Project
avrg: 12D miR HR13_24
Navigation
Downloads
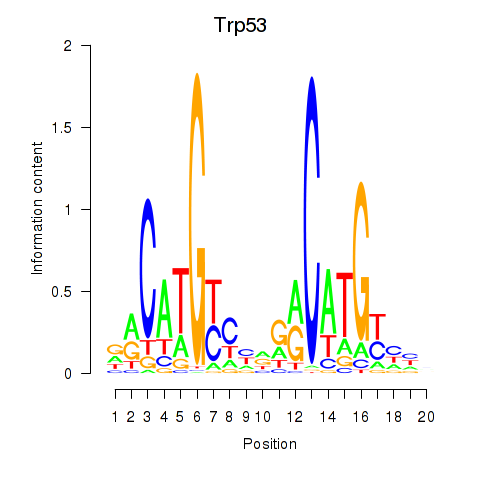
Results for Trp53
Z-value: 2.21
Transcription factors associated with Trp53
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Trp53
|
ENSMUSG00000059552.7 | transformation related protein 53 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Trp53 | mm10_v2_chr11_+_69580359_69580382 | 0.82 | 1.9e-03 | Click! |
Activity profile of Trp53 motif
Sorted Z-values of Trp53 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_-_40755201 | 3.96 |
ENSMUST00000020576.7
|
Ccng1
|
cyclin G1 |
| chr17_+_29090969 | 3.69 |
ENSMUST00000119901.1
|
Cdkn1a
|
cyclin-dependent kinase inhibitor 1A (P21) |
| chr5_-_38480131 | 1.76 |
ENSMUST00000143758.1
ENSMUST00000067886.5 |
Slc2a9
|
solute carrier family 2 (facilitated glucose transporter), member 9 |
| chr1_-_79858627 | 1.60 |
ENSMUST00000027467.4
|
Serpine2
|
serine (or cysteine) peptidase inhibitor, clade E, member 2 |
| chr15_+_62039216 | 1.58 |
ENSMUST00000183297.1
|
Pvt1
|
plasmacytoma variant translocation 1 |
| chr15_+_85859689 | 1.40 |
ENSMUST00000170629.1
|
Gtse1
|
G two S phase expressed protein 1 |
| chr8_-_22185758 | 1.35 |
ENSMUST00000046916.7
|
Ckap2
|
cytoskeleton associated protein 2 |
| chr2_+_152962485 | 1.33 |
ENSMUST00000099197.2
ENSMUST00000103155.3 |
Ttll9
|
tubulin tyrosine ligase-like family, member 9 |
| chr15_-_102350692 | 1.32 |
ENSMUST00000041208.7
|
Aaas
|
achalasia, adrenocortical insufficiency, alacrimia |
| chr7_+_16309577 | 1.30 |
ENSMUST00000002152.6
|
Bbc3
|
BCL2 binding component 3 |
| chr14_-_118925314 | 1.22 |
ENSMUST00000004055.8
|
Dzip1
|
DAZ interacting protein 1 |
| chr17_-_33890584 | 1.20 |
ENSMUST00000114361.2
ENSMUST00000173492.1 |
Kifc1
|
kinesin family member C1 |
| chr7_-_24545994 | 1.10 |
ENSMUST00000011776.6
|
Pinlyp
|
phospholipase A2 inhibitor and LY6/PLAUR domain containing |
| chr5_-_38502107 | 1.06 |
ENSMUST00000005238.6
|
Slc2a9
|
solute carrier family 2 (facilitated glucose transporter), member 9 |
| chr2_+_72285637 | 1.00 |
ENSMUST00000090824.5
ENSMUST00000135469.1 |
Zak
|
sterile alpha motif and leucine zipper containing kinase AZK |
| chr16_+_17144600 | 0.93 |
ENSMUST00000115702.1
|
Ydjc
|
YdjC homolog (bacterial) |
| chr17_-_33890539 | 0.92 |
ENSMUST00000173386.1
|
Kifc1
|
kinesin family member C1 |
| chr7_+_19851994 | 0.91 |
ENSMUST00000172815.1
|
Gm19345
|
predicted gene, 19345 |
| chr16_-_17144415 | 0.89 |
ENSMUST00000115709.1
|
Ccdc116
|
coiled-coil domain containing 116 |
| chr11_+_4267095 | 0.87 |
ENSMUST00000040750.3
|
Lif
|
leukemia inhibitory factor |
| chr9_+_66946057 | 0.71 |
ENSMUST00000040917.7
ENSMUST00000127896.1 |
Rps27l
|
ribosomal protein S27-like |
| chr11_-_95699143 | 0.70 |
ENSMUST00000062249.2
|
Gm9796
|
predicted gene 9796 |
| chr19_-_10974664 | 0.70 |
ENSMUST00000072748.6
|
Ms4a10
|
membrane-spanning 4-domains, subfamily A, member 10 |
| chr6_-_116461024 | 0.63 |
ENSMUST00000164547.1
ENSMUST00000170186.1 |
Alox5
|
arachidonate 5-lipoxygenase |
| chr2_+_129100995 | 0.63 |
ENSMUST00000103205.4
ENSMUST00000028874.7 |
Polr1b
|
polymerase (RNA) I polypeptide B |
| chr14_+_13454010 | 0.62 |
ENSMUST00000112656.2
|
Synpr
|
synaptoporin |
| chr4_+_139622842 | 0.62 |
ENSMUST00000039818.9
|
Aldh4a1
|
aldehyde dehydrogenase 4 family, member A1 |
| chr16_+_38458887 | 0.61 |
ENSMUST00000099816.2
|
Cd80
|
CD80 antigen |
| chr8_-_106136890 | 0.61 |
ENSMUST00000115979.2
|
Esrp2
|
epithelial splicing regulatory protein 2 |
| chr9_-_100486788 | 0.60 |
ENSMUST00000098458.3
|
Il20rb
|
interleukin 20 receptor beta |
| chr10_+_57631981 | 0.59 |
ENSMUST00000095668.3
ENSMUST00000075992.5 |
Pkib
|
protein kinase inhibitor beta, cAMP dependent, testis specific |
| chr3_+_79885930 | 0.58 |
ENSMUST00000029567.8
|
Fam198b
|
family with sequence similarity 198, member B |
| chr1_-_173982842 | 0.56 |
ENSMUST00000000266.7
|
Ifi202b
|
interferon activated gene 202B |
| chr7_+_47050628 | 0.56 |
ENSMUST00000010451.5
|
Tmem86a
|
transmembrane protein 86A |
| chr5_+_8798139 | 0.54 |
ENSMUST00000009058.5
|
Abcb1b
|
ATP-binding cassette, sub-family B (MDR/TAP), member 1B |
| chr13_-_12520377 | 0.53 |
ENSMUST00000179308.1
|
Edaradd
|
EDAR (ectodysplasin-A receptor)-associated death domain |
| chr5_+_92392585 | 0.52 |
ENSMUST00000126281.1
|
Art3
|
ADP-ribosyltransferase 3 |
| chr14_+_47001336 | 0.51 |
ENSMUST00000125113.1
|
Samd4
|
sterile alpha motif domain containing 4 |
| chr10_+_57632108 | 0.49 |
ENSMUST00000177438.1
|
Pkib
|
protein kinase inhibitor beta, cAMP dependent, testis specific |
| chr1_+_55237177 | 0.49 |
ENSMUST00000061334.8
|
Mars2
|
methionine-tRNA synthetase 2 (mitochondrial) |
| chr14_-_55944536 | 0.48 |
ENSMUST00000022834.6
|
Cma1
|
chymase 1, mast cell |
| chr1_+_133350510 | 0.46 |
ENSMUST00000094556.2
|
Ren1
|
renin 1 structural |
| chr13_-_22035589 | 0.46 |
ENSMUST00000091742.4
|
Hist1h2ah
|
histone cluster 1, H2ah |
| chr11_+_87582201 | 0.46 |
ENSMUST00000133202.1
|
Sept4
|
septin 4 |
| chr7_-_19458494 | 0.45 |
ENSMUST00000085715.5
|
Mark4
|
MAP/microtubule affinity-regulating kinase 4 |
| chr4_-_134254076 | 0.43 |
ENSMUST00000060050.5
|
Grrp1
|
glycine/arginine rich protein 1 |
| chr17_+_12584183 | 0.43 |
ENSMUST00000046959.7
|
Slc22a2
|
solute carrier family 22 (organic cation transporter), member 2 |
| chr17_+_3532554 | 0.42 |
ENSMUST00000168560.1
|
Cldn20
|
claudin 20 |
| chr11_-_102925086 | 0.41 |
ENSMUST00000021311.9
|
Kif18b
|
kinesin family member 18B |
| chr11_+_118476824 | 0.41 |
ENSMUST00000135383.2
|
Engase
|
endo-beta-N-acetylglucosaminidase |
| chr6_-_116461151 | 0.41 |
ENSMUST00000026795.6
|
Alox5
|
arachidonate 5-lipoxygenase |
| chr16_-_94856682 | 0.40 |
ENSMUST00000165538.1
|
Kcnj6
|
potassium inwardly-rectifying channel, subfamily J, member 6 |
| chr4_+_141213948 | 0.38 |
ENSMUST00000097813.2
|
Rsg1
|
REM2 and RAB-like small GTPase 1 |
| chr3_-_92573715 | 0.38 |
ENSMUST00000053107.4
|
Ivl
|
involucrin |
| chr5_-_116422858 | 0.38 |
ENSMUST00000036991.4
|
Hspb8
|
heat shock protein 8 |
| chr1_-_191907527 | 0.38 |
ENSMUST00000069573.5
|
1700034H15Rik
|
RIKEN cDNA 1700034H15 gene |
| chr9_-_106656081 | 0.38 |
ENSMUST00000023959.7
|
Grm2
|
glutamate receptor, metabotropic 2 |
| chr9_-_106158109 | 0.38 |
ENSMUST00000159809.1
ENSMUST00000162562.1 ENSMUST00000036382.6 ENSMUST00000112543.2 |
Glyctk
|
glycerate kinase |
| chr4_-_96591555 | 0.38 |
ENSMUST00000055693.8
|
Cyp2j9
|
cytochrome P450, family 2, subfamily j, polypeptide 9 |
| chr15_-_98898483 | 0.37 |
ENSMUST00000023737.4
|
Dhh
|
desert hedgehog |
| chr1_-_184883218 | 0.35 |
ENSMUST00000048308.5
|
C130074G19Rik
|
RIKEN cDNA C130074G19 gene |
| chr13_+_21754067 | 0.32 |
ENSMUST00000091709.2
|
Hist1h2bn
|
histone cluster 1, H2bn |
| chr16_-_11909398 | 0.32 |
ENSMUST00000127972.1
ENSMUST00000121750.1 ENSMUST00000096272.4 ENSMUST00000073371.6 |
Cpped1
|
calcineurin-like phosphoesterase domain containing 1 |
| chr14_-_59365410 | 0.31 |
ENSMUST00000161031.1
ENSMUST00000160425.1 |
Phf11d
|
PHD finger protein 11D |
| chr5_-_66080971 | 0.31 |
ENSMUST00000127275.1
ENSMUST00000113724.1 |
Rbm47
|
RNA binding motif protein 47 |
| chr17_+_26917091 | 0.31 |
ENSMUST00000078961.4
|
Kifc5b
|
kinesin family member C5B |
| chr14_+_13453937 | 0.31 |
ENSMUST00000153954.1
|
Synpr
|
synaptoporin |
| chr3_+_64081642 | 0.31 |
ENSMUST00000029406.4
|
Vmn2r1
|
vomeronasal 2, receptor 1 |
| chrX_-_73435332 | 0.30 |
ENSMUST00000033738.7
|
Trex2
|
three prime repair exonuclease 2 |
| chr17_+_28207778 | 0.30 |
ENSMUST00000002327.5
|
Def6
|
differentially expressed in FDCP 6 |
| chr14_-_59365465 | 0.30 |
ENSMUST00000095157.4
|
Phf11d
|
PHD finger protein 11D |
| chr16_+_57353093 | 0.30 |
ENSMUST00000159816.1
|
Filip1l
|
filamin A interacting protein 1-like |
| chr12_-_78906929 | 0.28 |
ENSMUST00000021544.7
|
Plek2
|
pleckstrin 2 |
| chr13_+_23574381 | 0.27 |
ENSMUST00000090776.4
|
Hist1h2ad
|
histone cluster 1, H2ad |
| chr13_+_22035821 | 0.27 |
ENSMUST00000110455.2
|
Hist1h2bk
|
histone cluster 1, H2bk |
| chr4_-_155056784 | 0.26 |
ENSMUST00000131173.2
|
Plch2
|
phospholipase C, eta 2 |
| chr19_+_4855129 | 0.25 |
ENSMUST00000119694.1
|
Ctsf
|
cathepsin F |
| chr17_+_27685197 | 0.25 |
ENSMUST00000097360.2
|
Pacsin1
|
protein kinase C and casein kinase substrate in neurons 1 |
| chr2_-_163397946 | 0.25 |
ENSMUST00000017961.4
ENSMUST00000109425.2 |
Jph2
|
junctophilin 2 |
| chr7_+_24884611 | 0.25 |
ENSMUST00000108428.1
|
Rps19
|
ribosomal protein S19 |
| chr5_+_52363925 | 0.25 |
ENSMUST00000101208.4
|
Sod3
|
superoxide dismutase 3, extracellular |
| chr2_+_29889217 | 0.24 |
ENSMUST00000123335.1
|
Odf2
|
outer dense fiber of sperm tails 2 |
| chr7_+_24884809 | 0.24 |
ENSMUST00000156372.1
ENSMUST00000124035.1 |
Rps19
|
ribosomal protein S19 |
| chr5_-_115194283 | 0.24 |
ENSMUST00000112113.1
|
Cabp1
|
calcium binding protein 1 |
| chr7_+_24884651 | 0.23 |
ENSMUST00000153451.2
ENSMUST00000108429.1 |
Rps19
|
ribosomal protein S19 |
| chr19_-_43524462 | 0.23 |
ENSMUST00000026196.7
|
Got1
|
glutamate oxaloacetate transaminase 1, soluble |
| chr17_-_45595842 | 0.23 |
ENSMUST00000164618.1
ENSMUST00000097317.3 ENSMUST00000170113.1 |
Slc29a1
|
solute carrier family 29 (nucleoside transporters), member 1 |
| chr19_-_47138280 | 0.23 |
ENSMUST00000140512.1
ENSMUST00000035822.1 |
Calhm2
|
calcium homeostasis modulator 2 |
| chr11_-_83429455 | 0.23 |
ENSMUST00000052521.2
|
Gas2l2
|
growth arrest-specific 2 like 2 |
| chr12_-_26415256 | 0.22 |
ENSMUST00000020971.6
ENSMUST00000062149.4 |
Rnf144a
|
ring finger protein 144A |
| chr1_-_22315792 | 0.22 |
ENSMUST00000164877.1
|
Rims1
|
regulating synaptic membrane exocytosis 1 |
| chr4_+_116596672 | 0.21 |
ENSMUST00000051869.7
|
Ccdc17
|
coiled-coil domain containing 17 |
| chr11_+_102835849 | 0.21 |
ENSMUST00000107073.1
|
Higd1b
|
HIG1 domain family, member 1B |
| chr13_-_98815408 | 0.21 |
ENSMUST00000040340.8
ENSMUST00000099277.4 ENSMUST00000179563.1 ENSMUST00000109403.1 |
Fcho2
|
FCH domain only 2 |
| chr9_-_59486323 | 0.21 |
ENSMUST00000165322.1
|
Arih1
|
ariadne ubiquitin-conjugating enzyme E2 binding protein homolog 1 (Drosophila) |
| chr1_+_51289106 | 0.21 |
ENSMUST00000051572.6
|
Sdpr
|
serum deprivation response |
| chr10_+_79822617 | 0.20 |
ENSMUST00000046833.4
|
Misp
|
mitotic spindle positioning |
| chr9_+_38773088 | 0.20 |
ENSMUST00000062124.3
|
Olfr921
|
olfactory receptor 921 |
| chr9_+_77636494 | 0.19 |
ENSMUST00000057781.7
|
Klhl31
|
kelch-like 31 |
| chr17_-_35066170 | 0.19 |
ENSMUST00000174190.1
ENSMUST00000097337.1 |
AU023871
|
expressed sequence AU023871 |
| chr14_-_20496780 | 0.19 |
ENSMUST00000022353.3
|
Mss51
|
MSS51 mitochondrial translational activator |
| chr5_-_38502079 | 0.18 |
ENSMUST00000147664.1
|
Slc2a9
|
solute carrier family 2 (facilitated glucose transporter), member 9 |
| chr2_-_155592567 | 0.18 |
ENSMUST00000155347.1
ENSMUST00000130881.1 ENSMUST00000079691.6 |
Gss
|
glutathione synthetase |
| chr7_-_24760311 | 0.18 |
ENSMUST00000063956.5
|
Cd177
|
CD177 antigen |
| chr4_-_9643638 | 0.17 |
ENSMUST00000108333.1
ENSMUST00000108334.1 ENSMUST00000108335.1 ENSMUST00000152526.1 ENSMUST00000103004.3 |
Asph
|
aspartate-beta-hydroxylase |
| chr4_-_156059414 | 0.17 |
ENSMUST00000184348.1
|
Ttll10
|
tubulin tyrosine ligase-like family, member 10 |
| chr3_-_132950043 | 0.16 |
ENSMUST00000117164.1
ENSMUST00000093971.4 ENSMUST00000042729.9 ENSMUST00000042744.9 ENSMUST00000117811.1 |
Npnt
|
nephronectin |
| chr7_+_130936172 | 0.16 |
ENSMUST00000006367.7
|
Htra1
|
HtrA serine peptidase 1 |
| chr9_+_65361049 | 0.16 |
ENSMUST00000147185.1
|
Gm514
|
predicted gene 514 |
| chr9_+_49518336 | 0.16 |
ENSMUST00000068730.3
|
Gm11149
|
predicted gene 11149 |
| chr2_+_86007778 | 0.16 |
ENSMUST00000062166.1
|
Olfr1032
|
olfactory receptor 1032 |
| chrX_+_136822781 | 0.16 |
ENSMUST00000113085.1
|
Plp1
|
proteolipid protein (myelin) 1 |
| chr10_+_110745433 | 0.16 |
ENSMUST00000174857.1
ENSMUST00000073781.5 ENSMUST00000173471.1 ENSMUST00000173634.1 |
E2f7
|
E2F transcription factor 7 |
| chr4_-_149099802 | 0.16 |
ENSMUST00000103217.4
|
Pex14
|
peroxisomal biogenesis factor 14 |
| chr1_+_6487231 | 0.15 |
ENSMUST00000140079.1
ENSMUST00000131494.1 |
St18
|
suppression of tumorigenicity 18 |
| chr17_+_78882003 | 0.15 |
ENSMUST00000180880.1
|
Gm26637
|
predicted gene, 26637 |
| chr14_-_103843685 | 0.15 |
ENSMUST00000172237.1
|
Ednrb
|
endothelin receptor type B |
| chr7_+_18718075 | 0.14 |
ENSMUST00000108481.1
ENSMUST00000051973.8 |
Psg22
|
pregnancy-specific glycoprotein 22 |
| chr2_+_136891501 | 0.14 |
ENSMUST00000141463.1
|
Slx4ip
|
SLX4 interacting protein |
| chr16_+_91391721 | 0.14 |
ENSMUST00000160764.1
|
Gm21970
|
predicted gene 21970 |
| chr8_-_88636117 | 0.14 |
ENSMUST00000034087.7
|
Snx20
|
sorting nexin 20 |
| chr2_+_10372426 | 0.14 |
ENSMUST00000114864.2
ENSMUST00000116594.2 ENSMUST00000041105.6 |
Sfmbt2
|
Scm-like with four mbt domains 2 |
| chr17_+_24840108 | 0.13 |
ENSMUST00000164251.1
|
Hagh
|
hydroxyacyl glutathione hydrolase |
| chr11_-_116168138 | 0.13 |
ENSMUST00000139020.1
ENSMUST00000103031.1 ENSMUST00000124828.1 |
Fbf1
|
Fas (TNFRSF6) binding factor 1 |
| chr4_+_43983496 | 0.13 |
ENSMUST00000095107.1
|
Ccin
|
calicin |
| chr1_-_52232296 | 0.13 |
ENSMUST00000114512.1
|
Gls
|
glutaminase |
| chr1_-_193130201 | 0.12 |
ENSMUST00000085555.1
|
Diexf
|
digestive organ expansion factor homolog (zebrafish) |
| chr14_-_78725089 | 0.12 |
ENSMUST00000074729.5
|
Dgkh
|
diacylglycerol kinase, eta |
| chrX_+_136822671 | 0.12 |
ENSMUST00000033800.6
|
Plp1
|
proteolipid protein (myelin) 1 |
| chr14_-_70653081 | 0.12 |
ENSMUST00000062629.4
|
Npm2
|
nucleophosmin/nucleoplasmin 2 |
| chr8_-_93079965 | 0.12 |
ENSMUST00000109582.1
|
Ces1b
|
carboxylesterase 1B |
| chr11_-_32267547 | 0.12 |
ENSMUST00000109389.2
ENSMUST00000129010.1 ENSMUST00000020530.5 |
Nprl3
|
nitrogen permease regulator-like 3 |
| chr8_-_71676674 | 0.11 |
ENSMUST00000066837.4
|
Gm9933
|
predicted gene 9933 |
| chr4_+_102430047 | 0.11 |
ENSMUST00000172616.1
|
Pde4b
|
phosphodiesterase 4B, cAMP specific |
| chr7_-_98309471 | 0.10 |
ENSMUST00000033020.7
|
Acer3
|
alkaline ceramidase 3 |
| chr7_-_141010759 | 0.10 |
ENSMUST00000026565.6
|
Ifitm3
|
interferon induced transmembrane protein 3 |
| chr4_-_135573623 | 0.10 |
ENSMUST00000105855.1
|
Grhl3
|
grainyhead-like 3 (Drosophila) |
| chr11_+_49280150 | 0.10 |
ENSMUST00000078932.1
|
Olfr1393
|
olfactory receptor 1393 |
| chr8_-_105851981 | 0.10 |
ENSMUST00000040776.4
|
Cenpt
|
centromere protein T |
| chr9_-_26999491 | 0.10 |
ENSMUST00000060513.7
ENSMUST00000120367.1 |
Acad8
|
acyl-Coenzyme A dehydrogenase family, member 8 |
| chr15_+_83791939 | 0.10 |
ENSMUST00000172115.1
ENSMUST00000172398.1 |
Mpped1
|
metallophosphoesterase domain containing 1 |
| chr7_+_142498832 | 0.10 |
ENSMUST00000078497.8
ENSMUST00000105953.3 ENSMUST00000179658.1 ENSMUST00000105954.3 ENSMUST00000105952.3 ENSMUST00000105955.1 ENSMUST00000074187.6 ENSMUST00000180152.1 ENSMUST00000105950.4 ENSMUST00000105957.3 ENSMUST00000169299.2 ENSMUST00000105958.3 ENSMUST00000105949.1 |
Tnnt3
|
troponin T3, skeletal, fast |
| chr6_-_38837224 | 0.09 |
ENSMUST00000160962.1
|
Hipk2
|
homeodomain interacting protein kinase 2 |
| chr3_-_146495115 | 0.09 |
ENSMUST00000093951.2
|
Spata1
|
spermatogenesis associated 1 |
| chr8_-_9977650 | 0.09 |
ENSMUST00000170033.1
|
Lig4
|
ligase IV, DNA, ATP-dependent |
| chrX_-_101734125 | 0.09 |
ENSMUST00000056614.6
|
Cxcr3
|
chemokine (C-X-C motif) receptor 3 |
| chr9_+_78395777 | 0.09 |
ENSMUST00000113367.1
|
Ddx43
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 43 |
| chr9_+_122117258 | 0.09 |
ENSMUST00000146832.1
ENSMUST00000139181.1 |
Snrk
|
SNF related kinase |
| chr7_+_64392645 | 0.08 |
ENSMUST00000037205.8
|
Mcee
|
methylmalonyl CoA epimerase |
| chr11_+_66911981 | 0.08 |
ENSMUST00000123434.2
|
Pirt
|
phosphoinositide-interacting regulator of transient receptor potential channels |
| chr16_-_45408875 | 0.08 |
ENSMUST00000023341.8
|
Cd200
|
CD200 antigen |
| chr13_-_23574196 | 0.08 |
ENSMUST00000105106.1
|
Hist1h2bf
|
histone cluster 1, H2bf |
| chr12_-_55492587 | 0.07 |
ENSMUST00000021413.7
|
Nfkbia
|
nuclear factor of kappa light polypeptide gene enhancer in B cells inhibitor, alpha |
| chr15_-_102246439 | 0.07 |
ENSMUST00000063339.7
|
Rarg
|
retinoic acid receptor, gamma |
| chr4_-_14796052 | 0.07 |
ENSMUST00000108276.1
ENSMUST00000023917.1 |
Lrrc69
|
leucine rich repeat containing 69 |
| chr8_+_9977707 | 0.06 |
ENSMUST00000139793.1
ENSMUST00000048216.5 |
Abhd13
|
abhydrolase domain containing 13 |
| chr16_-_20316750 | 0.06 |
ENSMUST00000182741.1
|
Cyp2ab1
|
cytochrome P450, family 2, subfamily ab, polypeptide 1 |
| chr7_-_92637079 | 0.06 |
ENSMUST00000056106.7
ENSMUST00000118157.1 |
Ankrd42
|
ankyrin repeat domain 42 |
| chr13_+_24376070 | 0.06 |
ENSMUST00000050859.5
|
Cmah
|
cytidine monophospho-N-acetylneuraminic acid hydroxylase |
| chr4_+_108459389 | 0.06 |
ENSMUST00000106673.1
ENSMUST00000043368.5 |
Zcchc11
|
zinc finger, CCHC domain containing 11 |
| chr13_+_59712284 | 0.06 |
ENSMUST00000165133.1
|
Spata31d1b
|
spermatogenesis associated 31 subfamily D, member 1B |
| chr16_-_45408955 | 0.06 |
ENSMUST00000163230.1
|
Cd200
|
CD200 antigen |
| chr17_-_24696147 | 0.06 |
ENSMUST00000046839.8
|
Gfer
|
growth factor, erv1 (S. cerevisiae)-like (augmenter of liver regeneration) |
| chr9_+_65346066 | 0.05 |
ENSMUST00000048184.2
|
Pdcd7
|
programmed cell death 7 |
| chr5_-_138272786 | 0.05 |
ENSMUST00000161279.1
ENSMUST00000161647.1 |
Gal3st4
|
galactose-3-O-sulfotransferase 4 |
| chr9_-_14718381 | 0.05 |
ENSMUST00000115643.1
|
Piwil4
|
piwi-like RNA-mediated gene silencing 4 |
| chr10_+_94550852 | 0.05 |
ENSMUST00000148910.1
ENSMUST00000117460.1 |
Tmcc3
|
transmembrane and coiled coil domains 3 |
| chr2_-_17731035 | 0.05 |
ENSMUST00000028080.5
|
Nebl
|
nebulette |
| chr7_-_64872993 | 0.04 |
ENSMUST00000094331.2
|
Ndnl2
|
necdin-like 2 |
| chr6_+_127446819 | 0.04 |
ENSMUST00000112191.1
|
Parp11
|
poly (ADP-ribose) polymerase family, member 11 |
| chr6_+_71707561 | 0.04 |
ENSMUST00000121469.1
|
Reep1
|
receptor accessory protein 1 |
| chr11_+_55469677 | 0.04 |
ENSMUST00000018727.3
|
G3bp1
|
GTPase activating protein (SH3 domain) binding protein 1 |
| chr7_+_30169861 | 0.04 |
ENSMUST00000085668.4
|
Gm5113
|
predicted gene 5113 |
| chr6_-_128891105 | 0.04 |
ENSMUST00000178918.1
ENSMUST00000160290.1 |
BC035044
|
cDNA sequence BC035044 |
| chr4_+_148558422 | 0.04 |
ENSMUST00000017408.7
ENSMUST00000076022.6 |
Exosc10
|
exosome component 10 |
| chr5_+_34999111 | 0.03 |
ENSMUST00000114283.1
|
Rgs12
|
regulator of G-protein signaling 12 |
| chr4_-_43030440 | 0.03 |
ENSMUST00000135660.1
|
Stoml2
|
stomatin (Epb7.2)-like 2 |
| chr1_+_161969179 | 0.03 |
ENSMUST00000111594.2
ENSMUST00000028021.6 |
Pigc
|
phosphatidylinositol glycan anchor biosynthesis, class C |
| chr9_-_70141484 | 0.03 |
ENSMUST00000034749.8
|
Fam81a
|
family with sequence similarity 81, member A |
| chr19_-_4793851 | 0.03 |
ENSMUST00000178615.1
ENSMUST00000179189.1 ENSMUST00000164376.2 ENSMUST00000164209.2 ENSMUST00000180248.1 |
Rbm4
|
RNA binding motif protein 4 |
| chr10_+_56377300 | 0.03 |
ENSMUST00000068581.7
|
Gja1
|
gap junction protein, alpha 1 |
| chr3_-_84304762 | 0.02 |
ENSMUST00000107692.1
|
Trim2
|
tripartite motif-containing 2 |
| chr9_+_108080436 | 0.02 |
ENSMUST00000035211.7
ENSMUST00000162886.1 |
Mst1
|
macrophage stimulating 1 (hepatocyte growth factor-like) |
| chr7_+_5034118 | 0.02 |
ENSMUST00000076791.3
|
4632433K11Rik
|
RIKEN cDNA 4632433K11 gene |
| chr5_+_34999046 | 0.02 |
ENSMUST00000114281.1
|
Rgs12
|
regulator of G-protein signaling 12 |
| chr2_+_136892168 | 0.01 |
ENSMUST00000099311.2
|
Slx4ip
|
SLX4 interacting protein |
| chr5_+_34999070 | 0.01 |
ENSMUST00000114280.1
|
Rgs12
|
regulator of G-protein signaling 12 |
| chr3_+_96172327 | 0.01 |
ENSMUST00000076372.4
|
Sf3b4
|
splicing factor 3b, subunit 4 |
| chr18_+_80253274 | 0.01 |
ENSMUST00000131780.1
ENSMUST00000157056.1 |
Pqlc1
|
PQ loop repeat containing 1 |
| chr17_+_32506446 | 0.01 |
ENSMUST00000165999.1
|
Cyp4f17
|
cytochrome P450, family 4, subfamily f, polypeptide 17 |
| chr7_+_24399921 | 0.01 |
ENSMUST00000108434.1
|
Smg9
|
smg-9 homolog, nonsense mediated mRNA decay factor (C. elegans) |
| chr10_+_80622677 | 0.01 |
ENSMUST00000079773.6
|
Csnk1g2
|
casein kinase 1, gamma 2 |
| chr17_+_52602700 | 0.01 |
ENSMUST00000039366.10
|
Kcnh8
|
potassium voltage-gated channel, subfamily H (eag-related), member 8 |
| chr18_-_35703108 | 0.01 |
ENSMUST00000025208.5
|
Dnajc18
|
DnaJ (Hsp40) homolog, subfamily C, member 18 |
| chr8_-_84893887 | 0.01 |
ENSMUST00000003907.7
ENSMUST00000182458.1 ENSMUST00000109745.1 ENSMUST00000142748.1 |
Gcdh
|
glutaryl-Coenzyme A dehydrogenase |
| chr8_+_35587780 | 0.00 |
ENSMUST00000037666.5
|
Mfhas1
|
malignant fibrous histiocytoma amplified sequence 1 |
| chr17_-_21933022 | 0.00 |
ENSMUST00000074295.7
|
Zfp942
|
zinc finger protein 942 |
| chr1_+_161969284 | 0.00 |
ENSMUST00000160881.1
ENSMUST00000159648.1 |
Pigc
|
phosphatidylinositol glycan anchor biosynthesis, class C |
| chr2_+_29890063 | 0.00 |
ENSMUST00000028128.6
|
Odf2
|
outer dense fiber of sperm tails 2 |
| chr11_+_88204396 | 0.00 |
ENSMUST00000118784.1
ENSMUST00000139170.1 ENSMUST00000107915.3 ENSMUST00000144070.1 |
Mrps23
|
mitochondrial ribosomal protein S23 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Trp53
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.1 | GO:0072382 | minus-end-directed vesicle transport along microtubule(GO:0072382) |
| 0.3 | 3.3 | GO:0071493 | cellular response to UV-B(GO:0071493) |
| 0.3 | 1.6 | GO:0061110 | dense core granule biogenesis(GO:0061110) |
| 0.3 | 1.0 | GO:0002540 | leukotriene production involved in inflammatory response(GO:0002540) |
| 0.3 | 1.0 | GO:1900740 | regulation of protein insertion into mitochondrial membrane involved in apoptotic signaling pathway(GO:1900739) positive regulation of protein insertion into mitochondrial membrane involved in apoptotic signaling pathway(GO:1900740) |
| 0.2 | 0.9 | GO:0072108 | positive regulation of mesenchymal to epithelial transition involved in metanephros morphogenesis(GO:0072108) |
| 0.2 | 0.6 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 0.2 | 0.6 | GO:0001807 | regulation of type IV hypersensitivity(GO:0001807) negative regulation of hypersensitivity(GO:0002884) |
| 0.2 | 3.0 | GO:0046415 | urate metabolic process(GO:0046415) |
| 0.2 | 0.7 | GO:0060265 | positive regulation of respiratory burst involved in inflammatory response(GO:0060265) |
| 0.2 | 0.5 | GO:0042222 | interleukin-1 biosynthetic process(GO:0042222) |
| 0.2 | 3.7 | GO:0007095 | mitotic G2 DNA damage checkpoint(GO:0007095) |
| 0.2 | 0.5 | GO:0002018 | renin-angiotensin regulation of aldosterone production(GO:0002018) |
| 0.1 | 0.5 | GO:0061153 | trachea submucosa development(GO:0061152) trachea gland development(GO:0061153) |
| 0.1 | 0.6 | GO:0007000 | nucleolus organization(GO:0007000) |
| 0.1 | 0.5 | GO:0014045 | establishment of endothelial blood-brain barrier(GO:0014045) |
| 0.1 | 0.2 | GO:0006533 | fumarate metabolic process(GO:0006106) glycerol biosynthetic process(GO:0006114) aspartate biosynthetic process(GO:0006532) aspartate catabolic process(GO:0006533) |
| 0.1 | 0.5 | GO:0030382 | sperm mitochondrion organization(GO:0030382) |
| 0.1 | 1.1 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.1 | 0.2 | GO:0042264 | peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 0.1 | 0.2 | GO:0010808 | positive regulation of synaptic vesicle priming(GO:0010808) |
| 0.1 | 0.2 | GO:0032877 | positive regulation of DNA endoreduplication(GO:0032877) |
| 0.1 | 0.2 | GO:0016561 | protein import into peroxisome matrix, translocation(GO:0016561) |
| 0.1 | 0.3 | GO:0060316 | positive regulation of ryanodine-sensitive calcium-release channel activity(GO:0060316) |
| 0.0 | 0.2 | GO:0097195 | pilomotor reflex(GO:0097195) positive regulation of transcription from RNA polymerase II promoter involved in smooth muscle cell differentiation(GO:2000721) |
| 0.0 | 1.6 | GO:0007026 | negative regulation of microtubule depolymerization(GO:0007026) |
| 0.0 | 0.7 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.0 | 0.2 | GO:0015862 | uridine transport(GO:0015862) |
| 0.0 | 0.3 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.0 | 0.4 | GO:0006517 | protein deglycosylation(GO:0006517) |
| 0.0 | 0.4 | GO:0015697 | organic cation transport(GO:0015695) quaternary ammonium group transport(GO:0015697) |
| 0.0 | 0.2 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 0.0 | 0.2 | GO:0060718 | chorionic trophoblast cell differentiation(GO:0060718) |
| 0.0 | 0.1 | GO:0014826 | vein smooth muscle contraction(GO:0014826) |
| 0.0 | 1.3 | GO:0051220 | cytoplasmic sequestering of protein(GO:0051220) |
| 0.0 | 0.1 | GO:0006543 | glutamine catabolic process(GO:0006543) |
| 0.0 | 0.3 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 0.0 | 0.4 | GO:0030238 | male sex determination(GO:0030238) |
| 0.0 | 0.1 | GO:0003430 | growth plate cartilage chondrocyte growth(GO:0003430) |
| 0.0 | 0.5 | GO:0000289 | nuclear-transcribed mRNA poly(A) tail shortening(GO:0000289) |
| 0.0 | 0.3 | GO:0019236 | response to pheromone(GO:0019236) |
| 0.0 | 0.4 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.0 | 0.3 | GO:0042759 | long-chain fatty acid biosynthetic process(GO:0042759) |
| 0.0 | 0.1 | GO:0033153 | somatic diversification of T cell receptor genes(GO:0002568) somatic recombination of T cell receptor gene segments(GO:0002681) T cell receptor V(D)J recombination(GO:0033153) double-strand break repair via classical nonhomologous end joining(GO:0097680) |
| 0.0 | 0.1 | GO:0043031 | negative regulation of macrophage activation(GO:0043031) |
| 0.0 | 0.1 | GO:0046381 | CMP-N-acetylneuraminate metabolic process(GO:0046381) |
| 0.0 | 0.1 | GO:0019243 | methylglyoxal catabolic process to D-lactate via S-lactoyl-glutathione(GO:0019243) methylglyoxal catabolic process(GO:0051596) methylglyoxal catabolic process to lactate(GO:0061727) |
| 0.0 | 0.0 | GO:0003347 | epicardial cell to mesenchymal cell transition(GO:0003347) |
| 0.0 | 0.6 | GO:0060441 | epithelial tube branching involved in lung morphogenesis(GO:0060441) |
| 0.0 | 0.2 | GO:2001269 | positive regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:2001269) |
| 0.0 | 0.6 | GO:0035458 | cellular response to interferon-beta(GO:0035458) |
| 0.0 | 0.1 | GO:0035456 | response to interferon-beta(GO:0035456) |
| 0.0 | 0.4 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.0 | 0.1 | GO:0050882 | voluntary musculoskeletal movement(GO:0050882) |
| 0.0 | 0.0 | GO:0090297 | positive regulation of mitochondrial DNA replication(GO:0090297) regulation of cardiolipin metabolic process(GO:1900208) positive regulation of cardiolipin metabolic process(GO:1900210) stress-induced mitochondrial fusion(GO:1990046) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 3.7 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 0.1 | 1.6 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.1 | 1.6 | GO:0097539 | ciliary transition fiber(GO:0097539) |
| 0.1 | 1.9 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.1 | 0.4 | GO:0000235 | astral microtubule(GO:0000235) |
| 0.1 | 0.2 | GO:0098833 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) |
| 0.1 | 1.0 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.1 | 0.5 | GO:0097227 | sperm annulus(GO:0097227) |
| 0.0 | 0.2 | GO:0098831 | presynaptic active zone cytoplasmic component(GO:0098831) |
| 0.0 | 0.5 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 0.9 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.0 | 2.7 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 0.2 | GO:1990415 | Pex17p-Pex14p docking complex(GO:1990415) peroxisomal importomer complex(GO:1990429) |
| 0.0 | 0.6 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.0 | 0.3 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.0 | 0.2 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.0 | 1.4 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.2 | GO:0097413 | Lewy body(GO:0097413) |
| 0.0 | 0.2 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 0.0 | 0.6 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.0 | 0.1 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.0 | 0.3 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.0 | 0.1 | GO:0005958 | DNA-dependent protein kinase-DNA ligase 4 complex(GO:0005958) DNA ligase IV complex(GO:0032807) |
| 0.0 | 1.4 | GO:0005643 | nuclear pore(GO:0005643) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 3.7 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 0.3 | 1.0 | GO:0004051 | arachidonate 5-lipoxygenase activity(GO:0004051) |
| 0.3 | 3.0 | GO:0015143 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.2 | 0.9 | GO:0005146 | leukemia inhibitory factor receptor binding(GO:0005146) |
| 0.2 | 0.6 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.2 | 1.1 | GO:0004859 | phospholipase inhibitor activity(GO:0004859) |
| 0.1 | 0.4 | GO:0005333 | norepinephrine transmembrane transporter activity(GO:0005333) |
| 0.1 | 0.7 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.1 | 0.5 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) |
| 0.1 | 0.2 | GO:0070546 | L-aspartate:2-oxoglutarate aminotransferase activity(GO:0004069) L-phenylalanine aminotransferase activity(GO:0070546) L-phenylalanine:2-oxoglutarate aminotransferase activity(GO:0080130) |
| 0.1 | 2.1 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.1 | 0.4 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.1 | 0.3 | GO:0008853 | exodeoxyribonuclease III activity(GO:0008853) |
| 0.1 | 0.4 | GO:0005113 | patched binding(GO:0005113) |
| 0.1 | 0.2 | GO:0070737 | protein-glycine ligase activity, elongating(GO:0070737) |
| 0.1 | 1.1 | GO:0004862 | cAMP-dependent protein kinase inhibitor activity(GO:0004862) |
| 0.0 | 0.2 | GO:0016721 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.0 | 0.1 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.0 | 0.4 | GO:0001640 | adenylate cyclase inhibiting G-protein coupled glutamate receptor activity(GO:0001640) |
| 0.0 | 0.6 | GO:0001054 | RNA polymerase I activity(GO:0001054) |
| 0.0 | 0.5 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.0 | 0.6 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.0 | 0.2 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.1 | GO:0030899 | calcium-dependent ATPase activity(GO:0030899) |
| 0.0 | 0.5 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.0 | 0.1 | GO:0004920 | interleukin-10 receptor activity(GO:0004920) |
| 0.0 | 0.5 | GO:0005123 | death receptor binding(GO:0005123) |
| 0.0 | 0.1 | GO:0004359 | glutaminase activity(GO:0004359) |
| 0.0 | 0.7 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.0 | 0.1 | GO:0030338 | CMP-N-acetylneuraminate monooxygenase activity(GO:0030338) |
| 0.0 | 0.5 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.0 | 0.5 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.0 | 0.2 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.1 | GO:0019958 | C-X-C chemokine binding(GO:0019958) |
| 0.0 | 0.1 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.0 | 0.1 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.0 | 0.1 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.0 | 0.4 | GO:0005549 | odorant binding(GO:0005549) |
| 0.0 | 0.1 | GO:0004416 | hydroxyacylglutathione hydrolase activity(GO:0004416) |
| 0.0 | 0.1 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.0 | 0.0 | GO:0034584 | piRNA binding(GO:0034584) |
| 0.0 | 0.1 | GO:0050265 | RNA uridylyltransferase activity(GO:0050265) |
| 0.0 | 0.3 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.0 | 1.1 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.1 | GO:0046790 | virion binding(GO:0046790) |
| 0.0 | 0.1 | GO:0003909 | DNA ligase activity(GO:0003909) DNA ligase (ATP) activity(GO:0003910) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 3.7 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.1 | 1.0 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 4.1 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.0 | 1.3 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 0.6 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.0 | 0.4 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 1.0 | PID P73PATHWAY | p73 transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.0 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.1 | 3.7 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.1 | 2.1 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.1 | 1.0 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.0 | 0.6 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 1.3 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.0 | 0.6 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.0 | 0.9 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.0 | 0.6 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |
| 0.0 | 0.5 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.0 | 0.4 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.0 | 0.5 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 0.2 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 0.7 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 0.1 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |