Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
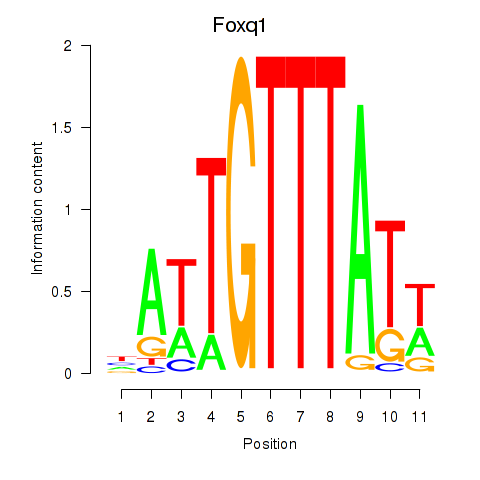
Results for Foxq1
Z-value: 0.50
Transcription factors associated with Foxq1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Foxq1
|
ENSMUSG00000038415.11 | Foxq1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Foxq1 | mm39_v1_chr13_+_31740117_31740117 | -0.33 | 5.3e-02 | Click! |
Activity profile of Foxq1 motif
Sorted Z-values of Foxq1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Foxq1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_87724533 | 1.76 |
ENSMUST00000126353.8
ENSMUST00000149357.8 |
Mllt3
|
myeloid/lymphoid or mixed-lineage leukemia; translocated to, 3 |
| chr7_+_110371811 | 1.57 |
ENSMUST00000005829.13
|
Ampd3
|
adenosine monophosphate deaminase 3 |
| chr4_-_87724512 | 1.38 |
ENSMUST00000148059.2
|
Mllt3
|
myeloid/lymphoid or mixed-lineage leukemia; translocated to, 3 |
| chr3_+_129326004 | 1.35 |
ENSMUST00000199910.5
ENSMUST00000197070.5 ENSMUST00000071402.7 |
Elovl6
|
ELOVL family member 6, elongation of long chain fatty acids (yeast) |
| chr10_+_115653152 | 1.13 |
ENSMUST00000080630.11
ENSMUST00000179196.3 ENSMUST00000035563.15 |
Tspan8
|
tetraspanin 8 |
| chr3_+_135531409 | 0.86 |
ENSMUST00000180196.8
|
Slc39a8
|
solute carrier family 39 (metal ion transporter), member 8 |
| chr14_+_26722319 | 0.86 |
ENSMUST00000035433.10
|
Hesx1
|
homeobox gene expressed in ES cells |
| chr11_-_82719850 | 0.68 |
ENSMUST00000021036.13
ENSMUST00000074515.11 ENSMUST00000103218.3 |
Rffl
|
ring finger and FYVE like domain containing protein |
| chr18_+_11790409 | 0.58 |
ENSMUST00000047322.8
|
Rbbp8
|
retinoblastoma binding protein 8, endonuclease |
| chr15_+_102368510 | 0.57 |
ENSMUST00000164688.2
|
Prr13
|
proline rich 13 |
| chr1_+_51328265 | 0.55 |
ENSMUST00000051572.8
|
Cavin2
|
caveolae associated 2 |
| chr7_+_78563513 | 0.55 |
ENSMUST00000038142.15
|
Isg20
|
interferon-stimulated protein |
| chr6_+_42241916 | 0.55 |
ENSMUST00000031895.13
|
Casp2
|
caspase 2 |
| chr9_-_100388857 | 0.54 |
ENSMUST00000112874.4
|
Nck1
|
non-catalytic region of tyrosine kinase adaptor protein 1 |
| chr1_+_134343107 | 0.53 |
ENSMUST00000027727.15
|
Adipor1
|
adiponectin receptor 1 |
| chr1_+_134343152 | 0.53 |
ENSMUST00000112237.2
|
Adipor1
|
adiponectin receptor 1 |
| chr9_+_56344700 | 0.52 |
ENSMUST00000239472.2
|
ENSMUSG00000118653.2
|
ubiquitin-conjugating enzyme E2S (Ube2s) retrogene |
| chr2_-_163486998 | 0.51 |
ENSMUST00000017851.4
|
Serinc3
|
serine incorporator 3 |
| chr9_-_105398346 | 0.50 |
ENSMUST00000176770.8
ENSMUST00000085133.13 |
Atp2c1
|
ATPase, Ca++-sequestering |
| chr13_-_59930059 | 0.50 |
ENSMUST00000225581.2
|
Gm49354
|
predicted gene, 49354 |
| chr15_+_54274151 | 0.50 |
ENSMUST00000036737.4
|
Colec10
|
collectin sub-family member 10 |
| chr19_+_45433899 | 0.49 |
ENSMUST00000224478.2
|
Btrc
|
beta-transducin repeat containing protein |
| chr10_-_116385007 | 0.47 |
ENSMUST00000164088.8
|
Cnot2
|
CCR4-NOT transcription complex, subunit 2 |
| chr11_-_62283431 | 0.47 |
ENSMUST00000151498.9
|
Ncor1
|
nuclear receptor co-repressor 1 |
| chr12_+_71936500 | 0.46 |
ENSMUST00000221317.2
|
Daam1
|
dishevelled associated activator of morphogenesis 1 |
| chr7_+_79836581 | 0.45 |
ENSMUST00000032754.9
|
Sema4b
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4B |
| chr5_-_115236354 | 0.45 |
ENSMUST00000100848.3
|
Gm10401
|
predicted gene 10401 |
| chr3_+_129326285 | 0.44 |
ENSMUST00000197235.5
|
Elovl6
|
ELOVL family member 6, elongation of long chain fatty acids (yeast) |
| chr8_-_54091980 | 0.43 |
ENSMUST00000047768.11
|
Neil3
|
nei like 3 (E. coli) |
| chr5_+_67473069 | 0.39 |
ENSMUST00000202770.2
|
Slc30a9
|
solute carrier family 30 (zinc transporter), member 9 |
| chr4_+_109200225 | 0.39 |
ENSMUST00000030281.12
|
Eps15
|
epidermal growth factor receptor pathway substrate 15 |
| chr6_+_58573656 | 0.39 |
ENSMUST00000031822.13
|
Abcg2
|
ATP binding cassette subfamily G member 2 (Junior blood group) |
| chr12_-_101924407 | 0.38 |
ENSMUST00000159883.2
ENSMUST00000160251.8 ENSMUST00000161011.8 ENSMUST00000021606.12 |
Atxn3
|
ataxin 3 |
| chr7_+_35148188 | 0.37 |
ENSMUST00000118383.8
|
Slc7a9
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 9 |
| chr7_+_35148461 | 0.36 |
ENSMUST00000118969.8
|
Slc7a9
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 9 |
| chr12_-_4957212 | 0.35 |
ENSMUST00000142867.8
|
Ubxn2a
|
UBX domain protein 2A |
| chr7_+_35148579 | 0.33 |
ENSMUST00000032703.10
|
Slc7a9
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 9 |
| chrX_+_138464065 | 0.33 |
ENSMUST00000113027.8
|
Rnf128
|
ring finger protein 128 |
| chr10_+_87926932 | 0.33 |
ENSMUST00000048621.8
|
Pmch
|
pro-melanin-concentrating hormone |
| chr5_-_138169253 | 0.32 |
ENSMUST00000139983.8
|
Mcm7
|
minichromosome maintenance complex component 7 |
| chr6_+_128376729 | 0.31 |
ENSMUST00000001561.12
|
Nrip2
|
nuclear receptor interacting protein 2 |
| chr10_-_5144699 | 0.30 |
ENSMUST00000215467.2
|
Syne1
|
spectrin repeat containing, nuclear envelope 1 |
| chrX_-_9335525 | 0.30 |
ENSMUST00000015484.10
|
Cybb
|
cytochrome b-245, beta polypeptide |
| chr2_+_60040231 | 0.30 |
ENSMUST00000102748.11
|
Marchf7
|
membrane associated ring-CH-type finger 7 |
| chr2_-_51039112 | 0.29 |
ENSMUST00000154545.2
ENSMUST00000017288.9 |
Rnd3
|
Rho family GTPase 3 |
| chr1_-_136273811 | 0.28 |
ENSMUST00000048309.12
|
Camsap2
|
calmodulin regulated spectrin-associated protein family, member 2 |
| chr1_-_82746169 | 0.27 |
ENSMUST00000027331.3
|
Tm4sf20
|
transmembrane 4 L six family member 20 |
| chr18_+_51250748 | 0.27 |
ENSMUST00000116639.4
|
Prr16
|
proline rich 16 |
| chr18_+_13107535 | 0.27 |
ENSMUST00000234035.2
ENSMUST00000235053.2 |
Impact
|
impact, RWD domain protein |
| chr9_+_108437485 | 0.27 |
ENSMUST00000081111.14
ENSMUST00000193421.2 |
Impdh2
|
inosine monophosphate dehydrogenase 2 |
| chr15_-_12549963 | 0.26 |
ENSMUST00000189324.2
|
Pdzd2
|
PDZ domain containing 2 |
| chr4_+_132366298 | 0.25 |
ENSMUST00000135299.8
ENSMUST00000020197.14 ENSMUST00000180250.8 ENSMUST00000081726.13 ENSMUST00000079157.11 |
Eya3
|
EYA transcriptional coactivator and phosphatase 3 |
| chr12_-_91745903 | 0.25 |
ENSMUST00000170077.2
|
Ston2
|
stonin 2 |
| chr8_+_46111703 | 0.25 |
ENSMUST00000134675.8
ENSMUST00000139869.8 ENSMUST00000126067.8 |
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr2_-_25498459 | 0.24 |
ENSMUST00000058137.9
|
Rabl6
|
RAB, member RAS oncogene family-like 6 |
| chr2_-_37537224 | 0.24 |
ENSMUST00000028279.10
|
Strbp
|
spermatid perinuclear RNA binding protein |
| chr6_+_128376844 | 0.24 |
ENSMUST00000120405.4
|
Nrip2
|
nuclear receptor interacting protein 2 |
| chr1_+_43137852 | 0.24 |
ENSMUST00000010434.8
|
AI597479
|
expressed sequence AI597479 |
| chr11_-_45845992 | 0.24 |
ENSMUST00000109254.2
|
Thg1l
|
tRNA-histidine guanylyltransferase 1-like (S. cerevisiae) |
| chr19_-_29721012 | 0.23 |
ENSMUST00000175764.9
|
9930021J03Rik
|
RIKEN cDNA 9930021J03 gene |
| chr2_-_37593287 | 0.23 |
ENSMUST00000072186.12
|
Strbp
|
spermatid perinuclear RNA binding protein |
| chr5_-_130284366 | 0.23 |
ENSMUST00000026387.11
|
Sbds
|
SBDS ribosome maturation factor |
| chrX_+_162694397 | 0.23 |
ENSMUST00000140845.2
|
Ap1s2
|
adaptor-related protein complex 1, sigma 2 subunit |
| chr8_+_46111778 | 0.22 |
ENSMUST00000143820.8
|
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr13_+_94954202 | 0.21 |
ENSMUST00000220825.2
|
Tbca
|
tubulin cofactor A |
| chr18_-_88945571 | 0.21 |
ENSMUST00000147313.2
|
Socs6
|
suppressor of cytokine signaling 6 |
| chr2_-_84652890 | 0.21 |
ENSMUST00000028471.6
|
Smtnl1
|
smoothelin-like 1 |
| chr6_+_136530970 | 0.21 |
ENSMUST00000189535.2
|
Atf7ip
|
activating transcription factor 7 interacting protein |
| chr1_-_44140820 | 0.20 |
ENSMUST00000152239.8
|
Tex30
|
testis expressed 30 |
| chr2_-_87868043 | 0.20 |
ENSMUST00000129056.3
|
Olfr73
|
olfactory receptor 73 |
| chr17_+_19724994 | 0.20 |
ENSMUST00000166081.3
ENSMUST00000231465.2 |
Vmn2r100
|
vomeronasal 2, receptor 100 |
| chr1_+_87332638 | 0.19 |
ENSMUST00000173152.2
ENSMUST00000173663.2 |
Gigyf2
|
GRB10 interacting GYF protein 2 |
| chr4_-_131565542 | 0.19 |
ENSMUST00000030741.9
ENSMUST00000105987.9 |
Ptpru
|
protein tyrosine phosphatase, receptor type, U |
| chr2_-_77349909 | 0.19 |
ENSMUST00000111830.9
|
Zfp385b
|
zinc finger protein 385B |
| chr17_-_34337446 | 0.19 |
ENSMUST00000095347.13
|
Brd2
|
bromodomain containing 2 |
| chr4_-_43584386 | 0.18 |
ENSMUST00000107884.3
|
Msmp
|
microseminoprotein, prostate associated |
| chr14_-_9015639 | 0.18 |
ENSMUST00000112656.4
|
Synpr
|
synaptoporin |
| chr12_+_25024913 | 0.18 |
ENSMUST00000066652.7
ENSMUST00000220459.2 ENSMUST00000222941.2 |
Kidins220
|
kinase D-interacting substrate 220 |
| chr2_+_87853118 | 0.17 |
ENSMUST00000214438.2
|
Olfr1161
|
olfactory receptor 1161 |
| chr12_-_85317359 | 0.17 |
ENSMUST00000166821.8
ENSMUST00000019378.8 ENSMUST00000220854.2 |
Mlh3
|
mutL homolog 3 |
| chr17_+_18269686 | 0.17 |
ENSMUST00000176802.3
|
Vmn2r124
|
vomeronasal 2, receptor 124 |
| chr3_+_68491487 | 0.17 |
ENSMUST00000182997.3
|
Schip1
|
schwannomin interacting protein 1 |
| chr7_+_16807965 | 0.17 |
ENSMUST00000071399.13
ENSMUST00000118367.3 |
Psg16
|
pregnancy specific glycoprotein 16 |
| chr16_-_88360037 | 0.16 |
ENSMUST00000049697.5
|
Cldn8
|
claudin 8 |
| chr13_+_28441511 | 0.16 |
ENSMUST00000223428.2
|
Rps18-ps5
|
ribosomal protein S18, pseudogene 5 |
| chr19_-_45619559 | 0.16 |
ENSMUST00000160718.9
|
Fbxw4
|
F-box and WD-40 domain protein 4 |
| chr2_+_60040291 | 0.16 |
ENSMUST00000102747.8
|
Marchf7
|
membrane associated ring-CH-type finger 7 |
| chr2_-_181223787 | 0.15 |
ENSMUST00000057816.15
|
Uckl1
|
uridine-cytidine kinase 1-like 1 |
| chr4_-_129015682 | 0.15 |
ENSMUST00000139450.8
ENSMUST00000125931.9 ENSMUST00000116444.10 |
Hpca
|
hippocalcin |
| chr16_+_43184507 | 0.15 |
ENSMUST00000148775.8
|
Zbtb20
|
zinc finger and BTB domain containing 20 |
| chr15_-_55411560 | 0.15 |
ENSMUST00000165356.2
|
Mrpl13
|
mitochondrial ribosomal protein L13 |
| chr8_-_34574970 | 0.15 |
ENSMUST00000118811.8
|
Dctn6
|
dynactin 6 |
| chr14_-_50380137 | 0.15 |
ENSMUST00000213685.2
|
Olfr728
|
olfactory receptor 728 |
| chrX_-_69408627 | 0.15 |
ENSMUST00000101509.9
|
Ids
|
iduronate 2-sulfatase |
| chr13_+_23715220 | 0.14 |
ENSMUST00000102972.6
|
H4c8
|
H4 clustered histone 8 |
| chr16_+_5703134 | 0.14 |
ENSMUST00000230658.2
|
Rbfox1
|
RNA binding protein, fox-1 homolog (C. elegans) 1 |
| chr15_-_50746202 | 0.14 |
ENSMUST00000184885.8
|
Trps1
|
transcriptional repressor GATA binding 1 |
| chr5_+_102629365 | 0.14 |
ENSMUST00000112854.8
|
Arhgap24
|
Rho GTPase activating protein 24 |
| chr2_-_45000389 | 0.14 |
ENSMUST00000201804.4
ENSMUST00000028229.13 ENSMUST00000202187.4 ENSMUST00000153561.6 ENSMUST00000201490.2 |
Zeb2
|
zinc finger E-box binding homeobox 2 |
| chr6_+_103487541 | 0.14 |
ENSMUST00000203912.3
|
Chl1
|
cell adhesion molecule L1-like |
| chr11_-_99511257 | 0.14 |
ENSMUST00000073853.3
|
Gm11562
|
predicted gene 11562 |
| chr16_-_78887971 | 0.13 |
ENSMUST00000023566.11
ENSMUST00000060402.6 |
Tmprss15
|
transmembrane protease, serine 15 |
| chr18_+_37622518 | 0.13 |
ENSMUST00000055949.4
|
Pcdhb18
|
protocadherin beta 18 |
| chr9_-_112016834 | 0.13 |
ENSMUST00000111872.9
ENSMUST00000164754.9 |
Arpp21
|
cyclic AMP-regulated phosphoprotein, 21 |
| chr18_+_37100672 | 0.13 |
ENSMUST00000193777.6
ENSMUST00000193389.2 |
Pcdha6
|
protocadherin alpha 6 |
| chr11_+_99770013 | 0.12 |
ENSMUST00000078442.4
|
Gm11567
|
predicted gene 11567 |
| chr4_+_108704982 | 0.11 |
ENSMUST00000102738.4
|
Kti12
|
KTI12 homolog, chromatin associated |
| chr1_-_132635042 | 0.11 |
ENSMUST00000043189.14
|
Nfasc
|
neurofascin |
| chr3_+_31150982 | 0.11 |
ENSMUST00000118204.2
|
Skil
|
SKI-like |
| chr10_+_69761784 | 0.11 |
ENSMUST00000181974.8
ENSMUST00000182795.8 ENSMUST00000182437.8 |
Ank3
|
ankyrin 3, epithelial |
| chr12_+_33364288 | 0.11 |
ENSMUST00000144586.2
|
Atxn7l1
|
ataxin 7-like 1 |
| chr4_-_3872105 | 0.11 |
ENSMUST00000105158.2
|
Mos
|
Moloney sarcoma oncogene |
| chr16_+_91188609 | 0.11 |
ENSMUST00000160764.2
|
Gm21970
|
predicted gene 21970 |
| chr6_-_53955647 | 0.11 |
ENSMUST00000204674.3
ENSMUST00000166545.2 ENSMUST00000203101.3 |
Cpvl
|
carboxypeptidase, vitellogenic-like |
| chr8_+_79236051 | 0.11 |
ENSMUST00000209992.2
|
Slc10a7
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 7 |
| chr4_+_109092459 | 0.11 |
ENSMUST00000106631.9
|
Calr4
|
calreticulin 4 |
| chr4_+_109092610 | 0.10 |
ENSMUST00000106628.8
|
Calr4
|
calreticulin 4 |
| chr2_-_7400690 | 0.10 |
ENSMUST00000182404.8
|
Celf2
|
CUGBP, Elav-like family member 2 |
| chr1_-_132635078 | 0.10 |
ENSMUST00000187861.7
|
Nfasc
|
neurofascin |
| chr2_+_109522781 | 0.10 |
ENSMUST00000111050.10
|
Bdnf
|
brain derived neurotrophic factor |
| chr17_+_30845909 | 0.10 |
ENSMUST00000236140.2
ENSMUST00000236118.2 ENSMUST00000235390.2 |
Dnah8
|
dynein, axonemal, heavy chain 8 |
| chr1_-_43235914 | 0.10 |
ENSMUST00000187357.2
|
Fhl2
|
four and a half LIM domains 2 |
| chr8_-_3904309 | 0.09 |
ENSMUST00000033888.5
|
Cd209e
|
CD209e antigen |
| chr6_+_88199250 | 0.09 |
ENSMUST00000061866.6
|
Dnajb8
|
DnaJ heat shock protein family (Hsp40) member B8 |
| chr14_-_54651442 | 0.09 |
ENSMUST00000227334.2
|
Slc7a7
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 7 |
| chr1_-_82748964 | 0.09 |
ENSMUST00000223838.2
|
Gm47791
|
predicted gene, 47791 |
| chr14_+_53698556 | 0.09 |
ENSMUST00000181728.3
|
Trav7-4
|
T cell receptor alpha variable 7-4 |
| chr15_-_12549350 | 0.09 |
ENSMUST00000190929.2
|
Pdzd2
|
PDZ domain containing 2 |
| chr18_+_57601541 | 0.09 |
ENSMUST00000091892.4
ENSMUST00000209782.2 |
Ctxn3
|
cortexin 3 |
| chr3_-_64252048 | 0.09 |
ENSMUST00000234658.2
|
Vmn2r128
|
vomeronasal 2 receptor 128 |
| chr4_-_45532470 | 0.08 |
ENSMUST00000147448.2
|
Shb
|
src homology 2 domain-containing transforming protein B |
| chr17_-_30845845 | 0.08 |
ENSMUST00000235547.2
ENSMUST00000188852.2 |
Glo1
1700097N02Rik
|
glyoxalase 1 RIKEN cDNA 1700097N02 gene |
| chr1_-_169938298 | 0.08 |
ENSMUST00000192312.6
|
Ddr2
|
discoidin domain receptor family, member 2 |
| chr3_+_53396120 | 0.08 |
ENSMUST00000029307.4
|
Stoml3
|
stomatin (Epb7.2)-like 3 |
| chr19_-_36896999 | 0.08 |
ENSMUST00000238948.2
ENSMUST00000057337.9 |
Fgfbp3
|
fibroblast growth factor binding protein 3 |
| chr1_-_169938060 | 0.08 |
ENSMUST00000027985.8
ENSMUST00000194690.6 |
Ddr2
|
discoidin domain receptor family, member 2 |
| chr3_-_108062172 | 0.08 |
ENSMUST00000062028.8
|
Gpr61
|
G protein-coupled receptor 61 |
| chr16_-_59138611 | 0.08 |
ENSMUST00000216261.2
|
Olfr204
|
olfactory receptor 204 |
| chr10_+_80591030 | 0.08 |
ENSMUST00000105336.9
|
Dot1l
|
DOT1-like, histone H3 methyltransferase (S. cerevisiae) |
| chr6_+_97968737 | 0.08 |
ENSMUST00000043628.13
ENSMUST00000203938.2 |
Mitf
|
melanogenesis associated transcription factor |
| chr2_-_87985537 | 0.07 |
ENSMUST00000216951.2
|
Olfr1167
|
olfactory receptor 1167 |
| chr17_+_12803019 | 0.07 |
ENSMUST00000046959.9
ENSMUST00000233066.2 |
Slc22a2
|
solute carrier family 22 (organic cation transporter), member 2 |
| chr9_-_39486831 | 0.07 |
ENSMUST00000216298.2
ENSMUST00000215194.2 |
Olfr959
|
olfactory receptor 959 |
| chr6_+_36364990 | 0.07 |
ENSMUST00000172278.8
|
Chrm2
|
cholinergic receptor, muscarinic 2, cardiac |
| chr9_-_118917275 | 0.07 |
ENSMUST00000214470.2
|
Plcd1
|
phospholipase C, delta 1 |
| chr18_+_37610858 | 0.07 |
ENSMUST00000051442.7
|
Pcdhb16
|
protocadherin beta 16 |
| chr12_+_8027640 | 0.07 |
ENSMUST00000171271.8
ENSMUST00000037811.13 |
Apob
|
apolipoprotein B |
| chr2_+_89708781 | 0.07 |
ENSMUST00000111519.3
|
Olfr1257
|
olfactory receptor 1257 |
| chrM_+_5319 | 0.07 |
ENSMUST00000082402.1
|
mt-Co1
|
mitochondrially encoded cytochrome c oxidase I |
| chr18_-_89787603 | 0.07 |
ENSMUST00000097495.5
|
Dok6
|
docking protein 6 |
| chr3_+_79791798 | 0.07 |
ENSMUST00000118853.8
ENSMUST00000145992.2 |
Gask1b
|
golgi associated kinase 1B |
| chr7_+_90075762 | 0.07 |
ENSMUST00000061391.9
|
Ccdc89
|
coiled-coil domain containing 89 |
| chr6_-_23655130 | 0.07 |
ENSMUST00000104979.2
|
Rnf148
|
ring finger protein 148 |
| chr5_-_124717146 | 0.07 |
ENSMUST00000031334.15
|
Eif2b1
|
eukaryotic translation initiation factor 2B, subunit 1 (alpha) |
| chr5_-_73787646 | 0.07 |
ENSMUST00000152215.3
|
Lrrc66
|
leucine rich repeat containing 66 |
| chr10_+_59942274 | 0.07 |
ENSMUST00000165024.3
|
Spock2
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan 2 |
| chr9_+_38259707 | 0.06 |
ENSMUST00000217063.2
|
Olfr898
|
olfactory receptor 898 |
| chr1_-_13442658 | 0.06 |
ENSMUST00000081713.11
|
Ncoa2
|
nuclear receptor coactivator 2 |
| chr7_-_133384449 | 0.06 |
ENSMUST00000063669.8
|
Dhx32
|
DEAH (Asp-Glu-Ala-His) box polypeptide 32 |
| chr9_-_56759229 | 0.06 |
ENSMUST00000055036.7
|
Odf3l1
|
outer dense fiber of sperm tails 3-like 1 |
| chr14_-_8146867 | 0.06 |
ENSMUST00000217035.2
ENSMUST00000206009.3 |
Olfr31
|
olfactory receptor 31 |
| chr16_-_26345493 | 0.06 |
ENSMUST00000165687.3
|
Tmem207
|
transmembrane protein 207 |
| chr3_-_141637245 | 0.06 |
ENSMUST00000106232.8
|
Bmpr1b
|
bone morphogenetic protein receptor, type 1B |
| chrY_+_13042891 | 0.06 |
ENSMUST00000194687.2
|
Gm20807
|
predicted gene, 20807 |
| chrY_-_10536042 | 0.06 |
ENSMUST00000187245.2
|
Gm20737
|
predicted gene, 20737 |
| chr2_-_104324035 | 0.05 |
ENSMUST00000111124.8
|
Hipk3
|
homeodomain interacting protein kinase 3 |
| chr1_+_6285082 | 0.05 |
ENSMUST00000160062.8
|
Rb1cc1
|
RB1-inducible coiled-coil 1 |
| chr1_+_156666485 | 0.05 |
ENSMUST00000111720.2
|
Angptl1
|
angiopoietin-like 1 |
| chr5_-_87847268 | 0.05 |
ENSMUST00000196869.5
ENSMUST00000199624.5 ENSMUST00000198057.5 ENSMUST00000082370.10 |
Csn2
|
casein beta |
| chr18_-_3280999 | 0.05 |
ENSMUST00000049942.13
|
Crem
|
cAMP responsive element modulator |
| chr6_-_130258891 | 0.05 |
ENSMUST00000112020.2
|
Klra10
|
killer cell lectin-like receptor subfamily A, member 10 |
| chr3_+_145693582 | 0.05 |
ENSMUST00000039517.11
|
Syde2
|
synapse defective 1, Rho GTPase, homolog 2 (C. elegans) |
| chr11_+_77656414 | 0.05 |
ENSMUST00000164315.2
|
Myo18a
|
myosin XVIIIA |
| chr1_+_13738967 | 0.05 |
ENSMUST00000088542.4
|
Xkr9
|
X-linked Kx blood group related 9 |
| chr10_-_85752932 | 0.05 |
ENSMUST00000218969.2
|
Prdm4
|
PR domain containing 4 |
| chr3_-_97329336 | 0.04 |
ENSMUST00000199297.4
|
Olfr1402
|
olfactory receptor 1402 |
| chrX_-_140979913 | 0.04 |
ENSMUST00000042530.4
|
Gucy2f
|
guanylate cyclase 2f |
| chr9_+_45230370 | 0.04 |
ENSMUST00000034597.8
|
Tmprss13
|
transmembrane protease, serine 13 |
| chr6_-_28134544 | 0.04 |
ENSMUST00000115323.8
|
Grm8
|
glutamate receptor, metabotropic 8 |
| chr19_+_23881821 | 0.04 |
ENSMUST00000237688.2
|
Apba1
|
amyloid beta (A4) precursor protein binding, family A, member 1 |
| chr17_-_37938000 | 0.04 |
ENSMUST00000223366.2
ENSMUST00000216128.2 |
Olfr115
Olfr116
|
olfactory receptor 115 olfactory receptor 116 |
| chr2_+_173579285 | 0.04 |
ENSMUST00000067530.6
|
Vapb
|
vesicle-associated membrane protein, associated protein B and C |
| chr8_-_37420293 | 0.03 |
ENSMUST00000179501.2
|
Dlc1
|
deleted in liver cancer 1 |
| chr13_-_67785843 | 0.03 |
ENSMUST00000225627.2
ENSMUST00000224814.2 |
Gm49345
|
predicted gene, 49345 |
| chr19_+_11872971 | 0.03 |
ENSMUST00000072784.4
|
Olfr1420
|
olfactory receptor 1420 |
| chr9_-_39188114 | 0.03 |
ENSMUST00000216698.2
|
Olfr945
|
olfactory receptor 945 |
| chr5_+_91175323 | 0.03 |
ENSMUST00000202724.4
ENSMUST00000041516.9 |
Epgn
|
epithelial mitogen |
| chr2_-_88963396 | 0.03 |
ENSMUST00000214027.2
ENSMUST00000215816.3 |
Olfr1222
|
olfactory receptor 1222 |
| chr2_+_24166920 | 0.03 |
ENSMUST00000168941.8
ENSMUST00000028360.8 ENSMUST00000123053.8 |
Il1f5
|
interleukin 1 family, member 5 (delta) |
| chr14_+_53691753 | 0.03 |
ENSMUST00000180711.3
ENSMUST00000184650.2 |
Trav6-4
|
T cell receptor alpha variable 6-4 |
| chr6_+_24733239 | 0.03 |
ENSMUST00000031690.6
|
Hyal6
|
hyaluronoglucosaminidase 6 |
| chrX_+_37689503 | 0.03 |
ENSMUST00000000365.3
|
Mcts1
|
malignant T cell amplified sequence 1 |
| chr13_+_22534534 | 0.03 |
ENSMUST00000226909.2
ENSMUST00000227167.2 ENSMUST00000226786.2 |
Vmn1r198
|
vomeronasal 1 receptor 198 |
| chr9_-_60048503 | 0.03 |
ENSMUST00000034829.6
|
Thsd4
|
thrombospondin, type I, domain containing 4 |
| chr6_+_123149847 | 0.03 |
ENSMUST00000088455.6
|
Clec4b2
|
C-type lectin domain family 4, member b2 |
| chr7_-_140597837 | 0.03 |
ENSMUST00000209328.2
|
Ifitm6
|
interferon induced transmembrane protein 6 |
| chr2_-_86061745 | 0.03 |
ENSMUST00000216056.2
|
Olfr1047
|
olfactory receptor 1047 |
| chr6_+_56933451 | 0.02 |
ENSMUST00000096612.4
|
Vmn1r4
|
vomeronasal 1 receptor 4 |
| chr7_+_101032021 | 0.02 |
ENSMUST00000141083.9
|
Arap1
|
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1 |
| chr2_+_112114906 | 0.02 |
ENSMUST00000053666.8
|
Slc12a6
|
solute carrier family 12, member 6 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.1 | GO:0015811 | L-cystine transport(GO:0015811) |
| 0.2 | 0.7 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.2 | 1.6 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.2 | 3.1 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.1 | 1.8 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.1 | 0.5 | GO:0002191 | cap-dependent translational initiation(GO:0002191) |
| 0.1 | 0.4 | GO:0019389 | glucuronoside metabolic process(GO:0019389) |
| 0.1 | 0.5 | GO:0030026 | cellular manganese ion homeostasis(GO:0030026) Golgi calcium ion homeostasis(GO:0032468) manganese ion homeostasis(GO:0055071) |
| 0.1 | 0.5 | GO:0072362 | regulation of glycolytic process by negative regulation of transcription from RNA polymerase II promoter(GO:0072362) |
| 0.1 | 0.3 | GO:0046005 | positive regulation of circadian sleep/wake cycle, REM sleep(GO:0046005) |
| 0.1 | 0.5 | GO:0009597 | detection of virus(GO:0009597) |
| 0.1 | 0.6 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.1 | 0.3 | GO:1901561 | cellular response to benomyl(GO:0072755) response to benomyl(GO:1901561) |
| 0.1 | 1.1 | GO:0033210 | leptin-mediated signaling pathway(GO:0033210) |
| 0.1 | 0.9 | GO:0070574 | cadmium ion transport(GO:0015691) cadmium ion transmembrane transport(GO:0070574) |
| 0.1 | 0.9 | GO:0030916 | otic vesicle formation(GO:0030916) |
| 0.1 | 0.6 | GO:0010792 | DNA double-strand break processing involved in repair via single-strand annealing(GO:0010792) |
| 0.1 | 0.2 | GO:0031283 | negative regulation of cGMP metabolic process(GO:0030824) negative regulation of cGMP biosynthetic process(GO:0030827) negative regulation of guanylate cyclase activity(GO:0031283) |
| 0.1 | 0.6 | GO:0035234 | ectopic germ cell programmed cell death(GO:0035234) |
| 0.0 | 0.3 | GO:0090292 | nuclear matrix organization(GO:0043578) nuclear matrix anchoring at nuclear membrane(GO:0090292) |
| 0.0 | 0.2 | GO:0007023 | post-chaperonin tubulin folding pathway(GO:0007023) |
| 0.0 | 0.5 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.0 | 0.3 | GO:0006177 | GMP biosynthetic process(GO:0006177) |
| 0.0 | 0.3 | GO:0097411 | hypoxia-inducible factor-1alpha signaling pathway(GO:0097411) |
| 0.0 | 0.1 | GO:1901994 | negative regulation of meiotic cell cycle phase transition(GO:1901994) |
| 0.0 | 0.4 | GO:0019065 | receptor-mediated endocytosis of virus by host cell(GO:0019065) endocytosis involved in viral entry into host cell(GO:0075509) |
| 0.0 | 0.1 | GO:0061193 | taste bud development(GO:0061193) |
| 0.0 | 0.2 | GO:0042256 | mature ribosome assembly(GO:0042256) |
| 0.0 | 0.4 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) |
| 0.0 | 0.3 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.0 | 0.4 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.0 | 0.1 | GO:0055011 | atrial cardiac muscle cell differentiation(GO:0055011) atrial cardiac muscle cell development(GO:0055014) |
| 0.0 | 0.2 | GO:0008655 | pyrimidine-containing compound salvage(GO:0008655) pyrimidine nucleoside salvage(GO:0043097) |
| 0.0 | 0.2 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.0 | 0.1 | GO:1904017 | response to Thyroglobulin triiodothyronine(GO:1904016) cellular response to Thyroglobulin triiodothyronine(GO:1904017) |
| 0.0 | 0.1 | GO:0034729 | histone H3-K79 methylation(GO:0034729) |
| 0.0 | 0.2 | GO:0007144 | female meiosis I(GO:0007144) |
| 0.0 | 0.1 | GO:0061723 | glycophagy(GO:0061723) |
| 0.0 | 0.1 | GO:1903487 | regulation of lactation(GO:1903487) |
| 0.0 | 1.1 | GO:1900047 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.0 | 0.2 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.0 | 0.6 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.0 | 0.0 | GO:1903027 | regulation of opsonization(GO:1903027) positive regulation of opsonization(GO:1903028) |
| 0.0 | 0.1 | GO:0097324 | melanocyte migration(GO:0097324) positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.0 | 0.6 | GO:0097320 | membrane tubulation(GO:0097320) |
| 0.0 | 0.1 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) phospholipase C-activating G-protein coupled acetylcholine receptor signaling pathway(GO:0007207) |
| 0.0 | 0.1 | GO:1990928 | response to amino acid starvation(GO:1990928) |
| 0.0 | 0.1 | GO:0032962 | positive regulation of inositol trisphosphate biosynthetic process(GO:0032962) |
| 0.0 | 0.1 | GO:0001550 | ovarian cumulus expansion(GO:0001550) fused antrum stage(GO:0048165) |
| 0.0 | 0.1 | GO:0000821 | regulation of arginine metabolic process(GO:0000821) |
| 0.0 | 0.2 | GO:0090091 | positive regulation of extracellular matrix disassembly(GO:0090091) |
| 0.0 | 0.3 | GO:0045793 | positive regulation of cell size(GO:0045793) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0005712 | chiasma(GO:0005712) |
| 0.0 | 3.1 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 0.5 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.0 | 0.2 | GO:0097454 | Schwann cell microvillus(GO:0097454) |
| 0.0 | 0.5 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 0.3 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.0 | 0.3 | GO:0042555 | MCM complex(GO:0042555) |
| 0.0 | 0.5 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.0 | 0.1 | GO:0034359 | mature chylomicron(GO:0034359) |
| 0.0 | 0.2 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.0 | 0.3 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.0 | 0.3 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.0 | 0.4 | GO:0030122 | AP-2 adaptor complex(GO:0030122) |
| 0.0 | 0.4 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.0 | 0.6 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 0.2 | GO:0044327 | dendritic spine head(GO:0044327) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.1 | GO:0015184 | L-cystine transmembrane transporter activity(GO:0015184) |
| 0.2 | 1.1 | GO:0097003 | adipokinetic hormone receptor activity(GO:0097003) |
| 0.2 | 1.6 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.1 | 1.8 | GO:0102337 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.1 | 0.5 | GO:0015410 | manganese-transporting ATPase activity(GO:0015410) |
| 0.1 | 0.6 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 0.1 | 0.5 | GO:0001226 | RNA polymerase II transcription corepressor binding(GO:0001226) |
| 0.1 | 0.2 | GO:0008192 | RNA guanylyltransferase activity(GO:0008192) |
| 0.1 | 0.5 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.1 | 0.4 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 0.1 | 0.5 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.0 | 0.4 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.3 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.0 | 0.4 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) |
| 0.0 | 0.2 | GO:0019237 | centromeric DNA binding(GO:0019237) |
| 0.0 | 0.6 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) |
| 0.0 | 0.5 | GO:0022889 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.0 | 0.6 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.1 | GO:0005169 | neurotrophin TRKB receptor binding(GO:0005169) |
| 0.0 | 0.3 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.0 | 0.2 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.0 | 0.2 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 0.0 | 0.1 | GO:1990763 | arrestin family protein binding(GO:1990763) |
| 0.0 | 0.1 | GO:0004920 | interleukin-10 receptor activity(GO:0004920) |
| 0.0 | 0.2 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.0 | 0.6 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.3 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.0 | 0.1 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.0 | 0.5 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.0 | 0.0 | GO:0033149 | FFAT motif binding(GO:0033149) |
| 0.0 | 0.5 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 0.3 | GO:0005521 | lamin binding(GO:0005521) |
| 0.0 | 0.4 | GO:0033130 | acetylcholine receptor binding(GO:0033130) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.0 | 0.6 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 0.7 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.0 | 0.5 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 1.8 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 0.5 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 1.1 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.0 | 0.6 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 0.3 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.0 | 0.7 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.0 | 0.5 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.0 | 0.3 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.0 | 0.2 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.0 | 0.5 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 0.5 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.6 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.0 | 0.2 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.0 | 0.1 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |