Project
avrg: 2D miR_HR1_12
Navigation
Downloads
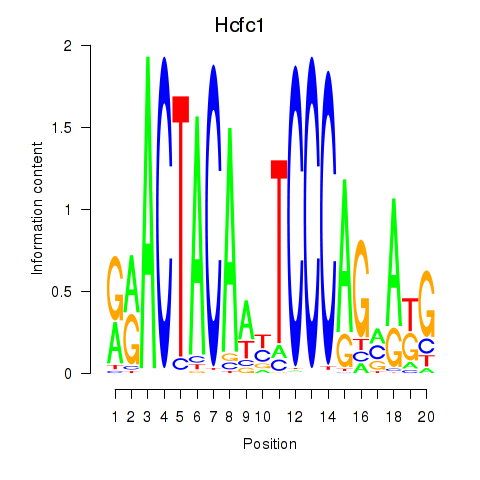
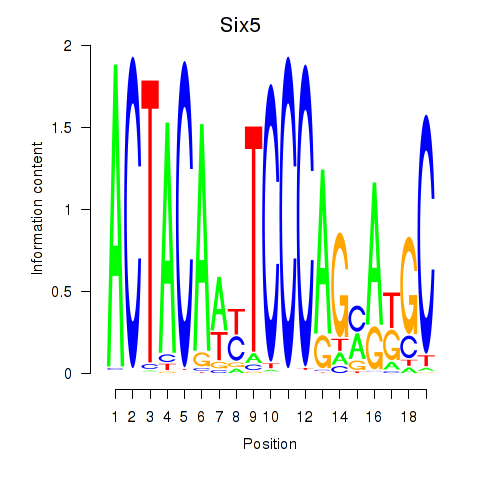
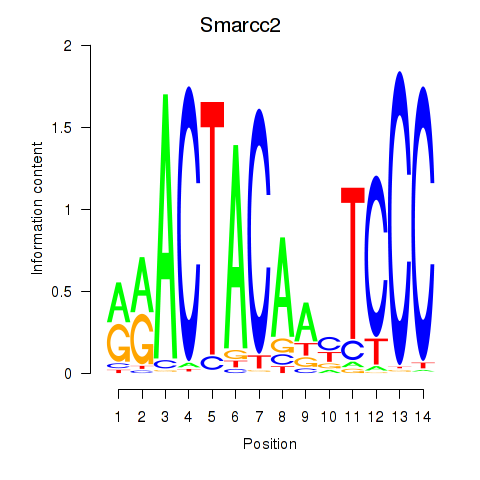
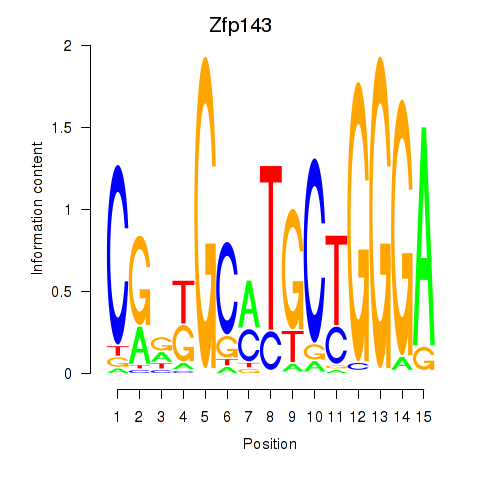
Results for Hcfc1_Six5_Smarcc2_Zfp143
Z-value: 4.59
Transcription factors associated with Hcfc1_Six5_Smarcc2_Zfp143
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hcfc1
|
ENSMUSG00000031386.8 | host cell factor C1 |
|
Six5
|
ENSMUSG00000040841.5 | sine oculis-related homeobox 5 |
|
Smarcc2
|
ENSMUSG00000025369.8 | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily c, member 2 |
|
Zfp143
|
ENSMUSG00000061079.7 | zinc finger protein 143 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hcfc1 | mm10_v2_chrX_-_73966329_73966376 | 0.92 | 2.8e-05 | Click! |
| Zfp143 | mm10_v2_chr7_+_110061702_110061732 | 0.86 | 3.1e-04 | Click! |
| Six5 | mm10_v2_chr7_+_19094594_19094633 | -0.79 | 2.2e-03 | Click! |
| Smarcc2 | mm10_v2_chr10_+_128459236_128459248 | 0.59 | 4.2e-02 | Click! |
Activity profile of Hcfc1_Six5_Smarcc2_Zfp143 motif
Sorted Z-values of Hcfc1_Six5_Smarcc2_Zfp143 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_118598209 | 20.95 |
ENSMUST00000038341.7
|
Bub1b
|
budding uninhibited by benzimidazoles 1 homolog, beta (S. cerevisiae) |
| chr6_+_124829540 | 16.49 |
ENSMUST00000150120.1
|
Cdca3
|
cell division cycle associated 3 |
| chr2_+_152847961 | 14.72 |
ENSMUST00000164120.1
ENSMUST00000178997.1 ENSMUST00000109816.1 |
Tpx2
|
TPX2, microtubule-associated protein homolog (Xenopus laevis) |
| chr5_+_33658123 | 13.86 |
ENSMUST00000074849.6
ENSMUST00000079534.4 |
Tacc3
|
transforming, acidic coiled-coil containing protein 3 |
| chr1_+_178187721 | 13.73 |
ENSMUST00000159284.1
|
Desi2
|
desumoylating isopeptidase 2 |
| chr2_+_152847993 | 13.66 |
ENSMUST00000028969.8
|
Tpx2
|
TPX2, microtubule-associated protein homolog (Xenopus laevis) |
| chr11_+_117849223 | 12.55 |
ENSMUST00000081387.4
|
Birc5
|
baculoviral IAP repeat-containing 5 |
| chr11_+_117849286 | 11.84 |
ENSMUST00000093906.4
|
Birc5
|
baculoviral IAP repeat-containing 5 |
| chr6_+_124829582 | 11.32 |
ENSMUST00000024270.7
|
Cdca3
|
cell division cycle associated 3 |
| chr11_-_101551837 | 11.22 |
ENSMUST00000017290.4
|
Brca1
|
breast cancer 1 |
| chr19_-_10203880 | 10.18 |
ENSMUST00000142241.1
ENSMUST00000116542.2 ENSMUST00000025651.5 ENSMUST00000156291.1 |
Fen1
|
flap structure specific endonuclease 1 |
| chr7_+_13278778 | 10.12 |
ENSMUST00000098814.4
ENSMUST00000146998.1 ENSMUST00000185145.1 |
Lig1
|
ligase I, DNA, ATP-dependent |
| chr13_-_35906324 | 10.09 |
ENSMUST00000174230.1
ENSMUST00000171686.2 |
Rpp40
|
ribonuclease P 40 subunit |
| chr8_+_71464910 | 9.62 |
ENSMUST00000048914.6
|
Mrpl34
|
mitochondrial ribosomal protein L34 |
| chr4_-_124936852 | 9.60 |
ENSMUST00000030690.5
ENSMUST00000084296.3 |
Cdca8
|
cell division cycle associated 8 |
| chr14_-_65833963 | 9.15 |
ENSMUST00000022613.9
|
Esco2
|
establishment of cohesion 1 homolog 2 (S. cerevisiae) |
| chr15_+_102296256 | 9.09 |
ENSMUST00000064924.4
|
Espl1
|
extra spindle poles-like 1 (S. cerevisiae) |
| chr4_-_117182623 | 9.03 |
ENSMUST00000065896.2
|
Kif2c
|
kinesin family member 2C |
| chr4_+_107367757 | 8.94 |
ENSMUST00000139560.1
|
Ndc1
|
NDC1 transmembrane nucleoporin |
| chr19_+_5024006 | 8.79 |
ENSMUST00000025826.5
|
Slc29a2
|
solute carrier family 29 (nucleoside transporters), member 2 |
| chr13_-_54590047 | 8.66 |
ENSMUST00000148222.1
ENSMUST00000026987.5 |
Nop16
|
NOP16 nucleolar protein |
| chr9_+_26999668 | 8.52 |
ENSMUST00000039161.8
|
Thyn1
|
thymocyte nuclear protein 1 |
| chr15_+_85859689 | 8.31 |
ENSMUST00000170629.1
|
Gtse1
|
G two S phase expressed protein 1 |
| chr12_-_91384403 | 8.25 |
ENSMUST00000141429.1
|
Cep128
|
centrosomal protein 128 |
| chr15_-_53902472 | 8.16 |
ENSMUST00000078673.6
|
Samd12
|
sterile alpha motif domain containing 12 |
| chr9_+_107587711 | 8.13 |
ENSMUST00000010192.5
|
Ifrd2
|
interferon-related developmental regulator 2 |
| chr3_+_88081997 | 7.96 |
ENSMUST00000071812.5
|
Iqgap3
|
IQ motif containing GTPase activating protein 3 |
| chr19_-_9899450 | 7.95 |
ENSMUST00000025562.7
|
Incenp
|
inner centromere protein |
| chr19_-_41802028 | 7.85 |
ENSMUST00000026150.8
ENSMUST00000177495.1 ENSMUST00000163265.1 |
Arhgap19
|
Rho GTPase activating protein 19 |
| chr4_-_41275091 | 7.82 |
ENSMUST00000030143.6
ENSMUST00000108068.1 |
Ubap2
|
ubiquitin-associated protein 2 |
| chr4_+_156109971 | 7.81 |
ENSMUST00000072554.6
ENSMUST00000169550.1 ENSMUST00000105576.1 |
9430015G10Rik
|
RIKEN cDNA 9430015G10 gene |
| chr15_+_78428564 | 7.71 |
ENSMUST00000166142.2
ENSMUST00000162517.1 ENSMUST00000089414.4 |
Kctd17
|
potassium channel tetramerisation domain containing 17 |
| chr8_+_85492568 | 7.50 |
ENSMUST00000034136.5
|
Gpt2
|
glutamic pyruvate transaminase (alanine aminotransferase) 2 |
| chrY_+_90784738 | 7.41 |
ENSMUST00000179483.1
|
Erdr1
|
erythroid differentiation regulator 1 |
| chr13_+_41249841 | 7.14 |
ENSMUST00000165561.2
|
Smim13
|
small integral membrane protein 13 |
| chr5_+_30711564 | 7.07 |
ENSMUST00000114729.1
|
Dpysl5
|
dihydropyrimidinase-like 5 |
| chr10_-_128565827 | 6.99 |
ENSMUST00000131728.1
ENSMUST00000026425.6 |
Pa2g4
|
proliferation-associated 2G4 |
| chr19_+_36083696 | 6.93 |
ENSMUST00000025714.7
|
Rpp30
|
ribonuclease P/MRP 30 subunit |
| chr4_+_134468320 | 6.93 |
ENSMUST00000030636.4
ENSMUST00000127279.1 ENSMUST00000105867.1 |
Stmn1
|
stathmin 1 |
| chr9_+_78109188 | 6.91 |
ENSMUST00000118869.1
ENSMUST00000125615.1 |
Ick
|
intestinal cell kinase |
| chr5_+_30711849 | 6.91 |
ENSMUST00000088081.4
ENSMUST00000101442.3 |
Dpysl5
|
dihydropyrimidinase-like 5 |
| chr9_+_106281061 | 6.81 |
ENSMUST00000072206.6
|
Poc1a
|
POC1 centriolar protein homolog A (Chlamydomonas) |
| chr10_+_77033260 | 6.70 |
ENSMUST00000136925.1
ENSMUST00000130703.1 |
Slc19a1
|
solute carrier family 19 (folate transporter), member 1 |
| chr17_-_35673517 | 6.59 |
ENSMUST00000162266.1
ENSMUST00000160734.1 ENSMUST00000159852.1 ENSMUST00000160039.1 |
Gtf2h4
|
general transcription factor II H, polypeptide 4 |
| chr5_+_108132885 | 6.57 |
ENSMUST00000047677.7
|
Ccdc18
|
coiled-coil domain containing 18 |
| chr15_-_81729864 | 6.45 |
ENSMUST00000171115.1
ENSMUST00000170134.1 ENSMUST00000052374.5 |
Rangap1
|
RAN GTPase activating protein 1 |
| chr5_+_21424934 | 6.45 |
ENSMUST00000056045.4
|
Fam185a
|
family with sequence similarity 185, member A |
| chr2_-_26021532 | 6.37 |
ENSMUST00000136750.1
|
Ubac1
|
ubiquitin associated domain containing 1 |
| chr2_+_119112793 | 6.30 |
ENSMUST00000140939.1
ENSMUST00000028795.3 |
Rad51
|
RAD51 homolog |
| chr1_+_175880775 | 6.28 |
ENSMUST00000039725.6
|
Exo1
|
exonuclease 1 |
| chr8_+_105860634 | 6.24 |
ENSMUST00000008594.7
|
Nutf2
|
nuclear transport factor 2 |
| chr2_-_26021679 | 6.23 |
ENSMUST00000036509.7
|
Ubac1
|
ubiquitin associated domain containing 1 |
| chr17_-_23673557 | 6.21 |
ENSMUST00000115489.1
|
Thoc6
|
THO complex 6 homolog (Drosophila) |
| chr10_+_40883819 | 6.19 |
ENSMUST00000105509.1
|
Wasf1
|
WAS protein family, member 1 |
| chr5_+_110839973 | 6.19 |
ENSMUST00000066160.1
|
Chek2
|
checkpoint kinase 2 |
| chr4_+_126556935 | 6.08 |
ENSMUST00000048391.8
|
Clspn
|
claspin |
| chr10_+_77033212 | 5.97 |
ENSMUST00000105410.3
|
Slc19a1
|
solute carrier family 19 (folate transporter), member 1 |
| chr11_+_16951371 | 5.95 |
ENSMUST00000109635.1
ENSMUST00000061327.1 |
Fbxo48
|
F-box protein 48 |
| chr2_+_129100995 | 5.93 |
ENSMUST00000103205.4
ENSMUST00000028874.7 |
Polr1b
|
polymerase (RNA) I polypeptide B |
| chr6_-_112696604 | 5.93 |
ENSMUST00000113182.1
ENSMUST00000113180.1 ENSMUST00000068487.5 ENSMUST00000077088.4 |
Rad18
|
RAD18 homolog (S. cerevisiae) |
| chr10_-_80918212 | 5.89 |
ENSMUST00000057623.7
ENSMUST00000179022.1 |
Lmnb2
|
lamin B2 |
| chr19_+_46056539 | 5.87 |
ENSMUST00000111899.1
ENSMUST00000099392.3 ENSMUST00000062322.4 |
Pprc1
|
peroxisome proliferative activated receptor, gamma, coactivator-related 1 |
| chr1_-_60098104 | 5.82 |
ENSMUST00000143342.1
|
Wdr12
|
WD repeat domain 12 |
| chr17_-_23673825 | 5.77 |
ENSMUST00000115490.1
ENSMUST00000047436.4 ENSMUST00000138190.1 ENSMUST00000095579.4 |
Thoc6
|
THO complex 6 homolog (Drosophila) |
| chr1_-_20820213 | 5.77 |
ENSMUST00000053266.9
|
Mcm3
|
minichromosome maintenance deficient 3 (S. cerevisiae) |
| chr19_-_47050823 | 5.77 |
ENSMUST00000026032.5
|
Pcgf6
|
polycomb group ring finger 6 |
| chr14_-_99099701 | 5.75 |
ENSMUST00000042471.9
|
Dis3
|
DIS3 mitotic control homolog (S. cerevisiae) |
| chr10_+_77033304 | 5.71 |
ENSMUST00000132984.1
|
Slc19a1
|
solute carrier family 19 (folate transporter), member 1 |
| chr10_-_81496329 | 5.62 |
ENSMUST00000020463.7
|
Ncln
|
nicalin homolog (zebrafish) |
| chr7_+_29303958 | 5.60 |
ENSMUST00000049977.6
|
Dpf1
|
D4, zinc and double PHD fingers family 1 |
| chr3_+_88532314 | 5.52 |
ENSMUST00000172699.1
|
Mex3a
|
mex3 homolog A (C. elegans) |
| chr18_-_77767752 | 5.46 |
ENSMUST00000048192.7
|
Haus1
|
HAUS augmin-like complex, subunit 1 |
| chr2_+_181680284 | 5.38 |
ENSMUST00000103042.3
|
Tcea2
|
transcription elongation factor A (SII), 2 |
| chr2_+_173021902 | 5.38 |
ENSMUST00000029014.9
|
Rbm38
|
RNA binding motif protein 38 |
| chr6_+_35177610 | 5.37 |
ENSMUST00000170234.1
|
Nup205
|
nucleoporin 205 |
| chr8_+_57488053 | 5.36 |
ENSMUST00000180690.1
|
2500002B13Rik
|
RIKEN cDNA 2500002B13 gene |
| chr3_+_79629074 | 5.36 |
ENSMUST00000029388.8
|
4930579G24Rik
|
RIKEN cDNA 4930579G24 gene |
| chr16_-_18811615 | 5.35 |
ENSMUST00000096990.3
|
Cdc45
|
cell division cycle 45 |
| chr9_+_81863744 | 5.33 |
ENSMUST00000057067.3
|
Mei4
|
meiosis-specific, MEI4 homolog (S. cerevisiae) |
| chr10_+_40883469 | 5.27 |
ENSMUST00000019975.7
|
Wasf1
|
WAS protein family, member 1 |
| chr11_-_4704334 | 5.26 |
ENSMUST00000058407.5
|
Uqcr10
|
ubiquinol-cytochrome c reductase, complex III subunit X |
| chr10_-_100487267 | 5.26 |
ENSMUST00000128009.1
|
Tmtc3
|
transmembrane and tetratricopeptide repeat containing 3 |
| chr3_+_85574109 | 5.26 |
ENSMUST00000127348.1
ENSMUST00000107672.1 ENSMUST00000107674.1 |
Pet112
|
PET112 homolog (S. cerevisiae) |
| chr15_+_78428650 | 5.24 |
ENSMUST00000159771.1
|
Kctd17
|
potassium channel tetramerisation domain containing 17 |
| chr10_+_128015157 | 5.18 |
ENSMUST00000178041.1
ENSMUST00000026461.7 |
Prim1
|
DNA primase, p49 subunit |
| chr9_+_83548309 | 5.18 |
ENSMUST00000113215.3
|
Sh3bgrl2
|
SH3 domain binding glutamic acid-rich protein like 2 |
| chr4_-_49521036 | 5.12 |
ENSMUST00000057829.3
|
Mrpl50
|
mitochondrial ribosomal protein L50 |
| chr1_-_60098135 | 5.03 |
ENSMUST00000141417.1
ENSMUST00000122038.1 |
Wdr12
|
WD repeat domain 12 |
| chr7_+_29307924 | 4.98 |
ENSMUST00000108230.1
ENSMUST00000065181.5 |
Dpf1
|
D4, zinc and double PHD fingers family 1 |
| chr10_-_41809607 | 4.93 |
ENSMUST00000019951.9
|
Cep57l1
|
centrosomal protein 57-like 1 |
| chr8_-_57487801 | 4.90 |
ENSMUST00000034022.3
|
Sap30
|
sin3 associated polypeptide |
| chr1_-_60097893 | 4.85 |
ENSMUST00000027173.8
|
Wdr12
|
WD repeat domain 12 |
| chr1_+_172482199 | 4.82 |
ENSMUST00000135267.1
ENSMUST00000052629.6 ENSMUST00000111235.2 |
Igsf9
|
immunoglobulin superfamily, member 9 |
| chr17_-_35673738 | 4.78 |
ENSMUST00000001565.8
|
Gtf2h4
|
general transcription factor II H, polypeptide 4 |
| chr2_+_152962485 | 4.74 |
ENSMUST00000099197.2
ENSMUST00000103155.3 |
Ttll9
|
tubulin tyrosine ligase-like family, member 9 |
| chr12_+_79297345 | 4.71 |
ENSMUST00000079533.5
ENSMUST00000171210.1 |
Rad51b
|
RAD51 homolog B |
| chr14_-_54517353 | 4.69 |
ENSMUST00000023873.5
|
Prmt5
|
protein arginine N-methyltransferase 5 |
| chr7_+_29303938 | 4.64 |
ENSMUST00000108231.1
|
Dpf1
|
D4, zinc and double PHD fingers family 1 |
| chr7_+_101663705 | 4.63 |
ENSMUST00000106998.1
|
Clpb
|
ClpB caseinolytic peptidase B |
| chr4_-_107178282 | 4.59 |
ENSMUST00000058585.7
|
Tceanc2
|
transcription elongation factor A (SII) N-terminal and central domain containing 2 |
| chr2_+_167062934 | 4.58 |
ENSMUST00000125674.1
|
1500012F01Rik
|
RIKEN cDNA 1500012F01 gene |
| chr19_+_53329413 | 4.56 |
ENSMUST00000025998.7
|
Mxi1
|
Max interacting protein 1 |
| chr16_+_35983424 | 4.54 |
ENSMUST00000173555.1
|
Kpna1
|
karyopherin (importin) alpha 1 |
| chr5_+_33658567 | 4.46 |
ENSMUST00000114426.3
|
Tacc3
|
transforming, acidic coiled-coil containing protein 3 |
| chr11_+_98907801 | 4.46 |
ENSMUST00000092706.6
|
Cdc6
|
cell division cycle 6 |
| chr5_-_34660068 | 4.45 |
ENSMUST00000041364.9
|
Nop14
|
NOP14 nucleolar protein |
| chr15_-_9140374 | 4.40 |
ENSMUST00000096482.3
ENSMUST00000110585.2 |
Skp2
|
S-phase kinase-associated protein 2 (p45) |
| chr5_-_134314637 | 4.40 |
ENSMUST00000173504.1
|
Gtf2i
|
general transcription factor II I |
| chr13_-_110280103 | 4.38 |
ENSMUST00000167824.1
|
Rab3c
|
RAB3C, member RAS oncogene family |
| chr4_+_116807714 | 4.37 |
ENSMUST00000102699.1
ENSMUST00000130359.1 |
Mutyh
|
mutY homolog (E. coli) |
| chr2_+_71055731 | 4.36 |
ENSMUST00000154704.1
ENSMUST00000135357.1 ENSMUST00000064141.5 ENSMUST00000112159.2 ENSMUST00000102701.3 |
Dcaf17
|
DDB1 and CUL4 associated factor 17 |
| chr10_-_81496313 | 4.29 |
ENSMUST00000118498.1
|
Ncln
|
nicalin homolog (zebrafish) |
| chr2_-_71055534 | 4.27 |
ENSMUST00000090849.5
ENSMUST00000100037.2 ENSMUST00000112186.2 |
Mettl8
|
methyltransferase like 8 |
| chr10_+_84917616 | 4.26 |
ENSMUST00000038523.7
ENSMUST00000095385.3 |
Ric8b
|
resistance to inhibitors of cholinesterase 8 homolog B (C. elegans) |
| chr7_+_101663633 | 4.23 |
ENSMUST00000001884.7
|
Clpb
|
ClpB caseinolytic peptidase B |
| chr18_-_42262053 | 4.20 |
ENSMUST00000097590.3
|
Lars
|
leucyl-tRNA synthetase |
| chr15_+_76904070 | 4.20 |
ENSMUST00000004072.8
|
Rpl8
|
ribosomal protein L8 |
| chrY_+_90785442 | 4.20 |
ENSMUST00000177591.1
ENSMUST00000177671.1 ENSMUST00000179077.1 |
Erdr1
|
erythroid differentiation regulator 1 |
| chr6_+_35177386 | 4.18 |
ENSMUST00000043815.9
|
Nup205
|
nucleoporin 205 |
| chr10_+_80142358 | 4.11 |
ENSMUST00000105366.1
|
Atp5d
|
ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit |
| chr10_+_80142295 | 4.08 |
ENSMUST00000003156.8
|
Atp5d
|
ATP synthase, H+ transporting, mitochondrial F1 complex, delta subunit |
| chr16_-_18811972 | 4.05 |
ENSMUST00000000028.7
ENSMUST00000115585.1 |
Cdc45
|
cell division cycle 45 |
| chr1_-_128103016 | 3.99 |
ENSMUST00000097597.2
|
Zranb3
|
zinc finger, RAN-binding domain containing 3 |
| chr9_-_98601642 | 3.94 |
ENSMUST00000035034.8
|
Mrps22
|
mitochondrial ribosomal protein S22 |
| chr5_+_136919137 | 3.90 |
ENSMUST00000181045.1
|
4933404O12Rik
|
RIKEN cDNA 4933404O12 gene |
| chrX_-_162829379 | 3.90 |
ENSMUST00000041370.4
ENSMUST00000112316.2 ENSMUST00000112315.1 |
Txlng
|
taxilin gamma |
| chr8_-_124721956 | 3.85 |
ENSMUST00000117624.1
ENSMUST00000041614.8 ENSMUST00000118134.1 |
Ttc13
|
tetratricopeptide repeat domain 13 |
| chr5_-_134314378 | 3.85 |
ENSMUST00000174867.1
|
Gtf2i
|
general transcription factor II I |
| chr11_+_116671658 | 3.84 |
ENSMUST00000106378.1
ENSMUST00000144049.1 |
1810032O08Rik
|
RIKEN cDNA 1810032O08 gene |
| chr1_+_181150926 | 3.82 |
ENSMUST00000134115.1
ENSMUST00000111059.1 |
Cnih4
|
cornichon homolog 4 (Drosophila) |
| chr10_-_128626464 | 3.82 |
ENSMUST00000026420.5
|
Rps26
|
ribosomal protein S26 |
| chr4_-_116708312 | 3.73 |
ENSMUST00000030453.4
|
Mmachc
|
methylmalonic aciduria cblC type, with homocystinuria |
| chr11_-_77489666 | 3.72 |
ENSMUST00000037593.7
ENSMUST00000092892.3 |
Ankrd13b
|
ankyrin repeat domain 13b |
| chr5_+_33658550 | 3.71 |
ENSMUST00000152847.1
|
Tacc3
|
transforming, acidic coiled-coil containing protein 3 |
| chr15_-_76639840 | 3.62 |
ENSMUST00000166974.1
ENSMUST00000168185.1 |
Tonsl
|
tonsoku-like, DNA repair protein |
| chr7_-_45830776 | 3.62 |
ENSMUST00000107723.2
ENSMUST00000131384.1 |
Grwd1
|
glutamate-rich WD repeat containing 1 |
| chr7_+_100227311 | 3.61 |
ENSMUST00000084935.3
|
Pgm2l1
|
phosphoglucomutase 2-like 1 |
| chr13_+_106947104 | 3.59 |
ENSMUST00000022203.8
|
Dimt1
|
DIM1 dimethyladenosine transferase 1-like (S. cerevisiae) |
| chrX_-_139998519 | 3.55 |
ENSMUST00000113007.1
ENSMUST00000033810.7 ENSMUST00000113011.2 ENSMUST00000087400.5 |
Rbm41
|
RNA binding motif protein 41 |
| chr16_+_43889936 | 3.54 |
ENSMUST00000151183.1
|
2610015P09Rik
|
RIKEN cDNA 2610015P09 gene |
| chr6_-_83054415 | 3.54 |
ENSMUST00000113962.1
ENSMUST00000089645.6 ENSMUST00000113963.1 |
Htra2
|
HtrA serine peptidase 2 |
| chr15_+_34495302 | 3.53 |
ENSMUST00000052290.7
ENSMUST00000079028.5 |
Pop1
|
processing of precursor 1, ribonuclease P/MRP family, (S. cerevisiae) |
| chr4_+_126556994 | 3.53 |
ENSMUST00000147675.1
|
Clspn
|
claspin |
| chr2_-_155074447 | 3.52 |
ENSMUST00000137242.1
ENSMUST00000054607.9 |
Ahcy
|
S-adenosylhomocysteine hydrolase |
| chr11_-_78183551 | 3.51 |
ENSMUST00000102483.4
|
Rpl23a
|
ribosomal protein L23A |
| chr17_+_24632671 | 3.50 |
ENSMUST00000047611.2
|
Nthl1
|
nth (endonuclease III)-like 1 (E.coli) |
| chr1_+_172481788 | 3.49 |
ENSMUST00000127052.1
|
Igsf9
|
immunoglobulin superfamily, member 9 |
| chr8_-_105707933 | 3.47 |
ENSMUST00000013299.9
|
Enkd1
|
enkurin domain containing 1 |
| chr6_-_120357440 | 3.46 |
ENSMUST00000112703.1
|
Ccdc77
|
coiled-coil domain containing 77 |
| chr12_+_87266696 | 3.43 |
ENSMUST00000021425.6
|
Ahsa1
|
AHA1, activator of heat shock protein ATPase 1 |
| chr19_+_5601854 | 3.40 |
ENSMUST00000025864.4
|
Rnaseh2c
|
ribonuclease H2, subunit C |
| chr6_-_120357342 | 3.40 |
ENSMUST00000163827.1
|
Ccdc77
|
coiled-coil domain containing 77 |
| chr12_+_40446050 | 3.37 |
ENSMUST00000037488.6
|
Dock4
|
dedicator of cytokinesis 4 |
| chr1_-_128102412 | 3.34 |
ENSMUST00000112538.1
ENSMUST00000086614.5 |
Zranb3
|
zinc finger, RAN-binding domain containing 3 |
| chr7_-_4445181 | 3.33 |
ENSMUST00000138798.1
|
Rdh13
|
retinol dehydrogenase 13 (all-trans and 9-cis) |
| chr1_+_179803376 | 3.30 |
ENSMUST00000097454.2
|
Gm10518
|
predicted gene 10518 |
| chr11_-_100712429 | 3.29 |
ENSMUST00000006973.5
ENSMUST00000103118.3 |
Kat2a
|
K(lysine) acetyltransferase 2A |
| chr17_-_25727364 | 3.28 |
ENSMUST00000170070.1
ENSMUST00000048054.7 |
Chtf18
|
CTF18, chromosome transmission fidelity factor 18 |
| chr4_+_62408770 | 3.26 |
ENSMUST00000084524.3
|
Prpf4
|
PRP4 pre-mRNA processing factor 4 homolog (yeast) |
| chr8_-_72443772 | 3.24 |
ENSMUST00000019876.5
|
Calr3
|
calreticulin 3 |
| chr2_-_69206146 | 3.24 |
ENSMUST00000127243.1
ENSMUST00000149643.1 ENSMUST00000167875.2 ENSMUST00000005365.8 |
Spc25
|
SPC25, NDC80 kinetochore complex component, homolog (S. cerevisiae) |
| chr8_-_92355764 | 3.22 |
ENSMUST00000180102.1
ENSMUST00000179421.1 ENSMUST00000179222.1 ENSMUST00000179029.1 |
4933436C20Rik
|
RIKEN cDNA 4933436C20 gene |
| chr9_-_57552760 | 3.11 |
ENSMUST00000034856.8
|
Mpi
|
mannose phosphate isomerase |
| chr13_-_54468805 | 3.10 |
ENSMUST00000026990.5
|
Thoc3
|
THO complex 3 |
| chr6_+_71909046 | 3.10 |
ENSMUST00000055296.8
|
Polr1a
|
polymerase (RNA) I polypeptide A |
| chr10_-_127522428 | 3.10 |
ENSMUST00000026470.4
|
Shmt2
|
serine hydroxymethyltransferase 2 (mitochondrial) |
| chr12_+_106010263 | 3.06 |
ENSMUST00000021539.8
ENSMUST00000085026.4 ENSMUST00000072040.5 |
Vrk1
|
vaccinia related kinase 1 |
| chr7_+_126861947 | 3.05 |
ENSMUST00000037248.3
|
Hirip3
|
HIRA interacting protein 3 |
| chr17_+_84626458 | 3.03 |
ENSMUST00000025101.8
|
Dync2li1
|
dynein cytoplasmic 2 light intermediate chain 1 |
| chr15_-_44428303 | 3.01 |
ENSMUST00000038719.6
|
Nudcd1
|
NudC domain containing 1 |
| chr6_-_128437653 | 3.00 |
ENSMUST00000151796.1
|
Fkbp4
|
FK506 binding protein 4 |
| chr6_+_108213086 | 2.98 |
ENSMUST00000032192.6
|
Itpr1
|
inositol 1,4,5-trisphosphate receptor 1 |
| chrX_-_8252304 | 2.96 |
ENSMUST00000115594.1
|
Ftsj1
|
FtsJ homolog 1 (E. coli) |
| chr2_-_77946180 | 2.96 |
ENSMUST00000111824.1
ENSMUST00000111819.1 ENSMUST00000128963.1 |
Cwc22
|
CWC22 spliceosome-associated protein homolog (S. cerevisiae) |
| chr2_-_69206133 | 2.95 |
ENSMUST00000112320.1
|
Spc25
|
SPC25, NDC80 kinetochore complex component, homolog (S. cerevisiae) |
| chr14_+_64950037 | 2.95 |
ENSMUST00000043914.5
|
Ints9
|
integrator complex subunit 9 |
| chr8_-_105851981 | 2.92 |
ENSMUST00000040776.4
|
Cenpt
|
centromere protein T |
| chr11_+_29547950 | 2.92 |
ENSMUST00000020753.3
|
Clhc1
|
clathrin heavy chain linker domain containing 1 |
| chr8_-_70897407 | 2.92 |
ENSMUST00000054220.8
|
Rpl18a
|
ribosomal protein L18A |
| chr7_-_99483645 | 2.91 |
ENSMUST00000107096.1
ENSMUST00000032998.6 |
Rps3
|
ribosomal protein S3 |
| chr6_-_87672142 | 2.91 |
ENSMUST00000032130.2
ENSMUST00000065997.2 |
Aplf
|
aprataxin and PNKP like factor |
| chr11_-_29547820 | 2.89 |
ENSMUST00000102844.3
|
Rps27a
|
ribosomal protein S27A |
| chr14_-_87141206 | 2.85 |
ENSMUST00000022599.7
|
Diap3
|
diaphanous homolog 3 (Drosophila) |
| chr4_-_126202335 | 2.85 |
ENSMUST00000142125.1
ENSMUST00000106141.2 |
Thrap3
|
thyroid hormone receptor associated protein 3 |
| chr1_-_179803625 | 2.84 |
ENSMUST00000027768.7
|
Ahctf1
|
AT hook containing transcription factor 1 |
| chr6_-_120357422 | 2.84 |
ENSMUST00000032283.5
|
Ccdc77
|
coiled-coil domain containing 77 |
| chr17_-_35838208 | 2.80 |
ENSMUST00000134978.2
|
Tubb5
|
tubulin, beta 5 class I |
| chr9_+_108296853 | 2.78 |
ENSMUST00000035230.5
|
Amt
|
aminomethyltransferase |
| chr8_+_124722139 | 2.77 |
ENSMUST00000034463.3
|
Arv1
|
ARV1 homolog (yeast) |
| chr2_-_27475600 | 2.75 |
ENSMUST00000147736.1
|
Brd3
|
bromodomain containing 3 |
| chr12_-_79192248 | 2.73 |
ENSMUST00000161204.1
|
Rdh11
|
retinol dehydrogenase 11 |
| chr6_+_21985903 | 2.72 |
ENSMUST00000137437.1
ENSMUST00000115383.2 |
Cped1
|
cadherin-like and PC-esterase domain containing 1 |
| chr4_-_139352538 | 2.69 |
ENSMUST00000102503.3
|
Mrto4
|
MRT4, mRNA turnover 4, homolog (S. cerevisiae) |
| chr1_-_177796451 | 2.69 |
ENSMUST00000016105.8
|
Adss
|
adenylosuccinate synthetase, non muscle |
| chr7_-_89941196 | 2.63 |
ENSMUST00000117354.1
|
l7Rn6
|
lethal, Chr 7, Rinchik 6 |
| chr18_+_34625009 | 2.60 |
ENSMUST00000166044.1
|
Kif20a
|
kinesin family member 20A |
| chr4_-_139352298 | 2.60 |
ENSMUST00000030513.6
ENSMUST00000155257.1 |
Mrto4
|
MRT4, mRNA turnover 4, homolog (S. cerevisiae) |
| chr11_-_70015346 | 2.59 |
ENSMUST00000018718.7
ENSMUST00000102574.3 |
Acadvl
|
acyl-Coenzyme A dehydrogenase, very long chain |
| chr7_+_122159422 | 2.59 |
ENSMUST00000033154.6
|
Plk1
|
polo-like kinase 1 |
| chr17_-_35838259 | 2.56 |
ENSMUST00000001566.8
|
Tubb5
|
tubulin, beta 5 class I |
Network of associatons between targets according to the STRING database.
