Project
avrg: 12D miR HR13_24
Navigation
Downloads
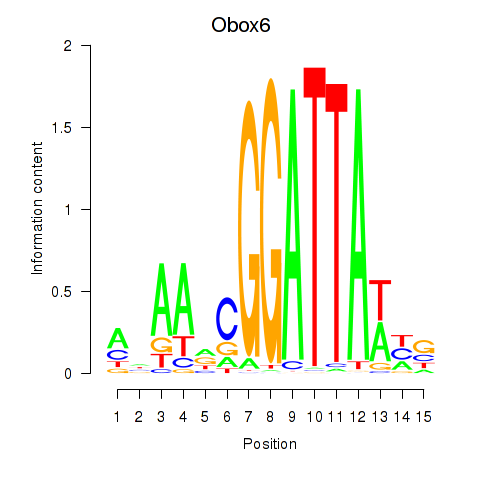
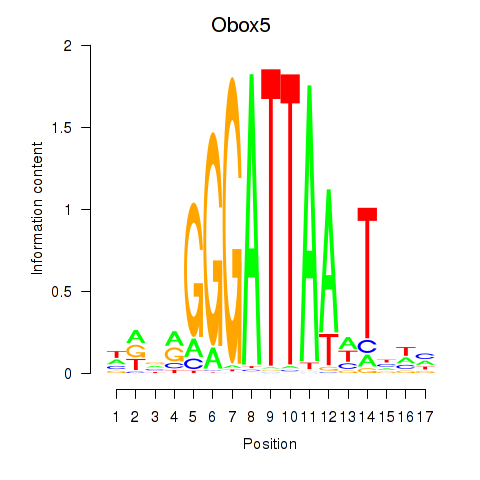
Results for Obox6_Obox5
Z-value: 2.42
Transcription factors associated with Obox6_Obox5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Obox6
|
ENSMUSG00000041583.7 | oocyte specific homeobox 6 |
|
Obox5
|
ENSMUSG00000074366.3 | oocyte specific homeobox 5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Obox6 | mm10_v2_chr7_-_15839678_15839678 | 0.31 | 3.6e-01 | Click! |
Activity profile of Obox6_Obox5 motif
Sorted Z-values of Obox6_Obox5 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_-_4752972 | 4.35 |
ENSMUST00000183971.1
ENSMUST00000182173.1 ENSMUST00000182738.1 ENSMUST00000184143.1 ENSMUST00000182111.1 ENSMUST00000182048.1 ENSMUST00000063324.7 |
Cox6b2
|
cytochrome c oxidase subunit VIb polypeptide 2 |
| chr16_-_63864114 | 3.15 |
ENSMUST00000064405.6
|
Epha3
|
Eph receptor A3 |
| chr2_+_181680284 | 3.09 |
ENSMUST00000103042.3
|
Tcea2
|
transcription elongation factor A (SII), 2 |
| chrX_-_51681856 | 2.97 |
ENSMUST00000114871.1
|
Hs6st2
|
heparan sulfate 6-O-sulfotransferase 2 |
| chrX_-_51681703 | 2.89 |
ENSMUST00000088172.5
|
Hs6st2
|
heparan sulfate 6-O-sulfotransferase 2 |
| chr4_+_141368116 | 2.65 |
ENSMUST00000006380.4
|
Fam131c
|
family with sequence similarity 131, member C |
| chr3_-_73708399 | 2.55 |
ENSMUST00000029367.5
|
Bche
|
butyrylcholinesterase |
| chr10_+_94198955 | 2.55 |
ENSMUST00000020209.9
ENSMUST00000179990.1 |
Ndufa12
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 12 |
| chr13_-_100786402 | 2.39 |
ENSMUST00000174038.1
ENSMUST00000091295.7 ENSMUST00000072119.8 |
Ccnb1
|
cyclin B1 |
| chrX_-_61185558 | 2.26 |
ENSMUST00000166381.1
|
Cdr1
|
cerebellar degeneration related antigen 1 |
| chr14_+_64588112 | 2.12 |
ENSMUST00000181808.1
|
A930011O12Rik
|
RIKEN cDNA A930011O12 gene |
| chr8_-_111992258 | 2.06 |
ENSMUST00000034427.5
ENSMUST00000139820.1 |
Adat1
|
adenosine deaminase, tRNA-specific 1 |
| chr6_+_113531675 | 2.01 |
ENSMUST00000036340.5
ENSMUST00000101051.2 |
Fancd2
|
Fanconi anemia, complementation group D2 |
| chr16_-_23127702 | 1.94 |
ENSMUST00000115338.1
ENSMUST00000115337.1 ENSMUST00000023598.8 |
Rfc4
|
replication factor C (activator 1) 4 |
| chr1_-_189688074 | 1.92 |
ENSMUST00000171929.1
ENSMUST00000165962.1 |
Cenpf
|
centromere protein F |
| chr15_+_93398344 | 1.77 |
ENSMUST00000109256.3
ENSMUST00000068457.7 ENSMUST00000049122.8 ENSMUST00000165935.1 |
Pphln1
|
periphilin 1 |
| chr7_-_99182681 | 1.66 |
ENSMUST00000033001.4
|
Dgat2
|
diacylglycerol O-acyltransferase 2 |
| chrX_+_20549780 | 1.60 |
ENSMUST00000023832.6
|
Rgn
|
regucalcin |
| chr3_+_145576196 | 1.58 |
ENSMUST00000098534.4
|
Znhit6
|
zinc finger, HIT type 6 |
| chr5_-_135251209 | 1.46 |
ENSMUST00000062572.2
|
Fzd9
|
frizzled homolog 9 (Drosophila) |
| chr7_-_45179539 | 1.46 |
ENSMUST00000179443.1
|
Gm581
|
predicted gene 581 |
| chr1_+_53061637 | 1.45 |
ENSMUST00000027269.5
|
Mstn
|
myostatin |
| chr7_+_16781341 | 1.42 |
ENSMUST00000108496.2
|
Slc1a5
|
solute carrier family 1 (neutral amino acid transporter), member 5 |
| chr15_-_101562889 | 1.41 |
ENSMUST00000023714.4
|
4732456N10Rik
|
RIKEN cDNA 4732456N10 gene |
| chr14_-_87141114 | 1.41 |
ENSMUST00000168889.1
|
Diap3
|
diaphanous homolog 3 (Drosophila) |
| chr14_-_87141206 | 1.38 |
ENSMUST00000022599.7
|
Diap3
|
diaphanous homolog 3 (Drosophila) |
| chrX_-_150657392 | 1.37 |
ENSMUST00000151403.2
ENSMUST00000087253.4 ENSMUST00000112709.1 ENSMUST00000163969.1 ENSMUST00000087258.3 |
Tro
|
trophinin |
| chr10_+_18407658 | 1.35 |
ENSMUST00000037341.7
|
Nhsl1
|
NHS-like 1 |
| chr8_-_4779513 | 1.33 |
ENSMUST00000022945.7
|
Shcbp1
|
Shc SH2-domain binding protein 1 |
| chr2_+_91922178 | 1.29 |
ENSMUST00000170432.1
|
Chrm4
|
cholinergic receptor, muscarinic 4 |
| chr8_-_13254154 | 1.28 |
ENSMUST00000033825.4
|
Adprhl1
|
ADP-ribosylhydrolase like 1 |
| chr3_-_63899437 | 1.27 |
ENSMUST00000159188.1
ENSMUST00000177143.1 |
Plch1
|
phospholipase C, eta 1 |
| chr17_-_25797032 | 1.18 |
ENSMUST00000165838.1
ENSMUST00000002344.6 |
Metrn
|
meteorin, glial cell differentiation regulator |
| chr4_+_134510999 | 1.17 |
ENSMUST00000105866.2
|
Aunip
|
aurora kinase A and ninein interacting protein |
| chr16_+_31422268 | 1.17 |
ENSMUST00000089759.2
|
Bdh1
|
3-hydroxybutyrate dehydrogenase, type 1 |
| chr6_+_119236507 | 1.16 |
ENSMUST00000037434.6
|
Cacna2d4
|
calcium channel, voltage-dependent, alpha 2/delta subunit 4 |
| chr8_-_13254068 | 1.16 |
ENSMUST00000168498.1
|
Adprhl1
|
ADP-ribosylhydrolase like 1 |
| chr19_+_44992127 | 1.16 |
ENSMUST00000179305.1
|
Sema4g
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4G |
| chr6_-_35308110 | 1.10 |
ENSMUST00000031868.4
|
Slc13a4
|
solute carrier family 13 (sodium/sulfate symporters), member 4 |
| chr2_-_110950923 | 1.08 |
ENSMUST00000099623.3
|
Ano3
|
anoctamin 3 |
| chr5_+_76656512 | 1.07 |
ENSMUST00000086909.4
|
Gm10430
|
predicted gene 10430 |
| chr8_-_123318553 | 1.07 |
ENSMUST00000118395.1
ENSMUST00000035495.8 |
Fanca
|
Fanconi anemia, complementation group A |
| chr5_-_150518164 | 1.07 |
ENSMUST00000118769.1
|
Zar1l
|
zygote arrest 1-like |
| chr11_+_3963970 | 1.06 |
ENSMUST00000020705.4
ENSMUST00000109985.1 |
Pes1
|
pescadillo homolog 1, containing BRCT domain (zebrafish) |
| chr8_-_70523085 | 1.04 |
ENSMUST00000137610.1
ENSMUST00000121623.1 ENSMUST00000093456.5 ENSMUST00000118850.1 |
Kxd1
|
KxDL motif containing 1 |
| chr3_-_123236134 | 1.03 |
ENSMUST00000106427.1
ENSMUST00000106426.1 ENSMUST00000051443.5 |
Synpo2
|
synaptopodin 2 |
| chr1_+_85575676 | 1.02 |
ENSMUST00000178024.1
|
G530012D18Rik
|
RIKEN cDNA G530012D1 gene |
| chr5_-_92435219 | 1.01 |
ENSMUST00000038514.8
|
Nup54
|
nucleoporin 54 |
| chr2_+_11705355 | 1.01 |
ENSMUST00000128156.2
|
Il15ra
|
interleukin 15 receptor, alpha chain |
| chr9_-_50617428 | 0.99 |
ENSMUST00000131351.1
ENSMUST00000171462.1 |
AU019823
|
expressed sequence AU019823 |
| chr2_+_121358591 | 0.98 |
ENSMUST00000000317.6
ENSMUST00000129130.1 |
Ckmt1
|
creatine kinase, mitochondrial 1, ubiquitous |
| chr16_+_55966275 | 0.97 |
ENSMUST00000023269.4
|
RPL24
|
60S ribosomal protein L24 |
| chr19_-_10101501 | 0.96 |
ENSMUST00000025567.7
|
Fads2
|
fatty acid desaturase 2 |
| chr11_-_78183551 | 0.96 |
ENSMUST00000102483.4
|
Rpl23a
|
ribosomal protein L23A |
| chr4_-_148130678 | 0.96 |
ENSMUST00000030862.4
|
Draxin
|
dorsal inhibitory axon guidance protein |
| chr1_-_71103146 | 0.94 |
ENSMUST00000027393.7
|
Bard1
|
BRCA1 associated RING domain 1 |
| chr2_+_72476159 | 0.92 |
ENSMUST00000102691.4
|
Cdca7
|
cell division cycle associated 7 |
| chr13_+_55399648 | 0.91 |
ENSMUST00000057167.7
|
Slc34a1
|
solute carrier family 34 (sodium phosphate), member 1 |
| chr19_-_41802028 | 0.91 |
ENSMUST00000026150.8
ENSMUST00000177495.1 ENSMUST00000163265.1 |
Arhgap19
|
Rho GTPase activating protein 19 |
| chr1_-_10038106 | 0.90 |
ENSMUST00000027050.3
|
Cops5
|
COP9 (constitutive photomorphogenic) homolog, subunit 5 (Arabidopsis thaliana) |
| chr10_+_57784914 | 0.90 |
ENSMUST00000165013.1
|
Fabp7
|
fatty acid binding protein 7, brain |
| chr11_-_87108656 | 0.89 |
ENSMUST00000051395.8
|
Prr11
|
proline rich 11 |
| chr8_+_123411424 | 0.89 |
ENSMUST00000071134.3
|
Tubb3
|
tubulin, beta 3 class III |
| chr11_-_86257553 | 0.89 |
ENSMUST00000132024.1
ENSMUST00000139285.1 |
Ints2
|
integrator complex subunit 2 |
| chr16_+_22920222 | 0.88 |
ENSMUST00000023587.4
ENSMUST00000116625.2 |
Fetub
|
fetuin beta |
| chr14_-_8172986 | 0.86 |
ENSMUST00000022268.8
|
Pdhb
|
pyruvate dehydrogenase (lipoamide) beta |
| chr5_+_110330697 | 0.86 |
ENSMUST00000112481.1
|
Pole
|
polymerase (DNA directed), epsilon |
| chr11_+_101330605 | 0.86 |
ENSMUST00000103105.3
|
Aoc3
|
amine oxidase, copper containing 3 |
| chrX_+_169685191 | 0.84 |
ENSMUST00000112104.1
ENSMUST00000112107.1 |
Mid1
|
midline 1 |
| chr1_+_135232045 | 0.84 |
ENSMUST00000110798.3
|
Gm4204
|
predicted gene 4204 |
| chr11_-_86257518 | 0.84 |
ENSMUST00000136469.1
ENSMUST00000018212.6 |
Ints2
|
integrator complex subunit 2 |
| chr9_-_50617228 | 0.82 |
ENSMUST00000147671.1
ENSMUST00000145139.1 ENSMUST00000155435.1 |
AU019823
|
expressed sequence AU019823 |
| chr15_-_83122756 | 0.81 |
ENSMUST00000018184.3
|
Rrp7a
|
ribosomal RNA processing 7 homolog A (S. cerevisiae) |
| chr16_-_46010212 | 0.80 |
ENSMUST00000130481.1
|
Plcxd2
|
phosphatidylinositol-specific phospholipase C, X domain containing 2 |
| chr8_-_95306585 | 0.80 |
ENSMUST00000119870.2
ENSMUST00000093268.4 |
Cngb1
|
cyclic nucleotide gated channel beta 1 |
| chr4_+_131843459 | 0.80 |
ENSMUST00000030742.4
ENSMUST00000137321.1 |
Mecr
|
mitochondrial trans-2-enoyl-CoA reductase |
| chr13_-_58354862 | 0.80 |
ENSMUST00000043605.5
|
Kif27
|
kinesin family member 27 |
| chr10_-_117063764 | 0.80 |
ENSMUST00000047672.7
|
Cct2
|
chaperonin containing Tcp1, subunit 2 (beta) |
| chr9_+_88581036 | 0.79 |
ENSMUST00000164661.2
|
Trim43a
|
tripartite motif-containing 43A |
| chr18_+_4920509 | 0.78 |
ENSMUST00000126977.1
|
Svil
|
supervillin |
| chr7_+_5056706 | 0.78 |
ENSMUST00000144802.1
|
Ccdc106
|
coiled-coil domain containing 106 |
| chr11_-_120796369 | 0.77 |
ENSMUST00000143139.1
ENSMUST00000129955.1 ENSMUST00000026151.4 ENSMUST00000167023.1 ENSMUST00000106133.1 ENSMUST00000106135.1 |
Dus1l
|
dihydrouridine synthase 1-like (S. cerevisiae) |
| chr7_-_7247328 | 0.77 |
ENSMUST00000170922.1
|
Vmn2r29
|
vomeronasal 2, receptor 29 |
| chr4_-_116167591 | 0.77 |
ENSMUST00000030465.3
ENSMUST00000143426.1 |
Tspan1
|
tetraspanin 1 |
| chr2_-_14056029 | 0.76 |
ENSMUST00000074854.7
|
Ptpla
|
protein tyrosine phosphatase-like (proline instead of catalytic arginine), member a |
| chr4_+_74013442 | 0.76 |
ENSMUST00000098006.2
ENSMUST00000084474.5 |
Frmd3
|
FERM domain containing 3 |
| chr8_-_107403197 | 0.76 |
ENSMUST00000003947.8
|
Nqo1
|
NAD(P)H dehydrogenase, quinone 1 |
| chr8_-_13254096 | 0.76 |
ENSMUST00000171619.1
|
Adprhl1
|
ADP-ribosylhydrolase like 1 |
| chr7_-_80403315 | 0.75 |
ENSMUST00000147150.1
|
Furin
|
furin (paired basic amino acid cleaving enzyme) |
| chr3_+_104781048 | 0.75 |
ENSMUST00000002298.6
|
Ppm1j
|
protein phosphatase 1J |
| chr4_-_137785371 | 0.74 |
ENSMUST00000133473.1
|
Alpl
|
alkaline phosphatase, liver/bone/kidney |
| chr4_+_85205120 | 0.73 |
ENSMUST00000107188.3
|
Sh3gl2
|
SH3-domain GRB2-like 2 |
| chr11_-_100472725 | 0.71 |
ENSMUST00000056665.3
|
Klhl11
|
kelch-like 11 |
| chr4_+_9269285 | 0.71 |
ENSMUST00000038841.7
|
Clvs1
|
clavesin 1 |
| chr4_+_155451570 | 0.71 |
ENSMUST00000135407.1
ENSMUST00000105619.1 |
C030017K20Rik
|
RIKEN cDNA C030017K20 gene |
| chr2_+_152962485 | 0.70 |
ENSMUST00000099197.2
ENSMUST00000103155.3 |
Ttll9
|
tubulin tyrosine ligase-like family, member 9 |
| chr3_+_84952146 | 0.69 |
ENSMUST00000029727.7
|
Fbxw7
|
F-box and WD-40 domain protein 7 |
| chr1_-_173599074 | 0.68 |
ENSMUST00000150649.1
ENSMUST00000180215.1 ENSMUST00000097462.2 |
Pydc4
|
pyrin domain containing 4 |
| chrX_-_85776606 | 0.68 |
ENSMUST00000142152.1
ENSMUST00000156390.1 ENSMUST00000113978.2 |
Gyk
|
glycerol kinase |
| chr1_+_118321834 | 0.68 |
ENSMUST00000027626.6
ENSMUST00000112688.3 |
Mki67ip
|
Mki67 (FHA domain) interacting nucleolar phosphoprotein |
| chr18_+_80206775 | 0.67 |
ENSMUST00000145963.1
ENSMUST00000025464.7 ENSMUST00000125127.1 ENSMUST00000025463.7 |
Txnl4a
Gm16286
|
thioredoxin-like 4A predicted gene 16286 |
| chr13_+_99344775 | 0.66 |
ENSMUST00000052249.5
|
Mrps27
|
mitochondrial ribosomal protein S27 |
| chr5_+_124552845 | 0.66 |
ENSMUST00000071057.7
|
Ddx55
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 55 |
| chr12_-_87444017 | 0.65 |
ENSMUST00000091090.4
|
2700073G19Rik
|
RIKEN cDNA 2700073G19 gene |
| chr9_+_108479849 | 0.64 |
ENSMUST00000065014.4
|
Lamb2
|
laminin, beta 2 |
| chr1_-_181144133 | 0.64 |
ENSMUST00000027797.7
|
Nvl
|
nuclear VCP-like |
| chr2_+_72297895 | 0.63 |
ENSMUST00000144111.1
|
Zak
|
sterile alpha motif and leucine zipper containing kinase AZK |
| chr8_+_45885479 | 0.63 |
ENSMUST00000034053.5
|
Pdlim3
|
PDZ and LIM domain 3 |
| chr4_-_88033328 | 0.63 |
ENSMUST00000078090.5
|
Mllt3
|
myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 3 |
| chr7_+_48789003 | 0.63 |
ENSMUST00000118927.1
ENSMUST00000125280.1 |
Zdhhc13
|
zinc finger, DHHC domain containing 13 |
| chr2_-_130629994 | 0.62 |
ENSMUST00000110262.1
ENSMUST00000028761.4 |
Fastkd5
Ubox5
|
FAST kinase domains 5 U box domain containing 5 |
| chr14_-_19977040 | 0.61 |
ENSMUST00000159028.1
|
Gng2
|
guanine nucleotide binding protein (G protein), gamma 2 |
| chr14_+_61138445 | 0.61 |
ENSMUST00000089394.3
ENSMUST00000119509.1 |
Sacs
|
sacsin |
| chr11_-_70459957 | 0.61 |
ENSMUST00000019064.2
|
Cxcl16
|
chemokine (C-X-C motif) ligand 16 |
| chr9_+_50617516 | 0.60 |
ENSMUST00000141366.1
|
Pih1d2
|
PIH1 domain containing 2 |
| chr15_-_102004305 | 0.60 |
ENSMUST00000023952.8
|
Krt8
|
keratin 8 |
| chr6_+_35177386 | 0.60 |
ENSMUST00000043815.9
|
Nup205
|
nucleoporin 205 |
| chr16_+_59471775 | 0.58 |
ENSMUST00000023407.5
ENSMUST00000120667.1 ENSMUST00000120674.1 |
Mina
|
myc induced nuclear antigen |
| chr2_-_29787622 | 0.58 |
ENSMUST00000177467.1
ENSMUST00000113807.3 |
Trub2
|
TruB pseudouridine (psi) synthase homolog 2 (E. coli) |
| chr14_+_13454010 | 0.58 |
ENSMUST00000112656.2
|
Synpr
|
synaptoporin |
| chr11_+_78920787 | 0.57 |
ENSMUST00000018610.6
|
Nos2
|
nitric oxide synthase 2, inducible |
| chr5_-_137613759 | 0.57 |
ENSMUST00000155251.1
ENSMUST00000124693.1 |
Pcolce
|
procollagen C-endopeptidase enhancer protein |
| chr4_+_48585193 | 0.57 |
ENSMUST00000107703.1
|
Tmeff1
|
transmembrane protein with EGF-like and two follistatin-like domains 1 |
| chr8_-_72435043 | 0.57 |
ENSMUST00000109974.1
|
Calr3
|
calreticulin 3 |
| chr17_-_50094277 | 0.56 |
ENSMUST00000113195.1
|
Rftn1
|
raftlin lipid raft linker 1 |
| chr13_+_4059565 | 0.56 |
ENSMUST00000041768.6
|
Akr1c14
|
aldo-keto reductase family 1, member C14 |
| chr13_-_118387224 | 0.56 |
ENSMUST00000022245.8
|
Mrps30
|
mitochondrial ribosomal protein S30 |
| chr1_-_93342734 | 0.55 |
ENSMUST00000027493.3
|
Pask
|
PAS domain containing serine/threonine kinase |
| chr1_+_181150926 | 0.54 |
ENSMUST00000134115.1
ENSMUST00000111059.1 |
Cnih4
|
cornichon homolog 4 (Drosophila) |
| chr11_+_67966442 | 0.54 |
ENSMUST00000021286.4
ENSMUST00000108675.1 |
Stx8
|
syntaxin 8 |
| chr13_+_21811737 | 0.53 |
ENSMUST00000104941.2
|
Hist1h4m
|
histone cluster 1, H4m |
| chr15_-_93398263 | 0.53 |
ENSMUST00000162160.1
ENSMUST00000076070.2 |
Zcrb1
|
zinc finger CCHC-type and RNA binding motif 1 |
| chr2_+_72476225 | 0.53 |
ENSMUST00000157019.1
|
Cdca7
|
cell division cycle associated 7 |
| chr15_+_11064764 | 0.52 |
ENSMUST00000061318.7
|
Adamts12
|
a disintegrin-like and metallopeptidase (reprolysin type) with thrombospondin type 1 motif, 12 |
| chr5_-_121660477 | 0.52 |
ENSMUST00000031412.5
ENSMUST00000111770.1 |
Acad10
|
acyl-Coenzyme A dehydrogenase family, member 10 |
| chr1_-_161979636 | 0.52 |
ENSMUST00000162676.1
|
4930558K02Rik
|
RIKEN cDNA 4930558K02 gene |
| chrX_-_102189371 | 0.51 |
ENSMUST00000033683.7
|
Rps4x
|
ribosomal protein S4, X-linked |
| chr2_-_84670727 | 0.51 |
ENSMUST00000117299.2
|
2700094K13Rik
|
RIKEN cDNA 2700094K13 gene |
| chr6_-_124814288 | 0.50 |
ENSMUST00000172132.2
|
Tpi1
|
triosephosphate isomerase 1 |
| chr7_+_5056856 | 0.49 |
ENSMUST00000131368.1
ENSMUST00000123956.1 |
Ccdc106
|
coiled-coil domain containing 106 |
| chr15_+_76343504 | 0.49 |
ENSMUST00000023210.6
|
Cyc1
|
cytochrome c-1 |
| chr11_-_80142164 | 0.49 |
ENSMUST00000050207.9
|
Tefm
|
transcription elongation factor, mitochondrial |
| chr11_-_68973840 | 0.49 |
ENSMUST00000038644.4
|
Rangrf
|
RAN guanine nucleotide release factor |
| chr11_+_23256566 | 0.49 |
ENSMUST00000136235.1
|
Xpo1
|
exportin 1, CRM1 homolog (yeast) |
| chr14_+_13453937 | 0.49 |
ENSMUST00000153954.1
|
Synpr
|
synaptoporin |
| chr17_-_35673738 | 0.48 |
ENSMUST00000001565.8
|
Gtf2h4
|
general transcription factor II H, polypeptide 4 |
| chr2_+_28496891 | 0.48 |
ENSMUST00000163121.1
|
Gbgt1
|
globoside alpha-1,3-N-acetylgalactosaminyltransferase 1 |
| chr2_-_166398124 | 0.48 |
ENSMUST00000151070.1
ENSMUST00000132465.1 |
Gm14268
|
predicted gene 14268 |
| chr7_-_133123409 | 0.47 |
ENSMUST00000170459.1
ENSMUST00000166400.1 |
Ctbp2
|
C-terminal binding protein 2 |
| chr4_+_48585135 | 0.47 |
ENSMUST00000030032.6
|
Tmeff1
|
transmembrane protein with EGF-like and two follistatin-like domains 1 |
| chr7_+_57590503 | 0.46 |
ENSMUST00000085240.4
|
Gabrb3
|
gamma-aminobutyric acid (GABA) A receptor, subunit beta 3 |
| chr17_+_28313513 | 0.46 |
ENSMUST00000114803.1
ENSMUST00000114801.1 ENSMUST00000114804.3 ENSMUST00000088007.4 |
Fance
|
Fanconi anemia, complementation group E |
| chr7_-_114276107 | 0.45 |
ENSMUST00000033008.9
|
Psma1
|
proteasome (prosome, macropain) subunit, alpha type 1 |
| chr15_+_91231578 | 0.45 |
ENSMUST00000109284.2
|
CN725425
|
cDNA sequence CN725425 |
| chr18_+_46597698 | 0.45 |
ENSMUST00000078079.3
ENSMUST00000168382.1 |
Eif1a
|
eukaryotic translation initiation factor 1A |
| chr15_+_81987835 | 0.45 |
ENSMUST00000165777.1
|
Xrcc6
|
X-ray repair complementing defective repair in Chinese hamster cells 6 |
| chr11_-_80142123 | 0.45 |
ENSMUST00000131601.1
|
Tefm
|
transcription elongation factor, mitochondrial |
| chr16_+_43889936 | 0.44 |
ENSMUST00000151183.1
|
2610015P09Rik
|
RIKEN cDNA 2610015P09 gene |
| chr8_+_11840474 | 0.44 |
ENSMUST00000033909.7
|
Tex29
|
testis expressed 29 |
| chr11_+_94044111 | 0.44 |
ENSMUST00000132079.1
|
Spag9
|
sperm associated antigen 9 |
| chr6_+_122707489 | 0.44 |
ENSMUST00000112581.1
ENSMUST00000112580.1 ENSMUST00000012540.4 |
Nanog
|
Nanog homeobox |
| chr17_+_74717743 | 0.44 |
ENSMUST00000024882.6
|
Ttc27
|
tetratricopeptide repeat domain 27 |
| chr10_+_62449489 | 0.44 |
ENSMUST00000181110.1
|
4930507D05Rik
|
RIKEN cDNA 4930507D05 gene |
| chr17_+_13526128 | 0.44 |
ENSMUST00000115649.2
|
Smok4a
|
sperm motility kinase 4A |
| chr11_-_66168505 | 0.44 |
ENSMUST00000080665.3
|
Dnah9
|
dynein, axonemal, heavy chain 9 |
| chr17_+_56613392 | 0.43 |
ENSMUST00000080492.5
|
Rpl36
|
ribosomal protein L36 |
| chr12_+_52097737 | 0.43 |
ENSMUST00000040090.9
|
Nubpl
|
nucleotide binding protein-like |
| chr5_-_49524764 | 0.42 |
ENSMUST00000172363.2
|
Kcnip4
|
Kv channel interacting protein 4 |
| chr14_-_19977151 | 0.42 |
ENSMUST00000055100.7
ENSMUST00000162425.1 |
Gng2
|
guanine nucleotide binding protein (G protein), gamma 2 |
| chr10_+_45067167 | 0.42 |
ENSMUST00000099858.2
|
Prep
|
prolyl endopeptidase |
| chr11_-_120086790 | 0.41 |
ENSMUST00000106227.1
ENSMUST00000106229.1 ENSMUST00000180242.1 |
Azi1
|
5-azacytidine induced gene 1 |
| chr18_+_23989632 | 0.41 |
ENSMUST00000074941.7
|
Zfp35
|
zinc finger protein 35 |
| chr1_+_71557149 | 0.41 |
ENSMUST00000027384.5
|
Atic
|
5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase |
| chr17_+_25786566 | 0.40 |
ENSMUST00000095500.4
|
Ccdc78
|
coiled-coil domain containing 78 |
| chr10_+_82378593 | 0.40 |
ENSMUST00000165906.1
|
Gm4924
|
predicted gene 4924 |
| chr13_+_4228682 | 0.40 |
ENSMUST00000118663.1
|
Akr1c19
|
aldo-keto reductase family 1, member C19 |
| chr5_-_92310003 | 0.40 |
ENSMUST00000031364.1
|
Sdad1
|
SDA1 domain containing 1 |
| chr6_+_41951625 | 0.40 |
ENSMUST00000031898.4
|
Sval1
|
seminal vesicle antigen-like 1 |
| chr11_+_70017199 | 0.39 |
ENSMUST00000133140.1
|
Dlg4
|
discs, large homolog 4 (Drosophila) |
| chr15_-_91191733 | 0.39 |
ENSMUST00000069511.6
|
Abcd2
|
ATP-binding cassette, sub-family D (ALD), member 2 |
| chr13_+_18717289 | 0.39 |
ENSMUST00000072961.4
|
Vps41
|
vacuolar protein sorting 41 (yeast) |
| chr8_-_105707933 | 0.38 |
ENSMUST00000013299.9
|
Enkd1
|
enkurin domain containing 1 |
| chr1_+_173673651 | 0.38 |
ENSMUST00000085876.4
|
Pydc3
|
pyrin domain containing 3 |
| chr9_-_100486788 | 0.38 |
ENSMUST00000098458.3
|
Il20rb
|
interleukin 20 receptor beta |
| chr15_+_76904070 | 0.38 |
ENSMUST00000004072.8
|
Rpl8
|
ribosomal protein L8 |
| chr11_+_88047302 | 0.37 |
ENSMUST00000139129.2
|
Srsf1
|
serine/arginine-rich splicing factor 1 |
| chr10_+_3973086 | 0.37 |
ENSMUST00000117291.1
ENSMUST00000120585.1 ENSMUST00000043735.7 |
Mthfd1l
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1-like |
| chr7_+_30650385 | 0.37 |
ENSMUST00000181529.1
|
Gm26610
|
predicted gene, 26610 |
| chr5_+_124552905 | 0.36 |
ENSMUST00000111438.1
|
Ddx55
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 55 |
| chr5_-_149053038 | 0.36 |
ENSMUST00000085546.6
|
Hmgb1
|
high mobility group box 1 |
| chr7_-_80324418 | 0.36 |
ENSMUST00000047362.4
ENSMUST00000121882.1 |
Rccd1
|
RCC1 domain containing 1 |
| chr16_+_17619341 | 0.36 |
ENSMUST00000006053.6
ENSMUST00000171435.1 ENSMUST00000163476.1 ENSMUST00000168101.1 ENSMUST00000165363.1 ENSMUST00000169662.1 ENSMUST00000090159.4 ENSMUST00000172182.1 ENSMUST00000163592.1 |
Smpd4
|
sphingomyelin phosphodiesterase 4 |
| chr5_+_124194894 | 0.36 |
ENSMUST00000159053.1
ENSMUST00000162577.1 |
Gm16338
|
predicted gene 16338 |
| chr7_+_111028951 | 0.36 |
ENSMUST00000005749.5
|
Ctr9
|
Ctr9, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) |
| chr6_+_123229843 | 0.36 |
ENSMUST00000112554.2
ENSMUST00000024118.4 ENSMUST00000117130.1 |
Clec4n
|
C-type lectin domain family 4, member n |
| chr16_+_43889896 | 0.36 |
ENSMUST00000122014.1
ENSMUST00000178400.1 |
2610015P09Rik
|
RIKEN cDNA 2610015P09 gene |
| chr5_-_139775631 | 0.35 |
ENSMUST00000072607.4
|
Ints1
|
integrator complex subunit 1 |
| chr18_+_80206887 | 0.35 |
ENSMUST00000127234.1
|
Gm16286
|
predicted gene 16286 |
| chr11_-_120598346 | 0.35 |
ENSMUST00000026125.2
|
Alyref
|
Aly/REF export factor |
| chr10_+_58394381 | 0.35 |
ENSMUST00000105468.1
|
Lims1
|
LIM and senescent cell antigen-like domains 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Obox6_Obox5
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 3.2 | GO:0097155 | fasciculation of sensory neuron axon(GO:0097155) |
| 1.0 | 5.9 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.8 | 2.4 | GO:0031660 | regulation of cyclin-dependent protein serine/threonine kinase activity involved in G2/M transition of mitotic cell cycle(GO:0031660) positive regulation of cyclin-dependent protein serine/threonine kinase activity involved in G2/M transition of mitotic cell cycle(GO:0031662) |
| 0.5 | 1.6 | GO:1901896 | positive regulation of calcium-transporting ATPase activity(GO:1901896) |
| 0.5 | 2.6 | GO:0014016 | neuroblast differentiation(GO:0014016) |
| 0.5 | 1.5 | GO:1990523 | bone regeneration(GO:1990523) |
| 0.5 | 3.2 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.4 | 1.5 | GO:0014732 | skeletal muscle atrophy(GO:0014732) |
| 0.4 | 1.4 | GO:0015825 | L-serine transport(GO:0015825) |
| 0.3 | 1.0 | GO:0036228 | protein targeting to nuclear inner membrane(GO:0036228) |
| 0.3 | 3.1 | GO:2000348 | regulation of CD40 signaling pathway(GO:2000348) |
| 0.3 | 0.9 | GO:2000118 | regulation of sodium-dependent phosphate transport(GO:2000118) |
| 0.3 | 0.9 | GO:0045004 | DNA replication proofreading(GO:0045004) |
| 0.3 | 1.6 | GO:0048254 | snoRNA localization(GO:0048254) |
| 0.3 | 1.3 | GO:0007207 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) phospholipase C-activating G-protein coupled acetylcholine receptor signaling pathway(GO:0007207) |
| 0.2 | 0.2 | GO:1903899 | positive regulation of PERK-mediated unfolded protein response(GO:1903899) |
| 0.2 | 1.9 | GO:1900264 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.2 | 0.6 | GO:0072244 | metanephric glomerular epithelium development(GO:0072244) |
| 0.2 | 1.7 | GO:0035356 | cellular triglyceride homeostasis(GO:0035356) |
| 0.2 | 1.0 | GO:0030920 | N-terminal peptidyl-serine acetylation(GO:0017198) N-terminal peptidyl-glutamic acid acetylation(GO:0018002) peptidyl-serine acetylation(GO:0030920) |
| 0.2 | 0.8 | GO:0051599 | response to hydrostatic pressure(GO:0051599) |
| 0.2 | 0.8 | GO:0090472 | dibasic protein processing(GO:0090472) |
| 0.2 | 1.5 | GO:2000298 | regulation of Rho-dependent protein serine/threonine kinase activity(GO:2000298) |
| 0.2 | 0.5 | GO:0046166 | alditol catabolic process(GO:0019405) glyceraldehyde-3-phosphate biosynthetic process(GO:0046166) |
| 0.2 | 0.8 | GO:0090666 | scaRNA localization to Cajal body(GO:0090666) |
| 0.2 | 1.1 | GO:0061591 | calcium activated phospholipid scrambling(GO:0061588) calcium activated phosphatidylcholine scrambling(GO:0061590) calcium activated galactosylceramide scrambling(GO:0061591) |
| 0.1 | 1.3 | GO:0097475 | motor neuron migration(GO:0097475) |
| 0.1 | 2.0 | GO:0034472 | snRNA 3'-end processing(GO:0034472) |
| 0.1 | 0.7 | GO:0071475 | cellular hyperosmotic salinity response(GO:0071475) |
| 0.1 | 0.7 | GO:1903378 | positive regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903378) |
| 0.1 | 0.9 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.1 | 0.5 | GO:1902202 | proteoglycan catabolic process(GO:0030167) regulation of hepatocyte growth factor receptor signaling pathway(GO:1902202) |
| 0.1 | 0.8 | GO:0046654 | tetrahydrofolate biosynthetic process(GO:0046654) |
| 0.1 | 1.3 | GO:0048386 | positive regulation of retinoic acid receptor signaling pathway(GO:0048386) |
| 0.1 | 0.4 | GO:0001807 | regulation of type IV hypersensitivity(GO:0001807) negative regulation of hypersensitivity(GO:0002884) |
| 0.1 | 0.9 | GO:0061732 | mitochondrial acetyl-CoA biosynthetic process from pyruvate(GO:0061732) |
| 0.1 | 0.4 | GO:0002270 | plasmacytoid dendritic cell activation(GO:0002270) |
| 0.1 | 0.4 | GO:2001160 | regulation of histone H3-K79 methylation(GO:2001160) |
| 0.1 | 1.1 | GO:0007000 | nucleolus organization(GO:0007000) |
| 0.1 | 0.9 | GO:0006390 | transcription from mitochondrial promoter(GO:0006390) |
| 0.1 | 0.6 | GO:0031284 | positive regulation of guanylate cyclase activity(GO:0031284) |
| 0.1 | 0.3 | GO:0006550 | isoleucine catabolic process(GO:0006550) |
| 0.1 | 0.4 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.1 | 0.6 | GO:0033762 | response to glucagon(GO:0033762) |
| 0.1 | 0.7 | GO:1990416 | cellular response to brain-derived neurotrophic factor stimulus(GO:1990416) |
| 0.1 | 0.9 | GO:0046826 | negative regulation of protein export from nucleus(GO:0046826) |
| 0.1 | 0.4 | GO:0000055 | ribosomal large subunit export from nucleus(GO:0000055) |
| 0.1 | 0.4 | GO:0002540 | leukotriene production involved in inflammatory response(GO:0002540) |
| 0.1 | 0.4 | GO:1902774 | late endosome to lysosome transport(GO:1902774) |
| 0.1 | 0.9 | GO:0035902 | response to immobilization stress(GO:0035902) |
| 0.1 | 0.6 | GO:0002457 | T cell antigen processing and presentation(GO:0002457) |
| 0.1 | 0.7 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.1 | 0.3 | GO:0032831 | CD4-positive, CD25-positive, alpha-beta regulatory T cell lineage commitment(GO:0002362) negative regulation of histone deacetylation(GO:0031064) positive regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032831) negative regulation of interferon-gamma biosynthetic process(GO:0045077) |
| 0.1 | 0.7 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 0.1 | 1.8 | GO:0031424 | keratinization(GO:0031424) |
| 0.1 | 0.8 | GO:0035372 | protein localization to microtubule(GO:0035372) |
| 0.1 | 0.3 | GO:0002182 | cytoplasmic translational elongation(GO:0002182) regulation of cytoplasmic translational elongation(GO:1900247) negative regulation of cytoplasmic translational elongation(GO:1900248) |
| 0.1 | 0.3 | GO:0006624 | vacuolar protein processing(GO:0006624) |
| 0.1 | 0.7 | GO:2000346 | negative regulation of hepatocyte proliferation(GO:2000346) |
| 0.1 | 0.3 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.1 | 0.9 | GO:1903894 | regulation of IRE1-mediated unfolded protein response(GO:1903894) |
| 0.1 | 0.2 | GO:1904049 | negative regulation of spontaneous neurotransmitter secretion(GO:1904049) |
| 0.1 | 0.2 | GO:0048022 | negative regulation of melanin biosynthetic process(GO:0048022) negative regulation of secondary metabolite biosynthetic process(GO:1900377) |
| 0.1 | 0.5 | GO:0086043 | bundle of His cell to Purkinje myocyte signaling(GO:0086028) bundle of His cell action potential(GO:0086043) regulation of cardiac muscle cell action potential involved in regulation of contraction(GO:0098909) |
| 0.1 | 0.1 | GO:0002780 | antimicrobial peptide biosynthetic process(GO:0002777) antibacterial peptide biosynthetic process(GO:0002780) |
| 0.1 | 0.8 | GO:0002862 | negative regulation of inflammatory response to antigenic stimulus(GO:0002862) |
| 0.1 | 0.2 | GO:0006532 | aspartate biosynthetic process(GO:0006532) |
| 0.1 | 0.2 | GO:0072221 | distal convoluted tubule development(GO:0072025) metanephric distal convoluted tubule development(GO:0072221) metanephric distal tubule development(GO:0072235) |
| 0.1 | 0.6 | GO:0000963 | mitochondrial RNA processing(GO:0000963) |
| 0.1 | 0.7 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 0.1 | 0.2 | GO:0010752 | signal complex assembly(GO:0007172) regulation of cGMP-mediated signaling(GO:0010752) regulation of B cell chemotaxis(GO:2000537) positive regulation of B cell chemotaxis(GO:2000538) |
| 0.1 | 0.2 | GO:0044313 | protein K6-linked deubiquitination(GO:0044313) protein K27-linked deubiquitination(GO:1990167) |
| 0.1 | 1.1 | GO:0008272 | sulfate transport(GO:0008272) |
| 0.1 | 0.4 | GO:0098535 | de novo centriole assembly(GO:0098535) |
| 0.1 | 0.2 | GO:0072174 | metanephric tubule formation(GO:0072174) |
| 0.1 | 0.7 | GO:0099628 | receptor diffusion trapping(GO:0098953) postsynaptic neurotransmitter receptor diffusion trapping(GO:0098970) neurotransmitter receptor diffusion trapping(GO:0099628) |
| 0.1 | 0.2 | GO:0060032 | notochord regression(GO:0060032) |
| 0.1 | 1.0 | GO:0021516 | dorsal spinal cord development(GO:0021516) |
| 0.1 | 2.8 | GO:0042775 | mitochondrial ATP synthesis coupled electron transport(GO:0042775) |
| 0.1 | 2.7 | GO:0021591 | ventricular system development(GO:0021591) |
| 0.1 | 0.3 | GO:0090273 | regulation of somatostatin secretion(GO:0090273) |
| 0.1 | 0.9 | GO:0060134 | prepulse inhibition(GO:0060134) |
| 0.1 | 0.6 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 0.1 | 0.6 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.1 | 0.2 | GO:0006478 | peptidyl-tyrosine sulfation(GO:0006478) |
| 0.1 | 0.4 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 0.0 | 0.5 | GO:0045719 | negative regulation of glycogen biosynthetic process(GO:0045719) |
| 0.0 | 0.3 | GO:0015842 | aminergic neurotransmitter loading into synaptic vesicle(GO:0015842) |
| 0.0 | 0.2 | GO:0060800 | astrocyte fate commitment(GO:0060018) regulation of cell differentiation involved in embryonic placenta development(GO:0060800) |
| 0.0 | 0.3 | GO:0006663 | platelet activating factor biosynthetic process(GO:0006663) |
| 0.0 | 0.6 | GO:0018206 | peptidyl-methionine modification(GO:0018206) |
| 0.0 | 0.4 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.0 | 0.2 | GO:2000048 | negative regulation of cell-cell adhesion mediated by cadherin(GO:2000048) |
| 0.0 | 0.3 | GO:0070127 | tRNA aminoacylation for mitochondrial protein translation(GO:0070127) |
| 0.0 | 0.1 | GO:0061642 | chemoattraction of axon(GO:0061642) |
| 0.0 | 1.0 | GO:0032825 | positive regulation of natural killer cell differentiation(GO:0032825) |
| 0.0 | 0.4 | GO:0042760 | very long-chain fatty acid catabolic process(GO:0042760) |
| 0.0 | 0.8 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.0 | 0.3 | GO:0061087 | positive regulation of histone H3-K27 methylation(GO:0061087) |
| 0.0 | 0.6 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.0 | 0.4 | GO:0035435 | cellular phosphate ion homeostasis(GO:0030643) phosphate ion transmembrane transport(GO:0035435) cellular divalent inorganic anion homeostasis(GO:0072501) cellular trivalent inorganic anion homeostasis(GO:0072502) |
| 0.0 | 0.8 | GO:0045653 | negative regulation of megakaryocyte differentiation(GO:0045653) |
| 0.0 | 1.0 | GO:0050908 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.0 | 0.5 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.0 | 1.2 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 0.2 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) |
| 0.0 | 0.3 | GO:0044789 | modulation by host of viral release from host cell(GO:0044789) positive regulation by host of viral release from host cell(GO:0044791) |
| 0.0 | 0.1 | GO:0070682 | proteasome regulatory particle assembly(GO:0070682) |
| 0.0 | 0.3 | GO:0006102 | isocitrate metabolic process(GO:0006102) |
| 0.0 | 0.2 | GO:0007597 | blood coagulation, intrinsic pathway(GO:0007597) |
| 0.0 | 0.3 | GO:0097646 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.0 | 0.2 | GO:0070560 | negative regulation of inositol phosphate biosynthetic process(GO:0010920) protein secretion by platelet(GO:0070560) |
| 0.0 | 0.3 | GO:0048664 | neuron fate determination(GO:0048664) |
| 0.0 | 0.2 | GO:0006551 | leucine metabolic process(GO:0006551) |
| 0.0 | 0.1 | GO:0042773 | ATP synthesis coupled electron transport(GO:0042773) |
| 0.0 | 0.5 | GO:0071420 | cellular response to histamine(GO:0071420) |
| 0.0 | 0.1 | GO:0009078 | alanine metabolic process(GO:0006522) pyruvate family amino acid metabolic process(GO:0009078) L-alanine metabolic process(GO:0042851) |
| 0.0 | 0.1 | GO:2000864 | estradiol secretion(GO:0035938) regulation of estradiol secretion(GO:2000864) |
| 0.0 | 3.5 | GO:0008033 | tRNA processing(GO:0008033) |
| 0.0 | 0.1 | GO:0006999 | nuclear pore organization(GO:0006999) |
| 0.0 | 0.6 | GO:0010818 | T cell chemotaxis(GO:0010818) |
| 0.0 | 0.2 | GO:0035810 | positive regulation of urine volume(GO:0035810) |
| 0.0 | 0.3 | GO:1901844 | regulation of cell communication by electrical coupling involved in cardiac conduction(GO:1901844) negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.0 | 0.2 | GO:1903690 | negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 0.0 | 0.1 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.0 | 0.2 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 0.0 | 0.8 | GO:0007339 | binding of sperm to zona pellucida(GO:0007339) |
| 0.0 | 0.5 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.0 | 0.1 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.0 | 0.3 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.0 | 0.2 | GO:1900364 | negative regulation of mRNA polyadenylation(GO:1900364) |
| 0.0 | 0.2 | GO:1990253 | cellular response to leucine starvation(GO:1990253) |
| 0.0 | 0.7 | GO:0008608 | attachment of spindle microtubules to kinetochore(GO:0008608) |
| 0.0 | 0.3 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 0.0 | 0.8 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.0 | 0.1 | GO:0031296 | B cell costimulation(GO:0031296) |
| 0.0 | 0.1 | GO:0045919 | positive regulation of cytolysis(GO:0045919) |
| 0.0 | 1.0 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.0 | 0.1 | GO:2000312 | regulation of kainate selective glutamate receptor activity(GO:2000312) |
| 0.0 | 0.2 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.0 | 0.2 | GO:0030718 | germ-line stem cell population maintenance(GO:0030718) |
| 0.0 | 0.8 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.0 | 0.2 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.0 | 0.2 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.0 | 0.4 | GO:0050832 | defense response to fungus(GO:0050832) |
| 0.0 | 0.4 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.0 | 0.1 | GO:0090435 | protein localization to nuclear envelope(GO:0090435) |
| 0.0 | 0.2 | GO:0006353 | DNA-templated transcription, termination(GO:0006353) |
| 0.0 | 1.3 | GO:0008543 | fibroblast growth factor receptor signaling pathway(GO:0008543) |
| 0.0 | 0.8 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.0 | 0.2 | GO:0032486 | Rap protein signal transduction(GO:0032486) |
| 0.0 | 0.1 | GO:0036112 | medium-chain fatty-acyl-CoA metabolic process(GO:0036112) |
| 0.0 | 0.1 | GO:0035404 | histone-serine phosphorylation(GO:0035404) |
| 0.0 | 0.2 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.0 | 0.1 | GO:1903755 | regulation of SUMO transferase activity(GO:1903182) positive regulation of SUMO transferase activity(GO:1903755) |
| 0.0 | 0.3 | GO:0051443 | positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
| 0.0 | 0.5 | GO:0008333 | endosome to lysosome transport(GO:0008333) |
| 0.0 | 1.0 | GO:0032091 | negative regulation of protein binding(GO:0032091) |
| 0.0 | 0.3 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 0.2 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.0 | 0.3 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.0 | 0.0 | GO:2000354 | regulation of ovarian follicle development(GO:2000354) |
| 0.0 | 0.3 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.4 | GO:0000942 | condensed nuclear chromosome outer kinetochore(GO:0000942) |
| 0.3 | 1.9 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 0.2 | 4.3 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.2 | 0.6 | GO:0005608 | laminin-3 complex(GO:0005608) |
| 0.2 | 1.9 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.2 | 0.6 | GO:1990047 | spindle matrix(GO:1990047) |
| 0.2 | 2.9 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.2 | 0.7 | GO:0043564 | Ku70:Ku80 complex(GO:0043564) |
| 0.2 | 0.7 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.2 | 0.9 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.2 | 0.9 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 0.1 | 1.6 | GO:0070761 | pre-snoRNP complex(GO:0070761) |
| 0.1 | 1.5 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.1 | 2.1 | GO:0032039 | integrator complex(GO:0032039) |
| 0.1 | 1.0 | GO:0031415 | NatA complex(GO:0031415) |
| 0.1 | 0.9 | GO:0045254 | pyruvate dehydrogenase complex(GO:0045254) |
| 0.1 | 0.5 | GO:0000438 | core TFIIH complex portion of holo TFIIH complex(GO:0000438) |
| 0.1 | 0.6 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.1 | 0.5 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 0.1 | 1.1 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.1 | 1.1 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.1 | 0.9 | GO:0045275 | respiratory chain complex III(GO:0045275) |
| 0.1 | 0.6 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) |
| 0.1 | 1.3 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 0.1 | 0.4 | GO:0098536 | deuterosome(GO:0098536) |
| 0.1 | 1.3 | GO:0097470 | ribbon synapse(GO:0097470) |
| 0.1 | 0.3 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.1 | 0.6 | GO:0005642 | annulate lamellae(GO:0005642) |
| 0.1 | 1.5 | GO:0031527 | filopodium membrane(GO:0031527) |
| 0.1 | 1.0 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.1 | 1.0 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.1 | 0.2 | GO:1990851 | Wnt-Frizzled-LRP5/6 complex(GO:1990851) |
| 0.1 | 0.8 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.1 | 0.2 | GO:1990761 | growth cone lamellipodium(GO:1990761) |
| 0.1 | 0.4 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.1 | 0.3 | GO:0071986 | Ragulator complex(GO:0071986) |
| 0.1 | 2.5 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.3 | GO:1903439 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.0 | 1.0 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.0 | 0.8 | GO:0012510 | trans-Golgi network transport vesicle membrane(GO:0012510) |
| 0.0 | 0.4 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 0.0 | 0.2 | GO:0014802 | terminal cisterna(GO:0014802) |
| 0.0 | 0.7 | GO:0065010 | extracellular membrane-bounded organelle(GO:0065010) |
| 0.0 | 0.2 | GO:0030981 | cortical microtubule cytoskeleton(GO:0030981) |
| 0.0 | 0.4 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.0 | 1.8 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 0.7 | GO:0044300 | juxtaparanode region of axon(GO:0044224) cerebellar mossy fiber(GO:0044300) |
| 0.0 | 0.6 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 1.6 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 0.2 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.0 | 0.2 | GO:0016035 | zeta DNA polymerase complex(GO:0016035) |
| 0.0 | 0.8 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.0 | 0.1 | GO:1990769 | proximal neuron projection(GO:1990769) |
| 0.0 | 0.6 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
| 0.0 | 0.6 | GO:0030660 | Golgi-associated vesicle membrane(GO:0030660) |
| 0.0 | 0.1 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.0 | 0.2 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.0 | 0.3 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.0 | 0.3 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.0 | 1.2 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.7 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.0 | 0.8 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 0.3 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 0.9 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 0.3 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 0.8 | GO:0043034 | costamere(GO:0043034) |
| 0.0 | 0.9 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.2 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.0 | 0.4 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.0 | 1.1 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 1.1 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 0.3 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.5 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.0 | 0.8 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 0.1 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.0 | 0.3 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.0 | 1.8 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 1.7 | GO:0005840 | ribosome(GO:0005840) |
| 0.0 | 3.5 | GO:0005769 | early endosome(GO:0005769) |
| 0.0 | 0.1 | GO:1990356 | sumoylated E2 ligase complex(GO:1990356) |
| 0.0 | 0.1 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.0 | 0.1 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.0 | 0.3 | GO:0030014 | CCR4-NOT complex(GO:0030014) messenger ribonucleoprotein complex(GO:1990124) |
| 0.0 | 1.0 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 0.2 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.0 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 5.9 | GO:0017095 | heparan sulfate 6-O-sulfotransferase activity(GO:0017095) |
| 0.8 | 3.2 | GO:0003875 | ADP-ribosylarginine hydrolase activity(GO:0003875) |
| 0.6 | 2.6 | GO:0004104 | cholinesterase activity(GO:0004104) |
| 0.6 | 2.4 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.6 | 1.7 | GO:0050252 | 2-acylglycerol O-acyltransferase activity(GO:0003846) retinol O-fatty-acyltransferase activity(GO:0050252) |
| 0.4 | 1.2 | GO:0003858 | 3-hydroxybutyrate dehydrogenase activity(GO:0003858) |
| 0.4 | 3.2 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) |
| 0.3 | 1.0 | GO:0050429 | calcium-dependent phospholipase C activity(GO:0050429) |
| 0.3 | 0.9 | GO:0030337 | DNA polymerase processivity factor activity(GO:0030337) |
| 0.3 | 0.9 | GO:0052594 | tryptamine:oxygen oxidoreductase (deaminating) activity(GO:0052593) aminoacetone:oxygen oxidoreductase(deaminating) activity(GO:0052594) aliphatic-amine oxidase activity(GO:0052595) phenethylamine:oxygen oxidoreductase (deaminating) activity(GO:0052596) |
| 0.3 | 0.8 | GO:0019166 | trans-2-enoyl-CoA reductase (NADPH) activity(GO:0019166) |
| 0.3 | 0.8 | GO:0047023 | androsterone dehydrogenase activity(GO:0047023) |
| 0.3 | 1.3 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.3 | 0.8 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 0.2 | 0.9 | GO:0015321 | sodium-dependent phosphate transmembrane transporter activity(GO:0015321) |
| 0.2 | 1.0 | GO:1990190 | peptide-glutamate-N-acetyltransferase activity(GO:1990190) |
| 0.2 | 1.0 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.2 | 1.0 | GO:0004768 | stearoyl-CoA 9-desaturase activity(GO:0004768) |
| 0.2 | 0.6 | GO:0034617 | nitric-oxide synthase activity(GO:0004517) tetrahydrobiopterin binding(GO:0034617) |
| 0.2 | 1.5 | GO:0001094 | TFIID-class transcription factor binding(GO:0001094) |
| 0.2 | 1.9 | GO:0033170 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.2 | 0.9 | GO:0004738 | pyruvate dehydrogenase activity(GO:0004738) pyruvate dehydrogenase [NAD(P)+] activity(GO:0034603) pyruvate dehydrogenase (NAD+) activity(GO:0034604) |
| 0.2 | 1.3 | GO:0030267 | hydroxypyruvate reductase activity(GO:0016618) glyoxylate reductase (NADP) activity(GO:0030267) |
| 0.2 | 0.8 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.1 | 2.1 | GO:0004000 | adenosine deaminase activity(GO:0004000) |
| 0.1 | 0.7 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.1 | 0.4 | GO:0004051 | arachidonate 5-lipoxygenase activity(GO:0004051) |
| 0.1 | 0.8 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.1 | 0.4 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.1 | 0.7 | GO:0044547 | DNA topoisomerase binding(GO:0044547) |
| 0.1 | 0.4 | GO:0000402 | open form four-way junction DNA binding(GO:0000401) crossed form four-way junction DNA binding(GO:0000402) |
| 0.1 | 1.0 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.1 | 0.6 | GO:0030942 | endoplasmic reticulum signal peptide binding(GO:0030942) |
| 0.1 | 0.9 | GO:0008310 | single-stranded DNA 3'-5' exodeoxyribonuclease activity(GO:0008310) |
| 0.1 | 0.7 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.1 | 1.4 | GO:0022889 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.1 | 2.8 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 0.4 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 0.1 | 0.4 | GO:0001010 | transcription factor activity, sequence-specific DNA binding transcription factor recruiting(GO:0001010) |
| 0.1 | 0.7 | GO:0031811 | G-protein coupled nucleotide receptor binding(GO:0031811) P2Y1 nucleotide receptor binding(GO:0031812) |
| 0.1 | 0.3 | GO:0004058 | aromatic-L-amino-acid decarboxylase activity(GO:0004058) L-dopa decarboxylase activity(GO:0036468) |
| 0.1 | 0.3 | GO:0000700 | mismatch base pair DNA N-glycosylase activity(GO:0000700) |
| 0.1 | 0.7 | GO:0051575 | 5'-deoxyribose-5-phosphate lyase activity(GO:0051575) |
| 0.1 | 0.5 | GO:0022851 | GABA-gated chloride ion channel activity(GO:0022851) |
| 0.1 | 2.0 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.1 | 1.5 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.1 | 1.7 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 0.2 | GO:0004069 | L-aspartate:2-oxoglutarate aminotransferase activity(GO:0004069) |
| 0.1 | 0.6 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.1 | 0.3 | GO:0004449 | isocitrate dehydrogenase (NAD+) activity(GO:0004449) |
| 0.1 | 1.5 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 0.9 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 0.3 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.1 | 0.2 | GO:0034211 | GTP-dependent protein kinase activity(GO:0034211) |
| 0.1 | 0.3 | GO:0003985 | acetyl-CoA C-acetyltransferase activity(GO:0003985) C-acetyltransferase activity(GO:0016453) |
| 0.1 | 0.8 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.1 | 1.1 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 0.1 | 1.9 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.1 | 0.2 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 0.1 | 0.8 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.1 | 0.2 | GO:0008476 | protein-tyrosine sulfotransferase activity(GO:0008476) |
| 0.1 | 0.4 | GO:0005000 | vasopressin receptor activity(GO:0005000) |
| 0.0 | 0.4 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.0 | 0.4 | GO:0008121 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.0 | 0.8 | GO:0043855 | cyclic nucleotide-gated ion channel activity(GO:0043855) |
| 0.0 | 0.3 | GO:0097643 | amylin receptor activity(GO:0097643) |
| 0.0 | 0.2 | GO:0015038 | glutathione disulfide oxidoreductase activity(GO:0015038) |
| 0.0 | 0.5 | GO:0016861 | intramolecular oxidoreductase activity, interconverting aldoses and ketoses(GO:0016861) |
| 0.0 | 1.0 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.0 | 0.6 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.0 | 0.4 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.2 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.0 | 1.4 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 0.1 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) |
| 0.0 | 0.3 | GO:0097027 | ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.0 | 0.5 | GO:0035615 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.0 | 0.3 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.0 | 0.6 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.0 | 0.6 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.2 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.0 | 0.2 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.0 | 0.6 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.3 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 0.0 | 0.3 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.0 | 0.1 | GO:0000171 | ribonuclease MRP activity(GO:0000171) |
| 0.0 | 0.1 | GO:1990188 | euchromatin binding(GO:1990188) |
| 0.0 | 0.2 | GO:0019864 | IgG binding(GO:0019864) |
| 0.0 | 1.3 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 0.4 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.0 | 0.5 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.0 | 0.6 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.0 | 0.1 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.0 | 1.5 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.0 | 0.8 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.0 | 1.2 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.0 | 0.2 | GO:0043423 | 3-phosphoinositide-dependent protein kinase binding(GO:0043423) |
| 0.0 | 0.1 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.0 | 0.5 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.0 | 0.5 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.0 | 0.1 | GO:0003831 | beta-N-acetylglucosaminylglycopeptide beta-1,4-galactosyltransferase activity(GO:0003831) |
| 0.0 | 1.3 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.0 | 0.3 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.0 | 0.8 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 1.6 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 0.1 | GO:0005550 | pheromone binding(GO:0005550) |
| 0.0 | 0.3 | GO:0070402 | NADPH binding(GO:0070402) |
| 0.0 | 0.4 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.0 | 0.1 | GO:0001851 | complement component C3b binding(GO:0001851) |
| 0.0 | 0.6 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.0 | 0.6 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 0.8 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 0.1 | GO:0016508 | long-chain-enoyl-CoA hydratase activity(GO:0016508) |
| 0.0 | 0.1 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.0 | 0.3 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 0.0 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.0 | 0.9 | GO:0009055 | electron carrier activity(GO:0009055) |
| 0.0 | 0.5 | GO:0008376 | acetylgalactosaminyltransferase activity(GO:0008376) |
| 0.0 | 0.2 | GO:0043008 | ATP-dependent protein binding(GO:0043008) |
| 0.0 | 0.4 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.0 | 0.3 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) dynein light chain binding(GO:0045503) |
| 0.0 | 1.0 | GO:0004004 | RNA helicase activity(GO:0003724) ATP-dependent RNA helicase activity(GO:0004004) |
| 0.0 | 0.6 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) |
| 0.0 | 0.2 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.0 | 0.2 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.0 | 0.2 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.0 | 0.1 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.0 | 0.6 | GO:0016504 | peptidase activator activity(GO:0016504) |
| 0.0 | 0.9 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.0 | 0.1 | GO:0061656 | SUMO conjugating enzyme activity(GO:0061656) |
| 0.0 | 1.6 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.0 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.0 | 1.0 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 0.0 | 1.6 | GO:0052689 | carboxylic ester hydrolase activity(GO:0052689) |
| 0.0 | 0.1 | GO:1990405 | protein antigen binding(GO:1990405) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 5.3 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.1 | 4.0 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 1.3 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.1 | 3.2 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.1 | 2.1 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 0.6 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 1.5 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.9 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.0 | 0.6 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.0 | 1.1 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.0 | 0.0 | PID AR NONGENOMIC PATHWAY | Nongenotropic Androgen signaling |
| 0.0 | 0.8 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 0.7 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 1.9 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 0.7 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.0 | 0.2 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 1.0 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 0.7 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 0.1 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 0.9 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 1.3 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 0.6 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.7 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.0 | 0.4 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.8 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 0.5 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 0.2 | PID AVB3 OPN PATHWAY | Osteopontin-mediated events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.6 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.2 | 2.0 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.2 | 2.8 | REACTOME REPAIR SYNTHESIS FOR GAP FILLING BY DNA POL IN TC NER | Genes involved in Repair synthesis for gap-filling by DNA polymerase in TC-NER |
| 0.2 | 2.7 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.2 | 5.9 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.1 | 1.5 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.1 | 0.3 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 0.1 | 1.7 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.1 | 2.4 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.1 | 0.9 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.1 | 0.8 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.0 | 3.2 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.5 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.0 | 1.5 | REACTOME PROSTACYCLIN SIGNALLING THROUGH PROSTACYCLIN RECEPTOR | Genes involved in Prostacyclin signalling through prostacyclin receptor |
| 0.0 | 3.9 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 1.3 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.0 | 0.7 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.0 | 0.4 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.0 | 1.4 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.0 | 2.0 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.4 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.0 | 0.6 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 0.5 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 0.4 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.0 | 0.1 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.0 | 0.6 | REACTOME SRP DEPENDENT COTRANSLATIONAL PROTEIN TARGETING TO MEMBRANE | Genes involved in SRP-dependent cotranslational protein targeting to membrane |
| 0.0 | 0.5 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.0 | 0.5 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.0 | 0.6 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.2 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.0 | 0.3 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.0 | 0.4 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.3 | REACTOME FGFR2C LIGAND BINDING AND ACTIVATION | Genes involved in FGFR2c ligand binding and activation |
| 0.0 | 0.6 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.0 | 0.3 | REACTOME ACYL CHAIN REMODELLING OF PI | Genes involved in Acyl chain remodelling of PI |
| 0.0 | 0.2 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.0 | 0.6 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.3 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.0 | 0.6 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 0.6 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 0.2 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.0 | 0.8 | REACTOME TRIGLYCERIDE BIOSYNTHESIS | Genes involved in Triglyceride Biosynthesis |
| 0.0 | 0.9 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 0.7 | REACTOME REGULATION OF ORNITHINE DECARBOXYLASE ODC | Genes involved in Regulation of ornithine decarboxylase (ODC) |
| 0.0 | 0.2 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.1 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.0 | 0.0 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |
| 0.0 | 0.2 | REACTOME SHC1 EVENTS IN ERBB4 SIGNALING | Genes involved in SHC1 events in ERBB4 signaling |