Project
DANIO-CODE
Navigation
Downloads
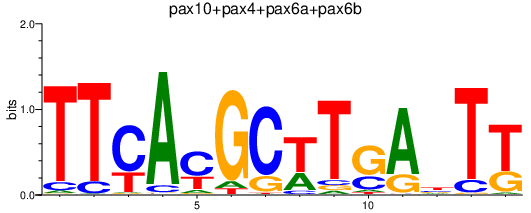
Results for pax10+pax4+pax6a+pax6b
Z-value: 0.99
Transcription factors associated with pax10+pax4+pax6a+pax6b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
pax4
|
ENSDARG00000021336 | paired box 4 |
|
pax6b
|
ENSDARG00000045936 | paired box 6b |
|
pax10
|
ENSDARG00000053364 | paired box 10 |
|
pax6a
|
ENSDARG00000103379 | paired box 6a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| pax6b | dr10_dc_chr7_+_15619409_15619438 | -0.61 | 1.2e-02 | Click! |
| pax6a | dr10_dc_chr25_-_14949028_14949084 | -0.58 | 1.9e-02 | Click! |
Activity profile of pax10+pax4+pax6a+pax6b motif
Sorted Z-values of pax10+pax4+pax6a+pax6b motif
Network of associatons between targets according to the STRING database.
First level regulatory network of pax10+pax4+pax6a+pax6b
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_+_52456064 | 2.92 |
ENSDART00000012758
|
zgc:77112
|
zgc:77112 |
| chr16_+_39209567 | 2.55 |
ENSDART00000121756
|
sybu
|
syntabulin (syntaxin-interacting) |
| chr5_+_57254393 | 2.06 |
ENSDART00000050949
|
btg4
|
B-cell translocation gene 4 |
| chr18_+_14308032 | 2.05 |
|
|
|
| chr17_-_30884905 | 2.01 |
ENSDART00000131633
ENSDART00000146824 |
evla
|
Enah/Vasp-like a |
| chr10_-_8088063 | 1.90 |
ENSDART00000099031
|
zgc:136254
|
zgc:136254 |
| chr21_-_2296253 | 1.85 |
ENSDART00000162867
|
zgc:66483
|
zgc:66483 |
| chr10_-_8094671 | 1.54 |
ENSDART00000099033
|
zgc:158494
|
zgc:158494 |
| chr21_-_2350174 | 1.37 |
ENSDART00000168946
|
si:ch211-241b2.4
|
si:ch211-241b2.4 |
| chr16_-_39209426 | 1.36 |
|
|
|
| chr25_-_8548041 | 1.34 |
ENSDART00000155280
|
GDPGP1
|
GDP-D-glucose phosphorylase 1 |
| chr9_+_8990576 | 1.31 |
ENSDART00000133899
|
ube2al
|
ubiquitin conjugating enzyme E2 A, like |
| chr19_-_4876806 | 1.28 |
ENSDART00000141336
ENSDART00000110551 |
st3gal1
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 1 |
| chr2_+_36110584 | 1.27 |
ENSDART00000163189
|
BX681417.4
|
ENSDARG00000101668 |
| chr12_-_35481361 | 1.26 |
ENSDART00000158658
ENSDART00000168958 ENSDART00000162175 |
sec24c
|
SEC24 homolog C, COPII coat complex component |
| chr4_-_72228164 | 1.25 |
|
|
|
| chr21_-_3548863 | 1.24 |
ENSDART00000086492
|
atp8b1
|
ATPase, aminophospholipid transporter, class I, type 8B, member 1 |
| chr18_-_20476969 | 1.24 |
ENSDART00000060311
|
paqr5a
|
progestin and adipoQ receptor family member Va |
| chr23_+_17999913 | 1.23 |
ENSDART00000012540
|
chia.4
|
chitinase, acidic.4 |
| chr21_-_2350090 | 1.22 |
ENSDART00000168946
|
si:ch211-241b2.4
|
si:ch211-241b2.4 |
| chr25_-_1243081 | 1.13 |
ENSDART00000156062
|
calml4b
|
calmodulin-like 4b |
| chr22_-_21021942 | 1.12 |
ENSDART00000133982
|
ssbp4
|
single stranded DNA binding protein 4 |
| chr9_+_8387050 | 1.09 |
ENSDART00000136847
|
si:dkey-90l23.2
|
si:dkey-90l23.2 |
| chr10_+_19610019 | 1.07 |
|
|
|
| chr25_+_7358777 | 0.99 |
ENSDART00000161593
|
ptdss2
|
phosphatidylserine synthase 2 |
| chr3_-_7487722 | 0.98 |
|
|
|
| chr14_-_14353487 | 0.97 |
ENSDART00000172241
|
nsdhl
|
NAD(P) dependent steroid dehydrogenase-like |
| chr14_+_21373857 | 0.95 |
ENSDART00000079649
ENSDART00000165795 |
ndufs8b
|
NADH dehydrogenase (ubiquinone) Fe-S protein 8b |
| chr25_-_9889107 | 0.93 |
ENSDART00000137407
|
AL929493.1
|
ENSDARG00000093575 |
| chr19_+_43090603 | 0.91 |
ENSDART00000018328
|
fbxl2
|
F-box and leucine-rich repeat protein 2 |
| chr5_+_47324215 | 0.89 |
ENSDART00000097429
|
BX470189.1
|
ENSDARG00000067658 |
| chr13_-_32595706 | 0.89 |
ENSDART00000012232
|
pdss2
|
prenyl (decaprenyl) diphosphate synthase, subunit 2 |
| chr14_+_31189527 | 0.88 |
ENSDART00000053026
|
fam122b
|
family with sequence similarity 122B |
| chr18_-_50439653 | 0.85 |
ENSDART00000151038
|
CU862078.1
|
ENSDARG00000096262 |
| chr13_-_49886891 | 0.84 |
ENSDART00000074230
|
pkz
|
protein kinase containing Z-DNA binding domains |
| chr7_+_26274744 | 0.83 |
ENSDART00000135313
|
tnk1
|
tyrosine kinase, non-receptor, 1 |
| chr14_-_30578373 | 0.83 |
ENSDART00000176631
|
si:ch211-126c2.4
|
si:ch211-126c2.4 |
| chr1_-_28849479 | 0.82 |
ENSDART00000109224
|
ENSDARG00000074117
|
ENSDARG00000074117 |
| chr12_+_23691261 | 0.82 |
ENSDART00000066331
|
svila
|
supervillin a |
| chr24_-_36383243 | 0.82 |
ENSDART00000155892
|
si:ch211-40k21.5
|
si:ch211-40k21.5 |
| chr7_-_30353095 | 0.81 |
ENSDART00000173828
|
rnf111
|
ring finger protein 111 |
| chr12_-_13692190 | 0.80 |
ENSDART00000152370
|
foxh1
|
forkhead box H1 |
| chr12_-_3042394 | 0.79 |
ENSDART00000002867
ENSDART00000126315 |
rfng
|
RFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| KN150663v1_-_3381 | 0.77 |
|
|
|
| chr3_-_54352535 | 0.77 |
ENSDART00000021977
ENSDART00000078973 |
dnmt1
|
DNA (cytosine-5-)-methyltransferase 1 |
| chr5_-_68751287 | 0.76 |
ENSDART00000112692
|
ENSDARG00000077155
|
ENSDARG00000077155 |
| chr10_+_24721086 | 0.76 |
ENSDART00000079549
|
tpte
|
transmembrane phosphatase with tensin homology |
| chr24_+_28304276 | 0.76 |
ENSDART00000018095
|
sh3glb1a
|
SH3-domain GRB2-like endophilin B1a |
| chr16_-_5205600 | 0.75 |
ENSDART00000148955
|
bckdhb
|
branched chain keto acid dehydrogenase E1, beta polypeptide |
| chr9_+_8990774 | 0.74 |
ENSDART00000133899
|
ube2al
|
ubiquitin conjugating enzyme E2 A, like |
| chr20_-_25587274 | 0.74 |
ENSDART00000141340
|
si:dkey-183n20.15
|
si:dkey-183n20.15 |
| chr19_+_43090664 | 0.74 |
ENSDART00000018328
|
fbxl2
|
F-box and leucine-rich repeat protein 2 |
| chr5_-_10966029 | 0.71 |
|
|
|
| chr11_-_11353309 | 0.70 |
ENSDART00000016677
|
zgc:77929
|
zgc:77929 |
| chr22_+_8973724 | 0.70 |
ENSDART00000106414
|
rnh1
|
ribonuclease/angiogenin inhibitor 1 |
| chr16_+_39209538 | 0.69 |
ENSDART00000121756
|
sybu
|
syntabulin (syntaxin-interacting) |
| chr3_-_7487630 | 0.68 |
|
|
|
| chr16_+_11888601 | 0.68 |
ENSDART00000133497
|
si:dkey-250k15.4
|
si:dkey-250k15.4 |
| chr21_-_33070005 | 0.67 |
|
|
|
| chr8_+_25873546 | 0.67 |
ENSDART00000137247
|
tmem115
|
transmembrane protein 115 |
| chr22_-_21021984 | 0.67 |
ENSDART00000133982
|
ssbp4
|
single stranded DNA binding protein 4 |
| chr20_-_48957424 | 0.66 |
ENSDART00000158993
|
BX663521.1
|
ENSDARG00000102751 |
| chr8_+_26064660 | 0.66 |
ENSDART00000099283
|
dalrd3
|
DALR anticodon binding domain containing 3 |
| chr1_+_51087450 | 0.66 |
ENSDART00000040397
|
prdx2
|
peroxiredoxin 2 |
| chr25_+_4508802 | 0.65 |
ENSDART00000021120
|
pgghg
|
protein-glucosylgalactosylhydroxylysine glucosidase |
| chr7_-_45579815 | 0.65 |
ENSDART00000170224
|
shcbp1
|
SHC SH2-domain binding protein 1 |
| chr15_+_20054306 | 0.65 |
ENSDART00000155199
|
zgc:112083
|
zgc:112083 |
| chr13_-_32595667 | 0.65 |
ENSDART00000012232
|
pdss2
|
prenyl (decaprenyl) diphosphate synthase, subunit 2 |
| chr21_-_39132507 | 0.64 |
ENSDART00000065143
|
unc119b
|
unc-119 homolog b (C. elegans) |
| chr7_-_41232765 | 0.64 |
ENSDART00000173577
|
ENSDARG00000105669
|
ENSDARG00000105669 |
| chr7_+_26274687 | 0.64 |
ENSDART00000135313
|
tnk1
|
tyrosine kinase, non-receptor, 1 |
| chr23_-_27579257 | 0.63 |
ENSDART00000137229
|
asb8
|
ankyrin repeat and SOCS box containing 8 |
| chr7_-_20330881 | 0.63 |
ENSDART00000169750
|
si:dkey-19b23.11
|
si:dkey-19b23.11 |
| chr20_-_47470364 | 0.63 |
ENSDART00000040679
|
dtnba
|
dystrobrevin, beta a |
| chr19_-_41933808 | 0.63 |
ENSDART00000062080
|
chrac1
|
chromatin accessibility complex 1 |
| chr7_-_30353179 | 0.62 |
ENSDART00000173828
|
rnf111
|
ring finger protein 111 |
| chr20_-_48957517 | 0.61 |
ENSDART00000158993
|
BX663521.1
|
ENSDARG00000102751 |
| chr8_-_44230474 | 0.61 |
ENSDART00000143020
|
stx2b
|
syntaxin 2b |
| chr13_+_8509859 | 0.61 |
ENSDART00000140489
|
epcam
|
epithelial cell adhesion molecule |
| chr4_+_11763234 | 0.60 |
ENSDART00000153714
ENSDART00000156950 |
BX465862.3
|
ENSDARG00000096099 |
| chr7_-_8492862 | 0.60 |
ENSDART00000172807
ENSDART00000173026 |
si:ch211-1o7.3
|
si:ch211-1o7.3 |
| chr19_-_42933592 | 0.57 |
ENSDART00000007642
|
zgc:110239
|
zgc:110239 |
| chr3_-_54352640 | 0.57 |
ENSDART00000128041
|
dnmt1
|
DNA (cytosine-5-)-methyltransferase 1 |
| chr11_+_42438405 | 0.57 |
ENSDART00000163269
|
zgc:194981
|
zgc:194981 |
| chr8_+_41004169 | 0.57 |
|
|
|
| chr23_-_27318468 | 0.57 |
|
|
|
| chr3_-_23382550 | 0.57 |
ENSDART00000170200
|
BX682558.1
|
ENSDARG00000101975 |
| chr19_-_11320648 | 0.55 |
ENSDART00000148697
|
ssr2
|
signal sequence receptor, beta |
| chr19_+_20579471 | 0.54 |
ENSDART00000090942
|
ccdc126
|
coiled-coil domain containing 126 |
| chr19_-_20578899 | 0.54 |
ENSDART00000151646
|
fam221a
|
family with sequence similarity 221, member A |
| chr8_+_41003546 | 0.54 |
ENSDART00000129344
|
gpat2
|
glycerol-3-phosphate acyltransferase 2, mitochondrial |
| chr6_-_51811024 | 0.53 |
ENSDART00000031597
|
psmf1
|
proteasome inhibitor subunit 1 |
| chr5_+_47324353 | 0.53 |
ENSDART00000097429
|
BX470189.1
|
ENSDARG00000067658 |
| chr25_-_8548210 | 0.53 |
ENSDART00000155280
|
GDPGP1
|
GDP-D-glucose phosphorylase 1 |
| chr5_+_31541677 | 0.53 |
ENSDART00000041504
|
tescb
|
tescalcin b |
| chr19_-_3785774 | 0.53 |
ENSDART00000161738
|
smim13
|
small integral membrane protein 13 |
| chr16_+_11888545 | 0.52 |
ENSDART00000133497
|
si:dkey-250k15.4
|
si:dkey-250k15.4 |
| chr18_+_36801847 | 0.52 |
ENSDART00000004129
|
si:ch211-160d20.3
|
si:ch211-160d20.3 |
| chr3_-_54414615 | 0.52 |
ENSDART00000053106
|
s1pr2
|
sphingosine-1-phosphate receptor 2 |
| chr12_+_23691397 | 0.52 |
ENSDART00000066331
|
svila
|
supervillin a |
| chr1_+_45148002 | 0.52 |
ENSDART00000148086
|
map2k7
|
mitogen-activated protein kinase kinase 7 |
| chr5_-_45986774 | 0.51 |
|
|
|
| chr23_-_18454945 | 0.50 |
ENSDART00000016891
|
hsd17b10
|
hydroxysteroid (17-beta) dehydrogenase 10 |
| chr15_-_743589 | 0.50 |
ENSDART00000156200
|
zgc:174574
|
zgc:174574 |
| chr22_+_22413803 | 0.49 |
ENSDART00000147825
|
kif14
|
kinesin family member 14 |
| chr25_+_19772873 | 0.48 |
ENSDART00000170493
|
gramd4b
|
GRAM domain containing 4b |
| chr13_+_50167246 | 0.47 |
ENSDART00000178572
|
CU570781.2
|
ENSDARG00000108623 |
| chr23_-_16160099 | 0.47 |
ENSDART00000042469
ENSDART00000146605 |
mrgbp
|
MRG/MORF4L binding protein |
| chr7_-_3986977 | 0.47 |
|
|
|
| KN150101v1_-_19017 | 0.47 |
|
|
|
| chr19_-_11097075 | 0.47 |
ENSDART00000010997
|
tpm3
|
tropomyosin 3 |
| chr24_-_26334771 | 0.47 |
ENSDART00000079984
ENSDART00000136871 |
rpl22l1
|
ribosomal protein L22-like 1 |
| chr21_-_39521698 | 0.46 |
ENSDART00000020174
|
dynll2b
|
dynein, light chain, LC8-type 2b |
| chr12_+_20685139 | 0.46 |
ENSDART00000153261
|
BX004803.1
|
ENSDARG00000096799 |
| chr5_-_23763146 | 0.46 |
|
|
|
| chr9_+_9956221 | 0.46 |
ENSDART00000168559
|
ttf2
|
transcription termination factor, RNA polymerase II |
| chr11_+_19440815 | 0.46 |
ENSDART00000005639
|
thoc7
|
THO complex 7 |
| chr5_+_25612385 | 0.46 |
ENSDART00000098514
|
oclnb
|
occludin b |
| chr21_-_39132951 | 0.46 |
ENSDART00000065143
|
unc119b
|
unc-119 homolog b (C. elegans) |
| chr5_+_38225036 | 0.45 |
ENSDART00000076835
|
mrpl1
|
mitochondrial ribosomal protein L1 |
| chr19_+_32614657 | 0.45 |
ENSDART00000022667
|
fam8a1a
|
family with sequence similarity 8, member A1a |
| chr14_-_14353451 | 0.44 |
ENSDART00000170355
ENSDART00000159888 |
nsdhl
|
NAD(P) dependent steroid dehydrogenase-like |
| chr5_-_47323844 | 0.44 |
ENSDART00000145665
ENSDART00000007057 |
ccnh
|
cyclin H |
| chr7_-_30353041 | 0.43 |
ENSDART00000173828
|
rnf111
|
ring finger protein 111 |
| chr3_+_36970806 | 0.42 |
ENSDART00000055228
|
psmc3ip
|
PSMC3 interacting protein |
| chr6_+_33092094 | 0.42 |
ENSDART00000122242
|
pomgnt1
|
protein O-linked mannose N-acetylglucosaminyltransferase 1 (beta 1,2-) |
| chr8_+_48954402 | 0.42 |
|
|
|
| chr5_+_61711614 | 0.41 |
ENSDART00000082965
|
ABR
|
active BCR-related |
| chr1_+_11381915 | 0.41 |
ENSDART00000055566
|
xpa
|
xeroderma pigmentosum, complementation group A |
| chr9_-_21677941 | 0.41 |
ENSDART00000121939
ENSDART00000080404 |
mphosph8
|
M-phase phosphoprotein 8 |
| chr25_-_19927083 | 0.41 |
ENSDART00000140182
|
cnot4a
|
CCR4-NOT transcription complex, subunit 4a |
| chr13_+_50167216 | 0.41 |
ENSDART00000178572
|
CU570781.2
|
ENSDARG00000108623 |
| chr25_-_8548430 | 0.41 |
ENSDART00000155280
|
GDPGP1
|
GDP-D-glucose phosphorylase 1 |
| chr14_+_7596207 | 0.41 |
ENSDART00000113299
|
zgc:110843
|
zgc:110843 |
| chr20_-_3220475 | 0.40 |
ENSDART00000123331
|
spint1b
|
serine peptidase inhibitor, Kunitz type 1 b |
| chr5_-_27416678 | 0.40 |
ENSDART00000078642
|
vps37b
|
vacuolar protein sorting 37 homolog B (S. cerevisiae) |
| chr23_+_44795356 | 0.39 |
ENSDART00000145905
|
eno3
|
enolase 3, (beta, muscle) |
| chr25_-_19926991 | 0.39 |
ENSDART00000140182
|
cnot4a
|
CCR4-NOT transcription complex, subunit 4a |
| chr13_+_28655433 | 0.39 |
ENSDART00000039028
|
nsmce4a
|
NSE4 homolog A, SMC5-SMC6 complex component |
| chr7_-_19274727 | 0.38 |
ENSDART00000114203
|
man2b2
|
mannosidase, alpha, class 2B, member 2 |
| chr18_-_21695887 | 0.38 |
ENSDART00000019861
ENSDART00000114292 |
mbtps1
|
membrane-bound transcription factor peptidase, site 1 |
| chr20_+_34248925 | 0.38 |
ENSDART00000145852
|
arpc5b
|
actin related protein 2/3 complex, subunit 5B |
| chr23_-_7191380 | 0.38 |
ENSDART00000176914
|
slco4a1
|
solute carrier organic anion transporter family, member 4A1 |
| chr1_+_10423544 | 0.38 |
ENSDART00000109858
|
knstrn
|
kinetochore-localized astrin/SPAG5 binding protein |
| chr10_+_24721139 | 0.38 |
ENSDART00000144710
|
tpte
|
transmembrane phosphatase with tensin homology |
| chr17_-_20159972 | 0.37 |
|
|
|
| chr19_-_27958005 | 0.37 |
ENSDART00000114301
|
si:ch211-152p11.4
|
si:ch211-152p11.4 |
| chr20_-_14784875 | 0.37 |
ENSDART00000063857
|
scrn2
|
secernin 2 |
| chr12_-_3042573 | 0.37 |
ENSDART00000002867
ENSDART00000126315 |
rfng
|
RFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr14_+_31189237 | 0.36 |
ENSDART00000053026
|
fam122b
|
family with sequence similarity 122B |
| chr2_-_31728072 | 0.36 |
ENSDART00000113498
|
lrrcc1
|
leucine rich repeat and coiled-coil centrosomal protein 1 |
| chr3_-_47294091 | 0.36 |
|
|
|
| chr16_+_22114192 | 0.36 |
ENSDART00000167919
|
znf687a
|
zinc finger protein 687a |
| chr25_-_24150237 | 0.35 |
ENSDART00000073527
|
spty2d1
|
SPT2 chromatin protein domain containing 1 |
| chr3_+_62116865 | 0.35 |
ENSDART00000112428
|
iqck
|
IQ motif containing K |
| chr24_-_36382704 | 0.35 |
ENSDART00000154858
|
si:ch211-40k21.5
|
si:ch211-40k21.5 |
| chr21_+_21871674 | 0.35 |
|
|
|
| chr23_-_27318392 | 0.35 |
|
|
|
| chr13_+_8509927 | 0.34 |
ENSDART00000140489
|
epcam
|
epithelial cell adhesion molecule |
| chr18_-_15300493 | 0.34 |
ENSDART00000048206
|
tmem263
|
transmembrane protein 263 |
| chr20_+_27813565 | 0.33 |
ENSDART00000008306
|
zbtb1
|
zinc finger and BTB domain containing 1 |
| chr25_-_8548290 | 0.33 |
ENSDART00000155280
|
GDPGP1
|
GDP-D-glucose phosphorylase 1 |
| chr19_-_6503643 | 0.33 |
|
|
|
| chr11_-_8116319 | 0.32 |
ENSDART00000101561
|
ttll7
|
tubulin tyrosine ligase-like family, member 7 |
| chr10_-_26202233 | 0.31 |
ENSDART00000136472
|
trim3b
|
tripartite motif containing 3b |
| chr14_+_6656429 | 0.31 |
ENSDART00000150050
|
hnrnpaba
|
heterogeneous nuclear ribonucleoprotein A/Ba |
| chr14_+_21874825 | 0.31 |
ENSDART00000114750
|
gabrb2
|
gamma-aminobutyric acid (GABA) A receptor, beta 2 |
| chr11_+_24060826 | 0.31 |
ENSDART00000138487
|
si:dkey-76p14.2
|
si:dkey-76p14.2 |
| chr14_-_23504183 | 0.30 |
ENSDART00000054264
|
nr3c1
|
nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) |
| chr23_-_24615681 | 0.30 |
ENSDART00000132265
|
atp13a2
|
ATPase type 13A2 |
| chr18_-_6011304 | 0.30 |
ENSDART00000122307
|
gcsha
|
glycine cleavage system protein H (aminomethyl carrier), a |
| chr15_+_40130393 | 0.29 |
ENSDART00000063786
|
cab39
|
calcium binding protein 39 |
| chr4_+_13932693 | 0.29 |
ENSDART00000142466
|
pphln1
|
periphilin 1 |
| chr19_-_20578863 | 0.29 |
ENSDART00000148179
|
fam221a
|
family with sequence similarity 221, member A |
| chr12_+_13306551 | 0.28 |
ENSDART00000089017
|
rnasen
|
ribonuclease type III, nuclear |
| chr20_-_3220522 | 0.28 |
ENSDART00000008077
|
spint1b
|
serine peptidase inhibitor, Kunitz type 1 b |
| chr6_-_40411469 | 0.28 |
ENSDART00000154520
|
BX936393.1
|
ENSDARG00000097645 |
| chr21_-_1618494 | 0.28 |
ENSDART00000124904
|
zgc:152948
|
zgc:152948 |
| chr13_+_12538980 | 0.27 |
ENSDART00000010517
|
eif4eb
|
eukaryotic translation initiation factor 4eb |
| chr7_+_8492338 | 0.27 |
|
|
|
| chr25_-_9889072 | 0.27 |
ENSDART00000137407
|
AL929493.1
|
ENSDARG00000093575 |
| chr1_-_25351767 | 0.26 |
ENSDART00000076120
ENSDART00000172737 |
gpank1
|
G patch domain and ankyrin repeats 1 |
| chr24_-_1014582 | 0.26 |
ENSDART00000114544
|
cdk13
|
cyclin-dependent kinase 13 |
| chr8_-_18480333 | 0.26 |
ENSDART00000149446
|
ENSDARG00000077696
|
ENSDARG00000077696 |
| chr19_+_12525407 | 0.26 |
ENSDART00000135706
|
ldlrad4a
|
low density lipoprotein receptor class A domain containing 4a |
| chr3_+_19490148 | 0.26 |
|
|
|
| chr25_-_24150270 | 0.26 |
ENSDART00000073527
|
spty2d1
|
SPT2 chromatin protein domain containing 1 |
| chr22_+_22413878 | 0.26 |
ENSDART00000147825
|
kif14
|
kinesin family member 14 |
| chr10_+_20222743 | 0.26 |
ENSDART00000167008
|
ppp3ccb
|
protein phosphatase 3, catalytic subunit, gamma isozyme, b |
| chr4_-_1968907 | 0.25 |
ENSDART00000067433
|
ube2nb
|
ubiquitin-conjugating enzyme E2Nb |
| chr6_-_8156471 | 0.25 |
ENSDART00000081561
|
ilf3a
|
interleukin enhancer binding factor 3a |
| chr13_-_9737104 | 0.25 |
ENSDART00000158381
|
myom2b
|
myomesin 2b |
| chr16_-_25765512 | 0.25 |
ENSDART00000132693
|
tomm40
|
translocase of outer mitochondrial membrane 40 homolog (yeast) |
| chr22_-_3165441 | 0.25 |
ENSDART00000158009
|
lonp1
|
lon peptidase 1, mitochondrial |
| chr12_-_3630622 | 0.24 |
ENSDART00000164707
|
coa3a
|
cytochrome C oxidase assembly factor 3a |
| chr8_+_41003629 | 0.24 |
ENSDART00000129344
|
gpat2
|
glycerol-3-phosphate acyltransferase 2, mitochondrial |
| chr7_+_21620944 | 0.24 |
ENSDART00000162252
|
pop7
|
POP7 homolog, ribonuclease P/MRP subunit |
| chr16_+_22113874 | 0.24 |
ENSDART00000170604
|
znf687a
|
zinc finger protein 687a |
| chr25_-_10694465 | 0.24 |
ENSDART00000127054
|
BX572619.1
|
ENSDARG00000089303 |
| chr2_+_34675356 | 0.24 |
|
|
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.2 | GO:0006032 | chitin metabolic process(GO:0006030) chitin catabolic process(GO:0006032) |
| 0.3 | 3.2 | GO:0019896 | axonal transport of mitochondrion(GO:0019896) |
| 0.3 | 0.8 | GO:1990511 | piRNA biosynthetic process(GO:1990511) |
| 0.3 | 0.8 | GO:0006420 | arginyl-tRNA aminoacylation(GO:0006420) |
| 0.2 | 0.7 | GO:0010847 | regulation of chromatin assembly(GO:0010847) |
| 0.2 | 1.9 | GO:0030579 | ubiquitin-dependent SMAD protein catabolic process(GO:0030579) |
| 0.2 | 1.1 | GO:2001287 | caveolin-mediated endocytosis(GO:0072584) negative regulation of clathrin-mediated endocytosis(GO:1900186) regulation of caveolin-mediated endocytosis(GO:2001286) negative regulation of caveolin-mediated endocytosis(GO:2001287) |
| 0.2 | 2.9 | GO:0005979 | regulation of glycogen biosynthetic process(GO:0005979) regulation of glucan biosynthetic process(GO:0010962) |
| 0.2 | 0.8 | GO:0010159 | specification of organ position(GO:0010159) |
| 0.1 | 1.0 | GO:0006659 | phosphatidylserine biosynthetic process(GO:0006659) |
| 0.1 | 0.4 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.1 | 0.5 | GO:0051414 | response to cortisol(GO:0051414) cellular response to cortisol stimulus(GO:0071387) |
| 0.1 | 0.4 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.1 | 1.3 | GO:0032776 | DNA methylation on cytosine(GO:0032776) |
| 0.1 | 1.2 | GO:0021592 | fourth ventricle development(GO:0021592) central nervous system segmentation(GO:0035283) brain segmentation(GO:0035284) |
| 0.1 | 1.3 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.1 | 2.0 | GO:0051289 | protein homotetramerization(GO:0051289) |
| 0.1 | 0.4 | GO:0032889 | regulation of vacuole fusion, non-autophagic(GO:0032889) |
| 0.1 | 0.7 | GO:0019430 | response to superoxide(GO:0000303) response to oxygen radical(GO:0000305) removal of superoxide radicals(GO:0019430) cellular response to oxygen radical(GO:0071450) cellular response to superoxide(GO:0071451) cellular oxidant detoxification(GO:0098869) cellular detoxification(GO:1990748) |
| 0.1 | 0.5 | GO:2000580 | positive regulation of microtubule motor activity(GO:2000576) regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000580) positive regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000582) |
| 0.1 | 0.3 | GO:0019464 | glycine catabolic process(GO:0006546) glycine decarboxylation via glycine cleavage system(GO:0019464) |
| 0.1 | 0.4 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.1 | 0.2 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.1 | 0.3 | GO:0010990 | SMAD protein complex assembly(GO:0007183) regulation of SMAD protein complex assembly(GO:0010990) negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.1 | 1.5 | GO:0006743 | ubiquinone metabolic process(GO:0006743) ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.1 | 0.7 | GO:0007584 | response to nutrient(GO:0007584) |
| 0.1 | 0.3 | GO:0021588 | cerebellum formation(GO:0021588) |
| 0.1 | 1.0 | GO:0010257 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.1 | 2.3 | GO:0016574 | histone ubiquitination(GO:0016574) |
| 0.1 | 0.2 | GO:0000967 | endonucleolytic cleavage to generate mature 5'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000472) endonucleolytic cleavage in 5'-ETS of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000480) RNA 5'-end processing(GO:0000966) rRNA 5'-end processing(GO:0000967) ncRNA 5'-end processing(GO:0034471) |
| 0.1 | 0.5 | GO:0034122 | negative regulation of toll-like receptor signaling pathway(GO:0034122) |
| 0.1 | 1.1 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.0 | 1.1 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.0 | 0.4 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.0 | 0.5 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.0 | 3.3 | GO:0008285 | negative regulation of cell proliferation(GO:0008285) |
| 0.0 | 0.4 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 0.0 | 1.0 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.0 | 0.2 | GO:0070131 | positive regulation of mitochondrial translation(GO:0070131) |
| 0.0 | 0.7 | GO:0043507 | activation of JUN kinase activity(GO:0007257) positive regulation of JUN kinase activity(GO:0043507) |
| 0.0 | 1.4 | GO:0001878 | response to yeast(GO:0001878) |
| 0.0 | 0.1 | GO:1902224 | cellular ketone body metabolic process(GO:0046950) ketone body metabolic process(GO:1902224) |
| 0.0 | 0.4 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.0 | 0.8 | GO:0051607 | defense response to virus(GO:0051607) |
| 0.0 | 1.0 | GO:0048920 | posterior lateral line neuromast primordium migration(GO:0048920) |
| 0.0 | 0.6 | GO:0031629 | synaptic vesicle fusion to presynaptic active zone membrane(GO:0031629) vesicle fusion to plasma membrane(GO:0099500) |
| 0.0 | 0.2 | GO:0006654 | phosphatidic acid biosynthetic process(GO:0006654) phosphatidic acid metabolic process(GO:0046473) |
| 0.0 | 0.2 | GO:0051131 | chaperone-mediated protein complex assembly(GO:0051131) |
| 0.0 | 0.6 | GO:0060319 | primitive erythrocyte differentiation(GO:0060319) |
| 0.0 | 1.6 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.0 | 0.2 | GO:0034975 | protein folding in endoplasmic reticulum(GO:0034975) |
| 0.0 | 0.1 | GO:0051877 | eye pigmentation(GO:0048069) pigment granule aggregation in cell center(GO:0051877) axis elongation involved in somitogenesis(GO:0090245) |
| 0.0 | 0.1 | GO:0045144 | meiotic sister chromatid segregation(GO:0045144) meiotic sister chromatid cohesion(GO:0051177) meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 0.0 | 0.1 | GO:0036065 | fucosylation(GO:0036065) |
| 0.0 | 0.9 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 1.2 | GO:0048477 | oogenesis(GO:0048477) |
| 0.0 | 0.5 | GO:0035825 | reciprocal DNA recombination(GO:0035825) |
| 0.0 | 1.2 | GO:0045087 | innate immune response(GO:0045087) |
| 0.0 | 0.5 | GO:0000470 | maturation of LSU-rRNA(GO:0000470) |
| 0.0 | 0.2 | GO:0090502 | RNA phosphodiester bond hydrolysis, endonucleolytic(GO:0090502) |
| 0.0 | 0.0 | GO:0006538 | glutamate catabolic process(GO:0006538) |
| 0.0 | 0.2 | GO:1903725 | regulation of lipid kinase activity(GO:0043550) regulation of phosphatidylinositol 3-kinase activity(GO:0043551) regulation of phospholipid metabolic process(GO:1903725) |
| 0.0 | 0.4 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.1 | GO:0033503 | HULC complex(GO:0033503) |
| 0.2 | 2.9 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.1 | 0.7 | GO:0005947 | mitochondrial alpha-ketoglutarate dehydrogenase complex(GO:0005947) mitochondrial tricarboxylic acid cycle enzyme complex(GO:0030062) |
| 0.1 | 0.3 | GO:0031371 | ubiquitin conjugating enzyme complex(GO:0031371) |
| 0.1 | 0.6 | GO:0008623 | CHRAC(GO:0008623) |
| 0.1 | 3.2 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 1.1 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.1 | 0.4 | GO:0070985 | TFIIK complex(GO:0070985) |
| 0.1 | 0.4 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.1 | 1.0 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.1 | 1.9 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 0.1 | 1.3 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.1 | 0.4 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 0.1 | 0.2 | GO:0000120 | RNA polymerase I transcription factor complex(GO:0000120) |
| 0.1 | 0.6 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.1 | 1.0 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.1 | 0.6 | GO:0019908 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) nuclear cyclin-dependent protein kinase holoenzyme complex(GO:0019908) |
| 0.0 | 1.9 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 0.5 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.0 | 0.4 | GO:0030915 | Smc5-Smc6 complex(GO:0030915) |
| 0.0 | 0.4 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 0.3 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.0 | 0.5 | GO:0043189 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 1.0 | GO:0045271 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 1.8 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 1.6 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 1.6 | GO:0030173 | integral component of Golgi membrane(GO:0030173) intrinsic component of Golgi membrane(GO:0031228) |
| 0.0 | 0.4 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.0 | 1.0 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 1.2 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.0 | 0.1 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.0 | 0.3 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.3 | GO:0003836 | beta-galactoside (CMP) alpha-2,3-sialyltransferase activity(GO:0003836) |
| 0.4 | 1.2 | GO:0004568 | chitinase activity(GO:0004568) |
| 0.4 | 1.9 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.3 | 1.2 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.3 | 0.8 | GO:0004814 | arginine-tRNA ligase activity(GO:0004814) |
| 0.3 | 2.0 | GO:0005522 | profilin binding(GO:0005522) |
| 0.2 | 3.0 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.2 | 2.9 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.2 | 1.3 | GO:0016314 | phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase activity(GO:0016314) |
| 0.2 | 1.4 | GO:0003854 | 3-beta-hydroxy-delta5-steroid dehydrogenase activity(GO:0003854) |
| 0.2 | 0.8 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.2 | 0.7 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.2 | 0.7 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.1 | 0.8 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.1 | 0.4 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.1 | 0.7 | GO:0008545 | JUN kinase kinase activity(GO:0008545) |
| 0.1 | 0.4 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.1 | 1.3 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.1 | 0.4 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.1 | 0.5 | GO:0070740 | protein-glutamic acid ligase activity(GO:0070739) tubulin-glutamic acid ligase activity(GO:0070740) |
| 0.1 | 1.5 | GO:0004659 | prenyltransferase activity(GO:0004659) |
| 0.1 | 0.3 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.1 | 1.9 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.1 | 0.1 | GO:0010521 | telomerase inhibitor activity(GO:0010521) |
| 0.1 | 0.6 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.0 | 0.2 | GO:0004630 | phospholipase D activity(GO:0004630) |
| 0.0 | 1.0 | GO:0003954 | NADH dehydrogenase activity(GO:0003954) |
| 0.0 | 1.0 | GO:0022884 | protein transmembrane transporter activity(GO:0008320) macromolecule transmembrane transporter activity(GO:0022884) |
| 0.0 | 0.4 | GO:0043035 | chromatin insulator sequence binding(GO:0043035) |
| 0.0 | 0.7 | GO:0015926 | glucosidase activity(GO:0015926) |
| 0.0 | 2.6 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 1.1 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 0.5 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.0 | 0.3 | GO:0004723 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.0 | 0.1 | GO:0008410 | CoA-transferase activity(GO:0008410) |
| 0.0 | 0.2 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.0 | 0.3 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 0.0 | 1.8 | GO:0005496 | steroid binding(GO:0005496) |
| 0.0 | 0.1 | GO:0047611 | acetylspermidine deacetylase activity(GO:0047611) |
| 0.0 | 1.2 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.2 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.0 | 1.8 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 1.3 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 0.4 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 0.3 | GO:0004559 | alpha-mannosidase activity(GO:0004559) |
| 0.0 | 0.9 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.6 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.0 | 0.2 | GO:0016671 | oxidoreductase activity, acting on a sulfur group of donors, disulfide as acceptor(GO:0016671) |
| 0.0 | 0.4 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 0.2 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.0 | 0.1 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 1.5 | GO:0000149 | SNARE binding(GO:0000149) |
| 0.0 | 0.3 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.0 | 0.3 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.1 | GO:0016857 | racemase and epimerase activity, acting on carbohydrates and derivatives(GO:0016857) |
| 0.0 | 0.2 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.0 | GO:0016639 | glutamate dehydrogenase (NAD+) activity(GO:0004352) glutamate dehydrogenase [NAD(P)+] activity(GO:0004353) oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.1 | 0.7 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.0 | 1.9 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.4 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 0.6 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.0 | 0.8 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 0.5 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.3 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.1 | 0.3 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.1 | 1.4 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 1.2 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 1.3 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.1 | 0.4 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.1 | 1.2 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 1.0 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.0 | 0.8 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 0.4 | REACTOME ACTIVATION OF CHAPERONES BY ATF6 ALPHA | Genes involved in Activation of Chaperones by ATF6-alpha |
| 0.0 | 0.7 | REACTOME JNK C JUN KINASES PHOSPHORYLATION AND ACTIVATION MEDIATED BY ACTIVATED HUMAN TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 0.0 | 0.4 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.8 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.0 | 0.3 | REACTOME ENERGY DEPENDENT REGULATION OF MTOR BY LKB1 AMPK | Genes involved in Energy dependent regulation of mTOR by LKB1-AMPK |
| 0.0 | 0.5 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.4 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 0.1 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.0 | 0.4 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 1.0 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 0.5 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 0.2 | REACTOME OXYGEN DEPENDENT PROLINE HYDROXYLATION OF HYPOXIA INDUCIBLE FACTOR ALPHA | Genes involved in Oxygen-dependent Proline Hydroxylation of Hypoxia-inducible Factor Alpha |
| 0.0 | 0.5 | REACTOME VIF MEDIATED DEGRADATION OF APOBEC3G | Genes involved in Vif-mediated degradation of APOBEC3G |
| 0.0 | 0.6 | REACTOME GLYCEROPHOSPHOLIPID BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.0 | 0.2 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |