Project
avrg: 2D miR_HR1_12
Navigation
Downloads
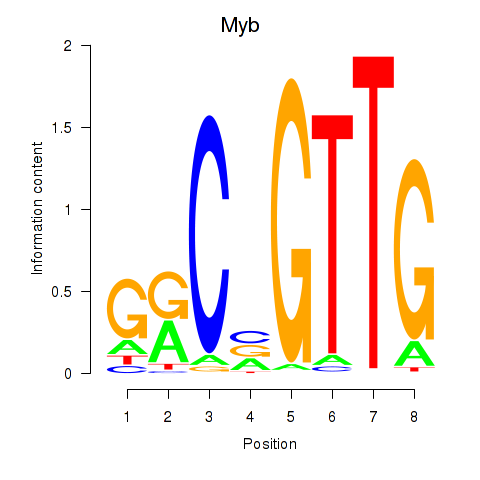
Results for Myb
Z-value: 5.05
Transcription factors associated with Myb
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Myb
|
ENSMUSG00000019982.8 | myeloblastosis oncogene |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Myb | mm10_v2_chr10_-_21160925_21160984 | 0.88 | 1.6e-04 | Click! |
Activity profile of Myb motif
Sorted Z-values of Myb motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_118437331 | 11.13 |
ENSMUST00000006565.6
|
Cdc20
|
cell division cycle 20 |
| chr2_+_118814195 | 9.92 |
ENSMUST00000110842.1
|
Knstrn
|
kinetochore-localized astrin/SPAG5 binding |
| chr2_+_118814237 | 8.94 |
ENSMUST00000028803.7
ENSMUST00000126045.1 |
Knstrn
|
kinetochore-localized astrin/SPAG5 binding |
| chr2_-_127831817 | 7.98 |
ENSMUST00000028858.7
|
Bub1
|
budding uninhibited by benzimidazoles 1 homolog (S. cerevisiae) |
| chr5_+_123749696 | 7.97 |
ENSMUST00000031366.7
|
Kntc1
|
kinetochore associated 1 |
| chr2_+_118813995 | 7.80 |
ENSMUST00000134661.1
|
Knstrn
|
kinetochore-localized astrin/SPAG5 binding |
| chr1_-_191575534 | 7.45 |
ENSMUST00000027933.5
|
Dtl
|
denticleless homolog (Drosophila) |
| chr17_-_33890584 | 6.51 |
ENSMUST00000114361.2
ENSMUST00000173492.1 |
Kifc1
|
kinesin family member C1 |
| chr6_+_124829540 | 6.40 |
ENSMUST00000150120.1
|
Cdca3
|
cell division cycle associated 3 |
| chr9_+_106281061 | 6.21 |
ENSMUST00000072206.6
|
Poc1a
|
POC1 centriolar protein homolog A (Chlamydomonas) |
| chrX_-_102157065 | 6.11 |
ENSMUST00000056904.2
|
Ercc6l
|
excision repair cross-complementing rodent repair deficiency complementation group 6 like |
| chrX_+_134308084 | 6.01 |
ENSMUST00000081064.5
ENSMUST00000101251.1 ENSMUST00000129782.1 |
Cenpi
|
centromere protein I |
| chr17_-_33890539 | 5.95 |
ENSMUST00000173386.1
|
Kifc1
|
kinesin family member C1 |
| chr6_+_124829582 | 5.91 |
ENSMUST00000024270.7
|
Cdca3
|
cell division cycle associated 3 |
| chr6_-_125191535 | 5.43 |
ENSMUST00000043848.4
|
Ncapd2
|
non-SMC condensin I complex, subunit D2 |
| chr17_+_26917091 | 5.37 |
ENSMUST00000078961.4
|
Kifc5b
|
kinesin family member C5B |
| chr11_+_103649498 | 5.14 |
ENSMUST00000057870.2
|
Rprml
|
reprimo-like |
| chr3_+_116594959 | 5.04 |
ENSMUST00000029571.8
|
Sass6
|
spindle assembly 6 homolog (C. elegans) |
| chr14_+_46760526 | 5.00 |
ENSMUST00000067426.4
|
Cdkn3
|
cyclin-dependent kinase inhibitor 3 |
| chr2_+_119047116 | 4.76 |
ENSMUST00000152380.1
ENSMUST00000099542.2 |
Casc5
|
cancer susceptibility candidate 5 |
| chr5_+_108132885 | 4.51 |
ENSMUST00000047677.7
|
Ccdc18
|
coiled-coil domain containing 18 |
| chr1_+_191821444 | 4.46 |
ENSMUST00000027931.7
|
Nek2
|
NIMA (never in mitosis gene a)-related expressed kinase 2 |
| chr13_+_51645232 | 4.42 |
ENSMUST00000075853.5
|
Cks2
|
CDC28 protein kinase regulatory subunit 2 |
| chr1_+_153425162 | 4.31 |
ENSMUST00000042373.5
|
Shcbp1l
|
Shc SH2-domain binding protein 1-like |
| chr2_-_172370506 | 4.29 |
ENSMUST00000109139.1
ENSMUST00000028997.7 ENSMUST00000109140.3 |
Aurka
|
aurora kinase A |
| chr5_-_8422582 | 4.15 |
ENSMUST00000168500.1
ENSMUST00000002368.9 |
Dbf4
|
DBF4 homolog (S. cerevisiae) |
| chr14_+_45351473 | 4.08 |
ENSMUST00000111835.2
|
Styx
|
serine/threonine/tyrosine interaction protein |
| chr5_-_8422695 | 4.08 |
ENSMUST00000171808.1
|
Dbf4
|
DBF4 homolog (S. cerevisiae) |
| chr2_+_119047129 | 4.05 |
ENSMUST00000153300.1
ENSMUST00000028799.5 |
Casc5
|
cancer susceptibility candidate 5 |
| chr19_-_41802028 | 3.91 |
ENSMUST00000026150.8
ENSMUST00000177495.1 ENSMUST00000163265.1 |
Arhgap19
|
Rho GTPase activating protein 19 |
| chr1_+_57995971 | 3.88 |
ENSMUST00000027202.8
|
Sgol2
|
shugoshin-like 2 (S. pombe) |
| chr18_+_34625009 | 3.86 |
ENSMUST00000166044.1
|
Kif20a
|
kinesin family member 20A |
| chr18_+_34624621 | 3.44 |
ENSMUST00000167161.1
|
Kif20a
|
kinesin family member 20A |
| chr11_-_69921057 | 3.41 |
ENSMUST00000108609.1
ENSMUST00000108608.1 ENSMUST00000164359.1 |
Eif5a
|
eukaryotic translation initiation factor 5A |
| chr1_-_131138232 | 3.39 |
ENSMUST00000016670.7
|
Dyrk3
|
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 3 |
| chr5_+_33658567 | 3.28 |
ENSMUST00000114426.3
|
Tacc3
|
transforming, acidic coiled-coil containing protein 3 |
| chr1_-_44101982 | 3.23 |
ENSMUST00000127923.1
|
Tex30
|
testis expressed 30 |
| chr17_-_6961156 | 3.12 |
ENSMUST00000063683.6
|
Tagap1
|
T cell activation GTPase activating protein 1 |
| chr8_+_83955507 | 3.12 |
ENSMUST00000005607.8
|
Asf1b
|
ASF1 anti-silencing function 1 homolog B (S. cerevisiae) |
| chr3_+_146404978 | 3.09 |
ENSMUST00000129978.1
|
Ssx2ip
|
synovial sarcoma, X breakpoint 2 interacting protein |
| chr7_+_126862431 | 3.09 |
ENSMUST00000132808.1
|
Hirip3
|
HIRA interacting protein 3 |
| chr11_-_69980468 | 3.06 |
ENSMUST00000143175.1
|
Elp5
|
elongator acetyltransferase complex subunit 5 |
| chr11_-_69921329 | 2.88 |
ENSMUST00000108613.3
ENSMUST00000043419.3 ENSMUST00000070996.4 |
Eif5a
|
eukaryotic translation initiation factor 5A |
| chr7_+_82174796 | 2.88 |
ENSMUST00000032874.7
|
Sh3gl3
|
SH3-domain GRB2-like 3 |
| chr11_-_77489666 | 2.88 |
ENSMUST00000037593.7
ENSMUST00000092892.3 |
Ankrd13b
|
ankyrin repeat domain 13b |
| chr3_+_146404631 | 2.78 |
ENSMUST00000106153.2
ENSMUST00000039021.4 ENSMUST00000106151.1 ENSMUST00000149262.1 |
Ssx2ip
|
synovial sarcoma, X breakpoint 2 interacting protein |
| chr6_-_131388417 | 2.77 |
ENSMUST00000032309.6
ENSMUST00000087865.2 |
Ybx3
|
Y box protein 3 |
| chr2_-_103796989 | 2.76 |
ENSMUST00000111147.1
|
Caprin1
|
cell cycle associated protein 1 |
| chr6_-_72439549 | 2.70 |
ENSMUST00000059472.8
|
Mat2a
|
methionine adenosyltransferase II, alpha |
| chr15_+_102073773 | 2.63 |
ENSMUST00000169681.1
|
Eif4b
|
eukaryotic translation initiation factor 4B |
| chr2_+_30807826 | 2.58 |
ENSMUST00000041830.3
ENSMUST00000152374.1 |
Ntmt1
|
N-terminal Xaa-Pro-Lys N-methyltransferase 1 |
| chr5_+_33658550 | 2.53 |
ENSMUST00000152847.1
|
Tacc3
|
transforming, acidic coiled-coil containing protein 3 |
| chr5_+_33658123 | 2.49 |
ENSMUST00000074849.6
ENSMUST00000079534.4 |
Tacc3
|
transforming, acidic coiled-coil containing protein 3 |
| chr8_+_83715177 | 2.48 |
ENSMUST00000019576.8
|
Ddx39
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 39 |
| chr14_-_46822232 | 2.46 |
ENSMUST00000111817.1
ENSMUST00000079314.5 |
Gmfb
|
glia maturation factor, beta |
| chr12_-_79192248 | 2.39 |
ENSMUST00000161204.1
|
Rdh11
|
retinol dehydrogenase 11 |
| chr8_-_31918203 | 2.39 |
ENSMUST00000073884.4
|
Nrg1
|
neuregulin 1 |
| chr1_-_44102414 | 2.33 |
ENSMUST00000143327.1
ENSMUST00000133677.1 |
Tex30
|
testis expressed 30 |
| chr4_-_83486178 | 2.33 |
ENSMUST00000130626.1
|
Psip1
|
PC4 and SFRS1 interacting protein 1 |
| chr11_-_69920892 | 2.28 |
ENSMUST00000152589.1
ENSMUST00000108612.1 ENSMUST00000108611.1 |
Eif5a
|
eukaryotic translation initiation factor 5A |
| chr1_-_57377476 | 2.25 |
ENSMUST00000181949.1
|
4930558J18Rik
|
RIKEN cDNA 4930558J18 gene |
| chr4_+_128993224 | 2.20 |
ENSMUST00000030583.6
ENSMUST00000102604.4 |
Ak2
|
adenylate kinase 2 |
| chr6_-_47594967 | 2.20 |
ENSMUST00000081721.6
ENSMUST00000114618.1 ENSMUST00000114616.1 |
Ezh2
|
enhancer of zeste homolog 2 (Drosophila) |
| chr7_+_126695942 | 2.16 |
ENSMUST00000106369.1
|
Bola2
|
bolA-like 2 (E. coli) |
| chr13_+_55464237 | 2.16 |
ENSMUST00000046533.7
|
Prr7
|
proline rich 7 (synaptic) |
| chr17_+_8165501 | 2.14 |
ENSMUST00000097419.3
ENSMUST00000024636.8 |
Fgfr1op
|
Fgfr1 oncogene partner |
| chr18_+_46597698 | 2.14 |
ENSMUST00000078079.3
ENSMUST00000168382.1 |
Eif1a
|
eukaryotic translation initiation factor 1A |
| chr3_-_88949906 | 2.06 |
ENSMUST00000172942.1
ENSMUST00000107491.4 |
Dap3
|
death associated protein 3 |
| chr19_-_9135603 | 2.05 |
ENSMUST00000049948.5
|
Asrgl1
|
asparaginase like 1 |
| chr3_+_146404844 | 2.03 |
ENSMUST00000106149.1
|
Ssx2ip
|
synovial sarcoma, X breakpoint 2 interacting protein |
| chr10_-_128565827 | 2.01 |
ENSMUST00000131728.1
ENSMUST00000026425.6 |
Pa2g4
|
proliferation-associated 2G4 |
| chr9_+_31030621 | 2.00 |
ENSMUST00000115222.2
|
Zbtb44
|
zinc finger and BTB domain containing 44 |
| chr17_-_46202576 | 1.99 |
ENSMUST00000024749.7
|
Polh
|
polymerase (DNA directed), eta (RAD 30 related) |
| chr1_-_44102362 | 1.98 |
ENSMUST00000147571.1
ENSMUST00000027215.5 ENSMUST00000147661.1 |
Tex30
|
testis expressed 30 |
| chr14_-_118923070 | 1.98 |
ENSMUST00000047208.5
|
Dzip1
|
DAZ interacting protein 1 |
| chr11_-_69921190 | 1.97 |
ENSMUST00000108607.1
|
Eif5a
|
eukaryotic translation initiation factor 5A |
| chr1_-_167285110 | 1.97 |
ENSMUST00000027839.8
|
Uck2
|
uridine-cytidine kinase 2 |
| chr7_-_45434590 | 1.97 |
ENSMUST00000107771.3
ENSMUST00000141761.1 |
Ruvbl2
|
RuvB-like protein 2 |
| chr1_-_44102433 | 1.93 |
ENSMUST00000129702.1
ENSMUST00000149502.1 ENSMUST00000156392.1 ENSMUST00000150911.1 |
Tex30
|
testis expressed 30 |
| chr10_+_128058974 | 1.92 |
ENSMUST00000084771.2
|
Ptges3
|
prostaglandin E synthase 3 (cytosolic) |
| chr7_+_126861947 | 1.92 |
ENSMUST00000037248.3
|
Hirip3
|
HIRA interacting protein 3 |
| chr9_+_47530173 | 1.90 |
ENSMUST00000114548.1
ENSMUST00000152459.1 ENSMUST00000143026.1 ENSMUST00000085909.2 ENSMUST00000114547.1 ENSMUST00000034581.3 |
Cadm1
|
cell adhesion molecule 1 |
| chr1_-_6215292 | 1.88 |
ENSMUST00000097832.1
|
4732440D04Rik
|
RIKEN cDNA 4732440D04 gene |
| chr3_-_88950271 | 1.87 |
ENSMUST00000174402.1
ENSMUST00000174077.1 |
Dap3
|
death associated protein 3 |
| chr8_+_83715504 | 1.85 |
ENSMUST00000109810.1
|
Ddx39
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 39 |
| chr1_-_44102341 | 1.82 |
ENSMUST00000128190.1
|
Tex30
|
testis expressed 30 |
| chr5_+_122372451 | 1.79 |
ENSMUST00000031420.4
|
Gpn3
|
GPN-loop GTPase 3 |
| chr4_-_116627921 | 1.78 |
ENSMUST00000030456.7
|
Nasp
|
nuclear autoantigenic sperm protein (histone-binding) |
| chr18_+_10617768 | 1.76 |
ENSMUST00000002551.3
|
Snrpd1
|
small nuclear ribonucleoprotein D1 |
| chr16_-_45724600 | 1.75 |
ENSMUST00000096057.4
|
Tagln3
|
transgelin 3 |
| chr17_+_27839974 | 1.72 |
ENSMUST00000071006.7
|
Snrpc
|
U1 small nuclear ribonucleoprotein C |
| chr2_-_104849465 | 1.71 |
ENSMUST00000126824.1
|
Prrg4
|
proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) |
| chr16_+_34690548 | 1.68 |
ENSMUST00000023532.6
|
Ccdc14
|
coiled-coil domain containing 14 |
| chr4_-_83486453 | 1.67 |
ENSMUST00000107214.2
ENSMUST00000107215.2 ENSMUST00000030207.8 |
Psip1
|
PC4 and SFRS1 interacting protein 1 |
| chr4_-_44167988 | 1.65 |
ENSMUST00000143337.1
|
Rnf38
|
ring finger protein 38 |
| chr10_+_128058947 | 1.64 |
ENSMUST00000052798.7
|
Ptges3
|
prostaglandin E synthase 3 (cytosolic) |
| chr18_+_67641589 | 1.64 |
ENSMUST00000025418.3
|
Psmg2
|
proteasome (prosome, macropain) assembly chaperone 2 |
| chr3_+_137864573 | 1.64 |
ENSMUST00000174561.1
ENSMUST00000173790.1 |
H2afz
|
H2A histone family, member Z |
| chr11_+_23666007 | 1.63 |
ENSMUST00000058163.4
|
Pus10
|
pseudouridylate synthase 10 |
| chr16_+_20674111 | 1.61 |
ENSMUST00000151679.1
|
Eif4g1
|
eukaryotic translation initiation factor 4, gamma 1 |
| chr3_+_82358056 | 1.59 |
ENSMUST00000091014.4
|
Map9
|
microtubule-associated protein 9 |
| chr16_+_64851991 | 1.58 |
ENSMUST00000067744.7
|
Cggbp1
|
CGG triplet repeat binding protein 1 |
| chr6_+_117906809 | 1.56 |
ENSMUST00000177918.1
ENSMUST00000163168.2 |
Hnrnpf
|
heterogeneous nuclear ribonucleoprotein F |
| chr18_+_69593361 | 1.54 |
ENSMUST00000114978.2
ENSMUST00000114977.1 |
Tcf4
|
transcription factor 4 |
| chr17_-_70851189 | 1.48 |
ENSMUST00000059775.8
|
Tgif1
|
TGFB-induced factor homeobox 1 |
| chr10_+_127677064 | 1.47 |
ENSMUST00000118612.1
ENSMUST00000048099.4 |
Tmem194
|
transmembrane protein 194 |
| chr18_+_56707725 | 1.47 |
ENSMUST00000025486.8
|
Lmnb1
|
lamin B1 |
| chr7_-_45204892 | 1.46 |
ENSMUST00000121017.2
|
Ccdc155
|
coiled-coil domain containing 155 |
| chr4_-_148626756 | 1.43 |
ENSMUST00000105699.1
|
Tardbp
|
TAR DNA binding protein |
| chr2_-_17731035 | 1.43 |
ENSMUST00000028080.5
|
Nebl
|
nebulette |
| chr10_-_41809607 | 1.43 |
ENSMUST00000019951.9
|
Cep57l1
|
centrosomal protein 57-like 1 |
| chr2_+_126152141 | 1.42 |
ENSMUST00000170908.1
|
Dtwd1
|
DTW domain containing 1 |
| chr6_+_89643982 | 1.41 |
ENSMUST00000000828.6
ENSMUST00000101171.1 |
Txnrd3
|
thioredoxin reductase 3 |
| chr7_+_29309429 | 1.41 |
ENSMUST00000137848.1
|
Dpf1
|
D4, zinc and double PHD fingers family 1 |
| chr11_+_23665615 | 1.37 |
ENSMUST00000109525.1
ENSMUST00000020520.4 |
Pus10
|
pseudouridylate synthase 10 |
| chr18_-_70530313 | 1.37 |
ENSMUST00000043286.8
|
Poli
|
polymerase (DNA directed), iota |
| chr4_+_149485260 | 1.35 |
ENSMUST00000030842.7
|
Lzic
|
leucine zipper and CTNNBIP1 domain containing |
| chr16_+_17070281 | 1.34 |
ENSMUST00000090199.3
|
Ypel1
|
yippee-like 1 (Drosophila) |
| chr4_-_116627478 | 1.33 |
ENSMUST00000081182.4
ENSMUST00000030457.5 |
Nasp
|
nuclear autoantigenic sperm protein (histone-binding) |
| chr4_+_149485215 | 1.32 |
ENSMUST00000124413.1
ENSMUST00000141293.1 |
Lzic
|
leucine zipper and CTNNBIP1 domain containing |
| chr3_+_103968588 | 1.31 |
ENSMUST00000156262.1
|
Phtf1
|
putative homeodomain transcription factor 1 |
| chr11_+_116434087 | 1.30 |
ENSMUST00000057676.6
|
Ubald2
|
UBA-like domain containing 2 |
| chrX_+_56786527 | 1.30 |
ENSMUST00000144600.1
|
Fhl1
|
four and a half LIM domains 1 |
| chr5_-_33652339 | 1.29 |
ENSMUST00000075670.6
|
Slbp
|
stem-loop binding protein |
| chr18_-_70530138 | 1.28 |
ENSMUST00000161542.1
ENSMUST00000159389.1 |
Poli
|
polymerase (DNA directed), iota |
| chr5_+_122158265 | 1.26 |
ENSMUST00000102528.4
ENSMUST00000086294.6 |
Ppp1cc
|
protein phosphatase 1, catalytic subunit, gamma isoform |
| chr7_+_44816088 | 1.26 |
ENSMUST00000057195.9
ENSMUST00000107891.1 |
Nup62
|
nucleoporin 62 |
| chr7_+_13024120 | 1.25 |
ENSMUST00000005705.7
|
Trim28
|
tripartite motif-containing 28 |
| chr5_+_135369942 | 1.24 |
ENSMUST00000000940.8
|
Nsun5
|
NOL1/NOP2/Sun domain family, member 5 |
| chr11_-_69920581 | 1.22 |
ENSMUST00000108610.1
|
Eif5a
|
eukaryotic translation initiation factor 5A |
| chrX_-_136741155 | 1.21 |
ENSMUST00000166930.1
ENSMUST00000113095.1 ENSMUST00000155207.1 ENSMUST00000080411.6 ENSMUST00000169418.1 |
Morf4l2
|
mortality factor 4 like 2 |
| chr17_-_80207299 | 1.19 |
ENSMUST00000063417.9
|
Srsf7
|
serine/arginine-rich splicing factor 7 |
| chr9_+_72438519 | 1.17 |
ENSMUST00000184604.1
|
Mns1
|
meiosis-specific nuclear structural protein 1 |
| chr15_+_57912199 | 1.16 |
ENSMUST00000022992.6
|
Tbc1d31
|
TBC1 domain family, member 31 |
| chr11_-_79962374 | 1.15 |
ENSMUST00000108241.1
ENSMUST00000043152.5 |
Utp6
|
UTP6, small subunit (SSU) processome component, homolog (yeast) |
| chr13_-_77131276 | 1.15 |
ENSMUST00000159300.1
|
Ankrd32
|
ankyrin repeat domain 32 |
| chr6_+_119479668 | 1.15 |
ENSMUST00000032094.5
|
Fbxl14
|
F-box and leucine-rich repeat protein 14 |
| chr8_+_83715239 | 1.14 |
ENSMUST00000172396.1
|
Ddx39
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 39 |
| chr10_-_22731336 | 1.14 |
ENSMUST00000127698.1
|
Tbpl1
|
TATA box binding protein-like 1 |
| chr1_+_10039762 | 1.14 |
ENSMUST00000122156.1
ENSMUST00000118263.1 ENSMUST00000119714.1 |
Cspp1
|
centrosome and spindle pole associated protein 1 |
| chr3_-_88950401 | 1.14 |
ENSMUST00000090938.4
|
Dap3
|
death associated protein 3 |
| chr2_-_92459709 | 1.13 |
ENSMUST00000136718.1
ENSMUST00000067631.6 |
Slc35c1
|
solute carrier family 35, member C1 |
| chr11_-_97782377 | 1.13 |
ENSMUST00000128801.1
|
Rpl23
|
ribosomal protein L23 |
| chr2_-_25224653 | 1.13 |
ENSMUST00000043584.4
|
Tubb4b
|
tubulin, beta 4B class IVB |
| chr8_+_84990585 | 1.12 |
ENSMUST00000064495.6
|
Hook2
|
hook homolog 2 (Drosophila) |
| chr2_-_5895319 | 1.12 |
ENSMUST00000026926.4
ENSMUST00000102981.3 |
Sec61a2
|
Sec61, alpha subunit 2 (S. cerevisiae) |
| chr19_-_44069736 | 1.12 |
ENSMUST00000172041.1
ENSMUST00000071698.6 ENSMUST00000112028.3 |
Erlin1
|
ER lipid raft associated 1 |
| chr2_+_29890534 | 1.11 |
ENSMUST00000113764.3
|
Odf2
|
outer dense fiber of sperm tails 2 |
| chr14_-_13961202 | 1.09 |
ENSMUST00000065865.8
|
Thoc7
|
THO complex 7 homolog (Drosophila) |
| chr2_-_165034821 | 1.09 |
ENSMUST00000153905.1
ENSMUST00000040381.8 |
Ncoa5
|
nuclear receptor coactivator 5 |
| chr11_+_104577281 | 1.06 |
ENSMUST00000106956.3
|
Myl4
|
myosin, light polypeptide 4 |
| chr6_-_28261907 | 1.05 |
ENSMUST00000115320.1
ENSMUST00000123098.1 ENSMUST00000115321.2 ENSMUST00000155494.1 |
Zfp800
|
zinc finger protein 800 |
| chr2_+_140395309 | 1.05 |
ENSMUST00000110067.1
ENSMUST00000110064.1 ENSMUST00000110063.1 ENSMUST00000110062.1 ENSMUST00000078027.5 ENSMUST00000043836.7 |
Macrod2
|
MACRO domain containing 2 |
| chr2_-_119477613 | 1.05 |
ENSMUST00000110808.1
ENSMUST00000049920.7 |
Ino80
|
INO80 homolog (S. cerevisiae) |
| chr2_+_68104671 | 1.04 |
ENSMUST00000042456.3
|
B3galt1
|
UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 1 |
| chr7_-_105399991 | 1.03 |
ENSMUST00000118726.1
ENSMUST00000074686.7 ENSMUST00000122327.1 ENSMUST00000179474.1 ENSMUST00000048079.6 |
Fam160a2
|
family with sequence similarity 160, member A2 |
| chr19_-_44069526 | 1.03 |
ENSMUST00000170801.1
|
Erlin1
|
ER lipid raft associated 1 |
| chr7_+_82175156 | 1.02 |
ENSMUST00000180243.1
|
Sh3gl3
|
SH3-domain GRB2-like 3 |
| chr11_-_97782409 | 1.02 |
ENSMUST00000103146.4
|
Rpl23
|
ribosomal protein L23 |
| chr4_+_125029992 | 0.99 |
ENSMUST00000030684.7
|
Gnl2
|
guanine nucleotide binding protein-like 2 (nucleolar) |
| chr15_+_100228229 | 0.98 |
ENSMUST00000171869.1
|
Atf1
|
activating transcription factor 1 |
| chr11_-_33513626 | 0.96 |
ENSMUST00000037522.7
|
Ranbp17
|
RAN binding protein 17 |
| chr1_+_171018920 | 0.95 |
ENSMUST00000078825.4
|
Fcgr4
|
Fc receptor, IgG, low affinity IV |
| chr11_-_120990871 | 0.95 |
ENSMUST00000154483.1
|
Csnk1d
|
casein kinase 1, delta |
| chr2_+_152687137 | 0.93 |
ENSMUST00000062148.6
|
Mcts2
|
malignant T cell amplified sequence 2 |
| chr10_-_22731918 | 0.92 |
ENSMUST00000095794.3
|
Tbpl1
|
TATA box binding protein-like 1 |
| chr9_-_44802951 | 0.92 |
ENSMUST00000044694.6
|
Ttc36
|
tetratricopeptide repeat domain 36 |
| chr18_-_67641329 | 0.90 |
ENSMUST00000097542.2
|
Cep76
|
centrosomal protein 76 |
| chr17_-_31855782 | 0.89 |
ENSMUST00000024839.4
|
Sik1
|
salt inducible kinase 1 |
| chrX_+_56894372 | 0.89 |
ENSMUST00000136396.1
|
Gpr112
|
G protein-coupled receptor 112 |
| chr6_+_48593883 | 0.88 |
ENSMUST00000154010.1
ENSMUST00000163452.1 ENSMUST00000118229.1 ENSMUST00000009420.8 |
Repin1
|
replication initiator 1 |
| chr11_+_22990519 | 0.88 |
ENSMUST00000173867.1
ENSMUST00000020562.4 |
Cct4
|
chaperonin containing Tcp1, subunit 4 (delta) |
| chr16_+_17070220 | 0.87 |
ENSMUST00000141959.1
|
Ypel1
|
yippee-like 1 (Drosophila) |
| chr14_-_62761112 | 0.86 |
ENSMUST00000053959.6
|
Ints6
|
integrator complex subunit 6 |
| chr2_-_62573905 | 0.86 |
ENSMUST00000102732.3
|
Fap
|
fibroblast activation protein |
| chr5_+_123252087 | 0.86 |
ENSMUST00000121964.1
|
Wdr66
|
WD repeat domain 66 |
| chr4_-_43010226 | 0.85 |
ENSMUST00000030165.4
|
Fancg
|
Fanconi anemia, complementation group G |
| chr6_+_116264186 | 0.85 |
ENSMUST00000036503.7
ENSMUST00000112900.3 |
Zfand4
|
zinc finger, AN1-type domain 4 |
| chr3_-_89214378 | 0.85 |
ENSMUST00000073572.4
|
Mtx1
|
metaxin 1 |
| chr17_-_35164891 | 0.83 |
ENSMUST00000025253.5
|
Prrc2a
|
proline-rich coiled-coil 2A |
| chr10_+_107271827 | 0.82 |
ENSMUST00000020057.8
ENSMUST00000105280.3 |
Lin7a
|
lin-7 homolog A (C. elegans) |
| chr10_-_8518801 | 0.82 |
ENSMUST00000061601.7
|
Ust
|
uronyl-2-sulfotransferase |
| chr3_-_90509450 | 0.81 |
ENSMUST00000107343.1
ENSMUST00000001043.7 ENSMUST00000107344.1 ENSMUST00000076639.4 ENSMUST00000107346.1 ENSMUST00000146740.1 ENSMUST00000107342.1 ENSMUST00000049937.6 |
Chtop
|
chromatin target of PRMT1 |
| chr10_+_84838143 | 0.80 |
ENSMUST00000095388.4
|
Rfx4
|
regulatory factor X, 4 (influences HLA class II expression) |
| chr2_+_60209887 | 0.79 |
ENSMUST00000102748.4
ENSMUST00000102747.1 |
March7
|
membrane-associated ring finger (C3HC4) 7 |
| chr15_+_80234071 | 0.78 |
ENSMUST00000023048.4
ENSMUST00000166030.1 |
Mief1
|
mitochondrial elongation factor 1 |
| chr14_-_31251194 | 0.77 |
ENSMUST00000022459.3
|
Phf7
|
PHD finger protein 7 |
| chr5_+_108065696 | 0.76 |
ENSMUST00000172045.1
|
Mtf2
|
metal response element binding transcription factor 2 |
| chr1_+_191025350 | 0.75 |
ENSMUST00000181050.1
|
A230020J21Rik
|
RIKEN cDNA A230020J21 gene |
| chr8_+_83389846 | 0.75 |
ENSMUST00000002259.6
|
Clgn
|
calmegin |
| chr17_+_46202740 | 0.74 |
ENSMUST00000087031.5
|
Xpo5
|
exportin 5 |
| chr4_+_152008803 | 0.74 |
ENSMUST00000097773.3
|
Klhl21
|
kelch-like 21 |
| chr2_-_165034770 | 0.74 |
ENSMUST00000122070.1
ENSMUST00000121377.1 |
Ncoa5
|
nuclear receptor coactivator 5 |
| chr4_-_95052188 | 0.73 |
ENSMUST00000107094.1
|
Jun
|
Jun oncogene |
| chr3_+_88297147 | 0.73 |
ENSMUST00000164166.1
ENSMUST00000168062.1 |
Cct3
|
chaperonin containing Tcp1, subunit 3 (gamma) |
| chr13_+_93308006 | 0.72 |
ENSMUST00000079086.6
|
Homer1
|
homer homolog 1 (Drosophila) |
| chr5_-_33652296 | 0.72 |
ENSMUST00000151081.1
ENSMUST00000101354.3 |
Slbp
|
stem-loop binding protein |
| chr17_+_31564749 | 0.72 |
ENSMUST00000175806.1
ENSMUST00000097352.3 |
Pknox1
|
Pbx/knotted 1 homeobox |
| chr3_-_89214329 | 0.71 |
ENSMUST00000118964.2
|
Mtx1
|
metaxin 1 |
| chr9_+_72438534 | 0.71 |
ENSMUST00000034746.8
|
Mns1
|
meiosis-specific nuclear structural protein 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Myb
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.2 | 12.5 | GO:0072382 | minus-end-directed vesicle transport along microtubule(GO:0072382) |
| 2.8 | 11.1 | GO:0098763 | mitotic cell cycle phase(GO:0098763) |
| 2.7 | 8.0 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 2.0 | 11.8 | GO:0045905 | translational frameshifting(GO:0006452) positive regulation of translational termination(GO:0045905) |
| 1.7 | 31.1 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 1.6 | 8.2 | GO:0007089 | traversing start control point of mitotic cell cycle(GO:0007089) |
| 1.4 | 4.3 | GO:0060282 | positive regulation of oocyte development(GO:0060282) |
| 1.4 | 4.1 | GO:0045204 | MAPK export from nucleus(GO:0045204) |
| 1.2 | 8.2 | GO:0035735 | intraciliary transport involved in cilium morphogenesis(GO:0035735) |
| 1.0 | 6.2 | GO:0003431 | growth plate cartilage chondrocyte development(GO:0003431) |
| 0.8 | 3.2 | GO:1902219 | negative regulation of intrinsic apoptotic signaling pathway in response to osmotic stress(GO:1902219) |
| 0.8 | 10.1 | GO:0019985 | translesion synthesis(GO:0019985) |
| 0.7 | 2.2 | GO:0071707 | immunoglobulin heavy chain V-D-J recombination(GO:0071707) |
| 0.7 | 5.7 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.7 | 5.4 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.7 | 3.4 | GO:0035617 | stress granule disassembly(GO:0035617) |
| 0.7 | 2.7 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.7 | 2.0 | GO:0009173 | UMP biosynthetic process(GO:0006222) pyrimidine ribonucleoside monophosphate metabolic process(GO:0009173) pyrimidine ribonucleoside monophosphate biosynthetic process(GO:0009174) UMP metabolic process(GO:0046049) |
| 0.6 | 2.6 | GO:0018201 | peptidyl-glycine modification(GO:0018201) |
| 0.6 | 1.9 | GO:0009826 | unidimensional cell growth(GO:0009826) susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.6 | 2.4 | GO:0016062 | adaptation of rhodopsin mediated signaling(GO:0016062) light adaption(GO:0036367) |
| 0.6 | 2.9 | GO:1902412 | regulation of mitotic cytokinesis(GO:1902412) |
| 0.5 | 2.1 | GO:0072717 | cellular response to actinomycin D(GO:0072717) |
| 0.5 | 2.1 | GO:0006235 | dTTP biosynthetic process(GO:0006235) pyrimidine deoxyribonucleoside triphosphate biosynthetic process(GO:0009212) dTTP metabolic process(GO:0046075) |
| 0.5 | 2.0 | GO:0071494 | cellular response to UV-C(GO:0071494) |
| 0.5 | 2.0 | GO:0071899 | regulation of estrogen receptor binding(GO:0071898) negative regulation of estrogen receptor binding(GO:0071899) |
| 0.5 | 8.3 | GO:0030953 | astral microtubule organization(GO:0030953) |
| 0.5 | 1.5 | GO:0090172 | microtubule cytoskeleton organization involved in homologous chromosome segregation(GO:0090172) |
| 0.4 | 1.8 | GO:1990022 | RNA polymerase II complex import to nucleus(GO:0044376) RNA polymerase III complex localization to nucleus(GO:1990022) |
| 0.4 | 1.2 | GO:1901535 | negative regulation of single stranded viral RNA replication via double stranded DNA intermediate(GO:0045869) regulation of DNA demethylation(GO:1901535) negative regulation of DNA demethylation(GO:1901536) |
| 0.4 | 7.3 | GO:0000920 | cell separation after cytokinesis(GO:0000920) |
| 0.4 | 1.5 | GO:1904009 | response to monosodium glutamate(GO:1904008) cellular response to monosodium glutamate(GO:1904009) |
| 0.4 | 2.2 | GO:0006172 | ADP biosynthetic process(GO:0006172) |
| 0.4 | 4.2 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 0.3 | 2.4 | GO:0045213 | neurotransmitter receptor metabolic process(GO:0045213) |
| 0.3 | 3.0 | GO:0031119 | tRNA pseudouridine synthesis(GO:0031119) |
| 0.3 | 2.1 | GO:0006528 | asparagine metabolic process(GO:0006528) |
| 0.3 | 0.9 | GO:0010710 | regulation of collagen catabolic process(GO:0010710) negative regulation of extracellular matrix disassembly(GO:0010716) |
| 0.3 | 2.1 | GO:0045541 | negative regulation of cholesterol biosynthetic process(GO:0045541) negative regulation of cholesterol metabolic process(GO:0090206) |
| 0.3 | 3.2 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.3 | 3.4 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.3 | 1.3 | GO:0061086 | negative regulation of histone H3-K27 methylation(GO:0061086) |
| 0.3 | 7.5 | GO:0007099 | centriole replication(GO:0007099) |
| 0.2 | 2.2 | GO:0043922 | negative regulation by host of viral transcription(GO:0043922) |
| 0.2 | 2.3 | GO:1904871 | protein localization to nuclear body(GO:1903405) protein localization to Cajal body(GO:1904867) regulation of protein localization to Cajal body(GO:1904869) positive regulation of protein localization to Cajal body(GO:1904871) protein localization to nucleoplasm(GO:1990173) |
| 0.2 | 2.0 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.2 | 0.6 | GO:1902951 | negative regulation of dendritic spine maintenance(GO:1902951) |
| 0.2 | 6.2 | GO:0034723 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.2 | 6.0 | GO:0034508 | centromere complex assembly(GO:0034508) |
| 0.2 | 0.7 | GO:1900368 | regulation of RNA interference(GO:1900368) |
| 0.2 | 0.4 | GO:0098989 | NMDA selective glutamate receptor signaling pathway(GO:0098989) |
| 0.2 | 0.9 | GO:0034627 | nicotinamide nucleotide biosynthetic process from aspartate(GO:0019355) 'de novo' NAD biosynthetic process(GO:0034627) 'de novo' NAD biosynthetic process from aspartate(GO:0034628) |
| 0.2 | 0.3 | GO:1903719 | regulation of I-kappaB phosphorylation(GO:1903719) positive regulation of I-kappaB phosphorylation(GO:1903721) |
| 0.2 | 0.5 | GO:0007315 | oocyte construction(GO:0007308) oocyte axis specification(GO:0007309) oocyte anterior/posterior axis specification(GO:0007314) pole plasm assembly(GO:0007315) maternal determination of anterior/posterior axis, embryo(GO:0008358) P granule organization(GO:0030719) |
| 0.2 | 1.5 | GO:0006682 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.2 | 0.8 | GO:1903361 | protein localization to basolateral plasma membrane(GO:1903361) |
| 0.2 | 0.5 | GO:0060830 | ciliary receptor clustering involved in smoothened signaling pathway(GO:0060830) |
| 0.2 | 0.8 | GO:0021914 | negative regulation of smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021914) |
| 0.2 | 3.0 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.2 | 1.2 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.2 | 0.5 | GO:0002353 | plasma kallikrein-kinin cascade(GO:0002353) |
| 0.2 | 1.1 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.1 | 0.4 | GO:0010615 | positive regulation of cardiac muscle adaptation(GO:0010615) positive regulation of cardiac muscle hypertrophy in response to stress(GO:1903244) |
| 0.1 | 0.4 | GO:1901978 | positive regulation of cell cycle checkpoint(GO:1901978) |
| 0.1 | 0.7 | GO:1903566 | positive regulation of protein localization to cilium(GO:1903566) |
| 0.1 | 1.6 | GO:2000507 | positive regulation of energy homeostasis(GO:2000507) |
| 0.1 | 1.2 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.1 | 0.4 | GO:0035521 | monoubiquitinated histone deubiquitination(GO:0035521) monoubiquitinated histone H2A deubiquitination(GO:0035522) |
| 0.1 | 1.3 | GO:0010972 | negative regulation of G2/M transition of mitotic cell cycle(GO:0010972) |
| 0.1 | 1.5 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.1 | 0.4 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.1 | 1.9 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.1 | 1.5 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 0.1 | 0.2 | GO:0061153 | trachea submucosa development(GO:0061152) trachea gland development(GO:0061153) |
| 0.1 | 0.6 | GO:1902966 | regulation of protein localization to early endosome(GO:1902965) positive regulation of protein localization to early endosome(GO:1902966) |
| 0.1 | 1.0 | GO:0010571 | positive regulation of nuclear cell cycle DNA replication(GO:0010571) |
| 0.1 | 1.6 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.1 | 1.1 | GO:0046784 | viral mRNA export from host cell nucleus(GO:0046784) |
| 0.1 | 4.6 | GO:0006406 | mRNA export from nucleus(GO:0006406) mRNA-containing ribonucleoprotein complex export from nucleus(GO:0071427) |
| 0.1 | 1.1 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.1 | 1.9 | GO:0034453 | microtubule anchoring(GO:0034453) |
| 0.1 | 1.5 | GO:0008608 | attachment of spindle microtubules to kinetochore(GO:0008608) |
| 0.1 | 0.3 | GO:0051866 | general adaptation syndrome(GO:0051866) |
| 0.1 | 1.6 | GO:0071392 | cellular response to estradiol stimulus(GO:0071392) |
| 0.1 | 2.2 | GO:0043153 | entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.1 | 0.7 | GO:0048752 | semicircular canal morphogenesis(GO:0048752) |
| 0.1 | 3.4 | GO:0045737 | positive regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045737) |
| 0.1 | 0.5 | GO:0043634 | polyadenylation-dependent ncRNA catabolic process(GO:0043634) |
| 0.1 | 1.2 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.1 | 0.2 | GO:0007529 | establishment of synaptic specificity at neuromuscular junction(GO:0007529) |
| 0.1 | 0.4 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.1 | 2.8 | GO:0061003 | positive regulation of dendritic spine morphogenesis(GO:0061003) |
| 0.1 | 0.4 | GO:0048208 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.1 | 0.7 | GO:0009158 | ribonucleoside monophosphate catabolic process(GO:0009158) purine ribonucleoside monophosphate catabolic process(GO:0009169) |
| 0.1 | 0.7 | GO:0044380 | protein localization to cytoskeleton(GO:0044380) |
| 0.1 | 5.9 | GO:0007093 | mitotic cell cycle checkpoint(GO:0007093) |
| 0.1 | 2.2 | GO:0045022 | early endosome to late endosome transport(GO:0045022) |
| 0.1 | 0.3 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) |
| 0.1 | 1.8 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 2.0 | GO:0051220 | cytoplasmic sequestering of protein(GO:0051220) |
| 0.0 | 0.7 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.0 | 0.8 | GO:0090141 | positive regulation of mitochondrial fission(GO:0090141) |
| 0.0 | 1.1 | GO:0032467 | positive regulation of cytokinesis(GO:0032467) |
| 0.0 | 0.8 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 0.4 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.0 | 1.5 | GO:0002026 | regulation of the force of heart contraction(GO:0002026) |
| 0.0 | 1.3 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.0 | 0.3 | GO:0051001 | negative regulation of nitric-oxide synthase activity(GO:0051001) |
| 0.0 | 1.4 | GO:0000305 | response to oxygen radical(GO:0000305) |
| 0.0 | 5.0 | GO:0008637 | apoptotic mitochondrial changes(GO:0008637) |
| 0.0 | 1.6 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.0 | 0.6 | GO:0001865 | NK T cell differentiation(GO:0001865) |
| 0.0 | 1.0 | GO:0042119 | neutrophil activation(GO:0042119) |
| 0.0 | 0.9 | GO:2001273 | regulation of glucose import in response to insulin stimulus(GO:2001273) |
| 0.0 | 1.8 | GO:0046627 | negative regulation of insulin receptor signaling pathway(GO:0046627) |
| 0.0 | 0.1 | GO:0097029 | mature conventional dendritic cell differentiation(GO:0097029) |
| 0.0 | 0.3 | GO:0000712 | resolution of meiotic recombination intermediates(GO:0000712) |
| 0.0 | 1.4 | GO:0000070 | mitotic sister chromatid segregation(GO:0000070) |
| 0.0 | 1.1 | GO:0000086 | G2/M transition of mitotic cell cycle(GO:0000086) |
| 0.0 | 3.8 | GO:0000082 | G1/S transition of mitotic cell cycle(GO:0000082) |
| 0.0 | 0.6 | GO:1902236 | negative regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signaling pathway(GO:1902236) |
| 0.0 | 13.8 | GO:0051301 | cell division(GO:0051301) |
| 0.0 | 0.2 | GO:0070166 | enamel mineralization(GO:0070166) |
| 0.0 | 0.5 | GO:0045116 | protein neddylation(GO:0045116) |
| 0.0 | 0.5 | GO:0050901 | leukocyte tethering or rolling(GO:0050901) |
| 0.0 | 1.0 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.0 | 0.6 | GO:0010867 | positive regulation of triglyceride biosynthetic process(GO:0010867) |
| 0.0 | 0.1 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 1.5 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.0 | 0.5 | GO:0002643 | regulation of tolerance induction(GO:0002643) |
| 0.0 | 1.9 | GO:0006413 | translational initiation(GO:0006413) |
| 0.0 | 0.3 | GO:0016226 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.0 | 4.5 | GO:0007018 | microtubule-based movement(GO:0007018) |
| 0.0 | 0.2 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.0 | 0.1 | GO:0038032 | termination of G-protein coupled receptor signaling pathway(GO:0038032) |
| 0.0 | 0.2 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.0 | 2.0 | GO:0006364 | rRNA processing(GO:0006364) |
| 0.0 | 0.6 | GO:0001541 | ovarian follicle development(GO:0001541) |
| 0.0 | 1.9 | GO:2001252 | positive regulation of chromosome organization(GO:2001252) |
| 0.0 | 1.8 | GO:0006626 | protein targeting to mitochondrion(GO:0006626) |
| 0.0 | 1.0 | GO:0042254 | ribosome biogenesis(GO:0042254) |
| 0.0 | 0.7 | GO:0008286 | insulin receptor signaling pathway(GO:0008286) |
| 0.0 | 0.1 | GO:0043982 | histone H4-K5 acetylation(GO:0043981) histone H4-K8 acetylation(GO:0043982) |
| 0.0 | 0.0 | GO:0006624 | vacuolar protein processing(GO:0006624) |
| 0.0 | 0.3 | GO:0008156 | negative regulation of DNA replication(GO:0008156) |
| 0.0 | 0.2 | GO:0046686 | response to cadmium ion(GO:0046686) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.7 | 8.0 | GO:1990423 | RZZ complex(GO:1990423) |
| 2.1 | 4.3 | GO:0042585 | germinal vesicle(GO:0042585) |
| 2.0 | 8.0 | GO:0000942 | condensed nuclear chromosome outer kinetochore(GO:0000942) |
| 1.8 | 5.4 | GO:0000799 | nuclear condensin complex(GO:0000799) |
| 1.2 | 13.0 | GO:0005642 | annulate lamellae(GO:0005642) |
| 1.1 | 7.4 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.9 | 2.7 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.8 | 5.0 | GO:0098536 | deuterosome(GO:0098536) |
| 0.7 | 26.7 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.7 | 12.5 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.5 | 4.2 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.5 | 1.5 | GO:1990047 | spindle matrix(GO:1990047) |
| 0.5 | 4.6 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.4 | 2.2 | GO:0097226 | sperm mitochondrial sheath(GO:0097226) |
| 0.4 | 3.1 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.4 | 10.2 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.4 | 1.6 | GO:0000235 | astral microtubule(GO:0000235) |
| 0.4 | 1.8 | GO:0034715 | pICln-Sm protein complex(GO:0034715) |
| 0.3 | 2.1 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.3 | 2.2 | GO:0070695 | FHF complex(GO:0070695) |
| 0.3 | 2.0 | GO:0071204 | histone pre-mRNA 3'end processing complex(GO:0071204) |
| 0.3 | 2.0 | GO:0097255 | R2TP complex(GO:0097255) |
| 0.2 | 0.7 | GO:0042272 | nucleocytoplasmic shuttling complex(GO:0031074) nuclear RNA export factor complex(GO:0042272) |
| 0.2 | 1.6 | GO:0001740 | Barr body(GO:0001740) |
| 0.2 | 5.9 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.2 | 12.7 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.2 | 1.5 | GO:0001940 | male pronucleus(GO:0001940) |
| 0.2 | 1.5 | GO:0005638 | lamin filament(GO:0005638) |
| 0.2 | 0.6 | GO:0071595 | Nem1-Spo7 phosphatase complex(GO:0071595) |
| 0.2 | 3.1 | GO:0097539 | ciliary transition fiber(GO:0097539) |
| 0.2 | 4.0 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 0.2 | 1.2 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.2 | 2.4 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.2 | 1.6 | GO:0000243 | commitment complex(GO:0000243) |
| 0.2 | 1.9 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.2 | 2.2 | GO:0045120 | pronucleus(GO:0045120) |
| 0.1 | 0.4 | GO:0000814 | ESCRT II complex(GO:0000814) |
| 0.1 | 5.1 | GO:0005763 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.1 | 2.3 | GO:0002199 | zona pellucida receptor complex(GO:0002199) |
| 0.1 | 12.6 | GO:0000922 | spindle pole(GO:0000922) |
| 0.1 | 0.9 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.1 | 1.1 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.1 | 0.8 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.1 | 0.3 | GO:0097057 | TRAF2-GSTP1 complex(GO:0097057) |
| 0.1 | 2.6 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
| 0.1 | 0.5 | GO:1990429 | Pex17p-Pex14p docking complex(GO:1990415) peroxisomal importomer complex(GO:1990429) |
| 0.1 | 1.1 | GO:0045298 | tubulin complex(GO:0045298) |
| 0.1 | 1.1 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.1 | 1.2 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.1 | 0.9 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.1 | 11.9 | GO:0000776 | kinetochore(GO:0000776) |
| 0.1 | 0.3 | GO:0031510 | SUMO activating enzyme complex(GO:0031510) |
| 0.1 | 0.4 | GO:0099522 | region of cytosol(GO:0099522) postsynaptic cytosol(GO:0099524) |
| 0.1 | 0.4 | GO:0005927 | muscle tendon junction(GO:0005927) |
| 0.1 | 0.3 | GO:1990590 | ATF1-ATF4 transcription factor complex(GO:1990590) |
| 0.1 | 4.8 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.1 | 5.9 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.1 | 9.9 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.1 | 0.5 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.1 | 0.9 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.1 | 0.4 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.1 | 2.4 | GO:0030673 | axolemma(GO:0030673) |
| 0.1 | 1.3 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.1 | 0.5 | GO:0097413 | Lewy body(GO:0097413) |
| 0.1 | 0.3 | GO:0071817 | MMXD complex(GO:0071817) CIA complex(GO:0097361) |
| 0.1 | 0.5 | GO:0071546 | pi-body(GO:0071546) |
| 0.1 | 0.3 | GO:1990745 | EARP complex(GO:1990745) |
| 0.1 | 0.3 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.1 | 1.1 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.0 | 1.8 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.0 | 0.1 | GO:0035061 | interchromatin granule(GO:0035061) |
| 0.0 | 0.8 | GO:0043189 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 2.3 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.0 | 1.9 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 1.8 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 0.4 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.0 | 1.5 | GO:0031672 | A band(GO:0031672) |
| 0.0 | 4.0 | GO:0022626 | cytosolic ribosome(GO:0022626) |
| 0.0 | 1.8 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.6 | GO:0005839 | proteasome core complex(GO:0005839) |
| 0.0 | 5.1 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 2.5 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 1.6 | GO:0000502 | proteasome complex(GO:0000502) |
| 0.0 | 0.7 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 0.5 | GO:0005921 | gap junction(GO:0005921) |
| 0.0 | 0.6 | GO:0043034 | costamere(GO:0043034) |
| 0.0 | 0.4 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.0 | 0.7 | GO:0005762 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 4.9 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.0 | 0.6 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.3 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 8.3 | GO:0005730 | nucleolus(GO:0005730) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 11.1 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 1.1 | 4.4 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 1.0 | 4.0 | GO:0097100 | supercoiled DNA binding(GO:0097100) |
| 0.9 | 3.6 | GO:0050220 | prostaglandin-E synthase activity(GO:0050220) |
| 0.7 | 11.8 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.7 | 2.0 | GO:0071207 | histone pre-mRNA stem-loop binding(GO:0071207) |
| 0.6 | 2.4 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.5 | 4.3 | GO:0035174 | histone serine kinase activity(GO:0035174) |
| 0.4 | 11.8 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.4 | 2.1 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.4 | 2.0 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.4 | 1.5 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.4 | 2.6 | GO:0033592 | RNA strand annealing activity(GO:0033592) |
| 0.4 | 1.5 | GO:0047275 | glucosaminylgalactosylglucosylceramide beta-galactosyltransferase activity(GO:0047275) |
| 0.4 | 3.3 | GO:1990446 | U1 snRNP binding(GO:1990446) |
| 0.3 | 2.4 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.3 | 2.0 | GO:0043141 | ATP-dependent 5'-3' DNA helicase activity(GO:0043141) |
| 0.3 | 2.2 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.3 | 1.2 | GO:0035851 | Krueppel-associated box domain binding(GO:0035851) |
| 0.3 | 1.1 | GO:0031800 | type 3 metabotropic glutamate receptor binding(GO:0031800) |
| 0.3 | 2.1 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.2 | 1.0 | GO:0019863 | IgE binding(GO:0019863) |
| 0.2 | 2.1 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.2 | 1.3 | GO:0051425 | PTB domain binding(GO:0051425) |
| 0.2 | 0.6 | GO:0019153 | protein-disulfide reductase (glutathione) activity(GO:0019153) |
| 0.2 | 0.9 | GO:0004515 | nicotinamide-nucleotide adenylyltransferase activity(GO:0000309) nicotinate-nucleotide adenylyltransferase activity(GO:0004515) |
| 0.2 | 2.2 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.2 | 4.9 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.2 | 1.8 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.2 | 3.0 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.2 | 12.7 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.2 | 0.5 | GO:0005128 | erythropoietin receptor binding(GO:0005128) |
| 0.1 | 2.1 | GO:0008242 | omega peptidase activity(GO:0008242) |
| 0.1 | 1.6 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.1 | 1.4 | GO:0004791 | thioredoxin-disulfide reductase activity(GO:0004791) |
| 0.1 | 0.8 | GO:0097016 | L27 domain binding(GO:0097016) |
| 0.1 | 1.8 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.1 | 0.6 | GO:0042610 | CD8 receptor binding(GO:0042610) |
| 0.1 | 1.3 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.1 | 5.0 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.1 | 1.5 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.1 | 0.3 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.1 | 0.3 | GO:0004949 | cannabinoid receptor activity(GO:0004949) |
| 0.1 | 0.7 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.1 | 1.3 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.1 | 4.5 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.1 | 1.8 | GO:0008432 | JUN kinase binding(GO:0008432) |
| 0.1 | 0.4 | GO:0033192 | calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.1 | 1.6 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.1 | 0.5 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.1 | 9.7 | GO:0004386 | helicase activity(GO:0004386) |
| 0.1 | 0.2 | GO:0008940 | nitrate reductase activity(GO:0008940) |
| 0.1 | 0.4 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.1 | 0.2 | GO:0003998 | acylphosphatase activity(GO:0003998) |
| 0.1 | 2.0 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.1 | 11.7 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 2.3 | GO:0042054 | histone methyltransferase activity(GO:0042054) |
| 0.0 | 1.3 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 1.2 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 1.5 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 2.5 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 0.4 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.0 | 0.2 | GO:0032564 | dATP binding(GO:0032564) |
| 0.0 | 2.0 | GO:0043531 | ADP binding(GO:0043531) |
| 0.0 | 1.4 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.3 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.0 | 1.5 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 3.1 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 1.3 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.7 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 5.8 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.2 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.0 | 1.5 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 16.4 | GO:0004674 | protein serine/threonine kinase activity(GO:0004674) |
| 0.0 | 2.6 | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups(GO:0016765) |
| 0.0 | 0.4 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.0 | 0.6 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.0 | 2.7 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 0.1 | GO:0005344 | catalase activity(GO:0004096) oxygen transporter activity(GO:0005344) |
| 0.0 | 0.3 | GO:0044388 | small protein activating enzyme binding(GO:0044388) |
| 0.0 | 0.3 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.0 | 1.9 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 1.2 | GO:0008173 | RNA methyltransferase activity(GO:0008173) |
| 0.0 | 0.7 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 7.7 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 0.4 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.0 | 0.8 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.0 | 0.6 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 2.3 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.0 | GO:0030235 | nitric-oxide synthase regulator activity(GO:0030235) |
| 0.0 | 1.0 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 0.2 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.7 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.0 | 0.3 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.0 | 1.1 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 37.9 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.2 | 8.3 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.2 | 6.4 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.1 | 6.1 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.1 | 2.4 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.1 | 4.8 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 2.0 | PID MYC PATHWAY | C-MYC pathway |
| 0.1 | 1.0 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.1 | 5.3 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.1 | 0.7 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.1 | 1.2 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.1 | 1.8 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.1 | 3.7 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 3.9 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.0 | 2.7 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 0.6 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 0.2 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.0 | 0.7 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.7 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 1.2 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.0 | 0.5 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 0.6 | ST GA13 PATHWAY | G alpha 13 Pathway |
| 0.0 | 0.3 | ST P38 MAPK PATHWAY | p38 MAPK Pathway |
| 0.0 | 0.8 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.0 | 0.6 | ST T CELL SIGNAL TRANSDUCTION | T Cell Signal Transduction |
| 0.0 | 0.6 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 19.7 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.6 | 14.7 | REACTOME APC CDC20 MEDIATED DEGRADATION OF NEK2A | Genes involved in APC-Cdc20 mediated degradation of Nek2A |
| 0.5 | 4.0 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.4 | 42.0 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.4 | 11.8 | REACTOME PTM GAMMA CARBOXYLATION HYPUSINE FORMATION AND ARYLSULFATASE ACTIVATION | Genes involved in PTM: gamma carboxylation, hypusine formation and arylsulfatase activation |
| 0.2 | 8.2 | REACTOME ACTIVATION OF THE PRE REPLICATIVE COMPLEX | Genes involved in Activation of the pre-replicative complex |
| 0.2 | 3.6 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.2 | 2.0 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.2 | 3.4 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.1 | 2.4 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.1 | 2.0 | REACTOME EXTENSION OF TELOMERES | Genes involved in Extension of Telomeres |
| 0.1 | 4.9 | REACTOME APC C CDH1 MEDIATED DEGRADATION OF CDC20 AND OTHER APC C CDH1 TARGETED PROTEINS IN LATE MITOSIS EARLY G1 | Genes involved in APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 |
| 0.1 | 1.1 | REACTOME ER PHAGOSOME PATHWAY | Genes involved in ER-Phagosome pathway |
| 0.1 | 2.2 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.1 | 3.0 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.1 | 1.5 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 0.1 | 0.6 | REACTOME ACTIVATION OF ATR IN RESPONSE TO REPLICATION STRESS | Genes involved in Activation of ATR in response to replication stress |
| 0.1 | 2.4 | REACTOME METABOLISM OF STEROID HORMONES AND VITAMINS A AND D | Genes involved in Metabolism of steroid hormones and vitamins A and D |
| 0.1 | 2.0 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.1 | 3.9 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.1 | 0.4 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.1 | 1.9 | REACTOME G BETA GAMMA SIGNALLING THROUGH PLC BETA | Genes involved in G beta:gamma signalling through PLC beta |
| 0.0 | 1.5 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.0 | 1.6 | REACTOME TELOMERE MAINTENANCE | Genes involved in Telomere Maintenance |
| 0.0 | 1.2 | REACTOME CD28 DEPENDENT PI3K AKT SIGNALING | Genes involved in CD28 dependent PI3K/Akt signaling |
| 0.0 | 1.5 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.5 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.7 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.0 | 1.3 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.0 | 1.1 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.6 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.0 | 2.7 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.0 | 1.6 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 0.4 | REACTOME PREFOLDIN MEDIATED TRANSFER OF SUBSTRATE TO CCT TRIC | Genes involved in Prefoldin mediated transfer of substrate to CCT/TriC |
| 0.0 | 0.5 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.0 | 2.4 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 0.3 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 2.3 | REACTOME DNA REPAIR | Genes involved in DNA Repair |
| 0.0 | 0.6 | REACTOME MICRORNA MIRNA BIOGENESIS | Genes involved in MicroRNA (miRNA) Biogenesis |
| 0.0 | 1.1 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 0.5 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.0 | 0.6 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.0 | 2.1 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 0.7 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.0 | 0.4 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.3 | REACTOME MAPK TARGETS NUCLEAR EVENTS MEDIATED BY MAP KINASES | Genes involved in MAPK targets/ Nuclear events mediated by MAP kinases |
| 0.0 | 0.4 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.0 | 0.4 | REACTOME S PHASE | Genes involved in S Phase |
| 0.0 | 0.8 | REACTOME CHONDROITIN SULFATE DERMATAN SULFATE METABOLISM | Genes involved in Chondroitin sulfate/dermatan sulfate metabolism |
| 0.0 | 0.4 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 1.1 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 1.3 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.7 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |