Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
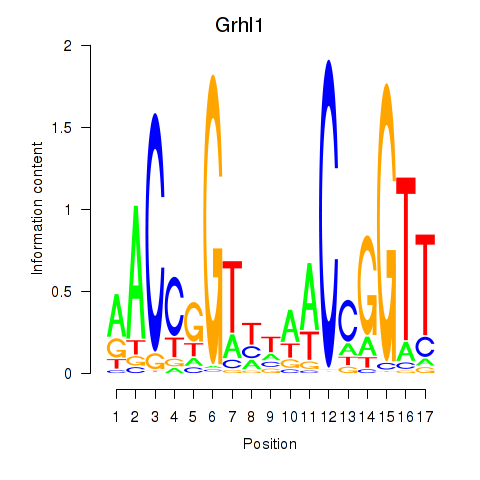
Results for Grhl1
Z-value: 3.16
Transcription factors associated with Grhl1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Grhl1
|
ENSMUSG00000020656.17 | Grhl1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Grhl1 | mm39_v1_chr12_+_24622274_24622307 | 0.10 | 5.8e-01 | Click! |
Activity profile of Grhl1 motif
Sorted Z-values of Grhl1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Grhl1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_-_93956143 | 20.32 |
ENSMUST00000176282.2
ENSMUST00000034173.14 |
Ces1e
|
carboxylesterase 1E |
| chr9_-_78254422 | 19.47 |
ENSMUST00000034902.12
|
Gsta2
|
glutathione S-transferase, alpha 2 (Yc2) |
| chr9_-_78254443 | 16.16 |
ENSMUST00000129247.2
|
Gsta2
|
glutathione S-transferase, alpha 2 (Yc2) |
| chrX_-_37545311 | 14.60 |
ENSMUST00000074913.12
ENSMUST00000016678.14 ENSMUST00000061755.9 |
Lamp2
|
lysosomal-associated membrane protein 2 |
| chr17_-_56312555 | 13.87 |
ENSMUST00000043785.8
|
Stap2
|
signal transducing adaptor family member 2 |
| chr11_+_72326391 | 12.71 |
ENSMUST00000100903.3
|
Ggt6
|
gamma-glutamyltransferase 6 |
| chr4_+_140970161 | 10.52 |
ENSMUST00000138096.8
ENSMUST00000006618.9 ENSMUST00000125392.8 |
Arhgef19
|
Rho guanine nucleotide exchange factor (GEF) 19 |
| chr17_+_56312672 | 10.22 |
ENSMUST00000133998.8
|
Mpnd
|
MPN domain containing |
| chr18_+_56565188 | 9.92 |
ENSMUST00000070166.6
|
Gramd3
|
GRAM domain containing 3 |
| chr9_+_78197205 | 9.77 |
ENSMUST00000119823.8
ENSMUST00000121273.2 |
Gsta5
|
glutathione S-transferase alpha 5 |
| chr7_+_100970435 | 8.50 |
ENSMUST00000210192.2
ENSMUST00000172630.8 |
Stard10
|
START domain containing 10 |
| chr19_+_18609291 | 7.90 |
ENSMUST00000042392.14
ENSMUST00000237347.2 |
Nmrk1
|
nicotinamide riboside kinase 1 |
| chr2_-_30095805 | 7.90 |
ENSMUST00000113663.9
ENSMUST00000044038.10 |
Kyat1
|
kynurenine aminotransferase 1 |
| chr14_-_73622638 | 7.77 |
ENSMUST00000228637.2
ENSMUST00000022704.9 |
Itm2b
|
integral membrane protein 2B |
| chr1_+_135746330 | 7.69 |
ENSMUST00000038760.10
|
Lad1
|
ladinin |
| chr2_-_30095784 | 7.67 |
ENSMUST00000113662.8
|
Kyat1
|
kynurenine aminotransferase 1 |
| chr15_-_75886166 | 7.67 |
ENSMUST00000060807.12
|
Fam83h
|
family with sequence similarity 83, member H |
| chr7_+_100971034 | 7.58 |
ENSMUST00000173270.8
|
Stard10
|
START domain containing 10 |
| chr13_+_8935537 | 7.42 |
ENSMUST00000169314.9
|
Idi1
|
isopentenyl-diphosphate delta isomerase |
| chr7_+_100970910 | 7.17 |
ENSMUST00000174291.8
ENSMUST00000167888.9 ENSMUST00000172662.2 |
Stard10
|
START domain containing 10 |
| chr7_+_65343156 | 5.79 |
ENSMUST00000032726.14
ENSMUST00000107495.5 ENSMUST00000143508.3 ENSMUST00000129166.3 ENSMUST00000206517.2 ENSMUST00000206837.2 ENSMUST00000206628.2 ENSMUST00000206361.2 |
Tm2d3
|
TM2 domain containing 3 |
| chr19_+_18609343 | 4.73 |
ENSMUST00000159572.9
|
Nmrk1
|
nicotinamide riboside kinase 1 |
| chr14_-_57370706 | 4.20 |
ENSMUST00000160703.2
|
Gjb6
|
gap junction protein, beta 6 |
| chr4_+_108691798 | 3.91 |
ENSMUST00000030296.9
|
Txndc12
|
thioredoxin domain containing 12 (endoplasmic reticulum) |
| chr16_+_43993599 | 3.62 |
ENSMUST00000119746.8
ENSMUST00000088356.10 ENSMUST00000169582.3 |
Usf3
|
upstream transcription factor family member 3 |
| chr3_+_79793237 | 2.98 |
ENSMUST00000029567.9
|
Gask1b
|
golgi associated kinase 1B |
| chr17_+_29487881 | 2.92 |
ENSMUST00000234845.2
ENSMUST00000235038.2 ENSMUST00000235050.2 ENSMUST00000120346.9 ENSMUST00000234377.2 ENSMUST00000235074.2 ENSMUST00000235040.2 ENSMUST00000234256.2 ENSMUST00000234459.2 |
BC004004
|
cDNA sequence BC004004 |
| chr4_-_155170738 | 2.68 |
ENSMUST00000030914.4
|
Rer1
|
retention in endoplasmic reticulum sorting receptor 1 |
| chr17_-_34847354 | 2.67 |
ENSMUST00000167097.9
|
Ppt2
|
palmitoyl-protein thioesterase 2 |
| chr17_+_29487762 | 2.51 |
ENSMUST00000064709.13
ENSMUST00000234711.2 |
BC004004
|
cDNA sequence BC004004 |
| chr1_-_52766615 | 2.44 |
ENSMUST00000156876.8
ENSMUST00000087701.4 |
Mfsd6
|
major facilitator superfamily domain containing 6 |
| chr7_-_101714251 | 2.23 |
ENSMUST00000130074.2
ENSMUST00000131104.3 ENSMUST00000096639.12 |
Rnf121
|
ring finger protein 121 |
| chr1_-_131207279 | 2.19 |
ENSMUST00000062108.10
|
Ikbke
|
inhibitor of kappaB kinase epsilon |
| chr3_-_10400710 | 2.02 |
ENSMUST00000078748.4
|
Slc10a5
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 5 |
| chr8_+_46081213 | 1.84 |
ENSMUST00000130850.8
|
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr4_-_19570073 | 1.82 |
ENSMUST00000029885.5
|
Cpne3
|
copine III |
| chr2_+_172392678 | 1.79 |
ENSMUST00000099058.10
|
Tfap2c
|
transcription factor AP-2, gamma |
| chr18_-_25886908 | 1.78 |
ENSMUST00000115816.3
ENSMUST00000223704.2 |
Celf4
|
CUGBP, Elav-like family member 4 |
| chr12_+_24622274 | 1.78 |
ENSMUST00000085553.13
|
Grhl1
|
grainyhead like transcription factor 1 |
| chr3_-_87171005 | 1.71 |
ENSMUST00000146512.2
|
Fcrls
|
Fc receptor-like S, scavenger receptor |
| chr2_+_84891281 | 1.71 |
ENSMUST00000238769.2
|
Tnks1bp1
|
tankyrase 1 binding protein 1 |
| chr5_-_86666408 | 1.66 |
ENSMUST00000140095.2
ENSMUST00000134179.8 |
Tmprss11g
|
transmembrane protease, serine 11g |
| chr4_+_155171034 | 1.65 |
ENSMUST00000030915.11
ENSMUST00000155775.8 ENSMUST00000127457.2 |
Morn1
|
MORN repeat containing 1 |
| chr5_-_117254146 | 1.62 |
ENSMUST00000166397.3
|
Suds3
|
suppressor of defective silencing 3 homolog (S. cerevisiae) |
| chr2_-_127383305 | 1.62 |
ENSMUST00000103214.3
|
Prom2
|
prominin 2 |
| chr17_-_34847013 | 1.51 |
ENSMUST00000166040.9
|
Ppt2
|
palmitoyl-protein thioesterase 2 |
| chr9_-_44179479 | 1.51 |
ENSMUST00000213803.2
|
Nlrx1
|
NLR family member X1 |
| chr2_-_179931672 | 1.39 |
ENSMUST00000038529.2
|
Rbbp8nl
|
RBBP8 N-terminal like |
| chr2_-_144174066 | 1.38 |
ENSMUST00000037423.4
|
Ovol2
|
ovo like zinc finger 2 |
| chr4_+_133793208 | 1.35 |
ENSMUST00000137053.2
|
Crybg2
|
crystallin beta-gamma domain containing 2 |
| chr7_+_12661337 | 1.25 |
ENSMUST00000045870.5
|
Rnf225
|
ring finger protein 225 |
| chr18_-_25887173 | 1.22 |
ENSMUST00000225477.2
|
Celf4
|
CUGBP, Elav-like family member 4 |
| chr17_+_7592045 | 1.19 |
ENSMUST00000095726.11
ENSMUST00000128533.8 ENSMUST00000129709.8 ENSMUST00000147803.8 ENSMUST00000140192.8 ENSMUST00000138222.8 ENSMUST00000144861.2 |
Tcp10a
|
t-complex protein 10a |
| chr17_+_35844091 | 1.19 |
ENSMUST00000025273.9
|
Psors1c2
|
psoriasis susceptibility 1 candidate 2 (human) |
| chr2_-_127383332 | 1.12 |
ENSMUST00000028855.14
|
Prom2
|
prominin 2 |
| chr5_-_86824205 | 1.01 |
ENSMUST00000038448.7
|
Tmprss11b
|
transmembrane protease, serine 11B |
| chr19_-_18609118 | 0.92 |
ENSMUST00000025631.7
ENSMUST00000236615.2 |
Ostf1
|
osteoclast stimulating factor 1 |
| chr12_+_76464372 | 0.86 |
ENSMUST00000220187.2
|
Ppp1r36
|
protein phosphatase 1, regulatory subunit 36 |
| chrX_+_99773784 | 0.82 |
ENSMUST00000113744.2
|
Gdpd2
|
glycerophosphodiester phosphodiesterase domain containing 2 |
| chr17_+_13574834 | 0.81 |
ENSMUST00000233944.2
ENSMUST00000097403.4 |
Tcp10c
|
t-complex protein 10c |
| chr19_-_5610628 | 0.78 |
ENSMUST00000025861.3
|
Ovol1
|
ovo like zinc finger 1 |
| chr3_-_87170903 | 0.75 |
ENSMUST00000090986.11
|
Fcrls
|
Fc receptor-like S, scavenger receptor |
| chr2_-_30857858 | 0.75 |
ENSMUST00000028200.9
|
Tor1a
|
torsin family 1, member A (torsin A) |
| chr4_+_135455427 | 0.73 |
ENSMUST00000102546.4
|
Il22ra1
|
interleukin 22 receptor, alpha 1 |
| chr2_+_24166920 | 0.73 |
ENSMUST00000168941.8
ENSMUST00000028360.8 ENSMUST00000123053.8 |
Il1f5
|
interleukin 1 family, member 5 (delta) |
| chr18_-_25886750 | 0.72 |
ENSMUST00000224553.2
ENSMUST00000025117.14 |
Celf4
|
CUGBP, Elav-like family member 4 |
| chr3_+_92292990 | 0.71 |
ENSMUST00000059845.2
ENSMUST00000191886.2 |
Sprr2h
|
small proline-rich protein 2H |
| chr9_-_44179758 | 0.71 |
ENSMUST00000034621.16
ENSMUST00000168499.9 |
Nlrx1
|
NLR family member X1 |
| chr6_+_139576475 | 0.67 |
ENSMUST00000187618.7
ENSMUST00000186585.3 |
Pik3c2g
|
phosphatidylinositol-4-phosphate 3-kinase catalytic subunit type 2 gamma |
| chr7_+_45204350 | 0.62 |
ENSMUST00000210300.2
|
Hsd17b14
|
hydroxysteroid (17-beta) dehydrogenase 14 |
| chr16_-_26808724 | 0.57 |
ENSMUST00000089832.6
|
Gmnc
|
geminin coiled-coil domain containing |
| chr3_+_92223927 | 0.55 |
ENSMUST00000061038.4
|
Sprr2b
|
small proline-rich protein 2B |
| chr2_+_28403255 | 0.54 |
ENSMUST00000028170.15
|
Ralgds
|
ral guanine nucleotide dissociation stimulator |
| chr19_+_4008645 | 0.54 |
ENSMUST00000179433.8
|
Aldh3b3
|
aldehyde dehydrogenase 3 family, member B3 |
| chr7_+_30487322 | 0.53 |
ENSMUST00000189673.7
ENSMUST00000190990.7 ENSMUST00000189962.7 ENSMUST00000187493.7 ENSMUST00000098559.3 |
Krtdap
|
keratinocyte differentiation associated protein |
| chr12_-_54703281 | 0.49 |
ENSMUST00000056228.8
|
Sptssa
|
serine palmitoyltransferase, small subunit A |
| chr8_+_124204598 | 0.48 |
ENSMUST00000001520.13
|
Afg3l1
|
AFG3-like AAA ATPase 1 |
| chrX_+_99773523 | 0.44 |
ENSMUST00000019503.14
|
Gdpd2
|
glycerophosphodiester phosphodiesterase domain containing 2 |
| chr2_+_149672814 | 0.43 |
ENSMUST00000137280.2
ENSMUST00000149705.2 |
Syndig1
|
synapse differentiation inducing 1 |
| chr6_-_113354468 | 0.42 |
ENSMUST00000099118.8
|
Tada3
|
transcriptional adaptor 3 |
| chr18_-_46413886 | 0.36 |
ENSMUST00000236999.2
|
Pggt1b
|
protein geranylgeranyltransferase type I, beta subunit |
| chr11_-_105346120 | 0.34 |
ENSMUST00000138977.8
|
Marchf10
|
membrane associated ring-CH-type finger 10 |
| chr7_-_29217967 | 0.30 |
ENSMUST00000181975.8
|
Sipa1l3
|
signal-induced proliferation-associated 1 like 3 |
| chr12_+_76464301 | 0.28 |
ENSMUST00000063977.9
|
Ppp1r36
|
protein phosphatase 1, regulatory subunit 36 |
| chr9_+_15150341 | 0.26 |
ENSMUST00000034413.8
|
Vstm5
|
V-set and transmembrane domain containing 5 |
| chr6_-_131293361 | 0.25 |
ENSMUST00000121078.2
|
Styk1
|
serine/threonine/tyrosine kinase 1 |
| chr15_-_63680596 | 0.22 |
ENSMUST00000110125.9
ENSMUST00000173503.3 |
Gsdmc
|
gasdermin C |
| chr18_-_64794338 | 0.19 |
ENSMUST00000025482.10
|
Atp8b1
|
ATPase, class I, type 8B, member 1 |
| chr17_+_37148015 | 0.18 |
ENSMUST00000179968.8
ENSMUST00000130367.8 ENSMUST00000053434.15 ENSMUST00000130801.8 ENSMUST00000144182.8 ENSMUST00000123715.8 |
Trim26
|
tripartite motif-containing 26 |
| chr4_+_112089442 | 0.18 |
ENSMUST00000038455.12
ENSMUST00000170945.2 |
Skint3
|
selection and upkeep of intraepithelial T cells 3 |
| chr9_-_7872970 | 0.17 |
ENSMUST00000115672.2
|
Birc3
|
baculoviral IAP repeat-containing 3 |
| chr3_+_92195873 | 0.17 |
ENSMUST00000090872.7
|
Sprr2a3
|
small proline-rich protein 2A3 |
| chr17_+_43700327 | 0.09 |
ENSMUST00000113599.2
ENSMUST00000224278.2 ENSMUST00000225466.2 |
Adgrf5
|
adhesion G protein-coupled receptor F5 |
| chr2_+_149672708 | 0.06 |
ENSMUST00000109935.8
|
Syndig1
|
synapse differentiation inducing 1 |
| chrX_-_111608339 | 0.04 |
ENSMUST00000039887.4
|
Pof1b
|
premature ovarian failure 1B |
| chr2_+_149672760 | 0.00 |
ENSMUST00000109934.2
ENSMUST00000140870.8 |
Syndig1
|
synapse differentiation inducing 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.2 | 15.6 | GO:0097052 | L-kynurenine metabolic process(GO:0097052) |
| 4.9 | 14.6 | GO:0071211 | protein targeting to vacuole involved in autophagy(GO:0071211) |
| 2.3 | 23.2 | GO:0035360 | positive regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035360) |
| 1.3 | 43.0 | GO:0035634 | response to stilbenoid(GO:0035634) |
| 1.0 | 7.8 | GO:0042985 | negative regulation of amyloid precursor protein biosynthetic process(GO:0042985) |
| 0.9 | 12.7 | GO:0006751 | glutathione catabolic process(GO:0006751) |
| 0.7 | 2.7 | GO:2001287 | negative regulation of caveolin-mediated endocytosis(GO:2001287) |
| 0.5 | 5.4 | GO:1901727 | positive regulation of histone deacetylase activity(GO:1901727) |
| 0.4 | 1.8 | GO:0060598 | dichotomous subdivision of terminal units involved in mammary gland duct morphogenesis(GO:0060598) |
| 0.4 | 12.6 | GO:0019363 | pyridine nucleotide biosynthetic process(GO:0019363) |
| 0.3 | 1.8 | GO:0038128 | ERBB2 signaling pathway(GO:0038128) |
| 0.3 | 3.7 | GO:1902866 | regulation of retina development in camera-type eye(GO:1902866) |
| 0.3 | 0.8 | GO:1901994 | negative regulation of meiotic cell cycle phase transition(GO:1901994) |
| 0.2 | 2.2 | GO:0039536 | negative regulation of RIG-I signaling pathway(GO:0039536) |
| 0.2 | 2.7 | GO:0071340 | skeletal muscle acetylcholine-gated channel clustering(GO:0071340) |
| 0.2 | 1.4 | GO:0060214 | endocardium formation(GO:0060214) |
| 0.1 | 1.8 | GO:0002934 | desmosome organization(GO:0002934) |
| 0.1 | 8.4 | GO:0045104 | intermediate filament cytoskeleton organization(GO:0045104) |
| 0.1 | 3.9 | GO:1902236 | negative regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signaling pathway(GO:1902236) |
| 0.1 | 3.6 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.1 | 0.7 | GO:1902714 | negative regulation of interferon-gamma secretion(GO:1902714) |
| 0.1 | 1.3 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.1 | 2.2 | GO:0010884 | positive regulation of lipid storage(GO:0010884) |
| 0.1 | 0.4 | GO:0051771 | negative regulation of nitric-oxide synthase biosynthetic process(GO:0051771) |
| 0.0 | 1.7 | GO:0031954 | positive regulation of protein autophosphorylation(GO:0031954) |
| 0.0 | 10.5 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.0 | 0.5 | GO:0034982 | mitochondrial protein processing(GO:0034982) |
| 0.0 | 1.3 | GO:0031424 | keratinization(GO:0031424) |
| 0.0 | 0.2 | GO:0021650 | vestibulocochlear nerve formation(GO:0021650) |
| 0.0 | 1.8 | GO:0003301 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.0 | 4.2 | GO:0042471 | ear morphogenesis(GO:0042471) |
| 0.0 | 0.7 | GO:0039703 | viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) |
| 0.0 | 1.5 | GO:0030968 | endoplasmic reticulum unfolded protein response(GO:0030968) |
| 0.0 | 0.5 | GO:0097091 | synaptic vesicle clustering(GO:0097091) |
| 0.0 | 1.7 | GO:0016575 | histone deacetylation(GO:0016575) |
| 0.0 | 0.6 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.0 | 1.1 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.7 | 14.6 | GO:0097637 | platelet dense granule membrane(GO:0031088) intrinsic component of autophagosome membrane(GO:0097636) integral component of autophagosome membrane(GO:0097637) |
| 1.3 | 23.2 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.5 | 2.7 | GO:0044393 | microspike(GO:0044393) |
| 0.2 | 7.7 | GO:0045095 | keratin filament(GO:0045095) |
| 0.2 | 4.2 | GO:0005922 | connexon complex(GO:0005922) |
| 0.2 | 7.8 | GO:0030660 | Golgi-associated vesicle membrane(GO:0030660) |
| 0.1 | 0.5 | GO:0005745 | m-AAA complex(GO:0005745) |
| 0.1 | 9.9 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 0.7 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.1 | 1.6 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.1 | 3.9 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.1 | 1.7 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.1 | 0.5 | GO:0017059 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.1 | 2.7 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.1 | 7.7 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 11.8 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 0.3 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.0 | 7.4 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.0 | 0.5 | GO:0042599 | lamellar body(GO:0042599) |
| 0.0 | 0.7 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 2.2 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 2.2 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.2 | 15.6 | GO:0047312 | L-phenylalanine:pyruvate aminotransferase activity(GO:0047312) glutamine-phenylpyruvate transaminase activity(GO:0047316) L-glutamine:pyruvate aminotransferase activity(GO:0047945) |
| 4.2 | 12.6 | GO:0061769 | ribosylnicotinamide kinase activity(GO:0050262) ribosylnicotinate kinase activity(GO:0061769) |
| 1.5 | 7.4 | GO:0004452 | isopentenyl-diphosphate delta-isomerase activity(GO:0004452) |
| 1.3 | 3.9 | GO:0019153 | protein-disulfide reductase (glutathione) activity(GO:0019153) |
| 1.3 | 7.7 | GO:1990254 | keratin filament binding(GO:1990254) |
| 1.2 | 35.6 | GO:0043295 | glutathione binding(GO:0043295) oligopeptide binding(GO:1900750) |
| 1.1 | 12.7 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 1.0 | 20.3 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.5 | 4.2 | GO:0008474 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.4 | 2.2 | GO:0008384 | IkappaB kinase activity(GO:0008384) |
| 0.3 | 3.6 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.3 | 2.0 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.2 | 1.7 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.2 | 0.7 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.2 | 0.7 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 0.1 | 1.3 | GO:0008889 | glycerophosphodiester phosphodiesterase activity(GO:0008889) |
| 0.1 | 7.8 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.1 | 0.5 | GO:0004758 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.1 | 2.7 | GO:0033130 | acetylcholine receptor binding(GO:0033130) |
| 0.1 | 0.7 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.1 | 10.5 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.1 | 0.5 | GO:0004030 | aldehyde dehydrogenase [NAD(P)+] activity(GO:0004030) |
| 0.1 | 0.4 | GO:0004661 | protein geranylgeranyltransferase activity(GO:0004661) |
| 0.1 | 3.7 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.0 | 0.7 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.0 | 2.7 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 1.6 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.0 | 23.9 | GO:0008289 | lipid binding(GO:0008289) |
| 0.0 | 1.1 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 1.8 | GO:0000980 | RNA polymerase II distal enhancer sequence-specific DNA binding(GO:0000980) |
| 0.0 | 12.6 | GO:0008233 | peptidase activity(GO:0008233) |
| 0.0 | 5.6 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 1.8 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.0 | 16.1 | GO:0019904 | protein domain specific binding(GO:0019904) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.8 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 45.4 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.3 | 7.8 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.2 | 7.4 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.2 | 4.2 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 14.6 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 2.2 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.1 | 2.2 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |
| 0.0 | 0.5 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.0 | 1.8 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |