Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
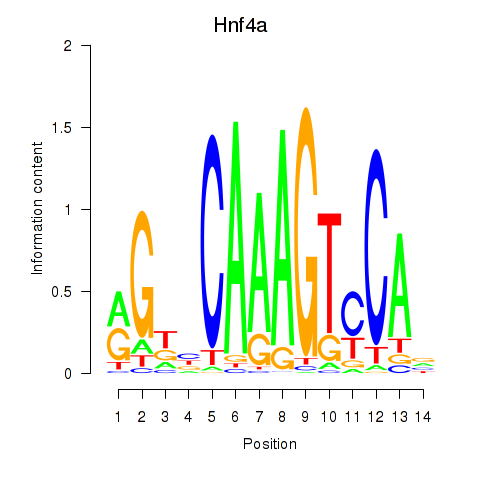
Results for Hnf4a
Z-value: 10.60
Transcription factors associated with Hnf4a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hnf4a
|
ENSMUSG00000017950.17 | Hnf4a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hnf4a | mm39_v1_chr2_+_163389068_163389108 | 0.61 | 7.7e-05 | Click! |
Activity profile of Hnf4a motif
Sorted Z-values of Hnf4a motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Hnf4a
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_39275518 | 160.92 |
ENSMUST00000003137.15
|
Cyp2c29
|
cytochrome P450, family 2, subfamily c, polypeptide 29 |
| chr4_-_62005498 | 128.85 |
ENSMUST00000107488.4
ENSMUST00000107472.8 ENSMUST00000084531.11 |
Mup3
|
major urinary protein 3 |
| chr17_-_46749370 | 103.46 |
ENSMUST00000087012.7
|
Slc22a7
|
solute carrier family 22 (organic anion transporter), member 7 |
| chr10_-_128796834 | 99.95 |
ENSMUST00000026398.5
|
Mettl7b
|
methyltransferase like 7B |
| chr15_-_82648376 | 97.82 |
ENSMUST00000055721.6
|
Cyp2d40
|
cytochrome P450, family 2, subfamily d, polypeptide 40 |
| chr19_-_40062174 | 93.25 |
ENSMUST00000048959.5
|
Cyp2c54
|
cytochrome P450, family 2, subfamily c, polypeptide 54 |
| chr2_+_172994841 | 80.60 |
ENSMUST00000029017.6
|
Pck1
|
phosphoenolpyruvate carboxykinase 1, cytosolic |
| chr2_-_25390625 | 74.10 |
ENSMUST00000040042.11
|
C8g
|
complement component 8, gamma polypeptide |
| chr3_+_122688721 | 69.88 |
ENSMUST00000023820.6
|
Fabp2
|
fatty acid binding protein 2, intestinal |
| chr4_-_60697274 | 69.46 |
ENSMUST00000117932.2
|
Mup12
|
major urinary protein 12 |
| chr19_+_12610668 | 67.05 |
ENSMUST00000044976.12
|
Glyat
|
glycine-N-acyltransferase |
| chr17_-_34962823 | 66.90 |
ENSMUST00000069507.9
|
C4b
|
complement component 4B (Chido blood group) |
| chr19_+_12610870 | 66.05 |
ENSMUST00000119960.2
|
Glyat
|
glycine-N-acyltransferase |
| chr19_+_39980868 | 65.62 |
ENSMUST00000049178.3
|
Cyp2c37
|
cytochrome P450, family 2. subfamily c, polypeptide 37 |
| chr19_-_8382424 | 64.02 |
ENSMUST00000064507.12
ENSMUST00000120540.2 ENSMUST00000096269.11 |
Slc22a30
|
solute carrier family 22, member 30 |
| chr7_-_99345016 | 62.30 |
ENSMUST00000107086.9
|
Slco2b1
|
solute carrier organic anion transporter family, member 2b1 |
| chr9_-_46146928 | 57.44 |
ENSMUST00000118649.8
|
Apoc3
|
apolipoprotein C-III |
| chr15_+_82336535 | 56.17 |
ENSMUST00000089129.7
ENSMUST00000229313.2 ENSMUST00000231136.2 |
Cyp2d9
|
cytochrome P450, family 2, subfamily d, polypeptide 9 |
| chr19_-_44396092 | 54.09 |
ENSMUST00000041331.4
|
Scd1
|
stearoyl-Coenzyme A desaturase 1 |
| chr19_+_38995463 | 53.52 |
ENSMUST00000025966.5
|
Cyp2c55
|
cytochrome P450, family 2, subfamily c, polypeptide 55 |
| chr11_+_101258368 | 50.13 |
ENSMUST00000019469.3
|
G6pc
|
glucose-6-phosphatase, catalytic |
| chr7_-_99344779 | 46.87 |
ENSMUST00000137914.2
ENSMUST00000207090.2 ENSMUST00000208225.2 |
Slco2b1
|
solute carrier organic anion transporter family, member 2b1 |
| chr11_+_69945157 | 45.25 |
ENSMUST00000108585.9
ENSMUST00000018699.13 |
Asgr1
|
asialoglycoprotein receptor 1 |
| chr3_-_121608809 | 45.18 |
ENSMUST00000197383.5
|
Abcd3
|
ATP-binding cassette, sub-family D (ALD), member 3 |
| chr19_-_44396269 | 44.34 |
ENSMUST00000235741.2
|
Scd1
|
stearoyl-Coenzyme A desaturase 1 |
| chr3_-_121608859 | 44.06 |
ENSMUST00000029770.8
|
Abcd3
|
ATP-binding cassette, sub-family D (ALD), member 3 |
| chrX_+_10118544 | 43.27 |
ENSMUST00000049910.13
|
Otc
|
ornithine transcarbamylase |
| chr17_-_46749320 | 41.90 |
ENSMUST00000233575.2
|
Slc22a7
|
solute carrier family 22 (organic anion transporter), member 7 |
| chr19_-_39637489 | 41.71 |
ENSMUST00000067328.7
|
Cyp2c67
|
cytochrome P450, family 2, subfamily c, polypeptide 67 |
| chr19_+_4771089 | 41.68 |
ENSMUST00000238976.3
|
Sptbn2
|
spectrin beta, non-erythrocytic 2 |
| chr17_-_74257164 | 40.90 |
ENSMUST00000024866.6
|
Xdh
|
xanthine dehydrogenase |
| chrX_+_10118600 | 40.66 |
ENSMUST00000115528.3
|
Otc
|
ornithine transcarbamylase |
| chr3_+_118355778 | 38.82 |
ENSMUST00000039177.12
|
Dpyd
|
dihydropyrimidine dehydrogenase |
| chr19_-_39451509 | 37.42 |
ENSMUST00000035488.3
|
Cyp2c38
|
cytochrome P450, family 2, subfamily c, polypeptide 38 |
| chr7_-_105249308 | 36.30 |
ENSMUST00000210531.2
ENSMUST00000033185.10 |
Hpx
|
hemopexin |
| chr9_+_46151994 | 36.16 |
ENSMUST00000034585.7
|
Apoa4
|
apolipoprotein A-IV |
| chr9_-_121745354 | 35.14 |
ENSMUST00000062474.5
|
Cyp8b1
|
cytochrome P450, family 8, subfamily b, polypeptide 1 |
| chr19_-_8109346 | 35.06 |
ENSMUST00000065651.5
|
Slc22a28
|
solute carrier family 22, member 28 |
| chr7_+_44114815 | 35.01 |
ENSMUST00000035929.11
ENSMUST00000146128.8 |
Aspdh
|
aspartate dehydrogenase domain containing |
| chr12_-_103871146 | 33.70 |
ENSMUST00000074051.6
|
Serpina1c
|
serine (or cysteine) peptidase inhibitor, clade A, member 1C |
| chr7_+_44114857 | 33.60 |
ENSMUST00000135624.2
|
Aspdh
|
aspartate dehydrogenase domain containing |
| chr7_-_99344832 | 33.52 |
ENSMUST00000145381.8
|
Slco2b1
|
solute carrier organic anion transporter family, member 2b1 |
| chr4_+_134124691 | 33.25 |
ENSMUST00000105870.8
|
Pafah2
|
platelet-activating factor acetylhydrolase 2 |
| chr8_-_122671588 | 33.02 |
ENSMUST00000057653.8
|
Car5a
|
carbonic anhydrase 5a, mitochondrial |
| chr17_-_35081129 | 33.00 |
ENSMUST00000154526.8
|
Cfb
|
complement factor B |
| chr8_-_93956143 | 32.86 |
ENSMUST00000176282.2
ENSMUST00000034173.14 |
Ces1e
|
carboxylesterase 1E |
| chr2_+_58645189 | 32.58 |
ENSMUST00000102755.4
ENSMUST00000230627.2 ENSMUST00000229923.2 |
Upp2
|
uridine phosphorylase 2 |
| chr16_+_37400590 | 32.35 |
ENSMUST00000159787.8
|
Hgd
|
homogentisate 1, 2-dioxygenase |
| chr17_-_35081456 | 32.28 |
ENSMUST00000025229.11
ENSMUST00000176203.9 ENSMUST00000128767.8 |
Cfb
|
complement factor B |
| chr16_+_37400500 | 31.11 |
ENSMUST00000160847.2
|
Hgd
|
homogentisate 1, 2-dioxygenase |
| chr15_+_3300249 | 30.69 |
ENSMUST00000082424.12
ENSMUST00000159158.9 ENSMUST00000159216.10 ENSMUST00000160311.3 |
Selenop
|
selenoprotein P |
| chr2_-_134396268 | 29.36 |
ENSMUST00000028704.3
|
Hao1
|
hydroxyacid oxidase 1, liver |
| chr15_+_101184488 | 29.21 |
ENSMUST00000229525.2
ENSMUST00000230525.2 |
Atg101
|
autophagy related 101 |
| chr1_+_74324089 | 28.14 |
ENSMUST00000113805.8
ENSMUST00000027370.13 ENSMUST00000087226.11 |
Pnkd
|
paroxysmal nonkinesiogenic dyskinesia |
| chr2_+_58644922 | 27.89 |
ENSMUST00000059102.13
|
Upp2
|
uridine phosphorylase 2 |
| chr7_+_143027473 | 27.66 |
ENSMUST00000052348.12
|
Slc22a18
|
solute carrier family 22 (organic cation transporter), member 18 |
| chr2_-_27138347 | 27.20 |
ENSMUST00000139312.8
|
Sardh
|
sarcosine dehydrogenase |
| chr8_+_13076024 | 27.13 |
ENSMUST00000033820.4
|
F7
|
coagulation factor VII |
| chr14_-_55995912 | 26.79 |
ENSMUST00000001497.9
|
Cideb
|
cell death-inducing DNA fragmentation factor, alpha subunit-like effector B |
| chr4_-_62069046 | 26.78 |
ENSMUST00000077719.4
|
Mup21
|
major urinary protein 21 |
| chr5_+_90708962 | 26.09 |
ENSMUST00000094615.8
ENSMUST00000200765.2 |
Albfm1
|
albumin superfamily member 1 |
| chr7_+_45546365 | 25.13 |
ENSMUST00000069772.16
ENSMUST00000210503.2 ENSMUST00000209350.2 |
Tmem143
|
transmembrane protein 143 |
| chr1_+_171041583 | 24.94 |
ENSMUST00000111328.8
|
Nr1i3
|
nuclear receptor subfamily 1, group I, member 3 |
| chr3_+_118355811 | 24.59 |
ENSMUST00000149101.3
|
Dpyd
|
dihydropyrimidine dehydrogenase |
| chr1_+_171041539 | 24.11 |
ENSMUST00000005820.11
ENSMUST00000075469.12 ENSMUST00000155126.8 |
Nr1i3
|
nuclear receptor subfamily 1, group I, member 3 |
| chr15_-_82291372 | 23.74 |
ENSMUST00000230198.2
ENSMUST00000230248.2 ENSMUST00000072776.5 ENSMUST00000229911.2 |
Cyp2d10
|
cytochrome P450, family 2, subfamily d, polypeptide 10 |
| chr4_+_141473983 | 22.68 |
ENSMUST00000038161.5
|
Agmat
|
agmatine ureohydrolase (agmatinase) |
| chr1_+_175459559 | 21.56 |
ENSMUST00000040250.15
|
Kmo
|
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chr6_-_128503666 | 21.52 |
ENSMUST00000143664.2
ENSMUST00000112132.8 |
Pzp
|
PZP, alpha-2-macroglobulin like |
| chr1_+_175459735 | 21.42 |
ENSMUST00000097458.4
|
Kmo
|
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chr15_-_82278223 | 21.34 |
ENSMUST00000170255.2
|
Cyp2d11
|
cytochrome P450, family 2, subfamily d, polypeptide 11 |
| chr8_+_87350672 | 21.32 |
ENSMUST00000034141.18
ENSMUST00000122188.10 |
Lonp2
|
lon peptidase 2, peroxisomal |
| chr15_+_10358611 | 20.64 |
ENSMUST00000110541.8
ENSMUST00000110542.8 |
Agxt2
|
alanine-glyoxylate aminotransferase 2 |
| chr8_+_13087805 | 20.61 |
ENSMUST00000128418.8
ENSMUST00000152034.2 |
F10
|
coagulation factor X |
| chr5_+_114284585 | 20.26 |
ENSMUST00000102582.8
|
Acacb
|
acetyl-Coenzyme A carboxylase beta |
| chr6_+_71176811 | 19.79 |
ENSMUST00000067492.8
|
Fabp1
|
fatty acid binding protein 1, liver |
| chr15_+_10358643 | 19.31 |
ENSMUST00000022858.8
|
Agxt2
|
alanine-glyoxylate aminotransferase 2 |
| chr8_+_13087292 | 19.25 |
ENSMUST00000063820.12
ENSMUST00000033821.11 |
F10
|
coagulation factor X |
| chr16_-_30086317 | 19.19 |
ENSMUST00000064856.9
|
Cpn2
|
carboxypeptidase N, polypeptide 2 |
| chr17_-_35100980 | 19.14 |
ENSMUST00000152417.8
ENSMUST00000146299.8 |
C2
Gm20547
|
complement component 2 (within H-2S) predicted gene 20547 |
| chr6_-_125357756 | 19.12 |
ENSMUST00000042647.7
|
Plekhg6
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 6 |
| chr19_+_44980565 | 19.03 |
ENSMUST00000179305.2
|
Sema4g
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4G |
| chr1_+_139429430 | 18.98 |
ENSMUST00000027615.7
|
F13b
|
coagulation factor XIII, beta subunit |
| chr11_+_73090270 | 18.96 |
ENSMUST00000006105.7
|
Shpk
|
sedoheptulokinase |
| chr11_-_77784922 | 18.81 |
ENSMUST00000017597.5
|
Pipox
|
pipecolic acid oxidase |
| chr1_+_87983099 | 18.31 |
ENSMUST00000138182.8
ENSMUST00000113142.10 |
Ugt1a10
|
UDP glycosyltransferase 1 family, polypeptide A10 |
| chr2_-_110136074 | 18.31 |
ENSMUST00000046233.9
|
Bbox1
|
butyrobetaine (gamma), 2-oxoglutarate dioxygenase 1 (gamma-butyrobetaine hydroxylase) |
| chr15_+_82439273 | 17.72 |
ENSMUST00000229103.2
ENSMUST00000068861.8 ENSMUST00000229904.2 |
Cyp2d12
|
cytochrome P450, family 2, subfamily d, polypeptide 12 |
| chr17_-_34822649 | 17.62 |
ENSMUST00000015622.8
|
Rnf5
|
ring finger protein 5 |
| chr19_-_39801188 | 17.53 |
ENSMUST00000162507.2
ENSMUST00000160476.9 ENSMUST00000239028.2 |
Cyp2c40
|
cytochrome P450, family 2, subfamily c, polypeptide 40 |
| chr1_-_136888118 | 17.09 |
ENSMUST00000192357.6
ENSMUST00000027649.14 |
Nr5a2
|
nuclear receptor subfamily 5, group A, member 2 |
| chr2_-_77000878 | 16.84 |
ENSMUST00000111833.3
|
Ccdc141
|
coiled-coil domain containing 141 |
| chr9_-_110576192 | 16.17 |
ENSMUST00000199791.2
|
Pth1r
|
parathyroid hormone 1 receptor |
| chr1_+_171052623 | 16.17 |
ENSMUST00000111321.8
ENSMUST00000005824.12 ENSMUST00000111320.8 ENSMUST00000111319.2 |
Apoa2
|
apolipoprotein A-II |
| chr11_+_78389913 | 16.07 |
ENSMUST00000017488.5
|
Vtn
|
vitronectin |
| chr11_-_61157986 | 15.98 |
ENSMUST00000066277.10
ENSMUST00000074127.14 ENSMUST00000108715.3 |
Aldh3a2
|
aldehyde dehydrogenase family 3, subfamily A2 |
| chr9_-_21913833 | 15.97 |
ENSMUST00000115336.10
|
Odad3
|
outer dynein arm docking complex subunit 3 |
| chr10_+_18720760 | 15.44 |
ENSMUST00000019998.9
|
Perp
|
PERP, TP53 apoptosis effector |
| chr1_+_87983189 | 15.41 |
ENSMUST00000173325.2
|
Ugt1a10
|
UDP glycosyltransferase 1 family, polypeptide A10 |
| chr2_+_92205651 | 15.38 |
ENSMUST00000028650.9
|
Pex16
|
peroxisomal biogenesis factor 16 |
| chr5_-_104125192 | 15.33 |
ENSMUST00000120320.8
|
Hsd17b13
|
hydroxysteroid (17-beta) dehydrogenase 13 |
| chr2_-_69172944 | 15.22 |
ENSMUST00000102709.8
ENSMUST00000102710.10 ENSMUST00000180142.2 |
Abcb11
|
ATP-binding cassette, sub-family B (MDR/TAP), member 11 |
| chr5_-_104125270 | 14.77 |
ENSMUST00000112803.3
|
Hsd17b13
|
hydroxysteroid (17-beta) dehydrogenase 13 |
| chr5_-_104125226 | 14.67 |
ENSMUST00000048118.15
|
Hsd17b13
|
hydroxysteroid (17-beta) dehydrogenase 13 |
| chr15_+_98997293 | 14.63 |
ENSMUST00000061295.7
|
Dnajc22
|
DnaJ heat shock protein family (Hsp40) member C22 |
| chr9_-_21913896 | 14.52 |
ENSMUST00000044926.6
|
Odad3
|
outer dynein arm docking complex subunit 3 |
| chr3_-_10400710 | 14.27 |
ENSMUST00000078748.4
|
Slc10a5
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 5 |
| chr7_-_45679703 | 14.25 |
ENSMUST00000002850.8
|
Abcc6
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 6 |
| chr12_+_37292029 | 14.24 |
ENSMUST00000160390.2
|
Agmo
|
alkylglycerol monooxygenase |
| chr4_+_118384183 | 14.08 |
ENSMUST00000106367.8
|
2610528J11Rik
|
RIKEN cDNA 2610528J11 gene |
| chr12_+_21366386 | 14.06 |
ENSMUST00000076813.8
ENSMUST00000221693.2 ENSMUST00000223345.2 ENSMUST00000222344.2 |
Iah1
|
isoamyl acetate-hydrolyzing esterase 1 homolog |
| chr7_-_30754792 | 13.72 |
ENSMUST00000206328.2
|
Fxyd1
|
FXYD domain-containing ion transport regulator 1 |
| chr1_+_157353696 | 13.67 |
ENSMUST00000111700.8
|
Sec16b
|
SEC16 homolog B (S. cerevisiae) |
| chr4_+_95467653 | 13.54 |
ENSMUST00000043335.11
|
Fggy
|
FGGY carbohydrate kinase domain containing |
| chr5_+_137568982 | 13.48 |
ENSMUST00000196471.5
ENSMUST00000198783.5 |
Tfr2
|
transferrin receptor 2 |
| chr15_-_76501525 | 13.21 |
ENSMUST00000230977.2
|
Slc39a4
|
solute carrier family 39 (zinc transporter), member 4 |
| chr17_-_46956054 | 13.05 |
ENSMUST00000003642.7
|
Klc4
|
kinesin light chain 4 |
| chrX_-_16683578 | 13.04 |
ENSMUST00000040820.13
|
Maob
|
monoamine oxidase B |
| chr7_+_29931735 | 12.80 |
ENSMUST00000108193.2
ENSMUST00000108192.2 |
Polr2i
|
polymerase (RNA) II (DNA directed) polypeptide I |
| chr7_-_30755007 | 12.59 |
ENSMUST00000206474.2
ENSMUST00000205807.2 ENSMUST00000039909.13 ENSMUST00000206305.2 ENSMUST00000205439.2 |
Fxyd1
|
FXYD domain-containing ion transport regulator 1 |
| chr19_-_11058452 | 12.48 |
ENSMUST00000025636.8
|
Ms4a8a
|
membrane-spanning 4-domains, subfamily A, member 8A |
| chr16_-_17745999 | 12.28 |
ENSMUST00000003622.16
|
Slc25a1
|
solute carrier family 25 (mitochondrial carrier, citrate transporter), member 1 |
| chr11_+_70410009 | 12.17 |
ENSMUST00000057685.3
|
Gltpd2
|
glycolipid transfer protein domain containing 2 |
| chr12_+_37291728 | 12.11 |
ENSMUST00000160768.8
|
Agmo
|
alkylglycerol monooxygenase |
| chr7_+_29931309 | 12.06 |
ENSMUST00000019882.16
ENSMUST00000149654.8 |
Polr2i
|
polymerase (RNA) II (DNA directed) polypeptide I |
| chr8_-_13250535 | 11.79 |
ENSMUST00000165605.4
ENSMUST00000209691.2 ENSMUST00000211128.2 ENSMUST00000210317.2 |
Grtp1
|
GH regulated TBC protein 1 |
| chr5_+_30437579 | 11.74 |
ENSMUST00000145167.9
|
Selenoi
|
selenoprotein I |
| chr3_+_59939175 | 11.59 |
ENSMUST00000029325.5
|
Aadac
|
arylacetamide deacetylase |
| chr12_+_37291632 | 11.50 |
ENSMUST00000049874.14
|
Agmo
|
alkylglycerol monooxygenase |
| chr9_+_44584523 | 11.39 |
ENSMUST00000034609.11
ENSMUST00000071219.12 |
Treh
|
trehalase (brush-border membrane glycoprotein) |
| chr6_+_72333209 | 11.23 |
ENSMUST00000206531.2
|
Tmem150a
|
transmembrane protein 150A |
| chr9_-_110576124 | 11.15 |
ENSMUST00000199862.5
ENSMUST00000198865.5 |
Pth1r
|
parathyroid hormone 1 receptor |
| chr4_+_97665992 | 11.07 |
ENSMUST00000107062.9
ENSMUST00000052018.12 ENSMUST00000107057.8 |
Nfia
|
nuclear factor I/A |
| chr4_+_95467701 | 11.05 |
ENSMUST00000150830.2
ENSMUST00000134012.9 |
Fggy
|
FGGY carbohydrate kinase domain containing |
| chr7_-_135130374 | 10.82 |
ENSMUST00000053716.8
|
Clrn3
|
clarin 3 |
| chr6_-_85114725 | 10.73 |
ENSMUST00000174769.2
ENSMUST00000174286.3 ENSMUST00000045986.8 |
Spr
|
sepiapterin reductase |
| chr9_-_55419442 | 10.64 |
ENSMUST00000034866.9
|
Etfa
|
electron transferring flavoprotein, alpha polypeptide |
| chr15_-_89064936 | 10.61 |
ENSMUST00000109331.9
|
Plxnb2
|
plexin B2 |
| chr12_+_59178072 | 10.44 |
ENSMUST00000176464.8
ENSMUST00000170992.9 ENSMUST00000176322.8 |
Mia2
|
MIA SH3 domain ER export factor 2 |
| chr5_-_24652775 | 10.41 |
ENSMUST00000123167.2
ENSMUST00000030799.15 |
Tmub1
|
transmembrane and ubiquitin-like domain containing 1 |
| chr12_+_59177780 | 10.39 |
ENSMUST00000177460.8
|
Mia2
|
MIA SH3 domain ER export factor 2 |
| chr1_-_24044688 | 10.32 |
ENSMUST00000027338.4
|
Sdhaf4
|
succinate dehydrogenase complex assembly factor 4 |
| chr9_+_21914083 | 10.10 |
ENSMUST00000216344.2
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr1_-_90897329 | 9.89 |
ENSMUST00000130042.2
ENSMUST00000027529.12 |
Rab17
|
RAB17, member RAS oncogene family |
| chr19_+_39499288 | 9.89 |
ENSMUST00000025968.5
|
Cyp2c39
|
cytochrome P450, family 2, subfamily c, polypeptide 39 |
| chr4_-_83242490 | 9.80 |
ENSMUST00000102823.10
|
Ttc39b
|
tetratricopeptide repeat domain 39B |
| chr19_-_8196196 | 9.76 |
ENSMUST00000113298.9
|
Slc22a29
|
solute carrier family 22. member 29 |
| chr12_+_59178258 | 9.45 |
ENSMUST00000177162.8
|
Mia2
|
MIA SH3 domain ER export factor 2 |
| chr11_+_74540284 | 9.29 |
ENSMUST00000117818.2
ENSMUST00000092915.12 |
Cluh
|
clustered mitochondria (cluA/CLU1) homolog |
| chr4_-_138053603 | 9.19 |
ENSMUST00000030536.13
|
Pink1
|
PTEN induced putative kinase 1 |
| chr4_-_138053545 | 9.08 |
ENSMUST00000105817.4
|
Pink1
|
PTEN induced putative kinase 1 |
| chr4_+_118384426 | 9.01 |
ENSMUST00000030261.6
|
2610528J11Rik
|
RIKEN cDNA 2610528J11 gene |
| chr9_+_21914334 | 8.90 |
ENSMUST00000115331.10
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr4_+_97665843 | 8.74 |
ENSMUST00000075448.13
ENSMUST00000092532.13 |
Nfia
|
nuclear factor I/A |
| chr10_+_81395242 | 8.58 |
ENSMUST00000002518.9
|
Tle5
|
TLE family member 5, transcriptional modulator |
| chr11_-_12362136 | 8.36 |
ENSMUST00000174874.8
|
Cobl
|
cordon-bleu WH2 repeat |
| chr5_+_8943943 | 8.20 |
ENSMUST00000196067.2
|
Abcb4
|
ATP-binding cassette, sub-family B (MDR/TAP), member 4 |
| chr19_-_39875192 | 8.17 |
ENSMUST00000168838.3
|
Cyp2c69
|
cytochrome P450, family 2, subfamily c, polypeptide 69 |
| chr2_-_130480014 | 7.85 |
ENSMUST00000089561.10
ENSMUST00000110260.8 |
Lzts3
|
leucine zipper, putative tumor suppressor family member 3 |
| chr4_+_126503611 | 7.79 |
ENSMUST00000097886.4
ENSMUST00000164362.2 |
5730409E04Rik
|
RIKEN cDNA 5730409E04Rik gene |
| chr9_+_21914296 | 7.77 |
ENSMUST00000003493.9
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr2_-_90748331 | 7.61 |
ENSMUST00000111461.13
|
Ptpmt1
|
protein tyrosine phosphatase, mitochondrial 1 |
| chr10_-_25412010 | 7.57 |
ENSMUST00000179685.3
|
Smlr1
|
small leucine-rich protein 1 |
| chr19_+_46140942 | 7.45 |
ENSMUST00000026254.14
|
Gbf1
|
golgi-specific brefeldin A-resistance factor 1 |
| chr12_+_59177552 | 7.05 |
ENSMUST00000175877.8
|
Mia2
|
MIA SH3 domain ER export factor 2 |
| chr15_-_90934021 | 6.75 |
ENSMUST00000109287.4
ENSMUST00000067205.16 ENSMUST00000088614.13 |
Kif21a
|
kinesin family member 21A |
| chr12_-_80159768 | 6.66 |
ENSMUST00000219642.2
ENSMUST00000165114.2 ENSMUST00000218835.2 ENSMUST00000021552.3 |
Zfp36l1
|
zinc finger protein 36, C3H type-like 1 |
| chr19_-_10581622 | 6.52 |
ENSMUST00000037678.7
|
Tkfc
|
triokinase, FMN cyclase |
| chr5_+_8943677 | 6.51 |
ENSMUST00000003717.13
|
Abcb4
|
ATP-binding cassette, sub-family B (MDR/TAP), member 4 |
| chr11_+_58090382 | 6.24 |
ENSMUST00000035266.11
ENSMUST00000094169.11 ENSMUST00000168280.2 ENSMUST00000058704.9 |
Igtp
Irgm2
|
interferon gamma induced GTPase immunity-related GTPase family M member 2 |
| chr15_-_90934059 | 6.22 |
ENSMUST00000109288.9
ENSMUST00000100304.11 |
Kif21a
|
kinesin family member 21A |
| chr5_+_31454787 | 6.03 |
ENSMUST00000201166.4
ENSMUST00000072228.9 |
Gckr
|
glucokinase regulatory protein |
| chr2_-_119060330 | 5.97 |
ENSMUST00000110820.3
|
Ppp1r14d
|
protein phosphatase 1, regulatory inhibitor subunit 14D |
| chr4_-_83242366 | 5.96 |
ENSMUST00000030205.14
ENSMUST00000048274.11 |
Ttc39b
|
tetratricopeptide repeat domain 39B |
| chr2_-_77000936 | 5.58 |
ENSMUST00000164114.9
ENSMUST00000049544.14 |
Ccdc141
|
coiled-coil domain containing 141 |
| chr3_-_79535966 | 5.53 |
ENSMUST00000120992.8
|
Etfdh
|
electron transferring flavoprotein, dehydrogenase |
| chr19_+_5138562 | 5.51 |
ENSMUST00000238093.2
ENSMUST00000025811.6 ENSMUST00000237025.2 |
Yif1a
|
Yip1 interacting factor homolog A (S. cerevisiae) |
| chr8_+_3637785 | 5.49 |
ENSMUST00000171962.3
ENSMUST00000207712.2 ENSMUST00000207970.2 ENSMUST00000207533.2 ENSMUST00000208240.2 ENSMUST00000207432.2 ENSMUST00000207077.2 |
Camsap3
|
calmodulin regulated spectrin-associated protein family, member 3 |
| chr2_-_17465410 | 5.43 |
ENSMUST00000145492.2
|
Nebl
|
nebulette |
| chr2_+_58991182 | 5.38 |
ENSMUST00000168631.8
ENSMUST00000102754.11 ENSMUST00000123908.8 |
Pkp4
|
plakophilin 4 |
| chr15_-_27630724 | 5.35 |
ENSMUST00000059662.8
|
Otulin
|
OTU deubiquitinase with linear linkage specificity |
| chr7_+_119773070 | 5.31 |
ENSMUST00000033201.7
|
Anks4b
|
ankyrin repeat and sterile alpha motif domain containing 4B |
| chr5_-_115109118 | 5.14 |
ENSMUST00000031535.12
|
Hnf1a
|
HNF1 homeobox A |
| chr5_+_92285748 | 5.09 |
ENSMUST00000031355.10
ENSMUST00000202155.2 |
Uso1
|
USO1 vesicle docking factor |
| chr4_-_119395966 | 5.04 |
ENSMUST00000079611.13
|
Frg2f1
|
FSHD region gene 2 family member 1 |
| chr2_+_3705824 | 5.04 |
ENSMUST00000115054.9
|
Fam107b
|
family with sequence similarity 107, member B |
| chr1_+_165958036 | 4.89 |
ENSMUST00000166860.2
|
Gpa33
|
glycoprotein A33 (transmembrane) |
| chr5_-_125371162 | 4.84 |
ENSMUST00000127148.2
|
Scarb1
|
scavenger receptor class B, member 1 |
| chr9_-_107482462 | 4.68 |
ENSMUST00000194433.6
|
Sema3b
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3B |
| chr1_+_165957784 | 4.67 |
ENSMUST00000060833.14
|
Gpa33
|
glycoprotein A33 (transmembrane) |
| chrX_-_104972844 | 4.59 |
ENSMUST00000198441.5
ENSMUST00000137453.8 |
Atrx
|
ATRX, chromatin remodeler |
| chr4_+_118266582 | 4.57 |
ENSMUST00000144577.2
|
Med8
|
mediator complex subunit 8 |
| chr10_-_13428855 | 4.56 |
ENSMUST00000105539.2
|
Pex3
|
peroxisomal biogenesis factor 3 |
| chr1_+_165957909 | 4.44 |
ENSMUST00000166159.2
|
Gpa33
|
glycoprotein A33 (transmembrane) |
| chr1_+_185187000 | 4.38 |
ENSMUST00000061093.7
|
Slc30a10
|
solute carrier family 30, member 10 |
| chr9_-_35030479 | 4.37 |
ENSMUST00000213526.2
ENSMUST00000215089.2 |
St3gal4
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 4 |
| chr9_+_21914513 | 4.37 |
ENSMUST00000215795.2
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr2_+_157870653 | 4.35 |
ENSMUST00000152452.8
|
Rprd1b
|
regulation of nuclear pre-mRNA domain containing 1B |
| chr14_-_52257113 | 4.32 |
ENSMUST00000166169.4
ENSMUST00000226605.2 |
Zfp219
|
zinc finger protein 219 |
| chr7_+_66339637 | 4.27 |
ENSMUST00000153007.2
ENSMUST00000121777.9 ENSMUST00000150071.8 ENSMUST00000077967.13 |
Lins1
|
lines homolog 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 32.8 | 98.4 | GO:1903699 | tarsal gland development(GO:1903699) |
| 28.5 | 142.7 | GO:0071718 | sodium-independent icosanoid transport(GO:0071718) |
| 28.0 | 83.9 | GO:0042450 | arginine biosynthetic process via ornithine(GO:0042450) |
| 22.3 | 89.2 | GO:0015910 | peroxisomal long-chain fatty acid import(GO:0015910) |
| 21.1 | 63.4 | GO:0019860 | uracil catabolic process(GO:0006212) uracil metabolic process(GO:0019860) |
| 20.2 | 80.6 | GO:0061402 | glycerol biosynthetic process(GO:0006114) positive regulation of transcription from RNA polymerase II promoter in response to acidic pH(GO:0061402) |
| 14.4 | 57.4 | GO:0010982 | regulation of high-density lipoprotein particle clearance(GO:0010982) |
| 14.2 | 469.5 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 13.3 | 40.0 | GO:0042853 | glycine biosynthetic process, by transamination of glyoxylate(GO:0019265) L-alanine catabolic process(GO:0042853) |
| 12.1 | 36.2 | GO:0034442 | regulation of lipoprotein oxidation(GO:0034442) negative regulation of lipoprotein oxidation(GO:0034443) multicellular organism lipid catabolic process(GO:0044240) |
| 11.4 | 68.6 | GO:0006742 | NADP catabolic process(GO:0006742) |
| 9.5 | 274.2 | GO:0035634 | response to stilbenoid(GO:0035634) |
| 9.1 | 18.3 | GO:1903383 | neuron intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:0036482) positive regulation of mitochondrial electron transport, NADH to ubiquinone(GO:1902958) regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903383) negative regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903384) |
| 9.1 | 27.2 | GO:1901053 | sarcosine metabolic process(GO:1901052) sarcosine catabolic process(GO:1901053) |
| 8.2 | 139.4 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 8.2 | 40.9 | GO:0009115 | xanthine catabolic process(GO:0009115) |
| 7.2 | 43.0 | GO:0019805 | quinolinate biosynthetic process(GO:0019805) |
| 7.1 | 63.5 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 6.8 | 102.2 | GO:0015747 | urate transport(GO:0015747) |
| 6.8 | 20.3 | GO:0046949 | fatty-acyl-CoA biosynthetic process(GO:0046949) malonyl-CoA metabolic process(GO:2001293) |
| 6.3 | 19.0 | GO:0035963 | response to interleukin-13(GO:0035962) cellular response to interleukin-13(GO:0035963) |
| 6.3 | 18.8 | GO:0006553 | lysine metabolic process(GO:0006553) |
| 6.1 | 42.5 | GO:0060332 | positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) |
| 5.9 | 29.4 | GO:0009441 | glycolate metabolic process(GO:0009441) |
| 5.4 | 16.2 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 5.3 | 16.0 | GO:0006714 | sesquiterpenoid metabolic process(GO:0006714) phytol metabolic process(GO:0033306) fatty alcohol metabolic process(GO:1903173) |
| 5.2 | 222.5 | GO:0019369 | arachidonic acid metabolic process(GO:0019369) |
| 5.0 | 50.1 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 4.9 | 14.7 | GO:1903412 | response to bile acid(GO:1903412) cellular response to bile acid(GO:1903413) |
| 4.6 | 18.3 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 4.5 | 27.1 | GO:0010641 | positive regulation of platelet-derived growth factor receptor signaling pathway(GO:0010641) |
| 3.8 | 30.7 | GO:0001887 | selenium compound metabolic process(GO:0001887) |
| 3.8 | 22.7 | GO:0008295 | spermidine biosynthetic process(GO:0008295) |
| 3.8 | 22.6 | GO:0016557 | peroxisome membrane biogenesis(GO:0016557) |
| 3.3 | 9.9 | GO:0002414 | immune response in mucosal-associated lymphoid tissue(GO:0002386) immunoglobulin transcytosis in epithelial cells(GO:0002414) |
| 3.2 | 19.1 | GO:2000427 | positive regulation of apoptotic cell clearance(GO:2000427) |
| 3.0 | 15.2 | GO:0046618 | drug export(GO:0046618) |
| 2.9 | 37.9 | GO:0046485 | ether lipid metabolic process(GO:0046485) |
| 2.8 | 28.1 | GO:0061727 | methylglyoxal catabolic process to D-lactate via S-lactoyl-glutathione(GO:0019243) methylglyoxal catabolic process(GO:0051596) methylglyoxal catabolic process to lactate(GO:0061727) |
| 2.8 | 33.7 | GO:0052696 | flavonoid glucuronidation(GO:0052696) xenobiotic glucuronidation(GO:0052697) |
| 2.6 | 13.2 | GO:0034224 | cellular response to zinc ion starvation(GO:0034224) |
| 2.6 | 13.0 | GO:0014063 | negative regulation of serotonin secretion(GO:0014063) |
| 2.5 | 7.4 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 2.5 | 12.3 | GO:0015746 | tricarboxylic acid transport(GO:0006842) citrate transport(GO:0015746) |
| 2.4 | 26.3 | GO:1903278 | positive regulation of sodium ion export(GO:1903275) positive regulation of sodium ion export from cell(GO:1903278) |
| 2.3 | 7.0 | GO:0071929 | alpha-tubulin acetylation(GO:0071929) |
| 2.3 | 94.5 | GO:0050892 | intestinal absorption(GO:0050892) |
| 2.3 | 24.9 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 2.2 | 6.7 | GO:1904582 | regulation of intracellular mRNA localization(GO:1904580) positive regulation of intracellular mRNA localization(GO:1904582) |
| 1.9 | 11.6 | GO:0010898 | positive regulation of triglyceride catabolic process(GO:0010898) |
| 1.9 | 9.5 | GO:0005984 | disaccharide metabolic process(GO:0005984) |
| 1.8 | 10.7 | GO:0006558 | L-phenylalanine metabolic process(GO:0006558) erythrose 4-phosphate/phosphoenolpyruvate family amino acid metabolic process(GO:1902221) |
| 1.5 | 4.4 | GO:0071579 | regulation of zinc ion transport(GO:0071579) |
| 1.4 | 27.3 | GO:0060732 | positive regulation of inositol phosphate biosynthetic process(GO:0060732) |
| 1.4 | 17.1 | GO:0061113 | pancreas morphogenesis(GO:0061113) |
| 1.4 | 13.7 | GO:0048208 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 1.4 | 4.1 | GO:1901668 | regulation of superoxide dismutase activity(GO:1901668) |
| 1.3 | 16.1 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 1.3 | 11.7 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 1.2 | 13.5 | GO:0097460 | ferrous iron import into cell(GO:0097460) ferrous iron import across plasma membrane(GO:0098707) |
| 1.2 | 18.2 | GO:0019321 | pentose metabolic process(GO:0019321) |
| 1.2 | 6.0 | GO:0009750 | response to fructose(GO:0009750) |
| 1.2 | 8.4 | GO:0001757 | somite specification(GO:0001757) |
| 1.2 | 9.3 | GO:0032049 | cardiolipin biosynthetic process(GO:0032049) |
| 1.1 | 6.5 | GO:0039534 | negative regulation of MDA-5 signaling pathway(GO:0039534) |
| 1.1 | 38.9 | GO:0006515 | misfolded or incompletely synthesized protein catabolic process(GO:0006515) |
| 1.0 | 31.1 | GO:0006491 | N-glycan processing(GO:0006491) |
| 1.0 | 15.4 | GO:0002934 | desmosome organization(GO:0002934) |
| 1.0 | 6.7 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.9 | 19.8 | GO:0072189 | ureter development(GO:0072189) |
| 0.9 | 4.5 | GO:1904970 | brush border assembly(GO:1904970) |
| 0.8 | 5.1 | GO:0014826 | regulation of systemic arterial blood pressure by endothelin(GO:0003100) vein smooth muscle contraction(GO:0014826) |
| 0.8 | 3.3 | GO:1900535 | medium-chain fatty-acyl-CoA catabolic process(GO:0036114) fatty-acyl-CoA catabolic process(GO:0036115) long-chain fatty-acyl-CoA catabolic process(GO:0036116) palmitic acid metabolic process(GO:1900533) palmitic acid biosynthetic process(GO:1900535) |
| 0.8 | 4.1 | GO:0090366 | DNA cytosine deamination(GO:0070383) positive regulation of mRNA modification(GO:0090366) |
| 0.8 | 17.8 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.8 | 2.4 | GO:0097156 | fasciculation of motor neuron axon(GO:0097156) |
| 0.8 | 4.6 | GO:1901580 | post-embryonic appendage morphogenesis(GO:0035120) post-embryonic limb morphogenesis(GO:0035127) post-embryonic forelimb morphogenesis(GO:0035128) telomeric repeat-containing RNA transcription(GO:0097393) telomeric repeat-containing RNA transcription from RNA pol II promoter(GO:0097394) regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901580) negative regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901581) positive regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901582) |
| 0.8 | 2.3 | GO:0060464 | lung lobe formation(GO:0060464) diaphragm morphogenesis(GO:0060540) |
| 0.7 | 27.7 | GO:0015893 | drug transport(GO:0015893) |
| 0.7 | 7.6 | GO:0097118 | neuroligin clustering involved in postsynaptic membrane assembly(GO:0097118) |
| 0.7 | 26.8 | GO:0090200 | positive regulation of release of cytochrome c from mitochondria(GO:0090200) |
| 0.7 | 4.7 | GO:0015862 | uridine transport(GO:0015862) |
| 0.6 | 5.0 | GO:0007403 | glial cell fate determination(GO:0007403) |
| 0.6 | 16.0 | GO:0021692 | cerebellar Purkinje cell layer morphogenesis(GO:0021692) |
| 0.6 | 2.3 | GO:0036367 | adaptation of rhodopsin mediated signaling(GO:0016062) light adaption(GO:0036367) |
| 0.6 | 5.1 | GO:0043691 | reverse cholesterol transport(GO:0043691) |
| 0.6 | 3.9 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.5 | 97.6 | GO:0009636 | response to toxic substance(GO:0009636) |
| 0.5 | 4.2 | GO:0043314 | negative regulation of neutrophil degranulation(GO:0043314) negative regulation of blood vessel remodeling(GO:0060313) negative regulation of neutrophil activation(GO:1902564) |
| 0.5 | 6.2 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.5 | 9.3 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.5 | 25.9 | GO:0006730 | one-carbon metabolic process(GO:0006730) |
| 0.5 | 1.9 | GO:1902683 | regulation of receptor localization to synapse(GO:1902683) |
| 0.4 | 27.8 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.4 | 18.4 | GO:0070328 | acylglycerol homeostasis(GO:0055090) triglyceride homeostasis(GO:0070328) |
| 0.4 | 5.4 | GO:0070423 | nucleotide-binding oligomerization domain containing signaling pathway(GO:0070423) nucleotide-binding oligomerization domain containing 2 signaling pathway(GO:0070431) |
| 0.4 | 60.5 | GO:0009166 | nucleotide catabolic process(GO:0009166) |
| 0.4 | 59.9 | GO:0006956 | complement activation(GO:0006956) |
| 0.4 | 5.1 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.3 | 2.3 | GO:0060406 | positive regulation of penile erection(GO:0060406) |
| 0.3 | 22.4 | GO:0021799 | cerebral cortex radially oriented cell migration(GO:0021799) |
| 0.3 | 21.5 | GO:0007566 | embryo implantation(GO:0007566) |
| 0.3 | 11.2 | GO:0046854 | phosphatidylinositol phosphorylation(GO:0046854) |
| 0.3 | 0.8 | GO:2000170 | positive regulation of peptidyl-cysteine S-nitrosylation(GO:2000170) |
| 0.3 | 4.0 | GO:0070932 | histone H3 deacetylation(GO:0070932) |
| 0.3 | 1.5 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 0.2 | 29.2 | GO:0000045 | autophagosome assembly(GO:0000045) |
| 0.2 | 1.0 | GO:0061215 | regulation of pronephros size(GO:0035565) mesonephros morphogenesis(GO:0061206) mesonephric nephron development(GO:0061215) mesonephric nephron morphogenesis(GO:0061228) mesenchymal stem cell maintenance involved in mesonephric nephron morphogenesis(GO:0061235) regulation of mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:0061295) negative regulation of mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:0061296) mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:1901146) |
| 0.2 | 0.7 | GO:1904954 | negative regulation of axon extension involved in axon guidance(GO:0048843) Spemann organizer formation(GO:0060061) negative regulation of axon guidance(GO:1902668) canonical Wnt signaling pathway involved in midbrain dopaminergic neuron differentiation(GO:1904954) |
| 0.2 | 2.1 | GO:0014816 | skeletal muscle satellite cell differentiation(GO:0014816) positive regulation of myoblast proliferation(GO:2000288) |
| 0.2 | 5.0 | GO:0030497 | fatty acid elongation(GO:0030497) |
| 0.2 | 1.3 | GO:0051256 | mitotic spindle midzone assembly(GO:0051256) |
| 0.2 | 2.2 | GO:0090141 | positive regulation of mitochondrial fission(GO:0090141) |
| 0.2 | 1.0 | GO:0070162 | adiponectin secretion(GO:0070162) regulation of adiponectin secretion(GO:0070163) negative regulation of adiponectin secretion(GO:0070164) |
| 0.2 | 2.2 | GO:0070940 | dephosphorylation of RNA polymerase II C-terminal domain(GO:0070940) |
| 0.2 | 80.7 | GO:0032259 | methylation(GO:0032259) |
| 0.2 | 16.8 | GO:0051897 | positive regulation of protein kinase B signaling(GO:0051897) |
| 0.2 | 1.2 | GO:0010571 | positive regulation of nuclear cell cycle DNA replication(GO:0010571) |
| 0.2 | 48.6 | GO:0016042 | lipid catabolic process(GO:0016042) |
| 0.2 | 3.5 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.2 | 0.5 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.1 | 11.8 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.1 | 10.6 | GO:0007405 | neuroblast proliferation(GO:0007405) |
| 0.1 | 5.4 | GO:0032467 | positive regulation of cytokinesis(GO:0032467) |
| 0.1 | 10.4 | GO:0030433 | ER-associated ubiquitin-dependent protein catabolic process(GO:0030433) |
| 0.1 | 2.2 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.1 | 28.0 | GO:0043401 | steroid hormone mediated signaling pathway(GO:0043401) |
| 0.1 | 1.8 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.1 | 9.4 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.1 | 4.6 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.1 | 0.6 | GO:0060137 | maternal process involved in parturition(GO:0060137) |
| 0.1 | 15.6 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.1 | 2.6 | GO:2000300 | regulation of synaptic vesicle exocytosis(GO:2000300) |
| 0.1 | 4.0 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.1 | 2.1 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.1 | 1.3 | GO:0010758 | regulation of macrophage chemotaxis(GO:0010758) |
| 0.1 | 0.7 | GO:0090331 | negative regulation of platelet aggregation(GO:0090331) |
| 0.1 | 1.1 | GO:0008272 | sulfate transport(GO:0008272) |
| 0.1 | 10.7 | GO:0031668 | cellular response to extracellular stimulus(GO:0031668) |
| 0.1 | 1.5 | GO:0030325 | adrenal gland development(GO:0030325) |
| 0.1 | 4.4 | GO:0042273 | ribosomal large subunit biogenesis(GO:0042273) |
| 0.1 | 9.6 | GO:0035304 | regulation of protein dephosphorylation(GO:0035304) |
| 0.0 | 15.5 | GO:0043434 | response to peptide hormone(GO:0043434) |
| 0.0 | 2.0 | GO:0032981 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.0 | 2.6 | GO:0045197 | establishment or maintenance of epithelial cell apical/basal polarity(GO:0045197) |
| 0.0 | 0.7 | GO:0030705 | cytoskeleton-dependent intracellular transport(GO:0030705) |
| 0.0 | 1.3 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.0 | 8.9 | GO:0007018 | microtubule-based movement(GO:0007018) |
| 0.0 | 0.3 | GO:0015074 | DNA integration(GO:0015074) |
| 0.0 | 0.7 | GO:0022011 | myelination in peripheral nervous system(GO:0022011) peripheral nervous system axon ensheathment(GO:0032292) |
| 0.0 | 0.4 | GO:0097094 | craniofacial suture morphogenesis(GO:0097094) |
| 0.0 | 3.6 | GO:0008360 | regulation of cell shape(GO:0008360) |
| 0.0 | 0.9 | GO:0010761 | fibroblast migration(GO:0010761) |
| 0.0 | 1.2 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) |
| 0.0 | 2.1 | GO:0006261 | DNA-dependent DNA replication(GO:0006261) |
| 0.0 | 0.1 | GO:0046838 | phosphorylated carbohydrate dephosphorylation(GO:0046838) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.2 | 74.1 | GO:0005579 | membrane attack complex(GO:0005579) |
| 8.2 | 89.7 | GO:0045179 | apical cortex(GO:0045179) |
| 6.2 | 31.1 | GO:0017177 | glucosidase II complex(GO:0017177) |
| 5.5 | 60.5 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 5.2 | 41.7 | GO:0008091 | spectrin(GO:0008091) |
| 4.0 | 16.2 | GO:0017133 | mitochondrial electron transfer flavoprotein complex(GO:0017133) electron transfer flavoprotein complex(GO:0045251) |
| 3.9 | 90.8 | GO:0042627 | chylomicron(GO:0042627) |
| 3.8 | 111.5 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 2.3 | 16.1 | GO:0048237 | rough endoplasmic reticulum lumen(GO:0048237) |
| 2.3 | 16.0 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 2.2 | 73.1 | GO:0044216 | other organism(GO:0044215) other organism cell(GO:0044216) other organism part(GO:0044217) |
| 1.7 | 13.5 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 1.7 | 29.9 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 1.5 | 18.3 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 1.5 | 139.6 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 1.2 | 8.4 | GO:1990357 | terminal web(GO:1990357) |
| 1.1 | 37.3 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 1.1 | 5.4 | GO:0071797 | LUBAC complex(GO:0071797) |
| 1.0 | 17.6 | GO:0036513 | Derlin-1 retrotranslocation complex(GO:0036513) |
| 0.9 | 24.9 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.9 | 157.4 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.8 | 4.1 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.8 | 22.6 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) |
| 0.8 | 4.6 | GO:1990421 | subtelomeric heterochromatin(GO:1990421) nuclear subtelomeric heterochromatin(GO:1990707) |
| 0.7 | 9.6 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.7 | 4.0 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.6 | 63.3 | GO:0005811 | lipid particle(GO:0005811) |
| 0.6 | 29.2 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.6 | 5.5 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.5 | 89.7 | GO:0005777 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.4 | 2.6 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.4 | 9.9 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.4 | 7.1 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.4 | 103.2 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.4 | 27.6 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.3 | 27.3 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.3 | 634.9 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.3 | 44.9 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.2 | 5.4 | GO:0030057 | desmosome(GO:0030057) |
| 0.2 | 7.6 | GO:0031305 | integral component of mitochondrial inner membrane(GO:0031305) |
| 0.2 | 14.2 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.2 | 9.2 | GO:0016592 | mediator complex(GO:0016592) |
| 0.2 | 5.1 | GO:0045120 | pronucleus(GO:0045120) |
| 0.1 | 282.6 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.1 | 4.4 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.1 | 20.9 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.1 | 0.7 | GO:0070695 | FHF complex(GO:0070695) |
| 0.1 | 2.6 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.1 | 135.5 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.1 | 3.5 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.1 | 2.1 | GO:0016460 | myosin II complex(GO:0016460) |
| 0.1 | 1.2 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.1 | 11.8 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.1 | 2.3 | GO:0016591 | DNA-directed RNA polymerase II, holoenzyme(GO:0016591) |
| 0.1 | 2.1 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.1 | 5.4 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.0 | 3.9 | GO:0031226 | intrinsic component of plasma membrane(GO:0031226) |
| 0.0 | 4.6 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 1.3 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.0 | 2.1 | GO:0001650 | fibrillar center(GO:0001650) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 36.7 | 146.8 | GO:0071614 | linoleic acid epoxygenase activity(GO:0071614) |
| 33.3 | 133.1 | GO:0047961 | glycine N-acyltransferase activity(GO:0047961) |
| 32.2 | 193.0 | GO:0034875 | oxidoreductase activity, acting on CH or CH2 groups, quinone or similar compound as acceptor(GO:0033695) caffeine oxidase activity(GO:0034875) |
| 26.9 | 80.6 | GO:0004611 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 21.1 | 63.4 | GO:0002061 | uracil binding(GO:0002058) pyrimidine nucleobase binding(GO:0002061) |
| 21.0 | 83.9 | GO:0016743 | carboxyl- or carbamoyltransferase activity(GO:0016743) |
| 19.9 | 178.9 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 18.4 | 73.6 | GO:0070653 | high-density lipoprotein particle receptor binding(GO:0070653) |
| 16.4 | 98.4 | GO:0032896 | palmitoyl-CoA 9-desaturase activity(GO:0032896) |
| 12.6 | 37.9 | GO:0050479 | glyceryl-ether monooxygenase activity(GO:0050479) |
| 12.5 | 50.1 | GO:0050309 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 11.3 | 45.3 | GO:0016899 | oxidoreductase activity, acting on the CH-OH group of donors, oxygen as acceptor(GO:0016899) |
| 10.1 | 60.5 | GO:0004850 | uridine phosphorylase activity(GO:0004850) |
| 9.3 | 390.3 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 9.1 | 27.2 | GO:0008480 | sarcosine dehydrogenase activity(GO:0008480) |
| 8.4 | 66.9 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 8.2 | 24.6 | GO:0019150 | D-ribulokinase activity(GO:0019150) |
| 8.2 | 40.9 | GO:0043546 | xanthine dehydrogenase activity(GO:0004854) oxidoreductase activity, acting on CH or CH2 groups, NAD or NADP as acceptor(GO:0016726) molybdenum ion binding(GO:0030151) molybdopterin cofactor binding(GO:0043546) |
| 7.0 | 188.2 | GO:0070330 | aromatase activity(GO:0070330) |
| 6.7 | 40.0 | GO:0008453 | alanine-glyoxylate transaminase activity(GO:0008453) |
| 6.1 | 36.3 | GO:0015232 | heme transporter activity(GO:0015232) |
| 6.0 | 36.2 | GO:0060228 | phosphatidylcholine-sterol O-acyltransferase activator activity(GO:0060228) |
| 5.3 | 21.3 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 4.5 | 199.0 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 4.1 | 74.1 | GO:0019841 | retinol binding(GO:0019841) |
| 4.1 | 20.3 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 3.9 | 43.0 | GO:0016174 | NAD(P)H oxidase activity(GO:0016174) |
| 3.9 | 27.3 | GO:0004991 | parathyroid hormone receptor activity(GO:0004991) |
| 3.8 | 49.0 | GO:0004887 | thyroid hormone receptor activity(GO:0004887) |
| 3.5 | 28.1 | GO:0004416 | hydroxyacylglutathione hydrolase activity(GO:0004416) |
| 3.1 | 18.8 | GO:0016647 | oxidoreductase activity, acting on the CH-NH group of donors, oxygen as acceptor(GO:0016647) |
| 2.9 | 11.7 | GO:0004307 | ethanolaminephosphotransferase activity(GO:0004307) |
| 2.8 | 81.7 | GO:0016638 | oxidoreductase activity, acting on the CH-NH2 group of donors(GO:0016638) |
| 2.7 | 13.5 | GO:0004998 | transferrin receptor activity(GO:0004998) |
| 2.5 | 29.5 | GO:0015125 | bile acid transmembrane transporter activity(GO:0015125) |
| 2.5 | 12.3 | GO:0015137 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 2.5 | 14.7 | GO:0090554 | xenobiotic-transporting ATPase activity(GO:0008559) phosphatidylcholine-translocating ATPase activity(GO:0090554) |
| 2.4 | 33.2 | GO:0003847 | 1-alkyl-2-acetylglycerophosphocholine esterase activity(GO:0003847) |
| 2.2 | 6.5 | GO:0034012 | glycerone kinase activity(GO:0004371) FAD-AMP lyase (cyclizing) activity(GO:0034012) triokinase activity(GO:0050354) |
| 2.1 | 22.7 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 2.0 | 18.3 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 2.0 | 8.1 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 1.9 | 37.1 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 1.7 | 30.7 | GO:0008430 | selenium binding(GO:0008430) |
| 1.7 | 21.5 | GO:0048403 | brain-derived neurotrophic factor binding(GO:0048403) |
| 1.5 | 50.8 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 1.5 | 128.8 | GO:0005550 | pheromone binding(GO:0005550) |
| 1.4 | 25.9 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 1.4 | 5.5 | GO:0016649 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 1.3 | 5.1 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 1.2 | 4.8 | GO:0070506 | high-density lipoprotein particle receptor activity(GO:0070506) |
| 1.2 | 27.7 | GO:0015238 | drug transmembrane transporter activity(GO:0015238) |
| 1.0 | 17.6 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.8 | 33.7 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.8 | 10.6 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.8 | 4.1 | GO:0004131 | cytosine deaminase activity(GO:0004131) |
| 0.8 | 2.3 | GO:0045030 | A1 adenosine receptor binding(GO:0031686) UTP-activated nucleotide receptor activity(GO:0045030) |
| 0.7 | 151.4 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.7 | 61.3 | GO:0038024 | cargo receptor activity(GO:0038024) |
| 0.7 | 4.6 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.6 | 7.0 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.6 | 6.2 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.6 | 4.4 | GO:0047288 | monosialoganglioside sialyltransferase activity(GO:0047288) |
| 0.6 | 5.0 | GO:0005166 | neurotrophin p75 receptor binding(GO:0005166) |
| 0.6 | 6.0 | GO:0070095 | fructose-6-phosphate binding(GO:0070095) |
| 0.6 | 4.2 | GO:0070699 | type II activin receptor binding(GO:0070699) |
| 0.6 | 7.6 | GO:0004439 | phosphatidylinositol-4,5-bisphosphate 5-phosphatase activity(GO:0004439) |
| 0.6 | 11.6 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.6 | 24.9 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) |
| 0.5 | 8.7 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.4 | 17.3 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.4 | 6.7 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.4 | 0.8 | GO:0008240 | tripeptidyl-peptidase activity(GO:0008240) |
| 0.4 | 26.3 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.3 | 91.7 | GO:0008168 | methyltransferase activity(GO:0008168) |
| 0.3 | 2.4 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.3 | 2.3 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.3 | 5.5 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.3 | 6.2 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.3 | 18.3 | GO:0016706 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, 2-oxoglutarate as one donor, and incorporation of one atom each of oxygen into both donors(GO:0016706) |
| 0.3 | 1.8 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.2 | 3.4 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 0.2 | 27.3 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.2 | 2.5 | GO:0030976 | thiamine pyrophosphate binding(GO:0030976) |
| 0.2 | 7.4 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.2 | 17.1 | GO:0098531 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) transcription factor activity, direct ligand regulated sequence-specific DNA binding(GO:0098531) |
| 0.2 | 3.3 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.2 | 10.1 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.2 | 10.6 | GO:0009055 | electron carrier activity(GO:0009055) |
| 0.2 | 8.4 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.2 | 12.1 | GO:0019003 | GDP binding(GO:0019003) |
| 0.2 | 25.1 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.1 | 2.6 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.1 | 1.3 | GO:0022851 | GABA-gated chloride ion channel activity(GO:0022851) |
| 0.1 | 12.6 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.1 | 23.1 | GO:0002020 | protease binding(GO:0002020) |
| 0.1 | 1.9 | GO:0033549 | MAP kinase phosphatase activity(GO:0033549) |
| 0.1 | 1.7 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.1 | 5.1 | GO:0001221 | transcription cofactor binding(GO:0001221) |
| 0.1 | 2.3 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.1 | 2.3 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.1 | 1.6 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.1 | 9.5 | GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds(GO:0004553) |
| 0.1 | 1.1 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.1 | 2.4 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.1 | 2.1 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.1 | 7.1 | GO:0001104 | RNA polymerase II transcription cofactor activity(GO:0001104) |
| 0.0 | 14.4 | GO:0008022 | protein C-terminus binding(GO:0008022) |
| 0.0 | 4.3 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 0.0 | 0.4 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.2 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.0 | 0.7 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.0 | 1.3 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.0 | 2.0 | GO:0019208 | phosphatase regulator activity(GO:0019208) |
| 0.0 | 4.4 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 1.2 | GO:0003678 | DNA helicase activity(GO:0003678) |
| 0.0 | 1.2 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 0.1 | GO:0016312 | inositol bisphosphate phosphatase activity(GO:0016312) |
| 0.0 | 7.0 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 2.6 | GO:0044325 | ion channel binding(GO:0044325) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 65.3 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 1.0 | 35.0 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 1.0 | 79.8 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.7 | 16.1 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.4 | 12.5 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.3 | 39.4 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.2 | 15.4 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.2 | 4.0 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.2 | 24.5 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.1 | 2.4 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.1 | 5.1 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.1 | 5.0 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.1 | 1.6 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.1 | 6.7 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.1 | 4.0 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.1 | 6.7 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.1 | 0.8 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.0 | 12.7 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 2.6 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 2.3 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 0.7 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.9 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 2.2 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 12.4 | 173.0 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 11.0 | 142.7 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 8.2 | 163.8 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 6.2 | 80.6 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 6.0 | 84.4 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 5.2 | 67.0 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 4.4 | 74.1 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 4.1 | 40.9 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 3.1 | 43.0 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 2.8 | 11.4 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 2.3 | 35.1 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 1.9 | 31.1 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 1.9 | 15.2 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 1.9 | 54.6 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 1.7 | 40.0 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 1.6 | 19.0 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 1.3 | 22.7 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 1.2 | 24.9 | REACTOME VIRAL MESSENGER RNA SYNTHESIS | Genes involved in Viral Messenger RNA Synthesis |
| 0.8 | 48.4 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.8 | 41.7 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.8 | 182.5 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.7 | 14.2 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.7 | 20.3 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.7 | 7.4 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.6 | 66.1 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.5 | 10.7 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.5 | 17.2 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |
| 0.5 | 12.9 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.5 | 24.6 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.4 | 13.1 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.4 | 6.7 | REACTOME DESTABILIZATION OF MRNA BY BRF1 | Genes involved in Destabilization of mRNA by Butyrate Response Factor 1 (BRF1) |
| 0.3 | 43.7 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.3 | 13.0 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.2 | 18.4 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.2 | 28.0 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.2 | 3.5 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.1 | 0.9 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.1 | 2.3 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.1 | 5.5 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.1 | 2.3 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.1 | 4.2 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.1 | 1.6 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 8.4 | REACTOME BIOLOGICAL OXIDATIONS | Genes involved in Biological oxidations |
| 0.1 | 3.7 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.1 | 2.7 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 2.1 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 1.3 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 4.0 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.0 | 5.7 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |
| 0.0 | 2.4 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |