Project
avrg: 12D miR HR13_24
Navigation
Downloads
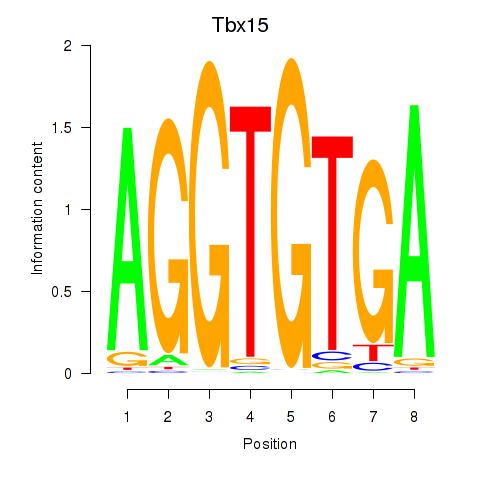
Results for Tbx15
Z-value: 2.98
Transcription factors associated with Tbx15
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Tbx15
|
ENSMUSG00000027868.5 | T-box 15 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Tbx15 | mm10_v2_chr3_+_99253754_99253807 | 0.83 | 1.4e-03 | Click! |
Activity profile of Tbx15 motif
Sorted Z-values of Tbx15 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_129121889 | 10.62 |
ENSMUST00000139450.1
ENSMUST00000125931.1 ENSMUST00000116444.2 |
Hpca
|
hippocalcin |
| chr12_+_109459843 | 8.60 |
ENSMUST00000173812.1
|
Dlk1
|
delta-like 1 homolog (Drosophila) |
| chr10_+_4710119 | 5.49 |
ENSMUST00000105588.1
ENSMUST00000105589.1 |
Esr1
|
estrogen receptor 1 (alpha) |
| chr15_+_103503261 | 4.71 |
ENSMUST00000023132.3
|
Pde1b
|
phosphodiesterase 1B, Ca2+-calmodulin dependent |
| chr5_+_120649188 | 4.34 |
ENSMUST00000156722.1
|
Rasal1
|
RAS protein activator like 1 (GAP1 like) |
| chr7_+_46396439 | 3.80 |
ENSMUST00000025202.6
|
Kcnc1
|
potassium voltage gated channel, Shaw-related subfamily, member 1 |
| chr4_-_129121699 | 3.76 |
ENSMUST00000135763.1
ENSMUST00000149763.1 ENSMUST00000164649.1 |
Hpca
|
hippocalcin |
| chr16_-_44333135 | 3.35 |
ENSMUST00000047446.6
|
Sidt1
|
SID1 transmembrane family, member 1 |
| chr16_-_44332925 | 3.31 |
ENSMUST00000136381.1
|
Sidt1
|
SID1 transmembrane family, member 1 |
| chr6_-_24956106 | 3.16 |
ENSMUST00000127247.2
|
Tmem229a
|
transmembrane protein 229A |
| chr16_+_87553313 | 3.14 |
ENSMUST00000026700.7
|
Map3k7cl
|
Map3k7 C-terminal like |
| chr11_+_3330781 | 3.04 |
ENSMUST00000136536.1
ENSMUST00000093399.4 |
Pik3ip1
|
phosphoinositide-3-kinase interacting protein 1 |
| chr7_+_141475240 | 3.00 |
ENSMUST00000026585.7
|
Tspan4
|
tetraspanin 4 |
| chr17_+_84511832 | 2.84 |
ENSMUST00000047206.5
|
Plekhh2
|
pleckstrin homology domain containing, family H (with MyTH4 domain) member 2 |
| chr11_-_69900949 | 2.62 |
ENSMUST00000102580.3
|
2810408A11Rik
|
RIKEN cDNA 2810408A11 gene |
| chr6_+_15196949 | 2.59 |
ENSMUST00000151301.1
ENSMUST00000131414.1 ENSMUST00000140557.1 ENSMUST00000115469.1 |
Foxp2
|
forkhead box P2 |
| chr2_+_91945703 | 2.48 |
ENSMUST00000178895.1
|
Gm9821
|
predicted gene 9821 |
| chr14_-_55116935 | 2.39 |
ENSMUST00000022819.5
|
Jph4
|
junctophilin 4 |
| chr9_+_47530173 | 2.39 |
ENSMUST00000114548.1
ENSMUST00000152459.1 ENSMUST00000143026.1 ENSMUST00000085909.2 ENSMUST00000114547.1 ENSMUST00000034581.3 |
Cadm1
|
cell adhesion molecule 1 |
| chr1_-_172329261 | 2.37 |
ENSMUST00000062387.2
|
Kcnj9
|
potassium inwardly-rectifying channel, subfamily J, member 9 |
| chr18_+_67289235 | 2.31 |
ENSMUST00000025403.6
|
Impa2
|
inositol (myo)-1(or 4)-monophosphatase 2 |
| chr1_-_105356658 | 2.27 |
ENSMUST00000058688.5
ENSMUST00000172299.1 |
Rnf152
|
ring finger protein 152 |
| chr4_-_82885148 | 2.20 |
ENSMUST00000048430.3
|
Cer1
|
cerberus 1 homolog (Xenopus laevis) |
| chr8_-_31918203 | 2.17 |
ENSMUST00000073884.4
|
Nrg1
|
neuregulin 1 |
| chr4_+_134468320 | 2.15 |
ENSMUST00000030636.4
ENSMUST00000127279.1 ENSMUST00000105867.1 |
Stmn1
|
stathmin 1 |
| chr14_+_62292475 | 2.07 |
ENSMUST00000166879.1
|
Rnaseh2b
|
ribonuclease H2, subunit B |
| chr1_+_6730135 | 1.97 |
ENSMUST00000155921.1
|
St18
|
suppression of tumorigenicity 18 |
| chr1_-_87394721 | 1.95 |
ENSMUST00000113212.3
|
Kcnj13
|
potassium inwardly-rectifying channel, subfamily J, member 13 |
| chr18_-_47333311 | 1.94 |
ENSMUST00000126684.1
ENSMUST00000156422.1 |
Sema6a
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6A |
| chr1_-_74749221 | 1.93 |
ENSMUST00000081636.6
|
Prkag3
|
protein kinase, AMP-activated, gamma 3 non-catatlytic subunit |
| chr8_-_4259257 | 1.90 |
ENSMUST00000053252.7
|
Ctxn1
|
cortexin 1 |
| chr11_-_69900930 | 1.89 |
ENSMUST00000018714.6
ENSMUST00000128046.1 |
2810408A11Rik
|
RIKEN cDNA 2810408A11 gene |
| chr13_-_43304153 | 1.89 |
ENSMUST00000055341.5
|
Gfod1
|
glucose-fructose oxidoreductase domain containing 1 |
| chr3_+_69004711 | 1.87 |
ENSMUST00000042901.8
|
Smc4
|
structural maintenance of chromosomes 4 |
| chr9_-_4796218 | 1.85 |
ENSMUST00000027020.6
ENSMUST00000063508.7 ENSMUST00000163309.1 |
Gria4
|
glutamate receptor, ionotropic, AMPA4 (alpha 4) |
| chr13_+_54503779 | 1.82 |
ENSMUST00000121401.1
ENSMUST00000118072.1 ENSMUST00000159721.1 |
Simc1
|
SUMO-interacting motifs containing 1 |
| chr9_-_44721383 | 1.82 |
ENSMUST00000148929.1
ENSMUST00000123406.1 |
Phldb1
|
pleckstrin homology-like domain, family B, member 1 |
| chr4_-_119658781 | 1.75 |
ENSMUST00000106309.2
ENSMUST00000044426.7 |
Guca2b
|
guanylate cyclase activator 2b (retina) |
| chr5_-_24392012 | 1.74 |
ENSMUST00000059401.6
|
Atg9b
|
autophagy related 9B |
| chr11_-_101551837 | 1.65 |
ENSMUST00000017290.4
|
Brca1
|
breast cancer 1 |
| chr7_+_105640448 | 1.62 |
ENSMUST00000058333.3
|
Timm10b
|
translocase of inner mitochondrial membrane 10B |
| chr7_+_105640522 | 1.61 |
ENSMUST00000106785.1
ENSMUST00000106786.1 ENSMUST00000106780.1 ENSMUST00000106784.1 |
Timm10b
|
translocase of inner mitochondrial membrane 10B |
| chrX_-_157492280 | 1.59 |
ENSMUST00000112529.1
|
Sms
|
spermine synthase |
| chrX_-_48034842 | 1.57 |
ENSMUST00000039026.7
|
Apln
|
apelin |
| chrX_-_150657392 | 1.57 |
ENSMUST00000151403.2
ENSMUST00000087253.4 ENSMUST00000112709.1 ENSMUST00000163969.1 ENSMUST00000087258.3 |
Tro
|
trophinin |
| chr8_+_79028587 | 1.57 |
ENSMUST00000119254.1
|
Zfp827
|
zinc finger protein 827 |
| chr7_-_116237767 | 1.56 |
ENSMUST00000182834.1
|
Plekha7
|
pleckstrin homology domain containing, family A member 7 |
| chr5_-_148399901 | 1.55 |
ENSMUST00000048116.8
|
Slc7a1
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 |
| chrX_-_162964557 | 1.54 |
ENSMUST00000038769.2
|
S100g
|
S100 calcium binding protein G |
| chr7_-_132813799 | 1.52 |
ENSMUST00000097998.2
|
Fam53b
|
family with sequence similarity 53, member B |
| chr7_+_130577334 | 1.50 |
ENSMUST00000059145.7
ENSMUST00000084513.4 |
Tacc2
|
transforming, acidic coiled-coil containing protein 2 |
| chr3_-_10208569 | 1.49 |
ENSMUST00000029041.4
|
Fabp4
|
fatty acid binding protein 4, adipocyte |
| chr7_-_141443989 | 1.49 |
ENSMUST00000026580.5
|
Lrdd
|
leucine-rich and death domain containing |
| chr7_-_132813715 | 1.48 |
ENSMUST00000134946.1
|
Fam53b
|
family with sequence similarity 53, member B |
| chr7_+_30291659 | 1.46 |
ENSMUST00000014065.8
ENSMUST00000150892.1 ENSMUST00000126216.1 |
Clip3
|
CAP-GLY domain containing linker protein 3 |
| chr7_+_48789003 | 1.45 |
ENSMUST00000118927.1
ENSMUST00000125280.1 |
Zdhhc13
|
zinc finger, DHHC domain containing 13 |
| chr14_+_45351473 | 1.43 |
ENSMUST00000111835.2
|
Styx
|
serine/threonine/tyrosine interaction protein |
| chrX_-_7947763 | 1.42 |
ENSMUST00000154244.1
|
Hdac6
|
histone deacetylase 6 |
| chr8_+_45885479 | 1.38 |
ENSMUST00000034053.5
|
Pdlim3
|
PDZ and LIM domain 3 |
| chr9_-_21760275 | 1.38 |
ENSMUST00000098942.4
|
Spc24
|
SPC24, NDC80 kinetochore complex component, homolog (S. cerevisiae) |
| chr4_+_150237694 | 1.37 |
ENSMUST00000141931.1
|
Eno1
|
enolase 1, alpha non-neuron |
| chr18_-_42899294 | 1.37 |
ENSMUST00000117687.1
|
Ppp2r2b
|
protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), beta isoform |
| chr7_+_18987518 | 1.36 |
ENSMUST00000063563.7
|
Nanos2
|
nanos homolog 2 (Drosophila) |
| chr5_+_115908644 | 1.34 |
ENSMUST00000141101.1
|
Cit
|
citron |
| chr11_-_69900886 | 1.34 |
ENSMUST00000108621.2
ENSMUST00000100969.2 |
2810408A11Rik
|
RIKEN cDNA 2810408A11 gene |
| chrX_-_150657366 | 1.34 |
ENSMUST00000148604.1
|
Tro
|
trophinin |
| chr12_-_31634592 | 1.33 |
ENSMUST00000020979.7
ENSMUST00000177962.1 |
Bcap29
|
B cell receptor associated protein 29 |
| chr7_+_28129459 | 1.25 |
ENSMUST00000059886.5
|
9530053A07Rik
|
RIKEN cDNA 9530053A07 gene |
| chr5_+_108065696 | 1.24 |
ENSMUST00000172045.1
|
Mtf2
|
metal response element binding transcription factor 2 |
| chr17_+_35861318 | 1.24 |
ENSMUST00000074259.8
ENSMUST00000174873.1 |
Nrm
|
nurim (nuclear envelope membrane protein) |
| chr6_+_50110186 | 1.23 |
ENSMUST00000166318.1
ENSMUST00000036236.8 ENSMUST00000036225.8 |
Mpp6
|
membrane protein, palmitoylated 6 (MAGUK p55 subfamily member 6) |
| chr4_+_148000722 | 1.20 |
ENSMUST00000103230.4
|
Nppa
|
natriuretic peptide type A |
| chr11_+_62077018 | 1.19 |
ENSMUST00000092415.5
|
Specc1
|
sperm antigen with calponin homology and coiled-coil domains 1 |
| chr11_+_23306910 | 1.19 |
ENSMUST00000137823.1
|
Usp34
|
ubiquitin specific peptidase 34 |
| chr19_-_10304867 | 1.19 |
ENSMUST00000039327.4
|
Dagla
|
diacylglycerol lipase, alpha |
| chr19_-_46327121 | 1.18 |
ENSMUST00000041391.4
ENSMUST00000096029.5 |
Psd
|
pleckstrin and Sec7 domain containing |
| chr14_-_118925314 | 1.18 |
ENSMUST00000004055.8
|
Dzip1
|
DAZ interacting protein 1 |
| chr1_+_153665274 | 1.17 |
ENSMUST00000152114.1
ENSMUST00000111812.1 |
Rgs8
|
regulator of G-protein signaling 8 |
| chr8_+_122422020 | 1.17 |
ENSMUST00000050963.3
|
Il17c
|
interleukin 17C |
| chr7_-_105640308 | 1.15 |
ENSMUST00000133519.1
ENSMUST00000084782.2 ENSMUST00000131446.1 |
Arfip2
|
ADP-ribosylation factor interacting protein 2 |
| chr4_-_41517326 | 1.13 |
ENSMUST00000030152.6
ENSMUST00000095126.4 |
1110017D15Rik
|
RIKEN cDNA 1110017D15 gene |
| chr17_+_32036098 | 1.12 |
ENSMUST00000081339.6
|
Rrp1b
|
ribosomal RNA processing 1 homolog B (S. cerevisiae) |
| chr19_-_28963863 | 1.11 |
ENSMUST00000161813.1
|
4430402I18Rik
|
RIKEN cDNA 4430402I18 gene |
| chr2_-_13011747 | 1.10 |
ENSMUST00000061545.5
|
C1ql3
|
C1q-like 3 |
| chr2_+_163820832 | 1.09 |
ENSMUST00000029188.7
|
Wisp2
|
WNT1 inducible signaling pathway protein 2 |
| chr16_-_18413452 | 1.09 |
ENSMUST00000165430.1
ENSMUST00000147720.1 |
Comt
|
catechol-O-methyltransferase |
| chr7_-_28302238 | 1.09 |
ENSMUST00000108315.3
|
Dll3
|
delta-like 3 (Drosophila) |
| chr5_-_148371525 | 1.07 |
ENSMUST00000138596.1
|
Slc7a1
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 |
| chr5_+_142702091 | 1.06 |
ENSMUST00000058418.7
|
Slc29a4
|
solute carrier family 29 (nucleoside transporters), member 4 |
| chr12_-_3426700 | 1.06 |
ENSMUST00000180149.1
|
1110002L01Rik
|
RIKEN cDNA 1110002L01 gene |
| chr12_+_102948843 | 1.06 |
ENSMUST00000101099.5
|
Unc79
|
unc-79 homolog (C. elegans) |
| chr6_+_50110837 | 1.04 |
ENSMUST00000167628.1
|
Mpp6
|
membrane protein, palmitoylated 6 (MAGUK p55 subfamily member 6) |
| chr4_+_42950369 | 1.04 |
ENSMUST00000084662.5
|
Dnajb5
|
DnaJ (Hsp40) homolog, subfamily B, member 5 |
| chr1_+_63273261 | 1.04 |
ENSMUST00000114132.1
ENSMUST00000126932.1 |
Zdbf2
|
zinc finger, DBF-type containing 2 |
| chr1_+_6730051 | 1.03 |
ENSMUST00000043578.6
ENSMUST00000131467.1 ENSMUST00000150761.1 ENSMUST00000151281.1 |
St18
|
suppression of tumorigenicity 18 |
| chr5_+_114444266 | 1.03 |
ENSMUST00000043760.8
ENSMUST00000112239.2 ENSMUST00000125650.1 |
Mvk
|
mevalonate kinase |
| chr7_-_132813528 | 1.03 |
ENSMUST00000097999.2
|
Fam53b
|
family with sequence similarity 53, member B |
| chr16_-_32810477 | 1.03 |
ENSMUST00000179384.2
|
Gm933
|
predicted gene 933 |
| chr2_+_156775409 | 1.02 |
ENSMUST00000088552.6
|
Myl9
|
myosin, light polypeptide 9, regulatory |
| chrX_-_7947553 | 1.02 |
ENSMUST00000133349.1
|
Hdac6
|
histone deacetylase 6 |
| chr7_+_28129804 | 1.01 |
ENSMUST00000150948.1
|
9530053A07Rik
|
RIKEN cDNA 9530053A07 gene |
| chr11_+_23306884 | 0.99 |
ENSMUST00000180046.1
|
Usp34
|
ubiquitin specific peptidase 34 |
| chr2_-_30178422 | 0.99 |
ENSMUST00000100220.4
ENSMUST00000179795.1 |
D2Wsu81e
|
DNA segment, Chr 2, Wayne State University 81, expressed |
| chr14_+_27238018 | 0.97 |
ENSMUST00000049206.5
|
Arhgef3
|
Rho guanine nucleotide exchange factor (GEF) 3 |
| chr4_+_11191726 | 0.97 |
ENSMUST00000029866.9
ENSMUST00000108324.3 |
Ccne2
|
cyclin E2 |
| chr1_+_153665666 | 0.96 |
ENSMUST00000111814.1
ENSMUST00000111810.1 |
Rgs8
|
regulator of G-protein signaling 8 |
| chr1_-_167285110 | 0.94 |
ENSMUST00000027839.8
|
Uck2
|
uridine-cytidine kinase 2 |
| chr3_-_88913885 | 0.93 |
ENSMUST00000107494.1
|
Msto1
|
misato homolog 1 (Drosophila) |
| chrX_+_133908418 | 0.93 |
ENSMUST00000033606.8
ENSMUST00000113303.1 ENSMUST00000165805.1 |
Srpx2
|
sushi-repeat-containing protein, X-linked 2 |
| chr10_-_120899067 | 0.93 |
ENSMUST00000092143.5
|
Msrb3
|
methionine sulfoxide reductase B3 |
| chrX_-_162829379 | 0.92 |
ENSMUST00000041370.4
ENSMUST00000112316.2 ENSMUST00000112315.1 |
Txlng
|
taxilin gamma |
| chr7_-_109170308 | 0.90 |
ENSMUST00000036992.7
|
Lmo1
|
LIM domain only 1 |
| chr7_+_140125651 | 0.90 |
ENSMUST00000026537.5
ENSMUST00000097967.3 |
Paox
|
polyamine oxidase (exo-N4-amino) |
| chr11_+_96323253 | 0.90 |
ENSMUST00000093944.3
|
Hoxb3
|
homeobox B3 |
| chr1_+_17601893 | 0.90 |
ENSMUST00000088476.2
|
Pi15
|
peptidase inhibitor 15 |
| chr11_+_98741871 | 0.89 |
ENSMUST00000103139.4
|
Thra
|
thyroid hormone receptor alpha |
| chr7_-_78577771 | 0.88 |
ENSMUST00000039438.7
|
Ntrk3
|
neurotrophic tyrosine kinase, receptor, type 3 |
| chr8_-_62123106 | 0.88 |
ENSMUST00000034052.6
ENSMUST00000034054.7 |
Anxa10
|
annexin A10 |
| chrX_-_7947848 | 0.88 |
ENSMUST00000115642.1
ENSMUST00000033501.8 ENSMUST00000145675.1 |
Hdac6
|
histone deacetylase 6 |
| chr5_-_96161742 | 0.88 |
ENSMUST00000129646.1
ENSMUST00000113005.2 ENSMUST00000154500.1 ENSMUST00000141383.1 |
Cnot6l
|
CCR4-NOT transcription complex, subunit 6-like |
| chr1_+_132298606 | 0.87 |
ENSMUST00000046071.4
|
Klhdc8a
|
kelch domain containing 8A |
| chr5_+_112343068 | 0.87 |
ENSMUST00000112359.2
ENSMUST00000035279.3 |
Hps4
|
Hermansky-Pudlak syndrome 4 homolog (human) |
| chr10_-_81472859 | 0.87 |
ENSMUST00000147524.1
ENSMUST00000119060.1 |
Celf5
|
CUGBP, Elav-like family member 5 |
| chr2_+_25242929 | 0.87 |
ENSMUST00000114355.1
ENSMUST00000060818.1 |
Rnf208
|
ring finger protein 208 |
| chr11_+_86683985 | 0.87 |
ENSMUST00000108022.1
ENSMUST00000108021.1 |
Ptrh2
|
peptidyl-tRNA hydrolase 2 |
| chr12_+_3891728 | 0.86 |
ENSMUST00000172689.1
ENSMUST00000111186.1 |
Dnmt3a
|
DNA methyltransferase 3A |
| chr14_-_55106547 | 0.86 |
ENSMUST00000036041.8
|
Ap1g2
|
adaptor protein complex AP-1, gamma 2 subunit |
| chr12_+_105336922 | 0.86 |
ENSMUST00000180503.1
|
2810011L19Rik
|
RIKEN cDNA 2810011L19 gene |
| chr12_-_56345862 | 0.85 |
ENSMUST00000021416.7
|
Mbip
|
MAP3K12 binding inhibitory protein 1 |
| chr17_+_46496753 | 0.84 |
ENSMUST00000046497.6
|
Dnph1
|
2'-deoxynucleoside 5'-phosphate N-hydrolase 1 |
| chr17_+_80944611 | 0.83 |
ENSMUST00000025092.4
|
Tmem178
|
transmembrane protein 178 |
| chr11_+_96316684 | 0.83 |
ENSMUST00000049241.7
|
Hoxb4
|
homeobox B4 |
| chr10_-_127263346 | 0.83 |
ENSMUST00000099172.3
|
Kif5a
|
kinesin family member 5A |
| chr11_-_86544754 | 0.83 |
ENSMUST00000138810.1
ENSMUST00000058286.2 ENSMUST00000154617.1 |
Rps6kb1
|
ribosomal protein S6 kinase, polypeptide 1 |
| chr9_+_108296853 | 0.82 |
ENSMUST00000035230.5
|
Amt
|
aminomethyltransferase |
| chr1_+_156558844 | 0.82 |
ENSMUST00000166172.2
ENSMUST00000027888.6 |
Abl2
|
v-abl Abelson murine leukemia viral oncogene 2 (arg, Abelson-related gene) |
| chr8_+_4243264 | 0.81 |
ENSMUST00000110996.1
|
Map2k7
|
mitogen-activated protein kinase kinase 7 |
| chr2_-_64975762 | 0.81 |
ENSMUST00000156765.1
|
Grb14
|
growth factor receptor bound protein 14 |
| chr3_-_82074639 | 0.80 |
ENSMUST00000029635.8
|
Gucy1b3
|
guanylate cyclase 1, soluble, beta 3 |
| chr8_+_79028317 | 0.80 |
ENSMUST00000087927.4
ENSMUST00000098614.2 |
Zfp827
|
zinc finger protein 827 |
| chr11_+_70018728 | 0.78 |
ENSMUST00000018700.6
ENSMUST00000134376.2 |
Dlg4
|
discs, large homolog 4 (Drosophila) |
| chr2_+_138278481 | 0.78 |
ENSMUST00000075410.4
|
Btbd3
|
BTB (POZ) domain containing 3 |
| chr12_+_51348265 | 0.77 |
ENSMUST00000119211.1
|
G2e3
|
G2/M-phase specific E3 ubiquitin ligase |
| chrX_+_75095854 | 0.77 |
ENSMUST00000033776.8
|
Dkc1
|
dyskeratosis congenita 1, dyskerin |
| chr11_+_70639118 | 0.77 |
ENSMUST00000055184.6
ENSMUST00000108551.2 |
Gp1ba
|
glycoprotein 1b, alpha polypeptide |
| chr7_+_111028951 | 0.76 |
ENSMUST00000005749.5
|
Ctr9
|
Ctr9, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) |
| chr4_-_57300362 | 0.76 |
ENSMUST00000153926.1
|
Ptpn3
|
protein tyrosine phosphatase, non-receptor type 3 |
| chr6_-_94700137 | 0.75 |
ENSMUST00000101126.2
ENSMUST00000032105.4 |
Lrig1
|
leucine-rich repeats and immunoglobulin-like domains 1 |
| chr8_-_125898291 | 0.75 |
ENSMUST00000047239.6
|
Pcnxl2
|
pecanex-like 2 (Drosophila) |
| chr1_+_170214826 | 0.73 |
ENSMUST00000159201.1
ENSMUST00000055830.1 |
4930500M09Rik
|
RIKEN cDNA 4930500M09 gene |
| chr10_-_128645784 | 0.73 |
ENSMUST00000065334.3
|
Ikzf4
|
IKAROS family zinc finger 4 |
| chr17_+_35135695 | 0.72 |
ENSMUST00000174478.1
ENSMUST00000174281.2 ENSMUST00000173550.1 |
Bag6
|
BCL2-associated athanogene 6 |
| chr16_+_57353093 | 0.72 |
ENSMUST00000159816.1
|
Filip1l
|
filamin A interacting protein 1-like |
| chr4_+_130047840 | 0.71 |
ENSMUST00000044565.8
ENSMUST00000132251.1 |
Col16a1
|
collagen, type XVI, alpha 1 |
| chr14_+_70077375 | 0.70 |
ENSMUST00000035908.1
|
Egr3
|
early growth response 3 |
| chr3_+_104781048 | 0.70 |
ENSMUST00000002298.6
|
Ppm1j
|
protein phosphatase 1J |
| chr19_+_6057888 | 0.70 |
ENSMUST00000043074.5
ENSMUST00000178310.1 |
Fau
|
Finkel-Biskis-Reilly murine sarcoma virus (FBR-MuSV) ubiquitously expressed (fox derived) |
| chr11_-_22286795 | 0.70 |
ENSMUST00000109563.2
ENSMUST00000180360.1 |
Ehbp1
|
EH domain binding protein 1 |
| chr1_+_161494649 | 0.69 |
ENSMUST00000086084.1
|
Tnfsf18
|
tumor necrosis factor (ligand) superfamily, member 18 |
| chr19_-_44146433 | 0.69 |
ENSMUST00000079033.4
|
Bloc1s2
|
biogenesis of lysosome-related organelles complex-1, subunit 2 |
| chrM_+_9452 | 0.69 |
ENSMUST00000082411.1
|
mt-Nd3
|
mitochondrially encoded NADH dehydrogenase 3 |
| chr4_-_132510493 | 0.68 |
ENSMUST00000030724.8
|
Sesn2
|
sestrin 2 |
| chr14_-_70177668 | 0.68 |
ENSMUST00000022681.4
|
Pdlim2
|
PDZ and LIM domain 2 |
| chr16_-_50330987 | 0.68 |
ENSMUST00000114488.1
|
Bbx
|
bobby sox homolog (Drosophila) |
| chr1_-_74304120 | 0.67 |
ENSMUST00000141560.1
|
Tmbim1
|
transmembrane BAX inhibitor motif containing 1 |
| chr4_+_11191354 | 0.67 |
ENSMUST00000170901.1
|
Ccne2
|
cyclin E2 |
| chr5_+_107331157 | 0.66 |
ENSMUST00000031215.8
ENSMUST00000112677.3 |
Brdt
|
bromodomain, testis-specific |
| chr11_-_97575210 | 0.66 |
ENSMUST00000107596.2
|
Srcin1
|
SRC kinase signaling inhibitor 1 |
| chr14_-_70524068 | 0.66 |
ENSMUST00000022692.3
|
Sftpc
|
surfactant associated protein C |
| chr2_+_25242227 | 0.66 |
ENSMUST00000154498.1
|
Rnf208
|
ring finger protein 208 |
| chr11_+_70018421 | 0.65 |
ENSMUST00000108588.1
|
Dlg4
|
discs, large homolog 4 (Drosophila) |
| chr12_+_51348370 | 0.65 |
ENSMUST00000121521.1
|
G2e3
|
G2/M-phase specific E3 ubiquitin ligase |
| chr7_-_98162318 | 0.64 |
ENSMUST00000107112.1
|
Capn5
|
calpain 5 |
| chr11_-_120731944 | 0.64 |
ENSMUST00000154565.1
ENSMUST00000026148.2 |
Cbr2
|
carbonyl reductase 2 |
| chr11_-_120630126 | 0.64 |
ENSMUST00000106180.1
|
Mafg
|
v-maf musculoaponeurotic fibrosarcoma oncogene family, protein G (avian) |
| chr12_+_73123709 | 0.64 |
ENSMUST00000021523.6
|
Mnat1
|
menage a trois 1 |
| chr3_-_145649970 | 0.63 |
ENSMUST00000029846.3
|
Cyr61
|
cysteine rich protein 61 |
| chr4_+_42114817 | 0.63 |
ENSMUST00000098123.3
|
Gm13304
|
predicted gene 13304 |
| chr19_-_43524462 | 0.63 |
ENSMUST00000026196.7
|
Got1
|
glutamate oxaloacetate transaminase 1, soluble |
| chr3_-_96905294 | 0.63 |
ENSMUST00000029738.7
|
Gpr89
|
G protein-coupled receptor 89 |
| chr15_-_12592556 | 0.62 |
ENSMUST00000075317.5
|
Pdzd2
|
PDZ domain containing 2 |
| chr12_+_33314277 | 0.62 |
ENSMUST00000133549.1
|
Atxn7l1
|
ataxin 7-like 1 |
| chr4_+_41903610 | 0.61 |
ENSMUST00000098128.3
|
Gm21541
|
predicted gene, 21541 |
| chr19_+_6057925 | 0.61 |
ENSMUST00000179142.1
|
Fau
|
Finkel-Biskis-Reilly murine sarcoma virus (FBR-MuSV) ubiquitously expressed (fox derived) |
| chrX_+_42151002 | 0.60 |
ENSMUST00000123245.1
|
Stag2
|
stromal antigen 2 |
| chr7_-_25072287 | 0.60 |
ENSMUST00000003468.8
|
Grik5
|
glutamate receptor, ionotropic, kainate 5 (gamma 2) |
| chr17_+_37529957 | 0.60 |
ENSMUST00000097325.3
|
Olfr111
|
olfactory receptor 111 |
| chr4_-_117156144 | 0.60 |
ENSMUST00000102696.4
|
Rps8
|
ribosomal protein S8 |
| chr17_-_6827990 | 0.60 |
ENSMUST00000181895.1
|
Gm2885
|
predicted gene 2885 |
| chr15_-_11905609 | 0.60 |
ENSMUST00000066529.3
|
Npr3
|
natriuretic peptide receptor 3 |
| chr9_+_109082485 | 0.60 |
ENSMUST00000026735.7
|
Ccdc51
|
coiled-coil domain containing 51 |
| chr13_+_29014399 | 0.59 |
ENSMUST00000146336.1
ENSMUST00000130109.1 |
A330102I10Rik
|
RIKEN cDNA A330102I10 gene |
| chr11_+_43682038 | 0.59 |
ENSMUST00000094294.4
|
Pwwp2a
|
PWWP domain containing 2A |
| chr17_+_47630690 | 0.58 |
ENSMUST00000024779.8
|
Usp49
|
ubiquitin specific peptidase 49 |
| chr6_+_115134899 | 0.58 |
ENSMUST00000009538.5
ENSMUST00000169345.1 |
Syn2
|
synapsin II |
| chr7_+_141228766 | 0.58 |
ENSMUST00000106027.2
|
Phrf1
|
PHD and ring finger domains 1 |
| chr15_-_43282695 | 0.58 |
ENSMUST00000022960.2
|
Eif3e
|
eukaryotic translation initiation factor 3, subunit E |
| chr15_+_100469034 | 0.58 |
ENSMUST00000037001.8
|
Letmd1
|
LETM1 domain containing 1 |
| chr1_-_44061936 | 0.57 |
ENSMUST00000168641.1
|
Gm8251
|
predicted gene 8251 |
| chr11_+_78032274 | 0.57 |
ENSMUST00000021187.5
|
Dhrs13
|
dehydrogenase/reductase (SDR family) member 13 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Tbx15
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.8 | 14.4 | GO:0031283 | negative regulation of cGMP metabolic process(GO:0030824) negative regulation of cGMP biosynthetic process(GO:0030827) negative regulation of guanylate cyclase activity(GO:0031283) |
| 1.1 | 3.3 | GO:0070843 | misfolded protein transport(GO:0070843) polyubiquitinated protein transport(GO:0070844) polyubiquitinated misfolded protein transport(GO:0070845) Hsp90 deacetylation(GO:0070846) |
| 1.1 | 5.5 | GO:0060523 | prostate epithelial cord elongation(GO:0060523) |
| 0.8 | 0.8 | GO:0099543 | trans-synaptic signaling by soluble gas(GO:0099543) |
| 0.8 | 2.4 | GO:0009826 | unidimensional cell growth(GO:0009826) susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.7 | 6.7 | GO:0033227 | dsRNA transport(GO:0033227) |
| 0.6 | 1.7 | GO:2000620 | positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 0.5 | 2.2 | GO:0035582 | sequestering of BMP in extracellular matrix(GO:0035582) |
| 0.5 | 1.6 | GO:1904020 | corticotropin-releasing hormone secretion(GO:0043396) regulation of corticotropin-releasing hormone secretion(GO:0043397) positive regulation of corticotropin-releasing hormone secretion(GO:0051466) regulation of G-protein coupled receptor internalization(GO:1904020) |
| 0.5 | 4.7 | GO:0036006 | response to macrophage colony-stimulating factor(GO:0036005) cellular response to macrophage colony-stimulating factor stimulus(GO:0036006) |
| 0.5 | 1.4 | GO:0017055 | negative regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0017055) |
| 0.5 | 1.4 | GO:0045204 | MAPK export from nucleus(GO:0045204) |
| 0.4 | 1.3 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.4 | 3.0 | GO:0043553 | negative regulation of phosphatidylinositol 3-kinase activity(GO:0043553) |
| 0.4 | 1.2 | GO:0098917 | retrograde trans-synaptic signaling(GO:0098917) |
| 0.4 | 5.0 | GO:1903818 | positive regulation of voltage-gated potassium channel activity(GO:1903818) |
| 0.4 | 1.1 | GO:0031335 | regulation of sulfur amino acid metabolic process(GO:0031335) |
| 0.3 | 1.7 | GO:0044805 | late nucleophagy(GO:0044805) |
| 0.3 | 1.3 | GO:1902045 | regulation of Fas signaling pathway(GO:1902044) negative regulation of Fas signaling pathway(GO:1902045) |
| 0.3 | 1.7 | GO:0061086 | negative regulation of histone H3-K27 methylation(GO:0061086) |
| 0.3 | 9.2 | GO:0046852 | positive regulation of bone resorption(GO:0045780) positive regulation of bone remodeling(GO:0046852) |
| 0.3 | 0.9 | GO:0006222 | UMP biosynthetic process(GO:0006222) pyrimidine ribonucleoside monophosphate metabolic process(GO:0009173) pyrimidine ribonucleoside monophosphate biosynthetic process(GO:0009174) UMP metabolic process(GO:0046049) |
| 0.3 | 2.5 | GO:0008215 | spermine metabolic process(GO:0008215) |
| 0.3 | 2.2 | GO:0045213 | neurotransmitter receptor metabolic process(GO:0045213) |
| 0.3 | 2.1 | GO:0070494 | regulation of thrombin receptor signaling pathway(GO:0070494) negative regulation of thrombin receptor signaling pathway(GO:0070495) |
| 0.3 | 1.2 | GO:0048687 | positive regulation of sprouting of injured axon(GO:0048687) positive regulation of axon extension involved in regeneration(GO:0048691) |
| 0.3 | 3.4 | GO:2001269 | positive regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:2001269) |
| 0.3 | 0.8 | GO:0098971 | anterograde dendritic transport of neurotransmitter receptor complex(GO:0098971) |
| 0.3 | 0.8 | GO:0006546 | glycine catabolic process(GO:0006546) glycine decarboxylation via glycine cleavage system(GO:0019464) |
| 0.3 | 0.8 | GO:1903538 | meiotic cell cycle process involved in oocyte maturation(GO:1903537) regulation of meiotic cell cycle process involved in oocyte maturation(GO:1903538) |
| 0.3 | 1.6 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.3 | 1.0 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.3 | 0.8 | GO:2001160 | regulation of histone H3-K79 methylation(GO:2001160) |
| 0.2 | 0.7 | GO:0032474 | otolith morphogenesis(GO:0032474) |
| 0.2 | 1.9 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.2 | 0.9 | GO:0030091 | protein repair(GO:0030091) |
| 0.2 | 0.7 | GO:2000328 | regulation of T-helper 17 cell lineage commitment(GO:2000328) |
| 0.2 | 0.7 | GO:0090526 | regulation of gluconeogenesis involved in cellular glucose homeostasis(GO:0090526) |
| 0.2 | 1.1 | GO:0007386 | compartment pattern specification(GO:0007386) |
| 0.2 | 0.6 | GO:0006114 | fumarate metabolic process(GO:0006106) glycerol biosynthetic process(GO:0006114) aspartate biosynthetic process(GO:0006532) aspartate catabolic process(GO:0006533) |
| 0.2 | 0.8 | GO:0051581 | negative regulation of neurotransmitter uptake(GO:0051581) negative regulation of serotonin uptake(GO:0051612) |
| 0.2 | 2.6 | GO:0060501 | positive regulation of epithelial cell proliferation involved in lung morphogenesis(GO:0060501) |
| 0.2 | 1.4 | GO:0044838 | cell quiescence(GO:0044838) |
| 0.2 | 2.3 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.2 | 0.4 | GO:1900157 | regulation of bone mineralization involved in bone maturation(GO:1900157) |
| 0.2 | 1.5 | GO:0072321 | chaperone-mediated protein transport(GO:0072321) |
| 0.2 | 0.5 | GO:1902277 | negative regulation of pancreatic amylase secretion(GO:1902277) |
| 0.2 | 0.9 | GO:0021615 | glossopharyngeal nerve morphogenesis(GO:0021615) |
| 0.2 | 0.5 | GO:2000547 | regulation of dendritic cell dendrite assembly(GO:2000547) |
| 0.2 | 2.6 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.2 | 1.5 | GO:0071285 | cellular response to lithium ion(GO:0071285) |
| 0.2 | 1.8 | GO:2001197 | regulation of basement membrane assembly involved in embryonic body morphogenesis(GO:1904259) positive regulation of basement membrane assembly involved in embryonic body morphogenesis(GO:1904261) basement membrane assembly involved in embryonic body morphogenesis(GO:2001197) |
| 0.2 | 2.6 | GO:0015809 | arginine transport(GO:0015809) |
| 0.2 | 0.5 | GO:0035934 | corticosterone secretion(GO:0035934) regulation of corticosterone secretion(GO:2000852) |
| 0.2 | 1.0 | GO:0060591 | chondroblast differentiation(GO:0060591) |
| 0.2 | 0.5 | GO:0034476 | U1 snRNA 3'-end processing(GO:0034473) U5 snRNA 3'-end processing(GO:0034476) |
| 0.2 | 0.8 | GO:0000454 | snoRNA guided rRNA pseudouridine synthesis(GO:0000454) telomerase RNA stabilization(GO:0090669) |
| 0.2 | 2.3 | GO:1904262 | negative regulation of TORC1 signaling(GO:1904262) |
| 0.1 | 0.4 | GO:0098735 | positive regulation of the force of heart contraction(GO:0098735) |
| 0.1 | 0.9 | GO:2000210 | positive regulation of anoikis(GO:2000210) |
| 0.1 | 0.4 | GO:2000504 | positive regulation of blood vessel remodeling(GO:2000504) |
| 0.1 | 0.6 | GO:0045903 | positive regulation of translational fidelity(GO:0045903) |
| 0.1 | 0.8 | GO:0048633 | positive regulation of skeletal muscle tissue growth(GO:0048633) |
| 0.1 | 3.4 | GO:1903861 | positive regulation of dendrite extension(GO:1903861) |
| 0.1 | 1.9 | GO:0007213 | G-protein coupled acetylcholine receptor signaling pathway(GO:0007213) regulation of dopamine receptor signaling pathway(GO:0060159) |
| 0.1 | 1.4 | GO:0099628 | receptor diffusion trapping(GO:0098953) postsynaptic neurotransmitter receptor diffusion trapping(GO:0098970) neurotransmitter receptor diffusion trapping(GO:0099628) |
| 0.1 | 0.5 | GO:0035522 | monoubiquitinated histone deubiquitination(GO:0035521) monoubiquitinated histone H2A deubiquitination(GO:0035522) |
| 0.1 | 1.4 | GO:0043653 | mitochondrial fragmentation involved in apoptotic process(GO:0043653) |
| 0.1 | 0.9 | GO:0044027 | hypermethylation of CpG island(GO:0044027) |
| 0.1 | 0.4 | GO:0044208 | 'de novo' AMP biosynthetic process(GO:0044208) |
| 0.1 | 0.2 | GO:0038095 | Fc-epsilon receptor signaling pathway(GO:0038095) |
| 0.1 | 0.8 | GO:0043545 | molybdopterin cofactor biosynthetic process(GO:0032324) molybdopterin cofactor metabolic process(GO:0043545) prosthetic group metabolic process(GO:0051189) |
| 0.1 | 0.4 | GO:2000041 | regulation of planar cell polarity pathway involved in axis elongation(GO:2000040) negative regulation of planar cell polarity pathway involved in axis elongation(GO:2000041) |
| 0.1 | 0.4 | GO:0045014 | carbon catabolite repression of transcription(GO:0045013) negative regulation of transcription by glucose(GO:0045014) |
| 0.1 | 0.6 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.1 | 1.5 | GO:0030953 | astral microtubule organization(GO:0030953) |
| 0.1 | 4.3 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.1 | 0.5 | GO:0044268 | multicellular organismal protein metabolic process(GO:0044268) |
| 0.1 | 1.1 | GO:0048149 | behavioral response to ethanol(GO:0048149) |
| 0.1 | 0.5 | GO:0021658 | rhombomere 3 morphogenesis(GO:0021658) |
| 0.1 | 0.4 | GO:0003273 | cell migration involved in endocardial cushion formation(GO:0003273) condensed mesenchymal cell proliferation(GO:0072137) |
| 0.1 | 2.1 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 0.6 | GO:0015867 | ATP transport(GO:0015867) |
| 0.1 | 0.6 | GO:0002158 | osteoclast proliferation(GO:0002158) |
| 0.1 | 0.2 | GO:0021569 | rhombomere 3 development(GO:0021569) |
| 0.1 | 0.9 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.1 | 0.4 | GO:0072108 | positive regulation of mesenchymal to epithelial transition involved in metanephros morphogenesis(GO:0072108) |
| 0.1 | 0.5 | GO:1903232 | melanosome assembly(GO:1903232) |
| 0.1 | 0.8 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.1 | 0.8 | GO:0009162 | deoxyribonucleoside monophosphate metabolic process(GO:0009162) |
| 0.1 | 0.6 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.1 | 1.3 | GO:0071712 | ER-associated misfolded protein catabolic process(GO:0071712) |
| 0.1 | 0.8 | GO:1900042 | positive regulation of interleukin-2 secretion(GO:1900042) |
| 0.1 | 0.8 | GO:2000671 | regulation of motor neuron apoptotic process(GO:2000671) |
| 0.1 | 0.5 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 0.1 | 0.5 | GO:0090214 | spongiotrophoblast layer developmental growth(GO:0090214) |
| 0.1 | 4.6 | GO:0043666 | regulation of phosphoprotein phosphatase activity(GO:0043666) |
| 0.1 | 0.3 | GO:0051866 | general adaptation syndrome(GO:0051866) |
| 0.1 | 1.9 | GO:2001224 | positive regulation of neuron migration(GO:2001224) |
| 0.1 | 0.5 | GO:0010626 | negative regulation of Schwann cell proliferation(GO:0010626) |
| 0.1 | 3.5 | GO:0035411 | catenin import into nucleus(GO:0035411) |
| 0.1 | 3.1 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.1 | 0.5 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.1 | 1.3 | GO:0050774 | negative regulation of dendrite morphogenesis(GO:0050774) |
| 0.1 | 0.2 | GO:1903588 | negative regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903588) |
| 0.1 | 0.2 | GO:0019043 | establishment of viral latency(GO:0019043) |
| 0.1 | 0.5 | GO:0097151 | positive regulation of inhibitory postsynaptic potential(GO:0097151) |
| 0.1 | 1.2 | GO:0030223 | neutrophil differentiation(GO:0030223) |
| 0.1 | 1.3 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 0.1 | 0.3 | GO:0060830 | ciliary receptor clustering involved in smoothened signaling pathway(GO:0060830) |
| 0.1 | 0.5 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.1 | 2.2 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
| 0.1 | 0.5 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.1 | 0.2 | GO:0001928 | regulation of exocyst assembly(GO:0001928) |
| 0.1 | 1.8 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 0.9 | GO:0071625 | vocalization behavior(GO:0071625) |
| 0.1 | 0.3 | GO:0051835 | positive regulation of synapse structural plasticity(GO:0051835) |
| 0.1 | 1.9 | GO:0051968 | positive regulation of synaptic transmission, glutamatergic(GO:0051968) |
| 0.1 | 0.3 | GO:1901069 | guanosine-containing compound catabolic process(GO:1901069) |
| 0.1 | 0.2 | GO:0001777 | T cell homeostatic proliferation(GO:0001777) regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.1 | 0.5 | GO:0014807 | regulation of somitogenesis(GO:0014807) |
| 0.1 | 0.6 | GO:1902416 | positive regulation of mRNA binding(GO:1902416) |
| 0.1 | 0.4 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.1 | 0.2 | GO:0006041 | glucosamine metabolic process(GO:0006041) |
| 0.1 | 2.5 | GO:0030835 | negative regulation of actin filament depolymerization(GO:0030835) |
| 0.1 | 0.6 | GO:0010944 | negative regulation of transcription by competitive promoter binding(GO:0010944) |
| 0.1 | 0.7 | GO:2000400 | positive regulation of T cell differentiation in thymus(GO:0033089) positive regulation of thymocyte aggregation(GO:2000400) |
| 0.0 | 0.3 | GO:0032071 | regulation of endodeoxyribonuclease activity(GO:0032071) |
| 0.0 | 0.4 | GO:0071493 | cellular response to UV-B(GO:0071493) |
| 0.0 | 0.5 | GO:0071372 | cellular response to follicle-stimulating hormone stimulus(GO:0071372) |
| 0.0 | 0.2 | GO:0042738 | exogenous drug catabolic process(GO:0042738) |
| 0.0 | 0.4 | GO:0050882 | voluntary musculoskeletal movement(GO:0050882) |
| 0.0 | 0.8 | GO:0060218 | hematopoietic stem cell differentiation(GO:0060218) |
| 0.0 | 1.0 | GO:0034315 | regulation of Arp2/3 complex-mediated actin nucleation(GO:0034315) |
| 0.0 | 1.6 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.0 | 0.2 | GO:0070318 | positive regulation of G0 to G1 transition(GO:0070318) |
| 0.0 | 0.1 | GO:0010536 | positive regulation of activation of Janus kinase activity(GO:0010536) |
| 0.0 | 0.9 | GO:0048311 | mitochondrion distribution(GO:0048311) |
| 0.0 | 0.5 | GO:0045792 | negative regulation of cell size(GO:0045792) |
| 0.0 | 0.3 | GO:0015879 | carnitine transport(GO:0015879) |
| 0.0 | 0.1 | GO:1900247 | cytoplasmic translational elongation(GO:0002182) regulation of cytoplasmic translational elongation(GO:1900247) negative regulation of cytoplasmic translational elongation(GO:1900248) |
| 0.0 | 0.4 | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 0.0 | 0.1 | GO:2000795 | negative regulation of epithelial cell proliferation involved in lung morphogenesis(GO:2000795) |
| 0.0 | 0.6 | GO:0007512 | adult heart development(GO:0007512) |
| 0.0 | 0.2 | GO:0006116 | NADH oxidation(GO:0006116) |
| 0.0 | 0.6 | GO:0033617 | mitochondrial respiratory chain complex IV assembly(GO:0033617) mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.0 | 0.8 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.0 | 0.3 | GO:0097119 | postsynaptic density protein 95 clustering(GO:0097119) |
| 0.0 | 0.1 | GO:2000686 | regulation of rubidium ion transmembrane transporter activity(GO:2000686) |
| 0.0 | 0.4 | GO:1901898 | negative regulation of relaxation of muscle(GO:1901078) regulation of cell communication by electrical coupling involved in cardiac conduction(GO:1901844) negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.0 | 0.7 | GO:0007141 | male meiosis I(GO:0007141) |
| 0.0 | 0.7 | GO:0099514 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.0 | 0.5 | GO:0045672 | positive regulation of osteoclast differentiation(GO:0045672) |
| 0.0 | 0.1 | GO:0003245 | cardiac muscle tissue growth involved in heart morphogenesis(GO:0003245) |
| 0.0 | 0.3 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 0.0 | 0.3 | GO:0071044 | histone mRNA catabolic process(GO:0071044) |
| 0.0 | 1.7 | GO:0007588 | excretion(GO:0007588) |
| 0.0 | 0.6 | GO:0002026 | regulation of the force of heart contraction(GO:0002026) |
| 0.0 | 0.2 | GO:1902775 | mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.0 | 0.5 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.0 | 0.6 | GO:2000300 | regulation of synaptic vesicle exocytosis(GO:2000300) |
| 0.0 | 0.4 | GO:0060394 | negative regulation of pathway-restricted SMAD protein phosphorylation(GO:0060394) |
| 0.0 | 0.2 | GO:0014816 | skeletal muscle satellite cell differentiation(GO:0014816) |
| 0.0 | 0.1 | GO:0045829 | negative regulation of isotype switching(GO:0045829) negative regulation of isotype switching to IgE isotypes(GO:0048294) |
| 0.0 | 0.8 | GO:0015986 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.0 | 1.1 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.0 | 0.1 | GO:1902626 | assembly of large subunit precursor of preribosome(GO:1902626) |
| 0.0 | 0.5 | GO:0034453 | microtubule anchoring(GO:0034453) |
| 0.0 | 1.9 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.0 | 0.3 | GO:0003283 | atrial septum development(GO:0003283) |
| 0.0 | 0.7 | GO:0051894 | positive regulation of focal adhesion assembly(GO:0051894) |
| 0.0 | 0.4 | GO:0009083 | branched-chain amino acid catabolic process(GO:0009083) |
| 0.0 | 0.0 | GO:0090194 | negative regulation of glomerular mesangial cell proliferation(GO:0072125) negative regulation of glomerulus development(GO:0090194) |
| 0.0 | 0.1 | GO:0015793 | glycerol transport(GO:0015793) |
| 0.0 | 1.1 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.0 | 0.1 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.0 | 0.4 | GO:0045671 | negative regulation of osteoclast differentiation(GO:0045671) |
| 0.0 | 0.6 | GO:0051452 | intracellular pH reduction(GO:0051452) |
| 0.0 | 0.3 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.0 | 0.5 | GO:0001921 | positive regulation of receptor recycling(GO:0001921) |
| 0.0 | 0.1 | GO:0043951 | negative regulation of cAMP-mediated signaling(GO:0043951) |
| 0.0 | 0.1 | GO:0033684 | positive regulation of gonadotropin secretion(GO:0032278) regulation of luteinizing hormone secretion(GO:0033684) |
| 0.0 | 0.4 | GO:0031571 | mitotic G1 DNA damage checkpoint(GO:0031571) |
| 0.0 | 0.5 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.0 | 0.1 | GO:0070162 | cellular triglyceride homeostasis(GO:0035356) adiponectin secretion(GO:0070162) regulation of adiponectin secretion(GO:0070163) |
| 0.0 | 0.3 | GO:0051923 | sulfation(GO:0051923) |
| 0.0 | 0.2 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.0 | 1.1 | GO:0015844 | monoamine transport(GO:0015844) |
| 0.0 | 0.7 | GO:0061098 | positive regulation of protein tyrosine kinase activity(GO:0061098) |
| 0.0 | 0.3 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.0 | 0.3 | GO:0032464 | positive regulation of protein homooligomerization(GO:0032464) |
| 0.0 | 0.7 | GO:0045604 | regulation of epidermal cell differentiation(GO:0045604) |
| 0.0 | 0.5 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 0.0 | 0.2 | GO:0045742 | positive regulation of epidermal growth factor receptor signaling pathway(GO:0045742) |
| 0.0 | 0.1 | GO:0006297 | nucleotide-excision repair, DNA gap filling(GO:0006297) |
| 0.0 | 0.2 | GO:0000183 | chromatin silencing at rDNA(GO:0000183) |
| 0.0 | 0.9 | GO:0070527 | platelet aggregation(GO:0070527) |
| 0.0 | 2.9 | GO:0030308 | negative regulation of cell growth(GO:0030308) |
| 0.0 | 1.4 | GO:2001243 | negative regulation of intrinsic apoptotic signaling pathway(GO:2001243) |
| 0.0 | 0.2 | GO:0090336 | positive regulation of brown fat cell differentiation(GO:0090336) |
| 0.0 | 0.9 | GO:0030500 | regulation of bone mineralization(GO:0030500) |
| 0.0 | 0.5 | GO:0045599 | negative regulation of fat cell differentiation(GO:0045599) |
| 0.0 | 0.1 | GO:0016479 | negative regulation of transcription from RNA polymerase I promoter(GO:0016479) negative regulation of transcription of nuclear large rRNA transcript from RNA polymerase I promoter(GO:1901837) |
| 0.0 | 0.4 | GO:0071385 | cellular response to corticosteroid stimulus(GO:0071384) cellular response to glucocorticoid stimulus(GO:0071385) |
| 0.0 | 0.3 | GO:0021952 | central nervous system projection neuron axonogenesis(GO:0021952) |
| 0.0 | 0.0 | GO:0021938 | smoothened signaling pathway involved in regulation of cerebellar granule cell precursor cell proliferation(GO:0021938) notochord regression(GO:0060032) |
| 0.0 | 0.3 | GO:0046597 | negative regulation of viral entry into host cell(GO:0046597) |
| 0.0 | 0.5 | GO:0001662 | behavioral fear response(GO:0001662) behavioral defense response(GO:0002209) fear response(GO:0042596) |
| 0.0 | 0.5 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.0 | 0.5 | GO:1901998 | toxin transport(GO:1901998) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 14.4 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.8 | 5.5 | GO:0097550 | transcriptional preinitiation complex(GO:0097550) |
| 0.6 | 3.2 | GO:0042719 | mitochondrial intermembrane space protein transporter complex(GO:0042719) |
| 0.5 | 1.6 | GO:0097135 | cyclin E2-CDK2 complex(GO:0097135) |
| 0.4 | 5.9 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.4 | 2.5 | GO:0032983 | kainate selective glutamate receptor complex(GO:0032983) |
| 0.3 | 2.1 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.3 | 1.2 | GO:0042827 | platelet dense granule(GO:0042827) |
| 0.3 | 2.4 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.3 | 1.7 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 0.2 | 1.4 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 0.2 | 4.1 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.2 | 0.8 | GO:0090661 | box H/ACA telomerase RNP complex(GO:0090661) |
| 0.2 | 1.9 | GO:0000796 | condensin complex(GO:0000796) |
| 0.2 | 3.6 | GO:0043196 | varicosity(GO:0043196) |
| 0.2 | 1.4 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.1 | 5.8 | GO:0032809 | neuronal cell body membrane(GO:0032809) |
| 0.1 | 0.4 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 0.1 | 1.6 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.1 | 1.8 | GO:0045180 | basal cortex(GO:0045180) |
| 0.1 | 1.1 | GO:0002177 | manchette(GO:0002177) |
| 0.1 | 1.1 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.1 | 1.0 | GO:0042629 | mast cell granule(GO:0042629) |
| 0.1 | 1.6 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.1 | 0.7 | GO:0045261 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 0.1 | 0.6 | GO:0072379 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.1 | 1.2 | GO:0097539 | ciliary transition fiber(GO:0097539) |
| 0.1 | 0.8 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.1 | 0.7 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.1 | 2.6 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.1 | 0.4 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.1 | 0.2 | GO:0030905 | retromer, tubulation complex(GO:0030905) |
| 0.1 | 0.5 | GO:0031415 | NatA complex(GO:0031415) |
| 0.1 | 1.4 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.1 | 1.7 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.1 | 3.2 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 0.5 | GO:0031315 | extrinsic component of mitochondrial outer membrane(GO:0031315) |
| 0.1 | 0.6 | GO:0005675 | holo TFIIH complex(GO:0005675) |
| 0.0 | 2.1 | GO:0030660 | Golgi-associated vesicle membrane(GO:0030660) |
| 0.0 | 0.3 | GO:0017071 | intracellular cyclic nucleotide activated cation channel complex(GO:0017071) |
| 0.0 | 1.5 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.0 | 2.5 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.5 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.0 | 0.8 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.0 | 2.5 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.7 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.0 | 0.4 | GO:0005750 | mitochondrial respiratory chain complex III(GO:0005750) |
| 0.0 | 0.8 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.0 | 0.8 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 1.3 | GO:0016460 | myosin II complex(GO:0016460) |
| 0.0 | 0.6 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.0 | 0.8 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.0 | 0.4 | GO:0030128 | clathrin coat of endocytic vesicle(GO:0030128) |
| 0.0 | 0.9 | GO:0001741 | XY body(GO:0001741) |
| 0.0 | 0.2 | GO:0030915 | Smc5-Smc6 complex(GO:0030915) |
| 0.0 | 0.4 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.0 | 1.2 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.0 | 0.8 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 0.2 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.0 | 0.3 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.0 | 1.1 | GO:0005747 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.9 | GO:0005798 | Golgi-associated vesicle(GO:0005798) |
| 0.0 | 2.0 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.0 | 0.2 | GO:0097208 | alveolar lamellar body(GO:0097208) |
| 0.0 | 0.5 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) |
| 0.0 | 6.5 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 0.5 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.0 | 0.2 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.0 | 0.2 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.0 | 0.4 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 0.3 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.0 | 2.7 | GO:0030017 | sarcomere(GO:0030017) |
| 0.0 | 0.7 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 0.4 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.0 | 0.5 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.0 | 0.1 | GO:0000815 | ESCRT III complex(GO:0000815) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 5.5 | GO:0031798 | type 1 metabotropic glutamate receptor binding(GO:0031798) RNA polymerase II transcription factor activity, estrogen-activated sequence-specific DNA binding(GO:0038052) |
| 1.1 | 6.7 | GO:0051033 | nucleic acid transmembrane transporter activity(GO:0051032) RNA transmembrane transporter activity(GO:0051033) |
| 0.8 | 3.3 | GO:0042903 | tubulin deacetylase activity(GO:0042903) |
| 0.8 | 3.0 | GO:0036313 | phosphatidylinositol 3-kinase catalytic subunit binding(GO:0036313) |
| 0.7 | 2.2 | GO:0016015 | morphogen activity(GO:0016015) |
| 0.7 | 4.7 | GO:0004117 | calmodulin-dependent cyclic-nucleotide phosphodiesterase activity(GO:0004117) calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.5 | 2.2 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.5 | 1.4 | GO:0002153 | steroid receptor RNA activator RNA binding(GO:0002153) |
| 0.5 | 2.3 | GO:0052832 | inositol monophosphate 1-phosphatase activity(GO:0008934) inositol monophosphate 3-phosphatase activity(GO:0052832) inositol monophosphate 4-phosphatase activity(GO:0052833) inositol monophosphate phosphatase activity(GO:0052834) |
| 0.4 | 2.4 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.4 | 1.1 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 0.3 | 1.7 | GO:0030250 | cyclase activator activity(GO:0010853) guanylate cyclase activator activity(GO:0030250) |
| 0.3 | 2.6 | GO:0015181 | arginine transmembrane transporter activity(GO:0015181) |
| 0.3 | 0.9 | GO:0046592 | polyamine oxidase activity(GO:0046592) spermine:oxygen oxidoreductase (spermidine-forming) activity(GO:0052901) |
| 0.3 | 2.1 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.3 | 0.8 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 0.3 | 1.8 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.3 | 1.8 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.2 | 1.5 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.2 | 0.9 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.2 | 0.6 | GO:0004069 | L-aspartate:2-oxoglutarate aminotransferase activity(GO:0004069) L-phenylalanine aminotransferase activity(GO:0070546) L-phenylalanine:2-oxoglutarate aminotransferase activity(GO:0080130) |
| 0.2 | 0.6 | GO:0071820 | N-box binding(GO:0071820) |
| 0.2 | 1.2 | GO:0005030 | neurotrophin receptor activity(GO:0005030) |
| 0.2 | 0.9 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.2 | 0.6 | GO:0071077 | coenzyme A transmembrane transporter activity(GO:0015228) adenosine 3',5'-bisphosphate transmembrane transporter activity(GO:0071077) AMP transmembrane transporter activity(GO:0080122) |
| 0.2 | 1.5 | GO:0043208 | glycosphingolipid binding(GO:0043208) |
| 0.2 | 1.4 | GO:0031811 | G-protein coupled nucleotide receptor binding(GO:0031811) P2Y1 nucleotide receptor binding(GO:0031812) |
| 0.2 | 0.5 | GO:0070736 | protein-glycine ligase activity, initiating(GO:0070736) |
| 0.2 | 0.8 | GO:0008545 | JUN kinase kinase activity(GO:0008545) |
| 0.2 | 0.6 | GO:0086038 | calcium:sodium antiporter activity involved in regulation of cardiac muscle cell membrane potential(GO:0086038) |
| 0.2 | 0.6 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.2 | 0.9 | GO:0033743 | peptide-methionine (R)-S-oxide reductase activity(GO:0033743) |
| 0.2 | 1.4 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.1 | 2.3 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.1 | 0.4 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 0.1 | 0.6 | GO:0004096 | catalase activity(GO:0004096) |
| 0.1 | 1.6 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.1 | 1.1 | GO:0008504 | monoamine transmembrane transporter activity(GO:0008504) |
| 0.1 | 5.8 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.1 | 0.3 | GO:0004534 | 5'-3' exoribonuclease activity(GO:0004534) |
| 0.1 | 3.3 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.1 | 0.7 | GO:0070728 | leucine binding(GO:0070728) |
| 0.1 | 3.1 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.1 | 0.6 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.1 | 0.8 | GO:0034513 | box H/ACA snoRNA binding(GO:0034513) |
| 0.1 | 0.9 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.1 | 1.9 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.1 | 2.0 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.1 | 0.4 | GO:0005146 | leukemia inhibitory factor receptor binding(GO:0005146) |
| 0.1 | 0.6 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
| 0.1 | 0.5 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 0.1 | 2.0 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.1 | 0.4 | GO:0005220 | inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.1 | 1.7 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.1 | 0.8 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.1 | 1.9 | GO:0005123 | death receptor binding(GO:0005123) |
| 0.1 | 0.5 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) |
| 0.1 | 1.3 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.1 | 1.4 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 0.3 | GO:0005223 | intracellular cGMP activated cation channel activity(GO:0005223) |
| 0.1 | 0.6 | GO:0031404 | chloride ion binding(GO:0031404) |
| 0.1 | 0.8 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
| 0.1 | 1.8 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.1 | 0.8 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.1 | 0.2 | GO:1990460 | leptin receptor binding(GO:1990460) |
| 0.1 | 1.6 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.1 | 0.3 | GO:1990188 | euchromatin binding(GO:1990188) |
| 0.0 | 0.8 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.3 | GO:0050694 | galactose 3-O-sulfotransferase activity(GO:0050694) |
| 0.0 | 0.9 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.3 | GO:0005124 | scavenger receptor binding(GO:0005124) |
| 0.0 | 0.2 | GO:0036033 | mediator complex binding(GO:0036033) |
| 0.0 | 1.9 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.7 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 3.4 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 0.1 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.0 | 0.4 | GO:0016634 | oxidoreductase activity, acting on the CH-CH group of donors, oxygen as acceptor(GO:0016634) |
| 0.0 | 1.1 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.0 | 1.0 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.1 | GO:0005128 | erythropoietin receptor binding(GO:0005128) |
| 0.0 | 0.3 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.0 | 1.7 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.8 | GO:0016799 | hydrolase activity, hydrolyzing N-glycosyl compounds(GO:0016799) |
| 0.0 | 0.7 | GO:0008483 | transaminase activity(GO:0008483) |
| 0.0 | 0.5 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.0 | 1.1 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 0.4 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.0 | 4.4 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 0.5 | GO:0031702 | type 1 angiotensin receptor binding(GO:0031702) |
| 0.0 | 0.9 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.0 | 1.2 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.5 | GO:0004862 | cAMP-dependent protein kinase inhibitor activity(GO:0004862) |
| 0.0 | 1.8 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 0.5 | GO:0051393 | alpha-actinin binding(GO:0051393) |
| 0.0 | 1.5 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.0 | 0.8 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 0.4 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.0 | 0.1 | GO:0019981 | interleukin-6 receptor activity(GO:0004915) interleukin-6 binding(GO:0019981) |
| 0.0 | 14.0 | GO:0003779 | actin binding(GO:0003779) |
| 0.0 | 0.7 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 1.9 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.0 | 1.0 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.0 | 0.8 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.0 | 0.0 | GO:0005143 | interleukin-12 receptor binding(GO:0005143) |
| 0.0 | 1.0 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 0.2 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.0 | 0.7 | GO:0030742 | GTP-dependent protein binding(GO:0030742) |
| 0.0 | 1.8 | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups(GO:0016765) |
| 0.0 | 0.1 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.0 | 0.4 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 0.4 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.0 | 0.1 | GO:0015254 | glycerol transmembrane transporter activity(GO:0015168) glycerol channel activity(GO:0015254) |
| 0.0 | 0.5 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.0 | 0.4 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 0.4 | GO:0008308 | voltage-gated anion channel activity(GO:0008308) |
| 0.0 | 0.4 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.0 | 0.1 | GO:0004331 | 6-phosphofructo-2-kinase activity(GO:0003873) fructose-2,6-bisphosphate 2-phosphatase activity(GO:0004331) |
| 0.0 | 1.1 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 2.7 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.3 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.3 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.0 | 0.6 | GO:0070063 | RNA polymerase binding(GO:0070063) |
| 0.0 | 0.1 | GO:0004315 | 3-oxoacyl-[acyl-carrier-protein] synthase activity(GO:0004315) |
| 0.0 | 0.9 | GO:0019905 | syntaxin binding(GO:0019905) |
| 0.0 | 0.9 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 1.3 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 1.1 | GO:0101005 | thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.0 | 3.3 | GO:0001047 | core promoter binding(GO:0001047) |
| 0.0 | 0.7 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 0.2 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.0 | 1.1 | GO:0003823 | antigen binding(GO:0003823) |
| 0.0 | 0.6 | GO:0043621 | protein self-association(GO:0043621) |
| 0.0 | 1.1 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.0 | 0.1 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.0 | 0.5 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.2 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.0 | 0.3 | GO:0017091 | AU-rich element binding(GO:0017091) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 5.8 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.1 | 7.7 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 8.2 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 2.2 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.1 | 1.6 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.1 | 2.1 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.1 | 2.7 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.1 | 2.5 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 2.7 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.1 | 3.4 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.1 | 0.4 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.1 | 3.7 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 0.8 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.0 | 1.9 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 0.6 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 0.8 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 2.1 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 1.4 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 0.9 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.0 | 1.4 | PID ATR PATHWAY | ATR signaling pathway |
| 0.0 | 1.0 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 0.6 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 1.0 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.5 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 1.2 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 0.6 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 1.3 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.0 | 0.5 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 0.7 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.0 | 0.8 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 1.1 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 0.5 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.0 | 0.4 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.0 | 0.7 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 0.8 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 0.2 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 0.6 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 0.2 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 8.6 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.2 | 3.1 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.2 | 7.7 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.1 | 4.7 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.1 | 0.8 | REACTOME NITRIC OXIDE STIMULATES GUANYLATE CYCLASE | Genes involved in Nitric oxide stimulates guanylate cyclase |
| 0.1 | 0.5 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GLP1 | Genes involved in Synthesis, Secretion, and Inactivation of Glucagon-like Peptide-1 (GLP-1) |
| 0.1 | 1.7 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.1 | 2.5 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.1 | 1.5 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.1 | 2.4 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 1.6 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.1 | 0.7 | REACTOME SIGNALING BY EGFR IN CANCER | Genes involved in Signaling by EGFR in Cancer |
| 0.1 | 0.8 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.1 | 3.0 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 0.4 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.1 | 0.8 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.1 | 0.8 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.1 | 1.4 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.6 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 3.3 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 2.4 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.0 | 1.1 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.0 | 0.8 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.0 | 1.0 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.6 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 0.0 | 0.4 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.0 | 0.8 | REACTOME JNK C JUN KINASES PHOSPHORYLATION AND ACTIVATION MEDIATED BY ACTIVATED HUMAN TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 0.0 | 1.2 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.8 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.0 | 1.0 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.0 | 0.5 | REACTOME DESTABILIZATION OF MRNA BY BRF1 | Genes involved in Destabilization of mRNA by Butyrate Response Factor 1 (BRF1) |
| 0.0 | 0.8 | REACTOME EXTENSION OF TELOMERES | Genes involved in Extension of Telomeres |
| 0.0 | 0.8 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 1.5 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.0 | 0.8 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.0 | 0.2 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.0 | 2.4 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.4 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.0 | 0.1 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 0.6 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.4 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.0 | 0.9 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 0.4 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 0.9 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.9 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.0 | 0.8 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 0.4 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 1.4 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 0.3 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.0 | 1.5 | REACTOME P75 NTR RECEPTOR MEDIATED SIGNALLING | Genes involved in p75 NTR receptor-mediated signalling |
| 0.0 | 0.5 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 0.8 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 0.5 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |
| 0.0 | 0.9 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.0 | 1.0 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 0.3 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 0.8 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |