Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
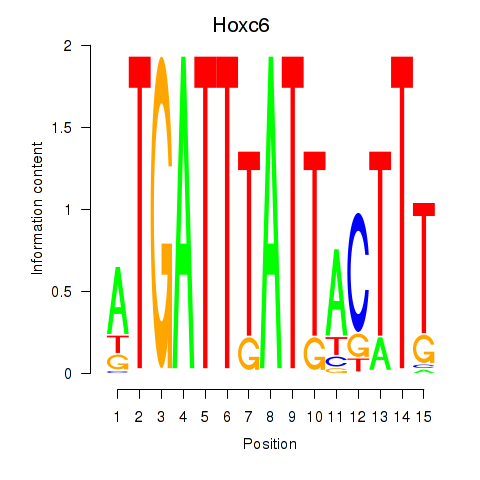
Results for Hoxc6
Z-value: 1.03
Transcription factors associated with Hoxc6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hoxc6
|
ENSMUSG00000001661.6 | Hoxc6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hoxc6 | mm39_v1_chr15_+_102917977_102918011 | -0.29 | 9.1e-02 | Click! |
Activity profile of Hoxc6 motif
Sorted Z-values of Hoxc6 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Hoxc6
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_39275518 | 9.45 |
ENSMUST00000003137.15
|
Cyp2c29
|
cytochrome P450, family 2, subfamily c, polypeptide 29 |
| chr7_+_51537645 | 2.39 |
ENSMUST00000208711.2
|
Gas2
|
growth arrest specific 2 |
| chr18_-_56705960 | 2.13 |
ENSMUST00000174518.8
|
Aldh7a1
|
aldehyde dehydrogenase family 7, member A1 |
| chr19_+_32597379 | 2.07 |
ENSMUST00000236290.2
ENSMUST00000025833.7 |
Papss2
|
3'-phosphoadenosine 5'-phosphosulfate synthase 2 |
| chr1_+_88062508 | 1.93 |
ENSMUST00000113134.8
ENSMUST00000140092.8 |
Ugt1a6a
|
UDP glucuronosyltransferase 1 family, polypeptide A6A |
| chr3_+_151143524 | 1.86 |
ENSMUST00000046977.12
|
Adgrl4
|
adhesion G protein-coupled receptor L4 |
| chr19_+_26725589 | 1.72 |
ENSMUST00000207812.2
ENSMUST00000175791.9 ENSMUST00000207118.2 ENSMUST00000209085.2 ENSMUST00000112637.10 ENSMUST00000207054.2 ENSMUST00000208589.2 ENSMUST00000176475.9 ENSMUST00000176698.9 ENSMUST00000207832.2 ENSMUST00000177252.9 ENSMUST00000208712.2 ENSMUST00000208186.2 ENSMUST00000208806.2 ENSMUST00000208027.2 |
Smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr1_+_88030951 | 1.72 |
ENSMUST00000113135.6
ENSMUST00000113138.8 |
Ugt1a7c
Ugt1a6b
|
UDP glucuronosyltransferase 1 family, polypeptide A7C UDP glucuronosyltransferase 1 family, polypeptide A6B |
| chr14_-_64654397 | 1.61 |
ENSMUST00000210428.2
|
Msra
|
methionine sulfoxide reductase A |
| chrM_-_14061 | 1.28 |
ENSMUST00000082419.1
|
mt-Nd6
|
mitochondrially encoded NADH dehydrogenase 6 |
| chrM_+_14138 | 1.26 |
ENSMUST00000082421.1
|
mt-Cytb
|
mitochondrially encoded cytochrome b |
| chr19_+_24853039 | 1.23 |
ENSMUST00000073080.7
|
Gm10053
|
predicted gene 10053 |
| chr14_-_64654592 | 1.11 |
ENSMUST00000210363.2
|
Msra
|
methionine sulfoxide reductase A |
| chr15_+_44482667 | 1.02 |
ENSMUST00000228648.2
ENSMUST00000226165.2 |
Ebag9
|
estrogen receptor-binding fragment-associated gene 9 |
| chrX_-_93256291 | 1.01 |
ENSMUST00000050328.15
|
Eif2s3x
|
eukaryotic translation initiation factor 2, subunit 3, structural gene X-linked |
| chrM_+_9459 | 1.00 |
ENSMUST00000082411.1
|
mt-Nd3
|
mitochondrially encoded NADH dehydrogenase 3 |
| chr15_+_44482944 | 0.94 |
ENSMUST00000022964.9
|
Ebag9
|
estrogen receptor-binding fragment-associated gene 9 |
| chr3_+_151143557 | 0.84 |
ENSMUST00000196970.3
|
Adgrl4
|
adhesion G protein-coupled receptor L4 |
| chr19_+_39499288 | 0.82 |
ENSMUST00000025968.5
|
Cyp2c39
|
cytochrome P450, family 2, subfamily c, polypeptide 39 |
| chr7_-_13856967 | 0.77 |
ENSMUST00000098809.4
|
Sult2a3
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 3 |
| chr14_+_79086492 | 0.74 |
ENSMUST00000040990.7
|
Vwa8
|
von Willebrand factor A domain containing 8 |
| chr1_+_171052623 | 0.71 |
ENSMUST00000111321.8
ENSMUST00000005824.12 ENSMUST00000111320.8 ENSMUST00000111319.2 |
Apoa2
|
apolipoprotein A-II |
| chr3_+_66893031 | 0.70 |
ENSMUST00000046542.13
ENSMUST00000162693.8 |
Rsrc1
|
arginine/serine-rich coiled-coil 1 |
| chr18_+_56705894 | 0.60 |
ENSMUST00000008445.7
|
Phax
|
phosphorylated adaptor for RNA export |
| chr2_-_121211410 | 0.58 |
ENSMUST00000038389.15
|
Strc
|
stereocilin |
| chr1_+_46464625 | 0.56 |
ENSMUST00000189749.7
|
Dnah7c
|
dynein, axonemal, heavy chain 7C |
| chr8_-_23196523 | 0.55 |
ENSMUST00000033939.13
ENSMUST00000063401.10 |
Ikbkb
|
inhibitor of kappaB kinase beta |
| chr11_-_23845207 | 0.54 |
ENSMUST00000102863.3
ENSMUST00000020513.10 |
Papolg
|
poly(A) polymerase gamma |
| chr9_+_40092216 | 0.51 |
ENSMUST00000218134.2
ENSMUST00000216720.2 ENSMUST00000214763.2 |
Olfr986
|
olfactory receptor 986 |
| chr3_+_66892979 | 0.50 |
ENSMUST00000162362.8
ENSMUST00000065074.14 ENSMUST00000065047.13 |
Rsrc1
|
arginine/serine-rich coiled-coil 1 |
| chr5_+_24618380 | 0.48 |
ENSMUST00000049346.10
|
Asic3
|
acid-sensing (proton-gated) ion channel 3 |
| chrX_-_93256244 | 0.46 |
ENSMUST00000113922.2
|
Eif2s3x
|
eukaryotic translation initiation factor 2, subunit 3, structural gene X-linked |
| chr14_+_75479727 | 0.46 |
ENSMUST00000022576.10
|
Cpb2
|
carboxypeptidase B2 (plasma) |
| chrX_-_104918911 | 0.41 |
ENSMUST00000200471.2
|
Atrx
|
ATRX, chromatin remodeler |
| chr13_+_93440572 | 0.41 |
ENSMUST00000109493.9
|
Homer1
|
homer scaffolding protein 1 |
| chr2_-_51039112 | 0.40 |
ENSMUST00000154545.2
ENSMUST00000017288.9 |
Rnd3
|
Rho family GTPase 3 |
| chrM_+_8603 | 0.40 |
ENSMUST00000082409.1
|
mt-Co3
|
mitochondrially encoded cytochrome c oxidase III |
| chr11_-_94673526 | 0.39 |
ENSMUST00000100554.8
|
Tmem92
|
transmembrane protein 92 |
| chr8_+_46111778 | 0.37 |
ENSMUST00000143820.8
|
Sorbs2
|
sorbin and SH3 domain containing 2 |
| chr10_-_112764879 | 0.37 |
ENSMUST00000099276.4
|
Atxn7l3b
|
ataxin 7-like 3B |
| chr15_-_50752437 | 0.36 |
ENSMUST00000183997.8
ENSMUST00000183757.8 |
Trps1
|
transcriptional repressor GATA binding 1 |
| chrX_-_135642025 | 0.35 |
ENSMUST00000155207.8
ENSMUST00000080411.13 ENSMUST00000169418.8 |
Morf4l2
|
mortality factor 4 like 2 |
| chr10_+_61010983 | 0.34 |
ENSMUST00000143207.8
ENSMUST00000148181.8 ENSMUST00000151886.8 |
Tbata
|
thymus, brain and testes associated |
| chr6_+_11926757 | 0.33 |
ENSMUST00000133776.2
|
Phf14
|
PHD finger protein 14 |
| chr1_-_85239791 | 0.31 |
ENSMUST00000162421.2
|
C130026I21Rik
|
RIKEN cDNA C130026I21 gene |
| chr3_-_144738526 | 0.31 |
ENSMUST00000029919.7
|
Clca1
|
chloride channel accessory 1 |
| chr17_+_3532455 | 0.31 |
ENSMUST00000227604.2
|
Tiam2
|
T cell lymphoma invasion and metastasis 2 |
| chr7_+_45204350 | 0.30 |
ENSMUST00000210300.2
|
Hsd17b14
|
hydroxysteroid (17-beta) dehydrogenase 14 |
| chr13_+_93441307 | 0.29 |
ENSMUST00000080127.12
|
Homer1
|
homer scaffolding protein 1 |
| chr5_-_129030367 | 0.28 |
ENSMUST00000111346.6
ENSMUST00000200470.5 |
Rimbp2
|
RIMS binding protein 2 |
| chr1_-_53745920 | 0.27 |
ENSMUST00000094964.7
|
Dnah7a
|
dynein, axonemal, heavy chain 7A |
| chrX_-_142716085 | 0.26 |
ENSMUST00000087313.10
|
Dcx
|
doublecortin |
| chr11_-_70560110 | 0.24 |
ENSMUST00000129434.2
ENSMUST00000018431.13 |
Spag7
|
sperm associated antigen 7 |
| chr1_-_22386016 | 0.23 |
ENSMUST00000164877.8
|
Rims1
|
regulating synaptic membrane exocytosis 1 |
| chr10_+_23814470 | 0.22 |
ENSMUST00000079134.5
|
Taar2
|
trace amine-associated receptor 2 |
| chr6_-_30693675 | 0.22 |
ENSMUST00000169422.8
ENSMUST00000115131.8 ENSMUST00000115130.3 ENSMUST00000031810.15 |
Cep41
|
centrosomal protein 41 |
| chr3_-_129518723 | 0.22 |
ENSMUST00000199615.5
ENSMUST00000197079.5 |
Egf
|
epidermal growth factor |
| chr17_-_29162794 | 0.21 |
ENSMUST00000232977.2
|
Pxt1
|
peroxisomal, testis specific 1 |
| chr6_-_123830422 | 0.20 |
ENSMUST00000162046.3
|
Vmn2r25
|
vomeronasal 2, receptor 25 |
| chr5_-_129030111 | 0.20 |
ENSMUST00000196085.5
|
Rimbp2
|
RIMS binding protein 2 |
| chr9_-_19088608 | 0.20 |
ENSMUST00000212127.3
|
Olfr839-ps1
|
olfactory receptor 839, pseudogene 1 |
| chr11_+_58556387 | 0.20 |
ENSMUST00000072030.4
|
Olfr322
|
olfactory receptor 322 |
| chrX_-_142716200 | 0.19 |
ENSMUST00000112851.8
ENSMUST00000112856.3 ENSMUST00000033642.10 |
Dcx
|
doublecortin |
| chr10_-_100425067 | 0.18 |
ENSMUST00000218821.2
ENSMUST00000054471.10 |
4930430F08Rik
|
RIKEN cDNA 4930430F08 gene |
| chr5_-_110194352 | 0.18 |
ENSMUST00000167969.2
|
Gm17655
|
predicted gene, 17655 |
| chr9_+_65268304 | 0.18 |
ENSMUST00000147185.3
|
Ubap1l
|
ubiquitin-associated protein 1-like |
| chr17_+_13094921 | 0.16 |
ENSMUST00000075296.4
|
Mrgprh
|
MAS-related GPR, member H |
| chr14_-_52616625 | 0.15 |
ENSMUST00000214980.2
|
Olfr1512
|
olfactory receptor 1512 |
| chr7_-_85974838 | 0.15 |
ENSMUST00000214977.2
|
Olfr308
|
olfactory receptor 308 |
| chr14_-_50586329 | 0.15 |
ENSMUST00000216634.2
|
Olfr735
|
olfactory receptor 735 |
| chrX_-_135641869 | 0.14 |
ENSMUST00000166930.8
ENSMUST00000113095.8 |
Morf4l2
|
mortality factor 4 like 2 |
| chr10_-_108535970 | 0.14 |
ENSMUST00000218023.2
|
Gm5136
|
predicted gene 5136 |
| chr8_-_55177510 | 0.13 |
ENSMUST00000175915.8
|
Wdr17
|
WD repeat domain 17 |
| chr10_-_70491764 | 0.13 |
ENSMUST00000162144.2
ENSMUST00000162793.8 |
Phyhipl
|
phytanoyl-CoA hydroxylase interacting protein-like |
| chr4_-_84464521 | 0.12 |
ENSMUST00000177040.2
|
Bnc2
|
basonuclin 2 |
| chr17_+_38082190 | 0.11 |
ENSMUST00000217119.2
|
Olfr122
|
olfactory receptor 122 |
| chr10_+_19588318 | 0.11 |
ENSMUST00000020185.5
|
Il20ra
|
interleukin 20 receptor, alpha |
| chr6_-_53955647 | 0.11 |
ENSMUST00000204674.3
ENSMUST00000166545.2 ENSMUST00000203101.3 |
Cpvl
|
carboxypeptidase, vitellogenic-like |
| chr7_-_16019379 | 0.11 |
ENSMUST00000174270.8
|
Ccdc9
|
coiled-coil domain containing 9 |
| chrX_+_111150171 | 0.10 |
ENSMUST00000164272.3
ENSMUST00000132037.2 |
4933403O08Rik
|
RIKEN cDNA 4933403O08 gene |
| chr7_+_108064533 | 0.09 |
ENSMUST00000217616.3
|
Olfr498
|
olfactory receptor 498 |
| chr3_+_5283606 | 0.09 |
ENSMUST00000026284.13
|
Zfhx4
|
zinc finger homeodomain 4 |
| chrX_-_104919201 | 0.05 |
ENSMUST00000198209.2
|
Atrx
|
ATRX, chromatin remodeler |
| chr5_+_17779273 | 0.05 |
ENSMUST00000030568.14
|
Sema3c
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3C |
| chr4_+_147445744 | 0.05 |
ENSMUST00000133078.8
ENSMUST00000154154.2 |
Zfp978
|
zinc finger protein 978 |
| chr6_+_36364990 | 0.05 |
ENSMUST00000172278.8
|
Chrm2
|
cholinergic receptor, muscarinic 2, cardiac |
| chr9_-_39161290 | 0.04 |
ENSMUST00000213472.2
|
Olfr1537
|
olfactory receptor 1537 |
| chr19_+_12972378 | 0.04 |
ENSMUST00000207997.3
|
Olfr1451
|
olfactory receptor 1451 |
| chr8_-_10026292 | 0.01 |
ENSMUST00000095476.6
|
Lig4
|
ligase IV, DNA, ATP-dependent |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.7 | GO:0006711 | estrogen catabolic process(GO:0006711) |
| 0.5 | 2.1 | GO:0034036 | purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.3 | 10.3 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.2 | 1.7 | GO:0035887 | aortic smooth muscle cell differentiation(GO:0035887) |
| 0.2 | 0.6 | GO:0051030 | snRNA transport(GO:0051030) |
| 0.2 | 1.9 | GO:0052697 | flavonoid glucuronidation(GO:0052696) xenobiotic glucuronidation(GO:0052697) |
| 0.2 | 0.7 | GO:0010901 | regulation of very-low-density lipoprotein particle remodeling(GO:0010901) diacylglycerol catabolic process(GO:0046340) |
| 0.2 | 1.3 | GO:0033762 | response to glucagon(GO:0033762) |
| 0.2 | 0.5 | GO:0003330 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 0.1 | 0.6 | GO:1903347 | negative regulation of bicellular tight junction assembly(GO:1903347) |
| 0.1 | 0.5 | GO:0050968 | sensory perception of sour taste(GO:0050915) detection of chemical stimulus involved in sensory perception of pain(GO:0050968) |
| 0.1 | 0.5 | GO:1901581 | post-embryonic appendage morphogenesis(GO:0035120) post-embryonic limb morphogenesis(GO:0035127) post-embryonic forelimb morphogenesis(GO:0035128) telomeric repeat-containing RNA transcription(GO:0097393) telomeric repeat-containing RNA transcription from RNA pol II promoter(GO:0097394) regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901580) negative regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901581) positive regulation of telomeric RNA transcription from RNA pol II promoter(GO:1901582) |
| 0.1 | 0.3 | GO:2000583 | regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000583) negative regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000584) |
| 0.1 | 0.2 | GO:0010808 | positive regulation of synaptic vesicle priming(GO:0010808) |
| 0.0 | 0.8 | GO:0036159 | inner dynein arm assembly(GO:0036159) |
| 0.0 | 0.7 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.0 | 0.2 | GO:0007262 | STAT protein import into nucleus(GO:0007262) negative regulation of cholesterol efflux(GO:0090370) |
| 0.0 | 0.4 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.0 | 0.6 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.0 | 1.2 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.0 | 0.5 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 1.0 | GO:0071385 | cellular response to glucocorticoid stimulus(GO:0071385) |
| 0.0 | 0.4 | GO:1902043 | positive regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902043) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.5 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.1 | 1.3 | GO:0045275 | respiratory chain complex III(GO:0045275) |
| 0.1 | 0.5 | GO:1990707 | subtelomeric heterochromatin(GO:1990421) nuclear subtelomeric heterochromatin(GO:1990707) |
| 0.1 | 1.7 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.1 | 0.8 | GO:0036156 | inner dynein arm(GO:0036156) |
| 0.0 | 0.6 | GO:0060091 | kinocilium(GO:0060091) |
| 0.0 | 0.7 | GO:0034366 | spherical high-density lipoprotein particle(GO:0034366) |
| 0.0 | 0.2 | GO:0098831 | presynaptic active zone cytoplasmic component(GO:0098831) |
| 0.0 | 0.6 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.0 | 2.3 | GO:0070469 | respiratory chain(GO:0070469) |
| 0.0 | 0.5 | GO:0043189 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 0.1 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.0 | 0.3 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.0 | 0.2 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.0 | 0.7 | GO:0043034 | costamere(GO:0043034) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 10.3 | GO:0034875 | oxidoreductase activity, acting on CH or CH2 groups, quinone or similar compound as acceptor(GO:0033695) caffeine oxidase activity(GO:0034875) |
| 0.7 | 2.7 | GO:0008113 | peptide-methionine (S)-S-oxide reductase activity(GO:0008113) |
| 0.5 | 2.1 | GO:0008802 | betaine-aldehyde dehydrogenase activity(GO:0008802) |
| 0.5 | 2.1 | GO:0004779 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 0.2 | 0.7 | GO:0070653 | high-density lipoprotein particle receptor binding(GO:0070653) |
| 0.2 | 1.3 | GO:0008121 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.1 | 0.8 | GO:0050656 | alcohol sulfotransferase activity(GO:0004027) 3'-phosphoadenosine 5'-phosphosulfate binding(GO:0050656) |
| 0.1 | 0.5 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.1 | 0.6 | GO:0008384 | IkappaB kinase activity(GO:0008384) |
| 0.1 | 3.7 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.1 | 2.3 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 0.5 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.1 | 0.7 | GO:0031802 | type 5 metabotropic glutamate receptor binding(GO:0031802) |
| 0.0 | 0.5 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.0 | 0.1 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.0 | 0.6 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.0 | 0.2 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.0 | 0.8 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.0 | 1.5 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.2 | GO:0001595 | angiotensin receptor activity(GO:0001595) |
| 0.0 | 1.1 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.0 | 0.4 | GO:0004129 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.0 | GO:1990763 | arrestin family protein binding(GO:1990763) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.4 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 0.6 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.0 | 1.7 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 2.0 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.1 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.1 | 2.4 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.6 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.0 | 0.7 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.2 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |