Project
avrg: 12D miR HR13_24
Navigation
Downloads
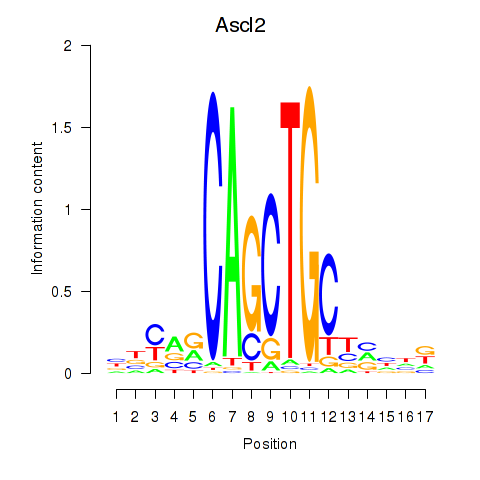
Results for Ascl2
Z-value: 3.10
Transcription factors associated with Ascl2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Ascl2
|
ENSMUSG00000009248.5 | achaete-scute family bHLH transcription factor 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Ascl2 | mm10_v2_chr7_-_142969238_142969264 | -0.53 | 9.0e-02 | Click! |
Activity profile of Ascl2 motif
Sorted Z-values of Ascl2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_-_128206346 | 8.52 |
ENSMUST00000033049.7
|
Cox6a2
|
cytochrome c oxidase subunit VIa polypeptide 2 |
| chr15_-_71727815 | 4.09 |
ENSMUST00000022953.8
|
Fam135b
|
family with sequence similarity 135, member B |
| chr12_-_17176888 | 3.97 |
ENSMUST00000170580.1
|
Kcnf1
|
potassium voltage-gated channel, subfamily F, member 1 |
| chr11_-_120648104 | 3.75 |
ENSMUST00000026134.2
|
Myadml2
|
myeloid-associated differentiation marker-like 2 |
| chr11_+_69965396 | 3.62 |
ENSMUST00000018713.6
|
Cldn7
|
claudin 7 |
| chr6_+_107529717 | 3.55 |
ENSMUST00000049285.8
|
Lrrn1
|
leucine rich repeat protein 1, neuronal |
| chr4_-_138367966 | 3.47 |
ENSMUST00000030535.3
|
Cda
|
cytidine deaminase |
| chr4_-_141846359 | 3.22 |
ENSMUST00000037059.10
|
Ctrc
|
chymotrypsin C (caldecrin) |
| chr7_-_143074037 | 3.22 |
ENSMUST00000136602.1
|
Trpm5
|
transient receptor potential cation channel, subfamily M, member 5 |
| chr10_+_127866457 | 3.17 |
ENSMUST00000092058.3
|
BC089597
|
cDNA sequence BC089597 |
| chr4_-_141846277 | 3.02 |
ENSMUST00000105781.1
|
Ctrc
|
chymotrypsin C (caldecrin) |
| chr9_-_62537036 | 2.83 |
ENSMUST00000048043.5
|
Coro2b
|
coronin, actin binding protein, 2B |
| chr10_+_69212634 | 2.80 |
ENSMUST00000020101.5
|
Rhobtb1
|
Rho-related BTB domain containing 1 |
| chr4_-_8239034 | 2.72 |
ENSMUST00000066674.7
|
Car8
|
carbonic anhydrase 8 |
| chr2_-_25196759 | 2.63 |
ENSMUST00000081869.6
|
Tor4a
|
torsin family 4, member A |
| chr5_+_91139591 | 2.51 |
ENSMUST00000031325.4
|
Areg
|
amphiregulin |
| chr9_-_45204083 | 2.48 |
ENSMUST00000034599.8
|
Tmprss4
|
transmembrane protease, serine 4 |
| chr9_-_62510498 | 2.34 |
ENSMUST00000164246.2
|
Coro2b
|
coronin, actin binding protein, 2B |
| chrX_+_143518671 | 2.29 |
ENSMUST00000134402.1
|
Pak3
|
p21 protein (Cdc42/Rac)-activated kinase 3 |
| chr16_-_22439570 | 2.28 |
ENSMUST00000170393.1
|
Etv5
|
ets variant gene 5 |
| chr4_-_130275542 | 2.27 |
ENSMUST00000154846.1
ENSMUST00000105996.1 |
Serinc2
|
serine incorporator 2 |
| chr15_-_101850778 | 2.25 |
ENSMUST00000023790.3
|
Krt1
|
keratin 1 |
| chr7_-_66427469 | 2.22 |
ENSMUST00000015278.7
|
Aldh1a3
|
aldehyde dehydrogenase family 1, subfamily A3 |
| chr4_-_130275213 | 2.18 |
ENSMUST00000122374.1
|
Serinc2
|
serine incorporator 2 |
| chr10_+_34483400 | 2.15 |
ENSMUST00000019913.7
ENSMUST00000170771.1 |
Frk
|
fyn-related kinase |
| chr16_-_32797413 | 2.14 |
ENSMUST00000115116.1
ENSMUST00000041123.8 |
Muc20
|
mucin 20 |
| chr1_-_156674290 | 2.10 |
ENSMUST00000079625.4
|
Tor3a
|
torsin family 3, member A |
| chr11_-_119086221 | 2.06 |
ENSMUST00000026665.7
|
Cbx4
|
chromobox 4 |
| chr12_+_26469204 | 2.04 |
ENSMUST00000020969.3
|
Cmpk2
|
cytidine monophosphate (UMP-CMP) kinase 2, mitochondrial |
| chrX_+_143518576 | 1.98 |
ENSMUST00000033640.7
|
Pak3
|
p21 protein (Cdc42/Rac)-activated kinase 3 |
| chr4_+_58943575 | 1.97 |
ENSMUST00000107554.1
|
Zkscan16
|
zinc finger with KRAB and SCAN domains 16 |
| chr8_-_41374602 | 1.93 |
ENSMUST00000110417.1
ENSMUST00000034000.8 ENSMUST00000143057.1 |
Asah1
|
N-acylsphingosine amidohydrolase 1 |
| chr17_+_5492558 | 1.82 |
ENSMUST00000089185.4
|
Zdhhc14
|
zinc finger, DHHC domain containing 14 |
| chr12_-_113422730 | 1.81 |
ENSMUST00000177715.1
ENSMUST00000103426.1 |
Ighm
|
immunoglobulin heavy constant mu |
| chr5_-_131538687 | 1.79 |
ENSMUST00000161374.1
|
Auts2
|
autism susceptibility candidate 2 |
| chr13_-_103096818 | 1.78 |
ENSMUST00000166336.1
|
Mast4
|
microtubule associated serine/threonine kinase family member 4 |
| chr2_+_34772089 | 1.78 |
ENSMUST00000028222.6
ENSMUST00000100171.2 |
Hspa5
|
heat shock protein 5 |
| chr18_-_82406777 | 1.76 |
ENSMUST00000065224.6
|
Galr1
|
galanin receptor 1 |
| chr5_-_69341699 | 1.75 |
ENSMUST00000054095.4
|
Kctd8
|
potassium channel tetramerisation domain containing 8 |
| chr5_+_102845007 | 1.75 |
ENSMUST00000070000.4
|
Arhgap24
|
Rho GTPase activating protein 24 |
| chr7_-_127993831 | 1.73 |
ENSMUST00000033056.3
|
Pycard
|
PYD and CARD domain containing |
| chr1_-_84696182 | 1.73 |
ENSMUST00000049126.6
|
Dner
|
delta/notch-like EGF-related receptor |
| chr18_-_61536522 | 1.72 |
ENSMUST00000171629.1
|
Arhgef37
|
Rho guanine nucleotide exchange factor (GEF) 37 |
| chr19_-_5349574 | 1.72 |
ENSMUST00000025764.5
|
Cst6
|
cystatin E/M |
| chr1_-_14918862 | 1.69 |
ENSMUST00000041447.4
|
Trpa1
|
transient receptor potential cation channel, subfamily A, member 1 |
| chr3_+_106486009 | 1.67 |
ENSMUST00000183271.1
ENSMUST00000061206.3 |
Dennd2d
|
DENN/MADD domain containing 2D |
| chr2_+_174760619 | 1.66 |
ENSMUST00000029030.2
|
Edn3
|
endothelin 3 |
| chr11_+_96929260 | 1.64 |
ENSMUST00000054311.5
ENSMUST00000107636.3 |
Prr15l
|
proline rich 15-like |
| chr10_-_128401218 | 1.62 |
ENSMUST00000042666.5
|
Slc39a5
|
solute carrier family 39 (metal ion transporter), member 5 |
| chr1_+_75549581 | 1.59 |
ENSMUST00000154101.1
|
Slc4a3
|
solute carrier family 4 (anion exchanger), member 3 |
| chr5_+_35757875 | 1.58 |
ENSMUST00000101280.3
ENSMUST00000054598.5 ENSMUST00000114205.1 ENSMUST00000114206.2 |
Ablim2
|
actin-binding LIM protein 2 |
| chr5_+_24428208 | 1.58 |
ENSMUST00000115049.2
|
Slc4a2
|
solute carrier family 4 (anion exchanger), member 2 |
| chr11_+_96929367 | 1.56 |
ENSMUST00000062172.5
|
Prr15l
|
proline rich 15-like |
| chr18_-_44662251 | 1.54 |
ENSMUST00000164666.1
|
Mcc
|
mutated in colorectal cancers |
| chr7_+_44207307 | 1.54 |
ENSMUST00000077354.4
|
Klk1b4
|
kallikrein 1-related pepidase b4 |
| chr15_-_32244632 | 1.53 |
ENSMUST00000181536.1
|
0610007N19Rik
|
RIKEN cDNA 0610007N19 |
| chr1_-_130715734 | 1.52 |
ENSMUST00000066863.6
ENSMUST00000050406.4 |
Pfkfb2
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2 |
| chr15_-_58364148 | 1.52 |
ENSMUST00000068515.7
|
Anxa13
|
annexin A13 |
| chr13_-_119408985 | 1.50 |
ENSMUST00000099149.3
ENSMUST00000069902.6 ENSMUST00000109204.1 |
Nnt
|
nicotinamide nucleotide transhydrogenase |
| chr4_-_148149684 | 1.49 |
ENSMUST00000126615.1
|
Fbxo6
|
F-box protein 6 |
| chr15_+_59315030 | 1.48 |
ENSMUST00000022977.7
|
Sqle
|
squalene epoxidase |
| chr10_-_61147659 | 1.48 |
ENSMUST00000092498.5
ENSMUST00000137833.1 ENSMUST00000155919.1 |
Sgpl1
|
sphingosine phosphate lyase 1 |
| chr11_+_53519920 | 1.48 |
ENSMUST00000147912.1
|
Sept8
|
septin 8 |
| chr1_+_120340569 | 1.47 |
ENSMUST00000037286.8
|
C1ql2
|
complement component 1, q subcomponent-like 2 |
| chr10_-_61147625 | 1.47 |
ENSMUST00000122259.1
|
Sgpl1
|
sphingosine phosphate lyase 1 |
| chr2_-_170406501 | 1.47 |
ENSMUST00000154650.1
|
Bcas1
|
breast carcinoma amplified sequence 1 |
| chr5_+_90759299 | 1.47 |
ENSMUST00000031318.4
|
Cxcl5
|
chemokine (C-X-C motif) ligand 5 |
| chr4_+_133553370 | 1.43 |
ENSMUST00000042706.2
|
Nr0b2
|
nuclear receptor subfamily 0, group B, member 2 |
| chr7_+_44225430 | 1.42 |
ENSMUST00000075162.3
|
Klk1
|
kallikrein 1 |
| chr11_+_53519871 | 1.40 |
ENSMUST00000120878.2
|
Sept8
|
septin 8 |
| chr15_+_59315088 | 1.39 |
ENSMUST00000100640.4
|
Sqle
|
squalene epoxidase |
| chr11_+_71749914 | 1.39 |
ENSMUST00000150531.1
|
Wscd1
|
WSC domain containing 1 |
| chr9_+_109096659 | 1.37 |
ENSMUST00000130366.1
|
Plxnb1
|
plexin B1 |
| chr7_-_131322292 | 1.37 |
ENSMUST00000046611.7
|
Cuzd1
|
CUB and zona pellucida-like domains 1 |
| chr15_-_98728120 | 1.34 |
ENSMUST00000003445.6
|
Fkbp11
|
FK506 binding protein 11 |
| chr13_-_52981027 | 1.32 |
ENSMUST00000071065.7
|
Nfil3
|
nuclear factor, interleukin 3, regulated |
| chr7_+_30977043 | 1.30 |
ENSMUST00000058093.4
|
Fam187b
|
family with sequence similarity 187, member B |
| chr17_-_87282793 | 1.29 |
ENSMUST00000146560.2
|
4833418N02Rik
|
RIKEN cDNA 4833418N02 gene |
| chr11_+_98664341 | 1.29 |
ENSMUST00000017348.2
|
Gsdma
|
gasdermin A |
| chr4_-_130275523 | 1.29 |
ENSMUST00000146478.1
|
Serinc2
|
serine incorporator 2 |
| chr6_+_125349699 | 1.29 |
ENSMUST00000032491.8
|
Tnfrsf1a
|
tumor necrosis factor receptor superfamily, member 1a |
| chr1_-_162866502 | 1.27 |
ENSMUST00000046049.7
|
Fmo1
|
flavin containing monooxygenase 1 |
| chr6_-_124733121 | 1.27 |
ENSMUST00000112484.3
|
Ptpn6
|
protein tyrosine phosphatase, non-receptor type 6 |
| chr19_-_6015152 | 1.26 |
ENSMUST00000025891.8
|
Capn1
|
calpain 1 |
| chr5_+_149411749 | 1.26 |
ENSMUST00000093110.5
|
Medag
|
mesenteric estrogen dependent adipogenesis |
| chr10_+_116986314 | 1.25 |
ENSMUST00000020378.4
|
Best3
|
bestrophin 3 |
| chr15_+_99591028 | 1.24 |
ENSMUST00000169082.1
|
Aqp5
|
aquaporin 5 |
| chr10_-_75560330 | 1.24 |
ENSMUST00000051129.9
|
Fam211b
|
family with sequence similarity 211, member B |
| chr11_+_115163333 | 1.24 |
ENSMUST00000021077.3
|
Slc9a3r1
|
solute carrier family 9 (sodium/hydrogen exchanger), member 3 regulator 1 |
| chr7_+_139213239 | 1.22 |
ENSMUST00000106104.1
|
Lrrc27
|
leucine rich repeat containing 27 |
| chr9_-_43239816 | 1.22 |
ENSMUST00000034512.5
|
Oaf
|
OAF homolog (Drosophila) |
| chr6_-_148444336 | 1.20 |
ENSMUST00000060095.8
ENSMUST00000100772.3 |
Tmtc1
|
transmembrane and tetratricopeptide repeat containing 1 |
| chr3_+_27182965 | 1.19 |
ENSMUST00000046515.8
|
Nceh1
|
neutral cholesterol ester hydrolase 1 |
| chr12_+_112678803 | 1.18 |
ENSMUST00000174780.1
ENSMUST00000169593.1 ENSMUST00000173942.1 |
Zbtb42
|
zinc finger and BTB domain containing 42 |
| chr7_+_44198191 | 1.17 |
ENSMUST00000085450.2
|
Klk1b3
|
kallikrein 1-related peptidase b3 |
| chr3_+_27182994 | 1.16 |
ENSMUST00000091284.4
|
Nceh1
|
neutral cholesterol ester hydrolase 1 |
| chr12_+_17690793 | 1.16 |
ENSMUST00000071858.3
|
Hpcal1
|
hippocalcin-like 1 |
| chr7_+_44188205 | 1.16 |
ENSMUST00000073713.6
|
Klk1b24
|
kallikrein 1-related peptidase b24 |
| chr14_+_33923582 | 1.15 |
ENSMUST00000168727.1
|
Gdf10
|
growth differentiation factor 10 |
| chr5_+_135168382 | 1.14 |
ENSMUST00000111187.3
ENSMUST00000111188.1 |
Bcl7b
|
B cell CLL/lymphoma 7B |
| chr8_+_62951195 | 1.14 |
ENSMUST00000118003.1
|
Spock3
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan 3 |
| chr13_-_62888282 | 1.13 |
ENSMUST00000092888.4
|
Fbp1
|
fructose bisphosphatase 1 |
| chr15_-_100599983 | 1.13 |
ENSMUST00000073837.6
|
Pou6f1
|
POU domain, class 6, transcription factor 1 |
| chr5_+_35757951 | 1.12 |
ENSMUST00000114204.1
ENSMUST00000129347.1 |
Ablim2
|
actin-binding LIM protein 2 |
| chr9_-_108567336 | 1.12 |
ENSMUST00000074208.4
|
Ndufaf3
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, assembly factor 3 |
| chrX_-_112698642 | 1.11 |
ENSMUST00000039887.3
|
Pof1b
|
premature ovarian failure 1B |
| chr1_-_88205674 | 1.08 |
ENSMUST00000119972.2
|
Dnajb3
|
DnaJ (Hsp40) homolog, subfamily B, member 3 |
| chr7_+_78913765 | 1.08 |
ENSMUST00000038142.8
|
Isg20
|
interferon-stimulated protein |
| chr4_-_62434722 | 1.07 |
ENSMUST00000107454.1
|
Rnf183
|
ring finger protein 183 |
| chr7_-_99626936 | 1.07 |
ENSMUST00000178124.1
|
Gm4980
|
predicted gene 4980 |
| chr1_-_167393826 | 1.07 |
ENSMUST00000028005.2
|
Mgst3
|
microsomal glutathione S-transferase 3 |
| chr17_-_21845759 | 1.06 |
ENSMUST00000084141.4
|
Zfp820
|
zinc finger protein 820 |
| chr17_-_57228003 | 1.06 |
ENSMUST00000177046.1
ENSMUST00000024988.8 |
C3
|
complement component 3 |
| chr6_+_56017489 | 1.06 |
ENSMUST00000052827.4
|
Ppp1r17
|
protein phosphatase 1, regulatory subunit 17 |
| chr8_+_3665747 | 1.06 |
ENSMUST00000014118.2
|
1810033B17Rik
|
RIKEN cDNA 1810033B17 gene |
| chr9_+_107975529 | 1.05 |
ENSMUST00000035216.4
|
Uba7
|
ubiquitin-like modifier activating enzyme 7 |
| chr7_+_80246375 | 1.05 |
ENSMUST00000058266.6
|
Ttll13
|
tubulin tyrosine ligase-like family, member 13 |
| chr2_+_174760781 | 1.04 |
ENSMUST00000140908.1
|
Edn3
|
endothelin 3 |
| chr8_-_111691002 | 1.04 |
ENSMUST00000034435.5
|
Ctrb1
|
chymotrypsinogen B1 |
| chr10_+_11343387 | 1.04 |
ENSMUST00000069106.4
|
Epm2a
|
epilepsy, progressive myoclonic epilepsy, type 2 gene alpha |
| chr1_+_74854954 | 1.04 |
ENSMUST00000160379.2
|
Cdk5r2
|
cyclin-dependent kinase 5, regulatory subunit 2 (p39) |
| chr15_+_102144362 | 1.04 |
ENSMUST00000023807.6
|
Igfbp6
|
insulin-like growth factor binding protein 6 |
| chr5_+_135168283 | 1.01 |
ENSMUST00000031692.5
|
Bcl7b
|
B cell CLL/lymphoma 7B |
| chr5_-_107726017 | 1.01 |
ENSMUST00000159263.2
|
Gfi1
|
growth factor independent 1 |
| chr4_-_133263042 | 1.01 |
ENSMUST00000105908.3
ENSMUST00000030674.7 |
Sytl1
|
synaptotagmin-like 1 |
| chr17_-_57247632 | 1.00 |
ENSMUST00000005975.6
|
Gpr108
|
G protein-coupled receptor 108 |
| chr2_-_26933781 | 0.99 |
ENSMUST00000154651.1
ENSMUST00000015011.3 |
Surf4
|
surfeit gene 4 |
| chr5_-_122502192 | 0.99 |
ENSMUST00000179939.1
ENSMUST00000177974.1 ENSMUST00000031423.8 |
Atp2a2
|
ATPase, Ca++ transporting, cardiac muscle, slow twitch 2 |
| chr11_-_59182810 | 0.98 |
ENSMUST00000108793.2
|
Gjc2
|
gap junction protein, gamma 2 |
| chr7_+_126950837 | 0.98 |
ENSMUST00000106332.1
|
Sez6l2
|
seizure related 6 homolog like 2 |
| chr7_+_122289297 | 0.98 |
ENSMUST00000064989.5
ENSMUST00000064921.4 |
Prkcb
|
protein kinase C, beta |
| chr7_-_67803489 | 0.97 |
ENSMUST00000181235.1
|
4833412C05Rik
|
RIKEN cDNA 4833412C05 gene |
| chr4_-_154160632 | 0.96 |
ENSMUST00000105639.3
ENSMUST00000030896.8 |
Tprgl
|
transformation related protein 63 regulated like |
| chr15_-_84855093 | 0.95 |
ENSMUST00000016768.5
|
Phf21b
|
PHD finger protein 21B |
| chr3_+_108284089 | 0.94 |
ENSMUST00000102632.4
|
Sort1
|
sortilin 1 |
| chr10_+_128270546 | 0.94 |
ENSMUST00000105238.3
ENSMUST00000085708.2 |
Stat2
|
signal transducer and activator of transcription 2 |
| chr19_-_56822161 | 0.94 |
ENSMUST00000118592.1
|
A630007B06Rik
|
RIKEN cDNA A630007B06 gene |
| chr12_+_73286868 | 0.93 |
ENSMUST00000153941.1
ENSMUST00000122920.1 ENSMUST00000101313.3 |
Slc38a6
|
solute carrier family 38, member 6 |
| chr3_-_129969989 | 0.93 |
ENSMUST00000146340.1
|
Ccdc109b
|
coiled-coil domain containing 109B |
| chr9_-_53975246 | 0.93 |
ENSMUST00000048409.7
|
Elmod1
|
ELMO/CED-12 domain containing 1 |
| chr6_-_29179584 | 0.93 |
ENSMUST00000159200.1
|
Prrt4
|
proline-rich transmembrane protein 4 |
| chr7_+_3290553 | 0.91 |
ENSMUST00000096744.5
|
Myadm
|
myeloid-associated differentiation marker |
| chr1_+_162639148 | 0.91 |
ENSMUST00000028020.9
|
Myoc
|
myocilin |
| chr1_+_160044564 | 0.90 |
ENSMUST00000168359.1
|
4930523C07Rik
|
RIKEN cDNA 4930523C07 gene |
| chr11_-_5837760 | 0.90 |
ENSMUST00000109837.1
|
Polm
|
polymerase (DNA directed), mu |
| chr10_-_128673896 | 0.89 |
ENSMUST00000054764.7
|
Suox
|
sulfite oxidase |
| chr2_+_121449362 | 0.89 |
ENSMUST00000110615.1
ENSMUST00000099475.5 |
Serf2
|
small EDRK-rich factor 2 |
| chr4_-_147642496 | 0.88 |
ENSMUST00000133006.1
ENSMUST00000037565.7 ENSMUST00000105720.1 |
2610305D13Rik
|
RIKEN cDNA 2610305D13 gene |
| chr19_-_41385070 | 0.88 |
ENSMUST00000059672.7
|
Pik3ap1
|
phosphoinositide-3-kinase adaptor protein 1 |
| chr3_-_129970152 | 0.88 |
ENSMUST00000029624.8
|
Ccdc109b
|
coiled-coil domain containing 109B |
| chr1_+_127868773 | 0.88 |
ENSMUST00000037649.5
|
Rab3gap1
|
RAB3 GTPase activating protein subunit 1 |
| chr1_+_180935022 | 0.87 |
ENSMUST00000037361.8
|
Lefty1
|
left right determination factor 1 |
| chr7_+_139212974 | 0.87 |
ENSMUST00000016124.8
|
Lrrc27
|
leucine rich repeat containing 27 |
| chr2_-_168741752 | 0.87 |
ENSMUST00000029060.4
|
Atp9a
|
ATPase, class II, type 9A |
| chr2_+_164403194 | 0.86 |
ENSMUST00000017151.1
|
Rbpjl
|
recombination signal binding protein for immunoglobulin kappa J region-like |
| chr4_-_147809788 | 0.86 |
ENSMUST00000105734.3
ENSMUST00000176201.1 |
Gm13157
Gm20707
|
predicted gene 13157 predicted gene 20707 |
| chr4_-_147702553 | 0.85 |
ENSMUST00000117638.1
|
Zfp534
|
zinc finger protein 534 |
| chrX_+_36328353 | 0.85 |
ENSMUST00000016383.3
|
Lonrf3
|
LON peptidase N-terminal domain and ring finger 3 |
| chr9_-_57836706 | 0.85 |
ENSMUST00000164010.1
ENSMUST00000171444.1 ENSMUST00000098686.3 |
Arid3b
|
AT rich interactive domain 3B (BRIGHT-like) |
| chr11_-_114795888 | 0.85 |
ENSMUST00000000206.3
|
Btbd17
|
BTB (POZ) domain containing 17 |
| chr3_+_132085281 | 0.84 |
ENSMUST00000029665.5
|
Dkk2
|
dickkopf homolog 2 (Xenopus laevis) |
| chr1_-_130462680 | 0.84 |
ENSMUST00000027650.6
|
Cd55
|
CD55 antigen |
| chr2_+_125136692 | 0.82 |
ENSMUST00000099452.2
|
Ctxn2
|
cortexin 2 |
| chr4_-_124850670 | 0.82 |
ENSMUST00000163946.1
ENSMUST00000106190.3 |
1110065P20Rik
|
RIKEN cDNA 1110065P20 gene |
| chr2_-_27072175 | 0.82 |
ENSMUST00000009358.2
|
Tmem8c
|
transmembrane protein 8C |
| chr7_+_142471838 | 0.82 |
ENSMUST00000038946.2
|
Lsp1
|
lymphocyte specific 1 |
| chr8_-_91133942 | 0.81 |
ENSMUST00000120213.1
ENSMUST00000109609.2 |
Aktip
|
thymoma viral proto-oncogene 1 interacting protein |
| chr7_+_139213003 | 0.81 |
ENSMUST00000156768.1
|
Lrrc27
|
leucine rich repeat containing 27 |
| chr5_+_120861421 | 0.80 |
ENSMUST00000072476.6
ENSMUST00000171820.1 |
Oas1h
|
2'-5' oligoadenylate synthetase 1H |
| chr11_+_121434913 | 0.79 |
ENSMUST00000026175.2
ENSMUST00000092302.4 ENSMUST00000103014.3 |
Fn3k
|
fructosamine 3 kinase |
| chr3_-_75270073 | 0.79 |
ENSMUST00000039047.4
|
Serpini2
|
serine (or cysteine) peptidase inhibitor, clade I, member 2 |
| chr6_-_52191695 | 0.79 |
ENSMUST00000101395.2
|
Hoxa4
|
homeobox A4 |
| chr6_-_56362356 | 0.79 |
ENSMUST00000044505.7
ENSMUST00000166102.1 ENSMUST00000164037.1 ENSMUST00000114327.2 |
Pde1c
|
phosphodiesterase 1C |
| chr16_+_17276662 | 0.79 |
ENSMUST00000069420.4
|
Tmem191c
|
transmembrane protein 191C |
| chr9_+_72985568 | 0.79 |
ENSMUST00000150826.2
ENSMUST00000085350.4 ENSMUST00000140675.1 |
Ccpg1
|
cell cycle progression 1 |
| chr1_+_194619815 | 0.78 |
ENSMUST00000027952.5
|
Plxna2
|
plexin A2 |
| chr6_-_72617000 | 0.78 |
ENSMUST00000070524.4
|
Tgoln1
|
trans-golgi network protein |
| chr4_-_156050465 | 0.78 |
ENSMUST00000184684.1
|
Ttll10
|
tubulin tyrosine ligase-like family, member 10 |
| chr4_-_124850652 | 0.78 |
ENSMUST00000125776.1
|
1110065P20Rik
|
RIKEN cDNA 1110065P20 gene |
| chr7_+_142472080 | 0.78 |
ENSMUST00000105966.1
|
Lsp1
|
lymphocyte specific 1 |
| chr8_+_92901387 | 0.78 |
ENSMUST00000104947.2
|
Capns2
|
calpain, small subunit 2 |
| chr2_-_77170592 | 0.77 |
ENSMUST00000164114.2
ENSMUST00000049544.7 |
Ccdc141
|
coiled-coil domain containing 141 |
| chr1_-_164458345 | 0.77 |
ENSMUST00000027863.7
|
Atp1b1
|
ATPase, Na+/K+ transporting, beta 1 polypeptide |
| chr10_-_120201558 | 0.77 |
ENSMUST00000020448.4
|
Irak3
|
interleukin-1 receptor-associated kinase 3 |
| chr6_+_105677768 | 0.77 |
ENSMUST00000089208.2
|
Cntn4
|
contactin 4 |
| chr17_-_22166190 | 0.77 |
ENSMUST00000080249.5
|
Zfp947
|
zinc finger protein 947 |
| chr4_-_156050785 | 0.77 |
ENSMUST00000051509.8
|
Ttll10
|
tubulin tyrosine ligase-like family, member 10 |
| chr7_+_121707189 | 0.76 |
ENSMUST00000065310.2
|
1700069B07Rik
|
RIKEN cDNA 1700069B07 gene |
| chr4_-_46536134 | 0.76 |
ENSMUST00000046897.6
|
Trim14
|
tripartite motif-containing 14 |
| chr16_-_74411292 | 0.76 |
ENSMUST00000117200.1
|
Robo2
|
roundabout homolog 2 (Drosophila) |
| chr5_+_114568016 | 0.76 |
ENSMUST00000043650.7
|
Fam222a
|
family with sequence similarity 222, member A |
| chr8_-_91134027 | 0.75 |
ENSMUST00000125257.1
|
Aktip
|
thymoma viral proto-oncogene 1 interacting protein |
| chr11_+_45980309 | 0.75 |
ENSMUST00000049038.3
|
Sox30
|
SRY-box containing gene 30 |
| chr6_+_105677745 | 0.75 |
ENSMUST00000113261.2
ENSMUST00000113264.2 |
Cntn4
|
contactin 4 |
| chr2_-_144011202 | 0.75 |
ENSMUST00000016072.5
ENSMUST00000037875.5 |
Rrbp1
|
ribosome binding protein 1 |
| chrX_+_95711641 | 0.75 |
ENSMUST00000150123.1
|
Zc3h12b
|
zinc finger CCCH-type containing 12B |
| chr13_-_62858364 | 0.74 |
ENSMUST00000021907.7
|
Fbp2
|
fructose bisphosphatase 2 |
| chr7_+_141461728 | 0.74 |
ENSMUST00000167491.1
ENSMUST00000165194.1 |
Efcab4a
|
EF-hand calcium binding domain 4A |
| chr15_-_75888754 | 0.74 |
ENSMUST00000184858.1
|
Mroh6
|
maestro heat-like repeat family member 6 |
| chr16_-_17576206 | 0.73 |
ENSMUST00000090165.4
ENSMUST00000164623.1 |
Slc7a4
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 4 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Ascl2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 3.5 | GO:0009972 | cytidine catabolic process(GO:0006216) cytidine deamination(GO:0009972) cytidine metabolic process(GO:0046087) |
| 0.8 | 5.7 | GO:1904222 | regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904217) positive regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904219) positive regulation of serine C-palmitoyltransferase activity(GO:1904222) |
| 0.7 | 2.2 | GO:0060166 | olfactory pit development(GO:0060166) |
| 0.7 | 2.0 | GO:0009196 | dUDP biosynthetic process(GO:0006227) pyrimidine nucleoside diphosphate metabolic process(GO:0009138) pyrimidine nucleoside diphosphate biosynthetic process(GO:0009139) pyrimidine deoxyribonucleoside diphosphate metabolic process(GO:0009196) pyrimidine deoxyribonucleoside diphosphate biosynthetic process(GO:0009197) dUDP metabolic process(GO:0046077) |
| 0.7 | 2.0 | GO:0050925 | negative regulation of negative chemotaxis(GO:0050925) |
| 0.7 | 8.5 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.6 | 2.5 | GO:0060598 | dichotomous subdivision of terminal units involved in mammary gland duct morphogenesis(GO:0060598) |
| 0.6 | 1.9 | GO:0046351 | disaccharide biosynthetic process(GO:0046351) |
| 0.6 | 1.8 | GO:0002343 | peripheral B cell selection(GO:0002343) B cell affinity maturation(GO:0002344) |
| 0.6 | 1.8 | GO:2000620 | positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 0.6 | 1.8 | GO:0021589 | hindbrain structural organization(GO:0021577) cerebellum structural organization(GO:0021589) |
| 0.6 | 1.8 | GO:0051464 | positive regulation of cortisol secretion(GO:0051464) |
| 0.6 | 1.7 | GO:0002588 | myeloid dendritic cell activation involved in immune response(GO:0002277) positive regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002582) positive regulation of antigen processing and presentation of peptide antigen via MHC class II(GO:0002588) |
| 0.5 | 1.6 | GO:0034224 | cellular response to zinc ion starvation(GO:0034224) |
| 0.5 | 1.5 | GO:0070948 | regulation of neutrophil mediated cytotoxicity(GO:0070948) regulation of neutrophil mediated killing of symbiont cell(GO:0070949) |
| 0.5 | 2.8 | GO:0050968 | detection of chemical stimulus involved in sensory perception of pain(GO:0050968) |
| 0.5 | 2.7 | GO:0003100 | regulation of systemic arterial blood pressure by endothelin(GO:0003100) |
| 0.4 | 1.2 | GO:0042939 | glutathione transport(GO:0034635) tripeptide transport(GO:0042939) |
| 0.4 | 2.3 | GO:0098728 | germ-line stem cell division(GO:0042078) male germ-line stem cell asymmetric division(GO:0048133) germline stem cell asymmetric division(GO:0098728) |
| 0.3 | 1.7 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.3 | 1.4 | GO:1900220 | semaphorin-plexin signaling pathway involved in bone trabecula morphogenesis(GO:1900220) |
| 0.3 | 0.9 | GO:0097051 | establishment of protein localization to endoplasmic reticulum membrane(GO:0097051) |
| 0.3 | 2.2 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.3 | 0.8 | GO:0030450 | regulation of complement activation, classical pathway(GO:0030450) |
| 0.3 | 1.4 | GO:0070103 | negative regulation of calcidiol 1-monooxygenase activity(GO:0010956) regulation of interleukin-6-mediated signaling pathway(GO:0070103) |
| 0.3 | 0.8 | GO:0021934 | hindbrain tangential cell migration(GO:0021934) |
| 0.3 | 1.0 | GO:0021586 | pons maturation(GO:0021586) |
| 0.3 | 0.8 | GO:1903279 | regulation of calcium:sodium antiporter activity(GO:1903279) |
| 0.3 | 1.5 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 0.2 | 0.7 | GO:0071314 | cellular response to cocaine(GO:0071314) |
| 0.2 | 0.9 | GO:0014734 | skeletal muscle hypertrophy(GO:0014734) |
| 0.2 | 1.1 | GO:0034421 | post-translational protein acetylation(GO:0034421) |
| 0.2 | 1.1 | GO:0021564 | vagus nerve development(GO:0021564) |
| 0.2 | 0.7 | GO:0006011 | UDP-glucose metabolic process(GO:0006011) |
| 0.2 | 1.1 | GO:0001970 | positive regulation of activation of membrane attack complex(GO:0001970) |
| 0.2 | 1.1 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.2 | 0.6 | GO:1903903 | positive regulation of keratinocyte apoptotic process(GO:1902174) regulation of establishment of T cell polarity(GO:1903903) |
| 0.2 | 0.6 | GO:0007439 | ectodermal digestive tract development(GO:0007439) embryonic ectodermal digestive tract development(GO:0048611) |
| 0.2 | 0.8 | GO:2001106 | regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.2 | 1.0 | GO:0035405 | histone-threonine phosphorylation(GO:0035405) |
| 0.2 | 1.8 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.2 | 0.8 | GO:0010936 | negative regulation of macrophage cytokine production(GO:0010936) |
| 0.2 | 3.6 | GO:0032463 | negative regulation of protein homooligomerization(GO:0032463) |
| 0.2 | 0.9 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.2 | 0.9 | GO:0015692 | vanadium ion transport(GO:0015676) lead ion transport(GO:0015692) |
| 0.2 | 0.9 | GO:0044849 | estrous cycle(GO:0044849) |
| 0.2 | 1.1 | GO:1903691 | positive regulation of wound healing, spreading of epidermal cells(GO:1903691) |
| 0.2 | 1.1 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 0.2 | 0.9 | GO:0007597 | blood coagulation, intrinsic pathway(GO:0007597) |
| 0.2 | 0.5 | GO:0009446 | putrescine biosynthetic process(GO:0009446) |
| 0.2 | 3.0 | GO:0033327 | Leydig cell differentiation(GO:0033327) |
| 0.2 | 0.2 | GO:0072098 | intermediate mesoderm development(GO:0048389) pattern specification involved in mesonephros development(GO:0061227) anterior/posterior pattern specification involved in kidney development(GO:0072098) |
| 0.2 | 0.8 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.2 | 0.5 | GO:0098917 | positive regulation of the force of heart contraction(GO:0098735) retrograde trans-synaptic signaling(GO:0098917) trans-synaptic signaling by soluble gas(GO:0099543) regulation of adrenergic receptor signaling pathway involved in heart process(GO:1901204) |
| 0.2 | 4.3 | GO:0010763 | positive regulation of fibroblast migration(GO:0010763) |
| 0.2 | 1.1 | GO:0098909 | regulation of cardiac muscle cell action potential involved in regulation of contraction(GO:0098909) |
| 0.2 | 0.5 | GO:0090004 | positive regulation of Golgi to plasma membrane protein transport(GO:0042998) positive regulation of establishment of protein localization to plasma membrane(GO:0090004) |
| 0.2 | 1.4 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.2 | 2.1 | GO:0048012 | hepatocyte growth factor receptor signaling pathway(GO:0048012) |
| 0.1 | 0.6 | GO:0021660 | rhombomere formation(GO:0021594) rhombomere 3 formation(GO:0021660) rhombomere 5 morphogenesis(GO:0021664) rhombomere 5 formation(GO:0021666) |
| 0.1 | 1.2 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 0.1 | 1.4 | GO:0000255 | allantoin metabolic process(GO:0000255) creatinine metabolic process(GO:0046449) |
| 0.1 | 1.1 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.1 | 0.4 | GO:0090157 | lysosomal microautophagy(GO:0016237) piecemeal microautophagy of nucleus(GO:0034727) suppression by virus of host autophagy(GO:0039521) negative regulation of sphingolipid biosynthesis involved in cellular sphingolipid homeostasis(GO:0090157) |
| 0.1 | 1.5 | GO:0033133 | positive regulation of glucokinase activity(GO:0033133) positive regulation of hexokinase activity(GO:1903301) |
| 0.1 | 0.1 | GO:0006713 | glucocorticoid catabolic process(GO:0006713) |
| 0.1 | 0.7 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.1 | 0.3 | GO:2000097 | regulation of lipoprotein oxidation(GO:0034442) negative regulation of lipoprotein oxidation(GO:0034443) regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 0.1 | 1.7 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.1 | 0.7 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.1 | 0.5 | GO:0009814 | defense response, incompatible interaction(GO:0009814) defense response to bacterium, incompatible interaction(GO:0009816) regulation of defense response to bacterium, incompatible interaction(GO:1902477) |
| 0.1 | 0.9 | GO:0034154 | toll-like receptor 7 signaling pathway(GO:0034154) |
| 0.1 | 0.5 | GO:0006868 | glutamine transport(GO:0006868) |
| 0.1 | 0.5 | GO:0001830 | trophectodermal cell fate commitment(GO:0001830) |
| 0.1 | 1.0 | GO:0046959 | habituation(GO:0046959) |
| 0.1 | 0.8 | GO:1900122 | positive regulation of receptor binding(GO:1900122) |
| 0.1 | 0.4 | GO:2000503 | positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 0.1 | 0.2 | GO:0061152 | trachea submucosa development(GO:0061152) trachea gland development(GO:0061153) |
| 0.1 | 1.3 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.1 | 0.2 | GO:0043686 | co-translational protein modification(GO:0043686) |
| 0.1 | 1.3 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.1 | 1.1 | GO:1903800 | positive regulation of intracellular estrogen receptor signaling pathway(GO:0033148) positive regulation of production of miRNAs involved in gene silencing by miRNA(GO:1903800) |
| 0.1 | 0.4 | GO:0060447 | peripheral nervous system axon regeneration(GO:0014012) bud outgrowth involved in lung branching(GO:0060447) |
| 0.1 | 0.4 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.1 | 0.3 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 0.1 | 0.7 | GO:0070493 | thrombin receptor signaling pathway(GO:0070493) |
| 0.1 | 0.3 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 0.1 | 0.3 | GO:0050915 | sensory perception of sour taste(GO:0050915) |
| 0.1 | 1.5 | GO:0006516 | glycoprotein catabolic process(GO:0006516) |
| 0.1 | 0.7 | GO:0010572 | positive regulation of platelet activation(GO:0010572) |
| 0.1 | 0.2 | GO:0008588 | release of cytoplasmic sequestered NF-kappaB(GO:0008588) |
| 0.1 | 0.6 | GO:0015871 | choline transport(GO:0015871) |
| 0.1 | 0.9 | GO:0016446 | somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 0.1 | 0.8 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.1 | 0.5 | GO:0035735 | intraciliary transport involved in cilium morphogenesis(GO:0035735) |
| 0.1 | 3.2 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.1 | 0.5 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.1 | 3.0 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.1 | 0.7 | GO:0014827 | intestine smooth muscle contraction(GO:0014827) |
| 0.1 | 0.7 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.1 | 0.6 | GO:0007196 | adenylate cyclase-inhibiting G-protein coupled glutamate receptor signaling pathway(GO:0007196) |
| 0.1 | 0.5 | GO:0040031 | snRNA modification(GO:0040031) |
| 0.1 | 0.5 | GO:0019336 | phenol-containing compound catabolic process(GO:0019336) |
| 0.1 | 0.1 | GO:1903972 | regulation of macrophage colony-stimulating factor signaling pathway(GO:1902226) regulation of response to macrophage colony-stimulating factor(GO:1903969) regulation of cellular response to macrophage colony-stimulating factor stimulus(GO:1903972) |
| 0.1 | 1.0 | GO:0010644 | cell communication by electrical coupling(GO:0010644) |
| 0.1 | 1.8 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.1 | 0.5 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.1 | 0.5 | GO:0007182 | common-partner SMAD protein phosphorylation(GO:0007182) |
| 0.1 | 0.2 | GO:0046341 | CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.1 | 0.5 | GO:1902004 | positive regulation of beta-amyloid formation(GO:1902004) |
| 0.1 | 0.6 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.1 | 0.6 | GO:0032119 | sequestering of zinc ion(GO:0032119) regulation of sequestering of zinc ion(GO:0061088) |
| 0.1 | 0.2 | GO:0098528 | skeletal muscle fiber differentiation(GO:0098528) |
| 0.1 | 0.2 | GO:0060083 | smooth muscle contraction involved in micturition(GO:0060083) |
| 0.1 | 0.2 | GO:0036118 | hyaluranon cable assembly(GO:0036118) regulation of hyaluranon cable assembly(GO:1900104) positive regulation of hyaluranon cable assembly(GO:1900106) |
| 0.1 | 0.7 | GO:0001574 | ganglioside biosynthetic process(GO:0001574) |
| 0.1 | 0.7 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.1 | 0.8 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.1 | 0.4 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.1 | 4.3 | GO:0060395 | SMAD protein signal transduction(GO:0060395) |
| 0.1 | 1.7 | GO:0006739 | NADP metabolic process(GO:0006739) |
| 0.1 | 0.8 | GO:0071941 | nitrogen cycle metabolic process(GO:0071941) |
| 0.1 | 1.1 | GO:0032897 | negative regulation of viral transcription(GO:0032897) |
| 0.1 | 0.6 | GO:0060770 | negative regulation of epithelial cell proliferation involved in prostate gland development(GO:0060770) |
| 0.1 | 0.1 | GO:0070296 | sarcoplasmic reticulum calcium ion transport(GO:0070296) |
| 0.1 | 0.7 | GO:1901550 | regulation of endothelial cell development(GO:1901550) regulation of establishment of endothelial barrier(GO:1903140) |
| 0.1 | 1.2 | GO:0006152 | purine nucleoside catabolic process(GO:0006152) purine ribonucleoside catabolic process(GO:0046130) |
| 0.0 | 0.2 | GO:0002036 | regulation of L-glutamate transport(GO:0002036) |
| 0.0 | 0.5 | GO:0045059 | positive thymic T cell selection(GO:0045059) |
| 0.0 | 0.3 | GO:1905146 | lysosomal protein catabolic process(GO:1905146) |
| 0.0 | 0.8 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.0 | 0.7 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.0 | 0.2 | GO:0010897 | negative regulation of triglyceride catabolic process(GO:0010897) |
| 0.0 | 1.0 | GO:0090036 | regulation of protein kinase C signaling(GO:0090036) |
| 0.0 | 0.6 | GO:0034312 | diol biosynthetic process(GO:0034312) |
| 0.0 | 0.1 | GO:0010760 | negative regulation of macrophage chemotaxis(GO:0010760) |
| 0.0 | 0.5 | GO:0042711 | maternal behavior(GO:0042711) |
| 0.0 | 3.8 | GO:0051453 | regulation of intracellular pH(GO:0051453) |
| 0.0 | 1.6 | GO:0045022 | early endosome to late endosome transport(GO:0045022) |
| 0.0 | 2.7 | GO:0016126 | sterol biosynthetic process(GO:0016126) |
| 0.0 | 0.4 | GO:1903715 | regulation of aerobic respiration(GO:1903715) |
| 0.0 | 0.7 | GO:0002115 | store-operated calcium entry(GO:0002115) |
| 0.0 | 0.4 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.0 | 0.1 | GO:0060139 | positive regulation by symbiont of host apoptotic process(GO:0052151) positive regulation of apoptotic process by virus(GO:0060139) |
| 0.0 | 0.3 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.0 | 0.7 | GO:0015991 | energy coupled proton transmembrane transport, against electrochemical gradient(GO:0015988) ATP hydrolysis coupled proton transport(GO:0015991) |
| 0.0 | 0.4 | GO:0030854 | positive regulation of granulocyte differentiation(GO:0030854) |
| 0.0 | 0.9 | GO:0070670 | response to interleukin-4(GO:0070670) cellular response to interleukin-4(GO:0071353) |
| 0.0 | 0.9 | GO:0060337 | type I interferon signaling pathway(GO:0060337) cellular response to type I interferon(GO:0071357) |
| 0.0 | 1.4 | GO:0051642 | centrosome localization(GO:0051642) |
| 0.0 | 0.6 | GO:0042492 | gamma-delta T cell differentiation(GO:0042492) |
| 0.0 | 0.2 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.0 | 0.7 | GO:0051573 | negative regulation of histone H3-K9 methylation(GO:0051573) |
| 0.0 | 0.5 | GO:0010457 | centriole-centriole cohesion(GO:0010457) |
| 0.0 | 0.1 | GO:0009181 | purine nucleoside diphosphate catabolic process(GO:0009137) purine ribonucleoside diphosphate catabolic process(GO:0009181) |
| 0.0 | 0.3 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.0 | 0.3 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.0 | 0.1 | GO:0006121 | mitochondrial electron transport, succinate to ubiquinone(GO:0006121) |
| 0.0 | 0.4 | GO:0006883 | cellular sodium ion homeostasis(GO:0006883) |
| 0.0 | 1.6 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.0 | 1.1 | GO:0010257 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.0 | 1.0 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 0.4 | GO:0060923 | cardiac muscle cell fate commitment(GO:0060923) |
| 0.0 | 1.5 | GO:0010633 | negative regulation of epithelial cell migration(GO:0010633) |
| 0.0 | 0.1 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.0 | 0.6 | GO:0044062 | regulation of excretion(GO:0044062) |
| 0.0 | 0.2 | GO:2000786 | positive regulation of autophagosome assembly(GO:2000786) |
| 0.0 | 1.7 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.0 | 0.9 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 0.1 | GO:0040009 | replicative cell aging(GO:0001302) regulation of growth rate(GO:0040009) G-quadruplex DNA unwinding(GO:0044806) |
| 0.0 | 0.3 | GO:0042487 | regulation of odontogenesis of dentin-containing tooth(GO:0042487) |
| 0.0 | 0.2 | GO:2000465 | regulation of glycogen (starch) synthase activity(GO:2000465) |
| 0.0 | 0.5 | GO:0034260 | negative regulation of GTPase activity(GO:0034260) |
| 0.0 | 0.1 | GO:0043552 | positive regulation of phosphatidylinositol 3-kinase activity(GO:0043552) |
| 0.0 | 0.3 | GO:1900017 | positive regulation of cytokine production involved in inflammatory response(GO:1900017) |
| 0.0 | 1.3 | GO:0061077 | chaperone-mediated protein folding(GO:0061077) |
| 0.0 | 0.9 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.0 | 0.5 | GO:1904376 | negative regulation of protein localization to plasma membrane(GO:1903077) negative regulation of protein localization to cell periphery(GO:1904376) |
| 0.0 | 0.3 | GO:0045109 | intermediate filament organization(GO:0045109) |
| 0.0 | 0.2 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.0 | 0.1 | GO:1901727 | positive regulation of histone deacetylase activity(GO:1901727) |
| 0.0 | 0.8 | GO:0006040 | amino sugar metabolic process(GO:0006040) |
| 0.0 | 2.8 | GO:0071805 | potassium ion transmembrane transport(GO:0071805) |
| 0.0 | 0.1 | GO:0015791 | polyol transport(GO:0015791) |
| 0.0 | 0.3 | GO:0001921 | positive regulation of receptor recycling(GO:0001921) |
| 0.0 | 0.6 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.0 | 0.1 | GO:0048208 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.0 | 1.1 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) |
| 0.0 | 0.1 | GO:0016340 | calcium-dependent cell-matrix adhesion(GO:0016340) |
| 0.0 | 0.5 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.0 | 0.7 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.0 | 1.3 | GO:0043271 | negative regulation of ion transport(GO:0043271) |
| 0.0 | 0.2 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.0 | 0.5 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.0 | 0.1 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.0 | 1.8 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.0 | 1.0 | GO:1903955 | positive regulation of protein targeting to mitochondrion(GO:1903955) |
| 0.0 | 3.5 | GO:0006874 | cellular calcium ion homeostasis(GO:0006874) |
| 0.0 | 0.2 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.0 | 0.4 | GO:0050848 | regulation of calcium-mediated signaling(GO:0050848) |
| 0.0 | 1.1 | GO:0007030 | Golgi organization(GO:0007030) |
| 0.0 | 0.3 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.0 | 1.0 | GO:0048015 | phosphatidylinositol-mediated signaling(GO:0048015) |
| 0.0 | 0.5 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.0 | 0.1 | GO:0002634 | regulation of germinal center formation(GO:0002634) |
| 0.0 | 0.9 | GO:0072332 | intrinsic apoptotic signaling pathway by p53 class mediator(GO:0072332) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 8.5 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.4 | 1.7 | GO:0097169 | AIM2 inflammasome complex(GO:0097169) |
| 0.4 | 1.2 | GO:0070195 | growth hormone receptor complex(GO:0070195) |
| 0.3 | 1.0 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.3 | 4.0 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.3 | 1.8 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.2 | 1.6 | GO:0070695 | FHF complex(GO:0070695) |
| 0.2 | 0.9 | GO:0070826 | paraferritin complex(GO:0070826) |
| 0.2 | 0.6 | GO:0032783 | ELL-EAF complex(GO:0032783) |
| 0.2 | 0.6 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.2 | 1.8 | GO:0090533 | cation-transporting ATPase complex(GO:0090533) |
| 0.2 | 0.5 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 0.2 | 1.4 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.1 | 1.0 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.1 | 3.3 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.1 | 1.8 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.1 | 0.6 | GO:1990745 | EARP complex(GO:1990745) |
| 0.1 | 1.5 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.1 | 2.1 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.1 | 0.6 | GO:0090498 | extrinsic component of Golgi membrane(GO:0090498) |
| 0.1 | 1.1 | GO:0071141 | SMAD protein complex(GO:0071141) |
| 0.1 | 1.0 | GO:0036454 | insulin-like growth factor binding protein complex(GO:0016942) growth factor complex(GO:0036454) |
| 0.1 | 1.4 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.1 | 1.5 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.1 | 0.6 | GO:0030478 | actin cap(GO:0030478) |
| 0.1 | 0.4 | GO:0070775 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.1 | 0.5 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.1 | 0.2 | GO:0043541 | UDP-N-acetylglucosamine transferase complex(GO:0043541) |
| 0.1 | 2.5 | GO:0030673 | axolemma(GO:0030673) |
| 0.1 | 1.2 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.1 | 0.4 | GO:1990462 | omegasome(GO:1990462) |
| 0.1 | 0.9 | GO:0035748 | myelin sheath abaxonal region(GO:0035748) |
| 0.1 | 0.2 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 0.1 | 0.5 | GO:0042582 | primary lysosome(GO:0005766) azurophil granule(GO:0042582) |
| 0.0 | 0.7 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 1.1 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 0.3 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.0 | 0.4 | GO:0031983 | vesicle lumen(GO:0031983) |
| 0.0 | 0.1 | GO:1990682 | CSF1-CSF1R complex(GO:1990682) |
| 0.0 | 1.0 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 0.4 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.0 | 0.7 | GO:0005767 | secondary lysosome(GO:0005767) |
| 0.0 | 4.2 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.0 | 0.3 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.0 | 0.4 | GO:0097197 | tetraspanin-enriched microdomain(GO:0097197) |
| 0.0 | 0.4 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.0 | 0.6 | GO:0036057 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.0 | 1.7 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 1.7 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 0.8 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) |
| 0.0 | 1.0 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.0 | 1.0 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 0.2 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.0 | 0.1 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.0 | 0.3 | GO:0005652 | nuclear lamina(GO:0005652) |
| 0.0 | 0.5 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 2.1 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 2.9 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 1.5 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 0.7 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.0 | 0.4 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 1.6 | GO:0005930 | axoneme(GO:0005930) |
| 0.0 | 0.7 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 0.2 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.0 | 0.5 | GO:0032421 | stereocilium bundle(GO:0032421) |
| 0.0 | 0.2 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.0 | 1.2 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 1.0 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 0.6 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 0.2 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 0.3 | GO:0005849 | mRNA cleavage factor complex(GO:0005849) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.2 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.7 | 2.0 | GO:0009041 | uridylate kinase activity(GO:0009041) |
| 0.7 | 2.7 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.6 | 1.9 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.6 | 3.5 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.5 | 1.5 | GO:0070737 | protein-glycine ligase activity, elongating(GO:0070737) |
| 0.5 | 2.8 | GO:0097603 | temperature-gated ion channel activity(GO:0097603) |
| 0.4 | 1.7 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 0.4 | 1.3 | GO:0043120 | tumor necrosis factor binding(GO:0043120) |
| 0.4 | 5.7 | GO:0015194 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.4 | 1.2 | GO:0004903 | growth hormone receptor activity(GO:0004903) |
| 0.4 | 2.2 | GO:0001758 | retinal dehydrogenase activity(GO:0001758) |
| 0.4 | 1.8 | GO:0004966 | galanin receptor activity(GO:0004966) |
| 0.4 | 1.1 | GO:0035500 | MH2 domain binding(GO:0035500) |
| 0.3 | 1.9 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.3 | 8.5 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.3 | 4.4 | GO:0019203 | carbohydrate phosphatase activity(GO:0019203) |
| 0.3 | 0.9 | GO:0038100 | nodal binding(GO:0038100) |
| 0.3 | 1.1 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.2 | 1.0 | GO:0047710 | bis(5'-adenosyl)-triphosphatase activity(GO:0047710) |
| 0.2 | 0.6 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.2 | 4.3 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.2 | 2.0 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.2 | 3.0 | GO:0016832 | aldehyde-lyase activity(GO:0016832) |
| 0.2 | 1.7 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.2 | 0.9 | GO:0015639 | cadmium ion transmembrane transporter activity(GO:0015086) lead ion transmembrane transporter activity(GO:0015094) vanadium ion transmembrane transporter activity(GO:0015100) ferrous iron uptake transmembrane transporter activity(GO:0015639) |
| 0.2 | 1.3 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.2 | 0.5 | GO:0004800 | thyroxine 5'-deiodinase activity(GO:0004800) |
| 0.2 | 0.5 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 0.2 | 0.7 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.2 | 0.5 | GO:0046870 | cadmium ion binding(GO:0046870) |
| 0.2 | 1.5 | GO:1901611 | phosphatidylglycerol binding(GO:1901611) |
| 0.2 | 2.4 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.2 | 3.2 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.2 | 1.0 | GO:1903763 | gap junction channel activity involved in cell communication by electrical coupling(GO:1903763) |
| 0.2 | 1.0 | GO:0035184 | histone threonine kinase activity(GO:0035184) |
| 0.2 | 0.9 | GO:0005030 | neurotrophin receptor activity(GO:0005030) |
| 0.2 | 2.2 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.2 | 1.1 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.1 | 0.9 | GO:0043546 | molybdopterin cofactor binding(GO:0043546) |
| 0.1 | 0.5 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.1 | 0.4 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.1 | 0.4 | GO:0019779 | Atg12 activating enzyme activity(GO:0019778) Atg8 activating enzyme activity(GO:0019779) |
| 0.1 | 0.4 | GO:0038052 | RNA polymerase II transcription factor activity, estrogen-activated sequence-specific DNA binding(GO:0038052) |
| 0.1 | 0.9 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 0.1 | 1.3 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.1 | 0.6 | GO:0004301 | epoxide hydrolase activity(GO:0004301) |
| 0.1 | 0.6 | GO:0001642 | group III metabotropic glutamate receptor activity(GO:0001642) |
| 0.1 | 0.8 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 2.9 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.1 | 1.5 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.1 | 0.8 | GO:0048101 | calmodulin-dependent cyclic-nucleotide phosphodiesterase activity(GO:0004117) calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.1 | 1.8 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.1 | 1.2 | GO:0048531 | beta-1,3-galactosyltransferase activity(GO:0048531) |
| 0.1 | 0.9 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.1 | 0.4 | GO:0042284 | sphingolipid delta-4 desaturase activity(GO:0042284) |
| 0.1 | 1.4 | GO:0004559 | alpha-mannosidase activity(GO:0004559) |
| 0.1 | 0.6 | GO:0043559 | insulin binding(GO:0043559) |
| 0.1 | 1.2 | GO:0015250 | water channel activity(GO:0015250) |
| 0.1 | 1.5 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.1 | 0.2 | GO:0047936 | glucose 1-dehydrogenase [NAD(P)] activity(GO:0047936) |
| 0.1 | 1.8 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.1 | 0.8 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.1 | 0.8 | GO:0008556 | sodium:potassium-exchanging ATPase activity(GO:0005391) potassium-transporting ATPase activity(GO:0008556) |
| 0.1 | 12.8 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 2.3 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.1 | 2.1 | GO:0032183 | SUMO binding(GO:0032183) |
| 0.1 | 1.3 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.1 | 3.2 | GO:0005272 | sodium channel activity(GO:0005272) |
| 0.1 | 0.2 | GO:0016309 | 1-phosphatidylinositol-5-phosphate 4-kinase activity(GO:0016309) |
| 0.1 | 0.5 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.1 | 0.3 | GO:0031800 | type 3 metabotropic glutamate receptor binding(GO:0031800) |
| 0.1 | 1.8 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.1 | 1.0 | GO:0044548 | S100 protein binding(GO:0044548) |
| 0.1 | 0.4 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.1 | 0.7 | GO:0070569 | uridylyltransferase activity(GO:0070569) |
| 0.1 | 0.4 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.1 | 0.2 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.1 | 1.4 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.1 | 2.5 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.1 | 1.3 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.1 | 0.9 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.1 | 0.7 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.1 | 0.7 | GO:0051378 | G-protein coupled serotonin receptor activity(GO:0004993) serotonin binding(GO:0051378) |
| 0.1 | 1.1 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.1 | 0.4 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.1 | 4.1 | GO:0005249 | voltage-gated potassium channel activity(GO:0005249) |
| 0.1 | 0.3 | GO:0042134 | rRNA primary transcript binding(GO:0042134) |
| 0.1 | 2.6 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.5 | GO:0003997 | acyl-CoA oxidase activity(GO:0003997) |
| 0.0 | 0.4 | GO:0019966 | interleukin-1 binding(GO:0019966) |
| 0.0 | 1.3 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 0.4 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.0 | 0.6 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 0.7 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 0.1 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 0.0 | 0.7 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 1.1 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.0 | 0.7 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.0 | 0.2 | GO:0034432 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.0 | 0.1 | GO:0098809 | nitrite reductase activity(GO:0098809) |
| 0.0 | 0.8 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.0 | 0.5 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.0 | 0.1 | GO:0072345 | NAADP-sensitive calcium-release channel activity(GO:0072345) |
| 0.0 | 0.5 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 1.0 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 1.9 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.0 | 0.7 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.0 | 0.5 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.0 | 0.8 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 0.1 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 0.0 | 0.2 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.0 | 4.1 | GO:0052689 | carboxylic ester hydrolase activity(GO:0052689) |
| 0.0 | 1.1 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 0.1 | GO:0001716 | L-amino-acid oxidase activity(GO:0001716) |
| 0.0 | 1.4 | GO:0008028 | monocarboxylic acid transmembrane transporter activity(GO:0008028) |
| 0.0 | 0.4 | GO:0008329 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 0.0 | 0.5 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 1.4 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.0 | 0.5 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 0.0 | 2.6 | GO:0000979 | RNA polymerase II core promoter sequence-specific DNA binding(GO:0000979) |
| 0.0 | 0.2 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.0 | 0.3 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.0 | 0.3 | GO:0031702 | type 1 angiotensin receptor binding(GO:0031702) |
| 0.0 | 0.2 | GO:0052813 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol bisphosphate kinase activity(GO:0052813) |
| 0.0 | 0.2 | GO:0071253 | connexin binding(GO:0071253) |
| 0.0 | 0.6 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 1.3 | GO:0051287 | NAD binding(GO:0051287) |
| 0.0 | 0.7 | GO:0051393 | alpha-actinin binding(GO:0051393) |
| 0.0 | 0.8 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.0 | 0.7 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 1.1 | GO:0061733 | peptide-lysine-N-acetyltransferase activity(GO:0061733) |
| 0.0 | 0.2 | GO:0004579 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.0 | 0.5 | GO:0017046 | peptide hormone binding(GO:0017046) |
| 0.0 | 0.2 | GO:0050780 | dopamine receptor binding(GO:0050780) |
| 0.0 | 0.4 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 0.1 | GO:0043262 | adenosine-diphosphatase activity(GO:0043262) |
| 0.0 | 0.5 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.0 | 1.4 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.0 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.2 | 2.9 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 2.5 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.1 | 6.6 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.1 | 1.3 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.1 | 0.8 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.1 | 1.4 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.1 | 2.8 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.1 | 0.9 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.1 | 4.0 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.1 | 1.0 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.1 | 2.2 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 1.3 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.1 | 0.4 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.1 | 2.1 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.0 | 0.7 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.0 | 1.5 | PID TXA2PATHWAY | Thromboxane A2 receptor signaling |
| 0.0 | 7.5 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 2.7 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 1.5 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.0 | 0.6 | SIG IL4RECEPTOR IN B LYPHOCYTES | Genes related to IL4 rceptor signaling in B lymphocytes |
| 0.0 | 1.5 | PID ERBB1 RECEPTOR PROXIMAL PATHWAY | EGF receptor (ErbB1) signaling pathway |
| 0.0 | 0.7 | PID CD8 TCR PATHWAY | TCR signaling in naïve CD8+ T cells |
| 0.0 | 2.0 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 1.4 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.0 | 0.6 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 0.4 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.0 | 0.5 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.0 | 0.6 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 2.8 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 0.8 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 0.2 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.0 | 0.3 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 0.2 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.0 | 0.3 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.8 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 1.5 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 0.4 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 0.6 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.0 | 0.1 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 0.4 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 0.4 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.0 | 0.8 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 0.4 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.0 | 0.2 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 0.7 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 0.1 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.0 | 0.4 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.0 | 0.6 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.8 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.1 | 1.9 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.1 | 2.9 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 1.2 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.1 | 1.8 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.1 | 3.5 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.1 | 5.2 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 2.3 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.1 | 1.3 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.1 | 1.3 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.1 | 4.0 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.1 | 1.6 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.1 | 0.6 | REACTOME ENDOGENOUS STEROLS | Genes involved in Endogenous sterols |
| 0.1 | 1.4 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.1 | 1.5 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.1 | 3.0 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.1 | 0.7 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.1 | 0.7 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.1 | 0.5 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.1 | 0.8 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.1 | 1.0 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 7.2 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.4 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 1.7 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 1.4 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.0 | 1.8 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.9 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.0 | 1.6 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 1.0 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.0 | 0.2 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.0 | 0.5 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 1.6 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.9 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.0 | 0.7 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.0 | 1.1 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 0.6 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.0 | 1.1 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 1.1 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 0.6 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.0 | 0.5 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.0 | 0.9 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.0 | 1.2 | REACTOME SPHINGOLIPID METABOLISM | Genes involved in Sphingolipid metabolism |
| 0.0 | 2.6 | REACTOME GLUCOSE METABOLISM | Genes involved in Glucose metabolism |
| 0.0 | 3.3 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 4.9 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.4 | REACTOME ACYL CHAIN REMODELLING OF PI | Genes involved in Acyl chain remodelling of PI |
| 0.0 | 0.4 | REACTOME DESTABILIZATION OF MRNA BY TRISTETRAPROLIN TTP | Genes involved in Destabilization of mRNA by Tristetraprolin (TTP) |
| 0.0 | 0.3 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 0.2 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.3 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.0 | 1.4 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.4 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 0.6 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.8 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.0 | 0.1 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.0 | 0.6 | REACTOME AMYLOIDS | Genes involved in Amyloids |