Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
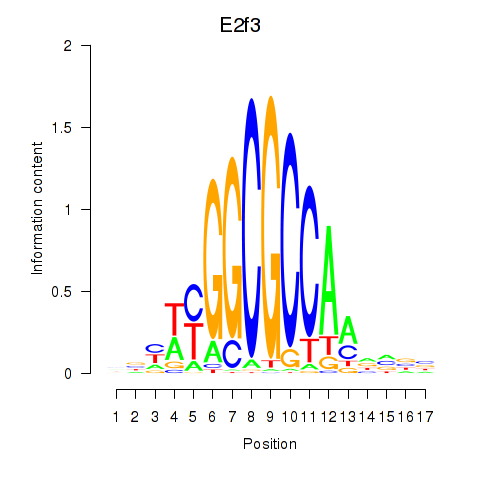
Results for E2f3
Z-value: 2.29
Transcription factors associated with E2f3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
E2f3
|
ENSMUSG00000016477.19 | E2f3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| E2f3 | mm39_v1_chr13_-_30170031_30170048 | 0.93 | 7.5e-16 | Click! |
Activity profile of E2f3 motif
Sorted Z-values of E2f3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of E2f3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_-_6733411 | 13.70 |
ENSMUST00000239104.2
ENSMUST00000051209.11 |
Peg3
|
paternally expressed 3 |
| chr3_-_84387700 | 12.85 |
ENSMUST00000194027.2
ENSMUST00000107689.7 |
Fhdc1
|
FH2 domain containing 1 |
| chr2_+_117942357 | 12.51 |
ENSMUST00000039559.9
|
Thbs1
|
thrombospondin 1 |
| chr2_-_126975804 | 10.48 |
ENSMUST00000110387.4
|
Ncaph
|
non-SMC condensin I complex, subunit H |
| chr12_+_24758968 | 9.68 |
ENSMUST00000154588.2
|
Rrm2
|
ribonucleotide reductase M2 |
| chr11_+_11635908 | 9.37 |
ENSMUST00000065433.12
|
Ikzf1
|
IKAROS family zinc finger 1 |
| chr12_-_4924341 | 9.20 |
ENSMUST00000137337.8
ENSMUST00000045921.14 |
Mfsd2b
|
major facilitator superfamily domain containing 2B |
| chr12_-_69274936 | 8.29 |
ENSMUST00000221411.2
ENSMUST00000021359.7 |
Pole2
|
polymerase (DNA directed), epsilon 2 (p59 subunit) |
| chr12_+_24758724 | 8.11 |
ENSMUST00000153058.8
|
Rrm2
|
ribonucleotide reductase M2 |
| chr5_-_113957362 | 8.06 |
ENSMUST00000202555.2
|
Selplg
|
selectin, platelet (p-selectin) ligand |
| chr15_+_84116231 | 7.98 |
ENSMUST00000023072.7
|
Parvb
|
parvin, beta |
| chr3_+_10077608 | 7.87 |
ENSMUST00000029046.9
|
Fabp5
|
fatty acid binding protein 5, epidermal |
| chr7_+_13012735 | 7.82 |
ENSMUST00000098814.13
ENSMUST00000146998.9 |
Lig1
|
ligase I, DNA, ATP-dependent |
| chr12_+_24758240 | 7.70 |
ENSMUST00000020980.12
|
Rrm2
|
ribonucleotide reductase M2 |
| chr10_+_20223516 | 7.54 |
ENSMUST00000169712.3
ENSMUST00000217608.2 |
Mtfr2
|
mitochondrial fission regulator 2 |
| chr4_+_43957401 | 7.49 |
ENSMUST00000030202.14
|
Glipr2
|
GLI pathogenesis-related 2 |
| chr9_+_64188857 | 7.44 |
ENSMUST00000215031.2
ENSMUST00000213165.2 ENSMUST00000213289.2 ENSMUST00000216594.2 ENSMUST00000034964.7 |
Tipin
|
timeless interacting protein |
| chr8_+_95712151 | 7.06 |
ENSMUST00000212799.2
|
Adgrg1
|
adhesion G protein-coupled receptor G1 |
| chr8_+_123294740 | 6.46 |
ENSMUST00000006760.3
|
Cdt1
|
chromatin licensing and DNA replication factor 1 |
| chr5_-_113957318 | 6.38 |
ENSMUST00000201194.4
|
Selplg
|
selectin, platelet (p-selectin) ligand |
| chr4_+_43957677 | 6.29 |
ENSMUST00000107855.2
|
Glipr2
|
GLI pathogenesis-related 2 |
| chr4_-_140805613 | 5.92 |
ENSMUST00000030760.15
|
Necap2
|
NECAP endocytosis associated 2 |
| chr10_-_93425553 | 5.22 |
ENSMUST00000020203.7
|
Snrpf
|
small nuclear ribonucleoprotein polypeptide F |
| chr15_-_83479312 | 5.19 |
ENSMUST00000016901.5
|
Ttll12
|
tubulin tyrosine ligase-like family, member 12 |
| chr5_-_148336711 | 5.07 |
ENSMUST00000048116.15
|
Slc7a1
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 |
| chrX_-_101232978 | 4.92 |
ENSMUST00000033683.8
|
Rps4x
|
ribosomal protein S4, X-linked |
| chr5_-_68004743 | 4.45 |
ENSMUST00000072971.13
ENSMUST00000113652.8 ENSMUST00000113651.8 ENSMUST00000037380.15 |
Atp8a1
|
ATPase, aminophospholipid transporter (APLT), class I, type 8A, member 1 |
| chr15_-_79967543 | 4.23 |
ENSMUST00000081650.15
|
Rpl3
|
ribosomal protein L3 |
| chr16_-_4536943 | 4.13 |
ENSMUST00000120056.8
ENSMUST00000074970.8 |
Nmral1
|
NmrA-like family domain containing 1 |
| chr11_-_101442663 | 4.11 |
ENSMUST00000017290.11
|
Brca1
|
breast cancer 1, early onset |
| chr16_-_4536992 | 4.06 |
ENSMUST00000115851.10
|
Nmral1
|
NmrA-like family domain containing 1 |
| chr3_+_68912302 | 3.80 |
ENSMUST00000136502.8
ENSMUST00000107803.7 |
Smc4
|
structural maintenance of chromosomes 4 |
| chr2_+_150628655 | 3.68 |
ENSMUST00000045441.8
|
Pygb
|
brain glycogen phosphorylase |
| chr4_+_132495636 | 3.67 |
ENSMUST00000102561.11
|
Rpa2
|
replication protein A2 |
| chr11_+_115455260 | 3.58 |
ENSMUST00000021085.11
|
Nup85
|
nucleoporin 85 |
| chr2_+_150751475 | 3.55 |
ENSMUST00000028948.5
|
Gins1
|
GINS complex subunit 1 (Psf1 homolog) |
| chr17_+_27136065 | 3.52 |
ENSMUST00000078961.6
|
Kifc5b
|
kinesin family member C5B |
| chrX_+_20570145 | 3.48 |
ENSMUST00000033383.3
|
Usp11
|
ubiquitin specific peptidase 11 |
| chr4_+_109137561 | 3.43 |
ENSMUST00000177089.8
ENSMUST00000175776.8 ENSMUST00000132165.9 |
Eps15
|
epidermal growth factor receptor pathway substrate 15 |
| chr18_+_31922173 | 3.18 |
ENSMUST00000025106.5
ENSMUST00000234146.2 |
Polr2d
|
polymerase (RNA) II (DNA directed) polypeptide D |
| chr18_+_67933169 | 3.11 |
ENSMUST00000025425.7
|
Cep192
|
centrosomal protein 192 |
| chr17_-_71782296 | 3.11 |
ENSMUST00000127430.2
|
Smchd1
|
SMC hinge domain containing 1 |
| chr10_-_80754016 | 3.11 |
ENSMUST00000057623.14
ENSMUST00000179022.8 |
Lmnb2
|
lamin B2 |
| chr5_-_36555434 | 3.04 |
ENSMUST00000037370.14
ENSMUST00000070720.8 |
Sorcs2
|
sortilin-related VPS10 domain containing receptor 2 |
| chr13_-_55477535 | 3.01 |
ENSMUST00000021941.8
|
Mxd3
|
Max dimerization protein 3 |
| chr7_+_89779421 | 2.96 |
ENSMUST00000207225.2
ENSMUST00000207484.2 ENSMUST00000209068.2 |
Picalm
|
phosphatidylinositol binding clathrin assembly protein |
| chr13_-_51947685 | 2.80 |
ENSMUST00000110040.9
ENSMUST00000021900.14 |
Sema4d
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4D |
| chr18_+_11790409 | 2.77 |
ENSMUST00000047322.8
|
Rbbp8
|
retinoblastoma binding protein 8, endonuclease |
| chr8_-_124586159 | 2.76 |
ENSMUST00000034452.12
|
Ccsap
|
centriole, cilia and spindle associated protein |
| chr1_-_128287347 | 2.74 |
ENSMUST00000190495.2
ENSMUST00000027601.11 |
Mcm6
|
minichromosome maintenance complex component 6 |
| chr2_+_109111083 | 2.65 |
ENSMUST00000028527.8
|
Kif18a
|
kinesin family member 18A |
| chr16_-_78173645 | 2.54 |
ENSMUST00000023570.14
|
Btg3
|
BTG anti-proliferation factor 3 |
| chr12_+_111505216 | 2.54 |
ENSMUST00000050993.11
|
Eif5
|
eukaryotic translation initiation factor 5 |
| chr5_-_65855511 | 2.45 |
ENSMUST00000201948.4
|
Pds5a
|
PDS5 cohesin associated factor A |
| chr8_+_106587212 | 2.43 |
ENSMUST00000008594.9
|
Nutf2
|
nuclear transport factor 2 |
| chr12_+_111505253 | 2.34 |
ENSMUST00000220803.2
|
Eif5
|
eukaryotic translation initiation factor 5 |
| chr8_+_106587268 | 2.31 |
ENSMUST00000212610.2
ENSMUST00000212484.2 ENSMUST00000212200.2 |
Nutf2
|
nuclear transport factor 2 |
| chr6_-_72322116 | 2.30 |
ENSMUST00000070345.5
|
Usp39
|
ubiquitin specific peptidase 39 |
| chr11_-_84020382 | 2.25 |
ENSMUST00000018795.13
|
Tada2a
|
transcriptional adaptor 2A |
| chr1_-_66902429 | 2.23 |
ENSMUST00000027153.6
|
Acadl
|
acyl-Coenzyme A dehydrogenase, long-chain |
| chr8_-_57940834 | 2.22 |
ENSMUST00000034022.4
|
Sap30
|
sin3 associated polypeptide |
| chr4_-_34614883 | 2.21 |
ENSMUST00000140334.2
ENSMUST00000108142.8 ENSMUST00000048706.10 |
Orc3
|
origin recognition complex, subunit 3 |
| chr8_-_120331936 | 2.20 |
ENSMUST00000093099.13
|
Taf1c
|
TATA-box binding protein associated factor, RNA polymerase I, C |
| chr4_+_109137462 | 2.18 |
ENSMUST00000102729.10
|
Eps15
|
epidermal growth factor receptor pathway substrate 15 |
| chrX_-_72703330 | 2.14 |
ENSMUST00000114473.8
ENSMUST00000002087.14 |
Pnck
|
pregnancy upregulated non-ubiquitously expressed CaM kinase |
| chr17_+_29251602 | 2.12 |
ENSMUST00000130216.3
|
Srsf3
|
serine and arginine-rich splicing factor 3 |
| chr10_-_61619790 | 2.11 |
ENSMUST00000020283.5
|
Macroh2a2
|
macroH2A.2 histone |
| chr17_+_36152383 | 2.05 |
ENSMUST00000082337.13
|
Mdc1
|
mediator of DNA damage checkpoint 1 |
| chr6_+_120440159 | 2.05 |
ENSMUST00000002976.5
|
Il17ra
|
interleukin 17 receptor A |
| chr8_+_95584146 | 2.01 |
ENSMUST00000211939.2
ENSMUST00000212124.2 |
Polr2c
|
polymerase (RNA) II (DNA directed) polypeptide C |
| chr5_-_100867520 | 2.00 |
ENSMUST00000112908.2
ENSMUST00000045617.15 |
Hpse
|
heparanase |
| chr2_+_115412148 | 1.94 |
ENSMUST00000166472.8
ENSMUST00000110918.3 |
Cdin1
|
CDAN1 interacting nuclease 1 |
| chr5_-_123038329 | 1.93 |
ENSMUST00000031435.14
|
Kdm2b
|
lysine (K)-specific demethylase 2B |
| chr18_-_52662728 | 1.90 |
ENSMUST00000025409.9
|
Lox
|
lysyl oxidase |
| chr17_-_67661382 | 1.87 |
ENSMUST00000223982.2
ENSMUST00000224091.2 |
Ptprm
|
protein tyrosine phosphatase, receptor type, M |
| chr14_-_30348153 | 1.87 |
ENSMUST00000112211.9
ENSMUST00000112210.11 |
Prkcd
|
protein kinase C, delta |
| chr3_-_51248032 | 1.86 |
ENSMUST00000062009.14
ENSMUST00000194641.6 |
Elf2
|
E74-like factor 2 |
| chr19_+_53588808 | 1.85 |
ENSMUST00000025930.10
|
Smc3
|
structural maintenance of chromosomes 3 |
| chr13_-_21715202 | 1.84 |
ENSMUST00000156674.3
ENSMUST00000110481.3 ENSMUST00000045228.12 |
Zkscan8
|
zinc finger with KRAB and SCAN domains 8 |
| chr2_+_31560725 | 1.83 |
ENSMUST00000038474.14
ENSMUST00000137156.2 |
Exosc2
|
exosome component 2 |
| chr16_+_91444730 | 1.81 |
ENSMUST00000119368.8
ENSMUST00000114037.9 ENSMUST00000114036.9 ENSMUST00000122302.8 |
Son
|
Son DNA binding protein |
| chr1_-_191307648 | 1.71 |
ENSMUST00000027933.11
|
Dtl
|
denticleless E3 ubiquitin protein ligase |
| chr4_+_11485945 | 1.71 |
ENSMUST00000055372.14
ENSMUST00000059914.13 |
Virma
|
vir like m6A methyltransferase associated |
| chr12_-_111638722 | 1.69 |
ENSMUST00000001304.9
|
Ckb
|
creatine kinase, brain |
| chr12_+_111504450 | 1.67 |
ENSMUST00000166123.9
ENSMUST00000222441.2 |
Eif5
|
eukaryotic translation initiation factor 5 |
| chr12_+_86725459 | 1.67 |
ENSMUST00000021681.4
|
Vash1
|
vasohibin 1 |
| chr15_+_99870661 | 1.67 |
ENSMUST00000100206.4
|
Larp4
|
La ribonucleoprotein domain family, member 4 |
| chr4_+_34614939 | 1.66 |
ENSMUST00000029968.14
ENSMUST00000148519.3 |
Rars2
|
arginyl-tRNA synthetase 2, mitochondrial |
| chr9_-_57673128 | 1.64 |
ENSMUST00000065330.8
|
Clk3
|
CDC-like kinase 3 |
| chr3_-_84063067 | 1.61 |
ENSMUST00000047368.8
|
Mnd1
|
meiotic nuclear divisions 1 |
| chr8_+_95584078 | 1.61 |
ENSMUST00000109521.4
|
Polr2c
|
polymerase (RNA) II (DNA directed) polypeptide C |
| chr4_+_130202388 | 1.60 |
ENSMUST00000070532.8
|
Fabp3
|
fatty acid binding protein 3, muscle and heart |
| chr10_-_80512117 | 1.60 |
ENSMUST00000200082.5
|
Mknk2
|
MAP kinase-interacting serine/threonine kinase 2 |
| chr2_+_32766126 | 1.59 |
ENSMUST00000028135.15
|
Niban2
|
niban apoptosis regulator 2 |
| chr5_-_68004702 | 1.59 |
ENSMUST00000135930.8
|
Atp8a1
|
ATPase, aminophospholipid transporter (APLT), class I, type 8A, member 1 |
| chr8_-_123303569 | 1.50 |
ENSMUST00000006764.9
|
Aprt
|
adenine phosphoribosyl transferase |
| chr8_-_123303352 | 1.48 |
ENSMUST00000211823.2
ENSMUST00000212093.2 |
Aprt
|
adenine phosphoribosyl transferase |
| chr8_-_70449018 | 1.42 |
ENSMUST00000065169.12
ENSMUST00000212478.2 |
Gatad2a
|
GATA zinc finger domain containing 2A |
| chr4_-_108436514 | 1.42 |
ENSMUST00000079213.6
|
Prpf38a
|
PRP38 pre-mRNA processing factor 38 (yeast) domain containing A |
| chr9_-_62888156 | 1.42 |
ENSMUST00000098651.6
ENSMUST00000214830.2 |
Pias1
|
protein inhibitor of activated STAT 1 |
| chr17_+_24633614 | 1.41 |
ENSMUST00000115371.9
ENSMUST00000088512.13 ENSMUST00000163717.2 |
Rnps1
|
RNA binding protein with serine rich domain 1 |
| chr5_-_34093678 | 1.39 |
ENSMUST00000030993.8
|
Nelfa
|
negative elongation factor complex member A, Whsc2 |
| chr12_+_80992294 | 1.38 |
ENSMUST00000110354.8
ENSMUST00000110352.10 ENSMUST00000110351.8 ENSMUST00000110356.3 |
Srsf5
|
serine and arginine-rich splicing factor 5 |
| chr11_+_55104609 | 1.35 |
ENSMUST00000108867.2
|
Slc36a1
|
solute carrier family 36 (proton/amino acid symporter), member 1 |
| chr5_-_148336574 | 1.29 |
ENSMUST00000202457.4
|
Slc7a1
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 |
| chr9_-_39514931 | 1.29 |
ENSMUST00000119722.8
|
AW551984
|
expressed sequence AW551984 |
| chrX_-_104973003 | 1.28 |
ENSMUST00000130980.2
ENSMUST00000113573.8 |
Atrx
|
ATRX, chromatin remodeler |
| chr1_+_127701901 | 1.26 |
ENSMUST00000112570.2
ENSMUST00000027587.15 |
Ccnt2
|
cyclin T2 |
| chr18_+_35252470 | 1.26 |
ENSMUST00000237154.2
|
Ctnna1
|
catenin (cadherin associated protein), alpha 1 |
| chr16_-_91443794 | 1.20 |
ENSMUST00000232367.2
ENSMUST00000231380.2 ENSMUST00000231444.2 ENSMUST00000232289.2 ENSMUST00000120450.2 ENSMUST00000023684.14 |
Gart
|
phosphoribosylglycinamide formyltransferase |
| chr9_+_56979307 | 1.19 |
ENSMUST00000169879.8
|
Sin3a
|
transcriptional regulator, SIN3A (yeast) |
| chr5_-_65855199 | 1.17 |
ENSMUST00000031104.7
|
Pds5a
|
PDS5 cohesin associated factor A |
| chr10_+_80062468 | 1.16 |
ENSMUST00000130260.2
|
Pwwp3a
|
PWWP domain containing 3A, DNA repair factor |
| chr4_+_11486002 | 1.16 |
ENSMUST00000108307.3
|
Virma
|
vir like m6A methyltransferase associated |
| chr10_+_80634488 | 1.15 |
ENSMUST00000151928.8
|
Sf3a2
|
splicing factor 3a, subunit 2 |
| chr10_+_75152705 | 1.12 |
ENSMUST00000105420.3
|
Adora2a
|
adenosine A2a receptor |
| chr6_-_97594498 | 1.12 |
ENSMUST00000113355.9
|
Frmd4b
|
FERM domain containing 4B |
| chr16_+_91282121 | 1.11 |
ENSMUST00000023689.11
ENSMUST00000117748.8 |
Ifnar1
|
interferon (alpha and beta) receptor 1 |
| chr9_+_113760376 | 1.08 |
ENSMUST00000214095.2
ENSMUST00000116492.3 ENSMUST00000216558.2 |
Ubp1
|
upstream binding protein 1 |
| chr18_+_44513547 | 1.07 |
ENSMUST00000202306.2
ENSMUST00000025350.10 |
Dcp2
|
decapping mRNA 2 |
| chr19_+_38384428 | 1.07 |
ENSMUST00000054098.4
|
Slc35g1
|
solute carrier family 35, member G1 |
| chr12_-_21423607 | 1.07 |
ENSMUST00000064536.13
|
Adam17
|
a disintegrin and metallopeptidase domain 17 |
| chr17_-_80369626 | 1.06 |
ENSMUST00000184635.8
|
Hnrnpll
|
heterogeneous nuclear ribonucleoprotein L-like |
| chr3_-_95125002 | 1.03 |
ENSMUST00000107209.8
|
Gabpb2
|
GA repeat binding protein, beta 2 |
| chr2_-_179867605 | 1.03 |
ENSMUST00000015791.6
|
Lama5
|
laminin, alpha 5 |
| chr18_+_9957906 | 1.03 |
ENSMUST00000025137.9
|
Thoc1
|
THO complex 1 |
| chr15_-_79430942 | 1.01 |
ENSMUST00000054014.9
ENSMUST00000229877.2 |
Ddx17
|
DEAD box helicase 17 |
| chr10_-_30476658 | 1.00 |
ENSMUST00000019927.7
|
Trmt11
|
tRNA methyltransferase 11 |
| chr3_-_65299967 | 1.00 |
ENSMUST00000119896.2
|
Ssr3
|
signal sequence receptor, gamma |
| chr8_-_70448872 | 1.00 |
ENSMUST00000177851.9
|
Gatad2a
|
GATA zinc finger domain containing 2A |
| chr12_-_21423551 | 0.98 |
ENSMUST00000101551.10
|
Adam17
|
a disintegrin and metallopeptidase domain 17 |
| chr7_-_5016237 | 0.97 |
ENSMUST00000208944.2
|
Fiz1
|
Flt3 interacting zinc finger protein 1 |
| chr3_-_41696906 | 0.91 |
ENSMUST00000026866.15
|
Sclt1
|
sodium channel and clathrin linker 1 |
| chr10_+_33780993 | 0.89 |
ENSMUST00000169670.8
|
Rsph4a
|
radial spoke head 4 homolog A (Chlamydomonas) |
| chr8_+_104837939 | 0.88 |
ENSMUST00000209911.2
|
Cdh5
|
cadherin 5 |
| chr7_+_100021425 | 0.88 |
ENSMUST00000098259.11
ENSMUST00000051777.15 |
C2cd3
|
C2 calcium-dependent domain containing 3 |
| chr15_+_99870787 | 0.87 |
ENSMUST00000231160.2
|
Larp4
|
La ribonucleoprotein domain family, member 4 |
| chr17_-_80369762 | 0.87 |
ENSMUST00000061331.14
|
Hnrnpll
|
heterogeneous nuclear ribonucleoprotein L-like |
| chr16_-_43709968 | 0.86 |
ENSMUST00000023387.14
|
Qtrt2
|
queuine tRNA-ribosyltransferase accessory subunit 2 |
| chr12_+_112112621 | 0.84 |
ENSMUST00000128402.3
|
Kif26a
|
kinesin family member 26A |
| chr1_+_156386414 | 0.82 |
ENSMUST00000166172.9
ENSMUST00000027888.13 |
Abl2
|
v-abl Abelson murine leukemia viral oncogene 2 (arg, Abelson-related gene) |
| chr17_-_48145466 | 0.81 |
ENSMUST00000066368.13
|
Mdfi
|
MyoD family inhibitor |
| chr3_+_66892979 | 0.80 |
ENSMUST00000162362.8
ENSMUST00000065074.14 ENSMUST00000065047.13 |
Rsrc1
|
arginine/serine-rich coiled-coil 1 |
| chr3_+_33854305 | 0.78 |
ENSMUST00000196139.5
ENSMUST00000200271.5 ENSMUST00000198529.5 ENSMUST00000117915.8 ENSMUST00000108210.9 ENSMUST00000196975.5 |
Ttc14
|
tetratricopeptide repeat domain 14 |
| chr3_-_95125190 | 0.76 |
ENSMUST00000136139.8
|
Gabpb2
|
GA repeat binding protein, beta 2 |
| chr11_-_86435579 | 0.75 |
ENSMUST00000138810.3
ENSMUST00000058286.9 ENSMUST00000154617.8 |
Rps6kb1
|
ribosomal protein S6 kinase, polypeptide 1 |
| chr4_+_116565784 | 0.74 |
ENSMUST00000138305.8
ENSMUST00000125671.8 ENSMUST00000130828.8 |
Ccdc163
|
coiled-coil domain containing 163 |
| chrX_-_104972938 | 0.74 |
ENSMUST00000198448.5
ENSMUST00000199233.5 ENSMUST00000134507.8 ENSMUST00000154866.8 ENSMUST00000128968.8 ENSMUST00000134381.8 ENSMUST00000150914.8 |
Atrx
|
ATRX, chromatin remodeler |
| chr13_+_54722833 | 0.73 |
ENSMUST00000156024.2
|
Arl10
|
ADP-ribosylation factor-like 10 |
| chr6_-_88818832 | 0.72 |
ENSMUST00000032169.8
|
Abtb1
|
ankyrin repeat and BTB (POZ) domain containing 1 |
| chr19_+_45006552 | 0.69 |
ENSMUST00000237043.2
ENSMUST00000178087.3 |
Lzts2
|
leucine zipper, putative tumor suppressor 2 |
| chr11_+_60066681 | 0.67 |
ENSMUST00000102688.2
|
Rai1
|
retinoic acid induced 1 |
| chr13_-_21586858 | 0.64 |
ENSMUST00000117721.8
ENSMUST00000070785.16 ENSMUST00000116433.2 ENSMUST00000223831.2 ENSMUST00000116434.11 ENSMUST00000224820.2 |
Zkscan3
|
zinc finger with KRAB and SCAN domains 3 |
| chr5_+_8943943 | 0.62 |
ENSMUST00000196067.2
|
Abcb4
|
ATP-binding cassette, sub-family B (MDR/TAP), member 4 |
| chr1_+_90531183 | 0.62 |
ENSMUST00000186750.2
|
Cops8
|
COP9 signalosome subunit 8 |
| chrX_-_72703652 | 0.61 |
ENSMUST00000114472.8
|
Pnck
|
pregnancy upregulated non-ubiquitously expressed CaM kinase |
| chr19_-_5416626 | 0.60 |
ENSMUST00000237167.2
|
Banf1
|
BAF nuclear assembly factor 1 |
| chr7_+_23781311 | 0.59 |
ENSMUST00000207002.2
ENSMUST00000068975.6 ENSMUST00000203854.3 |
Zfp180
|
zinc finger protein 180 |
| chr11_-_20199359 | 0.59 |
ENSMUST00000050611.14
ENSMUST00000109596.8 ENSMUST00000162811.2 |
Cep68
|
centrosomal protein 68 |
| chrX_-_8042129 | 0.59 |
ENSMUST00000143984.2
|
Tbc1d25
|
TBC1 domain family, member 25 |
| chr4_+_116565706 | 0.59 |
ENSMUST00000030452.13
ENSMUST00000106462.9 |
Ccdc163
|
coiled-coil domain containing 163 |
| chr12_-_69728572 | 0.57 |
ENSMUST00000183277.8
ENSMUST00000035773.14 |
Sos2
|
SOS Ras/Rho guanine nucleotide exchange factor 2 |
| chr14_-_55101505 | 0.52 |
ENSMUST00000142283.4
|
Homez
|
homeodomain leucine zipper-encoding gene |
| chr12_+_111504640 | 0.51 |
ENSMUST00000222375.2
ENSMUST00000222388.2 |
Eif5
|
eukaryotic translation initiation factor 5 |
| chr3_+_66893031 | 0.49 |
ENSMUST00000046542.13
ENSMUST00000162693.8 |
Rsrc1
|
arginine/serine-rich coiled-coil 1 |
| chr3_+_97920819 | 0.46 |
ENSMUST00000079812.8
|
Notch2
|
notch 2 |
| chr11_+_113539995 | 0.46 |
ENSMUST00000018805.15
|
Cog1
|
component of oligomeric golgi complex 1 |
| chr8_-_35432783 | 0.45 |
ENSMUST00000033929.6
|
Tnks
|
tankyrase, TRF1-interacting ankyrin-related ADP-ribose polymerase |
| chr1_-_135241429 | 0.45 |
ENSMUST00000134088.3
ENSMUST00000081104.10 |
Timm17a
|
translocase of inner mitochondrial membrane 17a |
| chr2_+_180240015 | 0.45 |
ENSMUST00000103059.2
|
Col9a3
|
collagen, type IX, alpha 3 |
| chr16_+_12927547 | 0.44 |
ENSMUST00000023206.14
|
Ercc4
|
excision repair cross-complementing rodent repair deficiency, complementation group 4 |
| chr11_-_62348599 | 0.44 |
ENSMUST00000127471.9
|
Ncor1
|
nuclear receptor co-repressor 1 |
| chr16_+_91282183 | 0.41 |
ENSMUST00000129878.2
|
Ifnar1
|
interferon (alpha and beta) receptor 1 |
| chr7_+_140774962 | 0.41 |
ENSMUST00000047093.11
|
Lrrc56
|
leucine rich repeat containing 56 |
| chr11_+_65698001 | 0.40 |
ENSMUST00000071465.9
ENSMUST00000018491.8 |
Zkscan6
|
zinc finger with KRAB and SCAN domains 6 |
| chr15_+_99870714 | 0.37 |
ENSMUST00000230956.2
|
Larp4
|
La ribonucleoprotein domain family, member 4 |
| chr18_-_13074834 | 0.35 |
ENSMUST00000122175.8
ENSMUST00000142467.2 ENSMUST00000074352.11 |
Osbpl1a
|
oxysterol binding protein-like 1A |
| chr18_+_11790888 | 0.35 |
ENSMUST00000234499.2
|
Rbbp8
|
retinoblastoma binding protein 8, endonuclease |
| chr1_-_4563821 | 0.35 |
ENSMUST00000191939.2
|
Sox17
|
SRY (sex determining region Y)-box 17 |
| chr9_+_55949141 | 0.35 |
ENSMUST00000114276.3
|
Rcn2
|
reticulocalbin 2 |
| chr1_-_95595280 | 0.34 |
ENSMUST00000043336.11
|
St8sia4
|
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 4 |
| chr12_-_21423524 | 0.34 |
ENSMUST00000232107.2
|
Adam17
|
a disintegrin and metallopeptidase domain 17 |
| chr18_-_84607615 | 0.33 |
ENSMUST00000125763.3
|
Zfp407
|
zinc finger protein 407 |
| chr17_-_33007238 | 0.33 |
ENSMUST00000159086.10
|
Zfp871
|
zinc finger protein 871 |
| chr9_-_78350486 | 0.32 |
ENSMUST00000070742.14
ENSMUST00000034898.14 |
Cgas
|
cyclic GMP-AMP synthase |
| chr8_+_84626715 | 0.30 |
ENSMUST00000141158.8
|
Adgrl1
|
adhesion G protein-coupled receptor L1 |
| chr16_-_22258469 | 0.29 |
ENSMUST00000079601.13
|
Etv5
|
ets variant 5 |
| chr12_-_100865783 | 0.28 |
ENSMUST00000053668.10
|
Gpr68
|
G protein-coupled receptor 68 |
| chr18_+_37637317 | 0.27 |
ENSMUST00000052179.8
|
Pcdhb20
|
protocadherin beta 20 |
| chr17_+_28075495 | 0.25 |
ENSMUST00000233809.2
|
Uhrf1bp1
|
UHRF1 (ICBP90) binding protein 1 |
| chr2_+_156154219 | 0.23 |
ENSMUST00000037096.9
|
Cnbd2
|
cyclic nucleotide binding domain containing 2 |
| chr10_+_43354807 | 0.23 |
ENSMUST00000167488.9
|
Bend3
|
BEN domain containing 3 |
| chr18_+_37544717 | 0.21 |
ENSMUST00000051126.4
|
Pcdhb10
|
protocadherin beta 10 |
| chr18_+_47245204 | 0.21 |
ENSMUST00000234633.2
|
Hspe1-rs1
|
heat shock protein 1 (chaperonin 10), related sequence 1 |
| chr2_+_34999497 | 0.17 |
ENSMUST00000028235.11
ENSMUST00000156933.8 ENSMUST00000028237.15 ENSMUST00000113032.8 |
Cntrl
|
centriolin |
| chr5_-_39801940 | 0.17 |
ENSMUST00000152057.2
ENSMUST00000053116.7 |
Hs3st1
|
heparan sulfate (glucosamine) 3-O-sulfotransferase 1 |
| chr6_+_71808427 | 0.15 |
ENSMUST00000171057.2
|
Immt
|
inner membrane protein, mitochondrial |
| chr2_-_120183575 | 0.15 |
ENSMUST00000028752.8
ENSMUST00000102501.10 |
Vps39
|
VPS39 HOPS complex subunit |
| chr3_+_41697046 | 0.14 |
ENSMUST00000120167.8
ENSMUST00000108065.9 ENSMUST00000146165.8 ENSMUST00000192193.6 ENSMUST00000119572.8 ENSMUST00000026867.14 ENSMUST00000026868.13 |
D3Ertd751e
|
DNA segment, Chr 3, ERATO Doi 751, expressed |
| chrX_+_134739783 | 0.14 |
ENSMUST00000173804.8
ENSMUST00000113136.8 |
Gprasp2
|
G protein-coupled receptor associated sorting protein 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.2 | 12.5 | GO:0010752 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) regulation of cGMP-mediated signaling(GO:0010752) |
| 3.1 | 9.4 | GO:0045660 | positive regulation of neutrophil differentiation(GO:0045660) |
| 2.9 | 14.4 | GO:0050902 | leukocyte adhesive activation(GO:0050902) |
| 2.6 | 10.5 | GO:2000373 | regulation of DNA topoisomerase (ATP-hydrolyzing) activity(GO:2000371) positive regulation of DNA topoisomerase (ATP-hydrolyzing) activity(GO:2000373) |
| 2.2 | 2.2 | GO:0036388 | pre-replicative complex assembly involved in nuclear cell cycle DNA replication(GO:0006267) pre-replicative complex assembly(GO:0036388) pre-replicative complex assembly involved in cell cycle DNA replication(GO:1902299) |
| 2.2 | 6.5 | GO:0036034 | mediator complex assembly(GO:0036034) DNA replication preinitiation complex assembly(GO:0071163) response to sorbitol(GO:0072708) regulation of mediator complex assembly(GO:2001176) positive regulation of mediator complex assembly(GO:2001178) |
| 2.0 | 8.0 | GO:0071963 | establishment or maintenance of cell polarity regulating cell shape(GO:0071963) |
| 1.7 | 8.3 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 1.5 | 7.4 | GO:0048478 | replication fork protection(GO:0048478) |
| 1.3 | 11.4 | GO:1902969 | mitotic DNA replication(GO:1902969) |
| 1.2 | 4.7 | GO:1904046 | negative regulation of vascular endothelial growth factor production(GO:1904046) |
| 1.1 | 25.5 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 1.1 | 3.2 | GO:0031990 | mRNA export from nucleus in response to heat stress(GO:0031990) |
| 1.0 | 3.1 | GO:0060821 | inactivation of X chromosome by DNA methylation(GO:0060821) |
| 0.8 | 4.1 | GO:2000620 | positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 0.7 | 3.0 | GO:0006168 | adenine salvage(GO:0006168) adenine metabolic process(GO:0046083) adenine biosynthetic process(GO:0046084) |
| 0.7 | 6.7 | GO:0061092 | regulation of phospholipid translocation(GO:0061091) positive regulation of phospholipid translocation(GO:0061092) |
| 0.7 | 2.8 | GO:1900220 | semaphorin-plexin signaling pathway involved in bone trabecula morphogenesis(GO:1900220) |
| 0.7 | 2.0 | GO:0030200 | heparan sulfate proteoglycan catabolic process(GO:0030200) |
| 0.5 | 5.6 | GO:0075509 | receptor-mediated endocytosis of virus by host cell(GO:0019065) endocytosis involved in viral entry into host cell(GO:0075509) |
| 0.5 | 3.0 | GO:1902963 | regulation of metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902962) negative regulation of metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902963) |
| 0.5 | 1.9 | GO:0021993 | initiation of neural tube closure(GO:0021993) |
| 0.5 | 1.9 | GO:0060697 | positive regulation of phospholipid catabolic process(GO:0060697) |
| 0.5 | 1.8 | GO:0071034 | CUT catabolic process(GO:0071034) CUT metabolic process(GO:0071043) |
| 0.4 | 3.8 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.4 | 1.3 | GO:0019085 | early viral transcription(GO:0019085) |
| 0.4 | 1.7 | GO:1901491 | negative regulation of lymphangiogenesis(GO:1901491) |
| 0.4 | 7.1 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.4 | 2.0 | GO:0032747 | positive regulation of interleukin-23 production(GO:0032747) |
| 0.4 | 1.6 | GO:2001245 | regulation of phosphatidylcholine biosynthetic process(GO:2001245) |
| 0.4 | 2.4 | GO:0055099 | response to high density lipoprotein particle(GO:0055099) |
| 0.4 | 6.4 | GO:0015809 | arginine transport(GO:0015809) |
| 0.4 | 2.2 | GO:0042758 | long-chain fatty acid catabolic process(GO:0042758) |
| 0.3 | 3.1 | GO:0010792 | DNA double-strand break processing involved in repair via single-strand annealing(GO:0010792) |
| 0.3 | 7.9 | GO:0006656 | phosphatidylcholine biosynthetic process(GO:0006656) |
| 0.3 | 1.7 | GO:0006420 | arginyl-tRNA aminoacylation(GO:0006420) |
| 0.3 | 1.2 | GO:1901675 | negative regulation of histone H3-K27 acetylation(GO:1901675) |
| 0.3 | 0.9 | GO:0061511 | centriole elongation(GO:0061511) |
| 0.3 | 7.1 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.2 | 2.4 | GO:0021995 | anterior neuropore closure(GO:0021506) neuropore closure(GO:0021995) |
| 0.2 | 1.2 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 0.2 | 4.7 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.2 | 0.5 | GO:1990705 | cholangiocyte proliferation(GO:1990705) |
| 0.2 | 2.7 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.2 | 0.8 | GO:0014057 | positive regulation of acetylcholine secretion, neurotransmission(GO:0014057) |
| 0.2 | 1.3 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.2 | 0.8 | GO:0001560 | regulation of cell growth by extracellular stimulus(GO:0001560) |
| 0.2 | 1.9 | GO:0044791 | modulation by host of viral release from host cell(GO:0044789) positive regulation by host of viral release from host cell(GO:0044791) |
| 0.2 | 0.6 | GO:0010387 | COP9 signalosome assembly(GO:0010387) |
| 0.2 | 2.9 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.2 | 0.6 | GO:0019043 | establishment of viral latency(GO:0019043) |
| 0.2 | 1.3 | GO:1904907 | negative regulation of maintenance of sister chromatid cohesion(GO:0034092) negative regulation of maintenance of mitotic sister chromatid cohesion(GO:0034183) maintenance of mitotic sister chromatid cohesion, telomeric(GO:0099403) mitotic sister chromatid cohesion, telomeric(GO:0099404) regulation of maintenance of mitotic sister chromatid cohesion, telomeric(GO:1904907) negative regulation of maintenance of mitotic sister chromatid cohesion, telomeric(GO:1904908) |
| 0.2 | 13.8 | GO:0010718 | positive regulation of epithelial to mesenchymal transition(GO:0010718) |
| 0.2 | 2.2 | GO:0090043 | regulation of tubulin deacetylation(GO:0090043) |
| 0.2 | 1.3 | GO:0015808 | L-alanine transport(GO:0015808) |
| 0.2 | 2.8 | GO:1901673 | regulation of mitotic spindle assembly(GO:1901673) |
| 0.2 | 1.9 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.2 | 1.7 | GO:0019985 | translesion synthesis(GO:0019985) |
| 0.2 | 3.1 | GO:0046599 | regulation of centriole replication(GO:0046599) |
| 0.2 | 1.1 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 0.1 | 0.4 | GO:0006296 | nucleotide-excision repair, DNA incision, 5'-to lesion(GO:0006296) |
| 0.1 | 3.6 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.1 | 2.1 | GO:0033120 | positive regulation of RNA splicing(GO:0033120) |
| 0.1 | 3.7 | GO:0009251 | glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.1 | 1.4 | GO:0034244 | negative regulation of transcription elongation from RNA polymerase II promoter(GO:0034244) |
| 0.1 | 1.5 | GO:0035457 | cellular response to interferon-alpha(GO:0035457) |
| 0.1 | 5.2 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.1 | 4.2 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.1 | 7.5 | GO:0000266 | mitochondrial fission(GO:0000266) |
| 0.1 | 2.7 | GO:0072520 | seminiferous tubule development(GO:0072520) |
| 0.1 | 0.4 | GO:0072362 | regulation of glycolytic process by negative regulation of transcription from RNA polymerase II promoter(GO:0072362) |
| 0.1 | 1.4 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.1 | 1.3 | GO:0060923 | cardiac muscle cell fate commitment(GO:0060923) |
| 0.1 | 0.6 | GO:2000973 | regulation of pro-B cell differentiation(GO:2000973) |
| 0.1 | 0.8 | GO:0048633 | positive regulation of skeletal muscle tissue growth(GO:0048633) |
| 0.1 | 1.7 | GO:0030644 | cellular chloride ion homeostasis(GO:0030644) |
| 0.1 | 0.9 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.1 | 0.3 | GO:0060807 | cardiogenic plate morphogenesis(GO:0003142) regulation of transcription from RNA polymerase II promoter involved in definitive endodermal cell fate specification(GO:0060807) regulation of cardiac cell fate specification(GO:2000043) |
| 0.1 | 1.1 | GO:0071044 | histone mRNA catabolic process(GO:0071044) |
| 0.1 | 1.6 | GO:0071243 | cellular response to arsenic-containing substance(GO:0071243) |
| 0.1 | 1.9 | GO:0031290 | retinal ganglion cell axon guidance(GO:0031290) |
| 0.1 | 0.9 | GO:0001955 | blood vessel maturation(GO:0001955) |
| 0.1 | 3.6 | GO:0048246 | macrophage chemotaxis(GO:0048246) |
| 0.1 | 1.0 | GO:0033148 | positive regulation of intracellular estrogen receptor signaling pathway(GO:0033148) |
| 0.1 | 0.3 | GO:2001206 | positive regulation of osteoclast development(GO:2001206) |
| 0.0 | 3.4 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 0.0 | 0.7 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.0 | 5.3 | GO:0048024 | regulation of mRNA splicing, via spliceosome(GO:0048024) |
| 0.0 | 0.6 | GO:0010457 | centriole-centriole cohesion(GO:0010457) |
| 0.0 | 0.6 | GO:1901096 | regulation of autophagosome maturation(GO:1901096) |
| 0.0 | 0.0 | GO:0035802 | adrenal cortex development(GO:0035801) adrenal cortex formation(GO:0035802) |
| 0.0 | 0.1 | GO:1902774 | late endosome to lysosome transport(GO:1902774) |
| 0.0 | 0.3 | GO:0042078 | germ-line stem cell division(GO:0042078) male germ-line stem cell asymmetric division(GO:0048133) germline stem cell asymmetric division(GO:0098728) |
| 0.0 | 0.8 | GO:0060707 | trophoblast giant cell differentiation(GO:0060707) |
| 0.0 | 0.1 | GO:0060023 | soft palate development(GO:0060023) |
| 0.0 | 0.4 | GO:2000773 | negative regulation of cellular senescence(GO:2000773) |
| 0.0 | 0.4 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.0 | 1.1 | GO:0090162 | establishment of epithelial cell polarity(GO:0090162) |
| 0.0 | 2.9 | GO:0045727 | positive regulation of translation(GO:0045727) |
| 0.0 | 2.2 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.0 | 1.0 | GO:0001755 | neural crest cell migration(GO:0001755) |
| 0.0 | 1.0 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.0 | 0.7 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 0.0 | 0.1 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.0 | 0.7 | GO:0006414 | translational elongation(GO:0006414) |
| 0.0 | 1.9 | GO:0008033 | tRNA processing(GO:0008033) |
| 0.0 | 0.3 | GO:0090129 | positive regulation of synapse maturation(GO:0090129) |
| 0.0 | 0.9 | GO:0003341 | cilium movement(GO:0003341) axoneme assembly(GO:0035082) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.2 | 25.5 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 2.2 | 2.2 | GO:0031261 | nuclear pre-replicative complex(GO:0005656) DNA replication preinitiation complex(GO:0031261) pre-replicative complex(GO:0036387) |
| 2.1 | 10.5 | GO:0000799 | nuclear condensin complex(GO:0000799) |
| 1.7 | 8.3 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 1.5 | 7.4 | GO:0031298 | replication fork protection complex(GO:0031298) |
| 1.4 | 12.5 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 1.2 | 3.5 | GO:0000811 | GINS complex(GO:0000811) |
| 0.9 | 5.2 | GO:0005683 | U7 snRNP(GO:0005683) |
| 0.8 | 14.4 | GO:0001931 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.8 | 3.1 | GO:0001740 | Barr body(GO:0001740) |
| 0.6 | 7.1 | GO:0097451 | glial limiting end-foot(GO:0097451) |
| 0.6 | 10.2 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.6 | 2.8 | GO:0061673 | mitotic spindle astral microtubule(GO:0061673) |
| 0.5 | 3.8 | GO:0000796 | condensin complex(GO:0000796) |
| 0.4 | 2.2 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.4 | 3.1 | GO:0005638 | lamin filament(GO:0005638) |
| 0.4 | 14.5 | GO:0030125 | clathrin vesicle coat(GO:0030125) |
| 0.4 | 4.7 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.4 | 2.9 | GO:0036396 | MIS complex(GO:0036396) |
| 0.4 | 4.1 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 0.4 | 2.2 | GO:0000120 | RNA polymerase I transcription factor complex(GO:0000120) |
| 0.3 | 1.4 | GO:0032021 | NELF complex(GO:0032021) |
| 0.3 | 2.7 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.3 | 1.9 | GO:0034991 | nuclear meiotic cohesin complex(GO:0034991) |
| 0.3 | 1.0 | GO:0043259 | laminin-10 complex(GO:0043259) |
| 0.3 | 3.6 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.3 | 6.8 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.2 | 1.7 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.2 | 0.4 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 0.2 | 2.9 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.2 | 3.8 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.2 | 2.7 | GO:0042555 | MCM complex(GO:0042555) |
| 0.2 | 1.8 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.1 | 1.4 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.1 | 1.3 | GO:0008024 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) |
| 0.1 | 7.8 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 1.0 | GO:0000445 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.1 | 0.4 | GO:0005594 | collagen type IX trimer(GO:0005594) |
| 0.1 | 3.1 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.1 | 2.4 | GO:0090545 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.1 | 0.9 | GO:0071439 | clathrin complex(GO:0071439) |
| 0.1 | 1.3 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.1 | 1.1 | GO:0016442 | RISC complex(GO:0016442) RNAi effector complex(GO:0031332) |
| 0.1 | 6.6 | GO:0005776 | autophagosome(GO:0005776) |
| 0.1 | 0.4 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.1 | 4.2 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.1 | 4.3 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 6.5 | GO:0000776 | kinetochore(GO:0000776) |
| 0.0 | 0.6 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.0 | 0.8 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 0.0 | 1.4 | GO:0071011 | precatalytic spliceosome(GO:0071011) |
| 0.0 | 0.1 | GO:0030123 | AP-3 adaptor complex(GO:0030123) |
| 0.0 | 0.9 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 0.6 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 3.1 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 8.8 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 1.9 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.0 | 7.8 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 3.7 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.0 | 0.6 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 1.6 | GO:0016528 | sarcoplasm(GO:0016528) |
| 0.0 | 1.3 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 4.7 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 1.4 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 1.6 | GO:0045111 | intermediate filament cytoskeleton(GO:0045111) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.2 | 25.5 | GO:0004748 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 3.1 | 12.5 | GO:0070052 | collagen V binding(GO:0070052) |
| 1.5 | 10.5 | GO:0044547 | DNA topoisomerase binding(GO:0044547) |
| 1.0 | 3.0 | GO:0002055 | adenine binding(GO:0002055) adenine phosphoribosyltransferase activity(GO:0003999) |
| 0.9 | 3.7 | GO:0008184 | glycogen phosphorylase activity(GO:0008184) |
| 0.9 | 2.7 | GO:0004637 | phosphoribosylamine-glycine ligase activity(GO:0004637) |
| 0.9 | 3.6 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.9 | 7.1 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.8 | 7.8 | GO:0003910 | DNA ligase activity(GO:0003909) DNA ligase (ATP) activity(GO:0003910) |
| 0.7 | 6.4 | GO:0015181 | arginine transmembrane transporter activity(GO:0015181) |
| 0.6 | 2.2 | GO:0004466 | long-chain-acyl-CoA dehydrogenase activity(GO:0004466) |
| 0.5 | 1.9 | GO:0004699 | calcium-independent protein kinase C activity(GO:0004699) TIR domain binding(GO:0070976) |
| 0.5 | 1.9 | GO:0036033 | mediator complex binding(GO:0036033) |
| 0.5 | 3.2 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.4 | 1.3 | GO:0005302 | L-tyrosine transmembrane transporter activity(GO:0005302) |
| 0.3 | 1.7 | GO:0004814 | arginine-tRNA ligase activity(GO:0004814) |
| 0.3 | 1.9 | GO:0004720 | protein-lysine 6-oxidase activity(GO:0004720) |
| 0.3 | 4.2 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.3 | 3.6 | GO:0001055 | RNA polymerase II activity(GO:0001055) |
| 0.3 | 3.7 | GO:0098505 | G-rich strand telomeric DNA binding(GO:0098505) |
| 0.3 | 3.6 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) |
| 0.2 | 9.4 | GO:0008187 | poly-pyrimidine tract binding(GO:0008187) |
| 0.2 | 8.3 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.2 | 1.7 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.2 | 2.4 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.2 | 2.0 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.2 | 1.1 | GO:0004534 | 5'-3' exoribonuclease activity(GO:0004534) |
| 0.2 | 3.5 | GO:0043138 | 3'-5' DNA helicase activity(GO:0043138) |
| 0.2 | 1.4 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.2 | 1.0 | GO:0004809 | tRNA (guanine-N2-)-methyltransferase activity(GO:0004809) |
| 0.2 | 2.0 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.2 | 3.0 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.2 | 6.7 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.2 | 0.8 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 0.2 | 1.6 | GO:0070538 | oleic acid binding(GO:0070538) |
| 0.1 | 1.9 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 2.8 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.1 | 2.2 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.1 | 2.2 | GO:0001163 | RNA polymerase I regulatory region sequence-specific DNA binding(GO:0001163) RNA polymerase I CORE element sequence-specific DNA binding(GO:0001164) |
| 0.1 | 2.9 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.1 | 7.9 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.1 | 0.8 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.1 | 4.7 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.1 | 1.3 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
| 0.1 | 2.7 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 3.2 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) |
| 0.1 | 0.9 | GO:0008479 | queuine tRNA-ribosyltransferase activity(GO:0008479) |
| 0.1 | 1.6 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.1 | 6.9 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.1 | 2.7 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.1 | 7.1 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.1 | 2.8 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.1 | 0.8 | GO:0001609 | G-protein coupled adenosine receptor activity(GO:0001609) |
| 0.1 | 4.8 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.0 | 0.4 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.0 | 0.4 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.0 | 1.3 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.0 | 4.4 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.0 | 0.1 | GO:0016434 | rRNA (cytosine) methyltransferase activity(GO:0016434) |
| 0.0 | 3.5 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 0.0 | 1.4 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 2.2 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.0 | 0.7 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.0 | 1.2 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.0 | 1.6 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.0 | 0.3 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.0 | 0.5 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.5 | GO:0043274 | phospholipase binding(GO:0043274) |
| 0.0 | 1.0 | GO:0030331 | estrogen receptor binding(GO:0030331) |
| 0.0 | 1.2 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 1.4 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 0.5 | GO:0051059 | NF-kappaB binding(GO:0051059) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 14.4 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.3 | 10.8 | PID ATM PATHWAY | ATM pathway |
| 0.3 | 14.3 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.2 | 11.1 | PID ATR PATHWAY | ATR signaling pathway |
| 0.2 | 14.4 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.2 | 20.0 | PID E2F PATHWAY | E2F transcription factor network |
| 0.1 | 1.5 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.1 | 1.4 | ST JAK STAT PATHWAY | Jak-STAT Pathway |
| 0.1 | 7.9 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.1 | 3.1 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.1 | 4.9 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.1 | 7.3 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.1 | 5.6 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.1 | 1.0 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.1 | 1.3 | ST INTERLEUKIN 4 PATHWAY | Interleukin 4 (IL-4) Pathway |
| 0.1 | 3.8 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 2.4 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.0 | 3.1 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 1.6 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 1.6 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.0 | 0.9 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 0.9 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 0.9 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.0 | 2.8 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.3 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.0 | 0.9 | PID NOTCH PATHWAY | Notch signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 29.1 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.9 | 17.3 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.6 | 8.3 | REACTOME REPAIR SYNTHESIS FOR GAP FILLING BY DNA POL IN TC NER | Genes involved in Repair synthesis for gap-filling by DNA polymerase in TC-NER |
| 0.4 | 6.3 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.4 | 5.2 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.3 | 6.8 | REACTOME VIRAL MESSENGER RNA SYNTHESIS | Genes involved in Viral Messenger RNA Synthesis |
| 0.3 | 8.0 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.2 | 2.2 | REACTOME E2F ENABLED INHIBITION OF PRE REPLICATION COMPLEX FORMATION | Genes involved in E2F-enabled inhibition of pre-replication complex formation |
| 0.2 | 2.9 | REACTOME RECEPTOR LIGAND BINDING INITIATES THE SECOND PROTEOLYTIC CLEAVAGE OF NOTCH RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 0.2 | 3.7 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.2 | 1.8 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.1 | 5.6 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.1 | 2.2 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 3.0 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 1.1 | REACTOME DESTABILIZATION OF MRNA BY BRF1 | Genes involved in Destabilization of mRNA by Butyrate Response Factor 1 (BRF1) |
| 0.1 | 1.5 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.1 | 8.5 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.1 | 13.5 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.1 | 3.5 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.1 | 1.3 | REACTOME ELONGATION ARREST AND RECOVERY | Genes involved in Elongation arrest and recovery |
| 0.1 | 6.7 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.1 | 0.8 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.1 | 2.2 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |
| 0.1 | 0.6 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.1 | 2.0 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.1 | 2.8 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 16.5 | REACTOME TRANSLATION | Genes involved in Translation |
| 0.1 | 1.7 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.1 | 1.2 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.1 | 3.5 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 1.8 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.1 | 2.1 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 1.4 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.1 | 3.4 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.1 | 1.9 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.6 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 3.0 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 3.2 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 0.4 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 1.2 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |