Project
avrg: 12D miR HR13_24
Navigation
Downloads
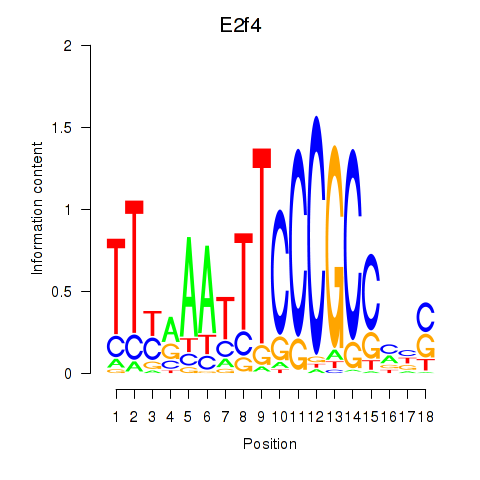
Results for E2f4
Z-value: 3.67
Transcription factors associated with E2f4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
E2f4
|
ENSMUSG00000014859.8 | E2F transcription factor 4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| E2f4 | mm10_v2_chr8_+_105297663_105297742 | 0.70 | 1.7e-02 | Click! |
Activity profile of E2f4 motif
Sorted Z-values of E2f4 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_6084983 | 7.83 |
ENSMUST00000025704.2
|
Cdca5
|
cell division cycle associated 5 |
| chr4_+_134510999 | 7.49 |
ENSMUST00000105866.2
|
Aunip
|
aurora kinase A and ninein interacting protein |
| chr8_-_53638945 | 7.02 |
ENSMUST00000047768.4
|
Neil3
|
nei like 3 (E. coli) |
| chr18_-_34751502 | 6.70 |
ENSMUST00000060710.7
|
Cdc25c
|
cell division cycle 25C |
| chr1_-_169531343 | 6.52 |
ENSMUST00000028000.7
|
Nuf2
|
NUF2, NDC80 kinetochore complex component, homolog (S. cerevisiae) |
| chr14_-_47418407 | 6.45 |
ENSMUST00000043296.3
|
Dlgap5
|
discs, large (Drosophila) homolog-associated protein 5 |
| chr6_+_124830217 | 6.40 |
ENSMUST00000131847.1
ENSMUST00000151674.1 |
Cdca3
|
cell division cycle associated 3 |
| chr1_-_169531447 | 6.08 |
ENSMUST00000111368.1
|
Nuf2
|
NUF2, NDC80 kinetochore complex component, homolog (S. cerevisiae) |
| chr1_-_189688074 | 5.75 |
ENSMUST00000171929.1
ENSMUST00000165962.1 |
Cenpf
|
centromere protein F |
| chr17_-_25727364 | 5.24 |
ENSMUST00000170070.1
ENSMUST00000048054.7 |
Chtf18
|
CTF18, chromosome transmission fidelity factor 18 |
| chr11_-_101551837 | 5.13 |
ENSMUST00000017290.4
|
Brca1
|
breast cancer 1 |
| chr10_-_88146867 | 4.54 |
ENSMUST00000164121.1
ENSMUST00000164803.1 ENSMUST00000168163.1 ENSMUST00000048518.9 |
Parpbp
|
PARP1 binding protein |
| chr12_+_117843873 | 4.51 |
ENSMUST00000176735.1
ENSMUST00000177339.1 |
Cdca7l
|
cell division cycle associated 7 like |
| chr4_-_132345686 | 4.43 |
ENSMUST00000030726.6
|
Rcc1
|
regulator of chromosome condensation 1 |
| chr13_-_73937761 | 4.39 |
ENSMUST00000022053.8
|
Trip13
|
thyroid hormone receptor interactor 13 |
| chr4_-_132345715 | 4.34 |
ENSMUST00000084250.4
|
Rcc1
|
regulator of chromosome condensation 1 |
| chr4_-_116123618 | 4.32 |
ENSMUST00000102704.3
ENSMUST00000102705.3 |
Rad54l
|
RAD54 like (S. cerevisiae) |
| chr17_-_24251382 | 4.15 |
ENSMUST00000115390.3
|
Ccnf
|
cyclin F |
| chr14_-_87141206 | 4.03 |
ENSMUST00000022599.7
|
Diap3
|
diaphanous homolog 3 (Drosophila) |
| chrX_+_164980592 | 3.97 |
ENSMUST00000101082.4
ENSMUST00000167446.1 ENSMUST00000057150.6 |
Fancb
|
Fanconi anemia, complementation group B |
| chr9_+_106477269 | 3.63 |
ENSMUST00000047721.8
|
Rrp9
|
RRP9, small subunit (SSU) processome component, homolog (yeast) |
| chr9_+_107950952 | 3.43 |
ENSMUST00000049348.3
|
Traip
|
TRAF-interacting protein |
| chr14_-_87141114 | 3.42 |
ENSMUST00000168889.1
|
Diap3
|
diaphanous homolog 3 (Drosophila) |
| chr4_+_11558914 | 3.40 |
ENSMUST00000178703.1
ENSMUST00000095145.5 ENSMUST00000108306.2 ENSMUST00000070755.6 |
Rad54b
|
RAD54 homolog B (S. cerevisiae) |
| chr16_-_90727329 | 3.37 |
ENSMUST00000099554.4
|
Mis18a
|
MIS18 kinetochore protein homolog A (S. pombe) |
| chr18_+_34751803 | 3.21 |
ENSMUST00000181453.1
ENSMUST00000181641.1 |
2010110K18Rik
|
RIKEN cDNA 2010110K18 gene |
| chr1_-_33669745 | 3.09 |
ENSMUST00000027312.9
|
Prim2
|
DNA primase, p58 subunit |
| chr7_-_92874196 | 3.01 |
ENSMUST00000032877.9
|
4632434I11Rik
|
RIKEN cDNA 4632434I11 gene |
| chr2_+_163054682 | 2.82 |
ENSMUST00000018005.3
|
Mybl2
|
myeloblastosis oncogene-like 2 |
| chr11_-_6606053 | 2.74 |
ENSMUST00000045713.3
|
Nacad
|
NAC alpha domain containing |
| chr5_-_112228934 | 2.74 |
ENSMUST00000181535.2
|
Miat
|
myocardial infarction associated transcript (non-protein coding) |
| chr5_+_33721724 | 2.69 |
ENSMUST00000067150.7
ENSMUST00000169212.2 ENSMUST00000114411.2 ENSMUST00000164207.3 |
Fgfr3
|
fibroblast growth factor receptor 3 |
| chr6_+_113531675 | 2.44 |
ENSMUST00000036340.5
ENSMUST00000101051.2 |
Fancd2
|
Fanconi anemia, complementation group D2 |
| chr1_-_57377476 | 2.25 |
ENSMUST00000181949.1
|
4930558J18Rik
|
RIKEN cDNA 4930558J18 gene |
| chr7_-_127260677 | 2.23 |
ENSMUST00000035276.4
|
Dctpp1
|
dCTP pyrophosphatase 1 |
| chr11_+_98907801 | 2.15 |
ENSMUST00000092706.6
|
Cdc6
|
cell division cycle 6 |
| chr7_-_38107490 | 1.99 |
ENSMUST00000108023.3
|
Ccne1
|
cyclin E1 |
| chr15_-_76639840 | 1.87 |
ENSMUST00000166974.1
ENSMUST00000168185.1 |
Tonsl
|
tonsoku-like, DNA repair protein |
| chr6_-_56704673 | 1.87 |
ENSMUST00000170382.2
|
Lsm5
|
LSM5 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr18_+_56707725 | 1.83 |
ENSMUST00000025486.8
|
Lmnb1
|
lamin B1 |
| chr9_-_123678782 | 1.83 |
ENSMUST00000170591.1
ENSMUST00000171647.1 |
Slc6a20a
|
solute carrier family 6 (neurotransmitter transporter), member 20A |
| chr8_+_18595131 | 1.82 |
ENSMUST00000039412.8
|
Mcph1
|
microcephaly, primary autosomal recessive 1 |
| chrX_-_8074720 | 1.80 |
ENSMUST00000115636.3
ENSMUST00000115638.3 |
Suv39h1
|
suppressor of variegation 3-9 homolog 1 (Drosophila) |
| chr15_+_55307743 | 1.76 |
ENSMUST00000023053.5
ENSMUST00000110221.2 ENSMUST00000110217.3 |
Col14a1
|
collagen, type XIV, alpha 1 |
| chr10_+_127677064 | 1.72 |
ENSMUST00000118612.1
ENSMUST00000048099.4 |
Tmem194
|
transmembrane protein 194 |
| chr2_+_30286406 | 1.72 |
ENSMUST00000138666.1
ENSMUST00000113634.2 |
Nup188
|
nucleoporin 188 |
| chr4_-_133967235 | 1.60 |
ENSMUST00000123234.1
|
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr1_+_157412352 | 1.58 |
ENSMUST00000061537.5
|
2810025M15Rik
|
RIKEN cDNA 2810025M15 gene |
| chr5_+_130257029 | 1.56 |
ENSMUST00000100662.3
ENSMUST00000040213.6 |
Tyw1
|
tRNA-yW synthesizing protein 1 homolog (S. cerevisiae) |
| chr10_-_117792663 | 1.55 |
ENSMUST00000167943.1
ENSMUST00000064848.5 |
Nup107
|
nucleoporin 107 |
| chr10_+_88146992 | 1.54 |
ENSMUST00000052355.7
|
Nup37
|
nucleoporin 37 |
| chr5_+_88764983 | 1.48 |
ENSMUST00000031311.9
|
Dck
|
deoxycytidine kinase |
| chr10_+_88147061 | 1.45 |
ENSMUST00000169309.1
|
Nup37
|
nucleoporin 37 |
| chr5_-_138172383 | 1.44 |
ENSMUST00000000505.9
|
Mcm7
|
minichromosome maintenance deficient 7 (S. cerevisiae) |
| chr11_+_16951371 | 1.44 |
ENSMUST00000109635.1
ENSMUST00000061327.1 |
Fbxo48
|
F-box protein 48 |
| chr8_+_124023394 | 1.41 |
ENSMUST00000034457.8
|
Urb2
|
URB2 ribosome biogenesis 2 homolog (S. cerevisiae) |
| chr7_+_97371604 | 1.41 |
ENSMUST00000098300.4
|
Alg8
|
asparagine-linked glycosylation 8 (alpha-1,3-glucosyltransferase) |
| chr5_-_138171813 | 1.37 |
ENSMUST00000155902.1
ENSMUST00000148879.1 |
Mcm7
|
minichromosome maintenance deficient 7 (S. cerevisiae) |
| chr9_+_72438519 | 1.30 |
ENSMUST00000184604.1
|
Mns1
|
meiosis-specific nuclear structural protein 1 |
| chr8_+_18595526 | 1.26 |
ENSMUST00000146819.1
|
Mcph1
|
microcephaly, primary autosomal recessive 1 |
| chr19_+_46075842 | 1.20 |
ENSMUST00000165017.1
|
Nolc1
|
nucleolar and coiled-body phosphoprotein 1 |
| chr9_+_44134562 | 1.19 |
ENSMUST00000034650.8
ENSMUST00000098852.2 |
Mcam
|
melanoma cell adhesion molecule |
| chrX_+_8271381 | 1.17 |
ENSMUST00000033512.4
|
Slc38a5
|
solute carrier family 38, member 5 |
| chr2_-_166713758 | 1.15 |
ENSMUST00000036719.5
|
Prex1
|
phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 1 |
| chr4_-_133967296 | 1.15 |
ENSMUST00000105893.1
|
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr18_-_60848911 | 1.15 |
ENSMUST00000177172.1
ENSMUST00000175934.1 ENSMUST00000176630.1 |
Tcof1
|
Treacher Collins Franceschetti syndrome 1, homolog |
| chr19_+_8735808 | 1.14 |
ENSMUST00000049424.9
|
Wdr74
|
WD repeat domain 74 |
| chr1_+_179803376 | 1.12 |
ENSMUST00000097454.2
|
Gm10518
|
predicted gene 10518 |
| chr2_+_181319714 | 1.12 |
ENSMUST00000098971.4
ENSMUST00000054622.8 ENSMUST00000108814.1 ENSMUST00000048608.9 ENSMUST00000108815.1 |
Rtel1
|
regulator of telomere elongation helicase 1 |
| chrX_+_8271133 | 1.12 |
ENSMUST00000127103.1
ENSMUST00000115591.1 |
Slc38a5
|
solute carrier family 38, member 5 |
| chrX_+_8271642 | 1.08 |
ENSMUST00000115590.1
|
Slc38a5
|
solute carrier family 38, member 5 |
| chr14_-_54517353 | 1.08 |
ENSMUST00000023873.5
|
Prmt5
|
protein arginine N-methyltransferase 5 |
| chr14_+_70555900 | 1.08 |
ENSMUST00000163060.1
|
Hr
|
hairless |
| chr14_+_55578123 | 1.07 |
ENSMUST00000174484.1
|
Psme1
|
proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) |
| chr7_-_16285454 | 1.07 |
ENSMUST00000174270.1
|
Ccdc9
|
coiled-coil domain containing 9 |
| chr5_-_33652339 | 1.04 |
ENSMUST00000075670.6
|
Slbp
|
stem-loop binding protein |
| chr2_-_3512746 | 1.02 |
ENSMUST00000056700.7
ENSMUST00000027961.5 |
Hspa14
Hspa14
|
heat shock protein 14 heat shock protein 14 |
| chr4_-_133967893 | 1.00 |
ENSMUST00000100472.3
ENSMUST00000136327.1 |
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr12_+_99884498 | 0.98 |
ENSMUST00000153627.1
|
Tdp1
|
tyrosyl-DNA phosphodiesterase 1 |
| chr4_-_59783800 | 0.98 |
ENSMUST00000107526.1
ENSMUST00000095063.4 |
Inip
|
INTS3 and NABP interacting protein |
| chrY_+_90784738 | 0.95 |
ENSMUST00000179483.1
|
Erdr1
|
erythroid differentiation regulator 1 |
| chr11_+_87405049 | 0.94 |
ENSMUST00000060835.5
|
Tex14
|
testis expressed gene 14 |
| chr3_+_88553716 | 0.93 |
ENSMUST00000008748.6
|
Ubqln4
|
ubiquilin 4 |
| chr17_+_34982099 | 0.92 |
ENSMUST00000007266.7
|
Lsm2
|
LSM2 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr4_-_132463873 | 0.91 |
ENSMUST00000102567.3
|
Med18
|
mediator of RNA polymerase II transcription, subunit 18 homolog (yeast) |
| chr17_+_34982154 | 0.86 |
ENSMUST00000173004.1
|
Lsm2
|
LSM2 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr17_+_50698525 | 0.85 |
ENSMUST00000061681.7
|
Gm7334
|
predicted gene 7334 |
| chrX_+_38600626 | 0.85 |
ENSMUST00000000365.2
|
Mcts1
|
malignant T cell amplified sequence 1 |
| chr14_-_33447142 | 0.84 |
ENSMUST00000111944.3
ENSMUST00000022504.5 ENSMUST00000111945.2 |
Mapk8
|
mitogen-activated protein kinase 8 |
| chr1_-_179803625 | 0.83 |
ENSMUST00000027768.7
|
Ahctf1
|
AT hook containing transcription factor 1 |
| chr4_-_3835595 | 0.81 |
ENSMUST00000138502.1
|
Rps20
|
ribosomal protein S20 |
| chr17_-_66101466 | 0.78 |
ENSMUST00000024909.8
ENSMUST00000147484.1 ENSMUST00000143987.1 |
Ndufv2
|
NADH dehydrogenase (ubiquinone) flavoprotein 2 |
| chr2_+_181319806 | 0.78 |
ENSMUST00000153112.1
|
Rtel1
|
regulator of telomere elongation helicase 1 |
| chr12_-_118198917 | 0.77 |
ENSMUST00000084806.6
|
Dnah11
|
dynein, axonemal, heavy chain 11 |
| chr2_-_172940299 | 0.73 |
ENSMUST00000009143.7
|
Bmp7
|
bone morphogenetic protein 7 |
| chr17_-_28350747 | 0.71 |
ENSMUST00000080572.7
ENSMUST00000156862.1 |
Tead3
|
TEA domain family member 3 |
| chr9_+_72438534 | 0.71 |
ENSMUST00000034746.8
|
Mns1
|
meiosis-specific nuclear structural protein 1 |
| chr15_+_57912199 | 0.70 |
ENSMUST00000022992.6
|
Tbc1d31
|
TBC1 domain family, member 31 |
| chr19_-_5366626 | 0.69 |
ENSMUST00000025762.8
|
Banf1
|
barrier to autointegration factor 1 |
| chr2_+_126152141 | 0.65 |
ENSMUST00000170908.1
|
Dtwd1
|
DTW domain containing 1 |
| chr4_-_133967953 | 0.64 |
ENSMUST00000102553.4
|
Hmgn2
|
high mobility group nucleosomal binding domain 2 |
| chr17_+_87975044 | 0.64 |
ENSMUST00000005503.3
|
Msh6
|
mutS homolog 6 (E. coli) |
| chr17_+_34981847 | 0.61 |
ENSMUST00000114011.4
|
Lsm2
|
LSM2 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr3_-_107969162 | 0.61 |
ENSMUST00000004136.8
ENSMUST00000106678.1 |
Gstm3
|
glutathione S-transferase, mu 3 |
| chr5_+_30232581 | 0.60 |
ENSMUST00000145167.1
|
Ept1
|
ethanolaminephosphotransferase 1 (CDP-ethanolamine-specific) |
| chr5_-_112228900 | 0.59 |
ENSMUST00000182509.1
|
Miat
|
myocardial infarction associated transcript (non-protein coding) |
| chr5_-_33652296 | 0.54 |
ENSMUST00000151081.1
ENSMUST00000101354.3 |
Slbp
|
stem-loop binding protein |
| chr9_+_57708534 | 0.50 |
ENSMUST00000043990.7
ENSMUST00000142807.1 |
Edc3
|
enhancer of mRNA decapping 3 homolog (S. cerevisiae) |
| chr18_-_39490649 | 0.49 |
ENSMUST00000115567.1
|
Nr3c1
|
nuclear receptor subfamily 3, group C, member 1 |
| chr7_+_58658181 | 0.48 |
ENSMUST00000168747.1
|
Atp10a
|
ATPase, class V, type 10A |
| chr7_+_45783686 | 0.47 |
ENSMUST00000118564.1
ENSMUST00000133428.1 |
Lmtk3
|
lemur tyrosine kinase 3 |
| chr7_+_6383310 | 0.46 |
ENSMUST00000081022.7
|
Zfp28
|
zinc finger protein 28 |
| chr13_-_69533839 | 0.45 |
ENSMUST00000044081.7
|
Papd7
|
PAP associated domain containing 7 |
| chr4_+_108834601 | 0.44 |
ENSMUST00000030296.8
|
Txndc12
|
thioredoxin domain containing 12 (endoplasmic reticulum) |
| chr1_+_57377593 | 0.42 |
ENSMUST00000042734.2
|
1700066M21Rik
|
RIKEN cDNA 1700066M21 gene |
| chr1_+_52630692 | 0.41 |
ENSMUST00000165859.1
|
Tmem194b
|
transmembrane protein 194B |
| chr1_+_86303221 | 0.38 |
ENSMUST00000113306.2
|
B3gnt7
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 7 |
| chr7_+_45783883 | 0.38 |
ENSMUST00000072580.5
|
Lmtk3
|
lemur tyrosine kinase 3 |
| chr11_-_106999369 | 0.38 |
ENSMUST00000106768.1
ENSMUST00000144834.1 |
Kpna2
|
karyopherin (importin) alpha 2 |
| chr7_+_24884651 | 0.36 |
ENSMUST00000153451.2
ENSMUST00000108429.1 |
Rps19
|
ribosomal protein S19 |
| chr7_+_24884611 | 0.32 |
ENSMUST00000108428.1
|
Rps19
|
ribosomal protein S19 |
| chr5_-_25705791 | 0.32 |
ENSMUST00000030773.7
|
Xrcc2
|
X-ray repair complementing defective repair in Chinese hamster cells 2 |
| chr12_+_106010263 | 0.30 |
ENSMUST00000021539.8
ENSMUST00000085026.4 ENSMUST00000072040.5 |
Vrk1
|
vaccinia related kinase 1 |
| chrX_+_105079761 | 0.30 |
ENSMUST00000119477.1
|
Pbdc1
|
polysaccharide biosynthesis domain containing 1 |
| chr10_-_81427114 | 0.29 |
ENSMUST00000078185.7
ENSMUST00000020461.8 ENSMUST00000105321.3 |
Nfic
|
nuclear factor I/C |
| chr7_-_16286010 | 0.27 |
ENSMUST00000145519.2
|
Ccdc9
|
coiled-coil domain containing 9 |
| chrX_+_105079735 | 0.26 |
ENSMUST00000033577.4
|
Pbdc1
|
polysaccharide biosynthesis domain containing 1 |
| chr4_+_44004438 | 0.24 |
ENSMUST00000107846.3
|
Clta
|
clathrin, light polypeptide (Lca) |
| chr2_+_140395446 | 0.23 |
ENSMUST00000110061.1
|
Macrod2
|
MACRO domain containing 2 |
| chr11_+_119602981 | 0.21 |
ENSMUST00000026671.6
|
Rptor
|
regulatory associated protein of MTOR, complex 1 |
| chr15_-_98677451 | 0.21 |
ENSMUST00000120997.1
ENSMUST00000109149.2 ENSMUST00000003451.4 |
Rnd1
|
Rho family GTPase 1 |
| chr15_+_12824841 | 0.20 |
ENSMUST00000090292.5
|
Drosha
|
drosha, ribonuclease type III |
| chr11_-_106999482 | 0.19 |
ENSMUST00000018506.6
|
Kpna2
|
karyopherin (importin) alpha 2 |
| chrX_-_12762069 | 0.19 |
ENSMUST00000096495.4
ENSMUST00000076016.5 |
Med14
|
mediator complex subunit 14 |
| chr3_+_121953213 | 0.17 |
ENSMUST00000037958.7
ENSMUST00000128366.1 |
Arhgap29
|
Rho GTPase activating protein 29 |
| chr4_-_43562397 | 0.16 |
ENSMUST00000030187.7
|
Tln1
|
talin 1 |
| chr2_+_101678403 | 0.16 |
ENSMUST00000004949.7
|
Traf6
|
TNF receptor-associated factor 6 |
| chr15_+_12824815 | 0.15 |
ENSMUST00000169061.1
|
Drosha
|
drosha, ribonuclease type III |
| chr7_+_126781483 | 0.15 |
ENSMUST00000172352.1
ENSMUST00000094037.4 |
Tbx6
|
T-box 6 |
| chr8_-_123158268 | 0.13 |
ENSMUST00000000755.7
|
Sult5a1
|
sulfotransferase family 5A, member 1 |
| chr12_-_87147883 | 0.13 |
ENSMUST00000037788.4
|
Pomt2
|
protein-O-mannosyltransferase 2 |
| chr6_-_148831395 | 0.13 |
ENSMUST00000145960.1
|
Ipo8
|
importin 8 |
| chr2_+_174076296 | 0.12 |
ENSMUST00000155000.1
ENSMUST00000134876.1 ENSMUST00000147038.1 |
Stx16
|
syntaxin 16 |
| chr12_-_4233958 | 0.10 |
ENSMUST00000111169.3
ENSMUST00000020981.5 |
Cenpo
|
centromere protein O |
| chr17_-_87025353 | 0.10 |
ENSMUST00000024957.6
|
Pigf
|
phosphatidylinositol glycan anchor biosynthesis, class F |
| chr1_+_132007606 | 0.09 |
ENSMUST00000086556.5
|
Elk4
|
ELK4, member of ETS oncogene family |
| chr11_-_88864534 | 0.09 |
ENSMUST00000018572.4
|
Akap1
|
A kinase (PRKA) anchor protein 1 |
| chr5_+_144100387 | 0.09 |
ENSMUST00000041804.7
|
Lmtk2
|
lemur tyrosine kinase 2 |
| chr9_-_44134481 | 0.08 |
ENSMUST00000180670.1
|
Gm10687
|
predicted gene 10687 |
| chr8_-_64849818 | 0.07 |
ENSMUST00000034017.7
|
Klhl2
|
kelch-like 2, Mayven |
| chr14_+_31019159 | 0.07 |
ENSMUST00000112094.1
ENSMUST00000144009.1 |
Pbrm1
|
polybromo 1 |
| chr11_+_108425192 | 0.06 |
ENSMUST00000150863.2
ENSMUST00000061287.5 ENSMUST00000149683.2 |
Cep112
|
centrosomal protein 112 |
| chr17_-_28350600 | 0.06 |
ENSMUST00000114799.1
|
Tead3
|
TEA domain family member 3 |
| chr7_+_97579868 | 0.03 |
ENSMUST00000042399.7
ENSMUST00000107153.1 |
Rsf1
|
remodeling and spacing factor 1 |
| chr17_+_88440711 | 0.03 |
ENSMUST00000112238.2
ENSMUST00000155640.1 |
Foxn2
|
forkhead box N2 |
| chr10_-_91123955 | 0.01 |
ENSMUST00000164505.1
ENSMUST00000170810.1 ENSMUST00000076694.6 |
Slc25a3
|
solute carrier family 25 (mitochondrial carrier, phosphate carrier), member 3 |
Network of associatons between targets according to the STRING database.
First level regulatory network of E2f4
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.6 | 7.8 | GO:0071922 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 1.7 | 5.1 | GO:2000620 | positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 1.0 | 4.0 | GO:1990414 | replication-born double-strand break repair via sister chromatid exchange(GO:1990414) |
| 0.9 | 2.7 | GO:0061144 | alveolar secondary septum development(GO:0061144) |
| 0.8 | 3.1 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.6 | 1.8 | GO:0031660 | regulation of cyclin-dependent protein serine/threonine kinase activity involved in G2/M transition of mitotic cell cycle(GO:0031660) positive regulation of cyclin-dependent protein serine/threonine kinase activity involved in G2/M transition of mitotic cell cycle(GO:0031662) |
| 0.6 | 4.2 | GO:0000320 | re-entry into mitotic cell cycle(GO:0000320) |
| 0.6 | 5.2 | GO:1900264 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.6 | 3.4 | GO:0061641 | CENP-A containing nucleosome assembly(GO:0034080) CENP-A containing chromatin organization(GO:0061641) |
| 0.6 | 2.2 | GO:0009223 | pyrimidine nucleotide catabolic process(GO:0006244) pyrimidine deoxyribonucleotide catabolic process(GO:0009223) |
| 0.5 | 4.4 | GO:0007144 | female meiosis I(GO:0007144) |
| 0.5 | 1.6 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 0.5 | 3.1 | GO:0060623 | regulation of chromosome condensation(GO:0060623) |
| 0.5 | 1.9 | GO:1904430 | mitotic telomere maintenance via semi-conservative replication(GO:1902990) negative regulation of t-circle formation(GO:1904430) |
| 0.5 | 1.8 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.4 | 4.5 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.4 | 1.5 | GO:0046125 | pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.4 | 13.5 | GO:0008608 | attachment of spindle microtubules to kinetochore(GO:0008608) |
| 0.3 | 1.7 | GO:0090521 | glomerular visceral epithelial cell migration(GO:0090521) |
| 0.3 | 2.8 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.3 | 8.4 | GO:0008340 | determination of adult lifespan(GO:0008340) |
| 0.3 | 1.1 | GO:0060161 | positive regulation of dopamine receptor signaling pathway(GO:0060161) |
| 0.3 | 3.4 | GO:0015816 | glycine transport(GO:0015816) |
| 0.2 | 2.4 | GO:2000348 | regulation of CD40 signaling pathway(GO:2000348) |
| 0.2 | 0.7 | GO:1900106 | hyaluranon cable assembly(GO:0036118) kidney mesenchyme morphogenesis(GO:0072131) metanephric mesenchyme morphogenesis(GO:0072133) regulation of hyaluranon cable assembly(GO:1900104) positive regulation of hyaluranon cable assembly(GO:1900106) |
| 0.2 | 3.4 | GO:0010804 | negative regulation of tumor necrosis factor-mediated signaling pathway(GO:0010804) |
| 0.2 | 6.7 | GO:1902751 | positive regulation of cell cycle G2/M phase transition(GO:1902751) |
| 0.2 | 0.8 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.2 | 0.8 | GO:2000017 | positive regulation of determination of dorsal identity(GO:2000017) |
| 0.2 | 1.9 | GO:0007000 | nucleolus organization(GO:0007000) |
| 0.2 | 2.4 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.2 | 3.3 | GO:0035881 | amacrine cell differentiation(GO:0035881) |
| 0.2 | 0.9 | GO:1901097 | negative regulation of autophagosome maturation(GO:1901097) |
| 0.2 | 1.8 | GO:0036123 | histone H3-K9 dimethylation(GO:0036123) |
| 0.2 | 1.6 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.1 | 7.0 | GO:0006284 | base-excision repair(GO:0006284) |
| 0.1 | 4.1 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.1 | 5.8 | GO:0051310 | metaphase plate congression(GO:0051310) |
| 0.1 | 11.6 | GO:0051225 | spindle assembly(GO:0051225) |
| 0.1 | 0.9 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.1 | 1.9 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.1 | 3.7 | GO:0040034 | regulation of development, heterochronic(GO:0040034) |
| 0.1 | 0.3 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) |
| 0.1 | 1.2 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.1 | 6.5 | GO:0007051 | spindle organization(GO:0007051) |
| 0.1 | 0.8 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.1 | 1.3 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.1 | 1.4 | GO:0006490 | oligosaccharide-lipid intermediate biosynthetic process(GO:0006490) |
| 0.1 | 0.7 | GO:0015074 | DNA integration(GO:0015074) |
| 0.1 | 0.5 | GO:1900170 | negative regulation of glucocorticoid mediated signaling pathway(GO:1900170) |
| 0.1 | 0.8 | GO:0003356 | regulation of cilium beat frequency(GO:0003356) |
| 0.1 | 0.3 | GO:0043987 | histone H3-S10 phosphorylation(GO:0043987) |
| 0.1 | 1.0 | GO:0000012 | single strand break repair(GO:0000012) |
| 0.1 | 0.8 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 0.1 | 0.5 | GO:0031087 | deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.1 | 0.4 | GO:0030422 | production of siRNA involved in RNA interference(GO:0030422) |
| 0.1 | 1.1 | GO:0033169 | histone H3-K9 demethylation(GO:0033169) |
| 0.1 | 1.8 | GO:0061050 | regulation of cell growth involved in cardiac muscle cell development(GO:0061050) |
| 0.0 | 0.1 | GO:1904100 | regulation of protein O-linked glycosylation(GO:1904098) positive regulation of protein O-linked glycosylation(GO:1904100) |
| 0.0 | 0.8 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 1.2 | GO:0019884 | antigen processing and presentation of exogenous antigen(GO:0019884) |
| 0.0 | 0.2 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.0 | 0.2 | GO:0071233 | response to leucine(GO:0043201) cellular response to leucine(GO:0071233) |
| 0.0 | 0.2 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.0 | 1.2 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.0 | 0.1 | GO:0014043 | negative regulation of neuron maturation(GO:0014043) |
| 0.0 | 0.6 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 1.2 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.2 | GO:0016322 | neuron remodeling(GO:0016322) |
| 0.0 | 1.0 | GO:0001825 | blastocyst formation(GO:0001825) |
| 0.0 | 0.1 | GO:0051534 | negative regulation of NFAT protein import into nucleus(GO:0051534) |
| 0.0 | 1.6 | GO:0008033 | tRNA processing(GO:0008033) |
| 0.0 | 0.8 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.0 | 3.0 | GO:0042254 | ribosome biogenesis(GO:0042254) |
| 0.0 | 0.8 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 12.6 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 0.9 | 7.8 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.9 | 5.1 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 0.6 | 1.9 | GO:0035101 | FACT complex(GO:0035101) |
| 0.6 | 3.1 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.6 | 4.3 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.6 | 5.2 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.6 | 2.8 | GO:0031523 | Myb complex(GO:0031523) |
| 0.5 | 5.8 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.5 | 3.6 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.5 | 5.4 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.4 | 4.0 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.4 | 1.8 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.3 | 1.7 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.3 | 6.4 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.3 | 1.1 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.3 | 1.8 | GO:0005638 | lamin filament(GO:0005638) |
| 0.2 | 4.1 | GO:0001673 | male germ cell nucleus(GO:0001673) |
| 0.2 | 1.6 | GO:0071204 | histone pre-mRNA 3'end processing complex(GO:0071204) |
| 0.2 | 2.8 | GO:0042555 | MCM complex(GO:0042555) |
| 0.2 | 1.0 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.2 | 1.1 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.1 | 1.1 | GO:0034709 | methylosome(GO:0034709) |
| 0.1 | 0.4 | GO:1903095 | microprocessor complex(GO:0070877) ribonuclease III complex(GO:1903095) |
| 0.1 | 0.9 | GO:0031595 | nuclear proteasome complex(GO:0031595) |
| 0.1 | 0.6 | GO:0032300 | mismatch repair complex(GO:0032300) |
| 0.1 | 9.6 | GO:0000922 | spindle pole(GO:0000922) |
| 0.1 | 0.2 | GO:0099631 | postsynaptic endocytic zone cytoplasmic component(GO:0099631) |
| 0.1 | 1.8 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.1 | 0.3 | GO:0033063 | Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.1 | 4.2 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 2.3 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 5.9 | GO:0000793 | condensed chromosome(GO:0000793) |
| 0.0 | 1.1 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 2.0 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 1.2 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 4.5 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 3.7 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 1.4 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 2.0 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.8 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.0 | 3.1 | GO:0000775 | chromosome, centromeric region(GO:0000775) |
| 0.0 | 0.2 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.0 | 0.8 | GO:0045271 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.8 | GO:0030286 | dynein complex(GO:0030286) |
| 0.0 | 10.0 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 8.8 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 1.4 | 7.0 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 0.5 | 1.6 | GO:0071207 | histone pre-mRNA stem-loop binding(GO:0071207) |
| 0.5 | 3.4 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.5 | 5.2 | GO:0033170 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.5 | 1.4 | GO:0000033 | alpha-1,3-mannosyltransferase activity(GO:0000033) |
| 0.5 | 3.6 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.4 | 4.3 | GO:0036310 | annealing helicase activity(GO:0036310) |
| 0.4 | 1.2 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.4 | 1.5 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.3 | 2.7 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.3 | 2.8 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.3 | 3.4 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.3 | 1.1 | GO:0035243 | protein-arginine omega-N symmetric methyltransferase activity(GO:0035243) |
| 0.3 | 1.8 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 0.2 | 1.8 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.2 | 2.2 | GO:0047429 | nucleoside-triphosphate diphosphatase activity(GO:0047429) |
| 0.2 | 0.6 | GO:0032142 | guanine/thymine mispair binding(GO:0032137) single guanine insertion binding(GO:0032142) |
| 0.2 | 0.8 | GO:0004705 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.2 | 5.8 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.1 | 0.4 | GO:0019153 | protein-disulfide reductase (glutathione) activity(GO:0019153) |
| 0.1 | 4.3 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.1 | 6.7 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.1 | 1.1 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.1 | 3.3 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 3.1 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 1.6 | GO:0010181 | FMN binding(GO:0010181) |
| 0.1 | 0.5 | GO:0038049 | glucocorticoid receptor activity(GO:0004883) transcription factor activity, ligand-activated RNA polymerase II transcription factor binding(GO:0038049) glucocorticoid-activated RNA polymerase II transcription factor binding transcription factor activity(GO:0038051) |
| 0.1 | 2.4 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.1 | 4.8 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.1 | 2.8 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.1 | 0.2 | GO:0001156 | TFIIIC-class transcription factor binding(GO:0001156) |
| 0.1 | 0.4 | GO:0032296 | ribonuclease III activity(GO:0004525) double-stranded RNA-specific ribonuclease activity(GO:0032296) |
| 0.1 | 1.8 | GO:0008432 | JUN kinase binding(GO:0008432) |
| 0.1 | 1.1 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.1 | 0.4 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.1 | 0.7 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.1 | 0.3 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.1 | 2.0 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.1 | 0.3 | GO:0035184 | nucleosomal histone binding(GO:0031493) histone threonine kinase activity(GO:0035184) |
| 0.0 | 1.2 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.0 | 0.8 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.0 | 0.8 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.7 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.0 | 0.1 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.0 | 4.7 | GO:0004721 | phosphoprotein phosphatase activity(GO:0004721) |
| 0.0 | 0.5 | GO:0016891 | endoribonuclease activity, producing 5'-phosphomonoesters(GO:0016891) |
| 0.0 | 0.2 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.0 | 0.6 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 0.9 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.0 | 7.8 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 0.2 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.0 | 0.6 | GO:0004364 | glutathione transferase activity(GO:0004364) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 13.9 | PID ATM PATHWAY | ATM pathway |
| 0.2 | 6.4 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.2 | 4.0 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.2 | 7.6 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 5.0 | PID ATR PATHWAY | ATR signaling pathway |
| 0.1 | 8.3 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.1 | 0.7 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.1 | 3.9 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 1.9 | ST GAQ PATHWAY | G alpha q Pathway |
| 0.0 | 1.8 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.0 | 1.8 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 2.7 | PID FGF PATHWAY | FGF signaling pathway |
| 0.0 | 0.8 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.0 | 1.8 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 1.6 | SIG PIP3 SIGNALING IN CARDIAC MYOCTES | Genes related to PIP3 signaling in cardiac myocytes |
| 0.0 | 0.6 | PID TCPTP PATHWAY | Signaling events mediated by TCPTP |
| 0.0 | 0.2 | PID ERBB1 RECEPTOR PROXIMAL PATHWAY | EGF receptor (ErbB1) signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 6.7 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.5 | 11.5 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.3 | 3.1 | REACTOME INHIBITION OF REPLICATION INITIATION OF DAMAGED DNA BY RB1 E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |
| 0.3 | 4.8 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.3 | 7.0 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.2 | 6.3 | REACTOME REGULATION OF GLUCOKINASE BY GLUCOKINASE REGULATORY PROTEIN | Genes involved in Regulation of Glucokinase by Glucokinase Regulatory Protein |
| 0.2 | 2.8 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.2 | 2.7 | REACTOME SIGNALING BY FGFR3 MUTANTS | Genes involved in Signaling by FGFR3 mutants |
| 0.1 | 1.6 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.1 | 8.8 | REACTOME LATE PHASE OF HIV LIFE CYCLE | Genes involved in Late Phase of HIV Life Cycle |
| 0.1 | 8.9 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.1 | 0.7 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.1 | 1.5 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 3.4 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.1 | 3.1 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.1 | 1.0 | REACTOME NRIF SIGNALS CELL DEATH FROM THE NUCLEUS | Genes involved in NRIF signals cell death from the nucleus |
| 0.1 | 1.1 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.1 | 1.8 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.1 | 0.4 | REACTOME MICRORNA MIRNA BIOGENESIS | Genes involved in MicroRNA (miRNA) Biogenesis |
| 0.0 | 1.4 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 1.0 | REACTOME DOUBLE STRAND BREAK REPAIR | Genes involved in Double-Strand Break Repair |
| 0.0 | 0.6 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.0 | 1.5 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 0.4 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 1.1 | REACTOME CDK MEDIATED PHOSPHORYLATION AND REMOVAL OF CDC6 | Genes involved in CDK-mediated phosphorylation and removal of Cdc6 |
| 0.0 | 0.8 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.5 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.0 | 1.1 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 1.1 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.2 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 0.4 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.0 | 1.2 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 0.2 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.0 | 0.8 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.5 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |