Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
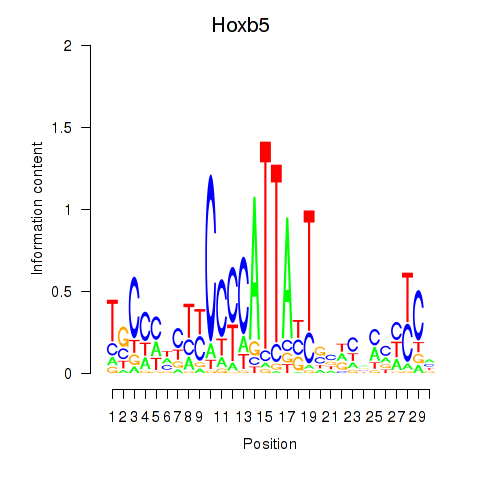
Results for Hoxb5
Z-value: 1.53
Transcription factors associated with Hoxb5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hoxb5
|
ENSMUSG00000038700.5 | Hoxb5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hoxb5 | mm39_v1_chr11_+_96194333_96194378 | 0.37 | 2.7e-02 | Click! |
Activity profile of Hoxb5 motif
Sorted Z-values of Hoxb5 motif
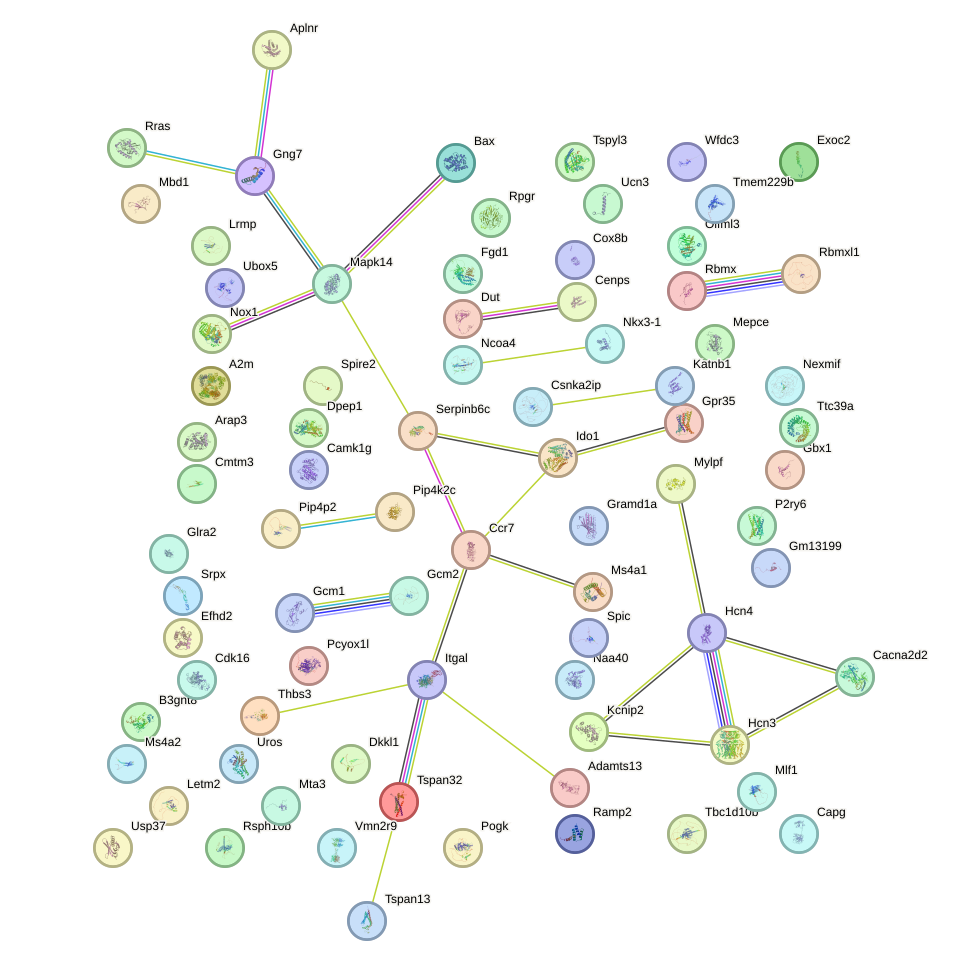
Network of associatons between targets according to the STRING database.
First level regulatory network of Hoxb5
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_+_121613177 | 16.26 |
ENSMUST00000032203.9
|
A2m
|
alpha-2-macroglobulin |
| chr7_+_142558837 | 8.33 |
ENSMUST00000207211.2
|
Tspan32
|
tetraspanin 32 |
| chr7_+_142559414 | 6.08 |
ENSMUST00000082008.12
ENSMUST00000105925.8 ENSMUST00000105924.8 |
Tspan32
|
tetraspanin 32 |
| chr17_+_47904355 | 5.80 |
ENSMUST00000182209.8
|
Ccnd3
|
cyclin D3 |
| chr17_+_47904441 | 5.70 |
ENSMUST00000182874.3
|
Ccnd3
|
cyclin D3 |
| chr2_+_125089110 | 5.48 |
ENSMUST00000082122.14
|
Dut
|
deoxyuridine triphosphatase |
| chr7_+_142559375 | 5.43 |
ENSMUST00000075172.12
ENSMUST00000105923.8 |
Tspan32
|
tetraspanin 32 |
| chr6_+_145067457 | 5.27 |
ENSMUST00000032396.13
|
Lrmp
|
lymphoid-restricted membrane protein |
| chr7_+_142558783 | 5.03 |
ENSMUST00000009396.13
|
Tspan32
|
tetraspanin 32 |
| chr12_-_36092475 | 4.84 |
ENSMUST00000020896.17
|
Tspan13
|
tetraspanin 13 |
| chr6_+_72526236 | 3.95 |
ENSMUST00000114071.8
|
Capg
|
capping protein (actin filament), gelsolin-like |
| chr7_+_126808016 | 3.72 |
ENSMUST00000206204.2
ENSMUST00000206772.2 |
Mylpf
|
myosin light chain, phosphorylatable, fast skeletal muscle |
| chr7_+_142559475 | 3.52 |
ENSMUST00000143512.3
|
Tspan32
|
tetraspanin 32 |
| chr10_+_128745214 | 3.13 |
ENSMUST00000220308.2
|
Cd63
|
CD63 antigen |
| chr10_-_88518878 | 2.93 |
ENSMUST00000004473.15
|
Spic
|
Spi-C transcription factor (Spi-1/PU.1 related) |
| chr14_+_31881822 | 2.92 |
ENSMUST00000163336.8
ENSMUST00000169722.8 ENSMUST00000168385.8 |
Ncoa4
|
nuclear receptor coactivator 4 |
| chr7_-_44181477 | 2.82 |
ENSMUST00000098483.9
ENSMUST00000035323.6 |
Spib
|
Spi-B transcription factor (Spi-1/PU.1 related) |
| chr4_-_149221998 | 2.81 |
ENSMUST00000176124.8
ENSMUST00000177408.2 ENSMUST00000105695.2 |
Cenps
|
centromere protein S |
| chr7_+_25327028 | 2.73 |
ENSMUST00000076034.8
|
B3gnt8
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 8 |
| chr12_-_113542610 | 2.68 |
ENSMUST00000195468.6
ENSMUST00000103442.3 |
Ighv5-2
|
immunoglobulin heavy variable 5-2 |
| chr7_-_133310779 | 2.68 |
ENSMUST00000124759.2
ENSMUST00000106144.8 ENSMUST00000106145.10 |
Uros
|
uroporphyrinogen III synthase |
| chr17_+_84013575 | 2.56 |
ENSMUST00000112350.8
ENSMUST00000112349.9 ENSMUST00000112352.10 ENSMUST00000067826.15 |
Mta3
|
metastasis associated 3 |
| chr8_+_13389656 | 2.55 |
ENSMUST00000210165.2
ENSMUST00000170909.2 |
Tfdp1
|
transcription factor Dp 1 |
| chr7_-_133310687 | 2.53 |
ENSMUST00000106146.8
|
Uros
|
uroporphyrinogen III synthase |
| chr8_+_95807814 | 2.48 |
ENSMUST00000034239.9
|
Katnb1
|
katanin p80 (WD40-containing) subunit B 1 |
| chr6_+_72526414 | 2.45 |
ENSMUST00000155705.8
|
Capg
|
capping protein (actin filament), gelsolin-like |
| chr6_-_128339775 | 2.41 |
ENSMUST00000112152.8
ENSMUST00000057421.15 ENSMUST00000112151.2 |
Rhno1
|
RAD9-HUS1-RAD1 interacting nuclear orphan 1 |
| chr4_-_149222057 | 2.39 |
ENSMUST00000030813.10
|
Cenps
|
centromere protein S |
| chr10_-_127047396 | 2.34 |
ENSMUST00000013970.9
|
Pip4k2c
|
phosphatidylinositol-5-phosphate 4-kinase, type II, gamma |
| chr7_-_126807581 | 2.32 |
ENSMUST00000120705.3
|
Tbc1d10b
|
TBC1 domain family, member 10b |
| chrX_+_20554193 | 2.13 |
ENSMUST00000115364.8
|
Cdk16
|
cyclin-dependent kinase 16 |
| chr19_-_45804446 | 2.10 |
ENSMUST00000079431.10
ENSMUST00000026247.13 ENSMUST00000162528.9 |
Kcnip2
|
Kv channel-interacting protein 2 |
| chr7_-_30850429 | 2.06 |
ENSMUST00000085636.13
ENSMUST00000001280.14 |
Gramd1a
|
GRAM domain containing 1A |
| chr12_-_79030250 | 2.02 |
ENSMUST00000070174.14
|
Tmem229b
|
transmembrane protein 229B |
| chr4_-_141602190 | 2.01 |
ENSMUST00000036854.4
|
Efhd2
|
EF hand domain containing 2 |
| chr2_-_164585102 | 1.91 |
ENSMUST00000103096.10
|
Wfdc3
|
WAP four-disulfide core domain 3 |
| chrX_+_149829131 | 1.89 |
ENSMUST00000112685.8
|
Fgd1
|
FYVE, RhoGEF and PH domain containing 1 |
| chr8_+_124059414 | 1.88 |
ENSMUST00000010298.7
|
Spire2
|
spire type actin nucleation factor 2 |
| chr7_-_45116316 | 1.87 |
ENSMUST00000033093.10
|
Bax
|
BCL2-associated X protein |
| chr19_-_7218363 | 1.74 |
ENSMUST00000236769.2
|
Naa40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit |
| chr7_-_45116216 | 1.72 |
ENSMUST00000210392.2
ENSMUST00000211365.2 |
Bax
|
BCL2-associated X protein |
| chr7_-_44861235 | 1.70 |
ENSMUST00000210741.2
ENSMUST00000209466.2 |
Dkkl1
|
dickkopf-like 1 |
| chr12_-_79027531 | 1.65 |
ENSMUST00000174072.8
|
Tmem229b
|
transmembrane protein 229B |
| chr7_-_44180700 | 1.50 |
ENSMUST00000205506.2
|
Spib
|
Spi-B transcription factor (Spi-1/PU.1 related) |
| chr2_-_153067297 | 1.46 |
ENSMUST00000099194.4
|
Tspyl3
|
TSPY-like 3 |
| chr7_-_45116197 | 1.44 |
ENSMUST00000211195.2
ENSMUST00000210019.2 |
Bax
|
BCL2-associated X protein |
| chr4_+_109272828 | 1.43 |
ENSMUST00000106618.8
|
Ttc39a
|
tetratricopeptide repeat domain 39A |
| chr2_+_84966569 | 1.39 |
ENSMUST00000057019.9
|
Aplnr
|
apelin receptor |
| chr19_-_7218512 | 1.38 |
ENSMUST00000025675.11
|
Naa40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit |
| chr7_-_100613579 | 1.27 |
ENSMUST00000060174.6
|
P2ry6
|
pyrimidinergic receptor P2Y, G-protein coupled, 6 |
| chr7_+_44667377 | 1.20 |
ENSMUST00000044111.10
|
Rras
|
related RAS viral (r-ras) oncogene |
| chr5_-_137784912 | 1.13 |
ENSMUST00000031740.16
|
Mepce
|
methylphosphate capping enzyme |
| chr11_+_101137786 | 1.05 |
ENSMUST00000107282.4
|
Ramp2
|
receptor (calcitonin) activity modifying protein 2 |
| chr3_+_67281449 | 1.04 |
ENSMUST00000061322.10
|
Mlf1
|
myeloid leukemia factor 1 |
| chr5_-_137784943 | 1.02 |
ENSMUST00000132726.2
|
Mepce
|
methylphosphate capping enzyme |
| chr8_+_105067159 | 0.96 |
ENSMUST00000212948.2
ENSMUST00000034343.5 |
Cmtm3
|
CKLF-like MARVEL transmembrane domain containing 3 |
| chr1_+_92878566 | 0.95 |
ENSMUST00000185421.2
|
Gpr35
|
G protein-coupled receptor 35 |
| chr17_+_28911529 | 0.83 |
ENSMUST00000114752.3
|
Mapk14
|
mitogen-activated protein kinase 14 |
| chrX_-_56438380 | 0.80 |
ENSMUST00000143310.2
ENSMUST00000098470.9 ENSMUST00000114726.8 |
Rbmx
|
RNA binding motif protein, X chromosome |
| chr13_-_31158031 | 0.80 |
ENSMUST00000021785.8
|
Exoc2
|
exocyst complex component 2 |
| chr8_-_26087475 | 0.79 |
ENSMUST00000210810.2
ENSMUST00000210616.2 ENSMUST00000079160.8 |
Letm2
|
leucine zipper-EF-hand containing transmembrane protein 2 |
| chr2_+_26863428 | 0.70 |
ENSMUST00000014996.14
|
Adamts13
|
a disintegrin-like and metallopeptidase (reprolysin type) with thrombospondin type 1 motif, 13 |
| chr9_+_77959206 | 0.67 |
ENSMUST00000024104.9
|
Gcm1
|
glial cells missing homolog 1 |
| chr18_-_61840654 | 0.64 |
ENSMUST00000025472.7
|
Pcyox1l
|
prenylcysteine oxidase 1 like |
| chr4_+_14864076 | 0.63 |
ENSMUST00000029875.4
|
Pip4p2
|
phosphatidylinositol-4,5-bisphosphate 4-phosphatase 2 |
| chr18_+_74401322 | 0.58 |
ENSMUST00000224047.2
|
Mbd1
|
methyl-CpG binding domain protein 1 |
| chr3_+_67281424 | 0.55 |
ENSMUST00000077916.12
|
Mlf1
|
myeloid leukemia factor 1 |
| chr16_-_64422716 | 0.55 |
ENSMUST00000209382.3
|
Csnka2ip
|
casein kinase 2, alpha prime interacting protein |
| chr18_-_38131766 | 0.51 |
ENSMUST00000236588.2
ENSMUST00000237272.2 ENSMUST00000236134.2 |
Arap3
|
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 3 |
| chrX_-_56438322 | 0.48 |
ENSMUST00000114730.8
|
Rbmx
|
RNA binding motif protein, X chromosome |
| chr5_-_103359117 | 0.47 |
ENSMUST00000112846.8
ENSMUST00000170792.9 ENSMUST00000112847.9 ENSMUST00000238446.3 ENSMUST00000133069.8 |
Mapk10
|
mitogen-activated protein kinase 10 |
| chr3_-_103645311 | 0.44 |
ENSMUST00000029440.10
|
Olfml3
|
olfactomedin-like 3 |
| chr3_+_89122477 | 0.43 |
ENSMUST00000029682.11
|
Thbs3
|
thrombospondin 3 |
| chr8_+_105066924 | 0.41 |
ENSMUST00000212081.2
|
Cmtm3
|
CKLF-like MARVEL transmembrane domain containing 3 |
| chr7_+_126895423 | 0.39 |
ENSMUST00000117762.8
|
Itgal
|
integrin alpha L |
| chr7_+_126895463 | 0.35 |
ENSMUST00000106306.9
ENSMUST00000120857.8 |
Itgal
|
integrin alpha L |
| chr1_-_74583434 | 0.34 |
ENSMUST00000189257.7
|
Usp37
|
ubiquitin specific peptidase 37 |
| chr19_-_11601007 | 0.31 |
ENSMUST00000164792.8
ENSMUST00000189641.2 ENSMUST00000186978.7 ENSMUST00000025583.12 |
Ms4a2
|
membrane-spanning 4-domains, subfamily A, member 2 |
| chr11_+_101137231 | 0.30 |
ENSMUST00000122006.8
ENSMUST00000151830.2 |
Ramp2
|
receptor (calcitonin) activity modifying protein 2 |
| chr7_+_126895531 | 0.24 |
ENSMUST00000170971.8
|
Itgal
|
integrin alpha L |
| chr1_-_193052568 | 0.23 |
ENSMUST00000016323.11
|
Camk1g
|
calcium/calmodulin-dependent protein kinase I gamma |
| chr14_-_40726472 | 0.23 |
ENSMUST00000153830.8
|
Prxl2a
|
peroxiredoxin like 2A |
| chr3_-_89067462 | 0.22 |
ENSMUST00000029686.4
|
Hcn3
|
hyperpolarization-activated, cyclic nucleotide-gated K+ 3 |
| chrX_-_9983836 | 0.21 |
ENSMUST00000115543.3
ENSMUST00000044789.10 ENSMUST00000115544.9 |
Srpx
|
sushi-repeat-containing protein |
| chr2_-_130471891 | 0.20 |
ENSMUST00000110262.3
ENSMUST00000028761.5 |
Fastkd5
Ubox5
|
FAST kinase domains 5 U box domain containing 5 |
| chr7_-_140480314 | 0.17 |
ENSMUST00000026561.10
|
Cox8b
|
cytochrome c oxidase subunit 8B |
| chrX_-_103244703 | 0.17 |
ENSMUST00000087879.11
|
Nexmif
|
neurite extension and migration factor |
| chr2_-_5867734 | 0.15 |
ENSMUST00000071016.3
|
Gm13199
|
predicted gene 13199 |
| chrX_-_133012457 | 0.14 |
ENSMUST00000159259.3
ENSMUST00000113275.10 |
Nox1
|
NADPH oxidase 1 |
| chr5_-_24732200 | 0.13 |
ENSMUST00000088311.6
|
Gbx1
|
gastrulation brain homeobox 1 |
| chrX_-_133012600 | 0.13 |
ENSMUST00000033610.13
|
Nox1
|
NADPH oxidase 1 |
| chr17_-_80885197 | 0.12 |
ENSMUST00000234602.2
|
Cdkl4
|
cyclin-dependent kinase-like 4 |
| chr8_-_25086976 | 0.09 |
ENSMUST00000033956.7
|
Ido1
|
indoleamine 2,3-dioxygenase 1 |
| chr8_-_123768984 | 0.08 |
ENSMUST00000212937.2
|
Ankrd11
|
ankyrin repeat domain 11 |
| chrX_-_103244784 | 0.08 |
ENSMUST00000118314.8
|
Nexmif
|
neurite extension and migration factor |
| chr8_-_26087552 | 0.07 |
ENSMUST00000210234.2
ENSMUST00000211422.2 |
Letm2
|
leucine zipper-EF-hand containing transmembrane protein 2 |
| chr9_+_107277058 | 0.07 |
ENSMUST00000085092.12
ENSMUST00000164988.9 |
Cacna2d2
|
calcium channel, voltage-dependent, alpha 2/delta subunit 2 |
| chr9_+_107277242 | 0.07 |
ENSMUST00000166799.8
ENSMUST00000170737.3 |
Cacna2d2
|
calcium channel, voltage-dependent, alpha 2/delta subunit 2 |
| chr1_-_166237341 | 0.05 |
ENSMUST00000135673.8
ENSMUST00000169324.8 ENSMUST00000128861.3 |
Pogk
|
pogo transposable element with KRAB domain |
| chr13_-_3995349 | 0.05 |
ENSMUST00000058610.8
|
Ucn3
|
urocortin 3 |
| chr15_-_34358018 | 0.05 |
ENSMUST00000227772.2
|
9430069I07Rik
|
RIKEN cDNA 9430069I07 gene |
| chr1_-_193052533 | 0.05 |
ENSMUST00000169907.8
|
Camk1g
|
calcium/calmodulin-dependent protein kinase I gamma |
| chr10_-_80850712 | 0.03 |
ENSMUST00000126317.2
ENSMUST00000092285.10 ENSMUST00000117805.8 |
Gng7
|
guanine nucleotide binding protein (G protein), gamma 7 |
| chr7_-_103674780 | 0.03 |
ENSMUST00000218535.2
|
Olfr640
|
olfactory receptor 640 |
| chr11_-_99045894 | 0.02 |
ENSMUST00000103134.4
|
Ccr7
|
chemokine (C-C motif) receptor 7 |
| chr5_+_143869875 | 0.02 |
ENSMUST00000166847.8
|
Rsph10b
|
radial spoke head 10 homolog B (Chlamydomonas) |
| chr8_+_105066980 | 0.02 |
ENSMUST00000211885.2
|
Cmtm3
|
CKLF-like MARVEL transmembrane domain containing 3 |
| chr9_+_8900460 | 0.02 |
ENSMUST00000070463.10
ENSMUST00000098986.4 |
Pgr
|
progesterone receptor |
| chr5_-_109000467 | 0.01 |
ENSMUST00000170419.3
ENSMUST00000233835.2 |
Vmn2r9
|
vomeronasal 2, receptor 9 |
| chrX_-_164110372 | 0.00 |
ENSMUST00000058787.9
|
Glra2
|
glycine receptor, alpha 2 subunit |
| chr2_-_151474391 | 0.00 |
ENSMUST00000137936.2
ENSMUST00000146172.8 ENSMUST00000094456.10 ENSMUST00000148755.8 ENSMUST00000109875.8 ENSMUST00000028951.14 ENSMUST00000109877.10 |
Snph
|
syntaphilin |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.4 | 16.3 | GO:0001868 | regulation of complement activation, lectin pathway(GO:0001868) negative regulation of complement activation, lectin pathway(GO:0001869) |
| 2.6 | 28.4 | GO:0030886 | negative regulation of myeloid dendritic cell activation(GO:0030886) |
| 1.7 | 5.2 | GO:0006780 | uroporphyrinogen III biosynthetic process(GO:0006780) |
| 1.7 | 5.0 | GO:1990117 | release of matrix enzymes from mitochondria(GO:0032976) B cell receptor apoptotic signaling pathway(GO:1990117) |
| 1.4 | 5.5 | GO:0009223 | dUMP biosynthetic process(GO:0006226) pyrimidine nucleotide catabolic process(GO:0006244) pyrimidine deoxyribonucleotide catabolic process(GO:0009223) |
| 0.7 | 2.7 | GO:0030309 | poly-N-acetyllactosamine metabolic process(GO:0030309) poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.6 | 2.5 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 0.5 | 3.1 | GO:2000680 | regulation of rubidium ion transport(GO:2000680) |
| 0.5 | 1.9 | GO:0070649 | formin-nucleated actin cable assembly(GO:0070649) |
| 0.4 | 11.5 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.3 | 2.1 | GO:0045163 | clustering of voltage-gated potassium channels(GO:0045163) |
| 0.3 | 1.4 | GO:0002030 | inhibitory G-protein coupled receptor phosphorylation(GO:0002030) |
| 0.3 | 1.0 | GO:1904456 | negative regulation of neuronal action potential(GO:1904456) |
| 0.3 | 5.2 | GO:0000712 | resolution of meiotic recombination intermediates(GO:0000712) |
| 0.2 | 2.1 | GO:0040031 | snRNA modification(GO:0040031) |
| 0.2 | 2.5 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.2 | 0.6 | GO:0030327 | prenylated protein catabolic process(GO:0030327) |
| 0.2 | 2.3 | GO:2000786 | positive regulation of autophagosome assembly(GO:2000786) |
| 0.2 | 1.4 | GO:0097646 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.2 | 0.7 | GO:0060800 | regulation of cell differentiation involved in embryonic placenta development(GO:0060800) |
| 0.2 | 6.4 | GO:0051016 | barbed-end actin filament capping(GO:0051016) |
| 0.1 | 1.3 | GO:0030321 | transepithelial chloride transport(GO:0030321) |
| 0.1 | 1.0 | GO:0002291 | T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:0002291) |
| 0.1 | 2.9 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.1 | 0.8 | GO:0001927 | exocyst assembly(GO:0001927) |
| 0.1 | 1.4 | GO:0050861 | positive regulation of B cell receptor signaling pathway(GO:0050861) |
| 0.1 | 0.8 | GO:2001184 | positive regulation of interleukin-12 secretion(GO:2001184) |
| 0.1 | 3.1 | GO:0006474 | N-terminal protein amino acid acetylation(GO:0006474) histone H2A acetylation(GO:0043968) |
| 0.1 | 1.6 | GO:0002318 | myeloid progenitor cell differentiation(GO:0002318) |
| 0.1 | 4.3 | GO:0030225 | macrophage differentiation(GO:0030225) |
| 0.1 | 2.6 | GO:0010971 | positive regulation of G2/M transition of mitotic cell cycle(GO:0010971) |
| 0.1 | 0.5 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.1 | 0.3 | GO:0045726 | positive regulation of integrin biosynthetic process(GO:0045726) |
| 0.1 | 0.5 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.1 | 2.1 | GO:0030252 | growth hormone secretion(GO:0030252) |
| 0.0 | 0.3 | GO:0038095 | Fc-epsilon receptor signaling pathway(GO:0038095) |
| 0.0 | 0.1 | GO:0097374 | proprioception(GO:0019230) sensory neuron axon guidance(GO:0097374) |
| 0.0 | 0.4 | GO:0060346 | bone trabecula formation(GO:0060346) |
| 0.0 | 0.2 | GO:0060244 | negative regulation of cell proliferation involved in contact inhibition(GO:0060244) |
| 0.0 | 0.1 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 5.3 | GO:0007338 | single fertilization(GO:0007338) |
| 0.0 | 0.6 | GO:0048712 | negative regulation of astrocyte differentiation(GO:0048712) |
| 0.0 | 1.3 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.0 | 2.3 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.0 | 2.4 | GO:0071479 | cellular response to ionizing radiation(GO:0071479) |
| 0.0 | 0.1 | GO:0060024 | rhythmic synaptic transmission(GO:0060024) |
| 0.0 | 0.3 | GO:0035871 | protein K11-linked deubiquitination(GO:0035871) |
| 0.0 | 1.7 | GO:0045600 | positive regulation of fat cell differentiation(GO:0045600) |
| 0.0 | 2.9 | GO:0001824 | blastocyst development(GO:0001824) |
| 0.0 | 2.7 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 0.2 | GO:0000963 | mitochondrial RNA processing(GO:0000963) |
| 0.0 | 3.8 | GO:1903169 | regulation of calcium ion transmembrane transport(GO:1903169) |
| 0.0 | 1.9 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.0 | 0.7 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.0 | 0.2 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.7 | 28.4 | GO:0070442 | integrin alphaIIb-beta3 complex(GO:0070442) |
| 1.7 | 5.0 | GO:0097144 | BAX complex(GO:0097144) |
| 1.3 | 5.2 | GO:0071821 | FANCM-MHF complex(GO:0071821) |
| 1.0 | 3.1 | GO:0031904 | endosome lumen(GO:0031904) |
| 0.4 | 6.4 | GO:0090543 | Flemming body(GO:0090543) |
| 0.2 | 1.4 | GO:1903440 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.2 | 1.3 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.2 | 1.0 | GO:0034687 | integrin alphaL-beta2 complex(GO:0034687) |
| 0.2 | 2.9 | GO:0044754 | autolysosome(GO:0044754) |
| 0.1 | 10.4 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.1 | 0.3 | GO:0032997 | Fc receptor complex(GO:0032997) Fc-epsilon receptor I complex(GO:0032998) |
| 0.0 | 14.1 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.0 | 0.2 | GO:0044316 | cone cell pedicle(GO:0044316) |
| 0.0 | 1.6 | GO:0097546 | ciliary base(GO:0097546) |
| 0.0 | 0.3 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.0 | 0.8 | GO:0000145 | exocyst(GO:0000145) |
| 0.0 | 1.9 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 3.7 | GO:0016459 | myosin complex(GO:0016459) |
| 0.0 | 2.6 | GO:0005776 | autophagosome(GO:0005776) |
| 0.0 | 3.3 | GO:0000922 | spindle pole(GO:0000922) |
| 0.0 | 2.1 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.0 | 2.5 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 0.1 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 0.0 | 2.4 | GO:0001726 | ruffle(GO:0001726) |
| 0.0 | 2.2 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.4 | 16.3 | GO:0019959 | interleukin-8 binding(GO:0019959) |
| 0.9 | 2.7 | GO:0016262 | protein N-acetylglucosaminyltransferase activity(GO:0016262) |
| 0.9 | 5.2 | GO:0004852 | uroporphyrinogen-III synthase activity(GO:0004852) |
| 0.8 | 2.3 | GO:0016309 | 1-phosphatidylinositol-5-phosphate 4-kinase activity(GO:0016309) |
| 0.5 | 5.0 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.5 | 5.5 | GO:0047429 | nucleoside-triphosphate diphosphatase activity(GO:0047429) |
| 0.4 | 2.1 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.4 | 2.5 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.3 | 3.1 | GO:1990189 | peptide-serine-N-acetyltransferase activity(GO:1990189) |
| 0.3 | 1.3 | GO:0015065 | uridine nucleotide receptor activity(GO:0015065) G-protein coupled pyrimidinergic nucleotide receptor activity(GO:0071553) |
| 0.2 | 13.7 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.2 | 1.4 | GO:0097643 | amylin receptor activity(GO:0097643) |
| 0.2 | 1.0 | GO:0030369 | ICAM-3 receptor activity(GO:0030369) |
| 0.1 | 1.0 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.1 | 0.5 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.1 | 0.6 | GO:0010385 | double-stranded methylated DNA binding(GO:0010385) |
| 0.1 | 4.4 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.1 | 0.6 | GO:0016670 | oxidoreductase activity, acting on a sulfur group of donors, oxygen as acceptor(GO:0016670) |
| 0.1 | 0.8 | GO:0051525 | mitogen-activated protein kinase p38 binding(GO:0048273) NFAT protein binding(GO:0051525) |
| 0.1 | 0.3 | GO:0019767 | IgE receptor activity(GO:0019767) |
| 0.1 | 4.3 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.1 | 3.7 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.3 | GO:0030171 | voltage-gated proton channel activity(GO:0030171) |
| 0.0 | 0.2 | GO:0005222 | intracellular cAMP activated cation channel activity(GO:0005222) |
| 0.0 | 0.1 | GO:0033754 | indoleamine 2,3-dioxygenase activity(GO:0033754) |
| 0.0 | 0.8 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.0 | 2.1 | GO:0008173 | RNA methyltransferase activity(GO:0008173) |
| 0.0 | 2.7 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 4.5 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.0 | GO:0051431 | corticotropin-releasing hormone receptor 2 binding(GO:0051431) |
| 0.0 | 1.9 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.5 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.0 | 0.9 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 1.3 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 11.5 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.2 | 5.0 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.2 | 17.1 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.0 | 1.2 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 2.5 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 1.9 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 2.9 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.0 | 1.0 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.0 | 3.7 | PID ERBB1 DOWNSTREAM PATHWAY | ErbB1 downstream signaling |
| 0.0 | 0.5 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.0 | 0.8 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 16.3 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.2 | 14.0 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.2 | 5.2 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.1 | 5.5 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.1 | 5.0 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.1 | 1.3 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.1 | 1.3 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.1 | 1.2 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 2.7 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.8 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 3.1 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 1.0 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 1.4 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |