Project
avrg: 12D miR HR13_24
Navigation
Downloads
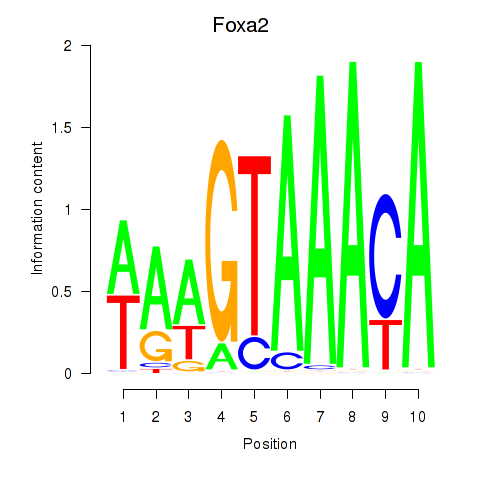
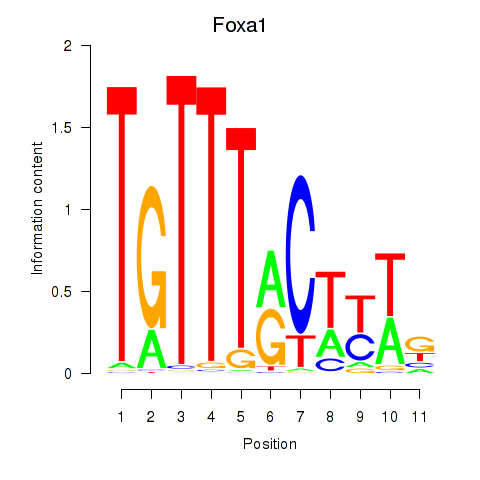
Results for Foxa2_Foxa1
Z-value: 3.57
Transcription factors associated with Foxa2_Foxa1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Foxa2
|
ENSMUSG00000037025.5 | forkhead box A2 |
|
Foxa1
|
ENSMUSG00000035451.6 | forkhead box A1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Foxa2 | mm10_v2_chr2_-_148045891_148045948 | -0.82 | 1.9e-03 | Click! |
| Foxa1 | mm10_v2_chr12_-_57546121_57546141 | -0.47 | 1.5e-01 | Click! |
Activity profile of Foxa2_Foxa1 motif
Sorted Z-values of Foxa2_Foxa1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_+_104766308 | 19.00 |
ENSMUST00000031663.3
|
C8b
|
complement component 8, beta polypeptide |
| chr4_+_104766334 | 18.22 |
ENSMUST00000065072.6
|
C8b
|
complement component 8, beta polypeptide |
| chr2_-_62483637 | 11.56 |
ENSMUST00000136686.1
ENSMUST00000102733.3 |
Gcg
|
glucagon |
| chr10_+_115817247 | 8.66 |
ENSMUST00000035563.7
ENSMUST00000080630.3 ENSMUST00000179196.1 |
Tspan8
|
tetraspanin 8 |
| chr4_-_57916283 | 8.23 |
ENSMUST00000063816.5
|
D630039A03Rik
|
RIKEN cDNA D630039A03 gene |
| chr5_-_86906937 | 7.63 |
ENSMUST00000031181.9
ENSMUST00000113333.1 |
Ugt2b34
|
UDP glucuronosyltransferase 2 family, polypeptide B34 |
| chrX_-_108664891 | 6.34 |
ENSMUST00000178160.1
|
Gm379
|
predicted gene 379 |
| chr3_-_113532288 | 6.26 |
ENSMUST00000132353.1
|
Amy2a1
|
amylase 2a1 |
| chr6_+_90619241 | 5.35 |
ENSMUST00000032177.8
|
Slc41a3
|
solute carrier family 41, member 3 |
| chr14_+_41131777 | 5.32 |
ENSMUST00000022314.3
ENSMUST00000170719.1 |
Sftpa1
|
surfactant associated protein A1 |
| chrX_+_164140447 | 5.04 |
ENSMUST00000073973.4
|
Ace2
|
angiotensin I converting enzyme (peptidyl-dipeptidase A) 2 |
| chr3_-_113258837 | 4.94 |
ENSMUST00000098673.3
|
Amy2a5
|
amylase 2a5 |
| chr11_+_96929260 | 4.79 |
ENSMUST00000054311.5
ENSMUST00000107636.3 |
Prr15l
|
proline rich 15-like |
| chr11_+_96929367 | 4.69 |
ENSMUST00000062172.5
|
Prr15l
|
proline rich 15-like |
| chr16_-_22439719 | 4.55 |
ENSMUST00000079601.6
|
Etv5
|
ets variant gene 5 |
| chr14_+_103513328 | 3.84 |
ENSMUST00000095576.3
|
Scel
|
sciellin |
| chr14_-_110755100 | 3.79 |
ENSMUST00000078386.2
|
Slitrk6
|
SLIT and NTRK-like family, member 6 |
| chr17_+_70522083 | 3.70 |
ENSMUST00000148486.1
ENSMUST00000133717.1 |
Dlgap1
|
discs, large (Drosophila) homolog-associated protein 1 |
| chr11_-_99993992 | 3.55 |
ENSMUST00000105049.1
|
Krtap17-1
|
keratin associated protein 17-1 |
| chr6_+_5390387 | 3.55 |
ENSMUST00000183358.1
|
Asb4
|
ankyrin repeat and SOCS box-containing 4 |
| chr14_-_41185188 | 3.54 |
ENSMUST00000077136.3
|
Sftpd
|
surfactant associated protein D |
| chr15_+_10223974 | 3.54 |
ENSMUST00000128450.1
ENSMUST00000148257.1 ENSMUST00000128921.1 |
Prlr
|
prolactin receptor |
| chr6_+_30541582 | 3.38 |
ENSMUST00000096066.4
|
Cpa2
|
carboxypeptidase A2, pancreatic |
| chr5_+_75152274 | 3.33 |
ENSMUST00000000476.8
|
Pdgfra
|
platelet derived growth factor receptor, alpha polypeptide |
| chrX_+_101376359 | 2.89 |
ENSMUST00000119080.1
|
Gjb1
|
gap junction protein, beta 1 |
| chr10_-_93310963 | 2.84 |
ENSMUST00000151153.1
|
Elk3
|
ELK3, member of ETS oncogene family |
| chr16_+_17331371 | 2.82 |
ENSMUST00000023450.6
ENSMUST00000161034.1 |
Serpind1
|
serine (or cysteine) peptidase inhibitor, clade D, member 1 |
| chr13_-_95478655 | 2.79 |
ENSMUST00000022186.3
|
S100z
|
S100 calcium binding protein, zeta |
| chr4_+_102570065 | 2.76 |
ENSMUST00000097950.2
|
Pde4b
|
phosphodiesterase 4B, cAMP specific |
| chr4_-_127354074 | 2.58 |
ENSMUST00000106090.1
ENSMUST00000060419.1 |
Gjb4
|
gap junction protein, beta 4 |
| chr5_+_102481374 | 2.56 |
ENSMUST00000094559.2
ENSMUST00000073302.5 |
Arhgap24
|
Rho GTPase activating protein 24 |
| chr6_-_136922169 | 2.48 |
ENSMUST00000032343.6
|
Erp27
|
endoplasmic reticulum protein 27 |
| chr17_-_31144271 | 2.48 |
ENSMUST00000024826.7
|
Tff2
|
trefoil factor 2 (spasmolytic protein 1) |
| chr5_+_102481546 | 2.47 |
ENSMUST00000112854.1
|
Arhgap24
|
Rho GTPase activating protein 24 |
| chr14_+_73237891 | 2.46 |
ENSMUST00000044405.6
|
Lpar6
|
lysophosphatidic acid receptor 6 |
| chr1_-_39720989 | 2.42 |
ENSMUST00000151913.1
|
Rfx8
|
regulatory factor X 8 |
| chr6_+_142298419 | 2.39 |
ENSMUST00000041993.2
|
Iapp
|
islet amyloid polypeptide |
| chr3_+_106486009 | 2.36 |
ENSMUST00000183271.1
ENSMUST00000061206.3 |
Dennd2d
|
DENN/MADD domain containing 2D |
| chr6_+_97991776 | 2.36 |
ENSMUST00000043628.6
|
Mitf
|
microphthalmia-associated transcription factor |
| chr2_+_24345282 | 2.30 |
ENSMUST00000114485.2
|
Il1rn
|
interleukin 1 receptor antagonist |
| chr1_+_157526127 | 2.29 |
ENSMUST00000111700.1
|
Sec16b
|
SEC16 homolog B (S. cerevisiae) |
| chr11_-_99986593 | 2.22 |
ENSMUST00000105050.2
|
Krtap16-1
|
keratin associated protein 16-1 |
| chr13_-_95525239 | 2.19 |
ENSMUST00000022185.8
|
F2rl1
|
coagulation factor II (thrombin) receptor-like 1 |
| chr2_+_58755177 | 2.19 |
ENSMUST00000102755.3
|
Upp2
|
uridine phosphorylase 2 |
| chr10_-_93311073 | 2.17 |
ENSMUST00000008542.5
|
Elk3
|
ELK3, member of ETS oncogene family |
| chr3_-_20275659 | 2.13 |
ENSMUST00000011607.5
|
Cpb1
|
carboxypeptidase B1 (tissue) |
| chr3_-_106167564 | 2.11 |
ENSMUST00000063062.8
|
Chi3l3
|
chitinase 3-like 3 |
| chr17_+_70522149 | 2.08 |
ENSMUST00000140728.1
|
Dlgap1
|
discs, large (Drosophila) homolog-associated protein 1 |
| chr7_+_131032061 | 2.08 |
ENSMUST00000084509.3
|
Dmbt1
|
deleted in malignant brain tumors 1 |
| chr5_-_108795352 | 2.04 |
ENSMUST00000004943.1
|
Tmed11
|
transmembrane emp24 protein transport domain containing |
| chr6_+_92940572 | 2.04 |
ENSMUST00000181145.1
ENSMUST00000181840.1 |
9530026P05Rik
|
RIKEN cDNA 9530026P05 gene |
| chr4_-_58499398 | 1.92 |
ENSMUST00000107570.1
|
Lpar1
|
lysophosphatidic acid receptor 1 |
| chr5_+_16553488 | 1.90 |
ENSMUST00000030683.3
|
Hgf
|
hepatocyte growth factor |
| chr2_+_58754910 | 1.88 |
ENSMUST00000059102.6
|
Upp2
|
uridine phosphorylase 2 |
| chr2_+_24345305 | 1.82 |
ENSMUST00000114482.1
|
Il1rn
|
interleukin 1 receptor antagonist |
| chr10_-_41611319 | 1.80 |
ENSMUST00000179614.1
|
Ccdc162
|
coiled-coil domain containing 162 |
| chr13_+_40859768 | 1.80 |
ENSMUST00000110191.2
|
Gcnt2
|
glucosaminyl (N-acetyl) transferase 2, I-branching enzyme |
| chr11_-_100146120 | 1.73 |
ENSMUST00000007317.7
|
Krt19
|
keratin 19 |
| chr8_+_107119110 | 1.65 |
ENSMUST00000046116.1
|
C630050I24Rik
|
RIKEN cDNA C630050I24 gene |
| chr5_-_123865491 | 1.62 |
ENSMUST00000057145.5
|
Niacr1
|
niacin receptor 1 |
| chr6_+_34863130 | 1.61 |
ENSMUST00000074949.3
|
Tmem140
|
transmembrane protein 140 |
| chr3_+_106113229 | 1.60 |
ENSMUST00000079132.5
ENSMUST00000139086.1 |
Chia
|
chitinase, acidic |
| chr7_-_100656953 | 1.59 |
ENSMUST00000107046.1
ENSMUST00000107045.1 ENSMUST00000139708.1 |
Plekhb1
|
pleckstrin homology domain containing, family B (evectins) member 1 |
| chr14_-_30943275 | 1.57 |
ENSMUST00000006704.8
ENSMUST00000163118.1 |
Itih1
|
inter-alpha trypsin inhibitor, heavy chain 1 |
| chr1_-_74885322 | 1.54 |
ENSMUST00000159232.1
ENSMUST00000068631.3 |
Fev
|
FEV (ETS oncogene family) |
| chr15_-_84855093 | 1.52 |
ENSMUST00000016768.5
|
Phf21b
|
PHD finger protein 21B |
| chr3_-_145032765 | 1.50 |
ENSMUST00000029919.5
|
Clca3
|
chloride channel calcium activated 3 |
| chr15_-_66560997 | 1.47 |
ENSMUST00000048372.5
|
Tmem71
|
transmembrane protein 71 |
| chr18_+_36952621 | 1.46 |
ENSMUST00000115661.2
|
Pcdha2
|
protocadherin alpha 2 |
| chr2_+_164403194 | 1.45 |
ENSMUST00000017151.1
|
Rbpjl
|
recombination signal binding protein for immunoglobulin kappa J region-like |
| chr12_+_71016658 | 1.44 |
ENSMUST00000125125.1
|
Arid4a
|
AT rich interactive domain 4A (RBP1-like) |
| chr19_-_58455398 | 1.42 |
ENSMUST00000026076.7
|
Gfra1
|
glial cell line derived neurotrophic factor family receptor alpha 1 |
| chr11_+_94565039 | 1.41 |
ENSMUST00000040418.8
|
Chad
|
chondroadherin |
| chr14_+_55853997 | 1.39 |
ENSMUST00000100529.3
|
Nynrin
|
NYN domain and retroviral integrase containing |
| chr2_-_51973219 | 1.37 |
ENSMUST00000028314.2
|
Nmi
|
N-myc (and STAT) interactor |
| chr16_-_22161450 | 1.37 |
ENSMUST00000115379.1
|
Igf2bp2
|
insulin-like growth factor 2 mRNA binding protein 2 |
| chr8_+_70373541 | 1.35 |
ENSMUST00000003659.7
|
Comp
|
cartilage oligomeric matrix protein |
| chr2_-_70825726 | 1.32 |
ENSMUST00000038584.8
|
Tlk1
|
tousled-like kinase 1 |
| chr11_+_110399115 | 1.32 |
ENSMUST00000020949.5
ENSMUST00000100260.1 |
Map2k6
|
mitogen-activated protein kinase kinase 6 |
| chr19_+_38264761 | 1.32 |
ENSMUST00000087252.5
|
Lgi1
|
leucine-rich repeat LGI family, member 1 |
| chr8_+_127064022 | 1.31 |
ENSMUST00000160272.1
ENSMUST00000079777.5 ENSMUST00000162907.1 |
Pard3
|
par-3 (partitioning defective 3) homolog (C. elegans) |
| chr14_+_55854115 | 1.28 |
ENSMUST00000168479.1
|
Nynrin
|
NYN domain and retroviral integrase containing |
| chr2_-_104742802 | 1.27 |
ENSMUST00000028595.7
|
Depdc7
|
DEP domain containing 7 |
| chr16_-_22657165 | 1.26 |
ENSMUST00000089925.3
|
Dgkg
|
diacylglycerol kinase, gamma |
| chr1_+_179546303 | 1.25 |
ENSMUST00000040706.8
|
Cnst
|
consortin, connexin sorting protein |
| chr2_+_24336846 | 1.24 |
ENSMUST00000114487.2
|
Il1rn
|
interleukin 1 receptor antagonist |
| chr19_+_55741810 | 1.23 |
ENSMUST00000111657.3
ENSMUST00000061496.9 ENSMUST00000041717.7 ENSMUST00000111662.4 |
Tcf7l2
|
transcription factor 7 like 2, T cell specific, HMG box |
| chr16_-_74411292 | 1.20 |
ENSMUST00000117200.1
|
Robo2
|
roundabout homolog 2 (Drosophila) |
| chr16_-_22657182 | 1.19 |
ENSMUST00000023578.7
|
Dgkg
|
diacylglycerol kinase, gamma |
| chr19_-_58455161 | 1.18 |
ENSMUST00000135730.1
ENSMUST00000152507.1 |
Gfra1
|
glial cell line derived neurotrophic factor family receptor alpha 1 |
| chr18_-_38918642 | 1.18 |
ENSMUST00000040647.4
|
Fgf1
|
fibroblast growth factor 1 |
| chr7_+_19359740 | 1.18 |
ENSMUST00000140836.1
|
Ppp1r13l
|
protein phosphatase 1, regulatory (inhibitor) subunit 13 like |
| chr11_+_87699897 | 1.16 |
ENSMUST00000040089.4
|
Rnf43
|
ring finger protein 43 |
| chr6_+_139843648 | 1.15 |
ENSMUST00000087657.6
|
Pik3c2g
|
phosphatidylinositol 3-kinase, C2 domain containing, gamma polypeptide |
| chr4_+_102254993 | 1.13 |
ENSMUST00000106908.2
|
Pde4b
|
phosphodiesterase 4B, cAMP specific |
| chr13_-_60177357 | 1.10 |
ENSMUST00000065086.4
|
Gas1
|
growth arrest specific 1 |
| chr14_+_61607455 | 1.09 |
ENSMUST00000051184.8
|
Kcnrg
|
potassium channel regulator |
| chr2_+_69219971 | 1.08 |
ENSMUST00000005364.5
ENSMUST00000112317.2 |
G6pc2
|
glucose-6-phosphatase, catalytic, 2 |
| chr2_-_51972990 | 1.08 |
ENSMUST00000145481.1
ENSMUST00000112705.2 |
Nmi
|
N-myc (and STAT) interactor |
| chr1_-_156034826 | 1.06 |
ENSMUST00000141878.1
ENSMUST00000123705.1 |
Tor1aip1
|
torsin A interacting protein 1 |
| chr19_+_55741884 | 1.05 |
ENSMUST00000111658.3
ENSMUST00000111654.1 |
Tcf7l2
|
transcription factor 7 like 2, T cell specific, HMG box |
| chr18_+_51117754 | 1.05 |
ENSMUST00000116639.2
|
Prr16
|
proline rich 16 |
| chr1_-_87573825 | 1.04 |
ENSMUST00000068681.5
|
Ngef
|
neuronal guanine nucleotide exchange factor |
| chr2_-_163645125 | 1.03 |
ENSMUST00000017851.3
|
Serinc3
|
serine incorporator 3 |
| chr1_-_156034800 | 1.02 |
ENSMUST00000169241.1
|
Tor1aip1
|
torsin A interacting protein 1 |
| chr15_+_5185700 | 1.02 |
ENSMUST00000081640.5
|
Ttc33
|
tetratricopeptide repeat domain 33 |
| chr2_+_30595037 | 1.01 |
ENSMUST00000102853.3
|
Cstad
|
CSA-conditional, T cell activation-dependent protein |
| chr11_+_114851142 | 1.00 |
ENSMUST00000133245.1
ENSMUST00000122967.2 |
Gprc5c
|
G protein-coupled receptor, family C, group 5, member C |
| chr6_+_29853746 | 1.00 |
ENSMUST00000064872.6
ENSMUST00000152581.1 ENSMUST00000176265.1 ENSMUST00000154079.1 |
Ahcyl2
|
S-adenosylhomocysteine hydrolase-like 2 |
| chr14_-_88471396 | 0.98 |
ENSMUST00000061628.5
|
Pcdh20
|
protocadherin 20 |
| chrX_+_159708593 | 0.98 |
ENSMUST00000080394.6
|
Sh3kbp1
|
SH3-domain kinase binding protein 1 |
| chr4_+_11123950 | 0.96 |
ENSMUST00000142297.1
|
Gm11827
|
predicted gene 11827 |
| chr16_-_38294774 | 0.95 |
ENSMUST00000023504.4
|
Nr1i2
|
nuclear receptor subfamily 1, group I, member 2 |
| chr17_+_68837062 | 0.95 |
ENSMUST00000178545.1
|
Tmem200c
|
transmembrane protein 200C |
| chr18_-_62756275 | 0.95 |
ENSMUST00000067450.1
ENSMUST00000048109.5 |
2700046A07Rik
|
RIKEN cDNA 2700046A07 gene |
| chr18_+_69925466 | 0.94 |
ENSMUST00000043929.4
|
Ccdc68
|
coiled-coil domain containing 68 |
| chr3_-_69859875 | 0.92 |
ENSMUST00000051239.7
ENSMUST00000171529.1 |
Sptssb
|
serine palmitoyltransferase, small subunit B |
| chr11_+_115824029 | 0.91 |
ENSMUST00000103032.4
ENSMUST00000133250.1 ENSMUST00000177736.1 |
Llgl2
|
lethal giant larvae homolog 2 (Drosophila) |
| chr12_-_31950535 | 0.91 |
ENSMUST00000172314.2
|
Hbp1
|
high mobility group box transcription factor 1 |
| chr4_+_150853919 | 0.91 |
ENSMUST00000073600.2
|
Errfi1
|
ERBB receptor feedback inhibitor 1 |
| chr13_-_74807913 | 0.90 |
ENSMUST00000065629.4
|
Cast
|
calpastatin |
| chr2_-_60722636 | 0.89 |
ENSMUST00000028348.2
ENSMUST00000112517.1 |
Itgb6
|
integrin beta 6 |
| chr13_+_111867931 | 0.89 |
ENSMUST00000128198.1
|
Gm15326
|
predicted gene 15326 |
| chr1_-_166409773 | 0.88 |
ENSMUST00000135673.1
ENSMUST00000079972.6 ENSMUST00000169324.1 ENSMUST00000111411.2 ENSMUST00000128861.1 |
Pogk
|
pogo transposable element with KRAB domain |
| chr3_-_113324052 | 0.88 |
ENSMUST00000179314.1
|
Amy2a3
|
amylase 2a3 |
| chr6_+_15185203 | 0.88 |
ENSMUST00000154448.1
|
Foxp2
|
forkhead box P2 |
| chr18_+_69925542 | 0.88 |
ENSMUST00000080050.5
|
Ccdc68
|
coiled-coil domain containing 68 |
| chr3_-_113291449 | 0.87 |
ENSMUST00000179568.1
|
Amy2a4
|
amylase 2a4 |
| chr15_-_3303521 | 0.87 |
ENSMUST00000165386.1
|
Ccdc152
|
coiled-coil domain containing 152 |
| chr9_+_107975529 | 0.86 |
ENSMUST00000035216.4
|
Uba7
|
ubiquitin-like modifier activating enzyme 7 |
| chr15_-_5108492 | 0.85 |
ENSMUST00000118365.2
|
Card6
|
caspase recruitment domain family, member 6 |
| chr5_-_51567717 | 0.85 |
ENSMUST00000127135.1
ENSMUST00000151104.1 |
Ppargc1a
|
peroxisome proliferative activated receptor, gamma, coactivator 1 alpha |
| chr2_-_170194033 | 0.84 |
ENSMUST00000180625.1
|
Gm17619
|
predicted gene, 17619 |
| chr7_+_78913401 | 0.84 |
ENSMUST00000118867.1
|
Isg20
|
interferon-stimulated protein |
| chr9_+_74848437 | 0.83 |
ENSMUST00000161862.1
ENSMUST00000162089.1 ENSMUST00000160017.1 ENSMUST00000160950.1 |
Gm16551
Gm20649
|
predicted gene 16551 predicted gene 20649 |
| chr14_-_93888732 | 0.83 |
ENSMUST00000068992.2
|
Pcdh9
|
protocadherin 9 |
| chr1_-_170867761 | 0.83 |
ENSMUST00000027974.6
|
Atf6
|
activating transcription factor 6 |
| chr18_-_84086379 | 0.83 |
ENSMUST00000060303.8
|
Tshz1
|
teashirt zinc finger family member 1 |
| chr19_+_55742056 | 0.82 |
ENSMUST00000111659.2
|
Tcf7l2
|
transcription factor 7 like 2, T cell specific, HMG box |
| chr2_+_160888101 | 0.82 |
ENSMUST00000109455.2
ENSMUST00000040872.6 |
Lpin3
|
lipin 3 |
| chr7_-_29168647 | 0.81 |
ENSMUST00000048923.6
|
Spred3
|
sprouty-related, EVH1 domain containing 3 |
| chr5_-_103211251 | 0.80 |
ENSMUST00000060871.5
ENSMUST00000112846.1 ENSMUST00000170792.1 ENSMUST00000112847.2 |
Mapk10
|
mitogen-activated protein kinase 10 |
| chr2_+_125136692 | 0.80 |
ENSMUST00000099452.2
|
Ctxn2
|
cortexin 2 |
| chr12_+_77238093 | 0.79 |
ENSMUST00000177595.1
ENSMUST00000171770.2 |
Fut8
|
fucosyltransferase 8 |
| chr6_-_137571007 | 0.78 |
ENSMUST00000100841.2
|
Eps8
|
epidermal growth factor receptor pathway substrate 8 |
| chr17_+_46161021 | 0.78 |
ENSMUST00000024748.7
ENSMUST00000172170.1 |
Gtpbp2
|
GTP binding protein 2 |
| chr8_-_45262012 | 0.78 |
ENSMUST00000034064.3
|
F11
|
coagulation factor XI |
| chr3_-_52104891 | 0.77 |
ENSMUST00000121440.1
|
Maml3
|
mastermind like 3 (Drosophila) |
| chr15_-_50890396 | 0.77 |
ENSMUST00000185183.1
|
Trps1
|
trichorhinophalangeal syndrome I (human) |
| chr9_-_99710063 | 0.77 |
ENSMUST00000035048.5
|
Cldn18
|
claudin 18 |
| chr1_+_9848270 | 0.76 |
ENSMUST00000171265.1
|
Sgk3
|
serum/glucocorticoid regulated kinase 3 |
| chr6_+_134830145 | 0.76 |
ENSMUST00000046303.5
|
Crebl2
|
cAMP responsive element binding protein-like 2 |
| chr5_+_95956916 | 0.75 |
ENSMUST00000023840.5
|
Cxcl13
|
chemokine (C-X-C motif) ligand 13 |
| chr4_+_98546710 | 0.75 |
ENSMUST00000102792.3
|
Inadl
|
InaD-like (Drosophila) |
| chr4_+_43669610 | 0.75 |
ENSMUST00000107866.1
|
Tmem8b
|
transmembrane protein 8B |
| chr13_+_112467504 | 0.74 |
ENSMUST00000183868.1
|
Il6st
|
interleukin 6 signal transducer |
| chr7_+_78913436 | 0.74 |
ENSMUST00000121645.1
|
Isg20
|
interferon-stimulated protein |
| chr3_+_137867675 | 0.74 |
ENSMUST00000090178.5
|
Dnajb14
|
DnaJ (Hsp40) homolog, subfamily B, member 14 |
| chr17_+_46161111 | 0.74 |
ENSMUST00000166563.1
|
Gtpbp2
|
GTP binding protein 2 |
| chr16_-_26526744 | 0.73 |
ENSMUST00000165687.1
|
Tmem207
|
transmembrane protein 207 |
| chr13_-_12464925 | 0.73 |
ENSMUST00000124888.1
|
Lgals8
|
lectin, galactose binding, soluble 8 |
| chr1_+_9848375 | 0.71 |
ENSMUST00000097826.4
|
Sgk3
|
serum/glucocorticoid regulated kinase 3 |
| chr18_-_3281036 | 0.71 |
ENSMUST00000049942.6
ENSMUST00000139537.1 ENSMUST00000124747.1 |
Crem
|
cAMP responsive element modulator |
| chr15_+_5185519 | 0.70 |
ENSMUST00000118193.1
ENSMUST00000022751.8 |
Ttc33
|
tetratricopeptide repeat domain 33 |
| chr9_+_86743641 | 0.70 |
ENSMUST00000179574.1
|
Prss35
|
protease, serine, 35 |
| chr7_+_90426312 | 0.69 |
ENSMUST00000061391.7
|
Ccdc89
|
coiled-coil domain containing 89 |
| chrX_+_136707976 | 0.69 |
ENSMUST00000055104.5
|
Tceal1
|
transcription elongation factor A (SII)-like 1 |
| chr7_+_78913765 | 0.69 |
ENSMUST00000038142.8
|
Isg20
|
interferon-stimulated protein |
| chr8_-_45294854 | 0.69 |
ENSMUST00000116473.2
|
Klkb1
|
kallikrein B, plasma 1 |
| chr5_-_17888884 | 0.67 |
ENSMUST00000169095.1
|
Cd36
|
CD36 antigen |
| chr6_-_41446062 | 0.67 |
ENSMUST00000095999.5
|
Gm10334
|
predicted gene 10334 |
| chr16_+_24448082 | 0.67 |
ENSMUST00000078988.2
|
Lpp
|
LIM domain containing preferred translocation partner in lipoma |
| chr3_+_135825648 | 0.66 |
ENSMUST00000180196.1
|
Slc39a8
|
solute carrier family 39 (metal ion transporter), member 8 |
| chr7_+_43351378 | 0.66 |
ENSMUST00000012798.7
ENSMUST00000122423.1 ENSMUST00000121494.1 |
Siglec5
|
sialic acid binding Ig-like lectin 5 |
| chr10_+_52391606 | 0.66 |
ENSMUST00000067085.4
|
Nepn
|
nephrocan |
| chr8_+_45658731 | 0.65 |
ENSMUST00000143820.1
ENSMUST00000132139.1 |
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr1_-_126830632 | 0.65 |
ENSMUST00000112583.1
ENSMUST00000094609.3 |
Nckap5
|
NCK-associated protein 5 |
| chr9_+_43744399 | 0.65 |
ENSMUST00000034510.7
|
Pvrl1
|
poliovirus receptor-related 1 |
| chr4_+_98546919 | 0.65 |
ENSMUST00000030290.7
|
Inadl
|
InaD-like (Drosophila) |
| chr7_-_34655500 | 0.64 |
ENSMUST00000032709.1
|
Kctd15
|
potassium channel tetramerisation domain containing 15 |
| chr5_-_66151903 | 0.63 |
ENSMUST00000167950.1
|
Rbm47
|
RNA binding motif protein 47 |
| chr3_+_27154020 | 0.63 |
ENSMUST00000181124.1
|
1700125G22Rik
|
RIKEN cDNA 1700125G22 gene |
| chr5_+_107437908 | 0.62 |
ENSMUST00000094541.2
|
Btbd8
|
BTB (POZ) domain containing 8 |
| chr8_-_8639363 | 0.60 |
ENSMUST00000152698.1
|
Efnb2
|
ephrin B2 |
| chr2_-_90479165 | 0.60 |
ENSMUST00000111495.2
|
Ptprj
|
protein tyrosine phosphatase, receptor type, J |
| chr19_+_26749726 | 0.59 |
ENSMUST00000175842.1
|
Smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr7_-_142661858 | 0.58 |
ENSMUST00000145896.2
|
Igf2
|
insulin-like growth factor 2 |
| chr2_+_52072823 | 0.57 |
ENSMUST00000112693.2
ENSMUST00000069794.5 |
Rif1
|
Rap1 interacting factor 1 homolog (yeast) |
| chr2_-_18048784 | 0.57 |
ENSMUST00000142856.1
|
Skida1
|
SKI/DACH domain containing 1 |
| chr1_-_138619687 | 0.56 |
ENSMUST00000027642.2
|
Nek7
|
NIMA (never in mitosis gene a)-related expressed kinase 7 |
| chr14_-_121915774 | 0.56 |
ENSMUST00000055475.7
|
Gpr18
|
G protein-coupled receptor 18 |
| chr18_+_37742088 | 0.55 |
ENSMUST00000003599.6
|
Pcdhga9
|
protocadherin gamma subfamily A, 9 |
| chr3_-_131344892 | 0.55 |
ENSMUST00000090246.4
ENSMUST00000126569.1 |
Sgms2
|
sphingomyelin synthase 2 |
| chr8_-_85380964 | 0.55 |
ENSMUST00000122452.1
|
Mylk3
|
myosin light chain kinase 3 |
| chr5_-_66054499 | 0.54 |
ENSMUST00000145625.1
|
Rbm47
|
RNA binding motif protein 47 |
| chr1_+_43445736 | 0.54 |
ENSMUST00000086421.5
ENSMUST00000114744.1 |
Nck2
|
non-catalytic region of tyrosine kinase adaptor protein 2 |
| chr19_+_26623419 | 0.54 |
ENSMUST00000176584.1
|
Smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr2_-_66256576 | 0.54 |
ENSMUST00000125446.2
ENSMUST00000102718.3 |
Ttc21b
|
tetratricopeptide repeat domain 21B |
| chr12_+_119443410 | 0.54 |
ENSMUST00000048880.6
|
Macc1
|
metastasis associated in colon cancer 1 |
| chr19_+_42090422 | 0.53 |
ENSMUST00000066778.4
|
Pi4k2a
|
phosphatidylinositol 4-kinase type 2 alpha |
Network of associatons between targets according to the STRING database.
First level regulatory network of Foxa2_Foxa1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.5 | 37.2 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 1.7 | 11.6 | GO:0035948 | positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) |
| 1.3 | 5.0 | GO:0015801 | aromatic amino acid transport(GO:0015801) tryptophan transport(GO:0015827) |
| 1.1 | 3.3 | GO:0072277 | metanephric glomerulus morphogenesis(GO:0072275) metanephric glomerulus vasculature morphogenesis(GO:0072276) metanephric glomerular capillary formation(GO:0072277) |
| 1.1 | 5.3 | GO:2000660 | negative regulation of interleukin-1-mediated signaling pathway(GO:2000660) |
| 1.0 | 8.9 | GO:0008228 | opsonization(GO:0008228) |
| 0.9 | 3.8 | GO:0060005 | vestibular reflex(GO:0060005) |
| 0.8 | 2.5 | GO:1902524 | positive regulation of protein K48-linked ubiquitination(GO:1902524) |
| 0.7 | 2.2 | GO:1900135 | positive regulation of eosinophil degranulation(GO:0043311) regulation of neutrophil mediated cytotoxicity(GO:0070948) regulation of neutrophil mediated killing of symbiont cell(GO:0070949) positive regulation of renin secretion into blood stream(GO:1900135) positive regulation of eosinophil activation(GO:1902568) |
| 0.7 | 4.0 | GO:0042078 | germ-line stem cell division(GO:0042078) male germ-line stem cell asymmetric division(GO:0048133) germline stem cell asymmetric division(GO:0098728) |
| 0.6 | 2.4 | GO:0097647 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.6 | 3.5 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 0.6 | 5.0 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.5 | 1.9 | GO:1904566 | response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904565) cellular response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904566) |
| 0.5 | 2.3 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.4 | 5.8 | GO:0070842 | aggresome assembly(GO:0070842) |
| 0.4 | 1.2 | GO:0090004 | positive regulation of Golgi to plasma membrane protein transport(GO:0042998) positive regulation of establishment of protein localization to plasma membrane(GO:0090004) |
| 0.4 | 2.5 | GO:0060455 | negative regulation of gastric acid secretion(GO:0060455) |
| 0.4 | 1.2 | GO:0050925 | negative regulation of negative chemotaxis(GO:0050925) |
| 0.4 | 1.5 | GO:0072344 | rescue of stalled ribosome(GO:0072344) |
| 0.3 | 3.1 | GO:0044334 | regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010908) positive regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010909) regulation of gluconeogenesis by regulation of transcription from RNA polymerase II promoter(GO:0035947) canonical Wnt signaling pathway involved in positive regulation of epithelial to mesenchymal transition(GO:0044334) embryonic hindgut morphogenesis(GO:0048619) |
| 0.3 | 4.4 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.3 | 2.7 | GO:0015074 | DNA integration(GO:0015074) |
| 0.3 | 3.5 | GO:2001214 | positive regulation of vasculogenesis(GO:2001214) |
| 0.3 | 1.6 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 0.3 | 1.5 | GO:1901509 | regulation of endothelial tube morphogenesis(GO:1901509) |
| 0.3 | 1.8 | GO:0036438 | maintenance of lens transparency(GO:0036438) |
| 0.3 | 2.9 | GO:0015868 | purine ribonucleotide transport(GO:0015868) |
| 0.3 | 0.8 | GO:2001205 | negative regulation of osteoclast development(GO:2001205) |
| 0.2 | 0.7 | GO:0070104 | negative regulation of interleukin-6-mediated signaling pathway(GO:0070104) |
| 0.2 | 1.4 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 0.2 | 1.0 | GO:0042738 | exogenous drug catabolic process(GO:0042738) |
| 0.2 | 2.4 | GO:0044336 | canonical Wnt signaling pathway involved in negative regulation of apoptotic process(GO:0044336) |
| 0.2 | 2.3 | GO:0048208 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.2 | 0.9 | GO:0016332 | establishment or maintenance of polarity of embryonic epithelium(GO:0016332) |
| 0.2 | 0.7 | GO:0070543 | response to linoleic acid(GO:0070543) |
| 0.2 | 1.3 | GO:0003383 | apical constriction(GO:0003383) |
| 0.2 | 0.8 | GO:2000182 | regulation of progesterone biosynthetic process(GO:2000182) |
| 0.2 | 2.1 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.2 | 1.0 | GO:0009597 | detection of virus(GO:0009597) |
| 0.2 | 0.8 | GO:0046368 | GDP-L-fucose metabolic process(GO:0046368) |
| 0.2 | 0.8 | GO:0070358 | actin polymerization-dependent cell motility(GO:0070358) |
| 0.2 | 1.2 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 0.2 | 1.5 | GO:0051611 | regulation of serotonin uptake(GO:0051611) |
| 0.2 | 0.6 | GO:2001034 | positive regulation of double-strand break repair via nonhomologous end joining(GO:2001034) |
| 0.2 | 0.8 | GO:0033634 | positive regulation of cell-cell adhesion mediated by integrin(GO:0033634) negative regulation of endothelial cell chemotaxis(GO:2001027) |
| 0.2 | 8.7 | GO:0030195 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.2 | 0.6 | GO:0002305 | gamma-delta intraepithelial T cell differentiation(GO:0002304) CD8-positive, gamma-delta intraepithelial T cell differentiation(GO:0002305) |
| 0.2 | 0.5 | GO:0045575 | basophil activation involved in immune response(GO:0002276) basophil activation(GO:0045575) |
| 0.2 | 0.7 | GO:0071630 | nucleus-associated proteasomal ubiquitin-dependent protein catabolic process(GO:0071630) |
| 0.2 | 1.9 | GO:0060665 | regulation of branching involved in salivary gland morphogenesis by mesenchymal-epithelial signaling(GO:0060665) regulation of tau-protein kinase activity(GO:1902947) |
| 0.2 | 2.6 | GO:0042048 | olfactory behavior(GO:0042048) |
| 0.2 | 0.5 | GO:0061144 | epithelial cell maturation involved in prostate gland development(GO:0060743) alveolar secondary septum development(GO:0061144) |
| 0.2 | 0.8 | GO:0060023 | soft palate development(GO:0060023) |
| 0.2 | 0.3 | GO:1903972 | regulation of macrophage colony-stimulating factor signaling pathway(GO:1902226) regulation of response to macrophage colony-stimulating factor(GO:1903969) regulation of cellular response to macrophage colony-stimulating factor stimulus(GO:1903972) |
| 0.2 | 0.5 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.2 | 0.9 | GO:1903243 | negative regulation of tumor necrosis factor biosynthetic process(GO:0042536) negative regulation of cardiac muscle hypertrophy in response to stress(GO:1903243) |
| 0.2 | 1.5 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.1 | 0.9 | GO:2000675 | negative regulation of type B pancreatic cell apoptotic process(GO:2000675) |
| 0.1 | 1.0 | GO:1904220 | regulation of serine C-palmitoyltransferase activity(GO:1904220) |
| 0.1 | 1.2 | GO:0035887 | aortic smooth muscle cell differentiation(GO:0035887) |
| 0.1 | 0.5 | GO:0002528 | regulation of vascular permeability involved in acute inflammatory response(GO:0002528) |
| 0.1 | 0.5 | GO:1903898 | positive regulation of translation in response to endoplasmic reticulum stress(GO:0036493) negative regulation of PERK-mediated unfolded protein response(GO:1903898) |
| 0.1 | 0.8 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.1 | 0.7 | GO:0032915 | positive regulation of transforming growth factor beta2 production(GO:0032915) |
| 0.1 | 0.9 | GO:0038044 | transforming growth factor-beta secretion(GO:0038044) |
| 0.1 | 1.0 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.1 | 0.7 | GO:0002317 | plasma cell differentiation(GO:0002317) |
| 0.1 | 1.3 | GO:0032308 | positive regulation of prostaglandin secretion(GO:0032308) |
| 0.1 | 0.4 | GO:2000503 | positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 0.1 | 0.3 | GO:0071895 | negative regulation of interleukin-13 production(GO:0032696) odontoblast differentiation(GO:0071895) |
| 0.1 | 0.5 | GO:0014846 | esophagus smooth muscle contraction(GO:0014846) |
| 0.1 | 1.6 | GO:0010745 | negative regulation of macrophage derived foam cell differentiation(GO:0010745) |
| 0.1 | 1.0 | GO:0042473 | outer ear morphogenesis(GO:0042473) |
| 0.1 | 0.3 | GO:0002784 | positive regulation of antimicrobial peptide production(GO:0002225) positive regulation of antimicrobial humoral response(GO:0002760) regulation of antimicrobial peptide production(GO:0002784) regulation of antibacterial peptide production(GO:0002786) positive regulation of antibacterial peptide production(GO:0002803) |
| 0.1 | 2.1 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.1 | 1.4 | GO:0060346 | bone trabecula formation(GO:0060346) |
| 0.1 | 0.9 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.1 | 1.0 | GO:0061002 | negative regulation of dendritic spine morphogenesis(GO:0061002) |
| 0.1 | 1.3 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.1 | 0.6 | GO:1903849 | regulation of aorta morphogenesis(GO:1903847) positive regulation of aorta morphogenesis(GO:1903849) |
| 0.1 | 0.3 | GO:2000297 | negative regulation of synapse maturation(GO:2000297) |
| 0.1 | 0.3 | GO:0002370 | natural killer cell cytokine production(GO:0002370) regulation of natural killer cell cytokine production(GO:0002727) |
| 0.1 | 4.2 | GO:0046834 | lipid phosphorylation(GO:0046834) |
| 0.1 | 0.4 | GO:0046909 | intermembrane transport(GO:0046909) protein transport from ciliary membrane to plasma membrane(GO:1903445) |
| 0.1 | 0.5 | GO:0061343 | cell adhesion involved in heart morphogenesis(GO:0061343) |
| 0.1 | 0.3 | GO:0055130 | D-serine catabolic process(GO:0036088) D-alanine family amino acid metabolic process(GO:0046144) D-alanine metabolic process(GO:0046436) D-alanine catabolic process(GO:0055130) |
| 0.1 | 0.3 | GO:0010248 | establishment or maintenance of transmembrane electrochemical gradient(GO:0010248) |
| 0.1 | 0.8 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.1 | 0.3 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 0.1 | 0.2 | GO:1904882 | pre-B cell allelic exclusion(GO:0002331) signal transduction involved in G2 DNA damage checkpoint(GO:0072425) signal transduction involved in mitotic G2 DNA damage checkpoint(GO:0072434) telomerase catalytic core complex assembly(GO:1904868) regulation of telomerase catalytic core complex assembly(GO:1904882) positive regulation of telomerase catalytic core complex assembly(GO:1904884) |
| 0.1 | 0.5 | GO:0050847 | progesterone receptor signaling pathway(GO:0050847) |
| 0.1 | 0.2 | GO:0002182 | cytoplasmic translational elongation(GO:0002182) regulation of cytoplasmic translational elongation(GO:1900247) negative regulation of cytoplasmic translational elongation(GO:1900248) |
| 0.1 | 0.4 | GO:0021993 | initiation of neural tube closure(GO:0021993) |
| 0.1 | 0.6 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 0.1 | 2.0 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.1 | 0.2 | GO:1904798 | negative regulation of interleukin-8 biosynthetic process(GO:0045415) positive regulation of core promoter binding(GO:1904798) |
| 0.1 | 0.8 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.1 | 0.7 | GO:0015691 | cadmium ion transport(GO:0015691) cadmium ion transmembrane transport(GO:0070574) |
| 0.1 | 0.2 | GO:1900106 | hyaluranon cable assembly(GO:0036118) monocyte aggregation(GO:0070487) kidney mesenchyme morphogenesis(GO:0072131) metanephric mesenchyme morphogenesis(GO:0072133) regulation of hyaluranon cable assembly(GO:1900104) positive regulation of hyaluranon cable assembly(GO:1900106) |
| 0.1 | 0.7 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 0.1 | 0.4 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.1 | 1.1 | GO:1902260 | negative regulation of delayed rectifier potassium channel activity(GO:1902260) |
| 0.1 | 0.2 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.1 | 0.8 | GO:1990440 | positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.1 | 0.2 | GO:0097401 | synaptic vesicle lumen acidification(GO:0097401) |
| 0.1 | 0.3 | GO:2000271 | positive regulation of fibroblast apoptotic process(GO:2000271) |
| 0.1 | 0.2 | GO:0015851 | nucleobase transport(GO:0015851) pyrimidine nucleobase transport(GO:0015855) |
| 0.0 | 0.3 | GO:0038128 | ERBB2 signaling pathway(GO:0038128) |
| 0.0 | 0.4 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.0 | 1.6 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.0 | 1.1 | GO:0051156 | glucose 6-phosphate metabolic process(GO:0051156) |
| 0.0 | 1.2 | GO:0003215 | cardiac right ventricle morphogenesis(GO:0003215) |
| 0.0 | 0.3 | GO:0045925 | positive regulation of female receptivity(GO:0045925) |
| 0.0 | 0.2 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.0 | 1.0 | GO:0045793 | positive regulation of cell size(GO:0045793) |
| 0.0 | 0.2 | GO:0061589 | calcium activated phosphatidylserine scrambling(GO:0061589) |
| 0.0 | 1.7 | GO:0060706 | cell differentiation involved in embryonic placenta development(GO:0060706) |
| 0.0 | 1.5 | GO:2001240 | negative regulation of signal transduction in absence of ligand(GO:1901099) negative regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001240) |
| 0.0 | 0.5 | GO:0051132 | NK T cell activation(GO:0051132) |
| 0.0 | 0.2 | GO:1904073 | regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 0.0 | 0.6 | GO:0043116 | negative regulation of vascular permeability(GO:0043116) |
| 0.0 | 4.3 | GO:0009166 | nucleotide catabolic process(GO:0009166) |
| 0.0 | 0.4 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.0 | 0.4 | GO:0001696 | gastric acid secretion(GO:0001696) |
| 0.0 | 0.5 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.0 | 0.5 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.0 | 0.4 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.0 | 0.3 | GO:0021559 | trigeminal nerve development(GO:0021559) |
| 0.0 | 0.3 | GO:0031179 | peptide modification(GO:0031179) |
| 0.0 | 0.6 | GO:1902043 | positive regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902043) |
| 0.0 | 0.1 | GO:0006447 | regulation of translational initiation by iron(GO:0006447) positive regulation of translational initiation by iron(GO:0045994) |
| 0.0 | 0.5 | GO:0044458 | motile cilium assembly(GO:0044458) |
| 0.0 | 0.2 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.0 | 0.3 | GO:0072189 | ureter development(GO:0072189) |
| 0.0 | 0.1 | GO:0010735 | positive regulation of transcription via serum response element binding(GO:0010735) |
| 0.0 | 0.2 | GO:0071280 | cellular response to copper ion(GO:0071280) |
| 0.0 | 0.0 | GO:0061030 | epithelial cell differentiation involved in mammary gland alveolus development(GO:0061030) |
| 0.0 | 0.5 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.0 | 0.2 | GO:0007343 | egg activation(GO:0007343) |
| 0.0 | 1.1 | GO:0045600 | positive regulation of fat cell differentiation(GO:0045600) |
| 0.0 | 0.1 | GO:0046116 | queuosine biosynthetic process(GO:0008616) queuosine metabolic process(GO:0046116) |
| 0.0 | 0.5 | GO:0002011 | morphogenesis of an epithelial sheet(GO:0002011) |
| 0.0 | 0.2 | GO:0015705 | thyroid hormone generation(GO:0006590) iodide transport(GO:0015705) |
| 0.0 | 0.2 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.0 | 1.6 | GO:0030216 | keratinocyte differentiation(GO:0030216) |
| 0.0 | 0.1 | GO:0043470 | regulation of carbohydrate catabolic process(GO:0043470) regulation of cellular carbohydrate catabolic process(GO:0043471) |
| 0.0 | 0.2 | GO:0002051 | osteoblast fate commitment(GO:0002051) |
| 0.0 | 0.1 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.0 | 2.5 | GO:0007596 | blood coagulation(GO:0007596) |
| 0.0 | 0.2 | GO:0048757 | endosome to melanosome transport(GO:0035646) endosome to pigment granule transport(GO:0043485) pigment granule maturation(GO:0048757) |
| 0.0 | 0.1 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.0 | 0.4 | GO:0044062 | regulation of excretion(GO:0044062) |
| 0.0 | 0.1 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 0.0 | 0.5 | GO:0007588 | excretion(GO:0007588) |
| 0.0 | 0.2 | GO:0086069 | bundle of His cell to Purkinje myocyte communication(GO:0086069) |
| 0.0 | 0.2 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.0 | 0.2 | GO:0031069 | hair follicle morphogenesis(GO:0031069) |
| 0.0 | 0.2 | GO:0016188 | synaptic vesicle maturation(GO:0016188) |
| 0.0 | 1.9 | GO:0042060 | wound healing(GO:0042060) |
| 0.0 | 1.0 | GO:0007006 | mitochondrial membrane organization(GO:0007006) |
| 0.0 | 0.2 | GO:0097094 | craniofacial suture morphogenesis(GO:0097094) |
| 0.0 | 0.7 | GO:0051591 | response to cAMP(GO:0051591) |
| 0.0 | 0.1 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.0 | 1.0 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.0 | 1.4 | GO:0051028 | mRNA transport(GO:0051028) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.7 | 37.1 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.5 | 4.1 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.5 | 3.5 | GO:0031466 | Cul5-RING ubiquitin ligase complex(GO:0031466) |
| 0.3 | 2.1 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.3 | 3.1 | GO:0070369 | beta-catenin-TCF7L2 complex(GO:0070369) |
| 0.3 | 0.9 | GO:0034685 | integrin alphav-beta6 complex(GO:0034685) |
| 0.3 | 5.8 | GO:0005922 | connexon complex(GO:0005922) |
| 0.2 | 0.7 | GO:0005900 | oncostatin-M receptor complex(GO:0005900) |
| 0.2 | 8.9 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.2 | 0.7 | GO:0044218 | other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 0.2 | 1.7 | GO:1990357 | terminal web(GO:1990357) |
| 0.2 | 0.3 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.2 | 4.4 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.2 | 1.3 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.1 | 0.9 | GO:0031211 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.1 | 2.2 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.1 | 2.5 | GO:0045095 | keratin filament(GO:0045095) |
| 0.1 | 0.6 | GO:0001940 | male pronucleus(GO:0001940) |
| 0.1 | 1.2 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.1 | 0.2 | GO:0016939 | kinesin II complex(GO:0016939) |
| 0.1 | 0.4 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.1 | 0.5 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.1 | 0.8 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.1 | 0.5 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) |
| 0.0 | 0.8 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 0.3 | GO:0001652 | granular component(GO:0001652) |
| 0.0 | 0.2 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 0.0 | 0.5 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 2.0 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 0.8 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 1.0 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 0.9 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.0 | 0.4 | GO:0005921 | gap junction(GO:0005921) |
| 0.0 | 0.5 | GO:0044292 | dendrite terminus(GO:0044292) |
| 0.0 | 3.6 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 0.2 | GO:0016035 | zeta DNA polymerase complex(GO:0016035) |
| 0.0 | 1.1 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.0 | 0.4 | GO:0036057 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.0 | 0.4 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.0 | 1.1 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 1.0 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 18.8 | GO:0009986 | cell surface(GO:0009986) |
| 0.0 | 0.9 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.0 | 0.1 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.0 | 0.7 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 1.2 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 0.6 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 0.2 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.0 | 0.2 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 5.3 | GO:0045352 | interleukin-1, Type II receptor binding(GO:0005151) interleukin-1 receptor antagonist activity(GO:0005152) interleukin-1 Type I receptor antagonist activity(GO:0045352) interleukin-1 Type II receptor antagonist activity(GO:0045353) |
| 1.3 | 5.0 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 1.0 | 6.7 | GO:0004556 | alpha-amylase activity(GO:0004556) |
| 0.8 | 4.1 | GO:0004850 | uridine phosphorylase activity(GO:0004850) |
| 0.7 | 5.0 | GO:0008241 | peptidyl-dipeptidase activity(GO:0008241) |
| 0.7 | 3.5 | GO:0004925 | prolactin receptor activity(GO:0004925) |
| 0.7 | 3.3 | GO:0038085 | vascular endothelial growth factor binding(GO:0038085) |
| 0.6 | 2.3 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 0.6 | 3.9 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.5 | 2.2 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.4 | 1.8 | GO:0008109 | N-acetyllactosaminide beta-1,6-N-acetylglucosaminyltransferase activity(GO:0008109) |
| 0.4 | 1.1 | GO:0004346 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 0.3 | 1.0 | GO:0005118 | sevenless binding(GO:0005118) |
| 0.3 | 3.6 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.2 | 0.7 | GO:0019981 | interleukin-6 receptor activity(GO:0004915) leukemia inhibitory factor receptor activity(GO:0004923) interleukin-6 binding(GO:0019981) |
| 0.2 | 0.9 | GO:0004758 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.2 | 7.7 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.2 | 0.7 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 0.2 | 0.9 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.2 | 5.5 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.2 | 1.6 | GO:0071253 | connexin binding(GO:0071253) |
| 0.2 | 0.8 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.2 | 2.8 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.2 | 4.0 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.2 | 2.9 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.2 | 0.5 | GO:0035651 | AP-3 adaptor complex binding(GO:0035651) |
| 0.1 | 1.0 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.1 | 0.8 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.1 | 4.4 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.1 | 1.2 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.1 | 0.4 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.1 | 11.3 | GO:0005179 | hormone activity(GO:0005179) |
| 0.1 | 0.6 | GO:0033188 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.1 | 0.5 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.1 | 0.3 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 0.1 | 3.0 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.1 | 0.5 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.1 | 0.5 | GO:0030942 | endoplasmic reticulum signal peptide binding(GO:0030942) |
| 0.1 | 1.4 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.1 | 0.3 | GO:0016715 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced ascorbate as one donor, and incorporation of one atom of oxygen(GO:0016715) |
| 0.1 | 0.5 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.1 | 2.1 | GO:0005521 | lamin binding(GO:0005521) |
| 0.1 | 1.1 | GO:0015194 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.1 | 10.3 | GO:0005178 | integrin binding(GO:0005178) |
| 0.1 | 1.5 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.1 | 0.8 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.1 | 0.2 | GO:0004677 | DNA-dependent protein kinase activity(GO:0004677) |
| 0.1 | 0.8 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.1 | 0.3 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.1 | 1.9 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.1 | 0.5 | GO:0005534 | galactose binding(GO:0005534) |
| 0.1 | 0.3 | GO:0003884 | D-amino-acid oxidase activity(GO:0003884) |
| 0.1 | 0.8 | GO:0008417 | fucosyltransferase activity(GO:0008417) |
| 0.1 | 0.2 | GO:0004947 | bradykinin receptor activity(GO:0004947) |
| 0.1 | 1.3 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.1 | 2.7 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.6 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.0 | 0.3 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 1.4 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.0 | 0.3 | GO:0033142 | progesterone receptor binding(GO:0033142) |
| 0.0 | 1.3 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.0 | 0.2 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.0 | 0.9 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 0.2 | GO:0001010 | transcription factor activity, sequence-specific DNA binding transcription factor recruiting(GO:0001010) |
| 0.0 | 0.2 | GO:0005350 | purine nucleobase transmembrane transporter activity(GO:0005345) pyrimidine nucleobase transmembrane transporter activity(GO:0005350) nucleobase transmembrane transporter activity(GO:0015205) glycerol channel activity(GO:0015254) |
| 0.0 | 0.9 | GO:0017081 | chloride channel regulator activity(GO:0017081) |
| 0.0 | 1.8 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.2 | GO:0031013 | troponin I binding(GO:0031013) |
| 0.0 | 0.2 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.0 | 1.6 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.0 | 0.4 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.0 | 0.4 | GO:0019865 | immunoglobulin binding(GO:0019865) |
| 0.0 | 1.5 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.1 | GO:0008479 | queuine tRNA-ribosyltransferase activity(GO:0008479) |
| 0.0 | 0.2 | GO:0015379 | potassium:chloride symporter activity(GO:0015379) potassium ion symporter activity(GO:0022820) |
| 0.0 | 1.3 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.0 | 0.3 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.0 | 1.2 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.8 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.0 | 1.4 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.0 | 1.0 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.2 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.0 | 0.7 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 28.5 | GO:0032403 | protein complex binding(GO:0032403) |
| 0.0 | 0.2 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 0.0 | 0.7 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.0 | 2.1 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 0.4 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.0 | 0.2 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.0 | 0.8 | GO:0048365 | Rac GTPase binding(GO:0048365) |
| 0.0 | 0.7 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 0.4 | GO:0051393 | alpha-actinin binding(GO:0051393) |
| 0.0 | 0.2 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.3 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.0 | 0.1 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.0 | 0.1 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.0 | 0.2 | GO:0004629 | phosphatidylinositol phospholipase C activity(GO:0004435) phospholipase C activity(GO:0004629) |
| 0.0 | 0.1 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.1 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.0 | 0.2 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 20.1 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 7.9 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.1 | 5.6 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 3.3 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 6.0 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.1 | 2.2 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.1 | 2.9 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 3.5 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.1 | 0.4 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.1 | 1.9 | PID TCPTP PATHWAY | Signaling events mediated by TCPTP |
| 0.0 | 0.9 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 0.8 | PID IGF1 PATHWAY | IGF1 pathway |
| 0.0 | 0.7 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.0 | 0.8 | PID GMCSF PATHWAY | GMCSF-mediated signaling events |
| 0.0 | 0.6 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.0 | 2.2 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 0.8 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.0 | 1.0 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.0 | 1.3 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 0.5 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.0 | 0.6 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.0 | 0.4 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.0 | 1.6 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 0.9 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 1.4 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.0 | 0.8 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.0 | 0.7 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 1.0 | PID FGF PATHWAY | FGF signaling pathway |
| 0.0 | 0.8 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 0.7 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 0.2 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.0 | 0.3 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.0 | 0.2 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 0.3 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.0 | 0.4 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 35.9 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.7 | 11.8 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.3 | 5.5 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.2 | 4.1 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.2 | 3.5 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.2 | 1.9 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.2 | 2.0 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.1 | 4.8 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.1 | 0.8 | REACTOME JNK C JUN KINASES PHOSPHORYLATION AND ACTIVATION MEDIATED BY ACTIVATED HUMAN TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 0.1 | 0.8 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.1 | 4.0 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.1 | 3.5 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.1 | 0.7 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.1 | 2.8 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.1 | 1.2 | REACTOME SIGNALING BY ACTIVATED POINT MUTANTS OF FGFR1 | Genes involved in Signaling by activated point mutants of FGFR1 |
| 0.1 | 1.5 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.1 | 3.4 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 0.7 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.0 | 0.8 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.0 | 1.6 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 2.3 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 1.9 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.0 | 1.7 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 1.2 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 0.7 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 1.0 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 0.4 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.0 | 0.7 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 0.8 | REACTOME N GLYCAN ANTENNAE ELONGATION IN THE MEDIAL TRANS GOLGI | Genes involved in N-glycan antennae elongation in the medial/trans-Golgi |
| 0.0 | 2.2 | REACTOME DOWNSTREAM SIGNAL TRANSDUCTION | Genes involved in Downstream signal transduction |
| 0.0 | 0.4 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 0.2 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.0 | 4.8 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.4 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 0.9 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 0.9 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.6 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.2 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.0 | 0.1 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.0 | 0.5 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.0 | 0.4 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 1.4 | REACTOME DIABETES PATHWAYS | Genes involved in Diabetes pathways |
| 0.0 | 3.7 | REACTOME CLASS I MHC MEDIATED ANTIGEN PROCESSING PRESENTATION | Genes involved in Class I MHC mediated antigen processing & presentation |
| 0.0 | 0.6 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 0.6 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 2.3 | REACTOME CLASS A1 RHODOPSIN LIKE RECEPTORS | Genes involved in Class A/1 (Rhodopsin-like receptors) |
| 0.0 | 0.1 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |