Project
avrg: 12D miR HR13_24
Navigation
Downloads
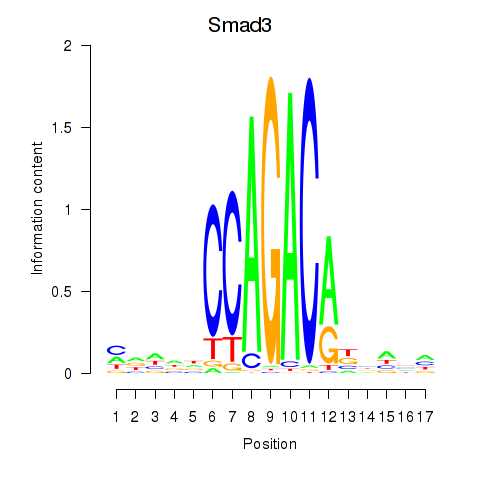
Results for Smad3
Z-value: 2.36
Transcription factors associated with Smad3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Smad3
|
ENSMUSG00000032402.6 | SMAD family member 3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Smad3 | mm10_v2_chr9_-_63757933_63757994 | -0.25 | 4.7e-01 | Click! |
Activity profile of Smad3 motif
Sorted Z-values of Smad3 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_+_100779074 | 4.22 |
ENSMUST00000110073.1
ENSMUST00000110070.1 |
9030617O03Rik
|
RIKEN cDNA 9030617O03 gene |
| chr12_+_100779088 | 3.87 |
ENSMUST00000110069.1
|
9030617O03Rik
|
RIKEN cDNA 9030617O03 gene |
| chr12_+_100779055 | 3.73 |
ENSMUST00000069782.4
|
9030617O03Rik
|
RIKEN cDNA 9030617O03 gene |
| chr2_-_152831665 | 2.47 |
ENSMUST00000156688.1
ENSMUST00000007803.5 |
Bcl2l1
|
BCL2-like 1 |
| chr2_-_152831112 | 2.32 |
ENSMUST00000128172.1
|
Bcl2l1
|
BCL2-like 1 |
| chr13_+_21735055 | 2.24 |
ENSMUST00000087714.4
|
Hist1h4j
|
histone cluster 1, H4j |
| chr12_-_76795489 | 2.08 |
ENSMUST00000082431.3
|
Gpx2
|
glutathione peroxidase 2 |
| chr13_+_23531044 | 1.91 |
ENSMUST00000102972.3
|
Hist1h4h
|
histone cluster 1, H4h |
| chr13_-_21750505 | 1.75 |
ENSMUST00000102983.1
|
Hist1h4k
|
histone cluster 1, H4k |
| chr13_-_21832194 | 1.72 |
ENSMUST00000102979.1
|
Hist1h4n
|
histone cluster 1, H4n |
| chr10_+_33905015 | 1.56 |
ENSMUST00000169670.1
|
Rsph4a
|
radial spoke head 4 homolog A (Chlamydomonas) |
| chr13_-_23551648 | 1.40 |
ENSMUST00000102971.1
|
Hist1h4f
|
histone cluster 1, H4f |
| chr13_+_22043189 | 1.32 |
ENSMUST00000110452.1
|
Hist1h2bj
|
histone cluster 1, H2bj |
| chr3_-_96263311 | 1.17 |
ENSMUST00000171473.1
|
Hist2h4
|
histone cluster 2, H4 |
| chr13_-_22042949 | 1.14 |
ENSMUST00000091741.4
|
Hist1h2ag
|
histone cluster 1, H2ag |
| chr13_-_22035589 | 1.06 |
ENSMUST00000091742.4
|
Hist1h2ah
|
histone cluster 1, H2ah |
| chr13_+_22035821 | 1.01 |
ENSMUST00000110455.2
|
Hist1h2bk
|
histone cluster 1, H2bk |
| chr11_-_99986593 | 0.99 |
ENSMUST00000105050.2
|
Krtap16-1
|
keratin associated protein 16-1 |
| chrX_+_143664365 | 0.96 |
ENSMUST00000126592.1
ENSMUST00000156449.1 ENSMUST00000155215.1 ENSMUST00000112865.1 |
Pak3
|
p21 protein (Cdc42/Rac)-activated kinase 3 |
| chrX_+_143664290 | 0.94 |
ENSMUST00000112868.1
|
Pak3
|
p21 protein (Cdc42/Rac)-activated kinase 3 |
| chr11_+_58954675 | 0.91 |
ENSMUST00000108817.3
ENSMUST00000047697.5 |
Hist3h2a
Trim17
|
histone cluster 3, H2a tripartite motif-containing 17 |
| chr13_+_21833736 | 0.90 |
ENSMUST00000180288.1
ENSMUST00000110467.1 |
Hist1h2br
|
histone cluster 1 H2br |
| chr6_+_116650674 | 0.83 |
ENSMUST00000067354.5
ENSMUST00000178241.1 |
8430408G22Rik
|
RIKEN cDNA 8430408G22 gene |
| chr7_-_44236098 | 0.83 |
ENSMUST00000037220.4
|
1700028J19Rik
|
RIKEN cDNA 1700028J19 gene |
| chr13_+_23581563 | 0.80 |
ENSMUST00000102968.1
|
Hist1h4d
|
histone cluster 1, H4d |
| chr13_-_21833575 | 0.74 |
ENSMUST00000081342.5
|
Hist1h2ap
|
histone cluster 1, H2ap |
| chr13_+_23533869 | 0.70 |
ENSMUST00000073261.2
|
Hist1h2af
|
histone cluster 1, H2af |
| chr7_-_100964371 | 0.70 |
ENSMUST00000060174.4
|
P2ry6
|
pyrimidinergic receptor P2Y, G-protein coupled, 6 |
| chr13_+_21787461 | 0.69 |
ENSMUST00000110473.2
ENSMUST00000102982.1 |
Hist1h2bp
|
histone cluster 1, H2bp |
| chr13_-_21810190 | 0.69 |
ENSMUST00000110469.1
ENSMUST00000091749.2 |
Hist1h2bq
|
histone cluster 1, H2bq |
| chr13_-_23571151 | 0.67 |
ENSMUST00000102969.3
|
Hist1h2ae
|
histone cluster 1, H2ae |
| chr5_-_116422858 | 0.67 |
ENSMUST00000036991.4
|
Hspb8
|
heat shock protein 8 |
| chr13_+_21810428 | 0.62 |
ENSMUST00000091745.5
|
Hist1h2ao
|
histone cluster 1, H2ao |
| chr6_+_136808248 | 0.62 |
ENSMUST00000074556.4
|
H2afj
|
H2A histone family, member J |
| chr5_-_72504202 | 0.61 |
ENSMUST00000005352.3
|
Corin
|
corin |
| chr13_+_23746734 | 0.58 |
ENSMUST00000099703.2
|
Hist1h2bb
|
histone cluster 1, H2bb |
| chr13_+_23571382 | 0.51 |
ENSMUST00000079251.5
|
Hist1h2bg
|
histone cluster 1, H2bg |
| chrX_+_153832225 | 0.51 |
ENSMUST00000148708.1
ENSMUST00000123264.1 ENSMUST00000049999.8 |
Spin2c
|
spindlin family, member 2C |
| chr17_+_56628118 | 0.50 |
ENSMUST00000112979.2
|
Catsperd
|
catsper channel auxiliary subunit delta |
| chr15_+_81936911 | 0.49 |
ENSMUST00000135663.1
|
Csdc2
|
cold shock domain containing C2, RNA binding |
| chr13_+_23684192 | 0.49 |
ENSMUST00000018246.4
|
Hist1h2bc
|
histone cluster 1, H2bc |
| chr14_-_79868398 | 0.46 |
ENSMUST00000179430.1
|
Gm10845
|
predicted gene 10845 |
| chr13_-_21787218 | 0.46 |
ENSMUST00000091751.2
|
Hist1h2an
|
histone cluster 1, H2an |
| chr14_+_119138415 | 0.45 |
ENSMUST00000065904.3
|
Hs6st3
|
heparan sulfate 6-O-sulfotransferase 3 |
| chrX_-_20291728 | 0.44 |
ENSMUST00000115393.2
|
Slc9a7
|
solute carrier family 9 (sodium/hydrogen exchanger), member 7 |
| chr1_-_126492900 | 0.44 |
ENSMUST00000161954.1
|
Nckap5
|
NCK-associated protein 5 |
| chr13_-_23683941 | 0.43 |
ENSMUST00000171127.1
|
Hist1h2ac
|
histone cluster 1, H2ac |
| chrX_-_20291776 | 0.42 |
ENSMUST00000072451.4
|
Slc9a7
|
solute carrier family 9 (sodium/hydrogen exchanger), member 7 |
| chr1_-_134235420 | 0.42 |
ENSMUST00000038191.6
ENSMUST00000086465.4 |
Adora1
|
adenosine A1 receptor |
| chrX_-_150814265 | 0.41 |
ENSMUST00000026302.6
ENSMUST00000129768.1 ENSMUST00000112699.2 |
Maged2
|
melanoma antigen, family D, 2 |
| chr2_-_148408146 | 0.40 |
ENSMUST00000099270.3
|
Thbd
|
thrombomodulin |
| chr1_-_75210732 | 0.38 |
ENSMUST00000113623.1
|
Glb1l
|
galactosidase, beta 1-like |
| chr5_+_92392585 | 0.36 |
ENSMUST00000126281.1
|
Art3
|
ADP-ribosyltransferase 3 |
| chr1_+_60909148 | 0.35 |
ENSMUST00000097720.3
|
Ctla4
|
cytotoxic T-lymphocyte-associated protein 4 |
| chrX_-_144688180 | 0.31 |
ENSMUST00000040184.3
|
Trpc5
|
transient receptor potential cation channel, subfamily C, member 5 |
| chr9_+_54286479 | 0.30 |
ENSMUST00000056740.5
|
Gldn
|
gliomedin |
| chr7_-_127588595 | 0.30 |
ENSMUST00000072155.3
|
Gm166
|
predicted gene 166 |
| chr14_+_32028989 | 0.30 |
ENSMUST00000022460.4
|
Galnt15
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 15 |
| chr1_+_131688766 | 0.28 |
ENSMUST00000129905.1
|
5430435G22Rik
|
RIKEN cDNA 5430435G22 gene |
| chr4_+_140986873 | 0.25 |
ENSMUST00000168047.1
ENSMUST00000037055.7 ENSMUST00000127833.2 |
Atp13a2
|
ATPase type 13A2 |
| chr14_+_57424054 | 0.25 |
ENSMUST00000122063.1
|
Ift88
|
intraflagellar transport 88 |
| chr7_-_119479249 | 0.24 |
ENSMUST00000033263.4
|
Umod
|
uromodulin |
| chr6_-_129533267 | 0.24 |
ENSMUST00000181594.1
|
1700101I11Rik
|
RIKEN cDNA 1700101I11 gene |
| chr1_-_126492683 | 0.23 |
ENSMUST00000162877.1
|
Nckap5
|
NCK-associated protein 5 |
| chr4_+_95557494 | 0.23 |
ENSMUST00000079223.4
ENSMUST00000177394.1 |
Fggy
|
FGGY carbohydrate kinase domain containing |
| chr15_-_103252810 | 0.23 |
ENSMUST00000154510.1
|
Nfe2
|
nuclear factor, erythroid derived 2 |
| chr8_-_72435043 | 0.23 |
ENSMUST00000109974.1
|
Calr3
|
calreticulin 3 |
| chr5_+_114774677 | 0.23 |
ENSMUST00000102578.4
|
Ankrd13a
|
ankyrin repeat domain 13a |
| chr10_+_79854658 | 0.22 |
ENSMUST00000171599.1
ENSMUST00000095457.4 |
Ptbp1
|
polypyrimidine tract binding protein 1 |
| chr2_+_178193075 | 0.22 |
ENSMUST00000103065.1
|
Phactr3
|
phosphatase and actin regulator 3 |
| chr6_+_120093348 | 0.22 |
ENSMUST00000112711.2
|
Ninj2
|
ninjurin 2 |
| chr3_-_107760221 | 0.22 |
ENSMUST00000153114.1
ENSMUST00000118593.1 ENSMUST00000120243.1 |
Csf1
|
colony stimulating factor 1 (macrophage) |
| chr7_+_44384803 | 0.21 |
ENSMUST00000120262.1
|
Syt3
|
synaptotagmin III |
| chr2_-_30194112 | 0.21 |
ENSMUST00000113659.1
ENSMUST00000113660.1 |
Ccbl1
|
cysteine conjugate-beta lyase 1 |
| chr9_+_44072196 | 0.20 |
ENSMUST00000176671.1
|
Usp2
|
ubiquitin specific peptidase 2 |
| chr13_-_96435952 | 0.19 |
ENSMUST00000181761.1
|
Ankdd1b
|
ankyrin repeat and death domain containing 1B |
| chr10_+_79854618 | 0.19 |
ENSMUST00000165704.1
|
Ptbp1
|
polypyrimidine tract binding protein 1 |
| chr5_+_145204523 | 0.18 |
ENSMUST00000085671.3
ENSMUST00000031601.7 |
Zkscan5
|
zinc finger with KRAB and SCAN domains 5 |
| chr19_-_23448322 | 0.17 |
ENSMUST00000036069.6
|
Mamdc2
|
MAM domain containing 2 |
| chr1_+_60908993 | 0.17 |
ENSMUST00000027164.2
|
Ctla4
|
cytotoxic T-lymphocyte-associated protein 4 |
| chr16_-_45158183 | 0.17 |
ENSMUST00000114600.1
|
Slc35a5
|
solute carrier family 35, member A5 |
| chr5_+_136038496 | 0.16 |
ENSMUST00000062606.6
|
Upk3b
|
uroplakin 3B |
| chr11_-_26591729 | 0.16 |
ENSMUST00000109504.1
|
Vrk2
|
vaccinia related kinase 2 |
| chr7_+_137437591 | 0.16 |
ENSMUST00000064404.6
|
Glrx3
|
glutaredoxin 3 |
| chr11_-_116110211 | 0.14 |
ENSMUST00000106441.1
ENSMUST00000021120.5 |
Trim47
|
tripartite motif-containing 47 |
| chr10_-_118868903 | 0.13 |
ENSMUST00000004281.8
|
Dyrk2
|
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 2 |
| chr9_+_109038565 | 0.13 |
ENSMUST00000112059.3
ENSMUST00000026737.5 |
Shisa5
|
shisa homolog 5 (Xenopus laevis) |
| chr2_-_132945906 | 0.13 |
ENSMUST00000038280.4
|
Fermt1
|
fermitin family homolog 1 (Drosophila) |
| chr7_+_123462274 | 0.12 |
ENSMUST00000033023.3
|
Aqp8
|
aquaporin 8 |
| chr17_-_31277327 | 0.12 |
ENSMUST00000024832.7
|
Rsph1
|
radial spoke head 1 homolog (Chlamydomonas) |
| chr4_+_45018583 | 0.12 |
ENSMUST00000133157.1
ENSMUST00000029999.8 ENSMUST00000107814.3 |
Polr1e
|
polymerase (RNA) I polypeptide E |
| chr15_-_101562889 | 0.12 |
ENSMUST00000023714.4
|
4732456N10Rik
|
RIKEN cDNA 4732456N10 gene |
| chr9_+_13621646 | 0.11 |
ENSMUST00000034401.8
|
Maml2
|
mastermind like 2 (Drosophila) |
| chr9_+_21196705 | 0.11 |
ENSMUST00000003395.9
|
Pde4a
|
phosphodiesterase 4A, cAMP specific |
| chr11_+_73240310 | 0.10 |
ENSMUST00000138853.1
|
Trpv1
|
transient receptor potential cation channel, subfamily V, member 1 |
| chr6_+_129533183 | 0.10 |
ENSMUST00000032264.6
|
Gabarapl1
|
gamma-aminobutyric acid (GABA) A receptor-associated protein-like 1 |
| chr11_-_103344651 | 0.10 |
ENSMUST00000041385.7
|
Arhgap27
|
Rho GTPase activating protein 27 |
| chr17_-_27623263 | 0.09 |
ENSMUST00000062397.6
ENSMUST00000176876.1 |
Nudt3
|
nudix (nucleotide diphosphate linked moiety X)-type motif 3 |
| chr17_-_27623441 | 0.09 |
ENSMUST00000025050.5
|
Nudt3
|
nudix (nucleotide diphosphate linked moiety X)-type motif 3 |
| chr7_-_104369782 | 0.08 |
ENSMUST00000164410.1
|
Trim30b
|
tripartite motif-containing 30B |
| chr7_-_24316590 | 0.08 |
ENSMUST00000108436.1
ENSMUST00000032673.8 |
Zfp94
|
zinc finger protein 94 |
| chr2_-_104257400 | 0.07 |
ENSMUST00000141159.1
|
D430041D05Rik
|
RIKEN cDNA D430041D05 gene |
| chr9_+_108490676 | 0.07 |
ENSMUST00000178075.1
ENSMUST00000085044.7 ENSMUST00000166103.2 ENSMUST00000006854.7 |
Usp19
|
ubiquitin specific peptidase 19 |
| chr19_-_47919269 | 0.07 |
ENSMUST00000095998.5
|
Itprip
|
inositol 1,4,5-triphosphate receptor interacting protein |
| chr19_+_55898553 | 0.06 |
ENSMUST00000148666.1
|
Tcf7l2
|
transcription factor 7 like 2, T cell specific, HMG box |
| chr2_-_17460610 | 0.06 |
ENSMUST00000145492.1
|
Nebl
|
nebulette |
| chr11_+_49523721 | 0.05 |
ENSMUST00000077143.4
|
Olfr1383
|
olfactory receptor 1383 |
| chr1_+_165485168 | 0.05 |
ENSMUST00000111440.1
ENSMUST00000027852.8 ENSMUST00000111439.1 |
Adcy10
|
adenylate cyclase 10 |
| chr7_-_45052865 | 0.04 |
ENSMUST00000057293.6
|
Prr12
|
proline rich 12 |
| chr1_+_146497614 | 0.04 |
ENSMUST00000132847.1
ENSMUST00000166814.1 |
Brinp3
|
bone morphogenetic protein/retinoic acid inducible neural specific 3 |
| chr9_-_20879718 | 0.04 |
ENSMUST00000043726.6
|
Angptl6
|
angiopoietin-like 6 |
| chr19_+_5740885 | 0.03 |
ENSMUST00000081496.5
|
Ltbp3
|
latent transforming growth factor beta binding protein 3 |
| chr7_+_127588698 | 0.02 |
ENSMUST00000033088.6
|
Rnf40
|
ring finger protein 40 |
| chr7_-_81454751 | 0.02 |
ENSMUST00000098331.3
ENSMUST00000178892.1 |
Cpeb1
|
cytoplasmic polyadenylation element binding protein 1 |
| chr14_+_20348159 | 0.02 |
ENSMUST00000090503.4
ENSMUST00000090499.5 ENSMUST00000037698.5 ENSMUST00000051915.6 |
Fam149b
|
family with sequence similarity 149, member B |
| chr4_+_130055010 | 0.01 |
ENSMUST00000123617.1
|
Col16a1
|
collagen, type XVI, alpha 1 |
| chr18_+_60526194 | 0.01 |
ENSMUST00000025505.5
|
Dctn4
|
dynactin 4 |
| chr1_-_23909687 | 0.01 |
ENSMUST00000129254.1
|
Smap1
|
small ArfGAP 1 |
| chr2_-_127143410 | 0.01 |
ENSMUST00000132773.1
|
Itpripl1
|
inositol 1,4,5-triphosphate receptor interacting protein-like 1 |
| chr11_+_46436925 | 0.01 |
ENSMUST00000152119.1
ENSMUST00000140027.1 ENSMUST00000020665.6 ENSMUST00000170928.1 ENSMUST00000109231.1 ENSMUST00000109232.3 ENSMUST00000128940.1 |
Med7
|
mediator complex subunit 7 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Smad3
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 4.8 | GO:0046898 | response to cycloheximide(GO:0046898) |
| 0.5 | 11.0 | GO:0045653 | negative regulation of megakaryocyte differentiation(GO:0045653) |
| 0.2 | 0.6 | GO:0003050 | regulation of systemic arterial blood pressure by atrial natriuretic peptide(GO:0003050) |
| 0.1 | 0.4 | GO:0032242 | positive regulation of nucleobase-containing compound transport(GO:0032241) regulation of nucleoside transport(GO:0032242) |
| 0.1 | 2.1 | GO:0002862 | negative regulation of inflammatory response to antigenic stimulus(GO:0002862) |
| 0.1 | 2.0 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 0.1 | 0.3 | GO:1902630 | regulation of membrane hyperpolarization(GO:1902630) |
| 0.1 | 0.7 | GO:0030321 | transepithelial chloride transport(GO:0030321) |
| 0.1 | 0.5 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 0.1 | 0.2 | GO:0007290 | spermatid nucleus elongation(GO:0007290) |
| 0.1 | 0.2 | GO:0072221 | distal convoluted tubule development(GO:0072025) metanephric distal convoluted tubule development(GO:0072221) metanephric distal tubule development(GO:0072235) |
| 0.1 | 0.5 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.1 | 0.2 | GO:1902226 | positive regulation of odontogenesis of dentin-containing tooth(GO:0042488) regulation of macrophage colony-stimulating factor signaling pathway(GO:1902226) regulation of response to macrophage colony-stimulating factor(GO:1903969) regulation of cellular response to macrophage colony-stimulating factor stimulus(GO:1903972) |
| 0.1 | 1.9 | GO:0010763 | positive regulation of fibroblast migration(GO:0010763) |
| 0.1 | 0.4 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.1 | 0.9 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.1 | 0.2 | GO:0018197 | peptidyl-aspartic acid modification(GO:0018197) |
| 0.1 | 0.2 | GO:0071544 | diphosphoinositol polyphosphate catabolic process(GO:0071544) |
| 0.1 | 0.9 | GO:0097369 | sodium ion import(GO:0097369) sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.0 | 5.7 | GO:0006342 | chromatin silencing(GO:0006342) |
| 0.0 | 0.1 | GO:0001180 | transcription initiation from RNA polymerase I promoter for nuclear large rRNA transcript(GO:0001180) |
| 0.0 | 0.3 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.0 | 0.4 | GO:0075522 | IRES-dependent viral translational initiation(GO:0075522) |
| 0.0 | 0.1 | GO:2001201 | regulation of transforming growth factor-beta secretion(GO:2001201) |
| 0.0 | 1.7 | GO:0035082 | axoneme assembly(GO:0035082) |
| 0.0 | 0.2 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.0 | 2.6 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.0 | 0.5 | GO:0048240 | sperm capacitation(GO:0048240) |
| 0.0 | 0.2 | GO:0034242 | negative regulation of syncytium formation by plasma membrane fusion(GO:0034242) |
| 0.0 | 0.2 | GO:2000659 | regulation of interleukin-1-mediated signaling pathway(GO:2000659) |
| 0.0 | 0.1 | GO:0015722 | canalicular bile acid transport(GO:0015722) |
| 0.0 | 0.1 | GO:0051534 | negative regulation of NFAT protein import into nucleus(GO:0051534) |
| 0.0 | 0.2 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.0 | 0.2 | GO:0019321 | pentose metabolic process(GO:0019321) |
| 0.0 | 0.3 | GO:0048642 | negative regulation of skeletal muscle tissue development(GO:0048642) |
| 0.0 | 0.0 | GO:1902460 | transforming growth factor beta activation(GO:0036363) regulation of mesenchymal stem cell proliferation(GO:1902460) positive regulation of mesenchymal stem cell proliferation(GO:1902462) |
| 0.0 | 0.1 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 4.8 | GO:0097136 | Bcl-2 family protein complex(GO:0097136) |
| 0.3 | 10.2 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.2 | 6.4 | GO:0000786 | nucleosome(GO:0000786) |
| 0.1 | 0.2 | GO:1990682 | CSF1-CSF1R complex(GO:1990682) |
| 0.1 | 0.2 | GO:0031310 | integral component of vacuolar membrane(GO:0031166) intrinsic component of vacuolar membrane(GO:0031310) |
| 0.1 | 0.5 | GO:0036128 | CatSper complex(GO:0036128) |
| 0.0 | 0.1 | GO:0046691 | intracellular canaliculus(GO:0046691) |
| 0.0 | 0.2 | GO:0002081 | outer acrosomal membrane(GO:0002081) |
| 0.0 | 1.0 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.9 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 0.4 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 0.0 | 0.4 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.0 | 0.5 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.0 | 0.1 | GO:0001520 | outer dense fiber(GO:0001520) |
| 0.0 | 1.6 | GO:0005930 | axoneme(GO:0005930) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 4.8 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.2 | 0.7 | GO:0015065 | uridine nucleotide receptor activity(GO:0015065) G-protein coupled pyrimidinergic nucleotide receptor activity(GO:0071553) |
| 0.1 | 0.5 | GO:0017095 | heparan sulfate 6-O-sulfotransferase activity(GO:0017095) |
| 0.1 | 1.9 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.1 | 2.1 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.1 | 0.4 | GO:0004565 | beta-galactosidase activity(GO:0004565) |
| 0.1 | 0.2 | GO:0019150 | D-ribulokinase activity(GO:0019150) |
| 0.1 | 0.2 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 0.1 | 0.4 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 0.1 | 0.9 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.1 | 0.4 | GO:0032795 | heterotrimeric G-protein binding(GO:0032795) |
| 0.0 | 10.8 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 0.2 | GO:0019864 | IgG binding(GO:0019864) |
| 0.0 | 0.3 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 0.0 | 0.2 | GO:0008486 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.0 | 0.4 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.0 | 0.3 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 0.1 | GO:0001179 | RNA polymerase I transcription factor binding(GO:0001179) |
| 0.0 | 0.3 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 0.6 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.2 | GO:0070300 | calcium-transporting ATPase activity(GO:0005388) phosphatidic acid binding(GO:0070300) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 4.8 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.0 | 1.9 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.0 | 1.5 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 0.2 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 13.1 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.2 | 4.8 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 0.1 | 1.1 | REACTOME NUCLEOTIDE LIKE PURINERGIC RECEPTORS | Genes involved in Nucleotide-like (purinergic) receptors |
| 0.0 | 0.4 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.3 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.0 | 0.5 | REACTOME CTLA4 INHIBITORY SIGNALING | Genes involved in CTLA4 inhibitory signaling |
| 0.0 | 0.2 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.0 | 0.5 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 0.1 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |