Project
avrg: 2D miR_HR1_12
Navigation
Downloads
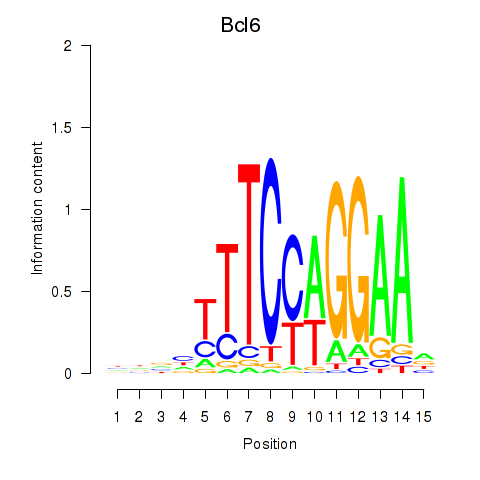
Results for Bcl6
Z-value: 3.38
Transcription factors associated with Bcl6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Bcl6
|
ENSMUSG00000022508.5 | B cell leukemia/lymphoma 6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Bcl6 | mm10_v2_chr16_-_23988852_23988852 | 0.12 | 7.2e-01 | Click! |
Activity profile of Bcl6 motif
Sorted Z-values of Bcl6 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chrX_+_164436987 | 6.44 |
ENSMUST00000036858.4
|
Asb11
|
ankyrin repeat and SOCS box-containing 11 |
| chr11_+_120530688 | 6.02 |
ENSMUST00000026119.7
|
Gcgr
|
glucagon receptor |
| chr9_+_107296843 | 4.97 |
ENSMUST00000167072.1
|
Cish
|
cytokine inducible SH2-containing protein |
| chr2_+_70563435 | 4.49 |
ENSMUST00000123330.1
|
Gad1
|
glutamate decarboxylase 1 |
| chr9_-_121792478 | 4.29 |
ENSMUST00000035110.4
|
Hhatl
|
hedgehog acyltransferase-like |
| chr2_+_70562854 | 3.96 |
ENSMUST00000130998.1
|
Gad1
|
glutamate decarboxylase 1 |
| chr9_+_107296682 | 3.73 |
ENSMUST00000168260.1
|
Cish
|
cytokine inducible SH2-containing protein |
| chr3_+_89418443 | 3.24 |
ENSMUST00000039110.5
ENSMUST00000125036.1 ENSMUST00000154791.1 ENSMUST00000128238.1 ENSMUST00000107417.2 |
Shc1
|
src homology 2 domain-containing transforming protein C1 |
| chr4_+_120666562 | 3.11 |
ENSMUST00000094814.4
|
Cited4
|
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 4 |
| chr5_-_123865491 | 3.05 |
ENSMUST00000057145.5
|
Niacr1
|
niacin receptor 1 |
| chr6_-_5496296 | 3.02 |
ENSMUST00000019721.4
|
Pdk4
|
pyruvate dehydrogenase kinase, isoenzyme 4 |
| chr7_+_43995833 | 2.79 |
ENSMUST00000007156.4
|
Klk1b11
|
kallikrein 1-related peptidase b11 |
| chr7_+_104244449 | 2.53 |
ENSMUST00000106849.2
ENSMUST00000060315.5 |
Trim34a
|
tripartite motif-containing 34A |
| chr16_-_21787796 | 2.41 |
ENSMUST00000023559.5
|
Ehhadh
|
enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase |
| chr1_-_169747634 | 2.32 |
ENSMUST00000027991.5
ENSMUST00000111357.1 |
Rgs4
|
regulator of G-protein signaling 4 |
| chr3_-_89093358 | 2.22 |
ENSMUST00000090929.5
ENSMUST00000052539.6 |
Rusc1
|
RUN and SH3 domain containing 1 |
| chr6_+_41521782 | 2.15 |
ENSMUST00000070380.4
|
Prss2
|
protease, serine, 2 |
| chr18_+_37484955 | 2.02 |
ENSMUST00000053856.4
|
Pcdhb17
|
protocadherin beta 17 |
| chr18_-_32139570 | 2.01 |
ENSMUST00000171765.1
|
Proc
|
protein C |
| chr4_+_115088708 | 1.99 |
ENSMUST00000171877.1
ENSMUST00000177647.1 ENSMUST00000106548.2 ENSMUST00000030488.2 |
Pdzk1ip1
|
PDZK1 interacting protein 1 |
| chr10_+_87521795 | 1.93 |
ENSMUST00000020241.8
|
Pah
|
phenylalanine hydroxylase |
| chr2_-_80447625 | 1.89 |
ENSMUST00000028389.3
|
Frzb
|
frizzled-related protein |
| chr9_-_39604124 | 1.84 |
ENSMUST00000042485.4
ENSMUST00000141370.1 |
AW551984
|
expressed sequence AW551984 |
| chr11_+_103133333 | 1.74 |
ENSMUST00000124928.1
ENSMUST00000062530.4 |
Hexim2
|
hexamethylene bis-acetamide inducible 2 |
| chrX_+_164140447 | 1.71 |
ENSMUST00000073973.4
|
Ace2
|
angiotensin I converting enzyme (peptidyl-dipeptidase A) 2 |
| chr7_+_104244465 | 1.69 |
ENSMUST00000106848.1
|
Trim34a
|
tripartite motif-containing 34A |
| chrX_+_101376359 | 1.68 |
ENSMUST00000119080.1
|
Gjb1
|
gap junction protein, beta 1 |
| chr12_-_40134175 | 1.68 |
ENSMUST00000078481.7
ENSMUST00000002640.5 |
Scin
|
scinderin |
| chr5_+_30466044 | 1.63 |
ENSMUST00000031078.3
ENSMUST00000114743.1 |
1700001C02Rik
|
RIKEN cDNA 1700001C02 gene |
| chr11_+_103133303 | 1.63 |
ENSMUST00000107037.1
|
Hexim2
|
hexamethylene bis-acetamide inducible 2 |
| chr14_-_51146757 | 1.60 |
ENSMUST00000080126.2
|
Rnase1
|
ribonuclease, RNase A family, 1 (pancreatic) |
| chr17_-_56121946 | 1.54 |
ENSMUST00000041357.7
|
Lrg1
|
leucine-rich alpha-2-glycoprotein 1 |
| chr6_-_13838432 | 1.53 |
ENSMUST00000115492.1
|
Gpr85
|
G protein-coupled receptor 85 |
| chr11_+_101245996 | 1.53 |
ENSMUST00000129680.1
|
Ramp2
|
receptor (calcitonin) activity modifying protein 2 |
| chr10_-_29144194 | 1.53 |
ENSMUST00000070359.2
|
Gm9996
|
predicted gene 9996 |
| chr12_-_12941827 | 1.52 |
ENSMUST00000043396.7
|
Mycn
|
v-myc myelocytomatosis viral related oncogene, neuroblastoma derived (avian) |
| chr11_+_93098404 | 1.52 |
ENSMUST00000107859.1
ENSMUST00000042943.6 ENSMUST00000107861.1 ENSMUST00000107858.2 |
Car10
|
carbonic anhydrase 10 |
| chr16_-_35871544 | 1.52 |
ENSMUST00000042665.8
|
Parp14
|
poly (ADP-ribose) polymerase family, member 14 |
| chr14_-_63245219 | 1.51 |
ENSMUST00000118022.1
ENSMUST00000067417.3 |
Gata4
|
GATA binding protein 4 |
| chr14_-_70627008 | 1.42 |
ENSMUST00000110984.2
|
Dmtn
|
dematin actin binding protein |
| chr16_-_36784924 | 1.41 |
ENSMUST00000168279.1
ENSMUST00000164579.1 ENSMUST00000023616.2 |
Slc15a2
|
solute carrier family 15 (H+/peptide transporter), member 2 |
| chr14_+_118137101 | 1.37 |
ENSMUST00000022728.2
|
Gpr180
|
G protein-coupled receptor 180 |
| chr10_+_95417352 | 1.37 |
ENSMUST00000181781.1
|
5730420D15Rik
|
RIKEN cDNA 5730420D15 gene |
| chr18_-_3281036 | 1.34 |
ENSMUST00000049942.6
ENSMUST00000139537.1 ENSMUST00000124747.1 |
Crem
|
cAMP responsive element modulator |
| chr10_+_87521920 | 1.34 |
ENSMUST00000142088.1
|
Pah
|
phenylalanine hydroxylase |
| chr1_+_125676969 | 1.31 |
ENSMUST00000027581.6
|
Gpr39
|
G protein-coupled receptor 39 |
| chr11_-_101175440 | 1.31 |
ENSMUST00000062759.3
|
Ccr10
|
chemokine (C-C motif) receptor 10 |
| chr8_+_119666498 | 1.30 |
ENSMUST00000024107.5
|
Wfdc1
|
WAP four-disulfide core domain 1 |
| chr19_-_43986552 | 1.28 |
ENSMUST00000026210.4
|
Cpn1
|
carboxypeptidase N, polypeptide 1 |
| chr11_+_87699897 | 1.26 |
ENSMUST00000040089.4
|
Rnf43
|
ring finger protein 43 |
| chr6_+_41392356 | 1.25 |
ENSMUST00000049079.7
|
Gm5771
|
predicted gene 5771 |
| chr7_-_30924169 | 1.24 |
ENSMUST00000074671.6
|
Hamp2
|
hepcidin antimicrobial peptide 2 |
| chr5_+_123015010 | 1.22 |
ENSMUST00000121652.1
ENSMUST00000051016.4 |
Orai1
|
ORAI calcium release-activated calcium modulator 1 |
| chr6_+_43265582 | 1.21 |
ENSMUST00000031750.7
|
Arhgef5
|
Rho guanine nucleotide exchange factor (GEF) 5 |
| chr5_+_90561102 | 1.20 |
ENSMUST00000094615.4
|
5830473C10Rik
|
RIKEN cDNA 5830473C10 gene |
| chr6_+_139621888 | 1.18 |
ENSMUST00000032353.8
|
Pik3c2g
|
phosphatidylinositol 3-kinase, C2 domain containing, gamma polypeptide |
| chr11_+_54438188 | 1.17 |
ENSMUST00000046835.7
|
Fnip1
|
folliculin interacting protein 1 |
| chr9_+_38718263 | 1.16 |
ENSMUST00000001544.5
ENSMUST00000118144.1 |
Vwa5a
|
von Willebrand factor A domain containing 5A |
| chr2_+_18698998 | 1.16 |
ENSMUST00000095132.3
|
BC061194
|
cDNA sequence BC061194 |
| chr3_+_96576984 | 1.15 |
ENSMUST00000148290.1
|
Gm16253
|
predicted gene 16253 |
| chr15_-_79141197 | 1.12 |
ENSMUST00000169604.1
|
1700088E04Rik
|
RIKEN cDNA 1700088E04 gene |
| chr9_+_45117813 | 1.12 |
ENSMUST00000170998.1
ENSMUST00000093855.3 |
Scn2b
|
sodium channel, voltage-gated, type II, beta |
| chr2_-_155729359 | 1.11 |
ENSMUST00000040833.4
|
Edem2
|
ER degradation enhancer, mannosidase alpha-like 2 |
| chr7_-_102759465 | 1.10 |
ENSMUST00000168007.1
ENSMUST00000060187.7 |
Olfr78
|
olfactory receptor 78 |
| chr16_-_5222257 | 1.08 |
ENSMUST00000050160.4
|
AU021092
|
expressed sequence AU021092 |
| chr10_+_112165676 | 1.08 |
ENSMUST00000170013.1
|
Caps2
|
calcyphosphine 2 |
| chr17_-_57247632 | 1.08 |
ENSMUST00000005975.6
|
Gpr108
|
G protein-coupled receptor 108 |
| chr11_+_66911981 | 1.07 |
ENSMUST00000123434.2
|
Pirt
|
phosphoinositide-interacting regulator of transient receptor potential channels |
| chr13_+_23555023 | 1.07 |
ENSMUST00000045301.6
|
Hist1h1d
|
histone cluster 1, H1d |
| chr14_-_21989475 | 1.06 |
ENSMUST00000043409.7
|
Zfp503
|
zinc finger protein 503 |
| chr10_-_128401218 | 1.05 |
ENSMUST00000042666.5
|
Slc39a5
|
solute carrier family 39 (metal ion transporter), member 5 |
| chr11_+_83703991 | 1.05 |
ENSMUST00000092836.5
|
Wfdc17
|
WAP four-disulfide core domain 17 |
| chr19_+_46152505 | 1.02 |
ENSMUST00000026254.7
|
Gbf1
|
golgi-specific brefeldin A-resistance factor 1 |
| chr16_-_36784784 | 1.01 |
ENSMUST00000165531.1
|
Slc15a2
|
solute carrier family 15 (H+/peptide transporter), member 2 |
| chr6_+_97929799 | 0.99 |
ENSMUST00000101123.3
|
Mitf
|
microphthalmia-associated transcription factor |
| chr7_-_127930066 | 0.99 |
ENSMUST00000032988.8
|
Prss8
|
protease, serine, 8 (prostasin) |
| chr2_-_66124994 | 0.98 |
ENSMUST00000028378.3
|
Galnt3
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 3 |
| chrX_+_101299143 | 0.97 |
ENSMUST00000118111.1
ENSMUST00000130555.1 ENSMUST00000151528.1 |
Nlgn3
|
neuroligin 3 |
| chr3_+_63295815 | 0.95 |
ENSMUST00000029400.1
|
Mme
|
membrane metallo endopeptidase |
| chr8_+_94532990 | 0.93 |
ENSMUST00000048653.2
ENSMUST00000109537.1 |
Cpne2
|
copine II |
| chr18_+_50053282 | 0.91 |
ENSMUST00000148159.2
|
Tnfaip8
|
tumor necrosis factor, alpha-induced protein 8 |
| chr8_+_3515378 | 0.90 |
ENSMUST00000004681.7
ENSMUST00000111070.2 |
Pnpla6
|
patatin-like phospholipase domain containing 6 |
| chr7_+_104244496 | 0.90 |
ENSMUST00000106854.1
ENSMUST00000143414.1 |
Trim34a
|
tripartite motif-containing 34A |
| chr10_+_87521954 | 0.89 |
ENSMUST00000143624.1
|
Pah
|
phenylalanine hydroxylase |
| chr3_+_96680093 | 0.89 |
ENSMUST00000130429.1
|
Ankrd35
|
ankyrin repeat domain 35 |
| chr3_+_142530329 | 0.89 |
ENSMUST00000171263.1
ENSMUST00000045097.9 |
Gbp7
|
guanylate binding protein 7 |
| chr9_+_50752758 | 0.88 |
ENSMUST00000034562.7
|
Cryab
|
crystallin, alpha B |
| chr14_-_30607808 | 0.87 |
ENSMUST00000112207.1
ENSMUST00000112206.1 ENSMUST00000112202.1 ENSMUST00000112203.1 |
Prkcd
|
protein kinase C, delta |
| chr7_-_126976092 | 0.86 |
ENSMUST00000181859.1
|
D830044I16Rik
|
RIKEN cDNA D830044I16 gene |
| chr6_-_54593139 | 0.85 |
ENSMUST00000046520.6
|
Fkbp14
|
FK506 binding protein 14 |
| chr3_+_95499273 | 0.83 |
ENSMUST00000015664.3
|
Ctsk
|
cathepsin K |
| chr13_-_67399738 | 0.81 |
ENSMUST00000181071.1
ENSMUST00000109732.1 |
Zfp429
|
zinc finger protein 429 |
| chr5_+_67260794 | 0.78 |
ENSMUST00000161369.1
|
Tmem33
|
transmembrane protein 33 |
| chr4_+_62559825 | 0.77 |
ENSMUST00000065870.7
|
Rgs3
|
regulator of G-protein signaling 3 |
| chr5_-_137116177 | 0.77 |
ENSMUST00000054384.5
ENSMUST00000152207.1 |
Trim56
|
tripartite motif-containing 56 |
| chr4_+_144892813 | 0.76 |
ENSMUST00000105744.1
ENSMUST00000171001.1 |
Dhrs3
|
dehydrogenase/reductase (SDR family) member 3 |
| chr6_+_129397478 | 0.76 |
ENSMUST00000112081.2
ENSMUST00000112079.2 |
Clec1b
|
C-type lectin domain family 1, member b |
| chr7_-_100371889 | 0.75 |
ENSMUST00000032963.8
|
Ppme1
|
protein phosphatase methylesterase 1 |
| chr4_-_118544010 | 0.74 |
ENSMUST00000128098.1
|
Tmem125
|
transmembrane protein 125 |
| chr1_+_193301953 | 0.74 |
ENSMUST00000016315.9
|
Lamb3
|
laminin, beta 3 |
| chr5_-_140702241 | 0.73 |
ENSMUST00000077890.5
ENSMUST00000041783.7 ENSMUST00000142081.1 |
Iqce
|
IQ motif containing E |
| chr14_+_122107119 | 0.72 |
ENSMUST00000171318.1
|
Tm9sf2
|
transmembrane 9 superfamily member 2 |
| chr11_+_78499087 | 0.72 |
ENSMUST00000017488.4
|
Vtn
|
vitronectin |
| chr14_-_54253907 | 0.71 |
ENSMUST00000128231.1
|
Dad1
|
defender against cell death 1 |
| chr11_-_90638062 | 0.70 |
ENSMUST00000020858.7
ENSMUST00000107875.1 ENSMUST00000107872.1 ENSMUST00000143203.1 |
Stxbp4
|
syntaxin binding protein 4 |
| chr7_-_24545994 | 0.70 |
ENSMUST00000011776.6
|
Pinlyp
|
phospholipase A2 inhibitor and LY6/PLAUR domain containing |
| chr19_+_56461629 | 0.69 |
ENSMUST00000178590.1
ENSMUST00000039666.6 |
Plekhs1
|
pleckstrin homology domain containing, family S member 1 |
| chr8_-_18741542 | 0.69 |
ENSMUST00000033846.6
|
Angpt2
|
angiopoietin 2 |
| chrX_+_101299207 | 0.69 |
ENSMUST00000065858.2
|
Nlgn3
|
neuroligin 3 |
| chr18_-_36726730 | 0.69 |
ENSMUST00000061829.6
|
Cd14
|
CD14 antigen |
| chr4_+_152274191 | 0.69 |
ENSMUST00000105650.1
ENSMUST00000105651.1 |
Gpr153
|
G protein-coupled receptor 153 |
| chr3_-_57294880 | 0.68 |
ENSMUST00000171384.1
|
Tm4sf1
|
transmembrane 4 superfamily member 1 |
| chr9_-_100571049 | 0.68 |
ENSMUST00000093792.2
|
Slc35g2
|
solute carrier family 35, member G2 |
| chr11_+_97840780 | 0.64 |
ENSMUST00000054783.4
|
B230217C12Rik
|
RIKEN cDNA B230217C12 gene |
| chr9_-_107541816 | 0.64 |
ENSMUST00000041459.3
|
Cyb561d2
|
cytochrome b-561 domain containing 2 |
| chr7_+_110768169 | 0.64 |
ENSMUST00000170374.1
|
Ampd3
|
adenosine monophosphate deaminase 3 |
| chr10_+_22158566 | 0.63 |
ENSMUST00000181645.1
ENSMUST00000105522.2 |
Raet1e
H60b
|
retinoic acid early transcript 1E histocompatibility 60b |
| chr14_+_51884982 | 0.63 |
ENSMUST00000167984.1
|
Mettl17
|
methyltransferase like 17 |
| chr2_+_164948219 | 0.63 |
ENSMUST00000017881.2
|
Mmp9
|
matrix metallopeptidase 9 |
| chr5_+_34915915 | 0.62 |
ENSMUST00000050535.1
|
Msantd1
|
Myb/SANT-like DNA-binding domain containing 1 |
| chr7_+_25077205 | 0.61 |
ENSMUST00000179556.1
ENSMUST00000053410.9 |
Zfp574
|
zinc finger protein 574 |
| chr1_-_175491130 | 0.61 |
ENSMUST00000027812.5
|
Rgs7
|
regulator of G protein signaling 7 |
| chr2_+_154656959 | 0.61 |
ENSMUST00000044277.9
|
Chmp4b
|
charged multivesicular body protein 4B |
| chr10_+_29143996 | 0.59 |
ENSMUST00000092629.2
|
Soga3
|
SOGA family member 3 |
| chr1_+_182954770 | 0.59 |
ENSMUST00000110997.1
|
Tlr5
|
toll-like receptor 5 |
| chr7_-_3629874 | 0.58 |
ENSMUST00000155592.1
ENSMUST00000108641.3 |
Tfpt
|
TCF3 (E2A) fusion partner |
| chr4_-_47010781 | 0.57 |
ENSMUST00000135777.1
|
Gm568
|
predicted gene 568 |
| chr7_-_142661305 | 0.56 |
ENSMUST00000105936.1
|
Igf2
|
insulin-like growth factor 2 |
| chr4_+_33062999 | 0.55 |
ENSMUST00000108162.1
ENSMUST00000024035.2 |
Gabrr2
|
gamma-aminobutyric acid (GABA) C receptor, subunit rho 2 |
| chr4_+_144893077 | 0.55 |
ENSMUST00000154208.1
|
Dhrs3
|
dehydrogenase/reductase (SDR family) member 3 |
| chr13_-_23369156 | 0.54 |
ENSMUST00000125328.1
ENSMUST00000145451.1 ENSMUST00000050101.2 |
Zfp322a
|
zinc finger protein 322A |
| chr10_-_128821576 | 0.54 |
ENSMUST00000026409.3
|
Ormdl2
|
ORM1-like 2 (S. cerevisiae) |
| chr19_-_37178011 | 0.54 |
ENSMUST00000133988.1
|
Cpeb3
|
cytoplasmic polyadenylation element binding protein 3 |
| chr15_-_36879816 | 0.54 |
ENSMUST00000100713.2
|
Gm10384
|
predicted gene 10384 |
| chrX_+_153832225 | 0.53 |
ENSMUST00000148708.1
ENSMUST00000123264.1 ENSMUST00000049999.8 |
Spin2c
|
spindlin family, member 2C |
| chr11_-_118401826 | 0.51 |
ENSMUST00000106290.3
ENSMUST00000043722.3 |
Lgals3bp
|
lectin, galactoside-binding, soluble, 3 binding protein |
| chr8_+_12915879 | 0.49 |
ENSMUST00000110876.2
ENSMUST00000110879.2 |
Mcf2l
|
mcf.2 transforming sequence-like |
| chr11_-_117969176 | 0.49 |
ENSMUST00000054002.3
|
Socs3
|
suppressor of cytokine signaling 3 |
| chr7_-_80232479 | 0.49 |
ENSMUST00000123279.1
|
Cib1
|
calcium and integrin binding 1 (calmyrin) |
| chr3_+_19957240 | 0.48 |
ENSMUST00000108325.2
|
Cp
|
ceruloplasmin |
| chr3_+_19957037 | 0.48 |
ENSMUST00000091309.5
ENSMUST00000108329.1 ENSMUST00000003714.6 |
Cp
|
ceruloplasmin |
| chr2_-_132578244 | 0.48 |
ENSMUST00000110142.1
|
Gpcpd1
|
glycerophosphocholine phosphodiesterase GDE1 homolog (S. cerevisiae) |
| chr5_-_38480131 | 0.48 |
ENSMUST00000143758.1
ENSMUST00000067886.5 |
Slc2a9
|
solute carrier family 2 (facilitated glucose transporter), member 9 |
| chr17_-_51826562 | 0.47 |
ENSMUST00000024720.4
ENSMUST00000129667.1 ENSMUST00000156051.1 ENSMUST00000169480.1 ENSMUST00000148559.1 |
Satb1
|
special AT-rich sequence binding protein 1 |
| chr10_+_23938572 | 0.47 |
ENSMUST00000079134.4
|
Taar2
|
trace amine-associated receptor 2 |
| chr7_+_129257027 | 0.46 |
ENSMUST00000094018.4
|
Ppapdc1a
|
phosphatidic acid phosphatase type 2 domain containing 1A |
| chr10_-_116473875 | 0.45 |
ENSMUST00000068233.4
|
Kcnmb4
|
potassium large conductance calcium-activated channel, subfamily M, beta member 4 |
| chr4_+_42917234 | 0.44 |
ENSMUST00000107976.2
ENSMUST00000069184.2 |
N28178
|
expressed sequence N28178 |
| chr16_-_90934506 | 0.44 |
ENSMUST00000142340.1
|
1110004E09Rik
|
RIKEN cDNA 1110004E09 gene |
| chr17_+_78882003 | 0.44 |
ENSMUST00000180880.1
|
Gm26637
|
predicted gene, 26637 |
| chr2_+_72054598 | 0.44 |
ENSMUST00000028525.5
|
Rapgef4
|
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr4_+_88779850 | 0.44 |
ENSMUST00000179425.1
|
Gm13272
|
predicted gene 13272 |
| chr3_-_30140407 | 0.44 |
ENSMUST00000108271.3
|
Mecom
|
MDS1 and EVI1 complex locus |
| chr7_+_110773658 | 0.43 |
ENSMUST00000143786.1
|
Ampd3
|
adenosine monophosphate deaminase 3 |
| chr4_+_144893127 | 0.43 |
ENSMUST00000142808.1
|
Dhrs3
|
dehydrogenase/reductase (SDR family) member 3 |
| chr11_+_55213783 | 0.43 |
ENSMUST00000108867.1
|
Slc36a1
|
solute carrier family 36 (proton/amino acid symporter), member 1 |
| chr9_-_18512885 | 0.42 |
ENSMUST00000034653.6
|
Muc16
|
mucin 16 |
| chr17_+_74489492 | 0.41 |
ENSMUST00000024873.6
|
Yipf4
|
Yip1 domain family, member 4 |
| chr5_-_6876523 | 0.41 |
ENSMUST00000164784.1
|
Zfp804b
|
zinc finger protein 804B |
| chr11_-_59228162 | 0.41 |
ENSMUST00000163300.1
ENSMUST00000061242.7 |
Arf1
|
ADP-ribosylation factor 1 |
| chr19_+_46341118 | 0.41 |
ENSMUST00000128041.1
|
Tmem180
|
transmembrane protein 180 |
| chr11_+_51763682 | 0.41 |
ENSMUST00000020653.5
|
Sar1b
|
SAR1 gene homolog B (S. cerevisiae) |
| chr2_+_155013531 | 0.40 |
ENSMUST00000029123.2
|
a
|
nonagouti |
| chr7_+_45514619 | 0.40 |
ENSMUST00000107761.1
|
Tulp2
|
tubby-like protein 2 |
| chr9_+_123150941 | 0.40 |
ENSMUST00000026890.4
|
Clec3b
|
C-type lectin domain family 3, member b |
| chr3_-_145032765 | 0.40 |
ENSMUST00000029919.5
|
Clca3
|
chloride channel calcium activated 3 |
| chr3_+_19957088 | 0.39 |
ENSMUST00000108328.1
|
Cp
|
ceruloplasmin |
| chr6_+_48448100 | 0.39 |
ENSMUST00000169350.2
ENSMUST00000043676.5 |
Sspo
|
SCO-spondin |
| chr6_+_129397297 | 0.39 |
ENSMUST00000032262.7
|
Clec1b
|
C-type lectin domain family 1, member b |
| chr17_-_47834682 | 0.38 |
ENSMUST00000066368.6
|
Mdfi
|
MyoD family inhibitor |
| chr3_+_97032416 | 0.38 |
ENSMUST00000132256.1
ENSMUST00000072600.6 |
Gja5
|
gap junction protein, alpha 5 |
| chr12_+_86082555 | 0.38 |
ENSMUST00000054565.6
|
Ift43
|
intraflagellar transport 43 homolog (Chlamydomonas) |
| chr12_+_28675220 | 0.37 |
ENSMUST00000020957.6
|
Adi1
|
acireductone dioxygenase 1 |
| chr11_-_75190458 | 0.37 |
ENSMUST00000044949.4
|
Dph1
|
DPH1 homolog (S. cerevisiae) |
| chr11_-_48871344 | 0.36 |
ENSMUST00000049519.3
|
Irgm1
|
immunity-related GTPase family M member 1 |
| chr3_-_89365233 | 0.35 |
ENSMUST00000070820.6
|
Dcst1
|
DC-STAMP domain containing 1 |
| chr1_+_135729147 | 0.34 |
ENSMUST00000027677.7
|
Csrp1
|
cysteine and glycine-rich protein 1 |
| chr15_+_73512559 | 0.34 |
ENSMUST00000043414.5
|
Dennd3
|
DENN/MADD domain containing 3 |
| chr8_-_87804411 | 0.34 |
ENSMUST00000165770.2
|
Zfp423
|
zinc finger protein 423 |
| chr17_+_8434423 | 0.34 |
ENSMUST00000074667.2
|
T
|
brachyury |
| chr7_+_66839752 | 0.32 |
ENSMUST00000107478.1
|
Adamts17
|
a disintegrin-like and metallopeptidase (reprolysin type) with thrombospondin type 1 motif, 17 |
| chr5_+_67260565 | 0.32 |
ENSMUST00000037918.5
ENSMUST00000162543.1 |
Tmem33
|
transmembrane protein 33 |
| chr5_+_67260696 | 0.32 |
ENSMUST00000161233.1
ENSMUST00000160352.1 |
Tmem33
|
transmembrane protein 33 |
| chr15_-_81104999 | 0.31 |
ENSMUST00000109579.2
|
Mkl1
|
MKL (megakaryoblastic leukemia)/myocardin-like 1 |
| chr4_-_32602760 | 0.31 |
ENSMUST00000056517.2
|
Gja10
|
gap junction protein, alpha 10 |
| chr10_-_95416850 | 0.31 |
ENSMUST00000020215.9
|
Socs2
|
suppressor of cytokine signaling 2 |
| chr12_-_103631404 | 0.31 |
ENSMUST00000121625.1
ENSMUST00000044231.5 |
Serpina10
|
serine (or cysteine) peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 10 |
| chr15_+_10952332 | 0.31 |
ENSMUST00000022853.8
ENSMUST00000110523.1 |
C1qtnf3
|
C1q and tumor necrosis factor related protein 3 |
| chr5_+_23434435 | 0.30 |
ENSMUST00000094962.2
ENSMUST00000115128.1 |
Kmt2e
|
lysine (K)-specific methyltransferase 2E |
| chr17_+_71019503 | 0.29 |
ENSMUST00000024847.7
|
Myom1
|
myomesin 1 |
| chr17_-_32886083 | 0.28 |
ENSMUST00000178401.1
|
Zfp870
|
zinc finger protein 870 |
| chr15_-_5108492 | 0.28 |
ENSMUST00000118365.2
|
Card6
|
caspase recruitment domain family, member 6 |
| chr11_+_49087022 | 0.28 |
ENSMUST00000046704.6
|
Ifi47
|
interferon gamma inducible protein 47 |
| chr17_+_28691342 | 0.28 |
ENSMUST00000114758.1
ENSMUST00000004990.6 ENSMUST00000062694.8 ENSMUST00000114754.1 |
Mapk14
|
mitogen-activated protein kinase 14 |
| chr8_+_68880491 | 0.27 |
ENSMUST00000015712.8
|
Lpl
|
lipoprotein lipase |
| chr8_+_71887264 | 0.27 |
ENSMUST00000034259.7
|
Zfp709
|
zinc finger protein 709 |
| chr11_+_96351632 | 0.27 |
ENSMUST00000100523.5
|
Hoxb2
|
homeobox B2 |
| chr1_-_134955908 | 0.27 |
ENSMUST00000045665.6
ENSMUST00000086444.4 ENSMUST00000112163.1 |
Ppp1r12b
|
protein phosphatase 1, regulatory (inhibitor) subunit 12B |
| chr13_-_23698454 | 0.26 |
ENSMUST00000102967.1
|
Hist1h4c
|
histone cluster 1, H4c |
| chr17_+_71019548 | 0.26 |
ENSMUST00000073211.5
ENSMUST00000179759.1 |
Myom1
|
myomesin 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Bcl6
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 4.2 | GO:0009073 | tyrosine biosynthetic process(GO:0006571) aromatic amino acid family biosynthetic process(GO:0009073) aromatic amino acid family biosynthetic process, prephenate pathway(GO:0009095) |
| 1.0 | 7.2 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 1.0 | 6.0 | GO:0033762 | response to glucagon(GO:0033762) |
| 0.9 | 4.3 | GO:1903059 | regulation of protein lipidation(GO:1903059) |
| 0.8 | 3.0 | GO:0010510 | regulation of acetyl-CoA biosynthetic process from pyruvate(GO:0010510) |
| 0.6 | 3.2 | GO:1990839 | response to endothelin(GO:1990839) |
| 0.6 | 1.7 | GO:2000331 | regulation of terminal button organization(GO:2000331) |
| 0.5 | 1.0 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.5 | 1.5 | GO:0061026 | cardiac muscle tissue regeneration(GO:0061026) negative regulation of connective tissue replacement(GO:1905204) |
| 0.5 | 2.4 | GO:0042938 | dipeptide transport(GO:0042938) |
| 0.5 | 1.4 | GO:0035585 | calcium-mediated signaling using extracellular calcium source(GO:0035585) |
| 0.5 | 1.4 | GO:1903371 | regulation of endoplasmic reticulum tubular network organization(GO:1903371) |
| 0.4 | 1.3 | GO:0035483 | gastric emptying(GO:0035483) |
| 0.4 | 1.7 | GO:0015801 | aromatic amino acid transport(GO:0015801) tryptophan transport(GO:0015827) |
| 0.4 | 0.4 | GO:1900133 | regulation of renin secretion into blood stream(GO:1900133) |
| 0.4 | 1.5 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 0.4 | 1.9 | GO:0070366 | regulation of hepatocyte differentiation(GO:0070366) |
| 0.4 | 1.1 | GO:0036509 | trimming of terminal mannose on B branch(GO:0036509) positive regulation of retrograde protein transport, ER to cytosol(GO:1904154) |
| 0.4 | 1.1 | GO:1900135 | positive regulation of renin secretion into blood stream(GO:1900135) |
| 0.4 | 1.1 | GO:0070315 | G1 to G0 transition involved in cell differentiation(GO:0070315) |
| 0.4 | 1.1 | GO:0034224 | cellular response to zinc ion starvation(GO:0034224) |
| 0.3 | 1.3 | GO:0030070 | insulin processing(GO:0030070) |
| 0.3 | 1.0 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.3 | 1.2 | GO:0002408 | myeloid dendritic cell chemotaxis(GO:0002408) |
| 0.3 | 1.2 | GO:0034760 | negative regulation of iron ion transport(GO:0034757) negative regulation of iron ion transmembrane transport(GO:0034760) |
| 0.3 | 2.0 | GO:1901552 | positive regulation of endothelial cell development(GO:1901552) positive regulation of establishment of endothelial barrier(GO:1903142) |
| 0.3 | 1.7 | GO:0045654 | positive regulation of megakaryocyte differentiation(GO:0045654) |
| 0.2 | 8.7 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.2 | 1.0 | GO:0007113 | endomitotic cell cycle(GO:0007113) thrombopoietin-mediated signaling pathway(GO:0038163) positive regulation of male germ cell proliferation(GO:2000256) |
| 0.2 | 0.7 | GO:0071725 | response to triacyl bacterial lipopeptide(GO:0071725) cellular response to triacyl bacterial lipopeptide(GO:0071727) |
| 0.2 | 1.1 | GO:0046684 | response to pyrethroid(GO:0046684) |
| 0.2 | 1.5 | GO:2000980 | regulation of auditory receptor cell differentiation(GO:0045607) regulation of mechanoreceptor differentiation(GO:0045631) regulation of inner ear receptor cell differentiation(GO:2000980) |
| 0.2 | 0.9 | GO:0006680 | glucosylceramide catabolic process(GO:0006680) |
| 0.2 | 1.3 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 0.2 | 1.5 | GO:0097647 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.2 | 0.5 | GO:1900060 | negative regulation of ceramide biosynthetic process(GO:1900060) |
| 0.2 | 0.5 | GO:1900247 | cytoplasmic translational elongation(GO:0002182) regulation of cytoplasmic translational elongation(GO:1900247) negative regulation of cytoplasmic translational elongation(GO:1900248) |
| 0.2 | 1.8 | GO:0060923 | cardiac muscle cell fate commitment(GO:0060923) |
| 0.2 | 1.0 | GO:0018242 | protein O-linked glycosylation via serine(GO:0018242) |
| 0.2 | 1.7 | GO:0015868 | purine ribonucleotide transport(GO:0015868) |
| 0.2 | 1.1 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.1 | 0.9 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.1 | 1.2 | GO:2000973 | positive regulation of B cell apoptotic process(GO:0002904) regulation of pro-B cell differentiation(GO:2000973) |
| 0.1 | 0.4 | GO:1904457 | positive regulation of neuronal action potential(GO:1904457) |
| 0.1 | 3.6 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.1 | 0.7 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.1 | 0.4 | GO:0036090 | cleavage furrow ingression(GO:0036090) lysosomal membrane organization(GO:0097212) |
| 0.1 | 0.4 | GO:0040030 | regulation of molecular function, epigenetic(GO:0040030) |
| 0.1 | 1.1 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.1 | 1.7 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.1 | 0.6 | GO:0034146 | toll-like receptor 5 signaling pathway(GO:0034146) |
| 0.1 | 3.4 | GO:0045736 | negative regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045736) negative regulation of cyclin-dependent protein kinase activity(GO:1904030) |
| 0.1 | 1.0 | GO:0044336 | canonical Wnt signaling pathway involved in negative regulation of apoptotic process(GO:0044336) |
| 0.1 | 0.6 | GO:0036438 | maintenance of lens transparency(GO:0036438) |
| 0.1 | 0.4 | GO:0043102 | L-methionine biosynthetic process from methylthioadenosine(GO:0019509) amino acid salvage(GO:0043102) L-methionine salvage(GO:0071267) |
| 0.1 | 0.4 | GO:0017182 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.1 | 0.3 | GO:0003257 | positive regulation of transcription from RNA polymerase II promoter involved in myocardial precursor cell differentiation(GO:0003257) |
| 0.1 | 1.1 | GO:0019236 | response to pheromone(GO:0019236) |
| 0.1 | 0.3 | GO:0010735 | positive regulation of transcription via serum response element binding(GO:0010735) |
| 0.1 | 0.2 | GO:1903984 | positive regulation of TRAIL-activated apoptotic signaling pathway(GO:1903984) |
| 0.1 | 0.7 | GO:0097421 | liver regeneration(GO:0097421) |
| 0.1 | 0.5 | GO:0060708 | spongiotrophoblast differentiation(GO:0060708) |
| 0.1 | 0.6 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 0.1 | 0.6 | GO:0060335 | positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) positive regulation of autophagosome maturation(GO:1901098) |
| 0.1 | 0.2 | GO:2000418 | positive regulation of eosinophil migration(GO:2000418) |
| 0.1 | 1.0 | GO:0070633 | transepithelial transport(GO:0070633) |
| 0.1 | 1.2 | GO:0002115 | store-operated calcium entry(GO:0002115) |
| 0.1 | 0.4 | GO:0015808 | L-alanine transport(GO:0015808) |
| 0.1 | 0.7 | GO:0097500 | receptor localization to nonmotile primary cilium(GO:0097500) |
| 0.1 | 0.3 | GO:0055095 | lipoprotein particle mediated signaling(GO:0055095) low-density lipoprotein particle mediated signaling(GO:0055096) |
| 0.1 | 0.4 | GO:0045409 | negative regulation of interleukin-6 biosynthetic process(GO:0045409) |
| 0.1 | 0.3 | GO:0070163 | adiponectin secretion(GO:0070162) regulation of adiponectin secretion(GO:0070163) negative regulation of monocyte chemotactic protein-1 production(GO:0071638) |
| 0.1 | 0.5 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.0 | 1.2 | GO:0039694 | viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) |
| 0.0 | 1.4 | GO:0046688 | response to copper ion(GO:0046688) |
| 0.0 | 1.5 | GO:0030511 | positive regulation of transforming growth factor beta receptor signaling pathway(GO:0030511) positive regulation of cellular response to transforming growth factor beta stimulus(GO:1903846) |
| 0.0 | 0.6 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 0.0 | 0.4 | GO:0010756 | positive regulation of plasminogen activation(GO:0010756) |
| 0.0 | 0.9 | GO:0044406 | adhesion of symbiont to host(GO:0044406) |
| 0.0 | 0.5 | GO:0030643 | cellular phosphate ion homeostasis(GO:0030643) cellular divalent inorganic anion homeostasis(GO:0072501) cellular trivalent inorganic anion homeostasis(GO:0072502) |
| 0.0 | 0.3 | GO:0021569 | rhombomere 3 development(GO:0021569) |
| 0.0 | 1.2 | GO:0048266 | behavioral response to pain(GO:0048266) |
| 0.0 | 0.6 | GO:1902187 | negative regulation of viral release from host cell(GO:1902187) |
| 0.0 | 1.8 | GO:0045744 | negative regulation of G-protein coupled receptor protein signaling pathway(GO:0045744) |
| 0.0 | 0.5 | GO:0046415 | urate metabolic process(GO:0046415) |
| 0.0 | 0.4 | GO:0090336 | positive regulation of brown fat cell differentiation(GO:0090336) |
| 0.0 | 0.2 | GO:0060718 | germ-line sex determination(GO:0018992) chorionic trophoblast cell differentiation(GO:0060718) |
| 0.0 | 0.4 | GO:0033141 | positive regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033141) |
| 0.0 | 0.4 | GO:0060707 | trophoblast giant cell differentiation(GO:0060707) |
| 0.0 | 1.5 | GO:0061045 | negative regulation of wound healing(GO:0061045) |
| 0.0 | 0.5 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.0 | 0.2 | GO:0032510 | endosome to lysosome transport via multivesicular body sorting pathway(GO:0032510) |
| 0.0 | 2.3 | GO:0043627 | response to estrogen(GO:0043627) |
| 0.0 | 0.4 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.0 | 1.3 | GO:0006687 | glycosphingolipid metabolic process(GO:0006687) |
| 0.0 | 0.1 | GO:0090234 | regulation of kinetochore assembly(GO:0090234) |
| 0.0 | 0.1 | GO:0030242 | pexophagy(GO:0030242) |
| 0.0 | 1.1 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.0 | 0.2 | GO:0048148 | behavioral response to cocaine(GO:0048148) |
| 0.0 | 0.4 | GO:0030220 | platelet formation(GO:0030220) |
| 0.0 | 1.6 | GO:0090501 | RNA phosphodiester bond hydrolysis(GO:0090501) |
| 0.0 | 0.6 | GO:0001913 | T cell mediated cytotoxicity(GO:0001913) |
| 0.0 | 0.2 | GO:0071318 | cellular response to ATP(GO:0071318) |
| 0.0 | 1.7 | GO:0006635 | fatty acid beta-oxidation(GO:0006635) |
| 0.0 | 0.7 | GO:0006482 | protein demethylation(GO:0006482) protein dealkylation(GO:0008214) |
| 0.0 | 3.5 | GO:0051607 | defense response to virus(GO:0051607) |
| 0.0 | 0.6 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.0 | 0.3 | GO:0050908 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.0 | 0.1 | GO:0045779 | negative regulation of bone resorption(GO:0045779) negative regulation of bone remodeling(GO:0046851) |
| 0.0 | 0.1 | GO:0014807 | regulation of somitogenesis(GO:0014807) |
| 0.0 | 0.2 | GO:0032119 | sequestering of zinc ion(GO:0032119) regulation of sequestering of zinc ion(GO:0061088) |
| 0.0 | 0.3 | GO:0042119 | neutrophil activation(GO:0042119) |
| 0.0 | 0.3 | GO:0035458 | cellular response to interferon-beta(GO:0035458) |
| 0.0 | 0.1 | GO:0072137 | condensed mesenchymal cell proliferation(GO:0072137) |
| 0.0 | 0.5 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.0 | 0.5 | GO:0046470 | phosphatidylcholine metabolic process(GO:0046470) |
| 0.0 | 0.1 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.0 | 0.2 | GO:0045653 | negative regulation of megakaryocyte differentiation(GO:0045653) |
| 0.0 | 0.4 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.0 | 1.2 | GO:0007586 | digestion(GO:0007586) |
| 0.0 | 0.9 | GO:0061077 | chaperone-mediated protein folding(GO:0061077) |
| 0.0 | 1.8 | GO:0007416 | synapse assembly(GO:0007416) |
| 0.0 | 2.2 | GO:0000209 | protein polyubiquitination(GO:0000209) |
| 0.0 | 0.3 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.4 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 0.3 | 10.3 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.3 | 1.5 | GO:1903439 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.2 | 0.7 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.1 | 0.4 | GO:0044316 | cone cell pedicle(GO:0044316) |
| 0.1 | 0.5 | GO:0035339 | SPOTS complex(GO:0035339) |
| 0.1 | 0.7 | GO:0048237 | rough endoplasmic reticulum lumen(GO:0048237) |
| 0.1 | 2.4 | GO:0005922 | connexon complex(GO:0005922) |
| 0.1 | 0.7 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.1 | 0.9 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.1 | 0.9 | GO:0020005 | symbiont-containing vacuole membrane(GO:0020005) |
| 0.1 | 0.6 | GO:0089717 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 0.1 | 1.1 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.1 | 0.6 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.1 | 0.4 | GO:0001652 | granular component(GO:0001652) |
| 0.0 | 0.1 | GO:0034272 | phosphatidylinositol 3-kinase complex, class III, type I(GO:0034271) phosphatidylinositol 3-kinase complex, class III, type II(GO:0034272) |
| 0.0 | 2.3 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 0.7 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.0 | 0.6 | GO:0044754 | autolysosome(GO:0044754) |
| 0.0 | 1.2 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 1.4 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 1.0 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.0 | 0.4 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.0 | 0.6 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.6 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.0 | 0.6 | GO:0044292 | dendrite terminus(GO:0044292) |
| 0.0 | 0.6 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 0.0 | 0.9 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.0 | 1.2 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.5 | GO:0030014 | CCR4-NOT complex(GO:0030014) messenger ribonucleoprotein complex(GO:1990124) |
| 0.0 | 1.1 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.0 | 0.1 | GO:0097447 | dendritic tree(GO:0097447) |
| 0.0 | 0.2 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 3.5 | GO:0010008 | endosome membrane(GO:0010008) |
| 0.0 | 0.3 | GO:0042627 | chylomicron(GO:0042627) |
| 0.0 | 2.8 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.0 | 1.4 | GO:0016363 | nuclear matrix(GO:0016363) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 6.0 | GO:0004967 | glucagon receptor activity(GO:0004967) |
| 1.2 | 8.4 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 0.8 | 4.2 | GO:0004505 | phenylalanine 4-monooxygenase activity(GO:0004505) |
| 0.8 | 3.2 | GO:0048408 | epidermal growth factor binding(GO:0048408) |
| 0.8 | 2.4 | GO:0042936 | dipeptide transporter activity(GO:0042936) |
| 0.8 | 3.0 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.5 | 2.4 | GO:0004165 | dodecenoyl-CoA delta-isomerase activity(GO:0004165) |
| 0.4 | 3.4 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
| 0.4 | 2.0 | GO:0070012 | oligopeptidase activity(GO:0070012) |
| 0.3 | 1.5 | GO:0097643 | amylin receptor activity(GO:0097643) |
| 0.2 | 1.7 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.2 | 1.7 | GO:0008241 | peptidyl-dipeptidase activity(GO:0008241) |
| 0.2 | 1.6 | GO:0016892 | endoribonuclease activity, producing 3'-phosphomonoesters(GO:0016892) |
| 0.2 | 0.9 | GO:0070976 | TIR domain binding(GO:0070976) |
| 0.2 | 1.2 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.2 | 1.0 | GO:0008427 | calcium-dependent protein kinase inhibitor activity(GO:0008427) |
| 0.2 | 1.4 | GO:0016724 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.2 | 1.1 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 0.2 | 2.4 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.1 | 0.4 | GO:0005302 | hydrogen:amino acid symporter activity(GO:0005280) L-tyrosine transmembrane transporter activity(GO:0005302) |
| 0.1 | 1.1 | GO:0003876 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.1 | 1.1 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.1 | 1.5 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.1 | 1.1 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.1 | 0.5 | GO:0047389 | glycerophosphocholine phosphodiesterase activity(GO:0047389) |
| 0.1 | 0.7 | GO:0004859 | phospholipase inhibitor activity(GO:0004859) |
| 0.1 | 1.3 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.1 | 0.7 | GO:0071723 | lipopeptide binding(GO:0071723) |
| 0.1 | 1.5 | GO:0070403 | NAD+ binding(GO:0070403) |
| 0.1 | 1.2 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.1 | 1.9 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 0.4 | GO:0031782 | melanocortin receptor binding(GO:0031779) type 3 melanocortin receptor binding(GO:0031781) type 4 melanocortin receptor binding(GO:0031782) |
| 0.1 | 1.2 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.1 | 9.5 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.1 | 1.7 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.1 | 0.2 | GO:0031728 | CCR3 chemokine receptor binding(GO:0031728) |
| 0.1 | 0.9 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 1.4 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 3.4 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.1 | 0.2 | GO:0015182 | L-asparagine transmembrane transporter activity(GO:0015182) |
| 0.1 | 0.7 | GO:0004576 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.1 | 0.6 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.5 | GO:1901702 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.0 | 0.9 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 1.0 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 2.3 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.6 | GO:0046703 | natural killer cell lectin-like receptor binding(GO:0046703) |
| 0.0 | 0.2 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 0.0 | 0.7 | GO:0005172 | vascular endothelial growth factor receptor binding(GO:0005172) |
| 0.0 | 0.2 | GO:0046790 | virion binding(GO:0046790) |
| 0.0 | 0.5 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.0 | 1.2 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.6 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.0 | 5.8 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.6 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 1.0 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.0 | 0.1 | GO:0004946 | bombesin receptor activity(GO:0004946) |
| 0.0 | 0.4 | GO:0016004 | phospholipase activator activity(GO:0016004) lipase activator activity(GO:0060229) |
| 0.0 | 0.6 | GO:0004622 | lysophospholipase activity(GO:0004622) |
| 0.0 | 0.4 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.0 | 0.6 | GO:0001871 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.0 | 0.5 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.0 | 1.5 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.0 | 0.1 | GO:0003953 | NAD+ nucleosidase activity(GO:0003953) |
| 0.0 | 0.5 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.0 | 0.6 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.0 | 0.8 | GO:0043394 | proteoglycan binding(GO:0043394) |
| 0.0 | 0.6 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 0.8 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.2 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 0.9 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.9 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.0 | 1.3 | GO:0043621 | protein self-association(GO:0043621) |
| 0.0 | 0.2 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 1.7 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.3 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.4 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.0 | 0.2 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.0 | 1.0 | GO:0008238 | exopeptidase activity(GO:0008238) |
| 0.0 | 1.7 | GO:0005057 | receptor signaling protein activity(GO:0005057) |
| 0.0 | 1.1 | GO:0003707 | steroid hormone receptor activity(GO:0003707) |
| 0.0 | 0.3 | GO:0005044 | scavenger receptor activity(GO:0005044) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 8.7 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.1 | 3.2 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.1 | 3.1 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.1 | 2.0 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.1 | 0.7 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 1.4 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 0.7 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 1.3 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.0 | 2.1 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 1.2 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.0 | 0.6 | PID AVB3 OPN PATHWAY | Osteopontin-mediated events |
| 0.0 | 1.0 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 0.8 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.7 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 1.2 | PID CD8 TCR PATHWAY | TCR signaling in naïve CD8+ T cells |
| 0.0 | 1.0 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 1.3 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.0 | 1.3 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 0.3 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.0 | 1.4 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 1.4 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 0.3 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.0 | 2.2 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 1.5 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 0.7 | PID P73PATHWAY | p73 transcription factor network |
| 0.0 | 0.6 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 8.4 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.2 | 2.2 | REACTOME DEFENSINS | Genes involved in Defensins |
| 0.2 | 3.0 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.2 | 9.5 | REACTOME GROWTH HORMONE RECEPTOR SIGNALING | Genes involved in Growth hormone receptor signaling |
| 0.2 | 3.2 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.2 | 2.2 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.1 | 2.4 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 1.2 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.1 | 1.4 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.1 | 6.0 | REACTOME GLUCAGON TYPE LIGAND RECEPTORS | Genes involved in Glucagon-type ligand receptors |
| 0.1 | 1.1 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.1 | 3.2 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.1 | 1.0 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.1 | 1.5 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.1 | 2.6 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.1 | 1.1 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.1 | 1.3 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |
| 0.1 | 1.1 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 0.7 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.0 | 0.5 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.0 | 0.6 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 0.7 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 2.0 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.0 | 1.4 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.5 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 0.4 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 0.6 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.0 | 5.5 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 0.9 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.0 | 0.4 | REACTOME REGULATION OF INSULIN SECRETION BY GLUCAGON LIKE PEPTIDE1 | Genes involved in Regulation of Insulin Secretion by Glucagon-like Peptide-1 |
| 0.0 | 0.6 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.7 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 0.6 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 0.4 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.0 | 0.7 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.0 | 1.6 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |