Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
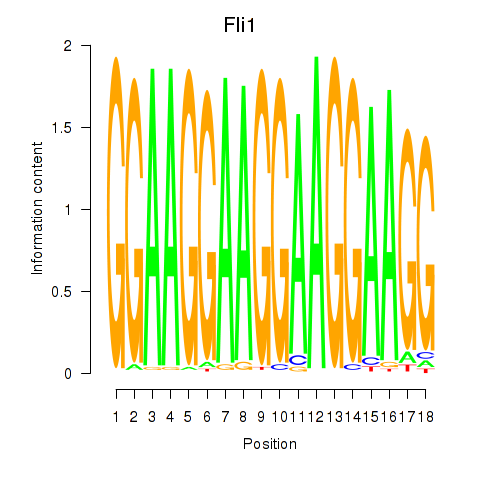
Results for Fli1
Z-value: 3.12
Transcription factors associated with Fli1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Fli1
|
ENSMUSG00000016087.14 | Fli1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Fli1 | mm39_v1_chr9_-_32452885_32452898 | -0.65 | 1.7e-05 | Click! |
Activity profile of Fli1 motif
Sorted Z-values of Fli1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Fli1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_-_40175709 | 54.83 |
ENSMUST00000051846.13
|
Cyp2c70
|
cytochrome P450, family 2, subfamily c, polypeptide 70 |
| chr9_-_57590926 | 40.45 |
ENSMUST00000034860.5
|
Cyp1a2
|
cytochrome P450, family 1, subfamily a, polypeptide 2 |
| chr8_-_25066313 | 33.13 |
ENSMUST00000121992.2
|
Ido2
|
indoleamine 2,3-dioxygenase 2 |
| chr5_-_89583469 | 25.48 |
ENSMUST00000200534.2
|
Gc
|
vitamin D binding protein |
| chr7_-_26638802 | 25.17 |
ENSMUST00000170227.3
|
Cyp2a22
|
cytochrome P450, family 2, subfamily a, polypeptide 22 |
| chr7_-_13571334 | 16.88 |
ENSMUST00000108522.5
|
Sult2a1
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 1 |
| chr17_-_32643131 | 16.80 |
ENSMUST00000236386.2
|
Pglyrp2
|
peptidoglycan recognition protein 2 |
| chr17_-_32643067 | 16.71 |
ENSMUST00000237130.2
|
Pglyrp2
|
peptidoglycan recognition protein 2 |
| chr13_-_4573312 | 11.53 |
ENSMUST00000221564.2
ENSMUST00000078239.5 ENSMUST00000080361.13 |
Akr1c20
|
aldo-keto reductase family 1, member C20 |
| chr11_-_5865124 | 8.69 |
ENSMUST00000109823.9
ENSMUST00000109822.8 |
Gck
|
glucokinase |
| chr15_-_96918203 | 8.12 |
ENSMUST00000166223.2
|
Slc38a4
|
solute carrier family 38, member 4 |
| chr19_+_46120327 | 7.76 |
ENSMUST00000043739.6
ENSMUST00000237098.2 |
Elovl3
|
elongation of very long chain fatty acids (FEN1/Elo2, SUR4/Elo3, yeast)-like 3 |
| chr7_+_13467422 | 7.75 |
ENSMUST00000086148.8
|
Sult2a2
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 2 |
| chr2_+_58644922 | 7.18 |
ENSMUST00000059102.13
|
Upp2
|
uridine phosphorylase 2 |
| chr6_-_146536025 | 6.87 |
ENSMUST00000037709.16
|
Tm7sf3
|
transmembrane 7 superfamily member 3 |
| chr5_-_73496291 | 6.71 |
ENSMUST00000200830.4
ENSMUST00000201908.4 ENSMUST00000200776.4 ENSMUST00000087195.9 |
Ociad2
|
OCIA domain containing 2 |
| chr13_+_4484305 | 6.71 |
ENSMUST00000021630.15
|
Akr1c6
|
aldo-keto reductase family 1, member C6 |
| chr17_+_79359617 | 6.33 |
ENSMUST00000233916.2
|
Qpct
|
glutaminyl-peptide cyclotransferase (glutaminyl cyclase) |
| chr11_+_16702203 | 6.31 |
ENSMUST00000102884.10
ENSMUST00000020329.13 |
Egfr
|
epidermal growth factor receptor |
| chr7_-_48493388 | 6.17 |
ENSMUST00000167786.4
|
Csrp3
|
cysteine and glycine-rich protein 3 |
| chr9_+_111268131 | 5.56 |
ENSMUST00000111879.5
|
Dclk3
|
doublecortin-like kinase 3 |
| chr15_-_96917804 | 5.45 |
ENSMUST00000231039.2
|
Slc38a4
|
solute carrier family 38, member 4 |
| chr5_-_73495895 | 5.19 |
ENSMUST00000202012.2
|
Ociad2
|
OCIA domain containing 2 |
| chr17_+_29077385 | 5.17 |
ENSMUST00000056866.8
|
Pnpla1
|
patatin-like phospholipase domain containing 1 |
| chr15_+_25940781 | 5.09 |
ENSMUST00000227275.2
|
Retreg1
|
reticulophagy regulator 1 |
| chr15_-_76010736 | 4.83 |
ENSMUST00000054022.12
ENSMUST00000089654.4 |
BC024139
|
cDNA sequence BC024139 |
| chr16_+_37400590 | 4.80 |
ENSMUST00000159787.8
|
Hgd
|
homogentisate 1, 2-dioxygenase |
| chr16_+_37400500 | 4.55 |
ENSMUST00000160847.2
|
Hgd
|
homogentisate 1, 2-dioxygenase |
| chr6_+_138117295 | 4.42 |
ENSMUST00000008684.11
|
Mgst1
|
microsomal glutathione S-transferase 1 |
| chr10_-_93375832 | 4.40 |
ENSMUST00000016034.3
|
Amdhd1
|
amidohydrolase domain containing 1 |
| chr6_+_138117519 | 4.30 |
ENSMUST00000120939.8
ENSMUST00000204628.3 ENSMUST00000140932.2 ENSMUST00000120302.8 |
Mgst1
|
microsomal glutathione S-transferase 1 |
| chr6_+_124470053 | 3.84 |
ENSMUST00000049124.10
|
C1rl
|
complement component 1, r subcomponent-like |
| chr9_+_78137927 | 3.80 |
ENSMUST00000098537.4
|
Gsta1
|
glutathione S-transferase, alpha 1 (Ya) |
| chr18_-_20192535 | 3.71 |
ENSMUST00000075214.9
ENSMUST00000039247.11 |
Dsc2
|
desmocollin 2 |
| chrX_-_135769285 | 3.68 |
ENSMUST00000058814.7
|
Rab9b
|
RAB9B, member RAS oncogene family |
| chr8_-_3770642 | 3.48 |
ENSMUST00000062037.7
|
Clec4g
|
C-type lectin domain family 4, member g |
| chr7_-_80055168 | 3.43 |
ENSMUST00000107362.10
ENSMUST00000135306.3 |
Furin
|
furin (paired basic amino acid cleaving enzyme) |
| chr6_-_119394634 | 3.02 |
ENSMUST00000032272.13
|
Adipor2
|
adiponectin receptor 2 |
| chr10_-_127457001 | 2.94 |
ENSMUST00000049149.15
|
Lrp1
|
low density lipoprotein receptor-related protein 1 |
| chr4_+_40722911 | 2.78 |
ENSMUST00000164233.8
ENSMUST00000137246.8 ENSMUST00000125442.8 |
Dnaja1
|
DnaJ heat shock protein family (Hsp40) member A1 |
| chr7_-_15978077 | 2.49 |
ENSMUST00000238893.2
ENSMUST00000171425.4 |
C5ar2
|
complement component 5a receptor 2 |
| chr4_+_148686985 | 2.46 |
ENSMUST00000105701.9
ENSMUST00000052060.7 |
Masp2
|
mannan-binding lectin serine peptidase 2 |
| chr17_-_45906768 | 2.39 |
ENSMUST00000164618.8
ENSMUST00000097317.10 ENSMUST00000170113.8 |
Slc29a1
|
solute carrier family 29 (nucleoside transporters), member 1 |
| chr14_+_53791444 | 2.35 |
ENSMUST00000198297.2
|
Trav14-1
|
T cell receptor alpha variable 14-1 |
| chr9_+_44966464 | 2.35 |
ENSMUST00000114664.8
|
Mpzl3
|
myelin protein zero-like 3 |
| chr8_-_71938598 | 2.30 |
ENSMUST00000093450.6
ENSMUST00000213382.2 |
Ano8
|
anoctamin 8 |
| chr12_+_87561085 | 2.28 |
ENSMUST00000110152.3
|
Eif1ad8
|
eukaryotic translation initiation factor 1A domain containing 8 |
| chr2_-_86940289 | 2.22 |
ENSMUST00000215828.3
|
Olfr259
|
olfactory receptor 259 |
| chr7_-_19556612 | 2.10 |
ENSMUST00000120537.8
|
Bcl3
|
B cell leukemia/lymphoma 3 |
| chr10_-_127456791 | 2.09 |
ENSMUST00000118455.2
ENSMUST00000121829.8 |
Lrp1
|
low density lipoprotein receptor-related protein 1 |
| chr17_-_31363245 | 2.07 |
ENSMUST00000024826.8
|
Tff2
|
trefoil factor 2 (spasmolytic protein 1) |
| chr7_-_140001552 | 2.07 |
ENSMUST00000213801.2
|
Olfr532
|
olfactory receptor 532 |
| chr11_-_100650566 | 2.02 |
ENSMUST00000107361.9
|
Kcnh4
|
potassium voltage-gated channel, subfamily H (eag-related), member 4 |
| chr9_-_64928927 | 2.02 |
ENSMUST00000036615.7
|
Hacd3
|
3-hydroxyacyl-CoA dehydratase 3 |
| chr5_+_43829631 | 1.97 |
ENSMUST00000125866.4
|
Cc2d2a
|
coiled-coil and C2 domain containing 2A |
| chr6_-_119394436 | 1.94 |
ENSMUST00000189710.2
|
Adipor2
|
adiponectin receptor 2 |
| chr18_-_75830595 | 1.89 |
ENSMUST00000165559.3
|
Ctif
|
CBP80/20-dependent translation initiation factor |
| chr6_-_57668992 | 1.85 |
ENSMUST00000053386.6
|
Pyurf
|
Pigy upstream reading frame |
| chr4_-_130031421 | 1.79 |
ENSMUST00000164887.3
|
Hcrtr1
|
hypocretin (orexin) receptor 1 |
| chrX_-_71699740 | 1.78 |
ENSMUST00000055966.13
|
Gabra3
|
gamma-aminobutyric acid (GABA) A receptor, subunit alpha 3 |
| chr15_-_76406602 | 1.69 |
ENSMUST00000096365.5
|
Scrt1
|
scratch family zinc finger 1 |
| chr14_+_53878158 | 1.59 |
ENSMUST00000179267.4
|
Trav14-2
|
T cell receptor alpha variable 14-2 |
| chr4_+_40723083 | 1.57 |
ENSMUST00000149794.2
|
Dnaja1
|
DnaJ heat shock protein family (Hsp40) member A1 |
| chr13_+_42862957 | 1.50 |
ENSMUST00000066928.12
ENSMUST00000148891.8 |
Phactr1
|
phosphatase and actin regulator 1 |
| chr7_-_6525801 | 1.48 |
ENSMUST00000213504.2
ENSMUST00000216447.2 ENSMUST00000213656.2 ENSMUST00000207820.3 |
Olfr1349
|
olfactory receptor 1349 |
| chr4_+_43406435 | 1.48 |
ENSMUST00000098106.9
ENSMUST00000139198.2 |
Rusc2
|
RUN and SH3 domain containing 2 |
| chr10_+_19810037 | 1.48 |
ENSMUST00000095806.10
ENSMUST00000120259.8 |
Map3k5
|
mitogen-activated protein kinase kinase kinase 5 |
| chr1_+_182591771 | 1.44 |
ENSMUST00000193660.6
|
Susd4
|
sushi domain containing 4 |
| chr17_-_36014863 | 1.43 |
ENSMUST00000148065.8
|
Ddr1
|
discoidin domain receptor family, member 1 |
| chr9_+_118307412 | 1.42 |
ENSMUST00000035020.15
|
Eomes
|
eomesodermin |
| chr9_+_118307918 | 1.42 |
ENSMUST00000150633.2
|
Eomes
|
eomesodermin |
| chr16_+_48104098 | 1.37 |
ENSMUST00000096045.9
ENSMUST00000050705.4 |
Dppa4
|
developmental pluripotency associated 4 |
| chr7_-_4606104 | 1.36 |
ENSMUST00000049113.14
|
Ptprh
|
protein tyrosine phosphatase, receptor type, H |
| chr14_-_55204023 | 1.35 |
ENSMUST00000124930.8
|
Myh6
|
myosin, heavy polypeptide 6, cardiac muscle, alpha |
| chr3_-_89820451 | 1.35 |
ENSMUST00000029559.7
ENSMUST00000197679.5 |
Il6ra
|
interleukin 6 receptor, alpha |
| chr5_-_135378896 | 1.34 |
ENSMUST00000201534.2
ENSMUST00000044972.11 |
Fkbp6
|
FK506 binding protein 6 |
| chr5_+_127709302 | 1.33 |
ENSMUST00000118139.3
|
Glt1d1
|
glycosyltransferase 1 domain containing 1 |
| chr10_+_84591919 | 1.31 |
ENSMUST00000060397.13
|
Rfx4
|
regulatory factor X, 4 (influences HLA class II expression) |
| chr14_-_55204054 | 1.31 |
ENSMUST00000226297.2
|
Myh6
|
myosin, heavy polypeptide 6, cardiac muscle, alpha |
| chr12_+_87665639 | 1.31 |
ENSMUST00000164838.3
|
Eif1ad6
|
eukaryotic translation initiation factor 1A domain containing 6 |
| chr17_-_24863956 | 1.31 |
ENSMUST00000019684.13
|
Slc9a3r2
|
solute carrier family 9 (sodium/hydrogen exchanger), member 3 regulator 2 |
| chrX_+_140258381 | 1.30 |
ENSMUST00000112931.8
ENSMUST00000112930.8 |
Col4a5
|
collagen, type IV, alpha 5 |
| chr3_+_144276154 | 1.28 |
ENSMUST00000082437.11
|
Selenof
|
selenoprotein F |
| chr6_-_77956635 | 1.26 |
ENSMUST00000161846.8
ENSMUST00000160894.8 |
Ctnna2
|
catenin (cadherin associated protein), alpha 2 |
| chr19_+_4264470 | 1.25 |
ENSMUST00000237171.2
|
Gm45928
|
predicted gene, 45928 |
| chr17_-_24863907 | 1.21 |
ENSMUST00000234505.2
|
Slc9a3r2
|
solute carrier family 9 (sodium/hydrogen exchanger), member 3 regulator 2 |
| chr2_-_121786573 | 1.21 |
ENSMUST00000104936.4
|
Mageb3
|
MAGE family member B3 |
| chr14_-_46628033 | 1.18 |
ENSMUST00000074077.12
|
Bmp4
|
bone morphogenetic protein 4 |
| chr4_+_33078796 | 1.14 |
ENSMUST00000131920.8
|
Gabrr2
|
gamma-aminobutyric acid (GABA) C receptor, subunit rho 2 |
| chr14_+_56640038 | 1.13 |
ENSMUST00000095793.3
ENSMUST00000223627.2 |
Rnf17
|
ring finger protein 17 |
| chr5_-_135378729 | 1.13 |
ENSMUST00000201784.4
ENSMUST00000201791.4 |
Fkbp6
|
FK506 binding protein 6 |
| chr14_+_55120777 | 1.12 |
ENSMUST00000022806.10
|
Bcl2l2
|
BCL2-like 2 |
| chr14_+_53404083 | 1.11 |
ENSMUST00000196639.2
ENSMUST00000177578.2 |
Trav14n-1
|
T cell receptor alpha variable 14N-1 |
| chr19_-_56378309 | 1.10 |
ENSMUST00000166203.2
ENSMUST00000167239.8 |
Nrap
|
nebulin-related anchoring protein |
| chr5_+_123390149 | 1.08 |
ENSMUST00000121964.8
|
Wdr66
|
WD repeat domain 66 |
| chr9_+_44018583 | 1.07 |
ENSMUST00000152956.8
ENSMUST00000114815.3 ENSMUST00000206295.2 ENSMUST00000206769.2 ENSMUST00000205500.2 |
C1qtnf5
|
C1q and tumor necrosis factor related protein 5 |
| chr2_+_119378178 | 1.03 |
ENSMUST00000014221.13
ENSMUST00000119172.2 |
Chp1
|
calcineurin-like EF hand protein 1 |
| chr9_+_118307250 | 1.03 |
ENSMUST00000111763.8
|
Eomes
|
eomesodermin |
| chr1_+_182591425 | 1.02 |
ENSMUST00000155229.7
ENSMUST00000153348.8 |
Susd4
|
sushi domain containing 4 |
| chr12_+_87783347 | 1.01 |
ENSMUST00000110149.2
|
Eif1ad2
|
eukaryotic translation initiation factor 1A domain containing 2 |
| chr10_+_26105605 | 0.99 |
ENSMUST00000218301.2
ENSMUST00000164660.8 ENSMUST00000060716.6 |
Samd3
|
sterile alpha motif domain containing 3 |
| chr14_-_46627957 | 0.98 |
ENSMUST00000100676.3
|
Bmp4
|
bone morphogenetic protein 4 |
| chr9_+_44018551 | 0.98 |
ENSMUST00000114821.9
ENSMUST00000114818.9 |
C1qtnf5
|
C1q and tumor necrosis factor related protein 5 |
| chr7_-_108471586 | 0.97 |
ENSMUST00000216500.2
|
Olfr517
|
olfactory receptor 517 |
| chr17_-_24428351 | 0.97 |
ENSMUST00000024931.6
|
Ntn3
|
netrin 3 |
| chr14_+_53093071 | 0.95 |
ENSMUST00000181038.3
ENSMUST00000187138.2 |
Trav14d-1
|
T cell receptor alpha variable 14D-1 |
| chr17_+_6157154 | 0.93 |
ENSMUST00000149756.8
|
Tulp4
|
tubby like protein 4 |
| chr11_+_87486472 | 0.91 |
ENSMUST00000134216.3
|
Mtmr4
|
myotubularin related protein 4 |
| chr9_+_44394080 | 0.91 |
ENSMUST00000220303.2
|
Bcl9l
|
B cell CLL/lymphoma 9-like |
| chr6_-_77956499 | 0.89 |
ENSMUST00000159626.8
ENSMUST00000075340.12 ENSMUST00000162273.2 |
Ctnna2
|
catenin (cadherin associated protein), alpha 2 |
| chr12_-_88290796 | 0.88 |
ENSMUST00000218054.3
|
Eif1ad15
|
eukaryotic translation initiation factor 1A domain containing 15 |
| chr19_+_4264437 | 0.88 |
ENSMUST00000235355.2
|
Clcf1
|
cardiotrophin-like cytokine factor 1 |
| chr10_+_84753480 | 0.84 |
ENSMUST00000038523.15
ENSMUST00000214693.2 ENSMUST00000095385.5 |
Ric8b
|
RIC8 guanine nucleotide exchange factor B |
| chr8_-_45863572 | 0.84 |
ENSMUST00000209651.2
ENSMUST00000211370.2 ENSMUST00000034056.12 ENSMUST00000167106.3 |
Tlr3
|
toll-like receptor 3 |
| chr10_+_21869776 | 0.83 |
ENSMUST00000092673.11
|
Sgk1
|
serum/glucocorticoid regulated kinase 1 |
| chr14_-_55204092 | 0.83 |
ENSMUST00000081857.14
|
Myh6
|
myosin, heavy polypeptide 6, cardiac muscle, alpha |
| chr16_-_10884005 | 0.81 |
ENSMUST00000162323.2
|
Litaf
|
LPS-induced TN factor |
| chr3_-_144275897 | 0.81 |
ENSMUST00000043325.9
|
Hs2st1
|
heparan sulfate 2-O-sulfotransferase 1 |
| chr2_-_119378108 | 0.80 |
ENSMUST00000060009.9
|
Exd1
|
exonuclease 3'-5' domain containing 1 |
| chr16_-_28571820 | 0.79 |
ENSMUST00000232352.2
|
Fgf12
|
fibroblast growth factor 12 |
| chr7_+_104506216 | 0.79 |
ENSMUST00000067695.8
|
Usp17la
|
ubiquitin specific peptidase 17-like A |
| chr1_+_54289833 | 0.79 |
ENSMUST00000027128.11
|
Ccdc150
|
coiled-coil domain containing 150 |
| chr1_+_170060318 | 0.78 |
ENSMUST00000162752.2
|
Sh2d1b2
|
SH2 domain containing 1B2 |
| chr3_+_156267429 | 0.77 |
ENSMUST00000074015.11
|
Negr1
|
neuronal growth regulator 1 |
| chr12_-_118265163 | 0.76 |
ENSMUST00000221844.2
|
Sp4
|
trans-acting transcription factor 4 |
| chr2_+_152804405 | 0.75 |
ENSMUST00000099197.9
ENSMUST00000103155.10 |
Ttll9
|
tubulin tyrosine ligase-like family, member 9 |
| chr2_+_79465696 | 0.75 |
ENSMUST00000111785.9
|
Itprid2
|
ITPR interacting domain containing 2 |
| chr19_+_59447950 | 0.73 |
ENSMUST00000174353.2
|
Emx2
|
empty spiracles homeobox 2 |
| chr5_-_66330394 | 0.73 |
ENSMUST00000201544.4
|
Rbm47
|
RNA binding motif protein 47 |
| chr6_+_57183497 | 0.72 |
ENSMUST00000227298.2
|
Vmn1r13
|
vomeronasal 1 receptor 13 |
| chr2_+_79465761 | 0.71 |
ENSMUST00000111788.8
ENSMUST00000111784.9 |
Itprid2
|
ITPR interacting domain containing 2 |
| chr12_-_115964081 | 0.71 |
ENSMUST00000103552.2
|
Ighv1-85
|
immunoglobulin heavy variable 1-85 |
| chr3_+_156267587 | 0.70 |
ENSMUST00000041425.12
ENSMUST00000106065.2 |
Negr1
|
neuronal growth regulator 1 |
| chr10_+_102348076 | 0.68 |
ENSMUST00000219445.2
|
Rassf9
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 9 |
| chr9_+_44018520 | 0.68 |
ENSMUST00000114816.8
|
C1qtnf5
|
C1q and tumor necrosis factor related protein 5 |
| chr6_-_88423464 | 0.66 |
ENSMUST00000204459.3
ENSMUST00000203213.3 ENSMUST00000205179.3 ENSMUST00000165242.4 |
Eefsec
|
eukaryotic elongation factor, selenocysteine-tRNA-specific |
| chr5_-_151157283 | 0.65 |
ENSMUST00000126770.2
|
Stard13
|
StAR-related lipid transfer (START) domain containing 13 |
| chr11_+_17207558 | 0.65 |
ENSMUST00000000594.9
ENSMUST00000156784.2 |
C1d
|
C1D nuclear receptor co-repressor |
| chr2_-_38177182 | 0.65 |
ENSMUST00000130472.8
|
Dennd1a
|
DENN/MADD domain containing 1A |
| chr10_-_3217767 | 0.63 |
ENSMUST00000170893.3
|
H60c
|
histocompatibility 60c |
| chr10_-_3217718 | 0.62 |
ENSMUST00000216211.2
|
H60c
|
histocompatibility 60c |
| chr18_+_47054867 | 0.62 |
ENSMUST00000234910.2
|
Arl14epl
|
ADP-ribosylation factor-like 14 effector protein-like |
| chr9_-_117843417 | 0.62 |
ENSMUST00000238919.2
|
Zcwpw2
|
zinc finger, CW type with PWWP domain 2 |
| chr1_+_42992109 | 0.61 |
ENSMUST00000179766.3
|
Gpr45
|
G protein-coupled receptor 45 |
| chr11_-_53525520 | 0.59 |
ENSMUST00000020650.2
|
Il13
|
interleukin 13 |
| chr1_-_126420659 | 0.59 |
ENSMUST00000161954.8
|
Nckap5
|
NCK-associated protein 5 |
| chr10_+_24471340 | 0.58 |
ENSMUST00000020171.12
|
Ccn2
|
cellular communication network factor 2 |
| chr15_+_102884874 | 0.58 |
ENSMUST00000173306.2
|
Hoxc9
|
homeobox C9 |
| chr7_+_34885782 | 0.57 |
ENSMUST00000135452.8
ENSMUST00000001854.12 |
Slc7a10
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 10 |
| chr14_-_7666348 | 0.56 |
ENSMUST00000224491.2
|
Nek10
|
NIMA (never in mitosis gene a)- related kinase 10 |
| chr3_-_129518723 | 0.56 |
ENSMUST00000199615.5
ENSMUST00000197079.5 |
Egf
|
epidermal growth factor |
| chr2_-_32976378 | 0.56 |
ENSMUST00000049618.9
|
Garnl3
|
GTPase activating RANGAP domain-like 3 |
| chr5_-_147244074 | 0.55 |
ENSMUST00000031650.4
|
Cdx2
|
caudal type homeobox 2 |
| chr4_-_150085722 | 0.54 |
ENSMUST00000153394.2
|
H6pd
|
hexose-6-phosphate dehydrogenase (glucose 1-dehydrogenase) |
| chr11_-_100583265 | 0.54 |
ENSMUST00000153494.2
|
Zfp385c
|
zinc finger protein 385C |
| chr10_-_62561865 | 0.54 |
ENSMUST00000133371.8
|
Stox1
|
storkhead box 1 |
| chr9_-_8004586 | 0.53 |
ENSMUST00000086580.12
ENSMUST00000065353.13 |
Yap1
|
yes-associated protein 1 |
| chr10_+_82190078 | 0.52 |
ENSMUST00000211099.2
|
Gm4924
|
predicted gene 4924 |
| chr7_-_104426677 | 0.51 |
ENSMUST00000211384.2
ENSMUST00000053464.9 |
Usp17le
|
ubiquitin specific peptidase 17-like E |
| chr14_-_101437750 | 0.51 |
ENSMUST00000187304.2
|
Prr30
|
proline rich 30 |
| chr15_-_98660873 | 0.50 |
ENSMUST00000156572.3
|
Arf3
|
ADP-ribosylation factor 3 |
| chr1_-_44118902 | 0.50 |
ENSMUST00000238662.2
|
Gm8251
|
predicted gene 8251 |
| chr16_+_91184661 | 0.49 |
ENSMUST00000139503.2
|
Ifnar2
|
interferon (alpha and beta) receptor 2 |
| chr4_+_42655251 | 0.49 |
ENSMUST00000177785.3
|
Ccl27b
|
chemokine (C-C motif) ligand 27b |
| chr13_-_99584091 | 0.48 |
ENSMUST00000223725.2
|
Map1b
|
microtubule-associated protein 1B |
| chr1_+_44590791 | 0.48 |
ENSMUST00000160854.8
|
Gulp1
|
GULP, engulfment adaptor PTB domain containing 1 |
| chr11_-_100244866 | 0.47 |
ENSMUST00000173630.8
|
Hap1
|
huntingtin-associated protein 1 |
| chr11_-_100650768 | 0.46 |
ENSMUST00000107363.3
|
Kcnh4
|
potassium voltage-gated channel, subfamily H (eag-related), member 4 |
| chr4_+_155874896 | 0.46 |
ENSMUST00000165000.8
|
Ankrd65
|
ankyrin repeat domain 65 |
| chr6_+_41512010 | 0.46 |
ENSMUST00000103288.2
|
Trbj1-5
|
T cell receptor beta joining 1-5 |
| chr9_-_109397316 | 0.46 |
ENSMUST00000198112.2
ENSMUST00000198397.5 ENSMUST00000056745.12 |
Fbxw15
|
F-box and WD-40 domain protein 15 |
| chr5_+_92285748 | 0.45 |
ENSMUST00000031355.10
ENSMUST00000202155.2 |
Uso1
|
USO1 vesicle docking factor |
| chr9_-_109024588 | 0.45 |
ENSMUST00000199102.2
|
Fbxw13
|
F-box and WD-40 domain protein 13 |
| chr6_+_145879839 | 0.44 |
ENSMUST00000032383.14
|
Sspn
|
sarcospan |
| chr2_+_121786892 | 0.42 |
ENSMUST00000110578.8
|
Ctdspl2
|
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase like 2 |
| chr16_+_3648742 | 0.41 |
ENSMUST00000214238.2
ENSMUST00000214590.2 |
Olfr15
|
olfactory receptor 15 |
| chrX_+_31839202 | 0.41 |
ENSMUST00000179991.3
|
Btbd35f2
|
BTB domain containing 35, family member 2 |
| chr5_-_151157233 | 0.41 |
ENSMUST00000129088.8
|
Stard13
|
StAR-related lipid transfer (START) domain containing 13 |
| chr11_+_56902658 | 0.41 |
ENSMUST00000094179.11
|
Gria1
|
glutamate receptor, ionotropic, AMPA1 (alpha 1) |
| chr2_+_121787131 | 0.40 |
ENSMUST00000110574.8
|
Ctdspl2
|
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase like 2 |
| chrX_+_72527208 | 0.40 |
ENSMUST00000033741.15
ENSMUST00000169489.2 |
Bgn
|
biglycan |
| chr2_+_65676111 | 0.40 |
ENSMUST00000122912.8
|
Csrnp3
|
cysteine-serine-rich nuclear protein 3 |
| chr10_+_98750268 | 0.40 |
ENSMUST00000219557.2
|
Atp2b1
|
ATPase, Ca++ transporting, plasma membrane 1 |
| chr15_-_76406102 | 0.39 |
ENSMUST00000164703.2
|
Scrt1
|
scratch family zinc finger 1 |
| chrX_+_33094635 | 0.39 |
ENSMUST00000177912.2
|
Btbd35f13
|
BTB domain containing 35, family member 13 |
| chrX_+_30768610 | 0.39 |
ENSMUST00000179532.2
|
Btbd35f29
|
BTB domain containing 35, family member 29 |
| chr4_+_42158092 | 0.39 |
ENSMUST00000098122.3
|
Gm13306
|
predicted gene 13306 |
| chr6_-_126141911 | 0.39 |
ENSMUST00000204542.3
|
Ntf3
|
neurotrophin 3 |
| chr6_-_28831746 | 0.39 |
ENSMUST00000062304.7
|
Lrrc4
|
leucine rich repeat containing 4 |
| chr11_+_56902624 | 0.39 |
ENSMUST00000036315.16
|
Gria1
|
glutamate receptor, ionotropic, AMPA1 (alpha 1) |
| chr2_-_38177359 | 0.39 |
ENSMUST00000102787.10
|
Dennd1a
|
DENN/MADD domain containing 1A |
| chrX_-_33014777 | 0.38 |
ENSMUST00000186329.2
|
Btbd35f15
|
BTB domain containing 35, family member 15 |
| chrX_-_33139812 | 0.38 |
ENSMUST00000105117.3
|
Btbd35f14
|
BTB domain containing 35, family member 14 |
| chr2_+_158344553 | 0.37 |
ENSMUST00000109484.2
|
Adig
|
adipogenin |
| chrX_-_31034822 | 0.37 |
ENSMUST00000238426.2
|
Btbd35f19
|
BTB domain containing 35, family member 19 |
| chrX_-_33580888 | 0.37 |
ENSMUST00000238632.2
|
Btbd35f9
|
BTB domain containing 35, family member 9 |
| chr6_+_122490577 | 0.37 |
ENSMUST00000118626.8
|
Mfap5
|
microfibrillar associated protein 5 |
| chr10_-_71121083 | 0.36 |
ENSMUST00000020085.7
|
Ube2d1
|
ubiquitin-conjugating enzyme E2D 1 |
| chrX_+_32070863 | 0.35 |
ENSMUST00000238237.2
|
Btbd35f1
|
BTB domain containing 35, family member 1 |
| chr14_-_20714634 | 0.35 |
ENSMUST00000119483.2
|
Synpo2l
|
synaptopodin 2-like |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.1 | 40.5 | GO:0018894 | dibenzo-p-dioxin metabolic process(GO:0018894) |
| 5.6 | 33.5 | GO:0032827 | peptidoglycan metabolic process(GO:0000270) peptidoglycan catabolic process(GO:0009253) negative regulation of natural killer cell differentiation(GO:0032824) negative regulation of natural killer cell differentiation involved in immune response(GO:0032827) |
| 3.0 | 33.1 | GO:0042436 | tryptophan catabolic process(GO:0006569) tryptophan catabolic process to kynurenine(GO:0019441) indole-containing compound catabolic process(GO:0042436) indolalkylamine catabolic process(GO:0046218) |
| 2.4 | 80.0 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 2.2 | 6.7 | GO:0071395 | response to jasmonic acid(GO:0009753) cellular response to jasmonic acid stimulus(GO:0071395) |
| 2.2 | 8.7 | GO:0071449 | cellular response to lipid hydroperoxide(GO:0071449) |
| 2.1 | 6.3 | GO:0018199 | peptidyl-glutamine modification(GO:0018199) |
| 2.1 | 6.2 | GO:1903920 | positive regulation of actin filament severing(GO:1903920) |
| 1.7 | 8.7 | GO:0009732 | detection of carbohydrate stimulus(GO:0009730) detection of hexose stimulus(GO:0009732) detection of monosaccharide stimulus(GO:0034287) detection of glucose(GO:0051594) |
| 1.3 | 3.9 | GO:0002302 | CD8-positive, alpha-beta T cell differentiation involved in immune response(GO:0002302) |
| 1.3 | 5.0 | GO:1905167 | positive regulation of lysosomal protein catabolic process(GO:1905167) |
| 1.2 | 25.5 | GO:0042359 | vitamin D metabolic process(GO:0042359) |
| 1.1 | 6.3 | GO:0070459 | prolactin secretion(GO:0070459) regulation of protein kinase C activity(GO:1900019) positive regulation of protein kinase C activity(GO:1900020) |
| 1.0 | 9.4 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 0.9 | 4.4 | GO:0006548 | histidine catabolic process(GO:0006548) |
| 0.9 | 3.5 | GO:0007522 | visceral muscle development(GO:0007522) |
| 0.9 | 3.4 | GO:0090472 | dibasic protein processing(GO:0090472) |
| 0.8 | 2.5 | GO:1902623 | negative regulation of neutrophil migration(GO:1902623) |
| 0.7 | 16.9 | GO:0051923 | sulfation(GO:0051923) |
| 0.7 | 2.2 | GO:1902460 | intermediate mesoderm development(GO:0048389) cloacal septation(GO:0060197) pattern specification involved in mesonephros development(GO:0061227) anterior/posterior pattern specification involved in kidney development(GO:0072098) regulation of mesenchymal stem cell proliferation(GO:1902460) positive regulation of mesenchymal stem cell proliferation(GO:1902462) cardiac jelly development(GO:1905072) negative regulation of cell proliferation involved in heart morphogenesis(GO:2000137) |
| 0.7 | 2.1 | GO:0045074 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.6 | 7.8 | GO:0019367 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.6 | 2.5 | GO:0030450 | regulation of complement activation, classical pathway(GO:0030450) |
| 0.6 | 5.1 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.5 | 4.4 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.5 | 5.0 | GO:0061042 | vascular wound healing(GO:0061042) |
| 0.4 | 1.3 | GO:0010536 | positive regulation of activation of Janus kinase activity(GO:0010536) |
| 0.4 | 3.7 | GO:0086073 | bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.4 | 2.0 | GO:1904491 | protein localization to ciliary transition zone(GO:1904491) |
| 0.3 | 2.4 | GO:0015862 | uridine transport(GO:0015862) |
| 0.3 | 2.0 | GO:0046726 | positive regulation by virus of viral protein levels in host cell(GO:0046726) |
| 0.3 | 2.1 | GO:0060455 | negative regulation of gastric acid secretion(GO:0060455) |
| 0.3 | 1.0 | GO:0032417 | positive regulation of sodium:proton antiporter activity(GO:0032417) |
| 0.2 | 14.1 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.2 | 1.3 | GO:0021914 | negative regulation of smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021914) |
| 0.2 | 1.1 | GO:0097498 | endothelial tube lumen extension(GO:0097498) |
| 0.2 | 0.8 | GO:0043396 | corticotropin-releasing hormone secretion(GO:0043396) regulation of corticotropin-releasing hormone secretion(GO:0043397) positive regulation of corticotropin-releasing hormone secretion(GO:0051466) |
| 0.2 | 0.6 | GO:0033861 | negative regulation of NAD(P)H oxidase activity(GO:0033861) |
| 0.2 | 2.5 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.2 | 1.0 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.2 | 0.8 | GO:0030202 | heparan sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0015014) heparin metabolic process(GO:0030202) |
| 0.2 | 0.8 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.2 | 3.5 | GO:0002710 | negative regulation of T cell mediated immunity(GO:0002710) |
| 0.2 | 1.1 | GO:0002282 | microglial cell activation involved in immune response(GO:0002282) regulation of dendritic cell cytokine production(GO:0002730) |
| 0.1 | 3.3 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.1 | 1.2 | GO:0060011 | Sertoli cell proliferation(GO:0060011) |
| 0.1 | 3.8 | GO:0035634 | response to stilbenoid(GO:0035634) |
| 0.1 | 2.1 | GO:0051823 | regulation of synapse structural plasticity(GO:0051823) |
| 0.1 | 2.5 | GO:0038063 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) |
| 0.1 | 0.5 | GO:0071500 | cellular response to nitrosative stress(GO:0071500) |
| 0.1 | 1.3 | GO:0035092 | sperm chromatin condensation(GO:0035092) |
| 0.1 | 0.4 | GO:1902528 | regulation of protein linear polyubiquitination(GO:1902528) positive regulation of protein linear polyubiquitination(GO:1902530) |
| 0.1 | 0.5 | GO:0032901 | positive regulation of neurotrophin production(GO:0032901) |
| 0.1 | 0.6 | GO:1900127 | positive regulation of hyaluronan biosynthetic process(GO:1900127) |
| 0.1 | 2.5 | GO:0014067 | negative regulation of phosphatidylinositol 3-kinase signaling(GO:0014067) |
| 0.1 | 0.3 | GO:0070268 | cornification(GO:0070268) |
| 0.1 | 1.4 | GO:0060484 | lung-associated mesenchyme development(GO:0060484) |
| 0.1 | 1.5 | GO:1902170 | cellular response to reactive nitrogen species(GO:1902170) |
| 0.1 | 0.4 | GO:0007403 | glial cell fate determination(GO:0007403) |
| 0.1 | 0.5 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.1 | 0.7 | GO:0006451 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.1 | 0.5 | GO:0072307 | metanephric nephron tubule epithelial cell differentiation(GO:0072257) regulation of metanephric nephron tubule epithelial cell differentiation(GO:0072307) |
| 0.1 | 2.9 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.1 | 0.3 | GO:0014886 | transition between slow and fast fiber(GO:0014886) |
| 0.1 | 0.4 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 0.1 | 0.7 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.1 | 6.4 | GO:1900181 | negative regulation of protein localization to nucleus(GO:1900181) |
| 0.1 | 0.7 | GO:1903546 | protein localization to photoreceptor outer segment(GO:1903546) |
| 0.1 | 0.8 | GO:0043402 | glucocorticoid mediated signaling pathway(GO:0043402) |
| 0.0 | 11.5 | GO:0006694 | steroid biosynthetic process(GO:0006694) |
| 0.0 | 7.8 | GO:0008202 | steroid metabolic process(GO:0008202) |
| 0.0 | 1.8 | GO:0045187 | regulation of circadian sleep/wake cycle, sleep(GO:0045187) |
| 0.0 | 6.7 | GO:0009166 | nucleotide catabolic process(GO:0009166) |
| 0.0 | 0.3 | GO:0060467 | negative regulation of fertilization(GO:0060467) |
| 0.0 | 0.6 | GO:0014807 | regulation of somitogenesis(GO:0014807) |
| 0.0 | 0.2 | GO:1903575 | cornified envelope assembly(GO:1903575) |
| 0.0 | 0.6 | GO:0007342 | fusion of sperm to egg plasma membrane(GO:0007342) |
| 0.0 | 0.2 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.0 | 0.7 | GO:0001911 | negative regulation of leukocyte mediated cytotoxicity(GO:0001911) negative regulation of natural killer cell mediated cytotoxicity(GO:0045953) |
| 0.0 | 1.3 | GO:0001913 | T cell mediated cytotoxicity(GO:0001913) |
| 0.0 | 3.7 | GO:0042147 | retrograde transport, endosome to Golgi(GO:0042147) |
| 0.0 | 0.4 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.0 | 0.5 | GO:0006098 | pentose-phosphate shunt(GO:0006098) |
| 0.0 | 0.7 | GO:0021542 | dentate gyrus development(GO:0021542) |
| 0.0 | 1.8 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.0 | 4.7 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.0 | 0.5 | GO:0061162 | establishment of monopolar cell polarity(GO:0061162) |
| 0.0 | 0.2 | GO:0010578 | regulation of adenylate cyclase activity involved in G-protein coupled receptor signaling pathway(GO:0010578) positive regulation of adenylate cyclase activity involved in G-protein coupled receptor signaling pathway(GO:0010579) |
| 0.0 | 0.1 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 0.0 | 0.4 | GO:1902916 | positive regulation of protein polyubiquitination(GO:1902916) |
| 0.0 | 0.8 | GO:0097435 | fibril organization(GO:0097435) |
| 0.0 | 2.1 | GO:2001222 | regulation of neuron migration(GO:2001222) |
| 0.0 | 0.3 | GO:0034551 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) |
| 0.0 | 0.9 | GO:0061512 | protein localization to cilium(GO:0061512) |
| 0.0 | 0.4 | GO:0097119 | postsynaptic density protein 95 clustering(GO:0097119) |
| 0.0 | 0.6 | GO:0000460 | maturation of 5.8S rRNA(GO:0000460) |
| 0.0 | 0.3 | GO:0046475 | glycerophospholipid catabolic process(GO:0046475) |
| 0.0 | 2.3 | GO:0006821 | chloride transport(GO:0006821) |
| 0.0 | 0.8 | GO:0060292 | long term synaptic depression(GO:0060292) |
| 0.0 | 0.7 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 1.0 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.0 | 0.9 | GO:0030512 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) |
| 0.0 | 0.3 | GO:0035455 | response to interferon-alpha(GO:0035455) |
| 0.0 | 0.4 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 0.0 | 0.0 | GO:1902774 | late endosome to lysosome transport(GO:1902774) |
| 0.0 | 0.2 | GO:0044458 | motile cilium assembly(GO:0044458) |
| 0.0 | 0.0 | GO:0060690 | epithelial cell differentiation involved in salivary gland development(GO:0060690) |
| 0.0 | 8.2 | GO:0007608 | sensory perception of smell(GO:0007608) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 6.3 | GO:0097487 | multivesicular body, internal vesicle(GO:0097487) |
| 0.7 | 7.2 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.5 | 8.7 | GO:0045180 | basal cortex(GO:0045180) |
| 0.4 | 2.1 | GO:0033257 | Bcl3/NF-kappaB2 complex(GO:0033257) |
| 0.3 | 1.3 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) |
| 0.2 | 1.5 | GO:1990604 | IRE1-TRAF2-ASK1 complex(GO:1990604) |
| 0.2 | 4.1 | GO:0012510 | trans-Golgi network transport vesicle membrane(GO:0012510) |
| 0.2 | 0.8 | GO:0098559 | cytoplasmic side of early endosome membrane(GO:0098559) |
| 0.2 | 1.3 | GO:0098645 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
| 0.2 | 0.8 | GO:0097058 | CRLF-CLCF1 complex(GO:0097058) |
| 0.2 | 0.8 | GO:0044308 | axonal spine(GO:0044308) |
| 0.2 | 1.2 | GO:0005927 | muscle tendon junction(GO:0005927) |
| 0.1 | 3.5 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 0.1 | 0.5 | GO:0071148 | TEAD-1-YAP complex(GO:0071148) TEAD-2-YAP complex(GO:0071149) |
| 0.1 | 3.7 | GO:0030057 | desmosome(GO:0030057) |
| 0.1 | 0.8 | GO:1990923 | PET complex(GO:1990923) |
| 0.1 | 2.9 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.1 | 2.0 | GO:0036038 | MKS complex(GO:0036038) |
| 0.1 | 1.8 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.1 | 1.2 | GO:0097136 | Bcl-2 family protein complex(GO:0097136) |
| 0.1 | 26.4 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.1 | 8.7 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.1 | 0.3 | GO:0034667 | integrin alpha3-beta1 complex(GO:0034667) |
| 0.1 | 12.1 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 2.7 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.0 | 4.2 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 3.1 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.0 | 0.4 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 1.4 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 1.0 | GO:0030665 | clathrin-coated vesicle membrane(GO:0030665) |
| 0.0 | 0.3 | GO:0005750 | mitochondrial respiratory chain complex III(GO:0005750) |
| 0.0 | 0.6 | GO:0000178 | exosome (RNase complex)(GO:0000178) |
| 0.0 | 1.3 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 11.9 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.0 | 9.5 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 48.8 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.0 | 3.6 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 0.2 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.0 | 0.5 | GO:0043196 | varicosity(GO:0043196) |
| 0.0 | 0.4 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 1.3 | GO:0032420 | stereocilium(GO:0032420) |
| 0.0 | 2.5 | GO:0000795 | synaptonemal complex(GO:0000795) |
| 0.0 | 0.6 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.0 | 0.7 | GO:0001917 | photoreceptor inner segment(GO:0001917) |
| 0.0 | 0.6 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 45.0 | GO:0070062 | extracellular exosome(GO:0070062) |
| 0.0 | 13.0 | GO:0031226 | intrinsic component of plasma membrane(GO:0031226) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 11.0 | 33.1 | GO:0033754 | indoleamine 2,3-dioxygenase activity(GO:0033754) |
| 6.7 | 40.5 | GO:0034875 | oxidoreductase activity, acting on CH or CH2 groups, quinone or similar compound as acceptor(GO:0033695) caffeine oxidase activity(GO:0034875) |
| 6.4 | 25.5 | GO:1902271 | D3 vitamins binding(GO:1902271) |
| 5.6 | 33.5 | GO:0016019 | N-acetylmuramoyl-L-alanine amidase activity(GO:0008745) peptidoglycan receptor activity(GO:0016019) |
| 2.5 | 80.0 | GO:0008392 | arachidonic acid epoxygenase activity(GO:0008392) |
| 2.4 | 16.9 | GO:0050656 | alcohol sulfotransferase activity(GO:0004027) 3'-phosphoadenosine 5'-phosphosulfate binding(GO:0050656) |
| 2.2 | 6.7 | GO:0047006 | 17-alpha,20-alpha-dihydroxypregn-4-en-3-one dehydrogenase activity(GO:0047006) |
| 1.6 | 6.3 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 1.2 | 7.2 | GO:0004850 | uridine phosphorylase activity(GO:0004850) |
| 1.0 | 5.0 | GO:0097003 | adipokinetic hormone receptor activity(GO:0097003) |
| 0.9 | 6.2 | GO:0061629 | RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 0.9 | 8.7 | GO:0004396 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) |
| 0.8 | 2.5 | GO:0001847 | opsonin receptor activity(GO:0001847) |
| 0.6 | 7.8 | GO:0102337 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.5 | 3.5 | GO:0030899 | calcium-dependent ATPase activity(GO:0030899) |
| 0.5 | 4.4 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 0.5 | 4.4 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.4 | 1.3 | GO:0019981 | interleukin-6 receptor activity(GO:0004915) interleukin-6 binding(GO:0019981) |
| 0.4 | 11.5 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.4 | 12.5 | GO:1900750 | glutathione binding(GO:0043295) oligopeptide binding(GO:1900750) |
| 0.4 | 2.0 | GO:0102344 | 3-hydroxy-behenoyl-CoA dehydratase activity(GO:0102344) 3-hydroxy-lignoceroyl-CoA dehydratase activity(GO:0102345) |
| 0.4 | 1.8 | GO:0016499 | orexin receptor activity(GO:0016499) |
| 0.3 | 5.0 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.3 | 9.4 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen(GO:0016701) oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.2 | 3.8 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.2 | 0.8 | GO:0030023 | extracellular matrix constituent conferring elasticity(GO:0030023) |
| 0.2 | 0.5 | GO:0047936 | glucose 1-dehydrogenase [NAD(P)] activity(GO:0047936) |
| 0.2 | 2.2 | GO:0039706 | co-receptor binding(GO:0039706) BMP receptor binding(GO:0070700) |
| 0.2 | 0.5 | GO:0031728 | CCR3 chemokine receptor binding(GO:0031728) |
| 0.1 | 2.1 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.1 | 2.7 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.1 | 8.6 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.1 | 2.5 | GO:0001846 | opsonin binding(GO:0001846) |
| 0.1 | 2.5 | GO:0005527 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.1 | 3.5 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.1 | 2.9 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.1 | 1.2 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.1 | 1.4 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 0.8 | GO:0043423 | 3-phosphoinositide-dependent protein kinase binding(GO:0043423) |
| 0.1 | 14.1 | GO:0015171 | amino acid transmembrane transporter activity(GO:0015171) |
| 0.1 | 0.8 | GO:0099583 | AMPA glutamate receptor activity(GO:0004971) neurotransmitter receptor activity involved in regulation of postsynaptic cytosolic calcium ion concentration(GO:0099583) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.1 | 1.3 | GO:0046703 | natural killer cell lectin-like receptor binding(GO:0046703) |
| 0.1 | 1.3 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.1 | 0.8 | GO:0005127 | ciliary neurotrophic factor receptor binding(GO:0005127) |
| 0.1 | 2.4 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 0.1 | 2.3 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.1 | 2.5 | GO:0050750 | low-density lipoprotein particle receptor binding(GO:0050750) |
| 0.1 | 0.7 | GO:0035368 | selenocysteine insertion sequence binding(GO:0035368) |
| 0.1 | 0.5 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 0.1 | 0.3 | GO:0004904 | interferon receptor activity(GO:0004904) type I interferon receptor activity(GO:0004905) type I interferon binding(GO:0019962) |
| 0.0 | 1.2 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.0 | 0.5 | GO:0043121 | neurotrophin binding(GO:0043121) |
| 0.0 | 1.2 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.0 | 0.3 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 0.0 | 0.3 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.0 | 2.7 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 1.5 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.3 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 0.0 | 0.3 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.6 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 3.9 | GO:0001191 | transcriptional repressor activity, RNA polymerase II transcription factor binding(GO:0001191) |
| 0.0 | 2.5 | GO:0005249 | voltage-gated potassium channel activity(GO:0005249) |
| 0.0 | 1.0 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.4 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 4.9 | GO:0005549 | odorant binding(GO:0005549) |
| 0.0 | 3.3 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.6 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 0.6 | GO:0043236 | laminin binding(GO:0043236) |
| 0.0 | 0.8 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.1 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) |
| 0.0 | 1.5 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 0.0 | 0.5 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.0 | 0.2 | GO:0030976 | thiamine pyrophosphate binding(GO:0030976) |
| 0.0 | 0.2 | GO:0016918 | retinal binding(GO:0016918) retinol binding(GO:0019841) |
| 0.0 | 0.8 | GO:0008408 | 3'-5' exonuclease activity(GO:0008408) |
| 0.0 | 0.8 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 0.4 | GO:0050840 | extracellular matrix binding(GO:0050840) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 6.3 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.2 | 10.2 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.1 | 16.9 | PID E2F PATHWAY | E2F transcription factor network |
| 0.1 | 5.4 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.1 | 0.6 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.1 | 0.7 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.1 | 2.1 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.1 | 4.4 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.1 | 4.2 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.1 | 1.5 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.1 | 3.9 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.0 | 2.3 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 2.2 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 1.2 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 6.2 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 2.5 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.0 | 0.5 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 0.8 | PID FOXO PATHWAY | FoxO family signaling |
| 0.0 | 0.5 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 13.5 | 40.5 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 2.4 | 33.1 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 1.2 | 16.9 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.6 | 25.5 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.4 | 7.2 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.3 | 2.5 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.2 | 8.7 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.2 | 3.4 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.2 | 7.8 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.2 | 11.5 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.2 | 14.1 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.2 | 5.9 | REACTOME SHC1 EVENTS IN EGFR SIGNALING | Genes involved in SHC1 events in EGFR signaling |
| 0.1 | 2.9 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.1 | 1.1 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.1 | 13.7 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.1 | 3.5 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.1 | 0.9 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |
| 0.0 | 2.5 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.0 | 2.4 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 1.3 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 2.5 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.0 | 0.8 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.0 | 0.4 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 0.6 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GLP1 | Genes involved in Synthesis, Secretion, and Inactivation of Glucagon-like Peptide-1 (GLP-1) |
| 0.0 | 0.3 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.0 | 0.8 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 0.4 | REACTOME PHOSPHORYLATION OF THE APC C | Genes involved in Phosphorylation of the APC/C |
| 0.0 | 0.5 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 1.0 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.3 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |