Project
avrg: 2D miR_HR1_12
Navigation
Downloads
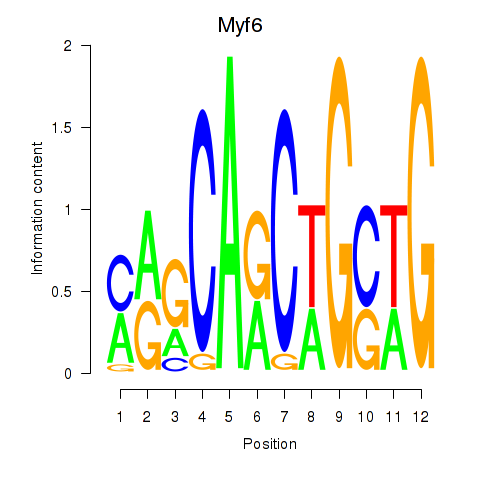
Results for Myf6
Z-value: 4.04
Transcription factors associated with Myf6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Myf6
|
ENSMUSG00000035923.3 | myogenic factor 6 |
Activity profile of Myf6 motif
Sorted Z-values of Myf6 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_119238794 | 4.62 |
ENSMUST00000026360.8
|
Itgb8
|
integrin beta 8 |
| chr5_+_30588078 | 4.61 |
ENSMUST00000066295.2
|
Kcnk3
|
potassium channel, subfamily K, member 3 |
| chr9_+_13621646 | 4.46 |
ENSMUST00000034401.8
|
Maml2
|
mastermind like 2 (Drosophila) |
| chr17_-_87282771 | 4.45 |
ENSMUST00000161759.1
|
4833418N02Rik
|
RIKEN cDNA 4833418N02 gene |
| chr6_+_107529717 | 4.21 |
ENSMUST00000049285.8
|
Lrrn1
|
leucine rich repeat protein 1, neuronal |
| chrX_-_108664891 | 4.14 |
ENSMUST00000178160.1
|
Gm379
|
predicted gene 379 |
| chr2_+_130295148 | 3.51 |
ENSMUST00000110288.2
|
Ebf4
|
early B cell factor 4 |
| chr6_-_112489808 | 3.50 |
ENSMUST00000053306.6
|
Oxtr
|
oxytocin receptor |
| chr17_-_87282793 | 3.48 |
ENSMUST00000146560.2
|
4833418N02Rik
|
RIKEN cDNA 4833418N02 gene |
| chr14_+_59625281 | 3.41 |
ENSMUST00000053949.5
|
Shisa2
|
shisa homolog 2 (Xenopus laevis) |
| chr16_+_17276662 | 3.37 |
ENSMUST00000069420.4
|
Tmem191c
|
transmembrane protein 191C |
| chr4_-_106799779 | 3.36 |
ENSMUST00000145061.1
ENSMUST00000102762.3 |
Acot11
|
acyl-CoA thioesterase 11 |
| chr10_+_127866457 | 3.08 |
ENSMUST00000092058.3
|
BC089597
|
cDNA sequence BC089597 |
| chr16_+_17276337 | 3.05 |
ENSMUST00000159065.1
ENSMUST00000159494.1 ENSMUST00000159811.1 |
Tmem191c
|
transmembrane protein 191C |
| chr11_+_69965396 | 2.94 |
ENSMUST00000018713.6
|
Cldn7
|
claudin 7 |
| chr4_-_138367966 | 2.94 |
ENSMUST00000030535.3
|
Cda
|
cytidine deaminase |
| chr17_+_24488773 | 2.62 |
ENSMUST00000024958.7
|
Caskin1
|
CASK interacting protein 1 |
| chr8_+_94152607 | 2.61 |
ENSMUST00000034211.8
|
Mt3
|
metallothionein 3 |
| chr12_+_108334341 | 2.55 |
ENSMUST00000021684.4
|
Cyp46a1
|
cytochrome P450, family 46, subfamily a, polypeptide 1 |
| chr8_+_62951195 | 2.53 |
ENSMUST00000118003.1
|
Spock3
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan 3 |
| chr9_+_45370185 | 2.49 |
ENSMUST00000085939.6
|
Fxyd6
|
FXYD domain-containing ion transport regulator 6 |
| chr18_-_82406777 | 2.38 |
ENSMUST00000065224.6
|
Galr1
|
galanin receptor 1 |
| chr18_-_36197343 | 2.37 |
ENSMUST00000115713.1
ENSMUST00000115712.1 |
Nrg2
|
neuregulin 2 |
| chr8_-_24948771 | 2.34 |
ENSMUST00000119720.1
ENSMUST00000121438.2 |
Adam32
|
a disintegrin and metallopeptidase domain 32 |
| chr8_-_70487314 | 2.32 |
ENSMUST00000045286.7
|
Tmem59l
|
transmembrane protein 59-like |
| chr5_+_140607334 | 2.28 |
ENSMUST00000031555.1
|
Lfng
|
LFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr17_-_46752170 | 2.27 |
ENSMUST00000121671.1
ENSMUST00000059844.6 |
Cnpy3
|
canopy 3 homolog (zebrafish) |
| chr2_-_24048857 | 2.27 |
ENSMUST00000114497.1
|
Hnmt
|
histamine N-methyltransferase |
| chr18_-_21652362 | 2.19 |
ENSMUST00000049105.4
|
Klhl14
|
kelch-like 14 |
| chr5_+_90786100 | 2.19 |
ENSMUST00000031326.8
|
Cxcl3
|
chemokine (C-X-C motif) ligand 3 |
| chr11_-_119086221 | 2.17 |
ENSMUST00000026665.7
|
Cbx4
|
chromobox 4 |
| chr10_+_17796256 | 2.13 |
ENSMUST00000037964.6
|
Txlnb
|
taxilin beta |
| chr10_+_63386550 | 2.05 |
ENSMUST00000043317.5
|
Dnajc12
|
DnaJ (Hsp40) homolog, subfamily C, member 12 |
| chr11_-_106487833 | 2.01 |
ENSMUST00000106801.1
|
Ern1
|
endoplasmic reticulum (ER) to nucleus signalling 1 |
| chr14_+_33923582 | 1.97 |
ENSMUST00000168727.1
|
Gdf10
|
growth differentiation factor 10 |
| chr6_+_125145235 | 1.94 |
ENSMUST00000119527.1
ENSMUST00000088276.6 ENSMUST00000051171.7 ENSMUST00000117675.1 |
Iffo1
|
intermediate filament family orphan 1 |
| chr4_+_58943575 | 1.93 |
ENSMUST00000107554.1
|
Zkscan16
|
zinc finger with KRAB and SCAN domains 16 |
| chr5_-_36398090 | 1.91 |
ENSMUST00000037370.7
ENSMUST00000070720.6 |
Sorcs2
|
sortilin-related VPS10 domain containing receptor 2 |
| chr1_-_121327776 | 1.90 |
ENSMUST00000160688.1
|
Insig2
|
insulin induced gene 2 |
| chr4_-_148149684 | 1.87 |
ENSMUST00000126615.1
|
Fbxo6
|
F-box protein 6 |
| chr11_+_78322965 | 1.87 |
ENSMUST00000017534.8
|
Aldoc
|
aldolase C, fructose-bisphosphate |
| chr14_+_55853997 | 1.87 |
ENSMUST00000100529.3
|
Nynrin
|
NYN domain and retroviral integrase containing |
| chr7_-_143649614 | 1.84 |
ENSMUST00000129476.1
ENSMUST00000084396.3 ENSMUST00000075588.6 ENSMUST00000146692.1 |
Tnfrsf22
|
tumor necrosis factor receptor superfamily, member 22 |
| chr6_+_45060036 | 1.83 |
ENSMUST00000114641.1
|
Cntnap2
|
contactin associated protein-like 2 |
| chrX_-_75084757 | 1.79 |
ENSMUST00000114104.1
ENSMUST00000114109.1 ENSMUST00000037374.4 |
Gab3
|
growth factor receptor bound protein 2-associated protein 3 |
| chrX_+_20703906 | 1.77 |
ENSMUST00000033383.2
|
Usp11
|
ubiquitin specific peptidase 11 |
| chr5_+_102845007 | 1.75 |
ENSMUST00000070000.4
|
Arhgap24
|
Rho GTPase activating protein 24 |
| chr1_-_121327734 | 1.74 |
ENSMUST00000160968.1
ENSMUST00000162582.1 |
Insig2
|
insulin induced gene 2 |
| chr11_-_106487796 | 1.71 |
ENSMUST00000001059.2
ENSMUST00000106799.1 ENSMUST00000106800.1 |
Ern1
|
endoplasmic reticulum (ER) to nucleus signalling 1 |
| chr14_+_55854115 | 1.70 |
ENSMUST00000168479.1
|
Nynrin
|
NYN domain and retroviral integrase containing |
| chr11_+_113619318 | 1.69 |
ENSMUST00000146390.2
ENSMUST00000106630.1 |
Sstr2
|
somatostatin receptor 2 |
| chr16_+_17276291 | 1.66 |
ENSMUST00000164950.1
ENSMUST00000159242.1 |
Tmem191c
|
transmembrane protein 191C |
| chr17_+_26933070 | 1.66 |
ENSMUST00000073724.5
|
Phf1
|
PHD finger protein 1 |
| chr14_+_65968483 | 1.65 |
ENSMUST00000022616.6
|
Clu
|
clusterin |
| chrX_-_7572843 | 1.65 |
ENSMUST00000132788.1
|
Ppp1r3f
|
protein phosphatase 1, regulatory (inhibitor) subunit 3F |
| chr1_-_121328024 | 1.64 |
ENSMUST00000003818.7
|
Insig2
|
insulin induced gene 2 |
| chr18_+_60803838 | 1.60 |
ENSMUST00000050487.8
ENSMUST00000097563.2 ENSMUST00000167610.1 |
Cd74
|
CD74 antigen (invariant polypeptide of major histocompatibility complex, class II antigen-associated) |
| chr11_+_87760533 | 1.60 |
ENSMUST00000039627.5
ENSMUST00000100644.3 |
Bzrap1
|
benzodiazepine receptor associated protein 1 |
| chr4_+_139380658 | 1.59 |
ENSMUST00000165860.1
ENSMUST00000097822.3 |
Ubr4
|
ubiquitin protein ligase E3 component n-recognin 4 |
| chr16_+_93683184 | 1.58 |
ENSMUST00000039620.6
|
Cbr3
|
carbonyl reductase 3 |
| chr11_-_3931789 | 1.58 |
ENSMUST00000109992.1
ENSMUST00000109988.1 |
Tcn2
|
transcobalamin 2 |
| chr2_+_19445632 | 1.57 |
ENSMUST00000028068.2
|
Ptf1a
|
pancreas specific transcription factor, 1a |
| chr6_-_126740151 | 1.54 |
ENSMUST00000112242.1
|
Kcna6
|
potassium voltage-gated channel, shaker-related, subfamily, member 6 |
| chr7_-_29168647 | 1.53 |
ENSMUST00000048923.6
|
Spred3
|
sprouty-related, EVH1 domain containing 3 |
| chr2_+_54436317 | 1.53 |
ENSMUST00000112636.1
ENSMUST00000112635.1 ENSMUST00000112634.1 |
Galnt13
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 13 |
| chr1_-_121327672 | 1.51 |
ENSMUST00000159085.1
ENSMUST00000159125.1 ENSMUST00000161818.1 |
Insig2
|
insulin induced gene 2 |
| chr4_+_102254993 | 1.49 |
ENSMUST00000106908.2
|
Pde4b
|
phosphodiesterase 4B, cAMP specific |
| chr6_+_125215551 | 1.47 |
ENSMUST00000032487.7
ENSMUST00000100942.2 ENSMUST00000063588.8 |
Vamp1
|
vesicle-associated membrane protein 1 |
| chr3_+_108284089 | 1.47 |
ENSMUST00000102632.4
|
Sort1
|
sortilin 1 |
| chrX_-_10117597 | 1.47 |
ENSMUST00000115543.2
ENSMUST00000044789.3 ENSMUST00000115544.2 |
Srpx
|
sushi-repeat-containing protein |
| chr7_-_127993831 | 1.46 |
ENSMUST00000033056.3
|
Pycard
|
PYD and CARD domain containing |
| chr6_+_56017489 | 1.45 |
ENSMUST00000052827.4
|
Ppp1r17
|
protein phosphatase 1, regulatory subunit 17 |
| chr2_-_181039286 | 1.45 |
ENSMUST00000067120.7
|
Chrna4
|
cholinergic receptor, nicotinic, alpha polypeptide 4 |
| chr17_-_63499983 | 1.44 |
ENSMUST00000024761.6
|
Fbxl17
|
F-box and leucine-rich repeat protein 17 |
| chr9_+_118506226 | 1.43 |
ENSMUST00000084820.4
|
Golga4
|
golgi autoantigen, golgin subfamily a, 4 |
| chr4_-_106800249 | 1.43 |
ENSMUST00000148688.1
|
Acot11
|
acyl-CoA thioesterase 11 |
| chr12_-_101819048 | 1.43 |
ENSMUST00000021603.8
|
Fbln5
|
fibulin 5 |
| chr4_-_109476666 | 1.43 |
ENSMUST00000030284.3
|
Rnf11
|
ring finger protein 11 |
| chr15_-_86033777 | 1.42 |
ENSMUST00000016172.7
|
Celsr1
|
cadherin, EGF LAG seven-pass G-type receptor 1 (flamingo homolog, Drosophila) |
| chr11_+_93099284 | 1.41 |
ENSMUST00000092780.3
ENSMUST00000107863.2 |
Car10
|
carbonic anhydrase 10 |
| chr9_-_39604124 | 1.41 |
ENSMUST00000042485.4
ENSMUST00000141370.1 |
AW551984
|
expressed sequence AW551984 |
| chr1_-_84696182 | 1.40 |
ENSMUST00000049126.6
|
Dner
|
delta/notch-like EGF-related receptor |
| chr2_-_164443177 | 1.39 |
ENSMUST00000017153.3
|
Sdc4
|
syndecan 4 |
| chr8_-_111259192 | 1.39 |
ENSMUST00000169020.1
ENSMUST00000003404.8 |
Glg1
|
golgi apparatus protein 1 |
| chr10_+_57784859 | 1.39 |
ENSMUST00000020024.5
|
Fabp7
|
fatty acid binding protein 7, brain |
| chr15_-_88819279 | 1.39 |
ENSMUST00000043087.8
ENSMUST00000162183.1 |
Alg12
|
asparagine-linked glycosylation 12 (alpha-1,6-mannosyltransferase) |
| chr6_-_5496296 | 1.38 |
ENSMUST00000019721.4
|
Pdk4
|
pyruvate dehydrogenase kinase, isoenzyme 4 |
| chr17_+_47436615 | 1.38 |
ENSMUST00000037701.6
|
AI661453
|
expressed sequence AI661453 |
| chr5_-_137611372 | 1.33 |
ENSMUST00000054564.6
|
Pcolce
|
procollagen C-endopeptidase enhancer protein |
| chr10_-_128525859 | 1.33 |
ENSMUST00000026427.6
|
Esyt1
|
extended synaptotagmin-like protein 1 |
| chr16_-_74411292 | 1.32 |
ENSMUST00000117200.1
|
Robo2
|
roundabout homolog 2 (Drosophila) |
| chrX_-_143827391 | 1.32 |
ENSMUST00000087316.5
|
Capn6
|
calpain 6 |
| chr6_+_118066356 | 1.31 |
ENSMUST00000164960.1
|
Rasgef1a
|
RasGEF domain family, member 1A |
| chr6_-_52204415 | 1.30 |
ENSMUST00000048794.6
|
Hoxa5
|
homeobox A5 |
| chr16_-_34095983 | 1.29 |
ENSMUST00000114973.1
ENSMUST00000114964.1 |
Kalrn
|
kalirin, RhoGEF kinase |
| chr17_+_79051906 | 1.28 |
ENSMUST00000040789.4
|
Qpct
|
glutaminyl-peptide cyclotransferase (glutaminyl cyclase) |
| chr9_+_107399858 | 1.28 |
ENSMUST00000085092.5
ENSMUST00000164988.2 |
Cacna2d2
|
calcium channel, voltage-dependent, alpha 2/delta subunit 2 |
| chr11_+_104231390 | 1.27 |
ENSMUST00000106992.3
|
Mapt
|
microtubule-associated protein tau |
| chr2_+_125136692 | 1.26 |
ENSMUST00000099452.2
|
Ctxn2
|
cortexin 2 |
| chr10_+_11343387 | 1.26 |
ENSMUST00000069106.4
|
Epm2a
|
epilepsy, progressive myoclonic epilepsy, type 2 gene alpha |
| chr7_+_4119556 | 1.26 |
ENSMUST00000079415.5
|
Ttyh1
|
tweety homolog 1 (Drosophila) |
| chr17_+_47436731 | 1.25 |
ENSMUST00000150819.2
|
AI661453
|
expressed sequence AI661453 |
| chr5_-_137600650 | 1.23 |
ENSMUST00000111007.1
ENSMUST00000133705.1 |
Mospd3
|
motile sperm domain containing 3 |
| chr7_+_4119525 | 1.22 |
ENSMUST00000119661.1
ENSMUST00000129423.1 |
Ttyh1
|
tweety homolog 1 (Drosophila) |
| chrX_+_159627265 | 1.22 |
ENSMUST00000112456.2
|
Sh3kbp1
|
SH3-domain kinase binding protein 1 |
| chr7_+_19083842 | 1.21 |
ENSMUST00000032568.7
ENSMUST00000122999.1 ENSMUST00000108473.3 ENSMUST00000108474.1 |
Dmpk
|
dystrophia myotonica-protein kinase |
| chr17_+_35342242 | 1.20 |
ENSMUST00000074806.5
|
H2-Q2
|
histocompatibility 2, Q region locus 2 |
| chr16_+_17797282 | 1.20 |
ENSMUST00000012161.3
|
Scarf2
|
scavenger receptor class F, member 2 |
| chr2_-_181039160 | 1.20 |
ENSMUST00000108851.1
|
Chrna4
|
cholinergic receptor, nicotinic, alpha polypeptide 4 |
| chr17_-_35979679 | 1.19 |
ENSMUST00000173724.1
ENSMUST00000172900.1 ENSMUST00000174849.1 |
Prr3
|
proline-rich polypeptide 3 |
| chr4_+_105157339 | 1.19 |
ENSMUST00000064139.7
|
Ppap2b
|
phosphatidic acid phosphatase type 2B |
| chr17_+_34931253 | 1.18 |
ENSMUST00000007253.5
|
Neu1
|
neuraminidase 1 |
| chr3_-_90389884 | 1.18 |
ENSMUST00000029541.5
|
Slc27a3
|
solute carrier family 27 (fatty acid transporter), member 3 |
| chr9_-_110408160 | 1.18 |
ENSMUST00000040021.7
|
Ptpn23
|
protein tyrosine phosphatase, non-receptor type 23 |
| chr8_+_125669818 | 1.17 |
ENSMUST00000053078.3
|
Map10
|
microtubule-associated protein 10 |
| chr1_+_167001417 | 1.17 |
ENSMUST00000165874.1
|
Fam78b
|
family with sequence similarity 78, member B |
| chr12_+_26469204 | 1.17 |
ENSMUST00000020969.3
|
Cmpk2
|
cytidine monophosphate (UMP-CMP) kinase 2, mitochondrial |
| chr5_-_137611429 | 1.16 |
ENSMUST00000031731.7
|
Pcolce
|
procollagen C-endopeptidase enhancer protein |
| chr18_+_65873478 | 1.16 |
ENSMUST00000025395.8
ENSMUST00000173530.1 |
Grp
|
gastrin releasing peptide |
| chr10_+_96616998 | 1.16 |
ENSMUST00000038377.7
|
Btg1
|
B cell translocation gene 1, anti-proliferative |
| chr17_+_29490812 | 1.15 |
ENSMUST00000024811.6
|
Pim1
|
proviral integration site 1 |
| chrX_+_73787062 | 1.15 |
ENSMUST00000002090.2
|
Ssr4
|
signal sequence receptor, delta |
| chr8_-_84011442 | 1.15 |
ENSMUST00000056686.5
|
2210011C24Rik
|
RIKEN cDNA 2210011C24 gene |
| chr5_+_117413977 | 1.14 |
ENSMUST00000180430.1
|
Ksr2
|
kinase suppressor of ras 2 |
| chr6_-_83121385 | 1.13 |
ENSMUST00000146328.1
ENSMUST00000113936.3 ENSMUST00000032111.4 |
Wbp1
|
WW domain binding protein 1 |
| chr16_-_97170707 | 1.13 |
ENSMUST00000056102.7
|
Dscam
|
Down syndrome cell adhesion molecule |
| chr3_+_65109343 | 1.12 |
ENSMUST00000159525.1
ENSMUST00000049230.8 |
Kcnab1
|
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr5_-_92505518 | 1.12 |
ENSMUST00000031377.7
|
Scarb2
|
scavenger receptor class B, member 2 |
| chr10_-_75560330 | 1.12 |
ENSMUST00000051129.9
|
Fam211b
|
family with sequence similarity 211, member B |
| chr9_-_21592805 | 1.11 |
ENSMUST00000034700.7
ENSMUST00000180365.1 ENSMUST00000078572.7 |
Yipf2
|
Yip1 domain family, member 2 |
| chr7_+_51879041 | 1.11 |
ENSMUST00000107591.2
|
Gas2
|
growth arrest specific 2 |
| chr5_-_115158169 | 1.10 |
ENSMUST00000053271.5
ENSMUST00000112121.1 |
Mlec
|
malectin |
| chr1_+_95313607 | 1.10 |
ENSMUST00000059975.6
|
Fam174a
|
family with sequence similarity 174, member A |
| chr2_+_21367532 | 1.10 |
ENSMUST00000055946.7
|
Gpr158
|
G protein-coupled receptor 158 |
| chr11_+_29692937 | 1.10 |
ENSMUST00000102843.3
ENSMUST00000102842.3 ENSMUST00000078830.4 ENSMUST00000170731.1 |
Rtn4
|
reticulon 4 |
| chr11_+_104231573 | 1.09 |
ENSMUST00000132977.1
ENSMUST00000132245.1 ENSMUST00000100347.4 |
Mapt
|
microtubule-associated protein tau |
| chr7_-_27337667 | 1.09 |
ENSMUST00000038618.6
ENSMUST00000108369.2 |
Ltbp4
|
latent transforming growth factor beta binding protein 4 |
| chr11_+_104231515 | 1.09 |
ENSMUST00000106993.3
|
Mapt
|
microtubule-associated protein tau |
| chr10_-_20548320 | 1.09 |
ENSMUST00000169404.1
|
Pde7b
|
phosphodiesterase 7B |
| chr14_-_123627059 | 1.09 |
ENSMUST00000000201.5
|
Nalcn
|
sodium leak channel, non-selective |
| chrX_-_112698642 | 1.08 |
ENSMUST00000039887.3
|
Pof1b
|
premature ovarian failure 1B |
| chr9_-_114844090 | 1.08 |
ENSMUST00000047013.3
|
Cmtm8
|
CKLF-like MARVEL transmembrane domain containing 8 |
| chr7_-_45136056 | 1.08 |
ENSMUST00000130628.1
|
Flt3l
|
FMS-like tyrosine kinase 3 ligand |
| chrX_+_159627534 | 1.08 |
ENSMUST00000073094.3
|
Sh3kbp1
|
SH3-domain kinase binding protein 1 |
| chrX_-_73880831 | 1.07 |
ENSMUST00000102871.3
|
L1cam
|
L1 cell adhesion molecule |
| chr7_-_67759735 | 1.07 |
ENSMUST00000074233.4
ENSMUST00000051389.8 |
Synm
|
synemin, intermediate filament protein |
| chr14_+_65971164 | 1.07 |
ENSMUST00000144619.1
|
Clu
|
clusterin |
| chr17_+_37046555 | 1.06 |
ENSMUST00000172789.1
|
Gabbr1
|
gamma-aminobutyric acid (GABA) B receptor, 1 |
| chr2_+_34772089 | 1.06 |
ENSMUST00000028222.6
ENSMUST00000100171.2 |
Hspa5
|
heat shock protein 5 |
| chr2_+_180598219 | 1.05 |
ENSMUST00000103059.1
|
Col9a3
|
collagen, type IX, alpha 3 |
| chr12_+_105453831 | 1.05 |
ENSMUST00000178224.1
|
D430019H16Rik
|
RIKEN cDNA D430019H16 gene |
| chr14_-_46390576 | 1.05 |
ENSMUST00000074077.5
|
Bmp4
|
bone morphogenetic protein 4 |
| chr1_+_180935022 | 1.04 |
ENSMUST00000037361.8
|
Lefty1
|
left right determination factor 1 |
| chr16_+_41532999 | 1.04 |
ENSMUST00000099761.3
|
Lsamp
|
limbic system-associated membrane protein |
| chr13_-_95444827 | 1.04 |
ENSMUST00000045583.7
|
Crhbp
|
corticotropin releasing hormone binding protein |
| chr13_-_58610877 | 1.04 |
ENSMUST00000022036.7
|
Slc28a3
|
solute carrier family 28 (sodium-coupled nucleoside transporter), member 3 |
| chr9_-_26806384 | 1.03 |
ENSMUST00000162702.1
ENSMUST00000040398.7 ENSMUST00000066560.6 |
Glb1l2
|
galactosidase, beta 1-like 2 |
| chr5_-_103211251 | 1.02 |
ENSMUST00000060871.5
ENSMUST00000112846.1 ENSMUST00000170792.1 ENSMUST00000112847.2 |
Mapk10
|
mitogen-activated protein kinase 10 |
| chr6_+_70844499 | 1.02 |
ENSMUST00000034093.8
ENSMUST00000162950.1 |
Eif2ak3
|
eukaryotic translation initiation factor 2 alpha kinase 3 |
| chr4_-_93335510 | 1.02 |
ENSMUST00000066774.4
|
Tusc1
|
tumor suppressor candidate 1 |
| chr1_-_75264195 | 1.02 |
ENSMUST00000027404.5
|
Ptprn
|
protein tyrosine phosphatase, receptor type, N |
| chr3_+_138065052 | 1.02 |
ENSMUST00000163080.2
|
1110002E22Rik
|
RIKEN cDNA 1110002E22 gene |
| chr16_-_52452654 | 1.02 |
ENSMUST00000168071.1
|
Alcam
|
activated leukocyte cell adhesion molecule |
| chr13_-_114932035 | 1.02 |
ENSMUST00000056117.8
|
Itga2
|
integrin alpha 2 |
| chr10_+_79997463 | 1.01 |
ENSMUST00000171637.1
ENSMUST00000043866.7 |
Abca7
|
ATP-binding cassette, sub-family A (ABC1), member 7 |
| chr11_-_99979053 | 1.01 |
ENSMUST00000105051.1
|
Krtap29-1
|
keratin associated protein 29-1 |
| chr10_+_87860030 | 1.01 |
ENSMUST00000062862.6
|
Igf1
|
insulin-like growth factor 1 |
| chr19_-_3686549 | 1.00 |
ENSMUST00000025856.10
ENSMUST00000176867.1 |
Lrp5
|
low density lipoprotein receptor-related protein 5 |
| chr17_+_35320529 | 1.00 |
ENSMUST00000105041.3
ENSMUST00000073208.5 |
H2-Q1
|
histocompatibility 2, Q region locus 1 |
| chr17_+_17831004 | 0.99 |
ENSMUST00000172097.2
|
4930546H06Rik
|
RIKEN cDNA 4930546H06 gene |
| chr10_+_69212634 | 0.99 |
ENSMUST00000020101.5
|
Rhobtb1
|
Rho-related BTB domain containing 1 |
| chr3_+_135826075 | 0.99 |
ENSMUST00000029810.5
|
Slc39a8
|
solute carrier family 39 (metal ion transporter), member 8 |
| chr10_-_81545175 | 0.99 |
ENSMUST00000043604.5
|
Gna11
|
guanine nucleotide binding protein, alpha 11 |
| chr5_+_134099704 | 0.98 |
ENSMUST00000016088.8
|
Gatsl2
|
GATS protein-like 2 |
| chr19_-_56822161 | 0.98 |
ENSMUST00000118592.1
|
A630007B06Rik
|
RIKEN cDNA A630007B06 gene |
| chr19_-_36919606 | 0.98 |
ENSMUST00000057337.7
|
Fgfbp3
|
fibroblast growth factor binding protein 3 |
| chr4_-_133263042 | 0.98 |
ENSMUST00000105908.3
ENSMUST00000030674.7 |
Sytl1
|
synaptotagmin-like 1 |
| chr13_+_64161862 | 0.97 |
ENSMUST00000021929.8
|
Habp4
|
hyaluronic acid binding protein 4 |
| chr1_+_74362108 | 0.97 |
ENSMUST00000097697.1
|
Gm216
|
predicted gene 216 |
| chr11_-_101894355 | 0.96 |
ENSMUST00000057054.7
|
Meox1
|
mesenchyme homeobox 1 |
| chr2_+_174330006 | 0.95 |
ENSMUST00000109085.1
ENSMUST00000109087.1 ENSMUST00000109084.1 |
Gnas
|
GNAS (guanine nucleotide binding protein, alpha stimulating) complex locus |
| chr6_+_125552948 | 0.94 |
ENSMUST00000112254.1
ENSMUST00000112253.1 ENSMUST00000001995.7 |
Vwf
|
Von Willebrand factor homolog |
| chr14_-_46390501 | 0.94 |
ENSMUST00000100676.2
|
Bmp4
|
bone morphogenetic protein 4 |
| chr7_+_51878967 | 0.93 |
ENSMUST00000051912.6
|
Gas2
|
growth arrest specific 2 |
| chr7_-_43660139 | 0.93 |
ENSMUST00000032667.8
|
Siglece
|
sialic acid binding Ig-like lectin E |
| chr17_+_26113286 | 0.92 |
ENSMUST00000025010.7
|
Tmem8
|
transmembrane protein 8 (five membrane-spanning domains) |
| chr11_+_50602072 | 0.91 |
ENSMUST00000040523.8
|
Adamts2
|
a disintegrin-like and metallopeptidase (reprolysin type) with thrombospondin type 1 motif, 2 |
| chr7_+_73740277 | 0.91 |
ENSMUST00000107456.2
|
Fam174b
|
family with sequence similarity 174, member B |
| chr10_+_77581720 | 0.91 |
ENSMUST00000009435.5
|
Pttg1ip
|
pituitary tumor-transforming 1 interacting protein |
| chr7_-_114562945 | 0.91 |
ENSMUST00000119712.1
ENSMUST00000032908.8 |
Cyp2r1
|
cytochrome P450, family 2, subfamily r, polypeptide 1 |
| chr5_+_43233928 | 0.91 |
ENSMUST00000114066.1
ENSMUST00000114065.1 |
Cpeb2
|
cytoplasmic polyadenylation element binding protein 2 |
| chr6_-_72617000 | 0.91 |
ENSMUST00000070524.4
|
Tgoln1
|
trans-golgi network protein |
| chr2_-_66784903 | 0.90 |
ENSMUST00000042792.6
|
Scn7a
|
sodium channel, voltage-gated, type VII, alpha |
| chr3_-_89402650 | 0.90 |
ENSMUST00000168325.1
ENSMUST00000057431.5 |
Lenep
|
lens epithelial protein |
| chr2_-_13793793 | 0.89 |
ENSMUST00000003509.8
|
St8sia6
|
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 6 |
| chr16_-_22657165 | 0.89 |
ENSMUST00000089925.3
|
Dgkg
|
diacylglycerol kinase, gamma |
| chr1_-_156674290 | 0.89 |
ENSMUST00000079625.4
|
Tor3a
|
torsin family 3, member A |
| chr1_+_162639148 | 0.89 |
ENSMUST00000028020.9
|
Myoc
|
myocilin |
| chr5_-_34187670 | 0.87 |
ENSMUST00000042701.6
ENSMUST00000119171.1 |
Mxd4
|
Max dimerization protein 4 |
| chr10_+_85928491 | 0.87 |
ENSMUST00000170396.1
|
Ascl4
|
achaete-scute complex homolog 4 (Drosophila) |
Network of associatons between targets according to the STRING database.
First level regulatory network of Myf6
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 2.9 | GO:0046087 | cytidine catabolic process(GO:0006216) cytidine deamination(GO:0009972) cytidine metabolic process(GO:0046087) |
| 1.0 | 4.8 | GO:0006987 | activation of signaling protein activity involved in unfolded protein response(GO:0006987) |
| 0.9 | 2.8 | GO:1903048 | regulation of acetylcholine-gated cation channel activity(GO:1903048) |
| 0.9 | 3.5 | GO:0001992 | regulation of systemic arterial blood pressure by vasopressin(GO:0001992) |
| 0.9 | 2.6 | GO:0097212 | lysosomal membrane organization(GO:0097212) |
| 0.8 | 2.4 | GO:0051464 | positive regulation of cortisol secretion(GO:0051464) |
| 0.8 | 2.3 | GO:0007386 | compartment pattern specification(GO:0007386) |
| 0.7 | 2.0 | GO:0018199 | peptidyl-glutamine modification(GO:0018199) |
| 0.7 | 2.0 | GO:0003130 | BMP signaling pathway involved in heart induction(GO:0003130) endodermal-mesodermal cell signaling(GO:0003133) endodermal-mesodermal cell signaling involved in heart induction(GO:0003134) intermediate mesoderm development(GO:0048389) cloacal septation(GO:0060197) regulation of transcription from RNA polymerase II promoter involved in kidney development(GO:0060994) pattern specification involved in mesonephros development(GO:0061227) anterior/posterior pattern specification involved in kidney development(GO:0072098) regulation of mesenchymal stem cell proliferation(GO:1902460) positive regulation of mesenchymal stem cell proliferation(GO:1902462) cardiac jelly development(GO:1905072) negative regulation of cell proliferation involved in heart morphogenesis(GO:2000137) |
| 0.6 | 1.9 | GO:0050925 | negative regulation of negative chemotaxis(GO:0050925) |
| 0.5 | 1.6 | GO:1901662 | phylloquinone metabolic process(GO:0042374) phylloquinone catabolic process(GO:0042376) quinone catabolic process(GO:1901662) |
| 0.5 | 6.8 | GO:0060363 | cranial suture morphogenesis(GO:0060363) |
| 0.5 | 1.5 | GO:0002588 | myeloid dendritic cell activation involved in immune response(GO:0002277) positive regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002582) positive regulation of antigen processing and presentation of peptide antigen via MHC class II(GO:0002588) |
| 0.5 | 1.4 | GO:0060488 | orthogonal dichotomous subdivision of terminal units involved in lung branching morphogenesis(GO:0060488) planar dichotomous subdivision of terminal units involved in lung branching morphogenesis(GO:0060489) lateral sprouting involved in lung morphogenesis(GO:0060490) |
| 0.5 | 5.2 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.5 | 2.3 | GO:0001692 | histamine metabolic process(GO:0001692) imidazole-containing compound catabolic process(GO:0052805) |
| 0.5 | 2.7 | GO:1902847 | regulation of neuronal signal transduction(GO:1902847) positive regulation of neurofibrillary tangle assembly(GO:1902998) |
| 0.4 | 3.6 | GO:0015074 | DNA integration(GO:0015074) |
| 0.4 | 1.3 | GO:0060574 | bronchiole development(GO:0060435) intestinal epithelial cell maturation(GO:0060574) |
| 0.4 | 3.9 | GO:1900034 | regulation of cellular response to heat(GO:1900034) |
| 0.4 | 0.4 | GO:1902996 | neurofibrillary tangle assembly(GO:1902988) regulation of neurofibrillary tangle assembly(GO:1902996) |
| 0.4 | 1.2 | GO:0071314 | cellular response to cocaine(GO:0071314) |
| 0.4 | 2.0 | GO:0045163 | clustering of voltage-gated potassium channels(GO:0045163) |
| 0.4 | 1.2 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.4 | 1.2 | GO:0046077 | dUDP biosynthetic process(GO:0006227) pyrimidine nucleoside diphosphate metabolic process(GO:0009138) pyrimidine nucleoside diphosphate biosynthetic process(GO:0009139) pyrimidine deoxyribonucleoside diphosphate metabolic process(GO:0009196) pyrimidine deoxyribonucleoside diphosphate biosynthetic process(GO:0009197) dUDP metabolic process(GO:0046077) |
| 0.4 | 0.4 | GO:2000040 | regulation of planar cell polarity pathway involved in axis elongation(GO:2000040) negative regulation of planar cell polarity pathway involved in axis elongation(GO:2000041) |
| 0.4 | 1.5 | GO:2000439 | positive regulation of monocyte extravasation(GO:2000439) |
| 0.4 | 1.1 | GO:0060060 | post-embryonic retina morphogenesis in camera-type eye(GO:0060060) |
| 0.4 | 7.4 | GO:0001573 | ganglioside metabolic process(GO:0001573) |
| 0.3 | 1.4 | GO:0010510 | regulation of acetyl-CoA biosynthetic process from pyruvate(GO:0010510) |
| 0.3 | 1.0 | GO:1901074 | regulation of engulfment of apoptotic cell(GO:1901074) |
| 0.3 | 1.7 | GO:0061086 | negative regulation of histone H3-K27 methylation(GO:0061086) |
| 0.3 | 1.6 | GO:0002901 | mature B cell apoptotic process(GO:0002901) regulation of mature B cell apoptotic process(GO:0002905) negative regulation of mature B cell apoptotic process(GO:0002906) |
| 0.3 | 3.4 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.3 | 1.2 | GO:0014722 | regulation of skeletal muscle contraction by calcium ion signaling(GO:0014722) |
| 0.3 | 1.5 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.3 | 1.5 | GO:0060244 | negative regulation of cell proliferation involved in contact inhibition(GO:0060244) |
| 0.3 | 2.9 | GO:0046959 | habituation(GO:0046959) |
| 0.3 | 1.2 | GO:0036343 | psychomotor behavior(GO:0036343) positive regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900738) |
| 0.3 | 0.9 | GO:0043323 | positive regulation of natural killer cell degranulation(GO:0043323) |
| 0.3 | 1.2 | GO:2000138 | positive regulation of cell proliferation involved in heart morphogenesis(GO:2000138) |
| 0.3 | 3.6 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.3 | 1.4 | GO:1903553 | positive regulation of extracellular exosome assembly(GO:1903553) |
| 0.3 | 1.7 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
| 0.3 | 0.8 | GO:0007161 | calcium-independent cell-matrix adhesion(GO:0007161) |
| 0.3 | 0.3 | GO:0042706 | eye photoreceptor cell fate commitment(GO:0042706) photoreceptor cell fate commitment(GO:0046552) camera-type eye photoreceptor cell fate commitment(GO:0060220) |
| 0.3 | 1.1 | GO:0060075 | regulation of resting membrane potential(GO:0060075) |
| 0.3 | 1.1 | GO:0031443 | fast-twitch skeletal muscle fiber contraction(GO:0031443) |
| 0.3 | 2.1 | GO:0045900 | negative regulation of translational elongation(GO:0045900) |
| 0.3 | 1.3 | GO:0035405 | histone-threonine phosphorylation(GO:0035405) |
| 0.3 | 1.6 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.3 | 1.0 | GO:0016095 | polyprenol catabolic process(GO:0016095) |
| 0.3 | 1.0 | GO:1900108 | negative regulation of nodal signaling pathway(GO:1900108) |
| 0.3 | 1.5 | GO:0031580 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 0.2 | 1.2 | GO:0002589 | regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002589) |
| 0.2 | 1.0 | GO:1904395 | positive regulation of skeletal muscle acetylcholine-gated channel clustering(GO:1904395) |
| 0.2 | 2.9 | GO:0032463 | negative regulation of protein homooligomerization(GO:0032463) |
| 0.2 | 1.0 | GO:0061056 | sclerotome development(GO:0061056) |
| 0.2 | 1.2 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.2 | 0.7 | GO:0090341 | negative regulation of secretion of lysosomal enzymes(GO:0090341) |
| 0.2 | 2.6 | GO:0006707 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.2 | 0.9 | GO:1905154 | negative regulation of membrane invagination(GO:1905154) |
| 0.2 | 1.6 | GO:0061074 | regulation of neural retina development(GO:0061074) |
| 0.2 | 0.7 | GO:0019074 | viral genome packaging(GO:0019072) viral RNA genome packaging(GO:0019074) |
| 0.2 | 0.9 | GO:0014734 | skeletal muscle hypertrophy(GO:0014734) |
| 0.2 | 1.3 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 0.2 | 0.7 | GO:0035585 | calcium-mediated signaling using extracellular calcium source(GO:0035585) |
| 0.2 | 0.2 | GO:0035801 | adrenal cortex development(GO:0035801) adrenal cortex formation(GO:0035802) |
| 0.2 | 0.4 | GO:0002485 | antigen processing and presentation of endogenous peptide antigen via MHC class I via ER pathway(GO:0002484) antigen processing and presentation of endogenous peptide antigen via MHC class I via ER pathway, TAP-dependent(GO:0002485) |
| 0.2 | 0.8 | GO:0097026 | dendritic cell dendrite assembly(GO:0097026) |
| 0.2 | 1.0 | GO:0015860 | purine nucleoside transmembrane transport(GO:0015860) nucleoside transmembrane transport(GO:1901642) |
| 0.2 | 0.4 | GO:0015675 | nickel cation transport(GO:0015675) |
| 0.2 | 0.4 | GO:0021577 | hindbrain structural organization(GO:0021577) cerebellum structural organization(GO:0021589) |
| 0.2 | 0.8 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 0.2 | 1.8 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.2 | 1.2 | GO:2000271 | positive regulation of fibroblast apoptotic process(GO:2000271) |
| 0.2 | 0.4 | GO:1903279 | regulation of calcium:sodium antiporter activity(GO:1903279) |
| 0.2 | 1.0 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) |
| 0.2 | 0.2 | GO:1901187 | regulation of ephrin receptor signaling pathway(GO:1901187) |
| 0.2 | 0.7 | GO:0009414 | response to water deprivation(GO:0009414) |
| 0.2 | 0.6 | GO:2000987 | positive regulation of fear response(GO:1903367) positive regulation of behavioral fear response(GO:2000987) |
| 0.2 | 1.1 | GO:0071787 | endoplasmic reticulum tubular network assembly(GO:0071787) |
| 0.2 | 1.4 | GO:0097647 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.2 | 0.7 | GO:0072014 | proximal tubule development(GO:0072014) |
| 0.2 | 0.4 | GO:1903334 | positive regulation of protein folding(GO:1903334) |
| 0.2 | 0.4 | GO:1903588 | negative regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903588) |
| 0.2 | 1.0 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.2 | 1.0 | GO:0070417 | cellular response to cold(GO:0070417) |
| 0.2 | 0.9 | GO:0032423 | regulation of mismatch repair(GO:0032423) |
| 0.2 | 0.2 | GO:0032831 | regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032829) positive regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032831) |
| 0.2 | 0.8 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.2 | 1.0 | GO:1904073 | regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 0.2 | 0.8 | GO:2000255 | negative regulation of male germ cell proliferation(GO:2000255) |
| 0.2 | 2.4 | GO:0098719 | sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.2 | 1.3 | GO:0060024 | rhythmic synaptic transmission(GO:0060024) |
| 0.2 | 0.5 | GO:1901256 | macrophage colony-stimulating factor production(GO:0036301) granulocyte colony-stimulating factor production(GO:0071611) regulation of granulocyte colony-stimulating factor production(GO:0071655) regulation of macrophage colony-stimulating factor production(GO:1901256) negative regulation of immunological synapse formation(GO:2000521) regulation of T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:2001188) negative regulation of T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:2001189) |
| 0.2 | 1.4 | GO:0060923 | cardiac muscle cell fate commitment(GO:0060923) |
| 0.2 | 0.6 | GO:0032747 | positive regulation of interleukin-23 production(GO:0032747) chemokine (C-X-C motif) ligand 1 production(GO:0072566) regulation of chemokine (C-X-C motif) ligand 1 production(GO:2000338) |
| 0.2 | 1.4 | GO:0033631 | cell-cell adhesion mediated by integrin(GO:0033631) |
| 0.2 | 1.1 | GO:0014053 | negative regulation of gamma-aminobutyric acid secretion(GO:0014053) |
| 0.1 | 1.6 | GO:2000465 | regulation of glycogen (starch) synthase activity(GO:2000465) |
| 0.1 | 5.5 | GO:0009409 | response to cold(GO:0009409) |
| 0.1 | 0.6 | GO:0070318 | positive regulation of G0 to G1 transition(GO:0070318) |
| 0.1 | 1.3 | GO:0045717 | negative regulation of fatty acid biosynthetic process(GO:0045717) |
| 0.1 | 1.0 | GO:0010694 | positive regulation of alkaline phosphatase activity(GO:0010694) |
| 0.1 | 1.6 | GO:0030432 | peristalsis(GO:0030432) |
| 0.1 | 0.6 | GO:0002265 | astrocyte activation involved in immune response(GO:0002265) positive regulation of lysosomal protein catabolic process(GO:1905167) |
| 0.1 | 1.9 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.1 | 0.3 | GO:0039534 | negative regulation of MDA-5 signaling pathway(GO:0039534) |
| 0.1 | 0.4 | GO:0061402 | glycerol biosynthetic process(GO:0006114) alditol biosynthetic process(GO:0019401) positive regulation of transcription from RNA polymerase II promoter in response to acidic pH(GO:0061402) |
| 0.1 | 0.4 | GO:0038027 | apolipoprotein A-I-mediated signaling pathway(GO:0038027) negative regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045715) |
| 0.1 | 0.6 | GO:0021993 | initiation of neural tube closure(GO:0021993) |
| 0.1 | 1.1 | GO:0003096 | renal sodium ion transport(GO:0003096) renal sodium ion absorption(GO:0070294) |
| 0.1 | 1.5 | GO:1901898 | negative regulation of relaxation of muscle(GO:1901078) negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.1 | 0.7 | GO:0071442 | positive regulation of histone H3-K14 acetylation(GO:0071442) |
| 0.1 | 0.4 | GO:0072708 | response to sorbitol(GO:0072708) |
| 0.1 | 0.5 | GO:0042373 | vitamin K metabolic process(GO:0042373) |
| 0.1 | 0.7 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.1 | 0.8 | GO:0048133 | germ-line stem cell division(GO:0042078) male germ-line stem cell asymmetric division(GO:0048133) germline stem cell asymmetric division(GO:0098728) |
| 0.1 | 0.7 | GO:0043314 | negative regulation of neutrophil degranulation(GO:0043314) negative regulation of blood vessel remodeling(GO:0060313) negative regulation of neutrophil activation(GO:1902564) |
| 0.1 | 0.7 | GO:0001970 | positive regulation of activation of membrane attack complex(GO:0001970) |
| 0.1 | 1.4 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.1 | 2.8 | GO:0090023 | positive regulation of neutrophil chemotaxis(GO:0090023) |
| 0.1 | 0.5 | GO:0072344 | rescue of stalled ribosome(GO:0072344) |
| 0.1 | 1.5 | GO:0043518 | negative regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043518) |
| 0.1 | 1.4 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.1 | 0.8 | GO:0015808 | L-alanine transport(GO:0015808) |
| 0.1 | 0.6 | GO:0033625 | positive regulation of integrin activation(GO:0033625) |
| 0.1 | 0.4 | GO:1903365 | regulation of fear response(GO:1903365) regulation of behavioral fear response(GO:2000822) |
| 0.1 | 3.1 | GO:0040037 | negative regulation of fibroblast growth factor receptor signaling pathway(GO:0040037) |
| 0.1 | 0.4 | GO:0090204 | protein localization to nuclear pore(GO:0090204) |
| 0.1 | 1.2 | GO:0071285 | cellular response to lithium ion(GO:0071285) |
| 0.1 | 0.5 | GO:0043602 | nitrate catabolic process(GO:0043602) nitric oxide catabolic process(GO:0046210) cellular organohalogen metabolic process(GO:0090345) cellular organofluorine metabolic process(GO:0090346) |
| 0.1 | 0.5 | GO:0003150 | muscular septum morphogenesis(GO:0003150) |
| 0.1 | 1.2 | GO:0051256 | mitotic spindle midzone assembly(GO:0051256) |
| 0.1 | 0.3 | GO:1905146 | lysosomal protein catabolic process(GO:1905146) |
| 0.1 | 0.7 | GO:0090315 | negative regulation of protein targeting to membrane(GO:0090315) |
| 0.1 | 0.7 | GO:0002317 | plasma cell differentiation(GO:0002317) |
| 0.1 | 0.5 | GO:0060849 | regulation of transcription involved in lymphatic endothelial cell fate commitment(GO:0060849) |
| 0.1 | 0.4 | GO:1904453 | regulation of hydrogen:potassium-exchanging ATPase activity(GO:1904451) positive regulation of hydrogen:potassium-exchanging ATPase activity(GO:1904453) |
| 0.1 | 0.4 | GO:0010808 | positive regulation of synaptic vesicle priming(GO:0010808) |
| 0.1 | 0.5 | GO:0060666 | dichotomous subdivision of terminal units involved in salivary gland branching(GO:0060666) |
| 0.1 | 1.9 | GO:0006516 | glycoprotein catabolic process(GO:0006516) |
| 0.1 | 0.8 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.1 | 1.4 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.1 | 1.0 | GO:0015691 | cadmium ion transport(GO:0015691) cadmium ion transmembrane transport(GO:0070574) |
| 0.1 | 1.3 | GO:0032494 | response to peptidoglycan(GO:0032494) |
| 0.1 | 0.3 | GO:0021649 | vestibulocochlear nerve structural organization(GO:0021649) ganglion morphogenesis(GO:0061552) dorsal root ganglion morphogenesis(GO:1904835) |
| 0.1 | 0.5 | GO:0032472 | Golgi calcium ion transport(GO:0032472) |
| 0.1 | 0.6 | GO:0048102 | autophagic cell death(GO:0048102) |
| 0.1 | 1.8 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.1 | 0.5 | GO:0015879 | carnitine transport(GO:0015879) |
| 0.1 | 0.3 | GO:0010847 | regulation of chromatin assembly(GO:0010847) |
| 0.1 | 1.7 | GO:0035493 | SNARE complex assembly(GO:0035493) |
| 0.1 | 0.6 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.1 | 0.6 | GO:0030242 | pexophagy(GO:0030242) |
| 0.1 | 0.8 | GO:0000255 | allantoin metabolic process(GO:0000255) |
| 0.1 | 0.2 | GO:0021698 | cerebellar cortex structural organization(GO:0021698) |
| 0.1 | 0.9 | GO:0043117 | positive regulation of vascular permeability(GO:0043117) |
| 0.1 | 0.3 | GO:0021664 | rhombomere formation(GO:0021594) rhombomere 3 formation(GO:0021660) rhombomere 5 morphogenesis(GO:0021664) rhombomere 5 formation(GO:0021666) |
| 0.1 | 1.0 | GO:0061469 | regulation of type B pancreatic cell proliferation(GO:0061469) |
| 0.1 | 0.3 | GO:0070829 | heterochromatin maintenance(GO:0070829) |
| 0.1 | 0.2 | GO:1904058 | positive regulation of sensory perception of pain(GO:1904058) |
| 0.1 | 0.2 | GO:0060466 | activation of meiosis involved in egg activation(GO:0060466) |
| 0.1 | 0.4 | GO:0002669 | positive regulation of T cell anergy(GO:0002669) positive regulation of lymphocyte anergy(GO:0002913) |
| 0.1 | 1.5 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.1 | 1.5 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.1 | 0.4 | GO:0010572 | positive regulation of platelet activation(GO:0010572) |
| 0.1 | 0.9 | GO:0052696 | flavonoid glucuronidation(GO:0052696) xenobiotic glucuronidation(GO:0052697) |
| 0.1 | 0.4 | GO:1902732 | positive regulation of chondrocyte proliferation(GO:1902732) |
| 0.1 | 0.5 | GO:0045196 | establishment or maintenance of neuroblast polarity(GO:0045196) establishment of neuroblast polarity(GO:0045200) regulation of cardiac muscle cell action potential involved in regulation of contraction(GO:0098909) |
| 0.1 | 0.2 | GO:0006713 | glucocorticoid catabolic process(GO:0006713) |
| 0.1 | 0.4 | GO:0050916 | sensory perception of sweet taste(GO:0050916) |
| 0.1 | 0.1 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.1 | 0.2 | GO:0070885 | negative regulation of calcineurin-NFAT signaling cascade(GO:0070885) |
| 0.1 | 1.1 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.1 | 0.3 | GO:0009814 | defense response, incompatible interaction(GO:0009814) defense response to bacterium, incompatible interaction(GO:0009816) regulation of defense response to bacterium, incompatible interaction(GO:1902477) |
| 0.1 | 1.1 | GO:1902260 | negative regulation of delayed rectifier potassium channel activity(GO:1902260) negative regulation of voltage-gated potassium channel activity(GO:1903817) |
| 0.1 | 0.4 | GO:0061462 | protein localization to lysosome(GO:0061462) |
| 0.1 | 0.7 | GO:0010748 | negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) |
| 0.1 | 0.1 | GO:0002645 | positive regulation of tolerance induction(GO:0002645) |
| 0.1 | 0.1 | GO:0097374 | sensory neuron axon guidance(GO:0097374) |
| 0.1 | 0.4 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.1 | 0.7 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 0.1 | 0.4 | GO:0060718 | chorionic trophoblast cell differentiation(GO:0060718) |
| 0.1 | 0.9 | GO:0060670 | branching involved in labyrinthine layer morphogenesis(GO:0060670) |
| 0.1 | 0.2 | GO:0048105 | establishment of body hair or bristle planar orientation(GO:0048104) establishment of body hair planar orientation(GO:0048105) |
| 0.1 | 0.5 | GO:0032926 | negative regulation of activin receptor signaling pathway(GO:0032926) |
| 0.1 | 0.9 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.1 | 0.3 | GO:0035360 | positive regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035360) |
| 0.1 | 0.9 | GO:1904262 | negative regulation of TORC1 signaling(GO:1904262) |
| 0.1 | 0.1 | GO:0032286 | central nervous system myelin maintenance(GO:0032286) |
| 0.1 | 0.2 | GO:0035630 | bone mineralization involved in bone maturation(GO:0035630) |
| 0.1 | 0.5 | GO:0090385 | phagosome-lysosome fusion(GO:0090385) |
| 0.1 | 1.5 | GO:0043001 | Golgi to plasma membrane protein transport(GO:0043001) |
| 0.1 | 0.3 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.0 | 0.1 | GO:0031392 | regulation of prostaglandin biosynthetic process(GO:0031392) positive regulation of prostaglandin biosynthetic process(GO:0031394) regulation of unsaturated fatty acid biosynthetic process(GO:2001279) positive regulation of unsaturated fatty acid biosynthetic process(GO:2001280) |
| 0.0 | 1.1 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.0 | 0.5 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.0 | 0.5 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 1.8 | GO:1902042 | negative regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902042) |
| 0.0 | 0.2 | GO:1903575 | cornified envelope assembly(GO:1903575) |
| 0.0 | 0.7 | GO:0042359 | vitamin D metabolic process(GO:0042359) |
| 0.0 | 0.3 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.0 | 0.2 | GO:0000432 | regulation of transcription from RNA polymerase II promoter by glucose(GO:0000430) positive regulation of transcription from RNA polymerase II promoter by glucose(GO:0000432) |
| 0.0 | 0.6 | GO:1900454 | positive regulation of long term synaptic depression(GO:1900454) |
| 0.0 | 0.5 | GO:0036295 | cellular response to increased oxygen levels(GO:0036295) |
| 0.0 | 0.5 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.0 | 1.6 | GO:0002474 | antigen processing and presentation of peptide antigen via MHC class I(GO:0002474) |
| 0.0 | 0.3 | GO:0035878 | nail development(GO:0035878) |
| 0.0 | 0.3 | GO:0042905 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.0 | 0.3 | GO:0071698 | olfactory placode formation(GO:0030910) olfactory placode development(GO:0071698) olfactory placode morphogenesis(GO:0071699) |
| 0.0 | 0.3 | GO:0072513 | positive regulation of secondary heart field cardioblast proliferation(GO:0072513) |
| 0.0 | 1.2 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 0.3 | GO:0034116 | positive regulation of heterotypic cell-cell adhesion(GO:0034116) regulation of Rho-dependent protein serine/threonine kinase activity(GO:2000298) |
| 0.0 | 0.2 | GO:1903849 | regulation of aorta morphogenesis(GO:1903847) positive regulation of aorta morphogenesis(GO:1903849) |
| 0.0 | 0.2 | GO:0035633 | cyclooxygenase pathway(GO:0019371) maintenance of blood-brain barrier(GO:0035633) |
| 0.0 | 2.1 | GO:0019228 | neuronal action potential(GO:0019228) |
| 0.0 | 0.7 | GO:0050901 | leukocyte tethering or rolling(GO:0050901) |
| 0.0 | 0.0 | GO:0001805 | type III hypersensitivity(GO:0001802) regulation of type III hypersensitivity(GO:0001803) positive regulation of type III hypersensitivity(GO:0001805) |
| 0.0 | 0.5 | GO:0006654 | phosphatidic acid biosynthetic process(GO:0006654) |
| 0.0 | 0.8 | GO:0036342 | post-anal tail morphogenesis(GO:0036342) |
| 0.0 | 0.1 | GO:0046005 | positive regulation of circadian sleep/wake cycle, REM sleep(GO:0046005) |
| 0.0 | 0.2 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 0.0 | 0.7 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.0 | 0.2 | GO:0090273 | regulation of somatostatin secretion(GO:0090273) |
| 0.0 | 1.4 | GO:0010955 | negative regulation of protein processing(GO:0010955) negative regulation of protein maturation(GO:1903318) |
| 0.0 | 0.5 | GO:0010759 | positive regulation of macrophage chemotaxis(GO:0010759) |
| 0.0 | 0.3 | GO:0030046 | parallel actin filament bundle assembly(GO:0030046) |
| 0.0 | 0.9 | GO:0006691 | leukotriene metabolic process(GO:0006691) |
| 0.0 | 2.0 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.1 | GO:0046951 | ketone body biosynthetic process(GO:0046951) |
| 0.0 | 0.1 | GO:0003358 | noradrenergic neuron development(GO:0003358) |
| 0.0 | 2.9 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.0 | 1.6 | GO:0006493 | protein O-linked glycosylation(GO:0006493) |
| 0.0 | 1.1 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 0.6 | GO:0035518 | histone H2A monoubiquitination(GO:0035518) |
| 0.0 | 0.1 | GO:0003220 | left ventricular cardiac muscle tissue morphogenesis(GO:0003220) |
| 0.0 | 0.7 | GO:0060445 | branching involved in salivary gland morphogenesis(GO:0060445) |
| 0.0 | 0.5 | GO:0051968 | positive regulation of synaptic transmission, glutamatergic(GO:0051968) |
| 0.0 | 0.8 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.2 | GO:0001835 | blastocyst hatching(GO:0001835) hatching(GO:0035188) organism emergence from protective structure(GO:0071684) |
| 0.0 | 2.3 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.0 | 1.0 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.0 | 0.3 | GO:0060159 | regulation of dopamine receptor signaling pathway(GO:0060159) |
| 0.0 | 0.6 | GO:0048505 | regulation of timing of cell differentiation(GO:0048505) |
| 0.0 | 0.1 | GO:0043652 | engulfment of apoptotic cell(GO:0043652) |
| 0.0 | 0.2 | GO:0050957 | equilibrioception(GO:0050957) |
| 0.0 | 2.2 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.0 | 0.1 | GO:0014028 | notochord formation(GO:0014028) |
| 0.0 | 0.5 | GO:0060314 | regulation of ryanodine-sensitive calcium-release channel activity(GO:0060314) |
| 0.0 | 0.4 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 0.0 | 1.0 | GO:0006308 | DNA catabolic process(GO:0006308) |
| 0.0 | 0.1 | GO:0006667 | sphinganine metabolic process(GO:0006667) |
| 0.0 | 0.3 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.0 | 0.1 | GO:0010626 | negative regulation of Schwann cell proliferation(GO:0010626) |
| 0.0 | 0.2 | GO:0002756 | MyD88-independent toll-like receptor signaling pathway(GO:0002756) |
| 0.0 | 0.6 | GO:0035066 | positive regulation of histone acetylation(GO:0035066) |
| 0.0 | 0.3 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.0 | 0.1 | GO:0002767 | immune response-inhibiting cell surface receptor signaling pathway(GO:0002767) |
| 0.0 | 0.1 | GO:0021691 | cerebellar Purkinje cell layer maturation(GO:0021691) |
| 0.0 | 0.2 | GO:0035887 | aortic smooth muscle cell differentiation(GO:0035887) |
| 0.0 | 0.8 | GO:0030225 | macrophage differentiation(GO:0030225) |
| 0.0 | 0.2 | GO:0002051 | osteoblast fate commitment(GO:0002051) |
| 0.0 | 0.1 | GO:0090160 | Golgi to lysosome transport(GO:0090160) |
| 0.0 | 0.2 | GO:0043568 | positive regulation of insulin-like growth factor receptor signaling pathway(GO:0043568) |
| 0.0 | 0.4 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.0 | 0.4 | GO:0007614 | short-term memory(GO:0007614) |
| 0.0 | 0.8 | GO:0021846 | cell proliferation in forebrain(GO:0021846) |
| 0.0 | 0.1 | GO:0046543 | development of secondary female sexual characteristics(GO:0046543) |
| 0.0 | 0.8 | GO:1903078 | positive regulation of protein localization to plasma membrane(GO:1903078) |
| 0.0 | 0.1 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 0.0 | 0.0 | GO:0010536 | positive regulation of activation of Janus kinase activity(GO:0010536) positive regulation of natural killer cell proliferation(GO:0032819) |
| 0.0 | 0.8 | GO:0010862 | positive regulation of pathway-restricted SMAD protein phosphorylation(GO:0010862) |
| 0.0 | 0.2 | GO:0043144 | snoRNA processing(GO:0043144) |
| 0.0 | 0.8 | GO:0010257 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.0 | 0.1 | GO:0006177 | GMP biosynthetic process(GO:0006177) |
| 0.0 | 0.5 | GO:0010666 | positive regulation of striated muscle cell apoptotic process(GO:0010663) positive regulation of cardiac muscle cell apoptotic process(GO:0010666) |
| 0.0 | 0.1 | GO:2000189 | positive regulation of cholesterol homeostasis(GO:2000189) |
| 0.0 | 0.9 | GO:0070373 | negative regulation of ERK1 and ERK2 cascade(GO:0070373) |
| 0.0 | 1.5 | GO:0007173 | epidermal growth factor receptor signaling pathway(GO:0007173) |
| 0.0 | 0.4 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.4 | GO:0033169 | histone H3-K9 demethylation(GO:0033169) |
| 0.0 | 0.9 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.0 | 0.1 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 0.0 | 0.0 | GO:0021853 | cerebral cortex GABAergic interneuron migration(GO:0021853) interneuron migration(GO:1904936) |
| 0.0 | 0.2 | GO:0048339 | paraxial mesoderm development(GO:0048339) |
| 0.0 | 0.0 | GO:0035523 | protein K29-linked deubiquitination(GO:0035523) protein K33-linked deubiquitination(GO:1990168) |
| 0.0 | 0.2 | GO:1901522 | positive regulation of transcription from RNA polymerase II promoter involved in cellular response to chemical stimulus(GO:1901522) |
| 0.0 | 0.0 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.0 | 0.1 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.0 | 1.2 | GO:0035383 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 0.5 | GO:0060395 | SMAD protein signal transduction(GO:0060395) |
| 0.0 | 0.4 | GO:0045124 | regulation of bone resorption(GO:0045124) |
| 0.0 | 0.5 | GO:0006695 | cholesterol biosynthetic process(GO:0006695) secondary alcohol biosynthetic process(GO:1902653) |
| 0.0 | 0.2 | GO:1900383 | regulation of synaptic plasticity by receptor localization to synapse(GO:1900383) |
| 0.0 | 0.2 | GO:0051904 | melanosome transport(GO:0032402) pigment granule transport(GO:0051904) |
| 0.0 | 0.2 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 0.0 | 0.3 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.0 | 0.4 | GO:0030514 | negative regulation of BMP signaling pathway(GO:0030514) |
| 0.0 | 0.2 | GO:0035094 | response to nicotine(GO:0035094) |
| 0.0 | 0.3 | GO:0045577 | regulation of B cell differentiation(GO:0045577) |
| 0.0 | 0.1 | GO:2000676 | positive regulation of type B pancreatic cell apoptotic process(GO:2000676) |
| 0.0 | 0.5 | GO:0007029 | endoplasmic reticulum organization(GO:0007029) |
| 0.0 | 0.7 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 0.0 | 0.2 | GO:0021702 | regulation of cytoplasmic mRNA processing body assembly(GO:0010603) cerebellar Purkinje cell differentiation(GO:0021702) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 6.8 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.6 | 3.7 | GO:1990604 | IRE1-TRAF2-ASK1 complex(GO:1990604) |
| 0.6 | 2.5 | GO:0030868 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.4 | 3.9 | GO:0045298 | tubulin complex(GO:0045298) |
| 0.4 | 1.5 | GO:0097169 | AIM2 inflammasome complex(GO:0097169) |
| 0.3 | 1.4 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.3 | 1.0 | GO:0034666 | integrin alpha2-beta1 complex(GO:0034666) |
| 0.3 | 1.5 | GO:0070820 | tertiary granule(GO:0070820) |
| 0.3 | 3.5 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.3 | 1.2 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.3 | 0.8 | GO:0070195 | growth hormone receptor complex(GO:0070195) |
| 0.2 | 1.4 | GO:1903439 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.2 | 1.2 | GO:0071256 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 0.2 | 0.9 | GO:0070436 | Grb2-EGFR complex(GO:0070436) |
| 0.2 | 1.1 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.2 | 3.1 | GO:0005797 | Golgi medial cisterna(GO:0005797) |
| 0.2 | 0.9 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) cytolytic granule(GO:0044194) |
| 0.2 | 3.2 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.2 | 1.1 | GO:0038039 | G-protein coupled receptor heterodimeric complex(GO:0038039) |
| 0.2 | 1.1 | GO:0005593 | FACIT collagen trimer(GO:0005593) |
| 0.2 | 0.6 | GO:0036488 | CHOP-C/EBP complex(GO:0036488) |
| 0.2 | 0.6 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.2 | 0.6 | GO:0032280 | symmetric synapse(GO:0032280) |
| 0.2 | 2.7 | GO:0034366 | spherical high-density lipoprotein particle(GO:0034366) |
| 0.2 | 0.7 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 0.2 | 1.4 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.2 | 1.3 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.1 | 0.4 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.1 | 1.9 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.1 | 5.5 | GO:0008305 | integrin complex(GO:0008305) |
| 0.1 | 1.1 | GO:0060053 | neurofilament cytoskeleton(GO:0060053) |
| 0.1 | 0.8 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.1 | 1.6 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.1 | 3.3 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.1 | 2.2 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.1 | 0.8 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.1 | 2.2 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.1 | 0.4 | GO:1990795 | lateral part of cell(GO:0097574) basolateral part of cell(GO:1990794) rod bipolar cell terminal bouton(GO:1990795) |
| 0.1 | 0.3 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 0.1 | 1.4 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.1 | 0.9 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.1 | 2.2 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.1 | 0.7 | GO:0005785 | signal recognition particle receptor complex(GO:0005785) |
| 0.1 | 1.2 | GO:1990023 | mitotic spindle midzone(GO:1990023) |
| 0.1 | 0.6 | GO:1990357 | terminal web(GO:1990357) |
| 0.1 | 1.3 | GO:0044232 | organelle membrane contact site(GO:0044232) |
| 0.1 | 0.3 | GO:0005713 | recombination nodule(GO:0005713) |
| 0.1 | 0.4 | GO:0098831 | presynaptic active zone cytoplasmic component(GO:0098831) |
| 0.1 | 1.0 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.1 | 0.6 | GO:0042825 | TAP complex(GO:0042825) |
| 0.1 | 1.0 | GO:0005767 | secondary lysosome(GO:0005767) |
| 0.1 | 0.6 | GO:0043219 | lateral loop(GO:0043219) |
| 0.1 | 0.7 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.1 | 3.2 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.1 | 0.4 | GO:0001652 | granular component(GO:0001652) |
| 0.1 | 1.3 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.1 | 0.1 | GO:0020018 | ciliary pocket(GO:0020016) ciliary pocket membrane(GO:0020018) |
| 0.1 | 1.9 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.1 | 0.7 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.1 | 0.5 | GO:0035748 | myelin sheath abaxonal region(GO:0035748) |
| 0.1 | 2.3 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 0.2 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.0 | 0.3 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.0 | 0.4 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.0 | 5.2 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 1.2 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.0 | 1.2 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 0.9 | GO:0097386 | glial cell projection(GO:0097386) |
| 0.0 | 1.0 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.0 | 0.2 | GO:0060187 | cell pole(GO:0060187) |
| 0.0 | 0.3 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 0.0 | 1.8 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 0.4 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.0 | 0.2 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.0 | 0.1 | GO:0016942 | insulin-like growth factor binding protein complex(GO:0016942) growth factor complex(GO:0036454) |
| 0.0 | 0.7 | GO:0016471 | vacuolar proton-transporting V-type ATPase complex(GO:0016471) |
| 0.0 | 1.0 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.0 | 0.9 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 0.4 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.0 | 0.6 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.0 | 0.8 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 0.1 | GO:0097208 | alveolar lamellar body(GO:0097208) |
| 0.0 | 0.4 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.0 | 0.9 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.4 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.0 | 0.4 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.0 | 0.4 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 0.0 | 0.9 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 7.1 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 0.9 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 0.3 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.0 | 0.4 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.0 | 3.2 | GO:0005765 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.0 | 2.0 | GO:0042641 | actomyosin(GO:0042641) |
| 0.0 | 0.1 | GO:0031045 | dense core granule(GO:0031045) |
| 0.0 | 0.2 | GO:0030126 | COPI vesicle coat(GO:0030126) COPI-coated vesicle membrane(GO:0030663) |
| 0.0 | 0.2 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.0 | 0.8 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 0.5 | GO:0060170 | ciliary membrane(GO:0060170) |
| 0.0 | 0.1 | GO:0031091 | platelet alpha granule(GO:0031091) |
| 0.0 | 0.1 | GO:0031211 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.0 | 1.2 | GO:0032587 | ruffle membrane(GO:0032587) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 6.1 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.8 | 2.4 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.6 | 2.5 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.6 | 0.6 | GO:0047389 | glycerophosphocholine phosphodiesterase activity(GO:0047389) |
| 0.6 | 2.4 | GO:0004966 | galanin receptor activity(GO:0004966) |
| 0.6 | 3.9 | GO:0099609 | microtubule lateral binding(GO:0099609) |
| 0.5 | 1.6 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.5 | 2.1 | GO:0004090 | carbonyl reductase (NADPH) activity(GO:0004090) |
| 0.5 | 3.5 | GO:0005000 | vasopressin receptor activity(GO:0005000) |
| 0.5 | 2.9 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.4 | 1.3 | GO:0004698 | calcium-dependent protein kinase C activity(GO:0004698) |
| 0.4 | 1.3 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.4 | 1.2 | GO:0009041 | uridylate kinase activity(GO:0009041) |
| 0.3 | 1.0 | GO:0003865 | 3-oxo-5-alpha-steroid 4-dehydrogenase activity(GO:0003865) cholestenone 5-alpha-reductase activity(GO:0047751) |
| 0.3 | 1.0 | GO:0015389 | pyrimidine- and adenine-specific:sodium symporter activity(GO:0015389) |
| 0.3 | 1.4 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.3 | 1.7 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 0.3 | 1.0 | GO:0090556 | apolipoprotein receptor activity(GO:0030226) apolipoprotein A-I receptor activity(GO:0034188) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.3 | 1.0 | GO:0071936 | coreceptor activity involved in Wnt signaling pathway(GO:0071936) |
| 0.3 | 1.0 | GO:0031826 | type 2A serotonin receptor binding(GO:0031826) |
| 0.3 | 4.1 | GO:0030881 | beta-2-microglobulin binding(GO:0030881) |
| 0.3 | 1.3 | GO:2001070 | starch binding(GO:2001070) |
| 0.3 | 1.2 | GO:0004308 | exo-alpha-sialidase activity(GO:0004308) alpha-sialidase activity(GO:0016997) exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.3 | 4.7 | GO:0022840 | leak channel activity(GO:0022840) narrow pore channel activity(GO:0022842) |
| 0.3 | 3.0 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.3 | 5.7 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.3 | 0.8 | GO:0038100 | nodal binding(GO:0038100) |
| 0.3 | 1.0 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.3 | 1.0 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.3 | 2.0 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.3 | 1.5 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.3 | 0.8 | GO:0004903 | growth hormone receptor activity(GO:0004903) |
| 0.3 | 0.8 | GO:0005280 | hydrogen:amino acid symporter activity(GO:0005280) L-tyrosine transmembrane transporter activity(GO:0005302) |
| 0.2 | 2.0 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.2 | 1.5 | GO:0005030 | neurotrophin receptor activity(GO:0005030) |
| 0.2 | 1.2 | GO:0098639 | collagen binding involved in cell-matrix adhesion(GO:0098639) |
| 0.2 | 1.4 | GO:0097643 | amylin receptor activity(GO:0097643) |
| 0.2 | 2.8 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.2 | 1.4 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 0.2 | 1.2 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.2 | 1.6 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.2 | 0.7 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.2 | 0.7 | GO:0048030 | disaccharide binding(GO:0048030) |
| 0.2 | 3.0 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) acetylcholine binding(GO:0042166) |
| 0.2 | 1.0 | GO:0004565 | beta-galactosidase activity(GO:0004565) |
| 0.2 | 0.6 | GO:0016964 | alpha-2 macroglobulin receptor activity(GO:0016964) |
| 0.2 | 1.1 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.2 | 2.4 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.2 | 1.5 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.2 | 2.7 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.2 | 1.6 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.2 | 2.2 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.2 | 1.2 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.2 | 0.7 | GO:0005093 | Rab GDP-dissociation inhibitor activity(GO:0005093) |
| 0.2 | 4.8 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.2 | 3.8 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.2 | 0.6 | GO:0070976 | TIR domain binding(GO:0070976) |
| 0.2 | 2.1 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.2 | 1.2 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.2 | 2.0 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.1 | 0.4 | GO:0030297 | transmembrane receptor protein tyrosine kinase activator activity(GO:0030297) |
| 0.1 | 0.9 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 0.1 | 1.3 | GO:0008499 | UDP-galactose:beta-N-acetylglucosamine beta-1,3-galactosyltransferase activity(GO:0008499) |
| 0.1 | 0.7 | GO:0032810 | sterol response element binding(GO:0032810) |
| 0.1 | 1.5 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.1 | 0.8 | GO:0045340 | mercury ion binding(GO:0045340) |
| 0.1 | 0.4 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.1 | 0.5 | GO:0015226 | amino-acid betaine transmembrane transporter activity(GO:0015199) carnitine transmembrane transporter activity(GO:0015226) |
| 0.1 | 0.5 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.1 | 0.6 | GO:0046980 | peptide antigen-transporting ATPase activity(GO:0015433) tapasin binding(GO:0046980) |
| 0.1 | 0.6 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.1 | 0.5 | GO:0016708 | nitric oxide dioxygenase activity(GO:0008941) oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of two atoms of oxygen into one donor(GO:0016708) iron-cytochrome-c reductase activity(GO:0047726) |
| 0.1 | 0.8 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.1 | 1.9 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.1 | 0.7 | GO:0051430 | corticotropin-releasing hormone receptor 1 binding(GO:0051430) |
| 0.1 | 0.9 | GO:0005168 | neurotrophin TRKA receptor binding(GO:0005168) |
| 0.1 | 1.3 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.1 | 0.8 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 0.3 | GO:0004489 | methylenetetrahydrofolate reductase (NAD(P)H) activity(GO:0004489) |
| 0.1 | 1.0 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.1 | 0.4 | GO:0016230 | sphingomyelin phosphodiesterase activator activity(GO:0016230) |
| 0.1 | 1.2 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.1 | 3.2 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.1 | 0.4 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 0.1 | 0.4 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.1 | 1.2 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.1 | 2.0 | GO:0000030 | mannosyltransferase activity(GO:0000030) |
| 0.1 | 0.9 | GO:0019865 | immunoglobulin binding(GO:0019865) |
| 0.1 | 0.7 | GO:0009374 | biotin binding(GO:0009374) |
| 0.1 | 0.5 | GO:0042285 | UDP-xylosyltransferase activity(GO:0035252) xylosyltransferase activity(GO:0042285) |
| 0.1 | 1.7 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.1 | 1.1 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.1 | 2.0 | GO:0016755 | transferase activity, transferring amino-acyl groups(GO:0016755) |
| 0.1 | 0.6 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.1 | 2.6 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.1 | 2.4 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.1 | 3.4 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.1 | 0.2 | GO:0004886 | 9-cis retinoic acid receptor activity(GO:0004886) |
| 0.1 | 0.3 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.1 | 1.1 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.1 | 0.4 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.1 | 0.8 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.1 | 0.5 | GO:1990380 | Lys63-specific deubiquitinase activity(GO:0061578) Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.1 | 0.4 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.1 | 1.1 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.1 | 1.9 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 0.1 | 0.3 | GO:0034235 | GPI anchor binding(GO:0034235) |
| 0.1 | 0.4 | GO:0015099 | nickel cation transmembrane transporter activity(GO:0015099) |
| 0.1 | 1.8 | GO:0005112 | Notch binding(GO:0005112) |
| 0.1 | 0.7 | GO:0015250 | water channel activity(GO:0015250) |
| 0.1 | 0.3 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.1 | 0.8 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.1 | 1.3 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 0.3 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.1 | 1.0 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.1 | 0.8 | GO:0051920 | peroxiredoxin activity(GO:0051920) |
| 0.0 | 0.3 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.0 | 1.1 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.0 | 0.6 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.0 | 0.5 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.0 | 1.9 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.0 | 0.5 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.0 | 0.2 | GO:0043533 | inositol 1,3,4,5 tetrakisphosphate binding(GO:0043533) |
| 0.0 | 1.2 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.0 | 0.3 | GO:0004645 | phosphorylase activity(GO:0004645) |
| 0.0 | 1.4 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 0.2 | GO:0019976 | interleukin-2 binding(GO:0019976) |
| 0.0 | 0.3 | GO:0070699 | type II activin receptor binding(GO:0070699) |
| 0.0 | 1.6 | GO:0016504 | peptidase activator activity(GO:0016504) |
| 0.0 | 0.7 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.9 | GO:0004697 | protein kinase C activity(GO:0004697) |
| 0.0 | 2.0 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 0.2 | GO:0004985 | opioid receptor activity(GO:0004985) |
| 0.0 | 0.4 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.0 | 0.5 | GO:0016274 | arginine N-methyltransferase activity(GO:0016273) protein-arginine N-methyltransferase activity(GO:0016274) |
| 0.0 | 0.3 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.0 | 4.3 | GO:0019905 | syntaxin binding(GO:0019905) |
| 0.0 | 0.4 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.0 | 1.0 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.0 | 1.3 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.0 | 0.7 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 3.6 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.2 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.0 | 0.2 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 0.0 | 0.4 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.6 | GO:0031418 | L-ascorbic acid binding(GO:0031418) |
| 0.0 | 1.1 | GO:0008373 | sialyltransferase activity(GO:0008373) |
| 0.0 | 0.6 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 0.6 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.0 | 1.0 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.3 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.0 | 0.5 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 0.5 | GO:0004445 | inositol-polyphosphate 5-phosphatase activity(GO:0004445) inositol trisphosphate phosphatase activity(GO:0046030) |
| 0.0 | 0.0 | GO:0043183 | vascular endothelial growth factor receptor 1 binding(GO:0043183) |
| 0.0 | 0.4 | GO:0015037 | peptide disulfide oxidoreductase activity(GO:0015037) |
| 0.0 | 1.0 | GO:0017046 | peptide hormone binding(GO:0017046) |
| 0.0 | 0.2 | GO:0050693 | LBD domain binding(GO:0050693) |
| 0.0 | 0.7 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.0 | 0.5 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.0 | 0.3 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.0 | 1.3 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.0 | 0.4 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.0 | 2.5 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.4 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.0 | 1.1 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.0 | 0.5 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 1.6 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.0 | 0.5 | GO:0016628 | oxidoreductase activity, acting on the CH-CH group of donors, NAD or NADP as acceptor(GO:0016628) |
| 0.0 | 0.2 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.0 | 0.1 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.0 | 0.2 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.0 | 0.1 | GO:0005219 | ryanodine-sensitive calcium-release channel activity(GO:0005219) |
| 0.0 | 0.5 | GO:0042171 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) lysophosphatidic acid acyltransferase activity(GO:0042171) |
| 0.0 | 0.8 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.0 | 0.2 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.0 | 0.3 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.0 | 0.1 | GO:0016454 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.0 | 0.8 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.0 | 0.1 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.0 | 0.3 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.0 | 0.1 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.0 | 0.3 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.0 | 0.1 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.0 | 0.5 | GO:0005248 | voltage-gated sodium channel activity(GO:0005248) |
| 0.0 | 0.1 | GO:0001162 | RNA polymerase II intronic transcription regulatory region sequence-specific DNA binding(GO:0001162) |
| 0.0 | 0.6 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 0.3 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 0.0 | 4.1 | GO:0003713 | transcription coactivator activity(GO:0003713) |
| 0.0 | 0.4 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.0 | 2.2 | GO:0005088 | Ras guanyl-nucleotide exchange factor activity(GO:0005088) |
| 0.0 | 1.9 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.0 | 0.2 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 0.1 | GO:0019828 | aspartic-type endopeptidase inhibitor activity(GO:0019828) |
| 0.0 | 0.2 | GO:0043236 | laminin binding(GO:0043236) |
| 0.0 | 0.5 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 0.4 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.0 | 1.0 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 0.2 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.0 | 0.2 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.2 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 5.9 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.2 | 3.7 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.1 | 1.5 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.1 | 7.0 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.1 | 3.6 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.1 | 2.1 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.1 | 0.7 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.1 | 1.1 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 1.3 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.1 | 2.2 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.1 | 0.6 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.1 | 1.1 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.1 | 1.5 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.1 | 3.0 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.1 | 1.3 | PID IGF1 PATHWAY | IGF1 pathway |
| 0.1 | 0.9 | PID ERBB1 RECEPTOR PROXIMAL PATHWAY | EGF receptor (ErbB1) signaling pathway |
| 0.1 | 1.3 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.1 | 1.2 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.0 | 0.5 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.0 | 3.9 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 1.4 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.0 | 2.1 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 1.9 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 1.0 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 1.2 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 0.9 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 0.4 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.0 | 1.8 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 1.6 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 1.0 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 2.2 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 0.7 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 0.4 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.0 | 0.4 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.0 | 0.5 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.0 | 0.7 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 0.3 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.0 | 0.5 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 0.6 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 1.0 | PID MET PATHWAY | Signaling events mediated by Hepatocyte Growth Factor Receptor (c-Met) |
| 0.0 | 0.3 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.0 | 4.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.4 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.4 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.0 | 0.5 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.0 | 0.8 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 0.2 | PID IFNG PATHWAY | IFN-gamma pathway |
| 0.0 | 0.7 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.4 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.0 | 0.3 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.0 | 0.2 | ST INTERLEUKIN 4 PATHWAY | Interleukin 4 (IL-4) Pathway |
| 0.0 | 0.3 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 1.1 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 0.4 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 0.4 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 0.2 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 2.0 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.4 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 0.2 | PID IL27 PATHWAY | IL27-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 3.6 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.3 | 2.6 | REACTOME ENDOGENOUS STEROLS | Genes involved in Endogenous sterols |
| 0.3 | 5.4 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.3 | 3.0 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.2 | 0.4 | REACTOME REGULATED PROTEOLYSIS OF P75NTR | Genes involved in Regulated proteolysis of p75NTR |
| 0.2 | 2.3 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.2 | 1.0 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.2 | 5.0 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.1 | 2.7 | REACTOME GRB2 SOS PROVIDES LINKAGE TO MAPK SIGNALING FOR INTERGRINS | Genes involved in GRB2:SOS provides linkage to MAPK signaling for Intergrins |
| 0.1 | 1.5 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.1 | 1.1 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.1 | 1.1 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.1 | 1.5 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.1 | 1.4 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.1 | 1.5 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.1 | 4.7 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 1.0 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.1 | 3.1 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.1 | 2.9 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.1 | 1.1 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.1 | 1.1 | REACTOME N GLYCAN TRIMMING IN THE ER AND CALNEXIN CALRETICULIN CYCLE | Genes involved in N-glycan trimming in the ER and Calnexin/Calreticulin cycle |
| 0.1 | 3.3 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.1 | 1.3 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.1 | 2.3 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.1 | 1.0 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.1 | 11.3 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.1 | 1.7 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.1 | 1.5 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.1 | 0.8 | REACTOME SOS MEDIATED SIGNALLING | Genes involved in SOS-mediated signalling |
| 0.1 | 1.0 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.1 | 2.0 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.1 | 3.7 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.1 | 1.0 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.1 | 0.6 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.1 | 1.3 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 4.7 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.1 | 0.5 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.1 | 2.0 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.1 | 1.1 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.1 | 2.1 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 0.8 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.1 | 4.1 | REACTOME UNFOLDED PROTEIN RESPONSE | Genes involved in Unfolded Protein Response |
| 0.0 | 1.1 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 1.4 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 0.4 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 1.3 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 3.9 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 1.0 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 0.6 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.0 | 1.8 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 1.6 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.0 | 0.3 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.9 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.9 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.8 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.0 | 0.1 | REACTOME SHC MEDIATED SIGNALLING | Genes involved in SHC-mediated signalling |
| 0.0 | 0.5 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 0.3 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.0 | 0.2 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 0.9 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.0 | 1.6 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 1.1 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.4 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 0.9 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 3.1 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 0.2 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.0 | 0.4 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 0.7 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.0 | 0.1 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.1 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.0 | 0.5 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.5 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.7 | REACTOME P75 NTR RECEPTOR MEDIATED SIGNALLING | Genes involved in p75 NTR receptor-mediated signalling |
| 0.0 | 4.1 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.1 | REACTOME PLATELET SENSITIZATION BY LDL | Genes involved in Platelet sensitization by LDL |
| 0.0 | 0.8 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.0 | 0.2 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 0.0 | 0.1 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 0.4 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.0 | 0.2 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 0.2 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.0 | 0.4 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.0 | 0.5 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.3 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 0.1 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |