Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
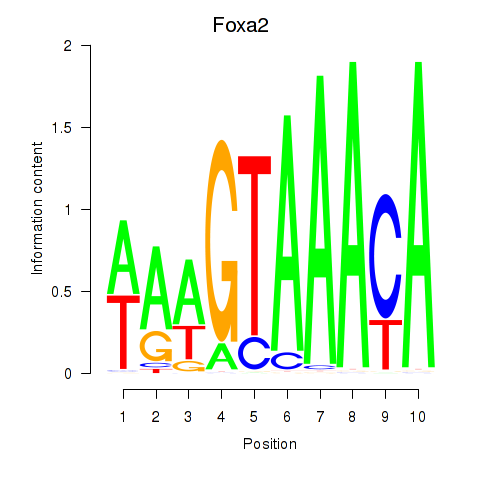
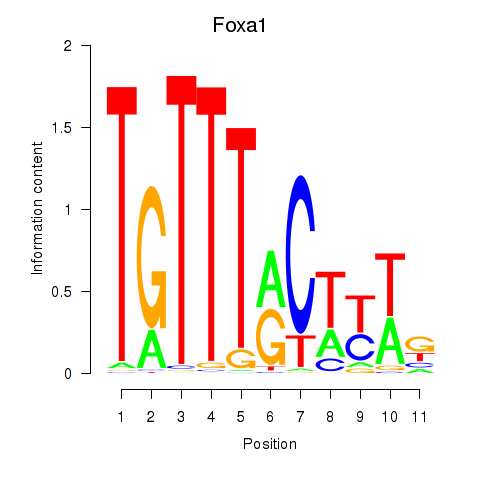
Results for Foxa2_Foxa1
Z-value: 2.75
Transcription factors associated with Foxa2_Foxa1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Foxa2
|
ENSMUSG00000037025.12 | Foxa2 |
|
Foxa1
|
ENSMUSG00000035451.8 | Foxa1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Foxa2 | mm39_v1_chr2_-_147888816_147888912 | 0.82 | 1.1e-09 | Click! |
| Foxa1 | mm39_v1_chr12_-_57592907_57592926 | 0.51 | 1.3e-03 | Click! |
Activity profile of Foxa2_Foxa1 motif
Sorted Z-values of Foxa2_Foxa1 motif
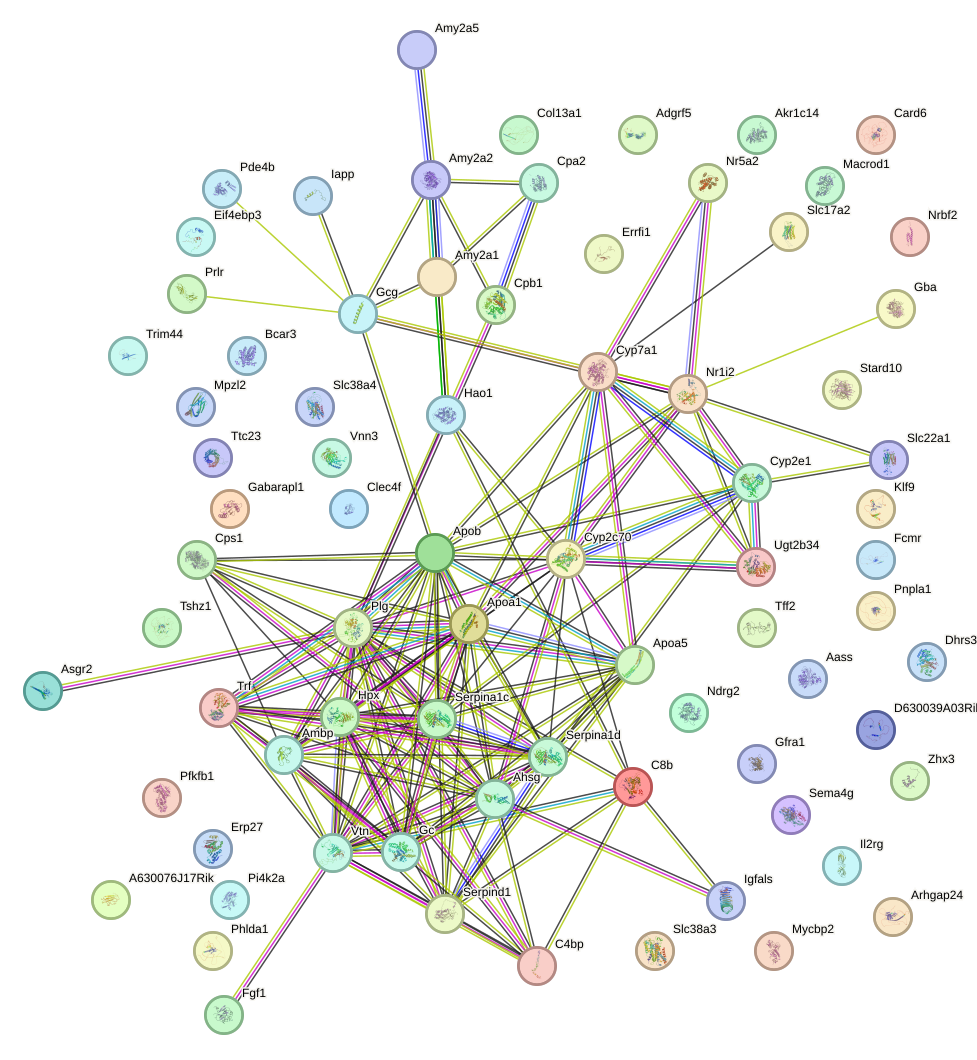
Network of associatons between targets according to the STRING database.
First level regulatory network of Foxa2_Foxa1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_-_113166153 | 25.35 |
ENSMUST00000098673.5
|
Amy2a5
|
amylase 2a5 |
| chr6_+_30541581 | 25.27 |
ENSMUST00000096066.5
|
Cpa2
|
carboxypeptidase A2, pancreatic |
| chr17_-_31363245 | 23.03 |
ENSMUST00000024826.8
|
Tff2
|
trefoil factor 2 (spasmolytic protein 1) |
| chr3_-_20329823 | 21.99 |
ENSMUST00000011607.6
|
Cpb1
|
carboxypeptidase B1 (tissue) |
| chr9_+_46179899 | 20.34 |
ENSMUST00000121598.8
|
Apoa5
|
apolipoprotein A-V |
| chr7_+_140343652 | 15.99 |
ENSMUST00000026552.9
ENSMUST00000209253.2 ENSMUST00000210235.2 |
Cyp2e1
|
cytochrome P450, family 2, subfamily e, polypeptide 1 |
| chr1_-_130589321 | 14.75 |
ENSMUST00000137276.3
|
C4bp
|
complement component 4 binding protein |
| chr14_-_52151537 | 14.51 |
ENSMUST00000227402.2
ENSMUST00000227237.2 |
Ndrg2
|
N-myc downstream regulated gene 2 |
| chr1_-_130589349 | 14.32 |
ENSMUST00000027657.14
|
C4bp
|
complement component 4 binding protein |
| chr7_+_130633776 | 13.88 |
ENSMUST00000084509.7
ENSMUST00000213064.3 ENSMUST00000208311.4 |
Dmbt1
|
deleted in malignant brain tumors 1 |
| chr4_-_57916283 | 13.00 |
ENSMUST00000063816.6
|
D630039A03Rik
|
RIKEN cDNA D630039A03 gene |
| chr3_-_113325938 | 12.32 |
ENSMUST00000132353.2
|
Amy2a1
|
amylase 2a1 |
| chr17_+_25097199 | 12.27 |
ENSMUST00000050714.8
|
Igfals
|
insulin-like growth factor binding protein, acid labile subunit |
| chr15_+_10224052 | 12.06 |
ENSMUST00000128450.8
ENSMUST00000148257.8 ENSMUST00000128921.8 |
Prlr
|
prolactin receptor |
| chr5_-_89605622 | 11.15 |
ENSMUST00000049209.13
|
Gc
|
vitamin D binding protein |
| chr2_-_134396268 | 10.40 |
ENSMUST00000028704.3
|
Hao1
|
hydroxyacid oxidase 1, liver |
| chr15_-_96929086 | 9.89 |
ENSMUST00000230086.2
|
Slc38a4
|
solute carrier family 38, member 4 |
| chr17_-_12894716 | 9.76 |
ENSMUST00000024596.10
|
Slc22a1
|
solute carrier family 22 (organic cation transporter), member 1 |
| chr4_-_6275629 | 9.71 |
ENSMUST00000029905.2
|
Cyp7a1
|
cytochrome P450, family 7, subfamily a, polypeptide 1 |
| chr6_-_136899167 | 9.17 |
ENSMUST00000032343.7
|
Erp27
|
endoplasmic reticulum protein 27 |
| chr17_+_12597490 | 8.96 |
ENSMUST00000014578.7
|
Plg
|
plasminogen |
| chr12_-_81014849 | 8.87 |
ENSMUST00000095572.5
|
Slc10a1
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 1 |
| chr12_-_81014755 | 8.80 |
ENSMUST00000218342.2
|
Slc10a1
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 1 |
| chr7_-_126651847 | 8.65 |
ENSMUST00000205424.2
|
Zg16
|
zymogen granule protein 16 |
| chr12_-_103871146 | 8.20 |
ENSMUST00000074051.6
|
Serpina1c
|
serine (or cysteine) peptidase inhibitor, clade A, member 1C |
| chr13_+_4099001 | 7.72 |
ENSMUST00000118717.10
|
Akr1c14
|
aldo-keto reductase family 1, member C14 |
| chr19_+_23118545 | 7.67 |
ENSMUST00000036884.3
|
Klf9
|
Kruppel-like factor 9 |
| chr4_+_150938376 | 7.54 |
ENSMUST00000073600.9
|
Errfi1
|
ERBB receptor feedback inhibitor 1 |
| chr2_+_58645189 | 7.41 |
ENSMUST00000102755.4
ENSMUST00000230627.2 ENSMUST00000229923.2 |
Upp2
|
uridine phosphorylase 2 |
| chr3_+_20011405 | 6.97 |
ENSMUST00000108325.9
|
Cp
|
ceruloplasmin |
| chr6_+_129510331 | 6.93 |
ENSMUST00000204956.2
ENSMUST00000204639.2 |
Gabarapl1
|
gamma-aminobutyric acid (GABA) A receptor-associated protein-like 1 |
| chr12_+_8027767 | 6.86 |
ENSMUST00000037520.14
|
Apob
|
apolipoprotein B |
| chr10_-_95678786 | 6.69 |
ENSMUST00000211096.2
|
Gm33543
|
predicted gene, 33543 |
| chr12_+_8027640 | 6.61 |
ENSMUST00000171271.8
ENSMUST00000037811.13 |
Apob
|
apolipoprotein B |
| chr7_-_105249308 | 6.59 |
ENSMUST00000210531.2
ENSMUST00000033185.10 |
Hpx
|
hemopexin |
| chr10_+_111342147 | 6.57 |
ENSMUST00000164773.2
|
Phlda1
|
pleckstrin homology like domain, family A, member 1 |
| chr15_-_96918203 | 6.57 |
ENSMUST00000166223.2
|
Slc38a4
|
solute carrier family 38, member 4 |
| chr16_+_22710785 | 6.34 |
ENSMUST00000023583.7
ENSMUST00000232098.2 |
Ahsg
|
alpha-2-HS-glycoprotein |
| chr10_+_87697155 | 6.26 |
ENSMUST00000122100.3
|
Igf1
|
insulin-like growth factor 1 |
| chr6_+_129510117 | 6.14 |
ENSMUST00000032264.9
|
Gabarapl1
|
gamma-aminobutyric acid (GABA) A receptor-associated protein-like 1 |
| chr2_+_58644922 | 6.04 |
ENSMUST00000059102.13
|
Upp2
|
uridine phosphorylase 2 |
| chr8_-_45747883 | 6.02 |
ENSMUST00000026907.6
|
Klkb1
|
kallikrein B, plasma 1 |
| chr3_+_107137924 | 6.00 |
ENSMUST00000179399.3
|
A630076J17Rik
|
RIKEN cDNA A630076J17 gene |
| chr10_-_95678748 | 5.99 |
ENSMUST00000210336.2
|
Gm33543
|
predicted gene, 33543 |
| chr6_+_129510145 | 5.77 |
ENSMUST00000204487.3
|
Gabarapl1
|
gamma-aminobutyric acid (GABA) A receptor-associated protein-like 1 |
| chr6_-_23132977 | 5.68 |
ENSMUST00000031707.14
|
Aass
|
aminoadipate-semialdehyde synthase |
| chr9_-_107546195 | 5.61 |
ENSMUST00000192990.6
|
Slc38a3
|
solute carrier family 38, member 3 |
| chr3_+_122277372 | 5.56 |
ENSMUST00000197073.2
|
Bcar3
|
breast cancer anti-estrogen resistance 3 |
| chr5_-_87054796 | 5.28 |
ENSMUST00000031181.16
ENSMUST00000113333.2 |
Ugt2b34
|
UDP glucuronosyltransferase 2 family, polypeptide B34 |
| chr10_+_23727325 | 5.17 |
ENSMUST00000020190.8
|
Vnn3
|
vanin 3 |
| chr15_-_96917804 | 5.15 |
ENSMUST00000231039.2
|
Slc38a4
|
solute carrier family 38, member 4 |
| chr3_-_146302343 | 5.08 |
ENSMUST00000029836.9
|
Dnase2b
|
deoxyribonuclease II beta |
| chr2_-_160701523 | 5.01 |
ENSMUST00000103112.8
|
Zhx3
|
zinc fingers and homeoboxes 3 |
| chr1_+_67162176 | 4.99 |
ENSMUST00000027144.8
|
Cps1
|
carbamoyl-phosphate synthetase 1 |
| chr15_+_4756684 | 4.93 |
ENSMUST00000161997.8
ENSMUST00000022788.15 |
C6
|
complement component 6 |
| chr3_+_20011251 | 4.93 |
ENSMUST00000108328.8
|
Cp
|
ceruloplasmin |
| chr18_-_39051695 | 4.92 |
ENSMUST00000040647.11
|
Fgf1
|
fibroblast growth factor 1 |
| chr11_-_59927688 | 4.88 |
ENSMUST00000102692.10
|
Pemt
|
phosphatidylethanolamine N-methyltransferase |
| chr12_-_103739847 | 4.86 |
ENSMUST00000078869.6
|
Serpina1d
|
serine (or cysteine) peptidase inhibitor, clade A, member 1D |
| chr9_-_71070506 | 4.79 |
ENSMUST00000074465.9
|
Aqp9
|
aquaporin 9 |
| chr9_-_107546166 | 4.70 |
ENSMUST00000177567.8
|
Slc38a3
|
solute carrier family 38, member 3 |
| chr2_-_62313981 | 4.63 |
ENSMUST00000136686.2
ENSMUST00000102733.10 |
Gcg
|
glucagon |
| chr15_+_4756657 | 4.62 |
ENSMUST00000162585.8
|
C6
|
complement component 6 |
| chrX_+_100419965 | 4.58 |
ENSMUST00000119080.8
|
Gjb1
|
gap junction protein, beta 1 |
| chr3_+_20011201 | 4.57 |
ENSMUST00000091309.12
ENSMUST00000108329.8 ENSMUST00000003714.13 |
Cp
|
ceruloplasmin |
| chr7_+_100966289 | 4.52 |
ENSMUST00000163799.9
ENSMUST00000164479.9 |
Stard10
|
START domain containing 10 |
| chr5_+_90708962 | 4.39 |
ENSMUST00000094615.8
ENSMUST00000200765.2 |
Albfm1
|
albumin superfamily member 1 |
| chr13_+_23991010 | 4.37 |
ENSMUST00000006786.11
ENSMUST00000099697.3 |
Slc17a2
|
solute carrier family 17 (sodium phosphate), member 2 |
| chr14_-_30665232 | 4.26 |
ENSMUST00000006704.17
ENSMUST00000163118.2 |
Itih1
|
inter-alpha trypsin inhibitor, heavy chain 1 |
| chr11_+_78389913 | 4.25 |
ENSMUST00000017488.5
|
Vtn
|
vitronectin |
| chr1_+_130793406 | 4.15 |
ENSMUST00000038829.7
|
Fcmr
|
Fc fragment of IgM receptor |
| chr6_-_83633064 | 3.99 |
ENSMUST00000014686.3
|
Clec4f
|
C-type lectin domain family 4, member f |
| chr16_+_17149235 | 3.76 |
ENSMUST00000023450.15
ENSMUST00000231884.2 |
Serpind1
|
serine (or cysteine) peptidase inhibitor, clade D, member 1 |
| chr19_+_42078859 | 3.74 |
ENSMUST00000235932.2
ENSMUST00000066778.6 |
Pi4k2a
|
phosphatidylinositol 4-kinase type 2 alpha |
| chr13_+_41013230 | 3.72 |
ENSMUST00000110191.10
|
Gcnt2
|
glucosaminyl (N-acetyl) transferase 2, I-branching enzyme |
| chr15_-_5137975 | 3.69 |
ENSMUST00000118365.3
|
Card6
|
caspase recruitment domain family, member 6 |
| chr19_-_58443593 | 3.69 |
ENSMUST00000135730.2
ENSMUST00000152507.8 |
Gfra1
|
glial cell line derived neurotrophic factor family receptor alpha 1 |
| chr16_-_38115172 | 3.69 |
ENSMUST00000023504.5
|
Nr1i2
|
nuclear receptor subfamily 1, group I, member 2 |
| chr15_+_102012782 | 3.66 |
ENSMUST00000230474.2
|
Tns2
|
tensin 2 |
| chr11_+_69983531 | 3.62 |
ENSMUST00000124721.2
|
Asgr2
|
asialoglycoprotein receptor 2 |
| chr11_+_69983459 | 3.58 |
ENSMUST00000102572.8
|
Asgr2
|
asialoglycoprotein receptor 2 |
| chr17_+_43671314 | 3.54 |
ENSMUST00000226087.2
|
Adgrf5
|
adhesion G protein-coupled receptor F5 |
| chr11_+_69983479 | 3.51 |
ENSMUST00000143772.8
|
Asgr2
|
asialoglycoprotein receptor 2 |
| chr19_-_40175709 | 3.42 |
ENSMUST00000051846.13
|
Cyp2c70
|
cytochrome P450, family 2, subfamily c, polypeptide 70 |
| chr7_+_119499322 | 3.33 |
ENSMUST00000106516.2
|
Lyrm1
|
LYR motif containing 1 |
| chr5_+_102629240 | 3.27 |
ENSMUST00000073302.12
ENSMUST00000094559.9 |
Arhgap24
|
Rho GTPase activating protein 24 |
| chr8_+_46111703 | 3.24 |
ENSMUST00000134675.8
ENSMUST00000139869.8 ENSMUST00000126067.8 |
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr19_-_58443830 | 3.23 |
ENSMUST00000026076.14
|
Gfra1
|
glial cell line derived neurotrophic factor family receptor alpha 1 |
| chr6_+_142244145 | 3.22 |
ENSMUST00000041993.3
|
Iapp
|
islet amyloid polypeptide |
| chr19_+_7034149 | 3.21 |
ENSMUST00000040261.7
|
Macrod1
|
mono-ADP ribosylhydrolase 1 |
| chr4_-_63072367 | 3.21 |
ENSMUST00000030041.5
|
Ambp
|
alpha 1 microglobulin/bikunin precursor |
| chr12_-_103704417 | 3.19 |
ENSMUST00000095450.11
ENSMUST00000187220.2 |
Serpina1b
|
serine (or cysteine) preptidase inhibitor, clade A, member 1B |
| chr4_+_144619647 | 3.16 |
ENSMUST00000154208.8
|
Dhrs3
|
dehydrogenase/reductase (SDR family) member 3 |
| chr6_+_34686543 | 3.15 |
ENSMUST00000031775.13
|
Cald1
|
caldesmon 1 |
| chr2_-_102231208 | 3.14 |
ENSMUST00000102573.8
|
Trim44
|
tripartite motif-containing 44 |
| chr12_-_103829810 | 3.11 |
ENSMUST00000085056.8
ENSMUST00000072876.12 ENSMUST00000124717.2 |
Serpina1a
|
serine (or cysteine) peptidase inhibitor, clade A, member 1A |
| chr4_+_144619696 | 3.03 |
ENSMUST00000142808.8
|
Dhrs3
|
dehydrogenase/reductase (SDR family) member 3 |
| chr15_+_10177709 | 2.99 |
ENSMUST00000124470.8
|
Prlr
|
prolactin receptor |
| chr4_+_102427247 | 2.89 |
ENSMUST00000097950.9
|
Pde4b
|
phosphodiesterase 4B, cAMP specific |
| chr15_-_5137951 | 2.83 |
ENSMUST00000141020.2
|
Card6
|
caspase recruitment domain family, member 6 |
| chr1_-_97904958 | 2.82 |
ENSMUST00000161567.8
|
Pam
|
peptidylglycine alpha-amidating monooxygenase |
| chr4_+_104623505 | 2.82 |
ENSMUST00000031663.10
ENSMUST00000065072.7 |
C8b
|
complement component 8, beta polypeptide |
| chr17_-_31852128 | 2.74 |
ENSMUST00000236909.2
|
Cbs
|
cystathionine beta-synthase |
| chr7_+_67305162 | 2.71 |
ENSMUST00000107470.2
|
Ttc23
|
tetratricopeptide repeat domain 23 |
| chr17_+_29077385 | 2.69 |
ENSMUST00000056866.8
|
Pnpla1
|
patatin-like phospholipase domain containing 1 |
| chr13_-_60325170 | 2.63 |
ENSMUST00000065086.6
|
Gas1
|
growth arrest specific 1 |
| chrX_+_149377416 | 2.61 |
ENSMUST00000112713.3
|
Pfkfb1
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 |
| chr10_-_67120959 | 2.53 |
ENSMUST00000159002.2
ENSMUST00000077839.13 |
Nrbf2
|
nuclear receptor binding factor 2 |
| chr19_+_44980565 | 2.51 |
ENSMUST00000179305.2
|
Sema4g
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4G |
| chr18_-_84104507 | 2.48 |
ENSMUST00000060303.10
|
Tshz1
|
teashirt zinc finger family member 1 |
| chr10_-_61814852 | 2.46 |
ENSMUST00000105453.8
ENSMUST00000105452.9 ENSMUST00000105454.3 |
Col13a1
|
collagen, type XIII, alpha 1 |
| chr18_+_36797113 | 2.42 |
ENSMUST00000036765.8
|
Eif4ebp3
|
eukaryotic translation initiation factor 4E binding protein 3 |
| chr1_-_136888118 | 2.36 |
ENSMUST00000192357.6
ENSMUST00000027649.14 |
Nr5a2
|
nuclear receptor subfamily 5, group A, member 2 |
| chr18_-_84104574 | 2.35 |
ENSMUST00000175783.3
|
Tshz1
|
teashirt zinc finger family member 1 |
| chr9_+_46139878 | 2.34 |
ENSMUST00000034588.9
ENSMUST00000132155.2 |
Apoa1
|
apolipoprotein A-I |
| chr9_+_44951075 | 2.33 |
ENSMUST00000217097.2
|
Mpzl2
|
myelin protein zero-like 2 |
| chr6_+_34686373 | 2.32 |
ENSMUST00000115021.8
|
Cald1
|
caldesmon 1 |
| chr4_+_148686985 | 2.31 |
ENSMUST00000105701.9
ENSMUST00000052060.7 |
Masp2
|
mannan-binding lectin serine peptidase 2 |
| chr8_-_122671588 | 2.27 |
ENSMUST00000057653.8
|
Car5a
|
carbonic anhydrase 5a, mitochondrial |
| chr15_+_3300249 | 2.24 |
ENSMUST00000082424.12
ENSMUST00000159158.9 ENSMUST00000159216.10 ENSMUST00000160311.3 |
Selenop
|
selenoprotein P |
| chr2_-_51862941 | 2.21 |
ENSMUST00000145481.8
ENSMUST00000112705.9 |
Nmi
|
N-myc (and STAT) interactor |
| chr2_-_25982160 | 2.17 |
ENSMUST00000114159.9
|
Nacc2
|
nucleus accumbens associated 2, BEN and BTB (POZ) domain containing |
| chr11_+_114741948 | 2.15 |
ENSMUST00000133245.2
ENSMUST00000122967.3 |
Gprc5c
|
G protein-coupled receptor, family C, group 5, member C |
| chr14_+_48358267 | 2.09 |
ENSMUST00000073150.6
|
Peli2
|
pellino 2 |
| chr19_-_58444336 | 2.07 |
ENSMUST00000131877.2
|
Gfra1
|
glial cell line derived neurotrophic factor family receptor alpha 1 |
| chr9_+_74860133 | 2.06 |
ENSMUST00000215370.2
|
Fam214a
|
family with sequence similarity 214, member A |
| chr2_-_104573179 | 1.98 |
ENSMUST00000028595.8
|
Depdc7
|
DEP domain containing 7 |
| chr14_+_73475335 | 1.98 |
ENSMUST00000044405.8
|
Lpar6
|
lysophosphatidic acid receptor 6 |
| chr9_-_44714263 | 1.97 |
ENSMUST00000044694.8
|
Ttc36
|
tetratricopeptide repeat domain 36 |
| chr13_+_112604037 | 1.94 |
ENSMUST00000183868.8
|
Il6st
|
interleukin 6 signal transducer |
| chr17_-_31383976 | 1.93 |
ENSMUST00000235870.2
|
Tff1
|
trefoil factor 1 |
| chr11_-_68277631 | 1.88 |
ENSMUST00000021284.4
|
Ntn1
|
netrin 1 |
| chr10_-_93146937 | 1.88 |
ENSMUST00000008542.12
|
Elk3
|
ELK3, member of ETS oncogene family |
| chr18_+_51250748 | 1.87 |
ENSMUST00000116639.4
|
Prr16
|
proline rich 16 |
| chr2_+_164245114 | 1.84 |
ENSMUST00000017151.2
|
Rbpjl
|
recombination signal binding protein for immunoglobulin kappa J region-like |
| chr5_+_102629365 | 1.83 |
ENSMUST00000112854.8
|
Arhgap24
|
Rho GTPase activating protein 24 |
| chr2_+_71884943 | 1.81 |
ENSMUST00000028525.6
|
Rapgef4
|
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr5_-_66238313 | 1.81 |
ENSMUST00000202700.4
ENSMUST00000094757.9 ENSMUST00000113724.6 |
Rbm47
|
RNA binding motif protein 47 |
| chr13_+_89687915 | 1.74 |
ENSMUST00000022108.9
|
Hapln1
|
hyaluronan and proteoglycan link protein 1 |
| chr10_+_96453408 | 1.73 |
ENSMUST00000218953.2
|
Btg1
|
BTG anti-proliferation factor 1 |
| chr1_+_87501721 | 1.73 |
ENSMUST00000166259.8
ENSMUST00000172222.8 ENSMUST00000163606.8 |
Neu2
|
neuraminidase 2 |
| chr15_+_59520199 | 1.73 |
ENSMUST00000067543.8
|
Trib1
|
tribbles pseudokinase 1 |
| chr1_-_87501548 | 1.71 |
ENSMUST00000068681.12
|
Ngef
|
neuronal guanine nucleotide exchange factor |
| chr7_-_100306160 | 1.67 |
ENSMUST00000107046.8
ENSMUST00000107045.9 ENSMUST00000139708.9 |
Plekhb1
|
pleckstrin homology domain containing, family B (evectins) member 1 |
| chr14_-_100522101 | 1.64 |
ENSMUST00000228216.2
|
Klf12
|
Kruppel-like factor 12 |
| chr2_+_158148413 | 1.63 |
ENSMUST00000109491.8
ENSMUST00000016168.9 |
Lbp
|
lipopolysaccharide binding protein |
| chr12_-_32000209 | 1.61 |
ENSMUST00000176084.2
ENSMUST00000176103.8 ENSMUST00000167458.9 |
Hbp1
|
high mobility group box transcription factor 1 |
| chr19_+_56414114 | 1.61 |
ENSMUST00000238892.2
|
Casp7
|
caspase 7 |
| chr13_-_60324755 | 1.55 |
ENSMUST00000223933.2
|
Gas1
|
growth arrest specific 1 |
| chr4_+_102112189 | 1.52 |
ENSMUST00000106908.9
|
Pde4b
|
phosphodiesterase 4B, cAMP specific |
| chr1_+_132226308 | 1.51 |
ENSMUST00000046071.5
|
Klhdc8a
|
kelch domain containing 8A |
| chr10_-_41487315 | 1.50 |
ENSMUST00000219054.2
|
Ccdc162
|
coiled-coil domain containing 162 |
| chr11_+_108814007 | 1.44 |
ENSMUST00000106711.2
|
Axin2
|
axin 2 |
| chrX_+_93278526 | 1.40 |
ENSMUST00000113908.8
ENSMUST00000113916.10 |
Klhl15
|
kelch-like 15 |
| chr12_-_32000169 | 1.37 |
ENSMUST00000176520.8
|
Hbp1
|
high mobility group box transcription factor 1 |
| chr1_-_170695328 | 1.37 |
ENSMUST00000027974.7
|
Atf6
|
activating transcription factor 6 |
| chrX_+_93278203 | 1.37 |
ENSMUST00000153900.8
|
Klhl15
|
kelch-like 15 |
| chr1_+_34275665 | 1.36 |
ENSMUST00000194192.3
|
Dst
|
dystonin |
| chr4_+_34893772 | 1.34 |
ENSMUST00000029975.10
ENSMUST00000135871.8 ENSMUST00000108130.2 |
Cga
|
glycoprotein hormones, alpha subunit |
| chr1_+_157353696 | 1.34 |
ENSMUST00000111700.8
|
Sec16b
|
SEC16 homolog B (S. cerevisiae) |
| chr4_-_103072343 | 1.33 |
ENSMUST00000150285.8
|
Slc35d1
|
solute carrier family 35 (UDP-glucuronic acid/UDP-N-acetylgalactosamine dual transporter), member D1 |
| chr6_-_87365859 | 1.30 |
ENSMUST00000032127.6
|
Gkn3
|
gastrokine 3 |
| chr11_-_46581135 | 1.29 |
ENSMUST00000169584.8
|
Timd2
|
T cell immunoglobulin and mucin domain containing 2 |
| chr3_-_131138541 | 1.29 |
ENSMUST00000090246.5
ENSMUST00000126569.2 |
Sgms2
|
sphingomyelin synthase 2 |
| chr12_-_108241597 | 1.29 |
ENSMUST00000222310.2
|
Ccdc85c
|
coiled-coil domain containing 85C |
| chr14_+_20732804 | 1.28 |
ENSMUST00000228545.2
|
Sec24c
|
Sec24 related gene family, member C (S. cerevisiae) |
| chr9_-_53447908 | 1.26 |
ENSMUST00000150244.2
|
Atm
|
ataxia telangiectasia mutated |
| chr1_+_75119472 | 1.26 |
ENSMUST00000189650.7
|
Retreg2
|
reticulophagy regulator family member 2 |
| chr6_-_59001455 | 1.25 |
ENSMUST00000089860.12
|
Fam13a
|
family with sequence similarity 13, member A |
| chr10_-_93146825 | 1.23 |
ENSMUST00000151153.2
|
Elk3
|
ELK3, member of ETS oncogene family |
| chr9_-_59260713 | 1.22 |
ENSMUST00000026265.8
|
Bbs4
|
Bardet-Biedl syndrome 4 (human) |
| chr7_+_28524627 | 1.22 |
ENSMUST00000066264.13
|
Ech1
|
enoyl coenzyme A hydratase 1, peroxisomal |
| chr3_-_82811269 | 1.20 |
ENSMUST00000029632.7
|
Lrat
|
lecithin-retinol acyltransferase (phosphatidylcholine-retinol-O-acyltransferase) |
| chr15_+_59520493 | 1.18 |
ENSMUST00000118228.2
|
Trib1
|
tribbles pseudokinase 1 |
| chr2_-_51863203 | 1.18 |
ENSMUST00000028314.9
|
Nmi
|
N-myc (and STAT) interactor |
| chr5_-_90788323 | 1.15 |
ENSMUST00000202784.4
ENSMUST00000031317.10 ENSMUST00000201370.2 |
Rassf6
|
Ras association (RalGDS/AF-6) domain family member 6 |
| chrX_+_93278588 | 1.12 |
ENSMUST00000096369.10
ENSMUST00000113911.9 |
Klhl15
|
kelch-like 15 |
| chr6_+_14901343 | 1.12 |
ENSMUST00000115477.8
|
Foxp2
|
forkhead box P2 |
| chr15_-_50753792 | 1.10 |
ENSMUST00000185183.2
|
Trps1
|
transcriptional repressor GATA binding 1 |
| chr17_-_34847354 | 1.10 |
ENSMUST00000167097.9
|
Ppt2
|
palmitoyl-protein thioesterase 2 |
| chr11_-_86884507 | 1.10 |
ENSMUST00000018571.5
|
Ypel2
|
yippee like 2 |
| chr12_+_112645237 | 1.09 |
ENSMUST00000174780.2
ENSMUST00000169593.2 ENSMUST00000173942.2 |
Zbtb42
|
zinc finger and BTB domain containing 42 |
| chr8_+_107329983 | 1.07 |
ENSMUST00000000312.12
ENSMUST00000167688.2 |
Cdh1
|
cadherin 1 |
| chr8_-_45715049 | 1.07 |
ENSMUST00000034064.5
|
F11
|
coagulation factor XI |
| chr9_+_44309727 | 1.06 |
ENSMUST00000213268.2
|
Slc37a4
|
solute carrier family 37 (glucose-6-phosphate transporter), member 4 |
| chr11_-_70350783 | 1.05 |
ENSMUST00000019064.9
|
Cxcl16
|
chemokine (C-X-C motif) ligand 16 |
| chr16_-_43836681 | 1.05 |
ENSMUST00000036174.10
|
Gramd1c
|
GRAM domain containing 1C |
| chr7_-_80053063 | 1.03 |
ENSMUST00000147150.2
|
Furin
|
furin (paired basic amino acid cleaving enzyme) |
| chr1_+_75119419 | 1.03 |
ENSMUST00000097694.11
ENSMUST00000190240.7 |
Retreg2
|
reticulophagy regulator family member 2 |
| chr3_+_85946145 | 1.02 |
ENSMUST00000238331.2
|
Sh3d19
|
SH3 domain protein D19 |
| chr2_+_120807498 | 1.00 |
ENSMUST00000067582.14
|
Tmem62
|
transmembrane protein 62 |
| chr17_+_48037758 | 1.00 |
ENSMUST00000024782.12
ENSMUST00000144955.2 |
Pgc
|
progastricsin (pepsinogen C) |
| chr15_-_3333003 | 0.99 |
ENSMUST00000165386.2
|
Ccdc152
|
coiled-coil domain containing 152 |
| chr1_-_74924481 | 0.98 |
ENSMUST00000159232.2
ENSMUST00000068631.4 |
Fev
|
FEV transcription factor, ETS family member |
| chr5_-_90788460 | 0.97 |
ENSMUST00000202704.4
|
Rassf6
|
Ras association (RalGDS/AF-6) domain family member 6 |
| chr6_-_134768288 | 0.97 |
ENSMUST00000149776.3
|
Dusp16
|
dual specificity phosphatase 16 |
| chr14_-_100521888 | 0.95 |
ENSMUST00000226774.2
|
Klf12
|
Kruppel-like factor 12 |
| chr7_-_65020655 | 0.93 |
ENSMUST00000032729.8
|
Tjp1
|
tight junction protein 1 |
| chr9_-_53448028 | 0.91 |
ENSMUST00000232179.2
|
Atm
|
ataxia telangiectasia mutated |
| chr12_-_103623418 | 0.91 |
ENSMUST00000044159.7
|
Serpina6
|
serine (or cysteine) peptidase inhibitor, clade A, member 6 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.7 | 22.7 | GO:0010901 | regulation of very-low-density lipoprotein particle remodeling(GO:0010901) |
| 3.4 | 10.3 | GO:0006867 | asparagine transport(GO:0006867) positive regulation of glutamine transport(GO:2000487) |
| 3.3 | 23.0 | GO:0060455 | negative regulation of gastric acid secretion(GO:0060455) |
| 3.2 | 16.1 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 2.5 | 15.0 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 2.4 | 14.5 | GO:0090361 | platelet-derived growth factor production(GO:0090360) regulation of platelet-derived growth factor production(GO:0090361) |
| 2.4 | 9.6 | GO:0001970 | positive regulation of activation of membrane attack complex(GO:0001970) |
| 2.1 | 10.4 | GO:0009441 | glycolate metabolic process(GO:0009441) |
| 1.9 | 9.7 | GO:1904253 | positive regulation of bile acid biosynthetic process(GO:0070859) positive regulation of bile acid metabolic process(GO:1904253) |
| 1.9 | 5.7 | GO:0006553 | lysine metabolic process(GO:0006553) |
| 1.6 | 9.8 | GO:0010248 | establishment or maintenance of transmembrane electrochemical gradient(GO:0010248) |
| 1.3 | 7.5 | GO:1903243 | negative regulation of cardiac muscle adaptation(GO:0010616) negative regulation of cardiac muscle hypertrophy in response to stress(GO:1903243) |
| 1.2 | 3.7 | GO:0002276 | basophil activation involved in immune response(GO:0002276) |
| 1.2 | 4.8 | GO:0015851 | nucleobase transport(GO:0015851) pyrimidine nucleobase transport(GO:0015855) |
| 1.2 | 18.9 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 1.1 | 3.4 | GO:1902524 | positive regulation of protein K48-linked ubiquitination(GO:1902524) |
| 1.1 | 15.4 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 1.1 | 7.7 | GO:0050847 | progesterone receptor signaling pathway(GO:0050847) |
| 1.0 | 2.9 | GO:0045643 | regulation of eosinophil differentiation(GO:0045643) positive regulation of eosinophil differentiation(GO:0045645) |
| 0.9 | 6.6 | GO:0002925 | positive regulation of humoral immune response mediated by circulating immunoglobulin(GO:0002925) heme transport(GO:0015886) positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) |
| 0.9 | 2.7 | GO:0006535 | cysteine biosynthetic process from serine(GO:0006535) |
| 0.9 | 6.3 | GO:1904073 | regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 0.8 | 22.8 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.7 | 3.7 | GO:0042908 | xenobiotic transport(GO:0042908) |
| 0.7 | 2.2 | GO:1900477 | negative regulation of G1/S transition of mitotic cell cycle by negative regulation of transcription from RNA polymerase II promoter(GO:1900477) |
| 0.7 | 2.2 | GO:0072425 | signal transduction involved in G2 DNA damage checkpoint(GO:0072425) signal transduction involved in mitotic G2 DNA damage checkpoint(GO:0072434) |
| 0.7 | 5.0 | GO:0019856 | 'de novo' pyrimidine nucleobase biosynthetic process(GO:0006207) pyrimidine nucleobase biosynthetic process(GO:0019856) |
| 0.7 | 5.0 | GO:1901509 | regulation of endothelial tube morphogenesis(GO:1901509) |
| 0.7 | 4.9 | GO:0050747 | positive regulation of lipoprotein metabolic process(GO:0050747) |
| 0.7 | 4.8 | GO:0060023 | soft palate development(GO:0060023) |
| 0.6 | 16.0 | GO:0017144 | drug metabolic process(GO:0017144) |
| 0.6 | 4.6 | GO:0035948 | positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) |
| 0.6 | 11.2 | GO:0042359 | vitamin D metabolic process(GO:0042359) |
| 0.6 | 3.9 | GO:0071630 | nucleus-associated proteasomal ubiquitin-dependent protein catabolic process(GO:0071630) |
| 0.5 | 0.5 | GO:0071393 | cellular response to progesterone stimulus(GO:0071393) |
| 0.5 | 1.6 | GO:0015920 | lipopolysaccharide transport(GO:0015920) |
| 0.5 | 13.9 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.5 | 3.7 | GO:0036438 | maintenance of lens transparency(GO:0036438) |
| 0.5 | 16.6 | GO:0046688 | response to copper ion(GO:0046688) |
| 0.5 | 5.1 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.5 | 1.0 | GO:0002760 | positive regulation of antimicrobial peptide production(GO:0002225) positive regulation of antimicrobial humoral response(GO:0002760) positive regulation of antibacterial peptide production(GO:0002803) |
| 0.5 | 3.2 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.5 | 1.8 | GO:1904457 | positive regulation of neuronal action potential(GO:1904457) |
| 0.4 | 4.6 | GO:0015868 | purine ribonucleotide transport(GO:0015868) |
| 0.4 | 3.2 | GO:0097647 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.4 | 1.2 | GO:0021529 | spinal cord oligodendrocyte cell differentiation(GO:0021529) spinal cord oligodendrocyte cell fate specification(GO:0021530) |
| 0.4 | 1.2 | GO:1990164 | histone H3-S28 phosphorylation(GO:0043988) histone H2A phosphorylation(GO:1990164) |
| 0.4 | 21.5 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.4 | 2.8 | GO:0031179 | peptide modification(GO:0031179) |
| 0.3 | 4.2 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.3 | 1.6 | GO:0072733 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 0.3 | 0.6 | GO:2000686 | regulation of rubidium ion transmembrane transporter activity(GO:2000686) |
| 0.3 | 1.2 | GO:0038108 | negative regulation of appetite by leptin-mediated signaling pathway(GO:0038108) |
| 0.3 | 3.5 | GO:0043031 | negative regulation of macrophage activation(GO:0043031) |
| 0.3 | 7.4 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.3 | 2.2 | GO:0001887 | selenium compound metabolic process(GO:0001887) |
| 0.3 | 3.5 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.3 | 5.1 | GO:0006309 | apoptotic DNA fragmentation(GO:0006309) |
| 0.3 | 1.1 | GO:0032915 | positive regulation of transforming growth factor beta2 production(GO:0032915) |
| 0.3 | 4.8 | GO:0060628 | regulation of ER to Golgi vesicle-mediated transport(GO:0060628) |
| 0.3 | 2.6 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.3 | 1.0 | GO:0090472 | dibasic protein processing(GO:0090472) |
| 0.2 | 2.7 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.2 | 1.4 | GO:0032423 | regulation of mismatch repair(GO:0032423) negative regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000054) |
| 0.2 | 2.5 | GO:0033564 | Cdc42 protein signal transduction(GO:0032488) anterior/posterior axon guidance(GO:0033564) |
| 0.2 | 3.2 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.2 | 1.3 | GO:0048207 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.2 | 33.7 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.2 | 0.7 | GO:0090187 | positive regulation of pancreatic juice secretion(GO:0090187) |
| 0.2 | 1.7 | GO:2000271 | positive regulation of fibroblast apoptotic process(GO:2000271) |
| 0.2 | 1.7 | GO:0006689 | ganglioside catabolic process(GO:0006689) oligosaccharide catabolic process(GO:0009313) |
| 0.2 | 4.1 | GO:1990001 | inhibition of cysteine-type endopeptidase activity involved in apoptotic process(GO:1990001) |
| 0.2 | 4.0 | GO:0051132 | NK T cell activation(GO:0051132) |
| 0.2 | 2.4 | GO:0061113 | pancreas morphogenesis(GO:0061113) |
| 0.2 | 1.0 | GO:0051611 | regulation of serotonin uptake(GO:0051611) |
| 0.2 | 2.4 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.2 | 1.1 | GO:0060066 | oviduct development(GO:0060066) |
| 0.2 | 0.7 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.2 | 6.3 | GO:0045780 | positive regulation of bone resorption(GO:0045780) positive regulation of bone remodeling(GO:0046852) |
| 0.2 | 0.9 | GO:0038165 | oncostatin-M-mediated signaling pathway(GO:0038165) |
| 0.2 | 1.3 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.1 | 0.6 | GO:0010159 | specification of organ position(GO:0010159) |
| 0.1 | 0.8 | GO:0060011 | Sertoli cell proliferation(GO:0060011) |
| 0.1 | 2.6 | GO:0010745 | negative regulation of macrophage derived foam cell differentiation(GO:0010745) |
| 0.1 | 1.3 | GO:0030206 | chondroitin sulfate biosynthetic process(GO:0030206) |
| 0.1 | 1.6 | GO:0032275 | luteinizing hormone secretion(GO:0032275) |
| 0.1 | 5.7 | GO:0002089 | lens morphogenesis in camera-type eye(GO:0002089) |
| 0.1 | 5.2 | GO:0006767 | water-soluble vitamin metabolic process(GO:0006767) |
| 0.1 | 0.4 | GO:0021759 | globus pallidus development(GO:0021759) |
| 0.1 | 0.6 | GO:1903445 | intermembrane transport(GO:0046909) protein transport from ciliary membrane to plasma membrane(GO:1903445) |
| 0.1 | 1.1 | GO:0035166 | post-embryonic hemopoiesis(GO:0035166) |
| 0.1 | 1.3 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.1 | 1.7 | GO:0060501 | positive regulation of epithelial cell proliferation involved in lung morphogenesis(GO:0060501) |
| 0.1 | 3.4 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.1 | 1.9 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.1 | 0.9 | GO:0060665 | regulation of branching involved in salivary gland morphogenesis by mesenchymal-epithelial signaling(GO:0060665) |
| 0.1 | 1.8 | GO:0014850 | response to muscle activity(GO:0014850) |
| 0.1 | 1.7 | GO:0061002 | negative regulation of dendritic spine morphogenesis(GO:0061002) |
| 0.1 | 2.4 | GO:0045947 | negative regulation of translational initiation(GO:0045947) |
| 0.1 | 1.4 | GO:1990440 | positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.1 | 0.8 | GO:0035887 | aortic smooth muscle cell differentiation(GO:0035887) |
| 0.1 | 13.4 | GO:0055088 | lipid homeostasis(GO:0055088) |
| 0.1 | 12.6 | GO:0009166 | nucleotide catabolic process(GO:0009166) |
| 0.1 | 0.3 | GO:0071698 | olfactory placode formation(GO:0030910) olfactory placode development(GO:0071698) olfactory placode morphogenesis(GO:0071699) |
| 0.1 | 1.6 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.1 | 0.4 | GO:0060346 | bone trabecula formation(GO:0060346) |
| 0.1 | 1.9 | GO:0045793 | positive regulation of cell size(GO:0045793) |
| 0.1 | 0.2 | GO:2000328 | regulation of T-helper 17 cell lineage commitment(GO:2000328) |
| 0.1 | 4.4 | GO:0003298 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.1 | 1.3 | GO:0072189 | ureter development(GO:0072189) |
| 0.1 | 1.5 | GO:0045792 | negative regulation of cell size(GO:0045792) |
| 0.1 | 0.9 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.1 | 5.5 | GO:0006940 | regulation of smooth muscle contraction(GO:0006940) |
| 0.1 | 0.3 | GO:0019262 | N-acetylneuraminate catabolic process(GO:0019262) |
| 0.1 | 0.5 | GO:0030578 | PML body organization(GO:0030578) positive regulation of leukocyte adhesion to vascular endothelial cell(GO:1904996) |
| 0.1 | 0.5 | GO:0007256 | activation of JNKK activity(GO:0007256) negative regulation of defense response to bacterium(GO:1900425) |
| 0.1 | 0.2 | GO:0071895 | odontoblast differentiation(GO:0071895) |
| 0.1 | 0.9 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.1 | 0.8 | GO:0015697 | quaternary ammonium group transport(GO:0015697) |
| 0.1 | 0.2 | GO:0021937 | cerebellar Purkinje cell-granule cell precursor cell signaling involved in regulation of granule cell precursor cell proliferation(GO:0021937) |
| 0.1 | 0.2 | GO:0016332 | establishment or maintenance of polarity of embryonic epithelium(GO:0016332) |
| 0.1 | 0.6 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.1 | 0.7 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 0.1 | 0.3 | GO:0010157 | response to chlorate(GO:0010157) |
| 0.1 | 0.4 | GO:0021798 | forebrain dorsal/ventral pattern formation(GO:0021798) |
| 0.1 | 0.5 | GO:0002669 | positive regulation of T cell anergy(GO:0002669) positive regulation of lymphocyte anergy(GO:0002913) |
| 0.1 | 0.5 | GO:0043116 | negative regulation of vascular permeability(GO:0043116) |
| 0.1 | 0.5 | GO:0008228 | opsonization(GO:0008228) |
| 0.1 | 0.2 | GO:0052564 | interleukin-15 production(GO:0032618) response to immune response of other organism involved in symbiotic interaction(GO:0052564) response to host immune response(GO:0052572) |
| 0.1 | 3.7 | GO:0045669 | positive regulation of osteoblast differentiation(GO:0045669) |
| 0.0 | 2.6 | GO:0001958 | endochondral ossification(GO:0001958) replacement ossification(GO:0036075) |
| 0.0 | 12.6 | GO:0010466 | negative regulation of peptidase activity(GO:0010466) |
| 0.0 | 0.8 | GO:0042048 | olfactory behavior(GO:0042048) |
| 0.0 | 1.3 | GO:0097286 | iron ion import(GO:0097286) |
| 0.0 | 0.2 | GO:0046604 | positive regulation of mitotic centrosome separation(GO:0046604) |
| 0.0 | 1.0 | GO:2000114 | regulation of establishment of cell polarity(GO:2000114) |
| 0.0 | 0.3 | GO:1904222 | regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904217) positive regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904219) positive regulation of serine C-palmitoyltransferase activity(GO:1904222) |
| 0.0 | 0.1 | GO:0097535 | lymphoid lineage cell migration(GO:0097534) lymphoid lineage cell migration into thymus(GO:0097535) |
| 0.0 | 2.1 | GO:0006094 | gluconeogenesis(GO:0006094) |
| 0.0 | 0.1 | GO:1904016 | response to Thyroglobulin triiodothyronine(GO:1904016) cellular response to Thyroglobulin triiodothyronine(GO:1904017) |
| 0.0 | 0.3 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.0 | 4.2 | GO:0035725 | sodium ion transmembrane transport(GO:0035725) |
| 0.0 | 0.5 | GO:0060746 | parental behavior(GO:0060746) |
| 0.0 | 0.4 | GO:1904778 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.0 | 0.3 | GO:0090336 | positive regulation of brown fat cell differentiation(GO:0090336) |
| 0.0 | 0.3 | GO:0021559 | trigeminal nerve development(GO:0021559) |
| 0.0 | 0.2 | GO:0070268 | cornification(GO:0070268) |
| 0.0 | 1.1 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.0 | 0.6 | GO:0007035 | vacuolar acidification(GO:0007035) |
| 0.0 | 0.0 | GO:0022601 | menstrual cycle phase(GO:0022601) |
| 0.0 | 0.5 | GO:1902230 | negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902230) |
| 0.0 | 1.0 | GO:0043550 | regulation of lipid kinase activity(GO:0043550) |
| 0.0 | 0.1 | GO:0033634 | positive regulation of cell-cell adhesion mediated by integrin(GO:0033634) negative regulation of endothelial cell chemotaxis(GO:2001027) |
| 0.0 | 0.3 | GO:0008211 | glucocorticoid metabolic process(GO:0008211) |
| 0.0 | 0.3 | GO:0003215 | cardiac right ventricle morphogenesis(GO:0003215) |
| 0.0 | 0.9 | GO:0051058 | negative regulation of Ras protein signal transduction(GO:0046580) negative regulation of small GTPase mediated signal transduction(GO:0051058) |
| 0.0 | 1.5 | GO:0033077 | T cell differentiation in thymus(GO:0033077) thymocyte aggregation(GO:0071594) |
| 0.0 | 0.3 | GO:0003351 | epithelial cilium movement(GO:0003351) |
| 0.0 | 0.2 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.0 | 1.9 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.0 | 0.0 | GO:0035038 | female pronucleus assembly(GO:0035038) |
| 0.0 | 0.1 | GO:0042706 | eye photoreceptor cell fate commitment(GO:0042706) photoreceptor cell fate commitment(GO:0046552) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.5 | 13.5 | GO:0034359 | mature chylomicron(GO:0034359) |
| 2.1 | 12.7 | GO:0044218 | other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 1.9 | 18.5 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 1.5 | 12.4 | GO:0005579 | membrane attack complex(GO:0005579) |
| 1.2 | 13.5 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 1.2 | 12.9 | GO:0005796 | Golgi lumen(GO:0005796) |
| 1.1 | 22.7 | GO:0042627 | chylomicron(GO:0042627) |
| 1.1 | 13.9 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.9 | 2.8 | GO:0005900 | oncostatin-M receptor complex(GO:0005900) |
| 0.9 | 2.6 | GO:0043540 | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase complex(GO:0043540) |
| 0.7 | 18.9 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 0.5 | 5.5 | GO:0030478 | actin cap(GO:0030478) |
| 0.4 | 1.8 | GO:0044316 | cone cell pedicle(GO:0044316) |
| 0.3 | 6.1 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.3 | 2.5 | GO:0035032 | phosphatidylinositol 3-kinase complex, class III(GO:0035032) |
| 0.3 | 19.2 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.2 | 4.9 | GO:0005922 | connexon complex(GO:0005922) |
| 0.1 | 1.5 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.1 | 0.5 | GO:0000221 | vacuolar proton-transporting V-type ATPase, V1 domain(GO:0000221) |
| 0.1 | 0.3 | GO:0016939 | kinesin II complex(GO:0016939) |
| 0.1 | 1.2 | GO:0034464 | BBSome(GO:0034464) |
| 0.1 | 14.7 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.1 | 37.1 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.1 | 5.0 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 2.2 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.1 | 14.4 | GO:0031227 | intrinsic component of endoplasmic reticulum membrane(GO:0031227) |
| 0.1 | 1.3 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.1 | 4.9 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.1 | 0.5 | GO:0031673 | H zone(GO:0031673) |
| 0.1 | 2.8 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 0.8 | GO:0097136 | Bcl-2 family protein complex(GO:0097136) |
| 0.1 | 1.0 | GO:0012510 | trans-Golgi network transport vesicle membrane(GO:0012510) |
| 0.1 | 0.9 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.1 | 0.7 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.1 | 5.2 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 11.6 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.1 | 0.9 | GO:0071437 | invadopodium(GO:0071437) |
| 0.1 | 107.9 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 1.2 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.0 | 0.9 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 2.2 | GO:1990391 | DNA repair complex(GO:1990391) |
| 0.0 | 0.4 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.0 | 0.8 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 6.8 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.0 | 2.2 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 0.3 | GO:0045178 | basal part of cell(GO:0045178) |
| 0.0 | 4.0 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 2.5 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 1.5 | GO:0005747 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.2 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.0 | 0.1 | GO:0001652 | granular component(GO:0001652) |
| 0.0 | 0.5 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.0 | 0.5 | GO:0030904 | retromer complex(GO:0030904) |
| 0.0 | 0.3 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 0.0 | 4.6 | GO:0005938 | cell cortex(GO:0005938) |
| 0.0 | 0.2 | GO:0033268 | node of Ranvier(GO:0033268) |
| 0.0 | 0.6 | GO:0031201 | SNARE complex(GO:0031201) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.6 | 25.3 | GO:0004556 | alpha-amylase activity(GO:0004556) |
| 3.5 | 10.6 | GO:1901375 | acetylcholine transmembrane transporter activity(GO:0005277) norepinephrine transmembrane transporter activity(GO:0005333) acetate ester transmembrane transporter activity(GO:1901375) |
| 3.4 | 20.3 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 3.0 | 15.0 | GO:0004925 | prolactin receptor activity(GO:0004925) |
| 2.8 | 11.2 | GO:1902271 | D3 vitamins binding(GO:1902271) |
| 2.6 | 10.4 | GO:0016899 | oxidoreductase activity, acting on the CH-OH group of donors, oxygen as acceptor(GO:0016899) |
| 2.6 | 10.3 | GO:0015182 | L-asparagine transmembrane transporter activity(GO:0015182) |
| 2.2 | 13.5 | GO:0004850 | uridine phosphorylase activity(GO:0004850) |
| 2.2 | 17.7 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 1.7 | 5.0 | GO:0004087 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 1.7 | 23.1 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 1.4 | 15.5 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 1.4 | 42.3 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 1.3 | 7.7 | GO:0047023 | androsterone dehydrogenase activity(GO:0047023) |
| 1.2 | 13.5 | GO:0035473 | lipase binding(GO:0035473) |
| 1.1 | 6.6 | GO:0015232 | heme transporter activity(GO:0015232) |
| 1.1 | 6.3 | GO:0030294 | receptor signaling protein tyrosine kinase inhibitor activity(GO:0030294) |
| 1.0 | 5.1 | GO:0004531 | deoxyribonuclease II activity(GO:0004531) |
| 1.0 | 4.8 | GO:0015205 | purine nucleobase transmembrane transporter activity(GO:0005345) pyrimidine nucleobase transmembrane transporter activity(GO:0005350) nucleobase transmembrane transporter activity(GO:0015205) glycerol channel activity(GO:0015254) |
| 0.9 | 18.9 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.9 | 3.7 | GO:0035651 | AP-3 adaptor complex binding(GO:0035651) |
| 0.9 | 3.7 | GO:0008109 | N-acetyllactosaminide beta-1,6-N-acetylglucosaminyltransferase activity(GO:0008109) |
| 0.9 | 2.7 | GO:0050421 | cystathionine beta-synthase activity(GO:0004122) oxidoreductase activity, acting on other nitrogenous compounds as donors, cytochrome as acceptor(GO:0016662) nitrite reductase (NO-forming) activity(GO:0050421) carbon monoxide binding(GO:0070025) nitric oxide binding(GO:0070026) |
| 0.9 | 6.2 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.8 | 3.2 | GO:0019862 | IgA binding(GO:0019862) |
| 0.8 | 2.3 | GO:0034191 | apolipoprotein A-I receptor binding(GO:0034191) |
| 0.8 | 3.1 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.7 | 2.2 | GO:0004677 | DNA-dependent protein kinase activity(GO:0004677) |
| 0.7 | 2.8 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.6 | 1.9 | GO:0004915 | interleukin-6 receptor activity(GO:0004915) interleukin-6 binding(GO:0019981) |
| 0.6 | 8.9 | GO:1990405 | protein antigen binding(GO:1990405) |
| 0.5 | 2.2 | GO:0005118 | sevenless binding(GO:0005118) |
| 0.5 | 19.4 | GO:0008392 | arachidonic acid epoxygenase activity(GO:0008392) |
| 0.5 | 4.0 | GO:0005534 | galactose binding(GO:0005534) |
| 0.4 | 4.9 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.4 | 1.3 | GO:0070287 | ferritin receptor activity(GO:0070287) |
| 0.4 | 12.3 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.4 | 1.1 | GO:0015152 | glucose-6-phosphate transmembrane transporter activity(GO:0015152) |
| 0.3 | 1.7 | GO:0052796 | exo-alpha-sialidase activity(GO:0004308) alpha-sialidase activity(GO:0016997) exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.3 | 19.2 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.3 | 2.6 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.3 | 4.6 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.3 | 4.4 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.3 | 1.6 | GO:0070891 | lipoteichoic acid binding(GO:0070891) |
| 0.2 | 4.9 | GO:0044548 | S100 protein binding(GO:0044548) |
| 0.2 | 9.7 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.2 | 0.9 | GO:0004924 | oncostatin-M receptor activity(GO:0004924) |
| 0.2 | 1.3 | GO:0047493 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.2 | 5.7 | GO:0016646 | oxidoreductase activity, acting on the CH-NH group of donors, NAD or NADP as acceptor(GO:0016646) |
| 0.2 | 29.0 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.2 | 2.3 | GO:0001846 | opsonin binding(GO:0001846) |
| 0.2 | 2.4 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.1 | 0.9 | GO:0016433 | rRNA (adenine) methyltransferase activity(GO:0016433) |
| 0.1 | 1.3 | GO:0015165 | pyrimidine nucleotide-sugar transmembrane transporter activity(GO:0015165) |
| 0.1 | 2.9 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.1 | 1.2 | GO:0034452 | dynactin binding(GO:0034452) |
| 0.1 | 2.7 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.1 | 1.8 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.1 | 2.3 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.1 | 16.7 | GO:0015171 | amino acid transmembrane transporter activity(GO:0015171) |
| 0.1 | 2.2 | GO:0008430 | selenium binding(GO:0008430) |
| 0.1 | 2.4 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.1 | 4.9 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.1 | 5.7 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.1 | 1.1 | GO:0098599 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.1 | 0.9 | GO:0071253 | connexin binding(GO:0071253) |
| 0.1 | 0.2 | GO:0030284 | estrogen receptor activity(GO:0030284) |
| 0.1 | 1.7 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 0.9 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.1 | 1.1 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.1 | 1.2 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.1 | 5.2 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.1 | 0.2 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.1 | 1.4 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.1 | 5.9 | GO:0098531 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) transcription factor activity, direct ligand regulated sequence-specific DNA binding(GO:0098531) |
| 0.1 | 2.7 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.1 | 0.2 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 0.1 | 4.1 | GO:0019213 | deacetylase activity(GO:0019213) |
| 0.1 | 0.4 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.1 | 0.5 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.1 | 0.5 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.0 | 1.5 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.1 | GO:0010698 | acetyltransferase activator activity(GO:0010698) |
| 0.0 | 2.8 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 0.1 | GO:0052597 | diamine oxidase activity(GO:0052597) histamine oxidase activity(GO:0052598) methylputrescine oxidase activity(GO:0052599) propane-1,3-diamine oxidase activity(GO:0052600) |
| 0.0 | 0.6 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.0 | 0.6 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.0 | 1.6 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.0 | 0.3 | GO:0005119 | smoothened binding(GO:0005119) hedgehog receptor activity(GO:0008158) |
| 0.0 | 0.5 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.0 | 1.5 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 0.7 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.0 | 1.0 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.0 | 0.8 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.0 | 5.6 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.4 | GO:0042731 | PH domain binding(GO:0042731) |
| 0.0 | 6.6 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.0 | 0.4 | GO:0044213 | intronic transcription regulatory region sequence-specific DNA binding(GO:0001161) intronic transcription regulatory region DNA binding(GO:0044213) |
| 0.0 | 0.1 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.0 | 1.1 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 0.3 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.0 | 7.8 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 0.3 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.2 | GO:0008174 | mRNA methyltransferase activity(GO:0008174) |
| 0.0 | 0.2 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.0 | 0.5 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.7 | GO:0050750 | low-density lipoprotein particle receptor binding(GO:0050750) |
| 0.0 | 0.6 | GO:0005521 | lamin binding(GO:0005521) |
| 0.0 | 6.7 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 0.6 | GO:0046961 | proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.0 | 0.7 | GO:0001221 | transcription cofactor binding(GO:0001221) |
| 0.0 | 0.8 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.0 | 0.9 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.8 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.0 | 0.5 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.0 | 9.8 | GO:0000977 | RNA polymerase II regulatory region sequence-specific DNA binding(GO:0000977) RNA polymerase II regulatory region DNA binding(GO:0001012) |
| 0.0 | 0.6 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.5 | GO:0016836 | hydro-lyase activity(GO:0016836) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 24.6 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.2 | 10.5 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.2 | 4.9 | PID IGF1 PATHWAY | IGF1 pathway |
| 0.2 | 15.0 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.2 | 1.6 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.2 | 8.6 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 0.9 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.1 | 12.9 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.1 | 3.7 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.1 | 9.3 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.1 | 11.2 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.1 | 21.9 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.1 | 6.1 | PID FGF PATHWAY | FGF signaling pathway |
| 0.1 | 3.3 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.1 | 1.4 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.1 | 1.9 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.1 | 5.2 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 2.2 | PID ATM PATHWAY | ATM pathway |
| 0.1 | 1.6 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.1 | 3.4 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 2.5 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 0.9 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 1.2 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.0 | 0.9 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.0 | 7.9 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.5 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 1.5 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 0.9 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.0 | 1.1 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 0.3 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 0.8 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 0.7 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.0 | 3.5 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.4 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.0 | 0.3 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 0.8 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.3 | 16.0 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 1.6 | 17.7 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 1.2 | 36.1 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.8 | 17.6 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.8 | 10.5 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.7 | 13.5 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.5 | 9.7 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 7ALPHA HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 7alpha-hydroxycholesterol |
| 0.5 | 31.9 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.5 | 14.3 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.4 | 15.3 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.3 | 4.8 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.3 | 7.1 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.3 | 12.2 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.3 | 15.0 | REACTOME GROWTH HORMONE RECEPTOR SIGNALING | Genes involved in Growth hormone receptor signaling |
| 0.3 | 5.3 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.2 | 1.9 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.2 | 4.8 | REACTOME FGFR4 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR4 ligand binding and activation |
| 0.2 | 4.6 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.2 | 2.5 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.2 | 3.6 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.2 | 2.2 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.1 | 1.2 | REACTOME METABOLISM OF STEROID HORMONES AND VITAMINS A AND D | Genes involved in Metabolism of steroid hormones and vitamins A and D |
| 0.1 | 1.4 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.1 | 4.6 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 1.3 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.1 | 2.4 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.1 | 1.0 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.1 | 3.5 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.1 | 5.5 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 4.4 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.1 | 1.7 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.1 | 2.0 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.1 | 5.6 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 0.6 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.1 | 1.1 | REACTOME IRAK1 RECRUITS IKK COMPLEX | Genes involved in IRAK1 recruits IKK complex |
| 0.1 | 2.7 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.1 | 18.3 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.1 | 1.3 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.1 | 6.1 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.1 | 2.6 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 2.8 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.1 | 1.8 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.1 | 0.9 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 1.6 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 1.2 | REACTOME NCAM SIGNALING FOR NEURITE OUT GROWTH | Genes involved in NCAM signaling for neurite out-growth |
| 0.0 | 3.4 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 0.5 | REACTOME NEF MEDIATES DOWN MODULATION OF CELL SURFACE RECEPTORS BY RECRUITING THEM TO CLATHRIN ADAPTERS | Genes involved in Nef-mediates down modulation of cell surface receptors by recruiting them to clathrin adapters |
| 0.0 | 0.5 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.0 | 1.7 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 7.4 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.7 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 1.3 | REACTOME SPHINGOLIPID METABOLISM | Genes involved in Sphingolipid metabolism |
| 0.0 | 0.4 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.0 | 0.7 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.0 | 1.1 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 0.8 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.4 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |