Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
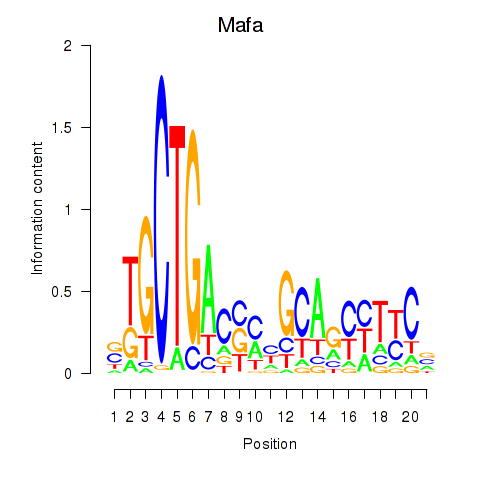
Results for Mafa
Z-value: 0.99
Transcription factors associated with Mafa
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Mafa
|
ENSMUSG00000047591.6 | Mafa |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Mafa | mm39_v1_chr15_-_75620060_75620087 | 0.17 | 3.1e-01 | Click! |
Activity profile of Mafa motif
Sorted Z-values of Mafa motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Mafa
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_-_142233270 | 6.38 |
ENSMUST00000162317.2
ENSMUST00000125933.2 ENSMUST00000105931.8 ENSMUST00000105930.8 ENSMUST00000105933.8 ENSMUST00000105932.2 ENSMUST00000000220.3 |
Ins2
|
insulin II |
| chr1_-_121255448 | 5.34 |
ENSMUST00000186915.2
ENSMUST00000160968.8 ENSMUST00000162582.2 |
Insig2
|
insulin induced gene 2 |
| chr1_-_121255753 | 5.20 |
ENSMUST00000003818.14
|
Insig2
|
insulin induced gene 2 |
| chr10_+_87695352 | 4.63 |
ENSMUST00000121952.8
ENSMUST00000126490.8 |
Igf1
|
insulin-like growth factor 1 |
| chr1_-_121255400 | 4.44 |
ENSMUST00000159085.8
ENSMUST00000159125.2 ENSMUST00000161818.2 |
Insig2
|
insulin induced gene 2 |
| chr19_+_52252735 | 4.39 |
ENSMUST00000039652.6
|
Ins1
|
insulin I |
| chr1_-_121255503 | 4.13 |
ENSMUST00000160688.2
|
Insig2
|
insulin induced gene 2 |
| chr9_-_107556823 | 2.16 |
ENSMUST00000010205.9
|
Gnat1
|
guanine nucleotide binding protein, alpha transducing 1 |
| chr1_+_171041583 | 1.96 |
ENSMUST00000111328.8
|
Nr1i3
|
nuclear receptor subfamily 1, group I, member 3 |
| chr1_+_171041539 | 1.93 |
ENSMUST00000005820.11
ENSMUST00000075469.12 ENSMUST00000155126.8 |
Nr1i3
|
nuclear receptor subfamily 1, group I, member 3 |
| chr4_+_102112189 | 1.80 |
ENSMUST00000106908.9
|
Pde4b
|
phosphodiesterase 4B, cAMP specific |
| chr1_-_169575203 | 1.59 |
ENSMUST00000027991.12
ENSMUST00000111357.2 |
Rgs4
|
regulator of G-protein signaling 4 |
| chr5_+_111565129 | 1.01 |
ENSMUST00000094463.5
|
Mn1
|
meningioma 1 |
| chr14_-_24054352 | 1.00 |
ENSMUST00000190339.2
|
Kcnma1
|
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr9_+_45817795 | 0.92 |
ENSMUST00000039059.8
|
Pcsk7
|
proprotein convertase subtilisin/kexin type 7 |
| chr3_+_108561223 | 0.92 |
ENSMUST00000106609.8
|
Clcc1
|
chloride channel CLIC-like 1 |
| chr5_-_24628483 | 0.90 |
ENSMUST00000198990.2
|
Cdk5
|
cyclin-dependent kinase 5 |
| chr5_-_24628514 | 0.85 |
ENSMUST00000030814.11
|
Cdk5
|
cyclin-dependent kinase 5 |
| chr4_+_116578117 | 0.78 |
ENSMUST00000045542.13
ENSMUST00000106459.8 |
Tesk2
|
testis-specific kinase 2 |
| chr6_-_23839419 | 0.68 |
ENSMUST00000115358.9
ENSMUST00000163871.9 |
Cadps2
|
Ca2+-dependent activator protein for secretion 2 |
| chr3_-_153650068 | 0.65 |
ENSMUST00000150070.2
|
Acadm
|
acyl-Coenzyme A dehydrogenase, medium chain |
| chr5_-_46014809 | 0.64 |
ENSMUST00000190036.7
ENSMUST00000189859.7 ENSMUST00000186633.3 ENSMUST00000016026.14 ENSMUST00000045586.13 ENSMUST00000238522.2 |
Lcorl
|
ligand dependent nuclear receptor corepressor-like |
| chr3_-_153650269 | 0.63 |
ENSMUST00000072697.13
|
Acadm
|
acyl-Coenzyme A dehydrogenase, medium chain |
| chr7_-_65020955 | 0.63 |
ENSMUST00000102592.10
|
Tjp1
|
tight junction protein 1 |
| chr16_+_17149235 | 0.62 |
ENSMUST00000023450.15
ENSMUST00000231884.2 |
Serpind1
|
serine (or cysteine) peptidase inhibitor, clade D, member 1 |
| chr11_-_66416790 | 0.62 |
ENSMUST00000066679.7
|
Shisa6
|
shisa family member 6 |
| chr9_-_105009023 | 0.61 |
ENSMUST00000035179.9
|
Nudt16
|
nudix (nucleoside diphosphate linked moiety X)-type motif 16 |
| chr11_+_98818640 | 0.56 |
ENSMUST00000107474.8
|
Rara
|
retinoic acid receptor, alpha |
| chr4_+_124696394 | 0.54 |
ENSMUST00000175875.2
|
Mtf1
|
metal response element binding transcription factor 1 |
| chr9_+_21746785 | 0.53 |
ENSMUST00000058777.8
|
Angptl8
|
angiopoietin-like 8 |
| chr4_+_155685830 | 0.49 |
ENSMUST00000105608.9
|
Slc35e2
|
solute carrier family 35, member E2 |
| chr11_-_66416621 | 0.47 |
ENSMUST00000123454.8
|
Shisa6
|
shisa family member 6 |
| chr6_+_85164420 | 0.47 |
ENSMUST00000045942.9
|
Emx1
|
empty spiracles homeobox 1 |
| chr1_-_135095344 | 0.47 |
ENSMUST00000027682.9
|
Gpr37l1
|
G protein-coupled receptor 37-like 1 |
| chr17_-_47321430 | 0.46 |
ENSMUST00000113337.10
ENSMUST00000113335.4 |
Ubr2
|
ubiquitin protein ligase E3 component n-recognin 2 |
| chr4_+_155686311 | 0.46 |
ENSMUST00000118607.2
|
Slc35e2
|
solute carrier family 35, member E2 |
| chr19_-_36097233 | 0.44 |
ENSMUST00000025718.10
|
Ankrd1
|
ankyrin repeat domain 1 (cardiac muscle) |
| chr9_-_97915036 | 0.43 |
ENSMUST00000162295.2
|
Clstn2
|
calsyntenin 2 |
| chr5_+_24628802 | 0.42 |
ENSMUST00000153274.2
ENSMUST00000141966.8 |
Slc4a2
|
solute carrier family 4 (anion exchanger), member 2 |
| chr9_-_105008972 | 0.42 |
ENSMUST00000186925.2
|
Nudt16
|
nudix (nucleoside diphosphate linked moiety X)-type motif 16 |
| chr11_+_87486472 | 0.41 |
ENSMUST00000134216.3
|
Mtmr4
|
myotubularin related protein 4 |
| chr5_-_124018055 | 0.40 |
ENSMUST00000164267.2
|
Hcar1
|
hydrocarboxylic acid receptor 1 |
| chr17_-_28299544 | 0.37 |
ENSMUST00000042692.13
ENSMUST00000114836.9 |
Tcp11
|
t-complex protein 11 |
| chr17_-_27732364 | 0.37 |
ENSMUST00000118161.3
|
Grm4
|
glutamate receptor, metabotropic 4 |
| chr2_-_73605684 | 0.36 |
ENSMUST00000112024.10
ENSMUST00000180045.8 |
Chn1
|
chimerin 1 |
| chr19_+_27194757 | 0.35 |
ENSMUST00000047645.13
ENSMUST00000167487.8 |
Vldlr
|
very low density lipoprotein receptor |
| chrX_-_20816841 | 0.34 |
ENSMUST00000009550.14
|
Elk1
|
ELK1, member of ETS oncogene family |
| chr13_-_43634695 | 0.33 |
ENSMUST00000144326.4
|
Ranbp9
|
RAN binding protein 9 |
| chrX_+_150961960 | 0.32 |
ENSMUST00000096275.5
|
Iqsec2
|
IQ motif and Sec7 domain 2 |
| chr4_+_43406435 | 0.31 |
ENSMUST00000098106.9
ENSMUST00000139198.2 |
Rusc2
|
RUN and SH3 domain containing 2 |
| chr19_+_34899811 | 0.31 |
ENSMUST00000223776.2
|
Kif20b
|
kinesin family member 20B |
| chr12_-_87346479 | 0.30 |
ENSMUST00000125733.8
|
Ism2
|
isthmin 2 |
| chr16_+_96268806 | 0.30 |
ENSMUST00000061739.9
|
Pcp4
|
Purkinje cell protein 4 |
| chr14_-_24054273 | 0.27 |
ENSMUST00000188285.7
ENSMUST00000190044.7 |
Kcnma1
|
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr7_+_24335969 | 0.27 |
ENSMUST00000080718.6
|
Lypd3
|
Ly6/Plaur domain containing 3 |
| chr9_-_44366273 | 0.27 |
ENSMUST00000047740.4
|
Upk2
|
uroplakin 2 |
| chr2_-_73605387 | 0.26 |
ENSMUST00000166199.9
|
Chn1
|
chimerin 1 |
| chr2_+_38821987 | 0.25 |
ENSMUST00000057279.6
|
Olfml2a
|
olfactomedin-like 2A |
| chr4_-_59138983 | 0.24 |
ENSMUST00000107547.2
|
Shoc1
|
shortage in chiasmata 1 |
| chr12_-_75224099 | 0.24 |
ENSMUST00000042299.4
|
Kcnh5
|
potassium voltage-gated channel, subfamily H (eag-related), member 5 |
| chr8_+_106099894 | 0.24 |
ENSMUST00000160650.2
|
Plekhg4
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 4 |
| chr11_+_105858764 | 0.24 |
ENSMUST00000001963.14
|
Ace
|
angiotensin I converting enzyme (peptidyl-dipeptidase A) 1 |
| chr5_+_124767114 | 0.24 |
ENSMUST00000037865.13
|
Atp6v0a2
|
ATPase, H+ transporting, lysosomal V0 subunit A2 |
| chr17_-_28299569 | 0.23 |
ENSMUST00000129046.9
ENSMUST00000043925.16 |
Tcp11
|
t-complex protein 11 |
| chr6_-_35285045 | 0.23 |
ENSMUST00000031868.5
|
Slc13a4
|
solute carrier family 13 (sodium/sulfate symporters), member 4 |
| chr9_-_97915227 | 0.22 |
ENSMUST00000035027.13
|
Clstn2
|
calsyntenin 2 |
| chr11_-_119119287 | 0.21 |
ENSMUST00000207655.2
ENSMUST00000036113.4 |
Tbc1d16
|
TBC1 domain family, member 16 |
| chr14_-_24054186 | 0.21 |
ENSMUST00000188991.7
ENSMUST00000224468.2 |
Kcnma1
|
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chrX_-_96420756 | 0.20 |
ENSMUST00000113832.2
ENSMUST00000037353.10 |
Eda2r
|
ectodysplasin A2 receptor |
| chr2_+_86127161 | 0.19 |
ENSMUST00000215090.2
|
Olfr1052
|
olfactory receptor 1052 |
| chrX_-_138683102 | 0.18 |
ENSMUST00000101217.4
|
Ripply1
|
ripply transcriptional repressor 1 |
| chr1_-_58735106 | 0.18 |
ENSMUST00000055313.14
ENSMUST00000188772.7 |
Flacc1
|
flagellum associated containing coiled-coil domains 1 |
| chr19_+_27194411 | 0.18 |
ENSMUST00000164746.8
ENSMUST00000172302.8 |
Vldlr
|
very low density lipoprotein receptor |
| chr7_-_12418918 | 0.17 |
ENSMUST00000167771.2
ENSMUST00000172743.9 ENSMUST00000233763.2 ENSMUST00000232880.2 |
Vmn2r55
|
vomeronasal 2, receptor 55 |
| chr17_+_31896781 | 0.16 |
ENSMUST00000228716.3
ENSMUST00000019192.8 ENSMUST00000228089.3 |
Cryaa
|
crystallin, alpha A |
| chr19_-_59158488 | 0.14 |
ENSMUST00000172821.4
|
Vax1
|
ventral anterior homeobox 1 |
| chr6_+_119213541 | 0.14 |
ENSMUST00000186622.3
|
Cacna2d4
|
calcium channel, voltage-dependent, alpha 2/delta subunit 4 |
| chr7_+_54485336 | 0.13 |
ENSMUST00000082373.8
|
Luzp2
|
leucine zipper protein 2 |
| chr16_+_64672334 | 0.12 |
ENSMUST00000067744.8
|
Cggbp1
|
CGG triplet repeat binding protein 1 |
| chr14_+_79825131 | 0.11 |
ENSMUST00000054908.10
|
Sugt1
|
SGT1, suppressor of G2 allele of SKP1 (S. cerevisiae) |
| chrX_+_103526380 | 0.11 |
ENSMUST00000087867.6
|
Uprt
|
uracil phosphoribosyltransferase |
| chr12_+_53294940 | 0.10 |
ENSMUST00000223358.3
|
Npas3
|
neuronal PAS domain protein 3 |
| chr14_-_24053994 | 0.10 |
ENSMUST00000225431.2
ENSMUST00000188210.8 ENSMUST00000224787.2 ENSMUST00000225315.2 ENSMUST00000225556.2 ENSMUST00000223727.2 ENSMUST00000223655.2 ENSMUST00000224077.2 ENSMUST00000224812.2 ENSMUST00000224285.2 ENSMUST00000225471.2 ENSMUST00000224232.2 ENSMUST00000223749.2 ENSMUST00000224025.2 |
Kcnma1
|
potassium large conductance calcium-activated channel, subfamily M, alpha member 1 |
| chr6_+_119213468 | 0.08 |
ENSMUST00000238905.2
ENSMUST00000037434.13 |
Cacna2d4
|
calcium channel, voltage-dependent, alpha 2/delta subunit 4 |
| chr7_-_103814019 | 0.08 |
ENSMUST00000154555.2
ENSMUST00000051137.15 |
Olfm5
|
olfactomedin 5 |
| chr16_-_17394495 | 0.08 |
ENSMUST00000231645.2
ENSMUST00000232226.2 ENSMUST00000232336.2 ENSMUST00000232385.2 ENSMUST00000231615.2 ENSMUST00000231283.2 ENSMUST00000172164.10 |
Slc7a4
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 4 |
| chr6_+_65358089 | 0.07 |
ENSMUST00000133352.8
|
Qrfprl
|
pyroglutamylated RFamide peptide receptor like |
| chr17_-_72910822 | 0.07 |
ENSMUST00000086639.6
|
Alk
|
anaplastic lymphoma kinase |
| chr11_+_119119925 | 0.06 |
ENSMUST00000053440.8
|
Ccdc40
|
coiled-coil domain containing 40 |
| chr4_-_141143313 | 0.04 |
ENSMUST00000006378.9
ENSMUST00000105788.2 |
Clcnkb
|
chloride channel, voltage-sensitive Kb |
| chr9_-_108888284 | 0.01 |
ENSMUST00000112053.2
|
Trex1
|
three prime repair exonuclease 1 |
| chr9_-_108888779 | 0.01 |
ENSMUST00000061973.5
|
Trex1
|
three prime repair exonuclease 1 |
| chr16_+_65612394 | 0.00 |
ENSMUST00000227997.2
|
Vgll3
|
vestigial like family member 3 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 6.4 | GO:0033861 | negative regulation of NAD(P)H oxidase activity(GO:0033861) neuron projection maintenance(GO:1990535) |
| 1.1 | 19.1 | GO:0060363 | cranial suture morphogenesis(GO:0060363) |
| 0.7 | 4.6 | GO:1904075 | regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 0.4 | 2.2 | GO:0050917 | sensory perception of umami taste(GO:0050917) regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051342) |
| 0.3 | 1.3 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.3 | 1.6 | GO:0034465 | response to carbon monoxide(GO:0034465) eye blink reflex(GO:0060082) |
| 0.2 | 1.0 | GO:0046707 | IDP metabolic process(GO:0046707) IDP catabolic process(GO:0046709) |
| 0.2 | 0.6 | GO:1902490 | regulation of sperm capacitation(GO:1902490) |
| 0.2 | 0.6 | GO:0060010 | Sertoli cell fate commitment(GO:0060010) |
| 0.2 | 1.7 | GO:0046826 | negative regulation of protein export from nucleus(GO:0046826) |
| 0.1 | 1.8 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.1 | 1.0 | GO:0001957 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.1 | 0.5 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
| 0.1 | 0.5 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.1 | 0.5 | GO:0034436 | glycoprotein transport(GO:0034436) |
| 0.1 | 0.7 | GO:1990504 | dense core granule exocytosis(GO:1990504) |
| 0.1 | 0.2 | GO:2000170 | positive regulation of peptidyl-cysteine S-nitrosylation(GO:2000170) |
| 0.0 | 1.1 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.0 | 0.2 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.0 | 0.5 | GO:0021940 | positive regulation of cerebellar granule cell precursor proliferation(GO:0021940) |
| 0.0 | 0.1 | GO:0052055 | modulation by symbiont of host molecular function(GO:0052055) modulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052203) modulation by host of symbiont catalytic activity(GO:0052422) |
| 0.0 | 0.4 | GO:0007196 | adenylate cyclase-inhibiting G-protein coupled glutamate receptor signaling pathway(GO:0007196) |
| 0.0 | 0.5 | GO:0060019 | radial glial cell differentiation(GO:0060019) |
| 0.0 | 0.6 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.0 | 0.2 | GO:0070309 | lens fiber cell morphogenesis(GO:0070309) |
| 0.0 | 0.1 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.1 | GO:0046049 | UMP biosynthetic process(GO:0006222) pyrimidine ribonucleoside monophosphate metabolic process(GO:0009173) pyrimidine ribonucleoside monophosphate biosynthetic process(GO:0009174) UMP metabolic process(GO:0046049) |
| 0.0 | 4.4 | GO:0015758 | glucose transport(GO:0015758) |
| 0.0 | 0.4 | GO:0043517 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) |
| 0.0 | 0.3 | GO:1900454 | positive regulation of long term synaptic depression(GO:1900454) |
| 0.0 | 0.2 | GO:0070072 | vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.0 | 0.3 | GO:0033603 | positive regulation of dopamine secretion(GO:0033603) |
| 0.0 | 3.9 | GO:0043401 | steroid hormone mediated signaling pathway(GO:0043401) |
| 0.0 | 0.6 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.0 | 0.1 | GO:0007406 | negative regulation of neuroblast proliferation(GO:0007406) |
| 0.0 | 0.2 | GO:0032525 | somite rostral/caudal axis specification(GO:0032525) |
| 0.0 | 0.2 | GO:0008272 | sulfate transport(GO:0008272) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.4 | 19.1 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.5 | 4.6 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.4 | 1.7 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.2 | 6.4 | GO:0005732 | small nucleolar ribonucleoprotein complex(GO:0005732) |
| 0.1 | 1.9 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.1 | 1.8 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.0 | 2.2 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 0.0 | 0.6 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 0.2 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.0 | 0.8 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.0 | 1.1 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 0.3 | GO:0005883 | neurofilament(GO:0005883) |
| 0.0 | 0.5 | GO:0034385 | very-low-density lipoprotein particle(GO:0034361) triglyceride-rich lipoprotein particle(GO:0034385) |
| 0.0 | 0.9 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 1.0 | GO:0034707 | chloride channel complex(GO:0034707) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.0 | GO:0035870 | dITP diphosphatase activity(GO:0035870) |
| 0.3 | 1.3 | GO:0070991 | medium-chain-acyl-CoA dehydrogenase activity(GO:0070991) |
| 0.3 | 3.9 | GO:0004887 | thyroid hormone receptor activity(GO:0004887) |
| 0.2 | 1.7 | GO:0030549 | acetylcholine receptor activator activity(GO:0030549) |
| 0.2 | 11.0 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.2 | 1.6 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.2 | 0.5 | GO:0034189 | very-low-density lipoprotein particle binding(GO:0034189) |
| 0.1 | 4.4 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.1 | 0.5 | GO:0070728 | leucine binding(GO:0070728) |
| 0.1 | 0.6 | GO:0071253 | connexin binding(GO:0071253) |
| 0.1 | 0.4 | GO:0061629 | RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 0.1 | 2.1 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.1 | 0.4 | GO:0001642 | group III metabotropic glutamate receptor activity(GO:0001642) |
| 0.0 | 0.2 | GO:0031711 | tripeptidyl-peptidase activity(GO:0008240) bradykinin receptor binding(GO:0031711) |
| 0.0 | 0.6 | GO:0035014 | phosphatidylinositol 3-kinase regulator activity(GO:0035014) |
| 0.0 | 1.8 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 0.2 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.0 | 1.6 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.4 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.0 | 19.4 | GO:0008134 | transcription factor binding(GO:0008134) |
| 0.0 | 0.1 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.0 | 0.1 | GO:0001162 | RNA polymerase II intronic transcription regulatory region sequence-specific DNA binding(GO:0001162) |
| 0.0 | 0.2 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.0 | 0.6 | GO:0046875 | ephrin receptor binding(GO:0046875) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 4.6 | PID IGF1 PATHWAY | IGF1 pathway |
| 0.1 | 2.2 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 2.3 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 0.3 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.0 | 3.9 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.0 | 1.3 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.6 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.0 | 0.6 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 0.2 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 3.5 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 2.1 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.0 | 4.5 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 1.6 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.0 | 0.6 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.0 | 0.4 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |