Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
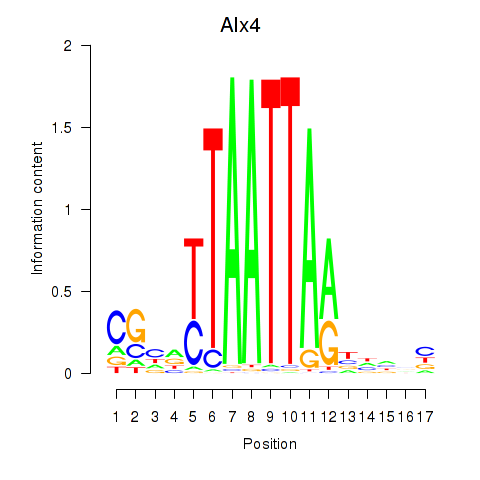
Results for Alx4
Z-value: 1.68
Transcription factors associated with Alx4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Alx4
|
ENSMUSG00000040310.13 | Alx4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Alx4 | mm39_v1_chr2_+_93472657_93472733 | 0.60 | 1.0e-04 | Click! |
Activity profile of Alx4 motif
Sorted Z-values of Alx4 motif
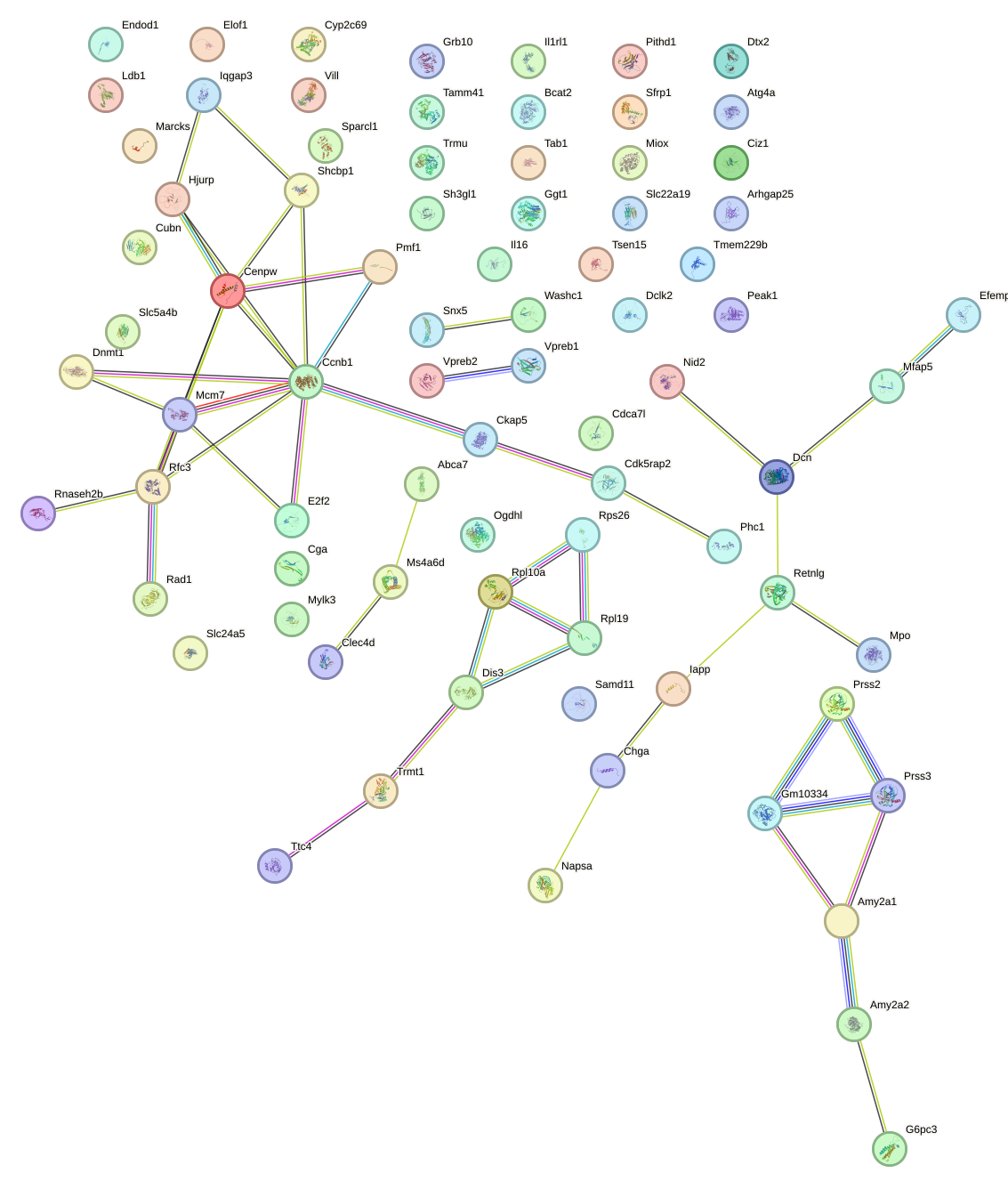
Network of associatons between targets according to the STRING database.
First level regulatory network of Alx4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_+_87684299 | 16.19 |
ENSMUST00000020779.11
|
Mpo
|
myeloperoxidase |
| chr5_-_138169253 | 9.54 |
ENSMUST00000139983.8
|
Mcm7
|
minichromosome maintenance complex component 7 |
| chr13_-_100922910 | 9.31 |
ENSMUST00000174038.2
ENSMUST00000091295.14 ENSMUST00000072119.15 |
Ccnb1
|
cyclin B1 |
| chr7_+_44221791 | 7.52 |
ENSMUST00000002274.10
|
Napsa
|
napsin A aspartic peptidase |
| chr15_+_89218601 | 7.22 |
ENSMUST00000023282.9
|
Miox
|
myo-inositol oxygenase |
| chr11_-_11920540 | 6.66 |
ENSMUST00000109653.8
|
Grb10
|
growth factor receptor bound protein 10 |
| chr6_-_41354538 | 6.60 |
ENSMUST00000096003.7
|
Prss3
|
protease, serine 3 |
| chr5_-_138169509 | 6.57 |
ENSMUST00000153867.8
|
Mcm7
|
minichromosome maintenance complex component 7 |
| chr16_+_48692976 | 6.57 |
ENSMUST00000065666.6
|
Retnlg
|
resistin like gamma |
| chr4_+_135899678 | 6.47 |
ENSMUST00000061721.6
|
E2f2
|
E2F transcription factor 2 |
| chr10_-_62215631 | 6.40 |
ENSMUST00000143236.8
ENSMUST00000133429.8 ENSMUST00000132926.8 ENSMUST00000116238.9 |
Hk1
|
hexokinase 1 |
| chr11_-_106205320 | 6.24 |
ENSMUST00000167143.2
|
Cd79b
|
CD79B antigen |
| chr16_-_16687119 | 5.87 |
ENSMUST00000075017.5
|
Vpreb1
|
pre-B lymphocyte gene 1 |
| chr9_-_20871081 | 5.65 |
ENSMUST00000177754.9
|
Dnmt1
|
DNA methyltransferase (cytosine-5) 1 |
| chr7_-_142209755 | 5.53 |
ENSMUST00000178921.2
|
Igf2
|
insulin-like growth factor 2 |
| chr19_-_46033353 | 5.00 |
ENSMUST00000026252.14
ENSMUST00000156585.9 ENSMUST00000185355.7 ENSMUST00000152946.8 |
Ldb1
|
LIM domain binding 1 |
| chr6_+_41498716 | 4.96 |
ENSMUST00000070380.5
|
Prss2
|
protease, serine 2 |
| chr2_-_131001916 | 4.66 |
ENSMUST00000103188.10
ENSMUST00000133602.8 ENSMUST00000028800.12 |
1700037H04Rik
|
RIKEN cDNA 1700037H04 gene |
| chr3_+_87989278 | 4.28 |
ENSMUST00000071812.11
|
Iqgap3
|
IQ motif containing GTPase activating protein 3 |
| chr6_-_41423004 | 4.27 |
ENSMUST00000095999.7
|
Gm10334
|
predicted gene 10334 |
| chr9_-_14292453 | 4.20 |
ENSMUST00000167549.2
|
Endod1
|
endonuclease domain containing 1 |
| chr6_-_115014777 | 3.89 |
ENSMUST00000174848.8
ENSMUST00000032461.12 |
Tamm41
|
TAM41 mitochondrial translocator assembly and maintenance homolog |
| chr10_+_75399920 | 3.86 |
ENSMUST00000141062.8
ENSMUST00000152657.8 |
Ggt1
|
gamma-glutamyltransferase 1 |
| chr7_-_24705320 | 3.83 |
ENSMUST00000102858.10
ENSMUST00000196684.2 ENSMUST00000080882.11 |
Atp1a3
|
ATPase, Na+/K+ transporting, alpha 3 polypeptide |
| chr8_-_86091946 | 3.50 |
ENSMUST00000034133.14
|
Mylk3
|
myosin light chain kinase 3 |
| chr4_-_131802561 | 3.49 |
ENSMUST00000105970.8
ENSMUST00000105975.8 |
Epb41
|
erythrocyte membrane protein band 4.1 |
| chr8_-_4829519 | 3.49 |
ENSMUST00000022945.9
|
Shcbp1
|
Shc SH2-domain binding protein 1 |
| chr5_-_138169476 | 3.38 |
ENSMUST00000147920.2
|
Mcm7
|
minichromosome maintenance complex component 7 |
| chr5_-_44139099 | 3.32 |
ENSMUST00000061299.9
|
Fgfbp1
|
fibroblast growth factor binding protein 1 |
| chr9_+_118892497 | 3.30 |
ENSMUST00000141185.8
ENSMUST00000126251.8 ENSMUST00000136561.2 |
Vill
|
villin-like |
| chr4_-_156340713 | 3.20 |
ENSMUST00000219393.2
|
Samd11
|
sterile alpha motif domain containing 11 |
| chr12_-_75678092 | 3.11 |
ENSMUST00000238938.2
|
Rplp2-ps1
|
ribosomal protein, large P2, pseudogene 1 |
| chr8_+_23901506 | 3.07 |
ENSMUST00000033952.8
|
Sfrp1
|
secreted frizzled-related protein 1 |
| chr3_-_113325938 | 2.88 |
ENSMUST00000132353.2
|
Amy2a1
|
amylase 2a1 |
| chr4_-_131802606 | 2.87 |
ENSMUST00000146021.8
|
Epb41
|
erythrocyte membrane protein band 4.1 |
| chr11_-_58059293 | 2.85 |
ENSMUST00000172035.8
ENSMUST00000035604.13 ENSMUST00000102711.9 |
Gemin5
|
gem nuclear organelle associated protein 5 |
| chr6_-_68609426 | 2.76 |
ENSMUST00000103328.3
|
Igkv10-96
|
immunoglobulin kappa variable 10-96 |
| chr2_+_124910037 | 2.73 |
ENSMUST00000070353.4
|
Slc24a5
|
solute carrier family 24, member 5 |
| chr1_+_40478787 | 2.72 |
ENSMUST00000097772.10
|
Il1rl1
|
interleukin 1 receptor-like 1 |
| chr2_+_91376650 | 2.70 |
ENSMUST00000099716.11
ENSMUST00000046769.16 ENSMUST00000111337.3 |
Ckap5
|
cytoskeleton associated protein 5 |
| chr12_-_79054050 | 2.67 |
ENSMUST00000056660.13
ENSMUST00000174721.8 |
Tmem229b
|
transmembrane protein 229B |
| chr5_-_44139121 | 2.64 |
ENSMUST00000199894.2
|
Fgfbp1
|
fibroblast growth factor binding protein 1 |
| chr10_-_128462616 | 2.63 |
ENSMUST00000026420.7
|
Rps26
|
ribosomal protein S26 |
| chr7_-_83304698 | 2.61 |
ENSMUST00000145610.8
|
Il16
|
interleukin 16 |
| chr6_+_123239076 | 2.55 |
ENSMUST00000032240.4
|
Clec4d
|
C-type lectin domain family 4, member d |
| chr12_+_117807224 | 2.54 |
ENSMUST00000021592.16
|
Cdca7l
|
cell division cycle associated 7 like |
| chr19_+_5524701 | 2.53 |
ENSMUST00000165485.8
ENSMUST00000166253.8 ENSMUST00000167371.8 ENSMUST00000167855.8 ENSMUST00000070118.14 |
Efemp2
|
epidermal growth factor-containing fibulin-like extracellular matrix protein 2 |
| chr9_+_21437440 | 2.50 |
ENSMUST00000086361.12
ENSMUST00000173769.3 |
AB124611
|
cDNA sequence AB124611 |
| chr7_-_142253247 | 2.40 |
ENSMUST00000105934.8
|
Ins2
|
insulin II |
| chr11_+_58839716 | 2.39 |
ENSMUST00000078267.5
|
H2bu2
|
H2B.U histone 2 |
| chr14_+_62529924 | 2.32 |
ENSMUST00000166879.8
|
Rnaseh2b
|
ribonuclease H2, subunit B |
| chr5_-_114911548 | 2.27 |
ENSMUST00000178440.8
ENSMUST00000043283.14 ENSMUST00000112185.9 ENSMUST00000155908.8 |
Git2
|
GIT ArfGAP 2 |
| chr11_+_97917520 | 2.27 |
ENSMUST00000092425.11
|
Rpl19
|
ribosomal protein L19 |
| chrX_+_139857640 | 2.25 |
ENSMUST00000112971.2
|
Atg4a
|
autophagy related 4A, cysteine peptidase |
| chr11_+_97917746 | 2.24 |
ENSMUST00000017548.7
|
Rpl19
|
ribosomal protein L19 |
| chr8_+_85415935 | 2.24 |
ENSMUST00000125370.10
ENSMUST00000175784.2 |
Trmt1
|
tRNA methyltransferase 1 |
| chr10_-_30076543 | 2.20 |
ENSMUST00000099985.6
|
Cenpw
|
centromere protein W |
| chr5_+_136023649 | 2.19 |
ENSMUST00000111142.9
ENSMUST00000111145.10 ENSMUST00000111144.8 ENSMUST00000199239.5 ENSMUST00000005072.10 ENSMUST00000130345.2 |
Dtx2
|
deltex 2, E3 ubiquitin ligase |
| chr17_+_28547445 | 2.14 |
ENSMUST00000042334.16
|
Rpl10a
|
ribosomal protein L10A |
| chr6_+_142244145 | 2.12 |
ENSMUST00000041993.3
|
Iapp
|
islet amyloid polypeptide |
| chr4_-_136329953 | 2.11 |
ENSMUST00000105847.8
ENSMUST00000116273.9 |
Kdm1a
|
lysine (K)-specific demethylase 1A |
| chr4_+_34893772 | 2.03 |
ENSMUST00000029975.10
ENSMUST00000135871.8 ENSMUST00000108130.2 |
Cga
|
glycoprotein hormones, alpha subunit |
| chr1_+_160898283 | 2.03 |
ENSMUST00000028035.14
ENSMUST00000111620.10 ENSMUST00000111618.8 |
Cenpl
|
centromere protein L |
| chr5_-_151574620 | 1.97 |
ENSMUST00000038131.10
|
Rfc3
|
replication factor C (activator 1) 3 |
| chr10_-_37014859 | 1.96 |
ENSMUST00000092584.6
|
Marcks
|
myristoylated alanine rich protein kinase C substrate |
| chr19_-_39875192 | 1.93 |
ENSMUST00000168838.3
|
Cyp2c69
|
cytochrome P450, family 2, subfamily c, polypeptide 69 |
| chr3_-_88317601 | 1.92 |
ENSMUST00000193338.6
ENSMUST00000056370.13 |
Pmf1
|
polyamine-modulated factor 1 |
| chrX_+_139857688 | 1.91 |
ENSMUST00000239541.1
|
Atg4a
|
autophagy related 4A, cysteine peptidase |
| chr1_-_152262425 | 1.85 |
ENSMUST00000015124.15
|
Tsen15
|
tRNA splicing endonuclease subunit 15 |
| chr8_+_85415876 | 1.81 |
ENSMUST00000109767.9
ENSMUST00000177084.8 ENSMUST00000109768.9 ENSMUST00000152301.9 ENSMUST00000177423.8 |
Trmt1
|
tRNA methyltransferase 1 |
| chr1_+_40478926 | 1.80 |
ENSMUST00000173514.8
|
Il1rl1
|
interleukin 1 receptor-like 1 |
| chr3_-_86827640 | 1.79 |
ENSMUST00000195561.6
|
Dclk2
|
doublecortin-like kinase 2 |
| chr15_+_80017315 | 1.75 |
ENSMUST00000023050.9
|
Tab1
|
TGF-beta activated kinase 1/MAP3K7 binding protein 1 |
| chr19_-_11582207 | 1.75 |
ENSMUST00000025582.11
|
Ms4a6d
|
membrane-spanning 4-domains, subfamily A, member 6D |
| chr7_+_89814713 | 1.71 |
ENSMUST00000207084.2
|
Picalm
|
phosphatidylinositol binding clathrin assembly protein |
| chr14_-_99337137 | 1.71 |
ENSMUST00000042471.11
|
Dis3
|
DIS3 homolog, exosome endoribonuclease and 3'-5' exoribonuclease |
| chr10_+_79832313 | 1.69 |
ENSMUST00000132517.8
|
Abca7
|
ATP-binding cassette, sub-family A (ABC1), member 7 |
| chr3_-_86827664 | 1.68 |
ENSMUST00000194452.2
ENSMUST00000191752.6 |
Dclk2
|
doublecortin-like kinase 2 |
| chr9_-_22028370 | 1.66 |
ENSMUST00000213233.2
|
Elof1
|
ELF1 homolog, elongation factor 1 |
| chr2_-_85966272 | 1.66 |
ENSMUST00000216566.3
ENSMUST00000214364.2 |
Olfr1039
|
olfactory receptor 1039 |
| chr17_-_56343531 | 1.63 |
ENSMUST00000233803.2
|
Sh3gl1
|
SH3-domain GRB2-like 1 |
| chr10_+_97315465 | 1.62 |
ENSMUST00000105287.11
|
Dcn
|
decorin |
| chr19_+_12647803 | 1.62 |
ENSMUST00000207341.3
ENSMUST00000208494.3 ENSMUST00000208657.3 |
Olfr1442
|
olfactory receptor 1442 |
| chr2_+_32253016 | 1.61 |
ENSMUST00000132028.8
ENSMUST00000136079.8 |
Ciz1
|
CDKN1A interacting zinc finger protein 1 |
| chr2_-_144112700 | 1.59 |
ENSMUST00000110030.10
|
Snx5
|
sorting nexin 5 |
| chr17_-_56343625 | 1.58 |
ENSMUST00000003268.11
|
Sh3gl1
|
SH3-domain GRB2-like 1 |
| chr17_+_66418525 | 1.58 |
ENSMUST00000072383.14
|
Washc1
|
WASH complex subunit 1 |
| chr7_+_45219766 | 1.57 |
ENSMUST00000120864.10
|
Bcat2
|
branched chain aminotransferase 2, mitochondrial |
| chr5_-_104261285 | 1.54 |
ENSMUST00000199947.2
|
Sparcl1
|
SPARC-like 1 |
| chr4_-_70328659 | 1.53 |
ENSMUST00000144099.8
|
Cdk5rap2
|
CDK5 regulatory subunit associated protein 2 |
| chr6_-_122317484 | 1.52 |
ENSMUST00000112600.9
|
Phc1
|
polyhomeotic 1 |
| chr6_-_87510200 | 1.50 |
ENSMUST00000113637.9
ENSMUST00000071024.7 |
Arhgap25
|
Rho GTPase activating protein 25 |
| chr9_-_56151334 | 1.46 |
ENSMUST00000188142.7
|
Peak1
|
pseudopodium-enriched atypical kinase 1 |
| chr4_-_135714465 | 1.44 |
ENSMUST00000105851.9
|
Pithd1
|
PITH (C-terminal proteasome-interacting domain of thioredoxin-like) domain containing 1 |
| chr14_+_19801333 | 1.44 |
ENSMUST00000022340.5
|
Nid2
|
nidogen 2 |
| chr14_-_104760051 | 1.43 |
ENSMUST00000022716.4
ENSMUST00000228448.2 ENSMUST00000227640.2 |
Obi1
|
ORC ubiquitin ligase 1 |
| chr1_-_88205233 | 1.43 |
ENSMUST00000065420.12
ENSMUST00000054674.15 |
Hjurp
|
Holliday junction recognition protein |
| chr2_+_32253692 | 1.41 |
ENSMUST00000113331.8
ENSMUST00000113338.9 |
Ciz1
|
CDKN1A interacting zinc finger protein 1 |
| chr4_-_106536063 | 1.40 |
ENSMUST00000106772.10
ENSMUST00000135676.2 ENSMUST00000026480.13 |
Ttc4
|
tetratricopeptide repeat domain 4 |
| chr5_-_114911509 | 1.39 |
ENSMUST00000086564.11
|
Git2
|
GIT ArfGAP 2 |
| chr9_-_22028419 | 1.39 |
ENSMUST00000214394.2
ENSMUST00000013966.8 |
Elof1
|
ELF1 homolog, elongation factor 1 |
| chr19_-_7688628 | 1.36 |
ENSMUST00000025666.8
|
Slc22a19
|
solute carrier family 22 (organic anion transporter), member 19 |
| chr14_+_32043944 | 1.35 |
ENSMUST00000022480.8
ENSMUST00000228529.2 |
Ogdhl
|
oxoglutarate dehydrogenase-like |
| chr1_-_152262339 | 1.34 |
ENSMUST00000162371.2
|
Tsen15
|
tRNA splicing endonuclease subunit 15 |
| chr10_-_75946790 | 1.34 |
ENSMUST00000120757.2
|
Slc5a4b
|
solute carrier family 5 (neutral amino acid transporters, system A), member 4b |
| chr10_-_129107354 | 1.33 |
ENSMUST00000204573.3
|
Olfr777
|
olfactory receptor 777 |
| chr6_-_57827328 | 1.32 |
ENSMUST00000203310.3
ENSMUST00000203488.3 |
Vmn1r21
|
vomeronasal 1 receptor 21 |
| chr14_+_51366512 | 1.32 |
ENSMUST00000095923.4
|
Rnase6
|
ribonuclease, RNase A family, 6 |
| chr2_-_13496624 | 1.30 |
ENSMUST00000091436.7
|
Cubn
|
cubilin (intrinsic factor-cobalamin receptor) |
| chr2_+_32253204 | 1.28 |
ENSMUST00000048964.14
ENSMUST00000113332.8 |
Ciz1
|
CDKN1A interacting zinc finger protein 1 |
| chr6_+_122490534 | 1.27 |
ENSMUST00000032210.14
ENSMUST00000148517.8 |
Mfap5
|
microfibrillar associated protein 5 |
| chr6_-_69282389 | 1.26 |
ENSMUST00000103350.3
|
Igkv4-68
|
immunoglobulin kappa variable 4-68 |
| chr11_+_102080446 | 1.25 |
ENSMUST00000070334.10
|
G6pc3
|
glucose 6 phosphatase, catalytic, 3 |
| chr13_+_21995906 | 1.24 |
ENSMUST00000104941.4
|
H4c17
|
H4 clustered histone 17 |
| chr3_-_72875187 | 1.22 |
ENSMUST00000167334.8
|
Sis
|
sucrase isomaltase (alpha-glucosidase) |
| chr9_+_19828161 | 1.22 |
ENSMUST00000217347.2
ENSMUST00000057596.10 |
Olfr77
|
olfactory receptor 77 |
| chr5_-_114911432 | 1.20 |
ENSMUST00000112183.8
|
Git2
|
GIT ArfGAP 2 |
| chr11_+_102080489 | 1.20 |
ENSMUST00000078975.8
|
G6pc3
|
glucose 6 phosphatase, catalytic, 3 |
| chr14_+_119025306 | 1.20 |
ENSMUST00000047761.13
ENSMUST00000071546.14 |
Cldn10
|
claudin 10 |
| chr19_+_34078333 | 1.19 |
ENSMUST00000025685.8
|
Lipm
|
lipase, family member M |
| chr17_+_66418598 | 1.18 |
ENSMUST00000116556.4
ENSMUST00000233354.2 |
Washc1
|
WASH complex subunit 1 |
| chr9_+_65368207 | 1.15 |
ENSMUST00000034955.8
ENSMUST00000213957.2 |
Spg21
|
SPG21, maspardin |
| chr3_-_105839980 | 1.15 |
ENSMUST00000098758.5
|
I830077J02Rik
|
RIKEN cDNA I830077J02 gene |
| chr12_-_69939762 | 1.14 |
ENSMUST00000110567.8
ENSMUST00000171211.8 |
Map4k5
|
mitogen-activated protein kinase kinase kinase kinase 5 |
| chr5_-_116162415 | 1.13 |
ENSMUST00000031486.14
ENSMUST00000111999.8 |
Prkab1
|
protein kinase, AMP-activated, beta 1 non-catalytic subunit |
| chr4_+_19818718 | 1.13 |
ENSMUST00000035890.8
|
Slc7a13
|
solute carrier family 7, (cationic amino acid transporter, y+ system) member 13 |
| chr2_-_73284262 | 1.12 |
ENSMUST00000102679.8
|
Wipf1
|
WAS/WASL interacting protein family, member 1 |
| chr7_+_128346655 | 1.12 |
ENSMUST00000042942.10
|
Sec23ip
|
Sec23 interacting protein |
| chr2_+_22785534 | 1.11 |
ENSMUST00000053729.14
|
Pdss1
|
prenyl (solanesyl) diphosphate synthase, subunit 1 |
| chr2_-_29677634 | 1.08 |
ENSMUST00000177467.8
ENSMUST00000113807.10 |
Trub2
|
TruB pseudouridine (psi) synthase family member 2 |
| chr7_+_101545547 | 1.06 |
ENSMUST00000035395.14
ENSMUST00000106973.8 ENSMUST00000144207.9 |
Anapc15
|
anaphase promoting complex C subunit 15 |
| chr7_+_126376099 | 1.06 |
ENSMUST00000038614.12
ENSMUST00000170882.8 ENSMUST00000106359.2 ENSMUST00000106357.8 ENSMUST00000145762.8 |
Ypel3
|
yippee like 3 |
| chr6_-_87312743 | 1.06 |
ENSMUST00000042025.12
ENSMUST00000205033.2 |
Antxr1
|
anthrax toxin receptor 1 |
| chr12_-_113790741 | 1.06 |
ENSMUST00000103457.3
ENSMUST00000192877.2 |
Ighv5-15
|
immunoglobulin heavy variable 5-15 |
| chr11_-_51497665 | 1.05 |
ENSMUST00000074669.10
ENSMUST00000101249.9 ENSMUST00000109103.4 |
Hnrnpab
|
heterogeneous nuclear ribonucleoprotein A/B |
| chr15_-_77840856 | 1.03 |
ENSMUST00000117725.2
ENSMUST00000016696.13 |
Foxred2
|
FAD-dependent oxidoreductase domain containing 2 |
| chr19_-_32173824 | 1.02 |
ENSMUST00000151822.2
|
Sgms1
|
sphingomyelin synthase 1 |
| chr19_-_24178000 | 1.02 |
ENSMUST00000233658.3
|
Tjp2
|
tight junction protein 2 |
| chr2_+_85838122 | 1.01 |
ENSMUST00000062166.2
|
Olfr1032
|
olfactory receptor 1032 |
| chr12_-_80690573 | 1.01 |
ENSMUST00000166931.2
ENSMUST00000218364.2 |
Erh
|
ERH mRNA splicing and mitosis factor |
| chr10_+_85858050 | 0.99 |
ENSMUST00000120344.8
ENSMUST00000117597.2 |
Fbxo7
|
F-box protein 7 |
| chr8_+_31640332 | 0.98 |
ENSMUST00000209851.2
ENSMUST00000098842.3 ENSMUST00000210129.2 |
Tti2
|
TELO2 interacting protein 2 |
| chr13_+_51562675 | 0.98 |
ENSMUST00000087978.5
|
S1pr3
|
sphingosine-1-phosphate receptor 3 |
| chr7_+_45271229 | 0.97 |
ENSMUST00000033100.5
|
Izumo1
|
izumo sperm-egg fusion 1 |
| chr2_-_88590813 | 0.96 |
ENSMUST00000124021.3
|
Olfr1199
|
olfactory receptor 1199 |
| chr2_+_24257576 | 0.96 |
ENSMUST00000140547.2
ENSMUST00000102942.8 |
Psd4
|
pleckstrin and Sec7 domain containing 4 |
| chr7_-_30259025 | 0.95 |
ENSMUST00000043975.11
ENSMUST00000156241.2 |
Lin37
|
lin-37 homolog (C. elegans) |
| chr1_-_88205185 | 0.95 |
ENSMUST00000147393.2
|
Hjurp
|
Holliday junction recognition protein |
| chr15_-_82678490 | 0.95 |
ENSMUST00000006094.6
|
Cyp2d26
|
cytochrome P450, family 2, subfamily d, polypeptide 26 |
| chr8_+_107757847 | 0.94 |
ENSMUST00000034388.10
|
Vps4a
|
vacuolar protein sorting 4A |
| chr5_+_136145485 | 0.93 |
ENSMUST00000111127.8
ENSMUST00000041366.14 ENSMUST00000111129.2 |
Polr2j
|
polymerase (RNA) II (DNA directed) polypeptide J |
| chr14_-_69945022 | 0.93 |
ENSMUST00000118374.8
|
R3hcc1
|
R3H domain and coiled-coil containing 1 |
| chrX_+_20529137 | 0.93 |
ENSMUST00000001989.9
|
Uba1
|
ubiquitin-like modifier activating enzyme 1 |
| chr17_-_57338468 | 0.93 |
ENSMUST00000007814.10
ENSMUST00000233480.2 |
Khsrp
|
KH-type splicing regulatory protein |
| chr15_-_79658608 | 0.92 |
ENSMUST00000229644.2
ENSMUST00000023055.8 |
Dnal4
|
dynein, axonemal, light chain 4 |
| chr14_-_54923517 | 0.92 |
ENSMUST00000125265.2
|
Acin1
|
apoptotic chromatin condensation inducer 1 |
| chr4_-_43710231 | 0.92 |
ENSMUST00000217544.2
ENSMUST00000107862.3 |
Olfr71
|
olfactory receptor 71 |
| chr3_-_66204228 | 0.91 |
ENSMUST00000029419.8
|
Veph1
|
ventricular zone expressed PH domain-containing 1 |
| chr1_-_127605660 | 0.91 |
ENSMUST00000160616.8
|
Tmem163
|
transmembrane protein 163 |
| chr13_+_76727787 | 0.91 |
ENSMUST00000126960.8
ENSMUST00000109583.9 |
Mctp1
|
multiple C2 domains, transmembrane 1 |
| chr12_-_114710326 | 0.89 |
ENSMUST00000103507.2
|
Ighv1-22
|
immunoglobulin heavy variable 1-22 |
| chr1_+_135768595 | 0.89 |
ENSMUST00000112087.9
ENSMUST00000178854.8 ENSMUST00000027671.12 ENSMUST00000179863.8 ENSMUST00000112085.9 ENSMUST00000112086.3 |
Tnnt2
|
troponin T2, cardiac |
| chrX_-_105264751 | 0.87 |
ENSMUST00000113495.9
|
Taf9b
|
TATA-box binding protein associated factor 9B |
| chr6_-_118456198 | 0.87 |
ENSMUST00000161170.2
|
Zfp9
|
zinc finger protein 9 |
| chr12_-_46863726 | 0.85 |
ENSMUST00000219330.2
|
Nova1
|
NOVA alternative splicing regulator 1 |
| chr14_-_69944942 | 0.85 |
ENSMUST00000121142.4
|
R3hcc1
|
R3H domain and coiled-coil containing 1 |
| chr11_+_62442502 | 0.84 |
ENSMUST00000136938.2
|
Ubb
|
ubiquitin B |
| chr7_-_110673269 | 0.82 |
ENSMUST00000163014.2
|
Eif4g2
|
eukaryotic translation initiation factor 4, gamma 2 |
| chr2_-_126342551 | 0.82 |
ENSMUST00000129187.2
|
Atp8b4
|
ATPase, class I, type 8B, member 4 |
| chr6_-_30936013 | 0.81 |
ENSMUST00000101589.5
|
Klf14
|
Kruppel-like factor 14 |
| chr2_+_65451100 | 0.78 |
ENSMUST00000144254.6
ENSMUST00000028377.14 |
Scn2a
|
sodium channel, voltage-gated, type II, alpha |
| chr9_-_123507937 | 0.78 |
ENSMUST00000040960.13
|
Slc6a20a
|
solute carrier family 6 (neurotransmitter transporter), member 20A |
| chr3_+_32762656 | 0.78 |
ENSMUST00000029214.14
|
Actl6a
|
actin-like 6A |
| chr8_+_22224506 | 0.78 |
ENSMUST00000080533.6
|
Defa24
|
defensin, alpha, 24 |
| chr7_-_30259253 | 0.78 |
ENSMUST00000108164.8
|
Lin37
|
lin-37 homolog (C. elegans) |
| chr9_+_123902143 | 0.78 |
ENSMUST00000168841.3
ENSMUST00000055918.7 |
Ccr2
|
chemokine (C-C motif) receptor 2 |
| chr8_-_70963202 | 0.77 |
ENSMUST00000125184.8
|
Uba52
|
ubiquitin A-52 residue ribosomal protein fusion product 1 |
| chr11_-_99134885 | 0.77 |
ENSMUST00000103132.10
ENSMUST00000038214.7 |
Krt222
|
keratin 222 |
| chr9_-_89586960 | 0.77 |
ENSMUST00000058488.9
|
Tmed3
|
transmembrane p24 trafficking protein 3 |
| chr5_-_118293443 | 0.77 |
ENSMUST00000049474.9
|
Fbxw8
|
F-box and WD-40 domain protein 8 |
| chr6_+_68279392 | 0.77 |
ENSMUST00000103322.3
|
Igkv2-109
|
immunoglobulin kappa variable 2-109 |
| chr12_+_38833454 | 0.75 |
ENSMUST00000161980.8
ENSMUST00000160701.8 |
Etv1
|
ets variant 1 |
| chrX_-_9123138 | 0.75 |
ENSMUST00000115553.3
|
Gm14862
|
predicted gene 14862 |
| chr2_-_88519531 | 0.74 |
ENSMUST00000213545.2
|
Olfr1195
|
olfactory receptor 1195 |
| chr13_-_103911092 | 0.74 |
ENSMUST00000074616.7
|
Srek1
|
splicing regulatory glutamine/lysine-rich protein 1 |
| chr2_+_87696836 | 0.72 |
ENSMUST00000213308.3
|
Olfr1152
|
olfactory receptor 1152 |
| chr2_-_111843053 | 0.72 |
ENSMUST00000213559.3
|
Olfr1310
|
olfactory receptor 1310 |
| chr18_+_37898633 | 0.72 |
ENSMUST00000044851.8
|
Pcdhga12
|
protocadherin gamma subfamily A, 12 |
| chr7_+_101546059 | 0.71 |
ENSMUST00000143835.8
|
Anapc15
|
anaphase promoting complex C subunit 15 |
| chr5_+_3593811 | 0.71 |
ENSMUST00000197082.5
ENSMUST00000115527.8 |
Fam133b
|
family with sequence similarity 133, member B |
| chr6_-_68887957 | 0.71 |
ENSMUST00000200454.2
|
Igkv4-86
|
immunoglobulin kappa variable 4-86 |
| chr11_+_106107752 | 0.70 |
ENSMUST00000021046.6
|
Ddx42
|
DEAD box helicase 42 |
| chr9_+_98873831 | 0.69 |
ENSMUST00000185472.2
|
Faim
|
Fas apoptotic inhibitory molecule |
| chr15_-_79658584 | 0.69 |
ENSMUST00000069877.12
|
Dnal4
|
dynein, axonemal, light chain 4 |
| chr2_-_86109346 | 0.67 |
ENSMUST00000217294.2
ENSMUST00000217245.2 ENSMUST00000216432.2 |
Olfr1051
|
olfactory receptor 1051 |
| chr4_-_147894245 | 0.67 |
ENSMUST00000105734.10
ENSMUST00000176201.2 |
Zfp984
Gm20707
|
zinc finger protein 984 predicted gene 20707 |
| chr8_-_70962972 | 0.66 |
ENSMUST00000140679.8
ENSMUST00000129909.8 ENSMUST00000081940.11 |
Uba52
|
ubiquitin A-52 residue ribosomal protein fusion product 1 |
| chr1_+_134333720 | 0.66 |
ENSMUST00000173908.8
|
Cyb5r1
|
cytochrome b5 reductase 1 |
| chr3_+_90200470 | 0.66 |
ENSMUST00000199754.5
|
Gatad2b
|
GATA zinc finger domain containing 2B |
| chr7_-_103094646 | 0.66 |
ENSMUST00000215417.2
|
Olfr605
|
olfactory receptor 605 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.4 | 16.2 | GO:0002149 | hypochlorous acid metabolic process(GO:0002148) hypochlorous acid biosynthetic process(GO:0002149) |
| 2.3 | 9.3 | GO:0031660 | regulation of cyclin-dependent protein serine/threonine kinase activity involved in G2/M transition of mitotic cell cycle(GO:0031660) positive regulation of cyclin-dependent protein serine/threonine kinase activity involved in G2/M transition of mitotic cell cycle(GO:0031662) |
| 1.7 | 5.0 | GO:0043973 | histone H3-K4 acetylation(GO:0043973) |
| 1.6 | 6.6 | GO:0002408 | myeloid dendritic cell chemotaxis(GO:0002408) |
| 1.6 | 19.5 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 1.4 | 5.6 | GO:0090116 | C-5 methylation of cytosine(GO:0090116) positive regulation of methylation-dependent chromatin silencing(GO:0090309) |
| 1.3 | 4.0 | GO:0002940 | tRNA N2-guanine methylation(GO:0002940) |
| 1.1 | 4.3 | GO:0032298 | positive regulation of DNA-dependent DNA replication initiation(GO:0032298) |
| 1.0 | 3.1 | GO:2000040 | regulation of midbrain dopaminergic neuron differentiation(GO:1904956) regulation of planar cell polarity pathway involved in axis elongation(GO:2000040) negative regulation of planar cell polarity pathway involved in axis elongation(GO:2000041) |
| 0.8 | 6.5 | GO:1990086 | lens fiber cell apoptotic process(GO:1990086) |
| 0.8 | 6.4 | GO:0061621 | NADH regeneration(GO:0006735) canonical glycolysis(GO:0061621) glucose catabolic process to pyruvate(GO:0061718) |
| 0.8 | 2.4 | GO:0033861 | negative regulation of NAD(P)H oxidase activity(GO:0033861) neuron projection maintenance(GO:1990535) |
| 0.7 | 2.7 | GO:0048022 | negative regulation of melanin biosynthetic process(GO:0048022) negative regulation of secondary metabolite biosynthetic process(GO:1900377) |
| 0.7 | 6.7 | GO:0045719 | negative regulation of glycogen biosynthetic process(GO:0045719) |
| 0.6 | 6.0 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
| 0.6 | 4.2 | GO:0051697 | protein delipidation(GO:0051697) |
| 0.6 | 3.5 | GO:0002528 | regulation of vascular permeability involved in acute inflammatory response(GO:0002528) |
| 0.6 | 1.7 | GO:1901074 | regulation of engulfment of apoptotic cell(GO:1901074) |
| 0.6 | 7.2 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.5 | 2.1 | GO:0046098 | guanine metabolic process(GO:0046098) |
| 0.5 | 1.6 | GO:0006550 | isoleucine catabolic process(GO:0006550) |
| 0.5 | 3.9 | GO:1901748 | peptide modification(GO:0031179) leukotriene D4 metabolic process(GO:1901748) leukotriene D4 biosynthetic process(GO:1901750) |
| 0.5 | 5.5 | GO:2000467 | positive regulation of glycogen (starch) synthase activity(GO:2000467) |
| 0.5 | 2.7 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.4 | 7.5 | GO:0043129 | surfactant homeostasis(GO:0043129) |
| 0.4 | 1.7 | GO:0071034 | CUT catabolic process(GO:0071034) CUT metabolic process(GO:0071043) |
| 0.4 | 2.4 | GO:0061641 | CENP-A containing nucleosome assembly(GO:0034080) CENP-A containing chromatin organization(GO:0061641) |
| 0.4 | 2.8 | GO:2000911 | positive regulation of cholesterol import(GO:1904109) positive regulation of sterol import(GO:2000911) |
| 0.4 | 4.5 | GO:0002826 | negative regulation of T-helper 1 type immune response(GO:0002826) |
| 0.4 | 1.4 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.4 | 3.9 | GO:0032049 | cardiolipin biosynthetic process(GO:0032049) |
| 0.4 | 6.4 | GO:1904778 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.3 | 2.5 | GO:0038094 | Fc-gamma receptor signaling pathway(GO:0038094) |
| 0.3 | 0.9 | GO:0044878 | mitotic cytokinesis checkpoint(GO:0044878) |
| 0.3 | 1.7 | GO:1902962 | regulation of metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902962) negative regulation of metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902963) |
| 0.3 | 1.9 | GO:0021886 | hypothalamus gonadotrophin-releasing hormone neuron differentiation(GO:0021886) hypothalamus gonadotrophin-releasing hormone neuron development(GO:0021888) |
| 0.3 | 3.2 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.3 | 2.1 | GO:0097647 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.3 | 0.8 | GO:1902567 | immune complex clearance(GO:0002434) immune complex clearance by monocytes and macrophages(GO:0002436) regulation of immune complex clearance by monocytes and macrophages(GO:0090264) positive regulation of immune complex clearance by monocytes and macrophages(GO:0090265) negative regulation of eosinophil activation(GO:1902567) positive regulation of CD8-positive, alpha-beta T cell extravasation(GO:2000451) |
| 0.3 | 3.8 | GO:0036376 | sodium ion export from cell(GO:0036376) |
| 0.2 | 2.4 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.2 | 3.2 | GO:0006388 | tRNA splicing, via endonucleolytic cleavage and ligation(GO:0006388) |
| 0.2 | 5.0 | GO:0031000 | response to caffeine(GO:0031000) |
| 0.2 | 4.3 | GO:0033601 | positive regulation of mammary gland epithelial cell proliferation(GO:0033601) |
| 0.2 | 2.0 | GO:1900262 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.2 | 1.3 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.2 | 0.4 | GO:0061346 | dichotomous subdivision of terminal units involved in lung branching(GO:0060448) non-canonical Wnt signaling pathway involved in heart development(GO:0061341) planar cell polarity pathway involved in heart morphogenesis(GO:0061346) |
| 0.2 | 1.6 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.2 | 1.5 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.2 | 0.6 | GO:0051892 | negative regulation of cardioblast differentiation(GO:0051892) |
| 0.2 | 1.7 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.2 | 1.4 | GO:0002238 | response to molecule of fungal origin(GO:0002238) |
| 0.2 | 1.0 | GO:1903054 | negative regulation of extracellular matrix organization(GO:1903054) |
| 0.1 | 1.6 | GO:2000582 | regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000580) positive regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000582) |
| 0.1 | 3.1 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.1 | 1.0 | GO:2001205 | negative regulation of osteoclast development(GO:2001205) |
| 0.1 | 2.0 | GO:0046884 | follicle-stimulating hormone secretion(GO:0046884) |
| 0.1 | 0.4 | GO:0070100 | negative regulation of chemokine-mediated signaling pathway(GO:0070100) |
| 0.1 | 2.6 | GO:0050930 | induction of positive chemotaxis(GO:0050930) |
| 0.1 | 0.5 | GO:0002752 | cell surface pattern recognition receptor signaling pathway(GO:0002752) positive regulation of acute inflammatory response to non-antigenic stimulus(GO:0002879) |
| 0.1 | 0.5 | GO:1903070 | negative regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903070) |
| 0.1 | 0.9 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.1 | 4.9 | GO:0048266 | behavioral response to pain(GO:0048266) |
| 0.1 | 1.0 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) |
| 0.1 | 2.2 | GO:0051382 | kinetochore assembly(GO:0051382) |
| 0.1 | 1.1 | GO:2000774 | positive regulation of cellular senescence(GO:2000774) |
| 0.1 | 2.3 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 1.1 | GO:0035878 | nail development(GO:0035878) |
| 0.1 | 0.5 | GO:0035063 | nuclear speck organization(GO:0035063) |
| 0.1 | 1.0 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.1 | 1.8 | GO:0090266 | regulation of mitotic cell cycle spindle assembly checkpoint(GO:0090266) regulation of mitotic spindle checkpoint(GO:1903504) |
| 0.1 | 2.8 | GO:0021860 | pyramidal neuron development(GO:0021860) |
| 0.1 | 1.3 | GO:1904659 | glucose transmembrane transport(GO:1904659) |
| 0.1 | 0.8 | GO:0090520 | sphingolipid mediated signaling pathway(GO:0090520) |
| 0.1 | 0.3 | GO:0032972 | regulation of muscle filament sliding speed(GO:0032972) |
| 0.1 | 0.2 | GO:2001200 | positive regulation of dendritic cell differentiation(GO:2001200) |
| 0.1 | 0.2 | GO:0006428 | isoleucyl-tRNA aminoacylation(GO:0006428) |
| 0.1 | 1.0 | GO:1903599 | positive regulation of mitophagy(GO:1903599) |
| 0.1 | 2.2 | GO:0097435 | fibril organization(GO:0097435) |
| 0.1 | 0.9 | GO:2000628 | regulation of miRNA metabolic process(GO:2000628) |
| 0.1 | 0.8 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.1 | 2.0 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.1 | 1.0 | GO:0045602 | negative regulation of endothelial cell differentiation(GO:0045602) |
| 0.1 | 0.4 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 0.1 | 2.1 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.1 | 2.1 | GO:0000470 | maturation of LSU-rRNA(GO:0000470) |
| 0.1 | 2.8 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.1 | 0.5 | GO:0043314 | negative regulation of neutrophil degranulation(GO:0043314) negative regulation of blood vessel remodeling(GO:0060313) negative regulation of neutrophil activation(GO:1902564) |
| 0.1 | 8.3 | GO:0050853 | B cell receptor signaling pathway(GO:0050853) |
| 0.1 | 0.2 | GO:2000481 | positive regulation of cAMP-dependent protein kinase activity(GO:2000481) |
| 0.1 | 7.7 | GO:0007586 | digestion(GO:0007586) |
| 0.1 | 7.0 | GO:0042100 | B cell proliferation(GO:0042100) |
| 0.1 | 1.1 | GO:0001522 | pseudouridine synthesis(GO:0001522) |
| 0.1 | 0.5 | GO:0031937 | positive regulation of chromatin silencing(GO:0031937) |
| 0.0 | 2.6 | GO:0033119 | negative regulation of RNA splicing(GO:0033119) |
| 0.0 | 1.1 | GO:0001675 | acrosome assembly(GO:0001675) |
| 0.0 | 1.5 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 1.6 | GO:0006907 | pinocytosis(GO:0006907) |
| 0.0 | 3.3 | GO:0051693 | actin filament capping(GO:0051693) |
| 0.0 | 0.7 | GO:0002756 | MyD88-independent toll-like receptor signaling pathway(GO:0002756) |
| 0.0 | 0.2 | GO:0045726 | positive regulation of integrin biosynthetic process(GO:0045726) cellular response to hyperoxia(GO:0071455) |
| 0.0 | 0.4 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.0 | 0.6 | GO:0044406 | adhesion of symbiont to host(GO:0044406) |
| 0.0 | 1.0 | GO:0006744 | ubiquinone biosynthetic process(GO:0006744) |
| 0.0 | 0.2 | GO:0006421 | asparaginyl-tRNA aminoacylation(GO:0006421) |
| 0.0 | 0.8 | GO:0048934 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.0 | 0.1 | GO:0042264 | peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 0.0 | 3.7 | GO:0008543 | fibroblast growth factor receptor signaling pathway(GO:0008543) |
| 0.0 | 0.5 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.0 | 0.5 | GO:0007342 | fusion of sperm to egg plasma membrane(GO:0007342) |
| 0.0 | 0.2 | GO:0030853 | negative regulation of granulocyte differentiation(GO:0030853) |
| 0.0 | 0.8 | GO:0060712 | spongiotrophoblast layer development(GO:0060712) |
| 0.0 | 0.1 | GO:0034371 | chylomicron remodeling(GO:0034371) |
| 0.0 | 0.1 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 0.0 | 0.8 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 0.7 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.0 | 0.2 | GO:0035726 | common myeloid progenitor cell proliferation(GO:0035726) |
| 0.0 | 1.0 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 0.1 | GO:0060316 | positive regulation of ryanodine-sensitive calcium-release channel activity(GO:0060316) |
| 0.0 | 1.5 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.0 | 0.1 | GO:2000583 | regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000583) negative regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000584) |
| 0.0 | 0.0 | GO:0060003 | copper ion export(GO:0060003) |
| 0.0 | 0.1 | GO:0072161 | mesenchymal cell differentiation involved in kidney development(GO:0072161) mesenchymal cell differentiation involved in renal system development(GO:2001012) |
| 0.0 | 0.6 | GO:0002183 | cytoplasmic translational initiation(GO:0002183) |
| 0.0 | 1.1 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.0 | 0.7 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.0 | 0.3 | GO:0070072 | vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.0 | 0.2 | GO:0048664 | neuron fate determination(GO:0048664) |
| 0.0 | 0.4 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.0 | 0.4 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.0 | 0.5 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.0 | 0.3 | GO:0035493 | SNARE complex assembly(GO:0035493) |
| 0.0 | 2.9 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 1.2 | GO:0043966 | histone H3 acetylation(GO:0043966) |
| 0.0 | 0.6 | GO:0006471 | protein ADP-ribosylation(GO:0006471) |
| 0.0 | 0.6 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 0.7 | GO:0043044 | ATP-dependent chromatin remodeling(GO:0043044) |
| 0.0 | 0.8 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.0 | 0.7 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.0 | 1.5 | GO:0034446 | substrate adhesion-dependent cell spreading(GO:0034446) |
| 0.0 | 0.3 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.0 | 0.6 | GO:0042273 | ribosomal large subunit biogenesis(GO:0042273) |
| 0.0 | 1.0 | GO:1903955 | positive regulation of protein targeting to mitochondrion(GO:1903955) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 9.3 | GO:0000942 | condensed nuclear chromosome outer kinetochore(GO:0000942) |
| 1.2 | 19.5 | GO:0042555 | MCM complex(GO:0042555) |
| 1.0 | 16.2 | GO:0005766 | primary lysosome(GO:0005766) azurophil granule(GO:0042582) |
| 0.6 | 7.5 | GO:0097208 | alveolar lamellar body(GO:0097208) |
| 0.6 | 6.2 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.4 | 2.3 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.4 | 1.9 | GO:0000444 | MIS12/MIND type complex(GO:0000444) |
| 0.3 | 3.8 | GO:0044326 | dendritic spine neck(GO:0044326) |
| 0.3 | 2.0 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 0.3 | 2.0 | GO:0042585 | germinal vesicle(GO:0042585) |
| 0.3 | 1.3 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.2 | 1.0 | GO:1990037 | Lewy body core(GO:1990037) |
| 0.2 | 1.7 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 0.2 | 2.8 | GO:0032797 | SMN complex(GO:0032797) |
| 0.2 | 1.6 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) tubular endosome(GO:0097422) |
| 0.2 | 2.8 | GO:0071203 | WASH complex(GO:0071203) |
| 0.2 | 1.9 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.1 | 5.8 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.1 | 0.4 | GO:0060187 | cell pole(GO:0060187) |
| 0.1 | 1.4 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.1 | 5.8 | GO:0097228 | sperm principal piece(GO:0097228) |
| 0.1 | 2.2 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.1 | 0.8 | GO:1990393 | 3M complex(GO:1990393) |
| 0.1 | 1.5 | GO:0001739 | sex chromatin(GO:0001739) |
| 0.1 | 4.8 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.1 | 8.1 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.1 | 0.9 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.1 | 2.4 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.1 | 1.0 | GO:0034709 | methylosome(GO:0034709) |
| 0.1 | 2.4 | GO:0005732 | small nucleolar ribonucleoprotein complex(GO:0005732) |
| 0.1 | 0.9 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.1 | 1.6 | GO:0098644 | complex of collagen trimers(GO:0098644) |
| 0.1 | 1.0 | GO:0000138 | Golgi trans cisterna(GO:0000138) |
| 0.1 | 0.3 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 0.1 | 1.2 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.1 | 0.8 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 1.8 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.1 | 1.0 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.1 | 0.6 | GO:0071541 | eukaryotic translation initiation factor 3 complex, eIF3m(GO:0071541) |
| 0.1 | 1.8 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.1 | 0.8 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.1 | 0.3 | GO:0070820 | tertiary granule(GO:0070820) |
| 0.1 | 1.7 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 0.3 | GO:1990584 | cardiac Troponin complex(GO:1990584) |
| 0.0 | 3.0 | GO:0099738 | cell cortex region(GO:0099738) |
| 0.0 | 0.1 | GO:0009328 | phenylalanine-tRNA ligase complex(GO:0009328) |
| 0.0 | 1.2 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.0 | 2.6 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.6 | GO:0020005 | symbiont-containing vacuole membrane(GO:0020005) |
| 0.0 | 4.3 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.0 | 0.5 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.0 | 2.5 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 0.9 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.0 | 0.9 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 7.2 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 0.8 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.0 | 0.8 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.0 | 1.2 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.0 | 1.1 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.0 | 1.0 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 1.0 | GO:0002080 | acrosomal membrane(GO:0002080) |
| 0.0 | 0.6 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 1.0 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.0 | 0.2 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.0 | 1.2 | GO:0030286 | dynein complex(GO:0030286) |
| 0.0 | 1.3 | GO:1990391 | DNA repair complex(GO:1990391) |
| 0.0 | 0.4 | GO:0000346 | transcription export complex(GO:0000346) |
| 0.0 | 10.0 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 0.8 | GO:0005921 | gap junction(GO:0005921) |
| 0.0 | 4.0 | GO:0000775 | chromosome, centromeric region(GO:0000775) |
| 0.0 | 0.3 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.0 | 2.4 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 2.1 | GO:0019814 | immunoglobulin complex(GO:0019814) |
| 0.0 | 5.8 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 5.0 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 0.9 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 0.2 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 9.3 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 1.5 | 4.5 | GO:0002113 | interleukin-33 binding(GO:0002113) |
| 1.3 | 3.9 | GO:0004605 | phosphatidate cytidylyltransferase activity(GO:0004605) |
| 1.1 | 4.5 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
| 0.9 | 2.8 | GO:0030622 | U4atac snRNA binding(GO:0030622) |
| 0.8 | 4.0 | GO:0004809 | tRNA (guanine-N2-)-methyltransferase activity(GO:0004809) |
| 0.8 | 6.4 | GO:0008865 | fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.8 | 3.2 | GO:0032093 | SAM domain binding(GO:0032093) |
| 0.6 | 2.4 | GO:0050309 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 0.6 | 4.3 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 0.6 | 1.7 | GO:0034188 | apolipoprotein receptor activity(GO:0030226) apolipoprotein A-I receptor activity(GO:0034188) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.6 | 2.2 | GO:0030023 | extracellular matrix constituent conferring elasticity(GO:0030023) |
| 0.6 | 7.2 | GO:0008199 | ferric iron binding(GO:0008199) |
| 0.5 | 3.2 | GO:0000213 | tRNA-intron endonuclease activity(GO:0000213) |
| 0.5 | 5.6 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.5 | 2.7 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 0.4 | 19.5 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.4 | 3.5 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.4 | 2.1 | GO:0034648 | histone demethylase activity (H3-dimethyl-K4 specific)(GO:0034648) |
| 0.3 | 5.0 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.3 | 2.3 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.3 | 3.9 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.3 | 3.8 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.3 | 1.2 | GO:0004574 | oligo-1,6-glucosidase activity(GO:0004574) |
| 0.3 | 1.1 | GO:0000010 | trans-hexaprenyltranstransferase activity(GO:0000010) trans-octaprenyltranstransferase activity(GO:0050347) |
| 0.3 | 3.6 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.3 | 0.8 | GO:0035717 | chemokine (C-C motif) ligand 7 binding(GO:0035717) |
| 0.2 | 8.1 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.2 | 7.5 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.2 | 8.9 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.2 | 1.6 | GO:0004084 | branched-chain-amino-acid transaminase activity(GO:0004084) L-leucine transaminase activity(GO:0052654) L-valine transaminase activity(GO:0052655) L-isoleucine transaminase activity(GO:0052656) |
| 0.2 | 1.4 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.2 | 0.9 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.2 | 1.3 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.2 | 2.0 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.2 | 11.9 | GO:0004601 | peroxidase activity(GO:0004601) |
| 0.2 | 1.0 | GO:0033188 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.2 | 2.7 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.1 | 6.0 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.1 | 6.7 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.1 | 3.1 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 2.7 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 1.3 | GO:0005412 | glucose:sodium symporter activity(GO:0005412) |
| 0.1 | 1.7 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.1 | 0.9 | GO:0030275 | LRR domain binding(GO:0030275) |
| 0.1 | 0.4 | GO:0003696 | satellite DNA binding(GO:0003696) |
| 0.1 | 1.4 | GO:0019534 | toxin transporter activity(GO:0019534) |
| 0.1 | 1.7 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.1 | 0.9 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 0.1 | 1.0 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 0.1 | 3.1 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.1 | 0.8 | GO:0034452 | dynactin binding(GO:0034452) |
| 0.1 | 10.3 | GO:0005179 | hormone activity(GO:0005179) |
| 0.1 | 0.2 | GO:0004822 | isoleucine-tRNA ligase activity(GO:0004822) |
| 0.1 | 4.3 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.1 | 0.8 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 0.1 | 0.4 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.1 | 15.1 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 2.8 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.1 | 1.0 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.1 | 1.1 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.1 | 3.5 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.1 | 0.9 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.1 | 0.2 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.1 | 0.7 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.1 | 1.6 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.1 | 1.0 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.0 | 0.3 | GO:0031013 | troponin C binding(GO:0030172) troponin I binding(GO:0031013) |
| 0.0 | 0.4 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 0.0 | 1.0 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.0 | 4.7 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 1.7 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.0 | 0.6 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.0 | 0.5 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.0 | 3.9 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.1 | GO:0004816 | asparagine-tRNA ligase activity(GO:0004816) |
| 0.0 | 0.2 | GO:0030171 | voltage-gated proton channel activity(GO:0030171) |
| 0.0 | 1.4 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.0 | 4.7 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 0.7 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.0 | 0.2 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.0 | 0.1 | GO:0035478 | chylomicron binding(GO:0035478) |
| 0.0 | 0.4 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.0 | 0.8 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 0.1 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.0 | 0.8 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.0 | 1.0 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.0 | 3.8 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.2 | GO:1901611 | phosphatidylglycerol binding(GO:1901611) |
| 0.0 | 0.4 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.0 | 0.5 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.0 | 5.8 | GO:0005549 | odorant binding(GO:0005549) |
| 0.0 | 5.9 | GO:0001047 | core promoter binding(GO:0001047) |
| 0.0 | 0.6 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 1.3 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 1.2 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 1.3 | GO:0015179 | L-amino acid transmembrane transporter activity(GO:0015179) |
| 0.0 | 0.3 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.0 | 2.4 | GO:0004519 | endonuclease activity(GO:0004519) |
| 0.0 | 6.6 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 0.5 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.5 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.0 | 0.1 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.0 | 0.5 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.0 | 0.5 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.0 | 0.0 | GO:0004008 | copper-exporting ATPase activity(GO:0004008) copper-transporting ATPase activity(GO:0043682) |
| 0.0 | 0.5 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 0.1 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.0 | 1.1 | GO:0004620 | phospholipase activity(GO:0004620) |
| 0.0 | 1.3 | GO:0043130 | ubiquitin binding(GO:0043130) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 21.9 | PID ATR PATHWAY | ATR signaling pathway |
| 0.4 | 6.5 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.3 | 16.2 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.2 | 9.7 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 6.4 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.1 | 6.7 | PID IGF1 PATHWAY | IGF1 pathway |
| 0.1 | 8.1 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 2.7 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 6.2 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.1 | 4.5 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.1 | 1.0 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.1 | 3.1 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 1.7 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.1 | 3.5 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.1 | 0.5 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.1 | 4.9 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.0 | 4.4 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 1.1 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 1.0 | PID S1P S1P3 PATHWAY | S1P3 pathway |
| 0.0 | 0.8 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.0 | 10.0 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.7 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.0 | 1.6 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 1.8 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 1.0 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.0 | 1.4 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 2.0 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.0 | 1.1 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.0 | 7.9 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 3.8 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.2 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.0 | 1.0 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 2.9 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.7 | PID CXCR4 PATHWAY | CXCR4-mediated signaling events |
| 0.0 | 0.7 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 19.5 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 1.1 | 6.5 | REACTOME CDC6 ASSOCIATION WITH THE ORC ORIGIN COMPLEX | Genes involved in CDC6 association with the ORC:origin complex |
| 0.8 | 9.3 | REACTOME E2F ENABLED INHIBITION OF PRE REPLICATION COMPLEX FORMATION | Genes involved in E2F-enabled inhibition of pre-replication complex formation |
| 0.5 | 6.8 | REACTOME DEFENSINS | Genes involved in Defensins |
| 0.3 | 3.2 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.3 | 6.7 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.3 | 5.5 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.2 | 2.0 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |
| 0.2 | 1.9 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.2 | 2.0 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.2 | 8.9 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.1 | 6.2 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.1 | 1.6 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.1 | 2.0 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 0.1 | 2.4 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.1 | 2.8 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.1 | 1.3 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.1 | 1.6 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.1 | 1.1 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.1 | 4.7 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.1 | 3.9 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 9.3 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.1 | 1.3 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.1 | 1.7 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.1 | 2.1 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.1 | 1.7 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.1 | 6.6 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.1 | 1.6 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.1 | 2.2 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 0.9 | REACTOME VIRAL MESSENGER RNA SYNTHESIS | Genes involved in Viral Messenger RNA Synthesis |
| 0.0 | 1.0 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 3.3 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.7 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.2 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.0 | 1.2 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.0 | 0.9 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.0 | 1.1 | REACTOME PROSTACYCLIN SIGNALLING THROUGH PROSTACYCLIN RECEPTOR | Genes involved in Prostacyclin signalling through prostacyclin receptor |
| 0.0 | 0.7 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.4 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 1.5 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 0.4 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.0 | 0.3 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.0 | 0.8 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.9 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.0 | 0.5 | REACTOME PREFOLDIN MEDIATED TRANSFER OF SUBSTRATE TO CCT TRIC | Genes involved in Prefoldin mediated transfer of substrate to CCT/TriC |
| 0.0 | 0.5 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |