Project
DANIO-CODE
Navigation
Downloads
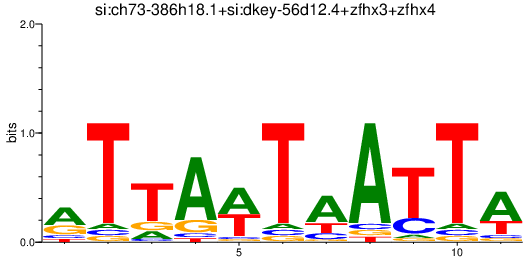
Results for si:ch73-386h18.1+si:dkey-56d12.4+zfhx3+zfhx4
Z-value: 0.45
Transcription factors associated with si:ch73-386h18.1+si:dkey-56d12.4+zfhx3+zfhx4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
si_dkey-56d12.4
|
ENSDARG00000070845 | si_dkey-56d12.4 |
|
si_ch73-386h18.1
|
ENSDARG00000073944 | si_ch73-386h18.1 |
|
zfhx4
|
ENSDARG00000075542 | zinc finger homeobox 4 |
|
zfhx3
|
ENSDARG00000103057 | zinc finger homeobox 3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| zfhx4 | dr10_dc_chr24_-_23174888_23174975 | 0.57 | 2.2e-02 | Click! |
Activity profile of si:ch73-386h18.1+si:dkey-56d12.4+zfhx3+zfhx4 motif
Sorted Z-values of si:ch73-386h18.1+si:dkey-56d12.4+zfhx3+zfhx4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of si:ch73-386h18.1+si:dkey-56d12.4+zfhx3+zfhx4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_+_28753020 | 1.16 |
ENSDART00000155815
|
nova2
|
neuro-oncological ventral antigen 2 |
| chr5_+_31745031 | 0.91 |
ENSDART00000147132
|
c9
|
complement component 9 |
| chr7_-_26035308 | 0.77 |
ENSDART00000131906
|
zgc:77439
|
zgc:77439 |
| chr4_+_9668755 | 0.71 |
ENSDART00000004604
|
si:dkey-153k10.9
|
si:dkey-153k10.9 |
| chr16_-_42990753 | 0.70 |
ENSDART00000149317
|
hfe2
|
hemochromatosis type 2 |
| chr23_+_21546553 | 0.69 |
ENSDART00000142921
|
si:ch73-21g5.7
|
si:ch73-21g5.7 |
| chr1_-_13547500 | 0.63 |
ENSDART00000044896
|
camk2d2
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II delta 2 |
| chr6_-_32362232 | 0.61 |
ENSDART00000140004
|
angptl3
|
angiopoietin-like 3 |
| chr20_-_9107294 | 0.60 |
ENSDART00000140792
|
OMA1
|
OMA1 zinc metallopeptidase |
| chr2_+_51087570 | 0.59 |
|
|
|
| chr25_+_24193604 | 0.58 |
ENSDART00000083407
|
b4galnt4a
|
beta-1,4-N-acetyl-galactosaminyl transferase 4a |
| chr11_+_23522743 | 0.54 |
ENSDART00000121874
|
nfasca
|
neurofascin homolog (chicken) a |
| chr1_-_25600988 | 0.53 |
ENSDART00000160381
|
cxxc4
|
CXXC finger 4 |
| chr22_+_20695983 | 0.50 |
ENSDART00000171321
|
si:dkey-211f22.5
|
si:dkey-211f22.5 |
| chr5_-_44229464 | 0.50 |
ENSDART00000019104
|
fbp2
|
fructose-1,6-bisphosphatase 2 |
| chr11_-_42096921 | 0.49 |
|
|
|
| chr4_-_5294280 | 0.48 |
ENSDART00000178921
|
SNAP91 (1 of many)
|
si:ch211-214j24.9 |
| chr8_+_15216833 | 0.48 |
ENSDART00000141185
|
slc5a9
|
solute carrier family 5 (sodium/sugar cotransporter), member 9 |
| chr19_+_14669633 | 0.47 |
ENSDART00000022076
|
fam46bb
|
family with sequence similarity 46, member Bb |
| chr23_+_24557956 | 0.45 |
|
|
|
| chr25_-_35094733 | 0.43 |
ENSDART00000153827
|
clpxb
|
caseinolytic mitochondrial matrix peptidase chaperone subunit b |
| chr16_+_28449134 | 0.42 |
ENSDART00000059044
|
itga8
|
integrin, alpha 8 |
| chr3_+_49277125 | 0.40 |
ENSDART00000161724
|
gas7a
|
growth arrest-specific 7a |
| chr2_+_21067708 | 0.39 |
ENSDART00000148400
ENSDART00000021168 |
rxrga
|
retinoid x receptor, gamma a |
| chr21_-_39132283 | 0.39 |
ENSDART00000065143
|
unc119b
|
unc-119 homolog b (C. elegans) |
| chr5_+_29578638 | 0.39 |
ENSDART00000134624
|
adamts15a
|
ADAM metallopeptidase with thrombospondin type 1 motif, 15a |
| chr9_+_21340251 | 0.37 |
ENSDART00000133903
|
hao2
|
hydroxyacid oxidase 2 (long chain) |
| chr24_-_38758093 | 0.36 |
|
|
|
| chr3_-_5318206 | 0.35 |
ENSDART00000137105
|
myh9b
|
myosin, heavy chain 9b, non-muscle |
| chr20_-_38714872 | 0.34 |
ENSDART00000050474
|
slc30a2
|
solute carrier family 30 (zinc transporter), member 2 |
| chr12_+_27035744 | 0.34 |
ENSDART00000025966
|
hoxb6b
|
homeobox B6b |
| chr20_-_25582221 | 0.34 |
ENSDART00000157559
|
si:dkey-183n20.15
|
si:dkey-183n20.15 |
| chr5_+_36332564 | 0.33 |
ENSDART00000037879
|
crx
|
cone-rod homeobox |
| chr12_-_19029801 | 0.33 |
ENSDART00000057124
|
tefa
|
thyrotrophic embryonic factor a |
| chr3_+_40028301 | 0.32 |
ENSDART00000011568
|
syngr3a
|
synaptogyrin 3a |
| chr24_-_2453987 | 0.31 |
ENSDART00000093331
|
rreb1a
|
ras responsive element binding protein 1a |
| chr16_-_9785057 | 0.30 |
ENSDART00000113724
|
mal2
|
mal, T-cell differentiation protein 2 (gene/pseudogene) |
| chr2_+_51087643 | 0.30 |
|
|
|
| chr21_-_20728623 | 0.29 |
ENSDART00000135940
ENSDART00000002231 |
ghrb
|
growth hormone receptor b |
| KN150001v1_+_14939 | 0.28 |
|
|
|
| chr23_-_20402258 | 0.28 |
ENSDART00000136204
|
CR749762.2
|
ENSDARG00000093223 |
| chr1_+_44137376 | 0.28 |
ENSDART00000083127
ENSDART00000162779 |
kdm2aa
|
lysine (K)-specific demethylase 2Aa |
| chr2_+_29265967 | 0.28 |
ENSDART00000099157
|
cdh18a
|
cadherin 18, type 2a |
| chr17_-_22047445 | 0.27 |
ENSDART00000156872
|
ttbk1b
|
tau tubulin kinase 1b |
| chr22_-_23641813 | 0.27 |
ENSDART00000159622
|
cfh
|
complement factor H |
| chr11_+_43136945 | 0.26 |
|
|
|
| chr24_-_21325777 | 0.25 |
ENSDART00000109848
|
atp8a2
|
ATPase, aminophospholipid transporter, class I, type 8A, member 2 |
| chr24_+_14569372 | 0.24 |
ENSDART00000134475
|
gdap1
|
ganglioside induced differentiation associated protein 1 |
| chr6_-_40061147 | 0.23 |
ENSDART00000103240
|
uroc1
|
urocanate hydratase 1 |
| chr8_+_18800414 | 0.23 |
ENSDART00000160732
|
mpnd
|
MPN domain containing |
| chr24_-_24306469 | 0.23 |
ENSDART00000154149
|
BX323067.1
|
ENSDARG00000097984 |
| chr22_+_13862110 | 0.20 |
ENSDART00000105711
|
sh3bp4a
|
SH3-domain binding protein 4a |
| chr9_+_2001849 | 0.19 |
ENSDART00000082329
|
evx2
|
even-skipped homeobox 2 |
| chr5_+_3721680 | 0.19 |
ENSDART00000140537
|
dhrs11a
|
dehydrogenase/reductase (SDR family) member 11a |
| chr3_-_27989221 | 0.19 |
ENSDART00000151143
|
rbfox1
|
RNA binding protein, fox-1 homolog (C. elegans) 1 |
| chr7_+_42124857 | 0.17 |
ENSDART00000004120
|
adamts18
|
ADAM metallopeptidase with thrombospondin type 1 motif, 18 |
| chr20_+_31173383 | 0.17 |
ENSDART00000136255
|
otofa
|
otoferlin a |
| chr18_+_1353857 | 0.16 |
ENSDART00000165301
|
rab27a
|
RAB27A, member RAS oncogene family |
| chr15_+_28753150 | 0.16 |
ENSDART00000060244
|
nova2
|
neuro-oncological ventral antigen 2 |
| chr14_-_24814149 | 0.16 |
ENSDART00000158108
|
si:rp71-1d10.8
|
si:rp71-1d10.8 |
| KN150001v1_+_15017 | 0.15 |
|
|
|
| chr4_+_14659537 | 0.15 |
|
|
|
| chr17_+_8642428 | 0.14 |
ENSDART00000105326
|
tonsl
|
tonsoku-like, DNA repair protein |
| chr12_-_36191048 | 0.13 |
ENSDART00000177986
|
cep131
|
centrosomal protein 131 |
| chr12_-_34621359 | 0.13 |
|
|
|
| chr23_-_19577146 | 0.13 |
ENSDART00000143288
|
asb14b
|
ankyrin repeat and SOCS box containing 14b |
| chr25_-_35094522 | 0.13 |
ENSDART00000153827
|
clpxb
|
caseinolytic mitochondrial matrix peptidase chaperone subunit b |
| chr23_+_44007440 | 0.12 |
ENSDART00000113757
|
ENSDARG00000076011
|
ENSDARG00000076011 |
| chr2_-_27953540 | 0.11 |
ENSDART00000040555
|
tgs1
|
trimethylguanosine synthase 1 |
| chr8_-_12253026 | 0.11 |
ENSDART00000091612
|
dab2ipa
|
DAB2 interacting protein a |
| chr3_-_27989425 | 0.11 |
ENSDART00000151143
|
rbfox1
|
RNA binding protein, fox-1 homolog (C. elegans) 1 |
| chr4_-_22617898 | 0.11 |
ENSDART00000131402
|
golgb1
|
golgin B1 |
| chr18_+_29971911 | 0.11 |
ENSDART00000146431
ENSDART00000099285 |
atmin
|
ATM interactor |
| chr1_-_32825274 | 0.10 |
ENSDART00000075632
|
creb1a
|
cAMP responsive element binding protein 1a |
| chr15_+_29842298 | 0.10 |
|
|
|
| chr9_+_31204464 | 0.10 |
|
|
|
| chr14_-_3836967 | 0.10 |
ENSDART00000055817
|
ENSDARG00000038270
|
ENSDARG00000038270 |
| chr24_+_26183653 | 0.10 |
ENSDART00000003884
|
mynn
|
myoneurin |
| chr11_-_26578973 | 0.09 |
|
|
|
| chr2_-_22173254 | 0.09 |
ENSDART00000137538
|
rab2a
|
RAB2A, member RAS oncogene family |
| chr25_+_34410545 | 0.08 |
ENSDART00000073441
|
sntb2
|
syntrophin, beta 2 |
| chr7_-_40734662 | 0.08 |
ENSDART00000013785
|
insig1
|
insulin induced gene 1 |
| chr21_+_13053889 | 0.08 |
ENSDART00000102256
|
spna2
|
spectrin alpha 2 |
| chr10_-_11303185 | 0.08 |
ENSDART00000146727
|
ptbp3
|
polypyrimidine tract binding protein 3 |
| chr1_+_53353287 | 0.08 |
ENSDART00000012104
|
dla
|
deltaA |
| chr17_+_9152727 | 0.07 |
|
|
|
| chr6_+_30469823 | 0.07 |
ENSDART00000121492
|
FP236735.1
|
ENSDARG00000087805 |
| chr19_-_41882385 | 0.06 |
|
|
|
| chr10_+_45056951 | 0.06 |
ENSDART00000158553
|
ENSDARG00000101143
|
ENSDARG00000101143 |
| chr10_-_8238422 | 0.05 |
ENSDART00000129467
|
dhx29
|
DEAH (Asp-Glu-Ala-His) box polypeptide 29 |
| chr17_+_8642373 | 0.04 |
ENSDART00000105326
|
tonsl
|
tonsoku-like, DNA repair protein |
| chr10_-_42302932 | 0.04 |
ENSDART00000076693
ENSDART00000073631 |
stambpa
|
STAM binding protein a |
| chr1_-_22160662 | 0.04 |
ENSDART00000054386
|
qdprb1
|
quinoid dihydropteridine reductase b1 |
| chr22_-_38493856 | 0.03 |
ENSDART00000168676
|
soul4
|
heme-binding protein soul4 |
| chr11_-_44738307 | 0.02 |
ENSDART00000167540
|
afmid
|
arylformamidase |
| chr13_+_45387478 | 0.02 |
ENSDART00000019113
|
tmem57b
|
transmembrane protein 57b |
| chr9_-_8035360 | 0.02 |
|
|
|
| chr1_-_9426640 | 0.01 |
|
|
|
| chr8_+_21974863 | 0.01 |
|
|
|
| chr16_+_24818799 | 0.01 |
ENSDART00000155217
|
si:dkey-79d12.4
|
si:dkey-79d12.4 |
| chr3_+_34513 | 0.01 |
|
|
|
| chr3_-_27989329 | 0.01 |
ENSDART00000151143
|
rbfox1
|
RNA binding protein, fox-1 homolog (C. elegans) 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0055091 | negative regulation of phospholipase activity(GO:0010519) phospholipid homeostasis(GO:0055091) |
| 0.2 | 0.9 | GO:0006958 | humoral immune response mediated by circulating immunoglobulin(GO:0002455) complement activation, classical pathway(GO:0006958) |
| 0.1 | 0.3 | GO:0061088 | sequestering of zinc ion(GO:0032119) regulation of sequestering of zinc ion(GO:0061088) |
| 0.1 | 0.2 | GO:0052805 | imidazole-containing compound metabolic process(GO:0052803) imidazole-containing compound catabolic process(GO:0052805) |
| 0.1 | 0.5 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.1 | 0.5 | GO:0019226 | transmission of nerve impulse(GO:0019226) |
| 0.1 | 0.3 | GO:0045056 | transcytosis(GO:0045056) |
| 0.0 | 0.2 | GO:0010996 | response to auditory stimulus(GO:0010996) |
| 0.0 | 0.3 | GO:0060416 | growth hormone receptor signaling pathway(GO:0060396) response to growth hormone(GO:0060416) cellular response to growth hormone stimulus(GO:0071378) |
| 0.0 | 0.2 | GO:0021521 | ventral spinal cord interneuron specification(GO:0021521) cell fate specification involved in pattern specification(GO:0060573) |
| 0.0 | 0.3 | GO:0070650 | actin filament bundle distribution(GO:0070650) |
| 0.0 | 0.1 | GO:0090317 | negative regulation of intracellular protein transport(GO:0090317) |
| 0.0 | 0.3 | GO:0007530 | sex determination(GO:0007530) |
| 0.0 | 0.4 | GO:0032526 | response to retinoic acid(GO:0032526) |
| 0.0 | 1.6 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.0 | 0.2 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.0 | 0.1 | GO:0055016 | hypochord development(GO:0055016) |
| 0.0 | 0.1 | GO:0030948 | negative regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030948) |
| 0.0 | 0.3 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.0 | 0.2 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.0 | 0.8 | GO:0071466 | cellular response to xenobiotic stimulus(GO:0071466) |
| 0.0 | 0.2 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.0 | 0.2 | GO:0061462 | protein localization to lysosome(GO:0061462) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.9 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.1 | 0.3 | GO:0070195 | growth hormone receptor complex(GO:0070195) |
| 0.0 | 0.2 | GO:0035101 | FACT complex(GO:0035101) |
| 0.0 | 0.2 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.0 | 0.3 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 0.1 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.0 | 0.2 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.0 | 0.5 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 0.4 | GO:0008305 | integrin complex(GO:0008305) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0033842 | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N-acetylgalactosaminyltransferase activity(GO:0033842) |
| 0.2 | 0.8 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.1 | 0.3 | GO:0004903 | growth hormone receptor activity(GO:0004903) |
| 0.1 | 0.4 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.1 | 0.5 | GO:0005355 | glucose transmembrane transporter activity(GO:0005355) monosaccharide transmembrane transporter activity(GO:0015145) hexose transmembrane transporter activity(GO:0015149) |
| 0.1 | 0.5 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.0 | 0.3 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.5 | GO:0050308 | carbohydrate phosphatase activity(GO:0019203) sugar-phosphatase activity(GO:0050308) |
| 0.0 | 0.6 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.0 | 0.4 | GO:0010181 | FMN binding(GO:0010181) |
| 0.0 | 0.2 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.0 | 0.1 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.0 | 0.3 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.2 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.0 | 0.3 | GO:0070122 | isopeptidase activity(GO:0070122) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.0 | 0.7 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 0.5 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.0 | 0.6 | PID AVB3 INTEGRIN PATHWAY | Integrins in angiogenesis |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.0 | 0.7 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.3 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 0.5 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 0.2 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.2 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |