258411 | 1.77 |
ENSMUST00000087497.11
ENSMUST00000131134.9 ENSMUST00000235819.2 ENSMUST00000114255.9 ENSMUST00000114252.9 ENSMUST00000237989.2 |
Col11a2
|
collagen, type XI, alpha 2 |
| chr2_-_91795910 | 1.73 |
ENSMUST00000239257.2
|
Dgkz
|
diacylglycerol kinase zeta |
| chr9_+_118881838 | 1.63 |
ENSMUST00000051386.13
ENSMUST00000074734.13 |
Vill
|
villin-like |
| chr7_+_28834276 | 1.58 |
ENSMUST00000161522.8
ENSMUST00000204845.3 ENSMUST00000205027.3 ENSMUST00000204194.3 ENSMUST00000203070.3 ENSMUST00000203380.3 |
Rasgrp4
|
RAS guanyl releasing protein 4 |
| chr18_-_80133177 | 1.54 |
ENSMUST00000178391.2
|
Gm21886
|
predicted gene, 21886 |
| chr9_+_62249341 | 1.53 |
ENSMUST00000135395.8
|
Anp32a
|
acidic (leucine-rich) nuclear phosphoprotein 32 family, member A |
| chr15_-_82291372 | 1.51 |
ENSMUST00000230198.2
ENSMUST00000230248.2 ENSMUST00000072776.5 ENSMUST00000229911.2 |
Cyp2d10
|
cytochrome P450, family 2, subfamily d, polypeptide 10 |
| chr9_-_106533279 | 1.50 |
ENSMUST00000023959.13
ENSMUST00000201681.2 |
Grm2
|
glutamate receptor, metabotropic 2 |
| chr5_+_110248276 | 1.50 |
ENSMUST00000141066.8
|
Plcxd1
|
phosphatidylinositol-specific phospholipase C, X domain containing 1 |
| chr7_+_111825063 | 1.47 |
ENSMUST00000050149.12
|
Mical2
|
microtubule associated monooxygenase, calponin and LIM domain containing 2 |
| chr10_+_90665270 | 1.44 |
ENSMUST00000182202.8
ENSMUST00000182966.8 |
Anks1b
|
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr4_-_155870321 | 1.43 |
ENSMUST00000097742.3
|
Tmem88b
|
transmembrane protein 88B |
| chr15_-_78456898 | 1.39 |
ENSMUST00000043214.8
|
Rac2
|
Rac family small GTPase 2 |
| chr16_-_17019352 | 1.33 |
ENSMUST00000090192.12
|
Ube2l3
|
ubiquitin-conjugating enzyme E2L 3 |
| chr7_-_126399778 | 1.32 |
ENSMUST00000141355.4
|
Aldoa
|
aldolase A, fructose-bisphosphate |
| chr10_+_90665639 | 1.29 |
ENSMUST00000179337.9
|
Anks1b
|
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr10_+_90665399 | 1.28 |
ENSMUST00000179694.9
|
Anks1b
|
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr2_-_167032068 | 1.27 |
ENSMUST00000059826.10
|
Kcnb1
|
potassium voltage gated channel, Shab-related subfamily, member 1 |
| chr6_-_144993451 | 1.26 |
ENSMUST00000123930.8
|
Bcat1
|
branched chain aminotransferase 1, cytosolic |
| chr1_+_171668173 | 1.26 |
ENSMUST00000136479.8
|
Cd84
|
CD84 antigen |
| chr18_-_61147272 | 1.26 |
ENSMUST00000025520.10
|
Slc6a7
|
solute carrier family 6 (neurotransmitter transporter, L-proline), member 7 |
| chr9_+_51124983 | 1.24 |
ENSMUST00000034554.9
|
Pou2af1
|
POU domain, class 2, associating factor 1 |
| chr2_+_172235702 | 1.20 |
ENSMUST00000099061.9
ENSMUST00000103073.9 |
Cass4
|
Cas scaffolding protein family member 4 |
| chr15_+_16778187 | 1.18 |
ENSMUST00000026432.8
|
Cdh9
|
cadherin 9 |
| chr2_-_104241358 | 1.17 |
ENSMUST00000230671.2
|
D430041D05Rik
|
RIKEN cDNA D430041D05 gene |
| chr4_+_20042045 | 1.16 |
ENSMUST00000098242.4
|
Ggh
|
gamma-glutamyl hydrolase |
| chrX_-_48886577 | 1.13 |
ENSMUST00000033442.14
ENSMUST00000114891.2 |
Igsf1
|
immunoglobulin superfamily, member 1 |
| chr15_+_82439273 | 1.13 |
ENSMUST00000229103.2
ENSMUST00000068861.8 ENSMUST00000229904.2 |
Cyp2d12
|
cytochrome P450, family 2, subfamily d, polypeptide 12 |
| chr10_+_80134917 | 1.12 |
ENSMUST00000154212.8
|
Apc2
|
APC regulator of WNT signaling pathway 2 |
| chr11_+_19874354 | 1.11 |
ENSMUST00000093299.13
|
Spred2
|
sprouty-related EVH1 domain containing 2 |
| chr5_+_125518609 | 1.10 |
ENSMUST00000049040.14
|
Bri3bp
|
Bri3 binding protein |
| chr15_-_74869684 | 1.10 |
ENSMUST00000190188.2
ENSMUST00000189068.7 ENSMUST00000186526.7 ENSMUST00000187171.2 ENSMUST00000187994.7 |
Ly6a
|
lymphocyte antigen 6 complex, locus A |
| chr9_+_62249177 | 1.07 |
ENSMUST00000128636.8
|
Anp32a
|
acidic (leucine-rich) nuclear phosphoprotein 32 family, member A |
| chr6_-_144993362 | 1.06 |
ENSMUST00000149769.6
|
Bcat1
|
branched chain aminotransferase 1, cytosolic |
| chr2_-_106804679 | 1.03 |
ENSMUST00000028536.13
|
Arl14ep
|
ADP-ribosylation factor-like 14 effector protein |
| chr16_-_10265204 | 1.01 |
ENSMUST00000051118.7
|
Tvp23a
|
trans-golgi network vesicle protein 23A |
| chr9_+_62248611 | 1.00 |
ENSMUST00000085519.13
|
Anp32a
|
acidic (leucine-rich) nuclear phosphoprotein 32 family, member A |
| chr6_+_57234937 | 1.00 |
ENSMUST00000228297.2
|
Vmn1r15
|
vomeronasal 1 receptor 15 |
| chr3_+_62245765 | 0.96 |
ENSMUST00000079300.13
|
Arhgef26
|
Rho guanine nucleotide exchange factor (GEF) 26 |
| chr5_-_131567526 | 0.96 |
ENSMUST00000161374.8
|
Auts2
|
autism susceptibility candidate 2 |
| chr1_+_50966670 | 0.94 |
ENSMUST00000081851.4
|
Tmeff2
|
transmembrane protein with EGF-like and two follistatin-like domains 2 |
| chr4_-_97472844 | 0.94 |
ENSMUST00000107067.8
ENSMUST00000107068.9 |
E130114P18Rik
|
RIKEN cDNA E130114P18 gene |
| chr17_+_47604995 | 0.93 |
ENSMUST00000190020.4
|
Trerf1
|
transcriptional regulating factor 1 |
| chr4_-_140805613 | 0.92 |
ENSMUST00000030760.15
|
Necap2
|
NECAP endocytosis associated 2 |
| chr11_-_47270201 | 0.91 |
ENSMUST00000077221.6
|
Sgcd
|
sarcoglycan, delta (dystrophin-associated glycoprotein) |
| chr17_-_54153367 | 0.90 |
ENSMUST00000023886.7
|
Sult1c2
|
sulfotransferase family, cytosolic, 1C, member 2 |
| chr7_+_15909010 | 0.88 |
ENSMUST00000176342.8
ENSMUST00000177540.8 |
Meis3
|
Meis homeobox 3 |
| chr12_-_101943134 | 0.84 |
ENSMUST00000221227.2
|
Ndufb1
|
NADH:ubiquinone oxidoreductase subunit B1 |
| chr15_-_74869483 | 0.84 |
ENSMUST00000023248.13
|
Ly6a
|
lymphocyte antigen 6 complex, locus A |
| chr10_-_79624758 | 0.82 |
ENSMUST00000020573.13
|
Prss57
|
protease, serine 57 |
| chr19_-_10502468 | 0.81 |
ENSMUST00000025570.8
ENSMUST00000236455.2 |
Sdhaf2
|
succinate dehydrogenase complex assembly factor 2 |
| chr17_+_28794615 | 0.81 |
ENSMUST00000232862.2
ENSMUST00000080780.8 |
Lhfpl5
|
lipoma HMGIC fusion partner-like 5 |
| chr18_-_31742946 | 0.80 |
ENSMUST00000060396.7
|
Slc25a46
|
solute carrier family 25, member 46 |
| chr12_-_17061545 | 0.78 |
ENSMUST00000190691.2
|
2410004P03Rik
|
RIKEN cDNA 2410004P03 gene |
| chr2_+_172235820 | 0.76 |
ENSMUST00000109136.3
ENSMUST00000228775.2 |
Cass4
|
Cas scaffolding protein family member 4 |
| chr5_+_64969679 | 0.73 |
ENSMUST00000166409.6
ENSMUST00000197879.2 |
Klf3
|
Kruppel-like factor 3 (basic) |
| chr4_+_73931679 | 0.71 |
ENSMUST00000098006.9
ENSMUST00000084474.6 |
Frmd3
|
FERM domain containing 3 |
| chr18_+_61238908 | 0.71 |
ENSMUST00000115268.4
|
Csf1r
|
colony stimulating factor 1 receptor |
| chr4_+_103476777 | 0.70 |
ENSMUST00000106827.8
|
Dab1
|
disabled 1 |
| chr7_-_132415257 | 0.70 |
ENSMUST00000097999.9
|
Fam53b
|
family with sequence similarity 53, member B |
| chr19_-_10502546 | 0.68 |
ENSMUST00000237827.2
|
Sdhaf2
|
succinate dehydrogenase complex assembly factor 2 |
| chr4_+_101276474 | 0.68 |
ENSMUST00000102780.8
ENSMUST00000106946.8 ENSMUST00000106945.8 |
Ak4
|
adenylate kinase 4 |
| chr11_+_43046476 | 0.67 |
ENSMUST00000238415.2
|
Atp10b
|
ATPase, class V, type 10B |
| chr2_-_121381832 | 0.66 |
ENSMUST00000212518.2
|
Frmd5
|
FERM domain containing 5 |
| chr4_-_134642286 | 0.66 |
ENSMUST00000105863.2
ENSMUST00000030626.12 |
Tmem50a
|
transmembrane protein 50A |
| chr11_+_19874403 | 0.62 |
ENSMUST00000093298.12
|
Spred2
|
sprouty-related EVH1 domain containing 2 |
| chr1_-_57008986 | 0.61 |
ENSMUST00000176759.2
ENSMUST00000177424.2 |
Satb2
|
special AT-rich sequence binding protein 2 |
| chr2_-_127571836 | 0.61 |
ENSMUST00000028856.3
|
Mall
|
mal, T cell differentiation protein-like |
| chr4_+_126156118 | 0.58 |
ENSMUST00000030660.9
|
Trappc3
|
trafficking protein particle complex 3 |
| chr8_+_85676489 | 0.58 |
ENSMUST00000121880.8
|
Rtbdn
|
retbindin |
| chr17_+_29042640 | 0.55 |
ENSMUST00000233088.2
ENSMUST00000233182.2 ENSMUST00000233520.2 |
Brpf3
|
bromodomain and PHD finger containing, 3 |
| chr2_+_134627987 | 0.55 |
ENSMUST00000131552.5
ENSMUST00000110116.8 |
Plcb1
|
phospholipase C, beta 1 |
| chr8_-_4375320 | 0.54 |
ENSMUST00000098950.6
|
Elavl1
|
ELAV (embryonic lethal, abnormal vision)-like 1 (Hu antigen R) |
| chr17_+_33743144 | 0.52 |
ENSMUST00000087623.13
ENSMUST00000234715.2 ENSMUST00000234497.2 |
Adamts10
|
a disintegrin-like and metallopeptidase (reprolysin type) with thrombospondin type 1 motif, 10 |
| chr11_+_53992054 | 0.51 |
ENSMUST00000135653.8
|
P4ha2
|
procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), alpha II polypeptide |
| chr5_-_134668152 | 0.50 |
ENSMUST00000036125.10
ENSMUST00000202622.4 |
Eif4h
|
eukaryotic translation initiation factor 4H |
| chr8_-_70169500 | 0.50 |
ENSMUST00000238408.2
ENSMUST00000238418.2 ENSMUST00000238753.2 |
Zfp869
|
zinc finger protein 869 |
| chr12_+_88327607 | 0.49 |
ENSMUST00000166940.3
|
Adck1
|
aarF domain containing kinase 1 |
| chr4_-_63414188 | 0.48 |
ENSMUST00000063650.10
ENSMUST00000102867.8 ENSMUST00000107393.8 ENSMUST00000084510.8 ENSMUST00000095038.8 ENSMUST00000119294.8 ENSMUST00000095037.2 ENSMUST00000063672.10 |
Whrn
|
whirlin |
| chr8_+_95712151 | 0.47 |
ENSMUST00000212799.2
|
Adgrg1
|
adhesion G protein-coupled receptor G1 |
| chr4_+_141171965 | 0.46 |
ENSMUST00000006377.13
|
Zbtb17
|
zinc finger and BTB domain containing 17 |
| chr5_-_22549688 | 0.46 |
ENSMUST00000062372.14
ENSMUST00000161356.8 |
Reln
|
reelin |
| chr12_+_116369017 | 0.46 |
ENSMUST00000084828.5
ENSMUST00000222469.2 ENSMUST00000221114.2 ENSMUST00000221970.2 |
Ncapg2
|
non-SMC condensin II complex, subunit G2 |
| chr18_+_31742565 | 0.45 |
ENSMUST00000164667.2
|
B930094E09Rik
|
RIKEN cDNA B930094E09 gene |
| chr17_+_25105617 | 0.43 |
ENSMUST00000117890.8
ENSMUST00000168265.8 ENSMUST00000120943.8 ENSMUST00000068508.13 ENSMUST00000119829.8 |
Spsb3
|
splA/ryanodine receptor domain and SOCS box containing 3 |
| chr19_+_10502612 | 0.41 |
ENSMUST00000237321.2
ENSMUST00000038379.5 |
Cpsf7
|
cleavage and polyadenylation specific factor 7 |
| chr8_+_70980374 | 0.41 |
ENSMUST00000119353.9
ENSMUST00000075491.14 ENSMUST00000119698.8 |
Fkbp8
|
FK506 binding protein 8 |
| chrX_-_77627486 | 0.39 |
ENSMUST00000114025.8
ENSMUST00000134602.8 ENSMUST00000114024.9 |
Prrg1
|
proline rich Gla (G-carboxyglutamic acid) 1 |
| chr5_-_140307280 | 0.38 |
ENSMUST00000031534.9
|
Mad1l1
|
MAD1 mitotic arrest deficient 1-like 1 |
| chr16_+_43582584 | 0.37 |
ENSMUST00000023390.5
|
Drd3
|
dopamine receptor D3 |
| chr11_-_72380730 | 0.36 |
ENSMUST00000045303.10
|
Spns2
|
spinster homolog 2 |
| chr15_-_82278223 | 0.36 |
ENSMUST00000170255.2
|
Cyp2d11
|
cytochrome P450, family 2, subfamily d, polypeptide 11 |
| chr14_-_40728839 | 0.35 |
ENSMUST00000152837.2
ENSMUST00000134715.8 |
Prxl2a
|
peroxiredoxin like 2A |
| chr5_-_113369096 | 0.35 |
ENSMUST00000211733.2
|
2900026A02Rik
|
RIKEN cDNA 2900026A02 gene |
| chr8_+_4303067 | 0.35 |
ENSMUST00000011981.5
ENSMUST00000208316.2 |
Snapc2
|
small nuclear RNA activating complex, polypeptide 2 |
| chr13_-_56283331 | 0.35 |
ENSMUST00000045788.9
ENSMUST00000016081.13 |
Macroh2a1
|
macroH2A.1 histone |
| chrX_-_77627388 | 0.33 |
ENSMUST00000177904.8
|
Prrg1
|
proline rich Gla (G-carboxyglutamic acid) 1 |
| chr10_+_81648020 | 0.33 |
ENSMUST00000188096.7
|
Gm4767
|
predicted gene 4767 |
| chr6_-_57367651 | 0.32 |
ENSMUST00000164732.2
|
Vmn1r18
|
vomeronasal 1 receptor 18 |
| chr2_+_131048998 | 0.32 |
ENSMUST00000153097.3
|
Ap5s1
|
adaptor-related protein 5 complex, sigma 1 subunit |
| chr13_-_43713583 | 0.31 |
ENSMUST00000021800.6
|
Mcur1
|
mitochondrial calcium uniporter regulator 1 |
| chr11_-_102588536 | 0.30 |
ENSMUST00000164506.3
ENSMUST00000092569.13 |
Ccdc43
|
coiled-coil domain containing 43 |
| chr7_-_118091135 | 0.30 |
ENSMUST00000178344.3
|
Itpripl2
|
inositol 1,4,5-triphosphate receptor interacting protein-like 2 |
| chr1_-_93563406 | 0.28 |
ENSMUST00000027498.14
|
Stk25
|
serine/threonine kinase 25 (yeast) |
| chrX_+_158155171 | 0.27 |
ENSMUST00000087143.7
|
Eif1ax
|
eukaryotic translation initiation factor 1A, X-linked |
| chr10_+_81484858 | 0.24 |
ENSMUST00000189672.7
|
Gm10778
|
predicted gene 10778 |
| chr12_+_84332006 | 0.24 |
ENSMUST00000123614.8
ENSMUST00000147363.8 ENSMUST00000135001.8 ENSMUST00000146377.8 |
Ptgr2
|
prostaglandin reductase 2 |
| chr19_-_53932581 | 0.19 |
ENSMUST00000236885.2
ENSMUST00000236098.2 ENSMUST00000236370.2 |
Bbip1
|
BBSome interacting protein 1 |
| chr6_-_116050081 | 0.19 |
ENSMUST00000173548.3
|
Tmcc1
|
transmembrane and coiled coil domains 1 |
| chr19_+_10914517 | 0.19 |
ENSMUST00000037261.4
|
Ptgdr2
|
prostaglandin D2 receptor 2 |
| chr9_+_118881926 | 0.18 |
ENSMUST00000131647.2
|
Vill
|
villin-like |
| chr17_+_33395390 | 0.18 |
ENSMUST00000112168.4
|
Olfr55
|
olfactory receptor 55 |
| chr7_-_132415408 | 0.15 |
ENSMUST00000134946.3
|
Fam53b
|
family with sequence similarity 53, member B |
| chr16_-_17019321 | 0.15 |
ENSMUST00000232668.2
ENSMUST00000231643.2 ENSMUST00000232035.2 |
Ube2l3
|
ubiquitin-conjugating enzyme E2L 3 |
| chrX_+_55917903 | 0.13 |
ENSMUST00000153784.8
|
Adgrg4
|
adhesion G protein-coupled receptor G4 |
| chr4_-_57300750 | 0.13 |
ENSMUST00000151964.2
|
Ptpn3
|
protein tyrosine phosphatase, non-receptor type 3 |
| chr11_-_78056347 | 0.12 |
ENSMUST00000017530.4
|
Traf4
|
TNF receptor associated factor 4 |
| chr19_-_53933052 | 0.11 |
ENSMUST00000135402.4
|
Bbip1
|
BBSome interacting protein 1 |
| chr13_+_68745528 | 0.11 |
ENSMUST00000022007.8
|
1700001L19Rik
|
RIKEN cDNA 1700001L19 gene |
| chr19_+_44321531 | 0.11 |
ENSMUST00000058856.9
|
Scd4
|
stearoyl-coenzyme A desaturase 4 |
| chr12_-_72711509 | 0.10 |
ENSMUST00000221750.2
|
Dhrs7
|
dehydrogenase/reductase (SDR family) member 7 |
| chr4_-_57300361 | 0.10 |
ENSMUST00000153926.8
|
Ptpn3
|
protein tyrosine phosphatase, non-receptor type 3 |
| chr8_-_71315651 | 0.09 |
ENSMUST00000212414.2
|
Kcnn1
|
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 1 |
| chr11_+_120348919 | 0.09 |
ENSMUST00000058370.14
ENSMUST00000175970.8 ENSMUST00000176120.2 |
Ccdc137
|
coiled-coil domain containing 137 |
| chr5_+_30869193 | 0.09 |
ENSMUST00000088081.11
ENSMUST00000101442.4 |
Dpysl5
|
dihydropyrimidinase-like 5 |
| chr12_-_72711533 | 0.08 |
ENSMUST00000021512.11
|
Dhrs7
|
dehydrogenase/reductase (SDR family) member 7 |
| chr8_+_47193275 | 0.08 |
ENSMUST00000207571.3
|
Irf2
|
interferon regulatory factor 2 |
| chr10_+_81919146 | 0.08 |
ENSMUST00000220287.2
|
BC024063
|
cDNA sequence BC024063 |
| chr11_+_69871952 | 0.07 |
ENSMUST00000108593.8
|
Ctdnep1
|
CTD nuclear envelope phosphatase 1 |
| chr10_+_81540659 | 0.07 |
ENSMUST00000200889.4
ENSMUST00000201819.4 |
Zfp433
|
zinc finger protein 433 |
| chr19_-_53932867 | 0.06 |
ENSMUST00000235688.2
ENSMUST00000235348.2 |
Bbip1
|
BBSome interacting protein 1 |
| chr2_-_73410632 | 0.04 |
ENSMUST00000028515.4
|
Chrna1
|
cholinergic receptor, nicotinic, alpha polypeptide 1 (muscle) |
| chr19_+_28941292 | 0.03 |
ENSMUST00000045674.4
|
Plpp6
|
phospholipid phosphatase 6 |
| chr8_-_70980584 | 0.03 |
ENSMUST00000138260.8
ENSMUST00000117580.8 |
Kxd1
|
KxDL motif containing 1 |
| chr19_+_53298906 | 0.03 |
ENSMUST00000003870.15
|
Mxi1
|
MAX interactor 1, dimerization protein |
| chr12_-_17061504 | 0.03 |
ENSMUST00000189479.2
|
2410004P03Rik
|
RIKEN cDNA 2410004P03 gene |
| chr13_+_50571889 | 0.03 |
ENSMUST00000046974.5
|
Fbxw17
|
F-box and WD-40 domain protein 17 |
| chr2_+_118428690 | 0.03 |
ENSMUST00000038341.8
|
Bub1b
|
BUB1B, mitotic checkpoint serine/threonine kinase |
| chr1_+_90531183 | 0.03 |
ENSMUST00000186750.2
|
Cops8
|
COP9 signalosome subunit 8 |
| chr10_+_81699768 | 0.03 |
ENSMUST00000220314.2
|
Gm32687
|
predicted gene, 32687 |
| chr10_-_81121596 | 0.02 |
ENSMUST00000218484.2
|
Tjp3
|
tight junction protein 3 |
| chr17_+_33418036 | 0.01 |
ENSMUST00000112165.4
|
Olfr239
|
olfactory receptor 239 |
| chr14_+_53707213 | 0.00 |
ENSMUST00000103643.4
|
Trav8-1
|
T cell receptor alpha variable 8-1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 5.5 | GO:0044208 | 'de novo' AMP biosynthetic process(GO:0044208) |
| 0.9 | 4.6 | GO:0097029 | mature conventional dendritic cell differentiation(GO:0097029) |
| 0.5 | 1.5 | GO:0019417 | sulfur oxidation(GO:0019417) |
| 0.4 | 1.5 | GO:0034553 | respiratory chain complex II assembly(GO:0034552) mitochondrial respiratory chain complex II assembly(GO:0034553) mitochondrial respiratory chain complex II biogenesis(GO:0097032) |
| 0.4 | 1.1 | GO:0032685 | negative regulation of granulocyte macrophage colony-stimulating factor production(GO:0032685) |
| 0.4 | 4.0 | GO:1900383 | regulation of synaptic plasticity by receptor localization to synapse(GO:1900383) |
| 0.3 | 1.3 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.3 | 2.4 | GO:0010626 | negative regulation of Schwann cell proliferation(GO:0010626) |
| 0.3 | 1.2 | GO:0097477 | lateral motor column neuron migration(GO:0097477) |
| 0.3 | 2.3 | GO:0009098 | branched-chain amino acid biosynthetic process(GO:0009082) leucine biosynthetic process(GO:0009098) valine biosynthetic process(GO:0009099) |
| 0.3 | 1.2 | GO:0046900 | tetrahydrofolylpolyglutamate metabolic process(GO:0046900) |
| 0.3 | 0.8 | GO:0090149 | mitochondrial membrane fission(GO:0090149) |
| 0.3 | 1.8 | GO:0060023 | soft palate development(GO:0060023) |
| 0.2 | 1.0 | GO:0098582 | innate vocalization behavior(GO:0098582) |
| 0.2 | 4.1 | GO:0097011 | cellular response to granulocyte macrophage colony-stimulating factor stimulus(GO:0097011) |
| 0.2 | 0.4 | GO:0032416 | negative regulation of sodium:proton antiporter activity(GO:0032416) |
| 0.2 | 1.9 | GO:0046959 | habituation(GO:0046959) |
| 0.2 | 0.5 | GO:0060466 | interleukin-12-mediated signaling pathway(GO:0035722) activation of meiosis involved in egg activation(GO:0060466) cellular response to interleukin-12(GO:0071349) response to fluoride(GO:1902617) response to vasopressin(GO:1904116) cellular response to vasopressin(GO:1904117) |
| 0.2 | 1.6 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
| 0.2 | 1.3 | GO:0010701 | positive regulation of norepinephrine secretion(GO:0010701) |
| 0.1 | 3.7 | GO:0031987 | locomotion involved in locomotory behavior(GO:0031987) |
| 0.1 | 0.9 | GO:0045113 | regulation of integrin biosynthetic process(GO:0045113) |
| 0.1 | 4.4 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.1 | 0.7 | GO:0038145 | macrophage colony-stimulating factor signaling pathway(GO:0038145) |
| 0.1 | 4.4 | GO:0044342 | type B pancreatic cell proliferation(GO:0044342) |
| 0.1 | 2.1 | GO:0070307 | lens fiber cell development(GO:0070307) |
| 0.1 | 2.3 | GO:0071420 | cellular response to histamine(GO:0071420) |
| 0.1 | 1.7 | GO:0043517 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) |
| 0.1 | 0.5 | GO:0070895 | transposon integration(GO:0070893) regulation of transposon integration(GO:0070894) negative regulation of transposon integration(GO:0070895) |
| 0.1 | 6.3 | GO:0019369 | arachidonic acid metabolic process(GO:0019369) |
| 0.1 | 1.7 | GO:0046339 | diacylglycerol metabolic process(GO:0046339) |
| 0.1 | 1.3 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.1 | 2.3 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.1 | 1.4 | GO:0060263 | regulation of respiratory burst(GO:0060263) |
| 0.1 | 1.1 | GO:0035414 | negative regulation of catenin import into nucleus(GO:0035414) |
| 0.1 | 3.2 | GO:0071158 | positive regulation of cell cycle arrest(GO:0071158) |
| 0.1 | 0.6 | GO:0031580 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 0.1 | 2.4 | GO:0022011 | myelination in peripheral nervous system(GO:0022011) peripheral nervous system axon ensheathment(GO:0032292) |
| 0.1 | 4.4 | GO:0051693 | actin filament capping(GO:0051693) |
| 0.1 | 0.5 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.1 | 0.4 | GO:0090235 | regulation of metaphase plate congression(GO:0090235) |
| 0.1 | 2.9 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.0 | 0.3 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 0.0 | 0.4 | GO:0048069 | sphingosine-1-phosphate signaling pathway(GO:0003376) eye pigmentation(GO:0048069) |
| 0.0 | 0.6 | GO:0021902 | commitment of neuronal cell to specific neuron type in forebrain(GO:0021902) |
| 0.0 | 1.5 | GO:0051443 | positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
| 0.0 | 0.5 | GO:0045217 | cell-cell junction maintenance(GO:0045217) |
| 0.0 | 0.9 | GO:0051923 | sulfation(GO:0051923) |
| 0.0 | 1.9 | GO:0035335 | peptidyl-tyrosine dephosphorylation(GO:0035335) |
| 0.0 | 4.6 | GO:0030500 | regulation of bone mineralization(GO:0030500) |
| 0.0 | 1.8 | GO:0002026 | regulation of the force of heart contraction(GO:0002026) |
| 0.0 | 1.0 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.0 | 0.7 | GO:0046033 | AMP metabolic process(GO:0046033) |
| 0.0 | 0.2 | GO:2000255 | negative regulation of male germ cell proliferation(GO:2000255) |
| 0.0 | 0.5 | GO:0060965 | negative regulation of gene silencing by miRNA(GO:0060965) |
| 0.0 | 0.5 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.0 | 0.2 | GO:0098902 | regulation of membrane depolarization during action potential(GO:0098902) |
| 0.0 | 0.3 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.0 | 0.8 | GO:0035411 | catenin import into nucleus(GO:0035411) |
| 0.0 | 1.5 | GO:0014047 | glutamate secretion(GO:0014047) |
| 0.0 | 0.5 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.0 | 0.4 | GO:0097500 | receptor localization to nonmotile primary cilium(GO:0097500) |
| 0.0 | 1.6 | GO:0046579 | positive regulation of Ras protein signal transduction(GO:0046579) |
| 0.0 | 0.5 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.0 | 0.1 | GO:1903964 | monounsaturated fatty acid metabolic process(GO:1903964) monounsaturated fatty acid biosynthetic process(GO:1903966) |
| 0.0 | 3.8 | GO:0050806 | positive regulation of synaptic transmission(GO:0050806) |
| 0.0 | 0.4 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.0 | 0.0 | GO:0010387 | COP9 signalosome assembly(GO:0010387) |
| 0.0 | 3.5 | GO:0071805 | potassium ion transmembrane transport(GO:0071805) |
| 0.0 | 5.6 | GO:0008380 | RNA splicing(GO:0008380) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0034774 | secretory granule lumen(GO:0034774) cytoplasmic membrane-bounded vesicle lumen(GO:0060205) |
| 0.2 | 5.6 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.2 | 3.8 | GO:0031045 | dense core granule(GO:0031045) |
| 0.2 | 2.4 | GO:0031315 | extrinsic component of mitochondrial outer membrane(GO:0031315) |
| 0.2 | 0.7 | GO:1990682 | CSF1-CSF1R complex(GO:1990682) |
| 0.1 | 0.6 | GO:0033165 | interphotoreceptor matrix(GO:0033165) |
| 0.1 | 0.6 | GO:0030125 | clathrin vesicle coat(GO:0030125) |
| 0.1 | 0.5 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 0.1 | 0.9 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.1 | 0.5 | GO:0002142 | stereocilia ankle link complex(GO:0002142) USH2 complex(GO:1990696) |
| 0.1 | 2.3 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.1 | 1.9 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.1 | 15.8 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.1 | 8.5 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.1 | 1.1 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.0 | 7.0 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.0 | 1.3 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.0 | 0.5 | GO:0097451 | glial limiting end-foot(GO:0097451) |
| 0.0 | 4.4 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 0.5 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.0 | 0.5 | GO:0000796 | condensin complex(GO:0000796) |
| 0.0 | 0.5 | GO:0001527 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.0 | 0.7 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.0 | 0.4 | GO:0034464 | BBSome(GO:0034464) |
| 0.0 | 0.6 | GO:0030008 | TRAPP complex(GO:0030008) |
| 0.0 | 0.1 | GO:0071595 | Nem1-Spo7 phosphatase complex(GO:0071595) |
| 0.0 | 3.6 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 1.6 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 1.8 | GO:0031672 | A band(GO:0031672) |
| 0.0 | 1.5 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 1.7 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 0.4 | GO:0005849 | mRNA cleavage factor complex(GO:0005849) |
| 0.0 | 4.0 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.0 | 1.6 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 3.1 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 0.1 | GO:0070775 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.0 | 1.3 | GO:0000118 | histone deacetylase complex(GO:0000118) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.2 | 9.0 | GO:0008147 | structural constituent of bone(GO:0008147) |
| 1.8 | 5.5 | GO:0004019 | adenylosuccinate synthase activity(GO:0004019) |
| 1.6 | 4.9 | GO:0051916 | granulocyte colony-stimulating factor binding(GO:0051916) |
| 0.4 | 3.7 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.3 | 1.7 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.3 | 2.3 | GO:0052655 | branched-chain-amino-acid transaminase activity(GO:0004084) L-leucine transaminase activity(GO:0052654) L-valine transaminase activity(GO:0052655) L-isoleucine transaminase activity(GO:0052656) |
| 0.3 | 1.9 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) ankyrin repeat binding(GO:0071532) |
| 0.3 | 2.3 | GO:0005237 | inhibitory extracellular ligand-gated ion channel activity(GO:0005237) |
| 0.2 | 0.7 | GO:0005011 | macrophage colony-stimulating factor receptor activity(GO:0005011) |
| 0.2 | 0.7 | GO:0046899 | nucleoside triphosphate adenylate kinase activity(GO:0046899) |
| 0.2 | 7.0 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.2 | 4.4 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.1 | 1.5 | GO:0001640 | adenylate cyclase inhibiting G-protein coupled glutamate receptor activity(GO:0001640) |
| 0.1 | 1.3 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.1 | 1.3 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 0.1 | 1.1 | GO:0034711 | inhibin binding(GO:0034711) |
| 0.1 | 1.5 | GO:0097027 | ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.1 | 1.9 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.1 | 1.7 | GO:0001727 | lipid kinase activity(GO:0001727) diacylglycerol kinase activity(GO:0004143) |
| 0.1 | 6.3 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.1 | 0.5 | GO:0004656 | procollagen-proline 4-dioxygenase activity(GO:0004656) |
| 0.1 | 0.5 | GO:0070326 | very-low-density lipoprotein particle receptor binding(GO:0070326) |
| 0.1 | 2.1 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 1.2 | GO:0008242 | omega peptidase activity(GO:0008242) |
| 0.1 | 0.2 | GO:0004958 | prostaglandin F receptor activity(GO:0004958) |
| 0.1 | 0.4 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.1 | 2.0 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 0.5 | GO:0033592 | RNA strand annealing activity(GO:0033592) |
| 0.1 | 0.4 | GO:0046624 | sphingolipid transporter activity(GO:0046624) |
| 0.1 | 0.2 | GO:0047522 | 13-prostaglandin reductase activity(GO:0036132) 15-oxoprostaglandin 13-oxidase activity(GO:0047522) |
| 0.1 | 4.0 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.0 | 2.4 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 0.6 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 1.5 | GO:0071949 | FAD binding(GO:0071949) |
| 0.0 | 1.6 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.0 | 1.8 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 1.8 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.5 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.0 | 2.2 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 0.5 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) phospholipase C activity(GO:0004629) |
| 0.0 | 2.9 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 7.1 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.0 | 0.4 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 0.7 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 0.1 | GO:0032896 | palmitoyl-CoA 9-desaturase activity(GO:0032896) |
| 0.0 | 0.3 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.0 | 1.2 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.0 | 4.1 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 1.4 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 3.7 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 1.5 | GO:0008081 | phosphoric diester hydrolase activity(GO:0008081) |
| 0.0 | 0.7 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.0 | 0.9 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.0 | 0.1 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 4.9 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.1 | 4.6 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.1 | 3.7 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.1 | 1.6 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.0 | 5.7 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 1.9 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.0 | 3.8 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
| 0.0 | 1.5 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.0 | 1.8 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 1.6 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 0.5 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 1.2 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 1.7 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.0 | 2.3 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 0.4 | PID NFAT 3PATHWAY | Role of Calcineurin-dependent NFAT signaling in lymphocytes |
| 0.0 | 1.3 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 1.8 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 0.9 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 5.5 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.2 | 2.2 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.1 | 7.0 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 1.9 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.1 | 2.3 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.1 | 2.3 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.1 | 1.5 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.1 | 4.0 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.1 | 0.9 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.1 | 1.3 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.0 | 1.4 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.0 | 1.2 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 3.5 | REACTOME REGULATION OF MRNA STABILITY BY PROTEINS THAT BIND AU RICH ELEMENTS | Genes involved in Regulation of mRNA Stability by Proteins that Bind AU-rich Elements |
| 0.0 | 1.8 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 1.8 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 1.3 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 0.4 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 0.7 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.4 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 3 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 3 Promoter |
| 0.0 | 0.8 | REACTOME ACTIVATION OF THE MRNA UPON BINDING OF THE CAP BINDING COMPLEX AND EIFS AND SUBSEQUENT BINDING TO 43S | Genes involved in Activation of the mRNA upon binding of the cap-binding complex and eIFs, and subsequent binding to 43S |