First level regulatory network of Hcfc1_Six5_Smarcc2_Zfp143
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.0 | 21.0 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 4.1 | 24.4 | GO:0031536 | positive regulation of exit from mitosis(GO:0031536) |
| 3.7 | 11.2 | GO:2000620 | positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 3.2 | 22.4 | GO:0033567 | DNA replication, Okazaki fragment processing(GO:0033567) |
| 3.1 | 9.4 | GO:1902299 | pre-replicative complex assembly involved in nuclear cell cycle DNA replication(GO:0006267) pre-replicative complex assembly(GO:0036388) DNA replication preinitiation complex assembly(GO:0071163) pre-replicative complex assembly involved in cell cycle DNA replication(GO:1902299) |
| 3.1 | 18.4 | GO:0098838 | reduced folate transmembrane transport(GO:0098838) |
| 2.5 | 7.5 | GO:0042851 | L-alanine metabolic process(GO:0042851) |
| 2.4 | 7.3 | GO:0036292 | DNA rewinding(GO:0036292) |
| 2.2 | 8.7 | GO:1990414 | replication-born double-strand break repair via sister chromatid exchange(GO:1990414) |
| 2.1 | 6.2 | GO:0090204 | protein localization to nuclear pore(GO:0090204) negative regulation of vascular endothelial growth factor production(GO:1904046) |
| 2.1 | 6.2 | GO:0051329 | interphase(GO:0051325) mitotic interphase(GO:0051329) |
| 1.9 | 9.4 | GO:0014886 | transition between slow and fast fiber(GO:0014886) |
| 1.8 | 22.1 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 1.8 | 9.2 | GO:0034421 | post-translational protein acetylation(GO:0034421) |
| 1.8 | 9.0 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 1.6 | 11.5 | GO:1990416 | cellular response to brain-derived neurotrophic factor stimulus(GO:1990416) |
| 1.6 | 22.7 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 1.5 | 2.9 | GO:0051106 | regulation of DNA ligation(GO:0051105) positive regulation of DNA ligation(GO:0051106) |
| 1.4 | 4.3 | GO:1905053 | regulation of base-excision repair(GO:1905051) positive regulation of base-excision repair(GO:1905053) |
| 1.4 | 5.8 | GO:0071043 | CUT catabolic process(GO:0071034) CUT metabolic process(GO:0071043) |
| 1.4 | 53.1 | GO:0060236 | regulation of mitotic spindle organization(GO:0060236) |
| 1.4 | 15.1 | GO:0046784 | viral mRNA export from host cell nucleus(GO:0046784) |
| 1.4 | 5.5 | GO:0099527 | postsynapse to nucleus signaling pathway(GO:0099527) |
| 1.3 | 17.0 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 1.3 | 6.5 | GO:0060447 | bud outgrowth involved in lung branching(GO:0060447) |
| 1.3 | 3.9 | GO:0002184 | cytoplasmic translational termination(GO:0002184) |
| 1.3 | 5.2 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 1.2 | 3.5 | GO:0006285 | base-excision repair, AP site formation(GO:0006285) nucleotide-excision repair, DNA incision, 5'-to lesion(GO:0006296) |
| 1.2 | 6.9 | GO:0035247 | peptidyl-arginine omega-N-methylation(GO:0035247) |
| 1.1 | 5.7 | GO:0035720 | intraciliary anterograde transport(GO:0035720) |
| 1.1 | 6.8 | GO:0003431 | growth plate cartilage chondrocyte development(GO:0003431) positive regulation of centrosome duplication(GO:0010825) |
| 1.1 | 11.2 | GO:1905098 | negative regulation of guanyl-nucleotide exchange factor activity(GO:1905098) |
| 1.1 | 3.3 | GO:0071929 | alpha-tubulin acetylation(GO:0071929) |
| 1.1 | 5.3 | GO:0042138 | meiotic DNA double-strand break formation(GO:0042138) |
| 1.1 | 4.2 | GO:0006438 | valyl-tRNA aminoacylation(GO:0006438) |
| 1.0 | 3.1 | GO:0009298 | GDP-mannose biosynthetic process(GO:0009298) |
| 0.9 | 3.7 | GO:1904694 | negative regulation of vascular smooth muscle contraction(GO:1904694) |
| 0.9 | 2.8 | GO:0019464 | glycine catabolic process(GO:0006546) glycine decarboxylation via glycine cleavage system(GO:0019464) |
| 0.9 | 5.4 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) |
| 0.9 | 2.7 | GO:0044208 | 'de novo' AMP biosynthetic process(GO:0044208) |
| 0.9 | 3.6 | GO:0002439 | chronic inflammatory response to antigenic stimulus(GO:0002439) |
| 0.9 | 9.6 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.8 | 3.3 | GO:0030974 | thiamine pyrophosphate transport(GO:0030974) |
| 0.8 | 7.1 | GO:0009644 | response to high light intensity(GO:0009644) |
| 0.8 | 3.8 | GO:0035405 | histone-threonine phosphorylation(GO:0035405) |
| 0.7 | 2.1 | GO:0018214 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.7 | 5.0 | GO:0035735 | intraciliary transport involved in cilium morphogenesis(GO:0035735) |
| 0.7 | 13.2 | GO:0007080 | mitotic metaphase plate congression(GO:0007080) |
| 0.7 | 4.8 | GO:0071169 | establishment of protein localization to chromatin(GO:0071169) |
| 0.7 | 2.7 | GO:0016062 | adaptation of rhodopsin mediated signaling(GO:0016062) light adaption(GO:0036367) |
| 0.6 | 3.1 | GO:1904749 | regulation of protein localization to nucleolus(GO:1904749) |
| 0.6 | 3.1 | GO:0006564 | L-serine biosynthetic process(GO:0006564) |
| 0.6 | 1.8 | GO:0046168 | glycerol-3-phosphate catabolic process(GO:0046168) |
| 0.6 | 2.4 | GO:0032053 | ciliary basal body organization(GO:0032053) |
| 0.6 | 6.6 | GO:0000920 | cell separation after cytokinesis(GO:0000920) |
| 0.6 | 4.2 | GO:0009235 | cobalamin metabolic process(GO:0009235) |
| 0.6 | 11.3 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.6 | 3.5 | GO:0060178 | regulation of exocyst localization(GO:0060178) |
| 0.6 | 8.0 | GO:0033601 | positive regulation of mammary gland epithelial cell proliferation(GO:0033601) |
| 0.6 | 2.8 | GO:1901525 | negative regulation of macromitophagy(GO:1901525) |
| 0.5 | 4.9 | GO:0000070 | mitotic sister chromatid segregation(GO:0000070) |
| 0.5 | 6.3 | GO:0016446 | somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 0.5 | 1.0 | GO:0071431 | tRNA export from nucleus(GO:0006409) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.5 | 1.0 | GO:0035246 | peptidyl-arginine N-methylation(GO:0035246) |
| 0.5 | 7.8 | GO:0000212 | meiotic spindle organization(GO:0000212) |
| 0.5 | 1.5 | GO:0042539 | hypotonic salinity response(GO:0042539) cellular hypotonic salinity response(GO:0071477) |
| 0.5 | 6.6 | GO:0006977 | DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest(GO:0006977) |
| 0.5 | 3.0 | GO:0030422 | production of siRNA involved in RNA interference(GO:0030422) |
| 0.5 | 2.5 | GO:0000454 | snoRNA guided rRNA pseudouridine synthesis(GO:0000454) |
| 0.5 | 4.0 | GO:0007000 | nucleolus organization(GO:0007000) |
| 0.5 | 1.5 | GO:1901097 | negative regulation of autophagosome maturation(GO:1901097) |
| 0.5 | 4.9 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.5 | 0.5 | GO:0000720 | pyrimidine dimer repair by nucleotide-excision repair(GO:0000720) |
| 0.5 | 5.3 | GO:0097033 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) mitochondrial respiratory chain complex III biogenesis(GO:0097033) |
| 0.5 | 3.3 | GO:1900262 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.5 | 2.3 | GO:1902775 | mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.5 | 6.0 | GO:0051984 | positive regulation of chromosome segregation(GO:0051984) |
| 0.5 | 2.3 | GO:0090666 | scaRNA localization to Cajal body(GO:0090666) |
| 0.4 | 1.3 | GO:1902990 | leading strand elongation(GO:0006272) mitotic telomere maintenance via semi-conservative replication(GO:1902990) |
| 0.4 | 0.9 | GO:0034085 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 0.4 | 3.5 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.4 | 13.7 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.4 | 4.3 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.4 | 2.6 | GO:0045917 | positive regulation of complement activation(GO:0045917) progesterone receptor signaling pathway(GO:0050847) positive regulation of protein activation cascade(GO:2000259) |
| 0.4 | 6.7 | GO:1902916 | positive regulation of protein polyubiquitination(GO:1902916) |
| 0.4 | 1.7 | GO:1990481 | mRNA pseudouridine synthesis(GO:1990481) |
| 0.4 | 1.2 | GO:0000350 | generation of catalytic spliceosome for second transesterification step(GO:0000350) |
| 0.4 | 10.5 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.4 | 0.8 | GO:0000389 | mRNA 3'-splice site recognition(GO:0000389) |
| 0.4 | 2.8 | GO:0032380 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 0.4 | 2.4 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 0.4 | 1.9 | GO:0000055 | ribosomal large subunit export from nucleus(GO:0000055) |
| 0.4 | 0.8 | GO:0072717 | cellular response to actinomycin D(GO:0072717) |
| 0.4 | 3.8 | GO:0042045 | epithelial fluid transport(GO:0042045) |
| 0.4 | 3.8 | GO:0003373 | dynamin polymerization involved in membrane fission(GO:0003373) dynamin polymerization involved in mitochondrial fission(GO:0003374) |
| 0.4 | 1.1 | GO:1902277 | negative regulation of pancreatic amylase secretion(GO:1902277) |
| 0.4 | 8.8 | GO:0015858 | nucleoside transport(GO:0015858) |
| 0.4 | 1.4 | GO:1990928 | response to amino acid starvation(GO:1990928) |
| 0.4 | 1.1 | GO:0098968 | neurotransmitter receptor transport postsynaptic membrane to endosome(GO:0098968) |
| 0.3 | 0.7 | GO:0016080 | synaptic vesicle targeting(GO:0016080) |
| 0.3 | 2.1 | GO:0090154 | positive regulation of sphingolipid biosynthetic process(GO:0090154) positive regulation of ceramide biosynthetic process(GO:2000304) |
| 0.3 | 12.2 | GO:0042273 | ribosomal large subunit biogenesis(GO:0042273) |
| 0.3 | 0.3 | GO:0010571 | positive regulation of nuclear cell cycle DNA replication(GO:0010571) positive regulation of DNA-dependent DNA replication(GO:2000105) |
| 0.3 | 0.6 | GO:0051902 | negative regulation of mitochondrial depolarization(GO:0051902) |
| 0.3 | 3.8 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 0.3 | 1.0 | GO:0043686 | co-translational protein modification(GO:0043686) |
| 0.3 | 2.8 | GO:0034501 | protein localization to kinetochore(GO:0034501) |
| 0.3 | 2.5 | GO:0007253 | cytoplasmic sequestering of NF-kappaB(GO:0007253) |
| 0.3 | 1.5 | GO:0014022 | neural plate elongation(GO:0014022) convergent extension involved in neural plate elongation(GO:0022007) |
| 0.3 | 0.3 | GO:0043465 | regulation of fermentation(GO:0043465) regulation of NAD metabolic process(GO:1902688) regulation of glucose catabolic process to lactate via pyruvate(GO:1904023) |
| 0.3 | 5.7 | GO:0000154 | rRNA modification(GO:0000154) |
| 0.3 | 7.5 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.3 | 0.9 | GO:2000616 | negative regulation of histone H3-K9 acetylation(GO:2000616) |
| 0.3 | 0.9 | GO:2000554 | regulation of T-helper 1 cell cytokine production(GO:2000554) positive regulation of T-helper 1 cell cytokine production(GO:2000556) |
| 0.3 | 1.3 | GO:0019276 | UDP-N-acetylgalactosamine metabolic process(GO:0019276) |
| 0.3 | 1.3 | GO:0075525 | viral translational termination-reinitiation(GO:0075525) |
| 0.3 | 4.3 | GO:0042407 | cristae formation(GO:0042407) |
| 0.3 | 0.8 | GO:0002276 | basophil activation involved in immune response(GO:0002276) |
| 0.3 | 2.1 | GO:0046826 | negative regulation of protein export from nucleus(GO:0046826) |
| 0.3 | 2.0 | GO:0034316 | negative regulation of Arp2/3 complex-mediated actin nucleation(GO:0034316) |
| 0.3 | 2.0 | GO:1903350 | response to dopamine(GO:1903350) cellular response to dopamine(GO:1903351) |
| 0.2 | 7.7 | GO:0045722 | positive regulation of gluconeogenesis(GO:0045722) |
| 0.2 | 0.7 | GO:1903588 | negative regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903588) |
| 0.2 | 2.4 | GO:0070940 | dephosphorylation of RNA polymerase II C-terminal domain(GO:0070940) |
| 0.2 | 0.7 | GO:0035928 | rRNA import into mitochondrion(GO:0035928) |
| 0.2 | 1.0 | GO:0046368 | GDP-L-fucose metabolic process(GO:0046368) |
| 0.2 | 1.0 | GO:0006624 | vacuolar protein processing(GO:0006624) |
| 0.2 | 7.1 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.2 | 3.1 | GO:0008608 | attachment of spindle microtubules to kinetochore(GO:0008608) |
| 0.2 | 0.7 | GO:0006659 | phosphatidylserine biosynthetic process(GO:0006659) |
| 0.2 | 1.6 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.2 | 2.1 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.2 | 8.6 | GO:0032469 | endoplasmic reticulum calcium ion homeostasis(GO:0032469) |
| 0.2 | 15.3 | GO:0051225 | spindle assembly(GO:0051225) |
| 0.2 | 3.6 | GO:0006337 | nucleosome disassembly(GO:0006337) |
| 0.2 | 0.9 | GO:1901069 | guanosine-containing compound catabolic process(GO:1901069) |
| 0.2 | 2.2 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.2 | 0.7 | GO:0060336 | negative regulation of response to interferon-gamma(GO:0060331) negative regulation of interferon-gamma-mediated signaling pathway(GO:0060336) |
| 0.2 | 2.4 | GO:0007342 | fusion of sperm to egg plasma membrane(GO:0007342) |
| 0.2 | 0.7 | GO:0006550 | isoleucine catabolic process(GO:0006550) |
| 0.2 | 0.6 | GO:0036090 | cleavage furrow ingression(GO:0036090) |
| 0.2 | 0.6 | GO:0006772 | thiamine metabolic process(GO:0006772) thiamine-containing compound metabolic process(GO:0042723) |
| 0.2 | 0.6 | GO:0031053 | primary miRNA processing(GO:0031053) |
| 0.2 | 1.9 | GO:0051974 | negative regulation of telomerase activity(GO:0051974) |
| 0.2 | 1.3 | GO:0061641 | CENP-A containing nucleosome assembly(GO:0034080) CENP-A containing chromatin organization(GO:0061641) |
| 0.2 | 2.5 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.2 | 3.1 | GO:0033211 | adiponectin-activated signaling pathway(GO:0033211) |
| 0.2 | 1.4 | GO:0017185 | peptidyl-lysine hydroxylation(GO:0017185) |
| 0.2 | 0.8 | GO:0061086 | negative regulation of histone H3-K27 methylation(GO:0061086) |
| 0.2 | 2.3 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.2 | 6.9 | GO:0033119 | negative regulation of RNA splicing(GO:0033119) |
| 0.2 | 4.9 | GO:0007052 | mitotic spindle organization(GO:0007052) |
| 0.2 | 0.9 | GO:1903265 | positive regulation of tumor necrosis factor-mediated signaling pathway(GO:1903265) |
| 0.2 | 0.6 | GO:0060912 | cardiac cell fate specification(GO:0060912) |
| 0.2 | 2.0 | GO:0016114 | terpenoid biosynthetic process(GO:0016114) |
| 0.2 | 0.5 | GO:1902774 | late endosome to lysosome transport(GO:1902774) |
| 0.2 | 1.1 | GO:0034227 | tRNA thio-modification(GO:0034227) |
| 0.2 | 11.6 | GO:0035019 | somatic stem cell population maintenance(GO:0035019) |
| 0.2 | 2.0 | GO:0048341 | paraxial mesoderm formation(GO:0048341) |
| 0.2 | 2.5 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.2 | 0.9 | GO:0045039 | protein import into mitochondrial inner membrane(GO:0045039) |
| 0.2 | 0.9 | GO:0030638 | polyketide metabolic process(GO:0030638) aminoglycoside antibiotic metabolic process(GO:0030647) daunorubicin metabolic process(GO:0044597) doxorubicin metabolic process(GO:0044598) |
| 0.2 | 0.5 | GO:0036091 | positive regulation of transcription from RNA polymerase II promoter in response to oxidative stress(GO:0036091) regulation of gluconeogenesis involved in cellular glucose homeostasis(GO:0090526) |
| 0.2 | 1.0 | GO:0097210 | axonal transport of mitochondrion(GO:0019896) response to gonadotropin-releasing hormone(GO:0097210) cellular response to gonadotropin-releasing hormone(GO:0097211) |
| 0.2 | 1.4 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.2 | 0.9 | GO:0031860 | telomeric 3' overhang formation(GO:0031860) |
| 0.2 | 0.7 | GO:0017182 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.2 | 0.7 | GO:0042796 | snRNA transcription from RNA polymerase III promoter(GO:0042796) |
| 0.2 | 2.2 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.2 | 2.6 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.2 | 2.0 | GO:0034351 | negative regulation of glial cell apoptotic process(GO:0034351) |
| 0.2 | 0.8 | GO:0002121 | inter-male aggressive behavior(GO:0002121) |
| 0.2 | 8.1 | GO:0030490 | maturation of SSU-rRNA(GO:0030490) |
| 0.2 | 0.8 | GO:0008616 | queuosine biosynthetic process(GO:0008616) queuosine metabolic process(GO:0046116) |
| 0.2 | 3.4 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.2 | 0.3 | GO:0060399 | positive regulation of growth hormone receptor signaling pathway(GO:0060399) |
| 0.2 | 0.8 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.2 | 2.2 | GO:0009083 | branched-chain amino acid catabolic process(GO:0009083) |
| 0.2 | 0.9 | GO:0070203 | regulation of establishment of protein localization to telomere(GO:0070203) positive regulation of establishment of protein localization to telomere(GO:1904851) |
| 0.1 | 8.5 | GO:0006364 | rRNA processing(GO:0006364) |
| 0.1 | 3.7 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.1 | 2.4 | GO:0052646 | alditol phosphate metabolic process(GO:0052646) |
| 0.1 | 2.0 | GO:0002176 | male germ cell proliferation(GO:0002176) |
| 0.1 | 5.4 | GO:0097031 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.1 | 0.3 | GO:0002362 | CD4-positive, CD25-positive, alpha-beta regulatory T cell lineage commitment(GO:0002362) |
| 0.1 | 2.7 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.1 | 8.2 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.1 | 5.5 | GO:0048255 | mRNA stabilization(GO:0048255) |
| 0.1 | 2.8 | GO:0031115 | negative regulation of microtubule polymerization(GO:0031115) |
| 0.1 | 0.4 | GO:0043461 | proton-transporting ATP synthase complex assembly(GO:0043461) proton-transporting ATP synthase complex biogenesis(GO:0070272) |
| 0.1 | 0.9 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.1 | 4.3 | GO:0006284 | base-excision repair(GO:0006284) |
| 0.1 | 0.9 | GO:0071971 | extracellular exosome assembly(GO:0071971) regulation of extracellular exosome assembly(GO:1903551) |
| 0.1 | 1.5 | GO:0018206 | peptidyl-methionine modification(GO:0018206) |
| 0.1 | 0.9 | GO:0043562 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.1 | 1.6 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.1 | 0.8 | GO:0035360 | positive regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035360) |
| 0.1 | 0.4 | GO:0000066 | mitochondrial ornithine transport(GO:0000066) |
| 0.1 | 0.1 | GO:0090365 | regulation of mRNA modification(GO:0090365) |
| 0.1 | 0.5 | GO:0071787 | endoplasmic reticulum tubular network assembly(GO:0071787) |
| 0.1 | 2.0 | GO:0045899 | positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.1 | 1.1 | GO:0098937 | dendritic transport(GO:0098935) anterograde dendritic transport(GO:0098937) |
| 0.1 | 0.8 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.1 | 0.9 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.1 | 1.5 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.1 | 0.3 | GO:1990927 | calcium ion regulated lysosome exocytosis(GO:1990927) |
| 0.1 | 0.5 | GO:0034197 | acylglycerol transport(GO:0034196) triglyceride transport(GO:0034197) |
| 0.1 | 0.3 | GO:2001160 | regulation of histone H3-K79 methylation(GO:2001160) |
| 0.1 | 1.3 | GO:0035067 | negative regulation of histone acetylation(GO:0035067) |
| 0.1 | 0.3 | GO:2000872 | positive regulation of progesterone secretion(GO:2000872) |
| 0.1 | 0.2 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 0.1 | 0.1 | GO:1904781 | positive regulation of protein localization to centrosome(GO:1904781) |
| 0.1 | 0.4 | GO:0070245 | positive regulation of thymocyte apoptotic process(GO:0070245) |
| 0.1 | 0.1 | GO:0060971 | embryonic heart tube left/right pattern formation(GO:0060971) |
| 0.1 | 0.5 | GO:0060613 | fat pad development(GO:0060613) |
| 0.1 | 1.5 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.1 | 0.4 | GO:0001306 | age-dependent response to oxidative stress(GO:0001306) age-dependent general metabolic decline(GO:0007571) |
| 0.1 | 0.5 | GO:0071955 | recycling endosome to Golgi transport(GO:0071955) |
| 0.1 | 0.5 | GO:0050747 | positive regulation of lipoprotein metabolic process(GO:0050747) |
| 0.1 | 0.9 | GO:0097034 | mitochondrial respiratory chain complex IV assembly(GO:0033617) mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.1 | 1.1 | GO:0090141 | positive regulation of mitochondrial fission(GO:0090141) |
| 0.1 | 0.3 | GO:0032914 | positive regulation of transforming growth factor beta1 production(GO:0032914) |
| 0.1 | 1.7 | GO:0060044 | negative regulation of cardiac muscle cell proliferation(GO:0060044) |
| 0.1 | 1.9 | GO:0042921 | glucocorticoid receptor signaling pathway(GO:0042921) |
| 0.1 | 0.4 | GO:0036112 | medium-chain fatty-acyl-CoA metabolic process(GO:0036112) |
| 0.1 | 0.7 | GO:0045046 | peroxisomal membrane transport(GO:0015919) protein import into peroxisome membrane(GO:0045046) |
| 0.1 | 9.2 | GO:0000910 | cytokinesis(GO:0000910) |
| 0.1 | 2.5 | GO:0034723 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.1 | 3.6 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.1 | 0.7 | GO:0035871 | protein K11-linked deubiquitination(GO:0035871) |
| 0.1 | 1.7 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.1 | 17.4 | GO:0000375 | RNA splicing, via transesterification reactions(GO:0000375) |
| 0.1 | 1.7 | GO:0019370 | leukotriene biosynthetic process(GO:0019370) |
| 0.1 | 0.5 | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 0.1 | 1.6 | GO:0071480 | cellular response to gamma radiation(GO:0071480) |
| 0.1 | 0.7 | GO:0006104 | succinyl-CoA metabolic process(GO:0006104) |
| 0.1 | 0.2 | GO:2001034 | positive regulation of double-strand break repair via nonhomologous end joining(GO:2001034) |
| 0.1 | 1.4 | GO:0031163 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.1 | 0.3 | GO:0072385 | minus-end-directed organelle transport along microtubule(GO:0072385) |
| 0.1 | 0.3 | GO:0036265 | 7-methylguanosine mRNA capping(GO:0006370) RNA (guanine-N7)-methylation(GO:0036265) |
| 0.1 | 0.9 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.1 | 0.2 | GO:0010040 | response to iron(II) ion(GO:0010040) |
| 0.1 | 0.2 | GO:0070200 | establishment of protein localization to telomere(GO:0070200) |
| 0.1 | 0.2 | GO:0034392 | negative regulation of smooth muscle cell apoptotic process(GO:0034392) |
| 0.1 | 1.0 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.1 | 0.6 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.1 | 3.3 | GO:0006401 | RNA catabolic process(GO:0006401) |
| 0.1 | 0.4 | GO:0033182 | regulation of histone ubiquitination(GO:0033182) |
| 0.1 | 0.7 | GO:0045719 | negative regulation of glycogen biosynthetic process(GO:0045719) |
| 0.1 | 1.1 | GO:0035024 | negative regulation of Rho protein signal transduction(GO:0035024) |
| 0.1 | 0.4 | GO:0010747 | positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) |
| 0.1 | 0.5 | GO:0090435 | protein localization to nuclear envelope(GO:0090435) |
| 0.1 | 0.9 | GO:1902751 | positive regulation of cell cycle G2/M phase transition(GO:1902751) |
| 0.1 | 0.1 | GO:0072236 | metanephric loop of Henle development(GO:0072236) |
| 0.1 | 2.6 | GO:0042254 | ribosome biogenesis(GO:0042254) |
| 0.1 | 0.7 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.1 | 0.2 | GO:2000210 | positive regulation of anoikis(GO:2000210) |
| 0.1 | 1.3 | GO:0042775 | mitochondrial ATP synthesis coupled electron transport(GO:0042775) |
| 0.1 | 0.6 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.1 | 0.2 | GO:0018197 | peptidyl-aspartic acid modification(GO:0018197) cellular response to manganese ion(GO:0071287) |
| 0.1 | 0.2 | GO:0000730 | DNA recombinase assembly(GO:0000730) double-strand break repair via synthesis-dependent strand annealing(GO:0045003) |
| 0.1 | 0.6 | GO:0060710 | chorio-allantoic fusion(GO:0060710) |
| 0.1 | 5.7 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.1 | 0.2 | GO:2000676 | positive regulation of type B pancreatic cell apoptotic process(GO:2000676) |
| 0.1 | 0.2 | GO:0009197 | dUDP biosynthetic process(GO:0006227) pyrimidine nucleoside diphosphate metabolic process(GO:0009138) pyrimidine nucleoside diphosphate biosynthetic process(GO:0009139) pyrimidine deoxyribonucleoside diphosphate metabolic process(GO:0009196) pyrimidine deoxyribonucleoside diphosphate biosynthetic process(GO:0009197) dUDP metabolic process(GO:0046077) |
| 0.1 | 0.8 | GO:2001241 | positive regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001241) |
| 0.1 | 0.5 | GO:0018344 | protein geranylgeranylation(GO:0018344) |
| 0.1 | 0.3 | GO:0019372 | lipoxygenase pathway(GO:0019372) |
| 0.1 | 4.6 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.1 | 0.4 | GO:0061002 | negative regulation of dendritic spine morphogenesis(GO:0061002) |
| 0.1 | 11.8 | GO:0007411 | axon guidance(GO:0007411) |
| 0.1 | 0.2 | GO:0048703 | embryonic viscerocranium morphogenesis(GO:0048703) |
| 0.1 | 0.7 | GO:0031100 | organ regeneration(GO:0031100) |
| 0.1 | 0.4 | GO:0010992 | ubiquitin homeostasis(GO:0010992) |
| 0.1 | 1.4 | GO:1901099 | negative regulation of signal transduction in absence of ligand(GO:1901099) negative regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001240) |
| 0.1 | 0.3 | GO:1900121 | negative regulation of receptor binding(GO:1900121) |
| 0.1 | 0.7 | GO:0030049 | muscle filament sliding(GO:0030049) |
| 0.1 | 0.4 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.0 | 0.7 | GO:0045943 | positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
| 0.0 | 21.3 | GO:0051301 | cell division(GO:0051301) |
| 0.0 | 0.6 | GO:0051085 | chaperone mediated protein folding requiring cofactor(GO:0051085) |
| 0.0 | 0.1 | GO:0061470 | T follicular helper cell differentiation(GO:0061470) |
| 0.0 | 0.7 | GO:0019430 | removal of superoxide radicals(GO:0019430) |
| 0.0 | 0.3 | GO:0051967 | negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.0 | 0.2 | GO:2001270 | regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001270) |
| 0.0 | 0.2 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.0 | 0.6 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.0 | 1.2 | GO:0006754 | ATP biosynthetic process(GO:0006754) |
| 0.0 | 0.3 | GO:0043568 | positive regulation of insulin-like growth factor receptor signaling pathway(GO:0043568) |
| 0.0 | 0.2 | GO:0033147 | negative regulation of intracellular estrogen receptor signaling pathway(GO:0033147) |
| 0.0 | 0.5 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.0 | 0.3 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.0 | 1.3 | GO:0006379 | mRNA cleavage(GO:0006379) |
| 0.0 | 0.2 | GO:0007062 | sister chromatid cohesion(GO:0007062) |
| 0.0 | 0.8 | GO:1900087 | positive regulation of G1/S transition of mitotic cell cycle(GO:1900087) |
| 0.0 | 1.2 | GO:0001541 | ovarian follicle development(GO:0001541) |
| 0.0 | 1.3 | GO:0043297 | apical junction assembly(GO:0043297) |
| 0.0 | 1.2 | GO:0034198 | cellular response to amino acid starvation(GO:0034198) |
| 0.0 | 0.0 | GO:0016441 | posttranscriptional gene silencing(GO:0016441) posttranscriptional gene silencing by RNA(GO:0035194) |
| 0.0 | 0.5 | GO:1904294 | positive regulation of ERAD pathway(GO:1904294) |
| 0.0 | 0.2 | GO:1903800 | positive regulation of production of miRNAs involved in gene silencing by miRNA(GO:1903800) |
| 0.0 | 16.0 | GO:0006412 | translation(GO:0006412) |
| 0.0 | 1.7 | GO:0018279 | peptidyl-asparagine modification(GO:0018196) protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.0 | 0.8 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.0 | 3.6 | GO:0019882 | antigen processing and presentation(GO:0019882) |
| 0.0 | 0.5 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.0 | 0.5 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 0.0 | 0.1 | GO:0009446 | putrescine biosynthetic process(GO:0009446) |
| 0.0 | 0.1 | GO:0006116 | NADH oxidation(GO:0006116) |
| 0.0 | 0.6 | GO:0045116 | protein neddylation(GO:0045116) |
| 0.0 | 0.4 | GO:0046520 | sphingosine biosynthetic process(GO:0046512) sphingoid biosynthetic process(GO:0046520) |
| 0.0 | 0.3 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.4 | GO:0007250 | activation of NF-kappaB-inducing kinase activity(GO:0007250) |
| 0.0 | 0.5 | GO:0070208 | protein heterotrimerization(GO:0070208) |
| 0.0 | 0.2 | GO:2000051 | negative regulation of non-canonical Wnt signaling pathway(GO:2000051) |
| 0.0 | 0.4 | GO:0090161 | Golgi ribbon formation(GO:0090161) |
| 0.0 | 0.1 | GO:0031293 | membrane protein intracellular domain proteolysis(GO:0031293) |
| 0.0 | 0.2 | GO:0010764 | negative regulation of fibroblast migration(GO:0010764) |
| 0.0 | 0.1 | GO:0006264 | mitochondrial DNA replication(GO:0006264) |
| 0.0 | 0.1 | GO:0006842 | tricarboxylic acid transport(GO:0006842) citrate transport(GO:0015746) |
| 0.0 | 0.1 | GO:0030579 | ubiquitin-dependent SMAD protein catabolic process(GO:0030579) |
| 0.0 | 0.9 | GO:0070206 | protein trimerization(GO:0070206) |
| 0.0 | 0.1 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 0.0 | 0.2 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.0 | 0.2 | GO:0034508 | centromere complex assembly(GO:0034508) |
| 0.0 | 0.3 | GO:0071361 | cellular response to ethanol(GO:0071361) |
| 0.0 | 0.2 | GO:0051443 | positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
| 0.0 | 0.3 | GO:0006105 | succinate metabolic process(GO:0006105) |
| 0.0 | 1.0 | GO:0002090 | regulation of receptor internalization(GO:0002090) |
| 0.0 | 0.0 | GO:0090343 | positive regulation of cell aging(GO:0090343) |
| 0.0 | 0.1 | GO:0097298 | regulation of nucleus size(GO:0097298) |
| 0.0 | 0.0 | GO:0090076 | relaxation of skeletal muscle(GO:0090076) |
| 0.0 | 1.1 | GO:0042475 | odontogenesis of dentin-containing tooth(GO:0042475) |
| 0.0 | 0.2 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.0 | 0.1 | GO:0036151 | phosphatidylcholine acyl-chain remodeling(GO:0036151) |
| 0.0 | 0.3 | GO:0097094 | craniofacial suture morphogenesis(GO:0097094) |
| 0.0 | 0.3 | GO:0006744 | ubiquinone metabolic process(GO:0006743) ubiquinone biosynthetic process(GO:0006744) |
| 0.0 | 1.1 | GO:0030500 | regulation of bone mineralization(GO:0030500) |
| 0.0 | 0.2 | GO:0090261 | positive regulation of inclusion body assembly(GO:0090261) |
| 0.0 | 2.1 | GO:1990823 | response to leukemia inhibitory factor(GO:1990823) cellular response to leukemia inhibitory factor(GO:1990830) |
| 0.0 | 0.3 | GO:0060632 | regulation of microtubule-based movement(GO:0060632) |
| 0.0 | 1.1 | GO:0050729 | positive regulation of inflammatory response(GO:0050729) |
| 0.0 | 0.2 | GO:0030889 | negative regulation of B cell proliferation(GO:0030889) |
| 0.0 | 0.4 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.1 | GO:0070234 | positive regulation of T cell apoptotic process(GO:0070234) |
| 0.0 | 0.0 | GO:0016185 | synaptic vesicle budding from presynaptic endocytic zone membrane(GO:0016185) synaptic vesicle budding(GO:0070142) |
| 0.0 | 0.3 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 0.1 | GO:2000270 | negative regulation of fibroblast apoptotic process(GO:2000270) |
| 0.0 | 0.3 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.0 | 0.1 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.0 | 2.0 | GO:0048167 | regulation of synaptic plasticity(GO:0048167) |
| 0.0 | 0.2 | GO:0010592 | positive regulation of lamellipodium assembly(GO:0010592) |
| 0.0 | 0.0 | GO:0042275 | error-free postreplication DNA repair(GO:0042275) |
| 0.0 | 0.2 | GO:0001709 | cell fate determination(GO:0001709) |
| 0.0 | 0.2 | GO:0030262 | apoptotic nuclear changes(GO:0030262) |
| 0.0 | 1.2 | GO:0007368 | determination of left/right symmetry(GO:0007368) |
| 0.0 | 0.1 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.0 | 0.0 | GO:0006999 | nuclear pore organization(GO:0006999) |
| 0.0 | 0.3 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 1.3 | GO:0002244 | hematopoietic progenitor cell differentiation(GO:0002244) |
| 0.0 | 0.0 | GO:0052422 | modulation by symbiont of host molecular function(GO:0052055) modulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052203) modulation by host of symbiont catalytic activity(GO:0052422) |
| 0.0 | 0.1 | GO:0080182 | histone H3-K4 trimethylation(GO:0080182) |
| 0.0 | 0.4 | GO:0045069 | regulation of viral genome replication(GO:0045069) |
| 0.0 | 0.1 | GO:0046007 | negative regulation of activated T cell proliferation(GO:0046007) |
| 0.0 | 0.8 | GO:0050728 | negative regulation of inflammatory response(GO:0050728) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.7 | 34.0 | GO:0032133 | chromosome passenger complex(GO:0032133) |
| 3.5 | 28.4 | GO:0005818 | aster(GO:0005818) |
| 3.1 | 9.4 | GO:0005656 | nuclear pre-replicative complex(GO:0005656) pre-replicative complex(GO:0036387) |
| 3.1 | 3.1 | GO:0070552 | BRISC complex(GO:0070552) |
| 3.0 | 20.9 | GO:0005655 | nucleolar ribonuclease P complex(GO:0005655) |
| 2.8 | 11.4 | GO:0000438 | core TFIIH complex portion of holo TFIIH complex(GO:0000438) |
| 2.2 | 6.5 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 1.9 | 9.5 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 1.9 | 11.4 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 1.8 | 23.5 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 1.7 | 17.0 | GO:0070545 | PeBoW complex(GO:0070545) |
| 1.5 | 15.1 | GO:0000445 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 1.3 | 3.9 | GO:0018444 | translation release factor complex(GO:0018444) |
| 1.2 | 6.2 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 1.2 | 3.6 | GO:0035101 | FACT complex(GO:0035101) |
| 1.1 | 9.0 | GO:0070652 | HAUS complex(GO:0070652) |
| 1.1 | 5.5 | GO:0071006 | U2-type catalytic step 1 spliceosome(GO:0071006) |
| 1.1 | 3.3 | GO:0071001 | U4/U6 snRNP(GO:0071001) |
| 1.0 | 23.0 | GO:0010369 | chromocenter(GO:0010369) |
| 1.0 | 5.2 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 1.0 | 8.2 | GO:0000275 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 1.0 | 3.0 | GO:1990423 | RZZ complex(GO:1990423) |
| 1.0 | 3.0 | GO:0031088 | platelet dense granule membrane(GO:0031088) |
| 1.0 | 4.9 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.9 | 4.7 | GO:0030689 | Noc complex(GO:0030689) |
| 0.9 | 5.5 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.9 | 11.4 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.8 | 5.9 | GO:0005638 | lamin filament(GO:0005638) |
| 0.8 | 4.0 | GO:0044615 | nuclear pore nuclear basket(GO:0044615) |
| 0.8 | 6.2 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.7 | 2.0 | GO:0098842 | postsynaptic early endosome(GO:0098842) |
| 0.7 | 2.0 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 0.6 | 1.9 | GO:0055087 | Ski complex(GO:0055087) |
| 0.6 | 10.2 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.6 | 2.5 | GO:0090661 | box H/ACA telomerase RNP complex(GO:0090661) |
| 0.6 | 4.3 | GO:0005956 | protein kinase CK2 complex(GO:0005956) |
| 0.6 | 1.8 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 0.6 | 6.1 | GO:0005750 | mitochondrial respiratory chain complex III(GO:0005750) |
| 0.6 | 15.2 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.6 | 6.6 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.5 | 5.9 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.5 | 5.4 | GO:0045298 | tubulin complex(GO:0045298) |
| 0.5 | 4.3 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.5 | 5.3 | GO:0034709 | methylosome(GO:0034709) |
| 0.5 | 3.3 | GO:0097255 | R2TP complex(GO:0097255) |
| 0.5 | 5.9 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.5 | 3.2 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.4 | 5.8 | GO:0042555 | MCM complex(GO:0042555) |
| 0.4 | 2.6 | GO:0000808 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.4 | 4.4 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.4 | 13.9 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.4 | 0.8 | GO:0005683 | U7 snRNP(GO:0005683) |
| 0.4 | 11.2 | GO:0000800 | lateral element(GO:0000800) |
| 0.4 | 25.2 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.4 | 1.1 | GO:0005673 | transcription factor TFIIE complex(GO:0005673) |
| 0.4 | 1.8 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.4 | 3.3 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.4 | 2.9 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.4 | 3.3 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.3 | 3.5 | GO:0000796 | condensin complex(GO:0000796) |
| 0.3 | 19.3 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.3 | 3.0 | GO:0070578 | RISC-loading complex(GO:0070578) |
| 0.3 | 2.0 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.3 | 5.8 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.3 | 3.6 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.3 | 2.1 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.3 | 1.5 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.3 | 14.8 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.3 | 2.4 | GO:0035748 | myelin sheath abaxonal region(GO:0035748) |
| 0.3 | 21.0 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.3 | 4.7 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.3 | 3.8 | GO:0008540 | proteasome regulatory particle, base subcomplex(GO:0008540) |
| 0.3 | 1.1 | GO:0033185 | dolichol-phosphate-mannose synthase complex(GO:0033185) |
| 0.3 | 2.8 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.3 | 1.9 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.3 | 1.3 | GO:0097361 | CIA complex(GO:0097361) |
| 0.3 | 1.3 | GO:0070044 | synaptobrevin 2-SNAP-25-syntaxin-1a complex(GO:0070044) |
| 0.3 | 6.9 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.2 | 2.0 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.2 | 4.3 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.2 | 9.2 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.2 | 0.7 | GO:0031251 | PAN complex(GO:0031251) |
| 0.2 | 3.5 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.2 | 0.9 | GO:0043259 | laminin-1 complex(GO:0005606) laminin-10 complex(GO:0043259) |
| 0.2 | 2.6 | GO:0005838 | proteasome regulatory particle(GO:0005838) |
| 0.2 | 6.9 | GO:0043596 | nuclear replication fork(GO:0043596) |
| 0.2 | 4.3 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.2 | 1.1 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.2 | 2.7 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.2 | 28.4 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.2 | 1.8 | GO:0071014 | post-mRNA release spliceosomal complex(GO:0071014) |
| 0.2 | 2.8 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.2 | 1.2 | GO:0030062 | mitochondrial tricarboxylic acid cycle enzyme complex(GO:0030062) |
| 0.2 | 4.2 | GO:0031305 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.2 | 1.8 | GO:0090543 | Flemming body(GO:0090543) |
| 0.2 | 2.9 | GO:0032039 | integrator complex(GO:0032039) |
| 0.2 | 2.6 | GO:0031315 | extrinsic component of mitochondrial outer membrane(GO:0031315) |
| 0.2 | 7.0 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.2 | 1.5 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.2 | 14.3 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.2 | 0.6 | GO:1990047 | spindle matrix(GO:1990047) |
| 0.2 | 2.0 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
| 0.2 | 1.3 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.2 | 17.1 | GO:0000922 | spindle pole(GO:0000922) |
| 0.2 | 3.3 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.2 | 6.2 | GO:0015030 | Cajal body(GO:0015030) |
| 0.2 | 0.4 | GO:0001739 | sex chromatin(GO:0001739) |
| 0.2 | 0.9 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.2 | 0.5 | GO:0070877 | microprocessor complex(GO:0070877) ribonuclease III complex(GO:1903095) |
| 0.2 | 3.1 | GO:0045277 | respiratory chain complex IV(GO:0045277) |
| 0.2 | 3.4 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.2 | 17.4 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.2 | 0.9 | GO:0005863 | striated muscle myosin thick filament(GO:0005863) |
| 0.2 | 4.7 | GO:0015935 | small ribosomal subunit(GO:0015935) |
| 0.2 | 4.7 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.2 | 2.0 | GO:0070938 | contractile ring(GO:0070938) |
| 0.2 | 1.4 | GO:0071541 | eukaryotic translation initiation factor 3 complex, eIF3m(GO:0071541) |
| 0.2 | 0.9 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.2 | 1.4 | GO:0030478 | actin cap(GO:0030478) |
| 0.1 | 0.6 | GO:0031523 | Myb complex(GO:0031523) |
| 0.1 | 0.7 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.1 | 0.9 | GO:0016461 | unconventional myosin complex(GO:0016461) |
| 0.1 | 0.4 | GO:0031372 | UBC13-MMS2 complex(GO:0031372) |
| 0.1 | 17.6 | GO:0000781 | chromosome, telomeric region(GO:0000781) |
| 0.1 | 4.2 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.1 | 2.5 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.1 | 6.0 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 1.5 | GO:0071004 | U2-type prespliceosome(GO:0071004) |
| 0.1 | 0.9 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.1 | 0.6 | GO:0000313 | organellar ribosome(GO:0000313) mitochondrial ribosome(GO:0005761) |
| 0.1 | 5.5 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 1.3 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.1 | 1.6 | GO:0030122 | AP-2 adaptor complex(GO:0030122) |
| 0.1 | 5.1 | GO:0005747 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.1 | 2.0 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.1 | 2.7 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.1 | 1.6 | GO:0098800 | inner mitochondrial membrane protein complex(GO:0098800) |
| 0.1 | 1.8 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.1 | 6.9 | GO:0005819 | spindle(GO:0005819) |
| 0.1 | 0.7 | GO:0031415 | NatA complex(GO:0031415) |
| 0.1 | 0.5 | GO:0005968 | Rab-protein geranylgeranyltransferase complex(GO:0005968) |
| 0.1 | 0.3 | GO:0031533 | mRNA cap methyltransferase complex(GO:0031533) |
| 0.1 | 1.6 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) |
| 0.1 | 1.6 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.1 | 0.8 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.1 | 0.5 | GO:0071256 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 0.1 | 0.7 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.1 | 6.9 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.1 | 2.4 | GO:0002080 | acrosomal membrane(GO:0002080) |
| 0.1 | 0.2 | GO:0033061 | DNA recombinase mediator complex(GO:0033061) |
| 0.1 | 0.2 | GO:0031166 | integral component of vacuolar membrane(GO:0031166) intrinsic component of vacuolar membrane(GO:0031310) |
| 0.1 | 1.6 | GO:0043196 | varicosity(GO:0043196) |
| 0.1 | 0.5 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 0.1 | 0.3 | GO:0089701 | U2AF(GO:0089701) |
| 0.1 | 3.0 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.1 | 0.5 | GO:0071203 | WASH complex(GO:0071203) |
| 0.1 | 0.9 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.1 | 0.7 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.1 | 0.5 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.1 | 0.3 | GO:0032983 | kainate selective glutamate receptor complex(GO:0032983) |
| 0.1 | 0.1 | GO:0071598 | neuronal ribonucleoprotein granule(GO:0071598) |
| 0.1 | 5.5 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.1 | 0.5 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.0 | 2.3 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 2.8 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 37.6 | GO:0005730 | nucleolus(GO:0005730) |
| 0.0 | 1.2 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 0.1 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.0 | 0.3 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.0 | 0.7 | GO:0030008 | TRAPP complex(GO:0030008) |
| 0.0 | 0.8 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 0.0 | 2.6 | GO:0000776 | kinetochore(GO:0000776) |
| 0.0 | 8.4 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 0.1 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.0 | 14.5 | GO:0005813 | centrosome(GO:0005813) |
| 0.0 | 0.2 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.0 | 1.1 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.0 | 0.5 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.0 | 0.5 | GO:0005839 | proteasome core complex(GO:0005839) |
| 0.0 | 2.4 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 2.4 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 0.1 | GO:0097543 | ciliary inversin compartment(GO:0097543) |
| 0.0 | 1.3 | GO:0031672 | A band(GO:0031672) |
| 0.0 | 0.8 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 37.4 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.0 | 0.3 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.3 | GO:1990204 | oxidoreductase complex(GO:1990204) |
| 0.0 | 0.8 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.4 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.0 | 0.3 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 0.3 | GO:0044665 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.0 | 0.0 | GO:0097165 | nuclear stress granule(GO:0097165) |
| 0.0 | 0.7 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 0.4 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.0 | 1.9 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.0 | 0.1 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.0 | 0.6 | GO:0032420 | stereocilium(GO:0032420) |
| 0.0 | 0.3 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.0 | 0.0 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 0.0 | 0.5 | GO:0030315 | T-tubule(GO:0030315) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.1 | 18.4 | GO:0008518 | reduced folate carrier activity(GO:0008518) |
| 3.0 | 26.7 | GO:0061676 | importin-alpha family protein binding(GO:0061676) |
| 2.4 | 16.5 | GO:0048256 | flap endonuclease activity(GO:0048256) |
| 2.0 | 12.2 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 1.9 | 20.8 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 1.6 | 11.2 | GO:0000150 | recombinase activity(GO:0000150) |
| 1.6 | 8.0 | GO:0044020 | histone methyltransferase activity (H4-R3 specific)(GO:0044020) |
| 1.4 | 5.7 | GO:0000700 | mismatch base pair DNA N-glycosylase activity(GO:0000700) |
| 1.3 | 8.0 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 1.3 | 6.4 | GO:0000702 | oxidized base lesion DNA N-glycosylase activity(GO:0000702) |
| 1.2 | 3.6 | GO:0000179 | rRNA (adenine-N6,N6-)-dimethyltransferase activity(GO:0000179) rRNA (adenine-N6-)-methyltransferase activity(GO:0008988) |
| 1.2 | 8.2 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 1.1 | 7.9 | GO:0000403 | Y-form DNA binding(GO:0000403) |
| 1.1 | 10.1 | GO:0003910 | DNA ligase activity(GO:0003909) DNA ligase (ATP) activity(GO:0003910) |
| 1.1 | 5.3 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 1.1 | 4.2 | GO:0004832 | valine-tRNA ligase activity(GO:0004832) |
| 1.0 | 8.1 | GO:0009019 | tRNA (guanine-N1-)-methyltransferase activity(GO:0009019) |
| 1.0 | 44.6 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 1.0 | 13.5 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.9 | 6.1 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.9 | 2.6 | GO:0031871 | proteinase activated receptor binding(GO:0031871) |
| 0.8 | 3.3 | GO:0010484 | H3 histone acetyltransferase activity(GO:0010484) |
| 0.8 | 18.6 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.8 | 3.0 | GO:0070883 | pre-miRNA binding(GO:0070883) |
| 0.7 | 3.0 | GO:0005220 | inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.7 | 2.2 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.7 | 2.1 | GO:0017099 | very-long-chain-acyl-CoA dehydrogenase activity(GO:0017099) |
| 0.7 | 10.2 | GO:0001054 | RNA polymerase I activity(GO:0001054) |
| 0.7 | 5.3 | GO:0008121 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.6 | 3.8 | GO:0035184 | histone threonine kinase activity(GO:0035184) |
| 0.6 | 3.0 | GO:0032767 | copper-dependent protein binding(GO:0032767) |
| 0.6 | 4.1 | GO:0003747 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.6 | 2.3 | GO:0004427 | inorganic diphosphatase activity(GO:0004427) |
| 0.6 | 14.8 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.6 | 6.1 | GO:0036310 | annealing helicase activity(GO:0036310) |
| 0.5 | 4.3 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.5 | 9.0 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.5 | 2.1 | GO:0003836 | beta-galactoside (CMP) alpha-2,3-sialyltransferase activity(GO:0003836) |
| 0.5 | 1.5 | GO:0035827 | rubidium ion transmembrane transporter activity(GO:0035827) |
| 0.5 | 1.5 | GO:0004140 | dephospho-CoA kinase activity(GO:0004140) |
| 0.5 | 5.4 | GO:0016427 | tRNA (cytosine) methyltransferase activity(GO:0016427) |
| 0.5 | 8.1 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.5 | 3.3 | GO:0015234 | thiamine transmembrane transporter activity(GO:0015234) |
| 0.5 | 0.9 | GO:0030620 | U2 snRNA binding(GO:0030620) |
| 0.5 | 4.2 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.5 | 4.6 | GO:0008174 | mRNA methyltransferase activity(GO:0008174) |
| 0.4 | 7.2 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.4 | 1.7 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
| 0.4 | 6.9 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.4 | 10.9 | GO:0008483 | transaminase activity(GO:0008483) |
| 0.4 | 9.2 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.4 | 1.3 | GO:0071209 | histone pre-mRNA DCP binding(GO:0071208) U7 snRNA binding(GO:0071209) |
| 0.4 | 1.2 | GO:0004793 | threonine aldolase activity(GO:0004793) L-allo-threonine aldolase activity(GO:0008732) |
| 0.4 | 0.8 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
| 0.4 | 2.7 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.4 | 3.8 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 0.4 | 12.0 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.4 | 1.8 | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity(GO:0004367) |
| 0.4 | 2.5 | GO:0034513 | box H/ACA snoRNA binding(GO:0034513) |
| 0.4 | 3.5 | GO:0097157 | pre-mRNA intronic binding(GO:0097157) |
| 0.4 | 2.8 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.3 | 1.7 | GO:0004301 | epoxide hydrolase activity(GO:0004301) |
| 0.3 | 2.1 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.3 | 1.3 | GO:0005483 | soluble NSF attachment protein activity(GO:0005483) |
| 0.3 | 3.3 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.3 | 2.1 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.3 | 8.8 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 0.3 | 0.9 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.3 | 1.1 | GO:0004582 | dolichyl-phosphate beta-D-mannosyltransferase activity(GO:0004582) |
| 0.3 | 2.0 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.3 | 2.5 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.3 | 1.6 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.3 | 1.8 | GO:0036402 | proteasome-activating ATPase activity(GO:0036402) |
| 0.3 | 1.0 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.3 | 6.2 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.3 | 2.1 | GO:0031749 | D2 dopamine receptor binding(GO:0031749) |
| 0.3 | 40.6 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.2 | 13.9 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.2 | 2.4 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.2 | 2.9 | GO:0003906 | DNA-(apurinic or apyrimidinic site) lyase activity(GO:0003906) |
| 0.2 | 1.4 | GO:0016300 | tRNA (uracil) methyltransferase activity(GO:0016300) |
| 0.2 | 0.7 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 0.2 | 0.7 | GO:0004774 | succinate-CoA ligase activity(GO:0004774) |
| 0.2 | 0.4 | GO:0031493 | nucleosomal histone binding(GO:0031493) |
| 0.2 | 2.2 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.2 | 0.9 | GO:0051880 | G-quadruplex DNA binding(GO:0051880) |
| 0.2 | 7.9 | GO:0031369 | translation initiation factor binding(GO:0031369) |
| 0.2 | 1.2 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.2 | 1.4 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.2 | 2.4 | GO:0000339 | RNA cap binding(GO:0000339) |
| 0.2 | 3.6 | GO:0048038 | quinone binding(GO:0048038) |
| 0.2 | 6.5 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.2 | 2.7 | GO:0001163 | RNA polymerase I regulatory region sequence-specific DNA binding(GO:0001163) RNA polymerase I CORE element sequence-specific DNA binding(GO:0001164) |
| 0.2 | 8.8 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.2 | 1.9 | GO:0015266 | protein channel activity(GO:0015266) |
| 0.2 | 4.5 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.2 | 13.7 | GO:0048365 | Rac GTPase binding(GO:0048365) |
| 0.2 | 0.9 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.2 | 2.0 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.2 | 0.8 | GO:0008061 | chitin binding(GO:0008061) |
| 0.2 | 1.1 | GO:1990446 | U1 snRNP binding(GO:1990446) |
| 0.2 | 1.8 | GO:0019213 | deacetylase activity(GO:0019213) |
| 0.2 | 1.5 | GO:0098599 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.2 | 4.1 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) |
| 0.2 | 4.1 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.1 | 1.5 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.1 | 5.3 | GO:0046966 | thyroid hormone receptor binding(GO:0046966) |
| 0.1 | 0.8 | GO:0004311 | farnesyltranstransferase activity(GO:0004311) |
| 0.1 | 1.7 | GO:0000182 | rDNA binding(GO:0000182) |
| 0.1 | 0.8 | GO:0008479 | queuine tRNA-ribosyltransferase activity(GO:0008479) |
| 0.1 | 4.8 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.1 | 0.3 | GO:0008177 | succinate dehydrogenase (ubiquinone) activity(GO:0008177) oxidoreductase activity, acting on the CH-CH group of donors, quinone or related compound as acceptor(GO:0016635) |
| 0.1 | 6.8 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.1 | 2.8 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.1 | 0.5 | GO:0032296 | ribonuclease III activity(GO:0004525) double-stranded RNA-specific ribonuclease activity(GO:0032296) |
| 0.1 | 0.8 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.1 | 0.9 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.1 | 4.7 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.1 | 1.2 | GO:0070034 | telomerase RNA binding(GO:0070034) |
| 0.1 | 0.5 | GO:0008297 | single-stranded DNA exodeoxyribonuclease activity(GO:0008297) |
| 0.1 | 3.1 | GO:0004129 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.1 | 2.3 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.1 | 7.1 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.1 | 2.3 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 1.7 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.1 | 0.2 | GO:0044547 | DNA topoisomerase binding(GO:0044547) |
| 0.1 | 0.4 | GO:0016508 | long-chain-enoyl-CoA hydratase activity(GO:0016508) |
| 0.1 | 2.4 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.1 | 1.0 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.1 | 1.7 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.1 | 0.9 | GO:0043208 | glycosphingolipid binding(GO:0043208) |
| 0.1 | 3.3 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.1 | 7.5 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.1 | 0.7 | GO:0017169 | CDP-alcohol phosphatidyltransferase activity(GO:0017169) |
| 0.1 | 2.0 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.1 | 0.3 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.1 | 14.0 | GO:0016810 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds(GO:0016810) |
| 0.1 | 3.6 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.1 | 1.7 | GO:0004697 | protein kinase C activity(GO:0004697) |
| 0.1 | 0.8 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 2.2 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.1 | 1.3 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.1 | 1.2 | GO:0046527 | glucosyltransferase activity(GO:0046527) |
| 0.1 | 0.9 | GO:0010181 | FMN binding(GO:0010181) |
| 0.1 | 0.4 | GO:0000064 | L-ornithine transmembrane transporter activity(GO:0000064) |
| 0.1 | 0.8 | GO:0030955 | potassium ion binding(GO:0030955) |
| 0.1 | 0.4 | GO:0004809 | tRNA (guanine-N2-)-methyltransferase activity(GO:0004809) |
| 0.1 | 1.0 | GO:0015368 | calcium:cation antiporter activity(GO:0015368) |
| 0.1 | 1.1 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.1 | 0.3 | GO:0034235 | GPI anchor binding(GO:0034235) |
| 0.1 | 0.3 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 0.1 | 1.0 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.1 | 0.6 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.1 | 2.7 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.1 | 2.2 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.1 | 2.9 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.1 | 0.2 | GO:0097003 | adipokinetic hormone receptor activity(GO:0097003) |
| 0.1 | 0.5 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.1 | 0.3 | GO:0034190 | apolipoprotein receptor binding(GO:0034190) |
| 0.1 | 0.2 | GO:0009041 | uridylate kinase activity(GO:0009041) |
| 0.1 | 0.1 | GO:0032135 | DNA insertion or deletion binding(GO:0032135) |
| 0.1 | 8.1 | GO:0004386 | helicase activity(GO:0004386) |
| 0.1 | 0.8 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.1 | 0.4 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.0 | 1.1 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.0 | 65.8 | GO:0003723 | RNA binding(GO:0003723) |
| 0.0 | 0.1 | GO:0000995 | transcription factor activity, core RNA polymerase III binding(GO:0000995) |
| 0.0 | 1.6 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 1.1 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.0 | 0.4 | GO:0031386 | protein tag(GO:0031386) |
| 0.0 | 1.0 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.0 | 0.3 | GO:0097027 | ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.0 | 0.3 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.0 | 0.3 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.0 | 0.5 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.0 | 2.5 | GO:0016651 | oxidoreductase activity, acting on NAD(P)H(GO:0016651) |
| 0.0 | 2.1 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.0 | 1.3 | GO:0008376 | acetylgalactosaminyltransferase activity(GO:0008376) |
| 0.0 | 0.8 | GO:0032452 | histone demethylase activity(GO:0032452) |
| 0.0 | 0.1 | GO:0030156 | benzodiazepine receptor binding(GO:0030156) |
| 0.0 | 0.4 | GO:0060229 | phospholipase activator activity(GO:0016004) lipase activator activity(GO:0060229) |
| 0.0 | 0.5 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.0 | 0.3 | GO:0004659 | prenyltransferase activity(GO:0004659) |
| 0.0 | 7.8 | GO:0015631 | tubulin binding(GO:0015631) |
| 0.0 | 0.5 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.0 | 3.7 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 0.3 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.0 | 1.8 | GO:0016879 | ligase activity, forming carbon-nitrogen bonds(GO:0016879) |
| 0.0 | 1.4 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.6 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.0 | 2.0 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.0 | 0.2 | GO:0015037 | peptide disulfide oxidoreductase activity(GO:0015037) |
| 0.0 | 0.3 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.0 | 0.1 | GO:0015137 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 0.0 | 2.3 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.3 | GO:0005234 | extracellular-glutamate-gated ion channel activity(GO:0005234) |
| 0.0 | 0.0 | GO:0000033 | alpha-1,3-mannosyltransferase activity(GO:0000033) |
| 0.0 | 0.7 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.4 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 14.9 | GO:0019904 | protein domain specific binding(GO:0019904) |
| 0.0 | 0.1 | GO:0004719 | protein-L-isoaspartate (D-aspartate) O-methyltransferase activity(GO:0004719) |
| 0.0 | 0.3 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 1.5 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.0 | 0.1 | GO:0019961 | interferon binding(GO:0019961) |
| 0.0 | 0.4 | GO:0016701 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen(GO:0016701) oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.0 | 3.6 | GO:0000287 | magnesium ion binding(GO:0000287) |
| 0.0 | 0.3 | GO:0015026 | coreceptor activity(GO:0015026) |
| 0.0 | 0.5 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 0.2 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 0.2 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.0 | 0.5 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.8 | GO:0008565 | protein transporter activity(GO:0008565) |
| 0.0 | 0.1 | GO:0008417 | fucosyltransferase activity(GO:0008417) |
| 0.0 | 3.0 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 0.1 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.0 | 1.0 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.2 | GO:0043175 | RNA polymerase core enzyme binding(GO:0043175) |
| 0.0 | 0.9 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 88.0 | PID AURORA A PATHWAY | Aurora A signaling |
| 1.0 | 41.8 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.5 | 27.3 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.4 | 6.5 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.3 | 7.2 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.3 | 27.8 | PID E2F PATHWAY | E2F transcription factor network |
| 0.3 | 7.9 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.3 | 10.8 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.2 | 0.8 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.1 | 5.6 | ST GAQ PATHWAY | G alpha q Pathway |
| 0.1 | 2.7 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.1 | 4.2 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 10.2 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.1 | 3.9 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.1 | 3.3 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.1 | 6.9 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.1 | 2.8 | PID PI3KCI AKT PATHWAY | Class I PI3K signaling events mediated by Akt |
| 0.1 | 2.3 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.1 | 2.1 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.1 | 0.9 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.1 | 0.9 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 3.3 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 5.6 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.1 | 2.2 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
| 0.1 | 3.1 | PID INSULIN PATHWAY | Insulin Pathway |
| 0.1 | 5.1 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.1 | 0.1 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.1 | 0.8 | ST ADRENERGIC | Adrenergic Pathway |
| 0.1 | 2.9 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.1 | 3.7 | PID P73PATHWAY | p73 transcription factor network |
| 0.1 | 2.6 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 0.8 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.1 | 1.8 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.0 | 1.0 | ST B CELL ANTIGEN RECEPTOR | B Cell Antigen Receptor |
| 0.0 | 0.8 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 0.6 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.0 | 0.6 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.0 | 0.9 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.0 | 1.3 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.0 | 0.2 | SIG CHEMOTAXIS | Genes related to chemotaxis |
| 0.0 | 0.6 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 0.7 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.0 | 0.3 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 1.3 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 0.5 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 0.6 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 0.3 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.0 | 0.2 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 27.4 | REACTOME PROCESSIVE SYNTHESIS ON THE LAGGING STRAND | Genes involved in Processive synthesis on the lagging strand |
| 1.3 | 18.4 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 1.2 | 25.1 | REACTOME INHIBITION OF THE PROTEOLYTIC ACTIVITY OF APC C REQUIRED FOR THE ONSET OF ANAPHASE BY MITOTIC SPINDLE CHECKPOINT COMPONENTS | Genes involved in Inhibition of the proteolytic activity of APC/C required for the onset of anaphase by mitotic spindle checkpoint components |
| 1.0 | 6.3 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 1.0 | 2.9 | REACTOME APC C CDC20 MEDIATED DEGRADATION OF CYCLIN B | Genes involved in APC/C:Cdc20 mediated degradation of Cyclin B |
| 0.9 | 15.2 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.9 | 2.6 | REACTOME PHOSPHORYLATION OF THE APC C | Genes involved in Phosphorylation of the APC/C |
| 0.9 | 22.2 | REACTOME TRANSPORT OF RIBONUCLEOPROTEINS INTO THE HOST NUCLEUS | Genes involved in Transport of Ribonucleoproteins into the Host Nucleus |
| 0.9 | 0.9 | REACTOME E2F ENABLED INHIBITION OF PRE REPLICATION COMPLEX FORMATION | Genes involved in E2F-enabled inhibition of pre-replication complex formation |
| 0.8 | 8.1 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.8 | 20.4 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |
| 0.7 | 7.9 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.7 | 70.6 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.6 | 11.0 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.6 | 2.4 | REACTOME FORMATION OF THE HIV1 EARLY ELONGATION COMPLEX | Genes involved in Formation of the HIV-1 Early Elongation Complex |
| 0.5 | 9.6 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.5 | 4.9 | REACTOME CDC6 ASSOCIATION WITH THE ORC ORIGIN COMPLEX | Genes involved in CDC6 association with the ORC:origin complex |
| 0.3 | 33.9 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.3 | 7.5 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.3 | 4.5 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.3 | 5.9 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.3 | 6.4 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.2 | 4.4 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.2 | 2.4 | REACTOME ELONGATION ARREST AND RECOVERY | Genes involved in Elongation arrest and recovery |
| 0.2 | 4.8 | REACTOME MICRORNA MIRNA BIOGENESIS | Genes involved in MicroRNA (miRNA) Biogenesis |
| 0.2 | 3.4 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.2 | 14.7 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.2 | 11.4 | REACTOME APC C CDH1 MEDIATED DEGRADATION OF CDC20 AND OTHER APC C CDH1 TARGETED PROTEINS IN LATE MITOSIS EARLY G1 | Genes involved in APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 |
| 0.2 | 0.8 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.2 | 2.7 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.2 | 4.0 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.2 | 3.7 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.2 | 4.5 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.2 | 4.5 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.2 | 1.9 | REACTOME P75NTR RECRUITS SIGNALLING COMPLEXES | Genes involved in p75NTR recruits signalling complexes |
| 0.2 | 2.2 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.2 | 2.6 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 0.3 | REACTOME MRNA CAPPING | Genes involved in mRNA Capping |
| 0.1 | 9.4 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.1 | 1.2 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.1 | 1.3 | REACTOME HOST INTERACTIONS OF HIV FACTORS | Genes involved in Host Interactions of HIV factors |
| 0.1 | 3.5 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.1 | 1.0 | REACTOME IRAK1 RECRUITS IKK COMPLEX | Genes involved in IRAK1 recruits IKK complex |
| 0.1 | 2.0 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.1 | 1.5 | REACTOME ACTIVATION OF THE MRNA UPON BINDING OF THE CAP BINDING COMPLEX AND EIFS AND SUBSEQUENT BINDING TO 43S | Genes involved in Activation of the mRNA upon binding of the cap-binding complex and eIFs, and subsequent binding to 43S |
| 0.1 | 2.4 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.1 | 0.7 | REACTOME ORC1 REMOVAL FROM CHROMATIN | Genes involved in Orc1 removal from chromatin |
| 0.1 | 1.9 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.1 | 1.9 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.1 | 1.5 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 1.2 | REACTOME EXTENSION OF TELOMERES | Genes involved in Extension of Telomeres |
| 0.1 | 1.9 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.1 | 2.1 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.1 | 1.4 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 0.1 | 2.6 | REACTOME METABOLISM OF STEROID HORMONES AND VITAMINS A AND D | Genes involved in Metabolism of steroid hormones and vitamins A and D |
| 0.1 | 7.3 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.1 | 2.6 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 1.3 | REACTOME RNA POL III TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase III Transcription Termination |
| 0.1 | 0.5 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 2 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 2 Promoter |
| 0.1 | 0.5 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.1 | 1.7 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.1 | 2.7 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.1 | 0.7 | REACTOME PREFOLDIN MEDIATED TRANSFER OF SUBSTRATE TO CCT TRIC | Genes involved in Prefoldin mediated transfer of substrate to CCT/TriC |
| 0.1 | 1.4 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 0.8 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.1 | 0.3 | REACTOME CYCLIN E ASSOCIATED EVENTS DURING G1 S TRANSITION | Genes involved in Cyclin E associated events during G1/S transition |
| 0.0 | 0.7 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.3 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.0 | 1.8 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 0.7 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.0 | 1.5 | REACTOME RNA POL I TRANSCRIPTION | Genes involved in RNA Polymerase I Transcription |
| 0.0 | 0.9 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.0 | 1.5 | REACTOME SIGNAL TRANSDUCTION BY L1 | Genes involved in Signal transduction by L1 |
| 0.0 | 1.2 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.0 | 0.5 | REACTOME NONSENSE MEDIATED DECAY ENHANCED BY THE EXON JUNCTION COMPLEX | Genes involved in Nonsense Mediated Decay Enhanced by the Exon Junction Complex |
| 0.0 | 1.0 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 0.8 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.0 | 0.9 | REACTOME FORMATION OF FIBRIN CLOT CLOTTING CASCADE | Genes involved in Formation of Fibrin Clot (Clotting Cascade) |
| 0.0 | 0.9 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.4 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.9 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 1.1 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.0 | 1.5 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 2.1 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.5 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 0.3 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 7ALPHA HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 7alpha-hydroxycholesterol |
| 0.0 | 0.2 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.0 | 2.7 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.0 | 4.1 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.1 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.0 | 0.3 | REACTOME PKA MEDIATED PHOSPHORYLATION OF CREB | Genes involved in PKA-mediated phosphorylation of CREB |
| 0.0 | 0.1 | REACTOME REGULATION OF ORNITHINE DECARBOXYLASE ODC | Genes involved in Regulation of ornithine decarboxylase (ODC) |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 1.0 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.0 | 0.1 | REACTOME TRANSCRIPTIONAL ACTIVITY OF SMAD2 SMAD3 SMAD4 HETEROTRIMER | Genes involved in Transcriptional activity of SMAD2/SMAD3:SMAD4 heterotrimer |
| 0.0 | 0.3 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 1.1 | REACTOME APOPTOTIC EXECUTION PHASE | Genes involved in Apoptotic execution phase |
| 0.0 | 0.1 | REACTOME DAG AND IP3 SIGNALING | Genes involved in DAG and IP3 signaling |
| 0.0 | 0.7 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 0.3 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |