Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
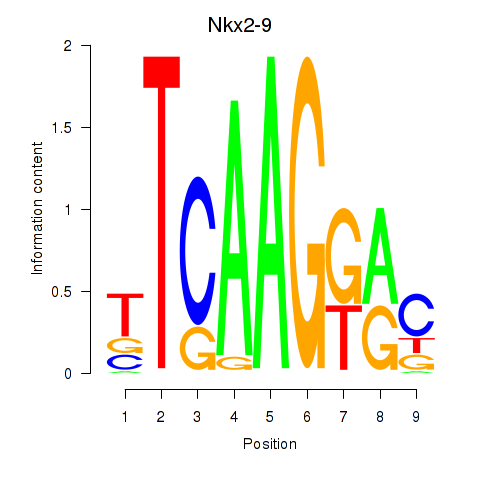
Results for Nkx2-9
Z-value: 2.42
Transcription factors associated with Nkx2-9
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Nkx2-9
|
ENSMUSG00000058669.8 | Nkx2-9 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Nkx2-9 | mm39_v1_chr12_-_56660054_56660113 | -0.57 | 3.2e-04 | Click! |
Activity profile of Nkx2-9 motif
Sorted Z-values of Nkx2-9 motif
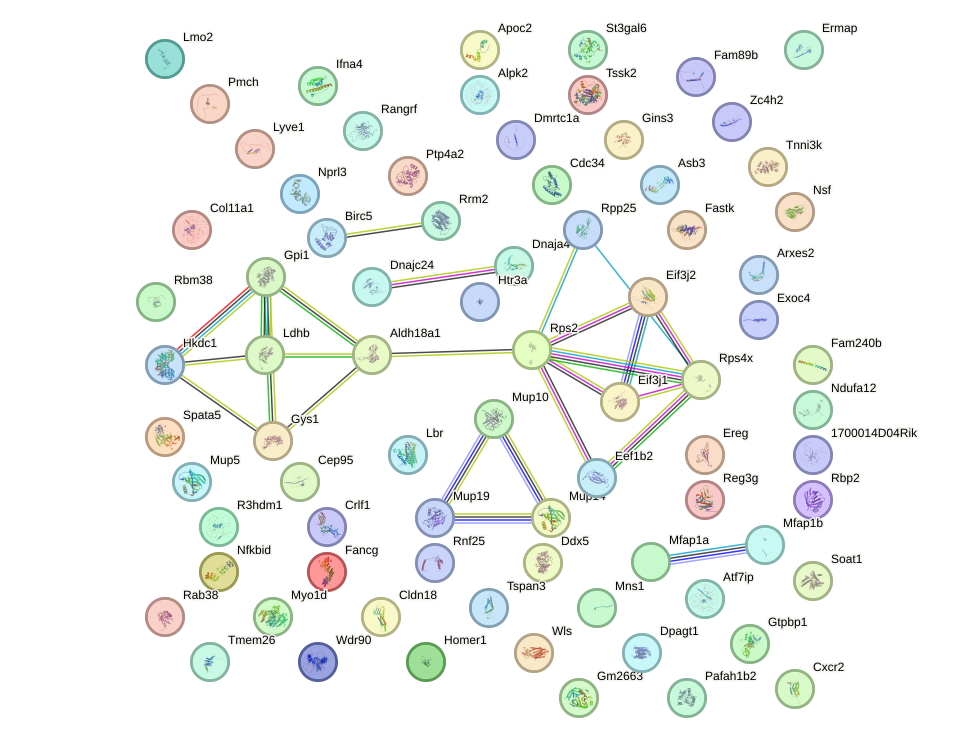
Network of associatons between targets according to the STRING database.
First level regulatory network of Nkx2-9
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_172864153 | 13.33 |
ENSMUST00000173997.2
|
Rbm38
|
RNA binding motif protein 38 |
| chr4_-_119047202 | 12.28 |
ENSMUST00000239029.2
ENSMUST00000138395.9 ENSMUST00000156746.3 |
Ermap
|
erythroblast membrane-associated protein |
| chr4_-_119047167 | 10.55 |
ENSMUST00000030396.15
|
Ermap
|
erythroblast membrane-associated protein |
| chr9_+_98372575 | 10.06 |
ENSMUST00000035029.3
|
Rbp2
|
retinol binding protein 2, cellular |
| chr4_-_119047180 | 8.61 |
ENSMUST00000150864.3
ENSMUST00000141227.9 |
Ermap
|
erythroblast membrane-associated protein |
| chr2_+_103799873 | 8.56 |
ENSMUST00000123437.8
|
Lmo2
|
LIM domain only 2 |
| chr4_-_119047146 | 8.51 |
ENSMUST00000124626.9
|
Ermap
|
erythroblast membrane-associated protein |
| chr11_+_117740111 | 7.56 |
ENSMUST00000093906.5
|
Birc5
|
baculoviral IAP repeat-containing 5 |
| chr9_+_54606832 | 4.57 |
ENSMUST00000070070.8
|
Dnaja4
|
DnaJ heat shock protein family (Hsp40) member A4 |
| chr10_-_62258195 | 4.19 |
ENSMUST00000020277.9
|
Hkdc1
|
hexokinase domain containing 1 |
| chr9_+_72345801 | 4.13 |
ENSMUST00000184604.8
ENSMUST00000034746.10 |
Mns1
|
meiosis-specific nuclear structural protein 1 |
| chr12_+_24758724 | 4.03 |
ENSMUST00000153058.8
|
Rrm2
|
ribonucleotide reductase M2 |
| chr11_+_117740077 | 3.95 |
ENSMUST00000081387.11
|
Birc5
|
baculoviral IAP repeat-containing 5 |
| chr12_+_24758968 | 3.67 |
ENSMUST00000154588.2
|
Rrm2
|
ribonucleotide reductase M2 |
| chr1_-_181669891 | 3.58 |
ENSMUST00000193028.2
ENSMUST00000191878.6 ENSMUST00000005003.12 |
Lbr
|
lamin B receptor |
| chrX_-_101232978 | 3.32 |
ENSMUST00000033683.8
|
Rps4x
|
ribosomal protein S4, X-linked |
| chr16_-_58344548 | 3.16 |
ENSMUST00000114357.10
|
St3gal6
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 6 |
| chr19_-_40576782 | 3.08 |
ENSMUST00000176939.8
|
Aldh18a1
|
aldehyde dehydrogenase 18 family, member A1 |
| chr11_-_68864666 | 3.01 |
ENSMUST00000038644.5
|
Rangrf
|
RAN guanine nucleotide release factor |
| chr16_-_58344602 | 2.90 |
ENSMUST00000114358.9
|
St3gal6
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 6 |
| chr4_-_61259997 | 2.83 |
ENSMUST00000071005.9
ENSMUST00000075206.12 |
Mup14
|
major urinary protein 14 |
| chrX_+_134894573 | 2.74 |
ENSMUST00000058119.9
|
Arxes2
|
adipocyte-related X-chromosome expressed sequence 2 |
| chr4_-_60777462 | 2.65 |
ENSMUST00000211875.2
|
Mup22
|
major urinary protein 22 |
| chr4_-_60222580 | 2.63 |
ENSMUST00000095058.5
ENSMUST00000163931.8 |
Mup8
|
major urinary protein 8 |
| chr15_+_79575046 | 2.61 |
ENSMUST00000046463.10
|
Gtpbp1
|
GTP binding protein 1 |
| chr4_-_61259801 | 2.56 |
ENSMUST00000125461.8
|
Mup14
|
major urinary protein 14 |
| chr4_-_61700450 | 2.50 |
ENSMUST00000107477.2
ENSMUST00000080606.9 |
Mup19
|
major urinary protein 19 |
| chr7_-_19411866 | 2.41 |
ENSMUST00000142352.9
|
Apoc2
|
apolipoprotein C-II |
| chr11_+_114559350 | 2.39 |
ENSMUST00000106602.10
ENSMUST00000077915.10 ENSMUST00000106599.8 ENSMUST00000082092.5 |
Rpl38
|
ribosomal protein L38 |
| chr1_+_63215976 | 2.23 |
ENSMUST00000129339.8
|
Eef1b2
|
eukaryotic translation elongation factor 1 beta 2 |
| chr7_+_45084257 | 2.23 |
ENSMUST00000003964.17
|
Gys1
|
glycogen synthase 1, muscle |
| chr6_-_40976413 | 2.19 |
ENSMUST00000166306.3
|
Gm2663
|
predicted gene 2663 |
| chr9_-_45923908 | 2.15 |
ENSMUST00000217514.2
|
Pafah1b2
|
platelet-activating factor acetylhydrolase, isoform 1b, subunit 2 |
| chr13_+_93441447 | 2.10 |
ENSMUST00000109497.8
ENSMUST00000109498.8 ENSMUST00000060490.11 ENSMUST00000109492.9 ENSMUST00000109496.8 ENSMUST00000109495.8 |
Homer1
|
homer scaffolding protein 1 |
| chr6_-_142453531 | 2.09 |
ENSMUST00000134191.3
ENSMUST00000239397.2 ENSMUST00000239395.2 ENSMUST00000032373.12 |
Ldhb
|
lactate dehydrogenase B |
| chr1_+_74193138 | 2.05 |
ENSMUST00000027372.8
ENSMUST00000106899.4 |
Cxcr2
|
chemokine (C-X-C motif) receptor 2 |
| chr1_+_128031055 | 1.92 |
ENSMUST00000188381.7
ENSMUST00000187900.7 ENSMUST00000036288.11 |
R3hdm1
|
R3H domain containing 1 |
| chr1_-_156301821 | 1.91 |
ENSMUST00000188027.2
ENSMUST00000187507.7 ENSMUST00000189661.7 |
Soat1
|
sterol O-acyltransferase 1 |
| chr7_+_45084300 | 1.90 |
ENSMUST00000211150.2
|
Gys1
|
glycogen synthase 1, muscle |
| chr9_-_99592116 | 1.84 |
ENSMUST00000035048.12
|
Cldn18
|
claudin 18 |
| chr16_-_58344497 | 1.82 |
ENSMUST00000151510.2
ENSMUST00000126978.8 ENSMUST00000123918.8 |
St3gal6
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 6 |
| chr2_+_121337226 | 1.75 |
ENSMUST00000099473.10
ENSMUST00000110602.9 |
Wdr76
|
WD repeat domain 76 |
| chr17_+_24939225 | 1.72 |
ENSMUST00000146867.2
|
Rps2
|
ribosomal protein S2 |
| chr7_-_110462446 | 1.67 |
ENSMUST00000033050.5
|
Lyve1
|
lymphatic vessel endothelial hyaluronan receptor 1 |
| chr19_-_5779648 | 1.61 |
ENSMUST00000116558.3
ENSMUST00000099955.4 ENSMUST00000161368.2 |
Fam89b
|
family with sequence similarity 89, member B |
| chr4_-_43010225 | 1.57 |
ENSMUST00000030165.5
|
Fancg
|
Fanconi anemia, complementation group G |
| chr3_+_113824181 | 1.51 |
ENSMUST00000123619.8
ENSMUST00000092155.12 |
Col11a1
|
collagen, type XI, alpha 1 |
| chr7_-_33929667 | 1.51 |
ENSMUST00000206415.2
|
Gpi1
|
glucose-6-phosphate isomerase 1 |
| chr11_-_80670815 | 1.46 |
ENSMUST00000041065.14
ENSMUST00000070997.6 |
Myo1d
|
myosin ID |
| chr7_-_33929708 | 1.40 |
ENSMUST00000038027.6
|
Gpi1
|
glucose-6-phosphate isomerase 1 |
| chr9_-_99592058 | 1.32 |
ENSMUST00000136429.8
|
Cldn18
|
claudin 18 |
| chr1_-_136273811 | 1.20 |
ENSMUST00000048309.12
|
Camsap2
|
calmodulin regulated spectrin-associated protein family, member 2 |
| chr6_-_113411489 | 1.18 |
ENSMUST00000133348.2
|
Cidec
|
cell death-inducing DFFA-like effector c |
| chr11_+_30904390 | 1.18 |
ENSMUST00000020551.13
ENSMUST00000117883.2 |
Asb3
|
ankyrin repeat and SOCS box-containing 3 |
| chr3_+_37474422 | 1.16 |
ENSMUST00000029277.13
ENSMUST00000198968.2 |
Spata5
|
spermatogenesis associated 5 |
| chr2_+_121337195 | 1.15 |
ENSMUST00000028676.12
|
Wdr76
|
WD repeat domain 76 |
| chr10_+_79518138 | 1.14 |
ENSMUST00000166603.2
ENSMUST00000219791.2 ENSMUST00000219930.2 ENSMUST00000218964.2 |
Cdc34
|
cell division cycle 34 |
| chr10_+_87926932 | 1.11 |
ENSMUST00000048621.8
|
Pmch
|
pro-melanin-concentrating hormone |
| chr12_-_113561594 | 1.06 |
ENSMUST00000103444.3
|
Ighv5-4
|
immunoglobulin heavy variable 5-4 |
| chr7_+_88079465 | 1.06 |
ENSMUST00000107256.4
|
Rab38
|
RAB38, member RAS oncogene family |
| chr12_-_113666198 | 1.01 |
ENSMUST00000103450.4
|
Ighv5-12
|
immunoglobulin heavy variable 5-12 |
| chr10_+_68559553 | 1.01 |
ENSMUST00000080995.8
|
Tmem26
|
transmembrane protein 26 |
| chr8_+_96360081 | 1.00 |
ENSMUST00000034094.11
ENSMUST00000212434.2 |
Gins3
|
GINS complex subunit 3 (Psf3 homolog) |
| chr18_-_43610829 | 0.96 |
ENSMUST00000057110.11
|
Eif3j2
|
eukaryotic translation initiation factor 3, subunit J2 |
| chr6_-_38331482 | 0.93 |
ENSMUST00000031850.10
ENSMUST00000114898.3 |
Zc3hav1
|
zinc finger CCCH type, antiviral 1 |
| chr13_-_59890564 | 0.90 |
ENSMUST00000055343.3
|
1700014D04Rik
|
RIKEN cDNA 1700014D04 gene |
| chr6_+_135339929 | 0.89 |
ENSMUST00000032330.16
|
Emp1
|
epithelial membrane protein 1 |
| chrX_+_41238193 | 0.89 |
ENSMUST00000115073.9
ENSMUST00000115072.8 |
Stag2
|
stromal antigen 2 |
| chr6_-_78445846 | 0.89 |
ENSMUST00000032089.3
|
Reg3g
|
regenerating islet-derived 3 gamma |
| chr5_-_138185438 | 0.88 |
ENSMUST00000110937.8
ENSMUST00000139276.2 ENSMUST00000048698.14 ENSMUST00000123415.8 |
Taf6
|
TATA-box binding protein associated factor 6 |
| chr9_-_56068282 | 0.88 |
ENSMUST00000034876.10
|
Tspan3
|
tetraspanin 3 |
| chr6_+_33226020 | 0.86 |
ENSMUST00000052266.15
ENSMUST00000090381.11 ENSMUST00000115080.2 |
Exoc4
|
exocyst complex component 4 |
| chr12_-_113589576 | 0.84 |
ENSMUST00000103446.2
|
Ighv5-6
|
immunoglobulin heavy variable 5-6 |
| chr9_+_57411279 | 0.81 |
ENSMUST00000080514.9
|
Rpp25
|
ribonuclease P/MRP 25 subunit |
| chr12_-_114443071 | 0.79 |
ENSMUST00000103492.2
|
Ighv10-1
|
immunoglobulin heavy variable 10-1 |
| chr12_-_113802603 | 0.78 |
ENSMUST00000103458.3
ENSMUST00000193652.2 |
Ighv5-16
|
immunoglobulin heavy variable 5-16 |
| chr8_-_81607140 | 0.76 |
ENSMUST00000210676.2
|
Gab1
|
growth factor receptor bound protein 2-associated protein 1 |
| chr11_-_103844870 | 0.75 |
ENSMUST00000103075.11
|
Nsf
|
N-ethylmaleimide sensitive fusion protein |
| chr18_-_60981981 | 0.75 |
ENSMUST00000177172.8
ENSMUST00000175934.8 ENSMUST00000176630.8 |
Tcof1
|
treacle ribosome biogenesis factor 1 |
| chr2_-_121337120 | 0.74 |
ENSMUST00000089926.6
|
Mfap1a
|
microfibrillar-associated protein 1A |
| chr8_-_81607109 | 0.71 |
ENSMUST00000034150.10
|
Gab1
|
growth factor receptor bound protein 2-associated protein 1 |
| chr11_-_32217547 | 0.68 |
ENSMUST00000109389.9
ENSMUST00000129010.2 ENSMUST00000020530.12 |
Nprl3
|
nitrogen permease regulator-like 3 |
| chr12_-_113700190 | 0.67 |
ENSMUST00000103452.3
ENSMUST00000192264.2 |
Ighv5-9-1
|
immunoglobulin heavy variable 5-9-1 |
| chr12_-_115825934 | 0.66 |
ENSMUST00000198777.2
|
Ighv1-77
|
immunoglobulin heavy variable 1-77 |
| chr8_+_70945806 | 0.63 |
ENSMUST00000008032.14
|
Crlf1
|
cytokine receptor-like factor 1 |
| chr10_+_94034817 | 0.59 |
ENSMUST00000020209.16
ENSMUST00000179990.8 |
Ndufa12
|
NADH:ubiquinone oxidoreductase subunit A12 |
| chr12_-_113823290 | 0.59 |
ENSMUST00000103459.5
|
Ighv5-17
|
immunoglobulin heavy variable 5-17 |
| chr19_+_12257218 | 0.55 |
ENSMUST00000207186.4
ENSMUST00000207915.2 |
Olfr1434
|
olfactory receptor 1434 |
| chr11_+_106680062 | 0.55 |
ENSMUST00000103068.10
ENSMUST00000018516.11 |
Cep95
|
centrosomal protein 95 |
| chr18_+_37102969 | 0.54 |
ENSMUST00000194544.2
|
Gm37013
|
predicted gene, 37013 |
| chr3_-_154760978 | 0.54 |
ENSMUST00000064076.6
|
Tnni3k
|
TNNI3 interacting kinase |
| chr2_-_105832353 | 0.52 |
ENSMUST00000155811.2
|
Dnajc24
|
DnaJ heat shock protein family (Hsp40) member C24 |
| chr5_-_138185686 | 0.52 |
ENSMUST00000110936.8
|
Taf6
|
TATA-box binding protein associated factor 6 |
| chr6_+_136530970 | 0.50 |
ENSMUST00000189535.2
|
Atf7ip
|
activating transcription factor 7 interacting protein |
| chr7_+_75259778 | 0.50 |
ENSMUST00000207923.2
|
Akap13
|
A kinase (PRKA) anchor protein 13 |
| chrX_-_94701983 | 0.49 |
ENSMUST00000119640.8
ENSMUST00000120620.8 ENSMUST00000044382.7 |
Zc4h2
|
zinc finger, C4H2 domain containing |
| chr4_-_61753422 | 0.49 |
ENSMUST00000082287.3
|
Mup5
|
major urinary protein 5 |
| chr11_-_106679983 | 0.49 |
ENSMUST00000129585.8
|
Ddx5
|
DEAD box helicase 5 |
| chr14_+_53007210 | 0.49 |
ENSMUST00000178768.4
|
Trav7d-4
|
T cell receptor alpha variable 7D-4 |
| chr16_+_17716480 | 0.48 |
ENSMUST00000055374.8
|
Tssk2
|
testis-specific serine kinase 2 |
| chrX_+_7744535 | 0.47 |
ENSMUST00000033495.15
|
Pim2
|
proviral integration site 2 |
| chr12_-_113860566 | 0.44 |
ENSMUST00000103474.5
|
Ighv7-1
|
immunoglobulin heavy variable 7-1 |
| chr11_+_69920849 | 0.43 |
ENSMUST00000143920.4
|
Dlg4
|
discs large MAGUK scaffold protein 4 |
| chr13_+_109397184 | 0.42 |
ENSMUST00000153234.8
|
Pde4d
|
phosphodiesterase 4D, cAMP specific |
| chr4_+_88760098 | 0.42 |
ENSMUST00000094973.5
|
Ifna4
|
interferon alpha 4 |
| chr4_+_129714494 | 0.41 |
ENSMUST00000165853.2
|
Ptp4a2
|
protein tyrosine phosphatase 4a2 |
| chr3_+_159545309 | 0.39 |
ENSMUST00000068952.10
ENSMUST00000198878.2 |
Wls
|
wntless WNT ligand secretion mediator |
| chrX_-_101950075 | 0.39 |
ENSMUST00000120808.8
ENSMUST00000121197.8 |
Dmrtc1a
|
DMRT-like family C1a |
| chr17_-_26080429 | 0.38 |
ENSMUST00000079461.15
ENSMUST00000176923.9 |
Wdr90
|
WD repeat domain 90 |
| chr19_-_12147438 | 0.37 |
ENSMUST00000207679.3
ENSMUST00000219261.2 |
Olfr1555-ps1
|
olfactory receptor 1555, pseudogene 1 |
| chr11_+_69920956 | 0.37 |
ENSMUST00000232115.2
|
Dlg4
|
discs large MAGUK scaffold protein 4 |
| chr12_-_114502585 | 0.37 |
ENSMUST00000103496.4
|
Ighv1-7
|
immunoglobulin heavy variable V1-7 |
| chr11_+_69920542 | 0.37 |
ENSMUST00000232266.2
ENSMUST00000132597.5 |
Dlg4
|
discs large MAGUK scaffold protein 4 |
| chr7_+_30121147 | 0.37 |
ENSMUST00000108176.9
|
Nfkbid
|
nuclear factor of kappa light polypeptide gene enhancer in B cells inhibitor, delta |
| chr14_+_54440591 | 0.35 |
ENSMUST00000103725.2
|
Traj16
|
T cell receptor alpha joining 16 |
| chr3_-_75389047 | 0.34 |
ENSMUST00000193989.4
ENSMUST00000203169.3 |
Wdr49
|
WD repeat domain 49 |
| chr9_+_44238089 | 0.33 |
ENSMUST00000054708.5
|
Dpagt1
|
dolichyl-phosphate (UDP-N-acetylglucosamine) acetylglucosaminephosphotransferase 1 (GlcNAc-1-P transferase) |
| chr1_-_74639723 | 0.33 |
ENSMUST00000127938.8
ENSMUST00000154874.8 |
Rnf25
|
ring finger protein 25 |
| chr18_-_65527078 | 0.32 |
ENSMUST00000035548.16
|
Alpk2
|
alpha-kinase 2 |
| chr5_-_24650164 | 0.31 |
ENSMUST00000115043.8
ENSMUST00000115041.2 ENSMUST00000030800.13 |
Fastk
|
Fas-activated serine/threonine kinase |
| chr12_-_115410489 | 0.30 |
ENSMUST00000194581.2
|
Ighv1-62-2
|
immunoglobulin heavy variable 1-62-2 |
| chr1_+_59296065 | 0.30 |
ENSMUST00000160662.8
ENSMUST00000114248.3 |
Cdk15
|
cyclin-dependent kinase 15 |
| chr9_-_48822399 | 0.29 |
ENSMUST00000003826.8
|
Htr3a
|
5-hydroxytryptamine (serotonin) receptor 3A |
| chr12_-_114487525 | 0.29 |
ENSMUST00000103495.3
|
Ighv10-3
|
immunoglobulin heavy variable V10-3 |
| chr13_-_64645606 | 0.28 |
ENSMUST00000180282.2
|
Fam240b
|
family with sequence similarity 240 member B |
| chr10_-_129951252 | 0.26 |
ENSMUST00000213145.2
|
Olfr823
|
olfactory receptor 823 |
| chr11_+_69917640 | 0.25 |
ENSMUST00000135916.9
ENSMUST00000232659.2 |
Dlg4
|
discs large MAGUK scaffold protein 4 |
| chr5_+_91222470 | 0.25 |
ENSMUST00000031324.6
|
Ereg
|
epiregulin |
| chr7_-_101667346 | 0.25 |
ENSMUST00000209844.2
ENSMUST00000211502.2 ENSMUST00000094134.5 |
Il18bp
|
interleukin 18 binding protein |
| chr9_+_108368032 | 0.25 |
ENSMUST00000166103.9
ENSMUST00000085044.14 ENSMUST00000193678.6 ENSMUST00000178075.8 |
Usp19
|
ubiquitin specific peptidase 19 |
| chr12_-_75463555 | 0.24 |
ENSMUST00000051079.4
|
Gphb5
|
glycoprotein hormone beta 5 |
| chr3_-_59009231 | 0.23 |
ENSMUST00000085040.5
|
Gpr171
|
G protein-coupled receptor 171 |
| chr11_+_76792977 | 0.22 |
ENSMUST00000102495.8
|
Tmigd1
|
transmembrane and immunoglobulin domain containing 1 |
| chrX_+_13499008 | 0.21 |
ENSMUST00000096492.4
|
Gpr34
|
G protein-coupled receptor 34 |
| chr1_+_61017057 | 0.20 |
ENSMUST00000027162.12
ENSMUST00000102827.4 |
Icos
|
inducible T cell co-stimulator |
| chr2_+_67004178 | 0.20 |
ENSMUST00000239009.2
ENSMUST00000238912.2 |
Xirp2
|
xin actin-binding repeat containing 2 |
| chr17_+_35878332 | 0.20 |
ENSMUST00000044326.5
|
2300002M23Rik
|
RIKEN cDNA 2300002M23 gene |
| chr17_-_26204621 | 0.20 |
ENSMUST00000041641.9
ENSMUST00000212520.2 |
Capn15
|
calpain 15 |
| chr11_-_22932090 | 0.19 |
ENSMUST00000160826.2
ENSMUST00000093270.6 ENSMUST00000071068.9 ENSMUST00000159081.8 |
Commd1b
Commd1
|
COMM domain containing 1B COMM domain containing 1 |
| chr13_-_32960379 | 0.19 |
ENSMUST00000230119.2
|
Mylk4
|
myosin light chain kinase family, member 4 |
| chr15_-_98816012 | 0.18 |
ENSMUST00000023736.10
|
Lmbr1l
|
limb region 1 like |
| chr11_-_106679312 | 0.18 |
ENSMUST00000021062.12
|
Ddx5
|
DEAD box helicase 5 |
| chr3_-_37180093 | 0.17 |
ENSMUST00000029275.6
|
Il2
|
interleukin 2 |
| chr6_-_141954471 | 0.17 |
ENSMUST00000111832.3
|
Gm6614
|
predicted gene 6614 |
| chr5_-_35259847 | 0.16 |
ENSMUST00000153664.2
|
Lrpap1
|
low density lipoprotein receptor-related protein associated protein 1 |
| chr16_-_48232770 | 0.16 |
ENSMUST00000212197.2
|
Gm5485
|
predicted gene 5485 |
| chr12_-_114793177 | 0.15 |
ENSMUST00000103511.2
ENSMUST00000195735.2 |
Ighv1-31
|
immunoglobulin heavy variable 1-31 |
| chr7_-_133966588 | 0.10 |
ENSMUST00000172947.8
|
D7Ertd443e
|
DNA segment, Chr 7, ERATO Doi 443, expressed |
| chr17_-_37548054 | 0.07 |
ENSMUST00000213328.2
ENSMUST00000214848.2 ENSMUST00000214622.2 ENSMUST00000073667.6 ENSMUST00000216225.2 |
Olfr97
|
olfactory receptor 97 |
| chr6_-_42453259 | 0.07 |
ENSMUST00000204324.3
ENSMUST00000203396.3 |
Olfr457
|
olfactory receptor 457 |
| chr7_+_34885782 | 0.07 |
ENSMUST00000135452.8
ENSMUST00000001854.12 |
Slc7a10
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 10 |
| chr16_+_24266829 | 0.06 |
ENSMUST00000078988.10
|
Lpp
|
LIM domain containing preferred translocation partner in lipoma |
| chr12_-_114579763 | 0.06 |
ENSMUST00000103500.2
|
Ighv1-12
|
immunoglobulin heavy variable V1-12 |
| chr4_-_88602149 | 0.05 |
ENSMUST00000105147.2
|
Ifna2
|
interferon alpha 2 |
| chr12_+_103281188 | 0.05 |
ENSMUST00000189885.2
ENSMUST00000179363.2 |
Fam181a
|
family with sequence similarity 181, member A |
| chr17_+_34062059 | 0.05 |
ENSMUST00000002379.15
|
Cd320
|
CD320 antigen |
| chr11_-_102837514 | 0.05 |
ENSMUST00000057849.6
|
C1ql1
|
complement component 1, q subcomponent-like 1 |
| chr17_-_32491339 | 0.04 |
ENSMUST00000237008.2
|
Brd4
|
bromodomain containing 4 |
| chr1_+_24216691 | 0.03 |
ENSMUST00000054588.15
|
Col9a1
|
collagen, type IX, alpha 1 |
| chr12_-_115567853 | 0.03 |
ENSMUST00000103538.3
ENSMUST00000198646.2 |
Ighv1-67
|
immunoglobulin heavy variable V1-67 |
| chr6_+_42263644 | 0.02 |
ENSMUST00000163936.8
|
Clcn1
|
chloride channel, voltage-sensitive 1 |
| chr7_-_103814019 | 0.02 |
ENSMUST00000154555.2
ENSMUST00000051137.15 |
Olfm5
|
olfactomedin 5 |
| chr6_+_42263609 | 0.02 |
ENSMUST00000238845.2
ENSMUST00000031894.13 |
Clcn1
|
chloride channel, voltage-sensitive 1 |
| chr5_+_117552042 | 0.01 |
ENSMUST00000180430.2
|
Ksr2
|
kinase suppressor of ras 2 |
| chr8_+_67150055 | 0.01 |
ENSMUST00000039303.7
|
Npy1r
|
neuropeptide Y receptor Y1 |
| chr14_+_53210216 | 0.01 |
ENSMUST00000103599.3
|
Trav3d-3
|
T cell receptor alpha variable 3D-3 |
| chr6_-_71609881 | 0.00 |
ENSMUST00000065509.11
ENSMUST00000207023.2 |
Kdm3a
|
lysine (K)-specific demethylase 3A |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 11.5 | GO:0031536 | positive regulation of exit from mitosis(GO:0031536) |
| 1.1 | 10.1 | GO:0006776 | vitamin A metabolic process(GO:0006776) |
| 1.0 | 3.1 | GO:0006592 | ornithine biosynthetic process(GO:0006592) |
| 0.7 | 13.3 | GO:0006977 | DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest(GO:0006977) |
| 0.6 | 1.9 | GO:0042986 | positive regulation of amyloid precursor protein biosynthetic process(GO:0042986) |
| 0.6 | 2.4 | GO:0060697 | positive regulation of phospholipid catabolic process(GO:0060697) |
| 0.5 | 8.6 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 0.5 | 3.2 | GO:2001205 | negative regulation of osteoclast development(GO:2001205) |
| 0.4 | 1.1 | GO:0046005 | positive regulation of circadian sleep/wake cycle, REM sleep(GO:0046005) |
| 0.4 | 2.9 | GO:0061718 | NADH regeneration(GO:0006735) canonical glycolysis(GO:0061621) glucose catabolic process to pyruvate(GO:0061718) |
| 0.3 | 2.1 | GO:0033030 | negative regulation of neutrophil apoptotic process(GO:0033030) |
| 0.3 | 7.7 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 0.3 | 3.0 | GO:0098909 | regulation of cardiac muscle cell action potential involved in regulation of contraction(GO:0098909) |
| 0.3 | 4.6 | GO:0090084 | negative regulation of inclusion body assembly(GO:0090084) |
| 0.2 | 2.4 | GO:0048318 | axial mesoderm development(GO:0048318) |
| 0.2 | 4.1 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.2 | 1.5 | GO:0035989 | tendon development(GO:0035989) |
| 0.2 | 4.2 | GO:0046835 | carbohydrate phosphorylation(GO:0046835) |
| 0.2 | 2.1 | GO:0019244 | lactate biosynthetic process from pyruvate(GO:0019244) |
| 0.2 | 1.1 | GO:0090383 | platelet dense granule organization(GO:0060155) phagosome acidification(GO:0090383) melanosome assembly(GO:1903232) |
| 0.1 | 2.9 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 0.4 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.1 | 7.9 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.1 | 0.9 | GO:0051715 | cytolysis in other organism(GO:0051715) |
| 0.1 | 1.4 | GO:0099628 | receptor diffusion trapping(GO:0098953) postsynaptic neurotransmitter receptor diffusion trapping(GO:0098970) neurotransmitter receptor diffusion trapping(GO:0099628) |
| 0.1 | 2.1 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.1 | 1.7 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.1 | 1.1 | GO:0043951 | negative regulation of cAMP-mediated signaling(GO:0043951) |
| 0.1 | 1.4 | GO:0006027 | glycosaminoglycan catabolic process(GO:0006027) |
| 0.1 | 3.5 | GO:0061014 | positive regulation of mRNA catabolic process(GO:0061014) |
| 0.1 | 0.9 | GO:0048341 | paraxial mesoderm formation(GO:0048341) |
| 0.1 | 0.7 | GO:1903800 | positive regulation of production of miRNAs involved in gene silencing by miRNA(GO:1903800) |
| 0.1 | 0.8 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.1 | 0.7 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.1 | 1.5 | GO:0035728 | response to hepatocyte growth factor(GO:0035728) |
| 0.1 | 0.2 | GO:1902071 | regulation of hypoxia-inducible factor-1alpha signaling pathway(GO:1902071) |
| 0.1 | 0.6 | GO:2000672 | negative regulation of motor neuron apoptotic process(GO:2000672) |
| 0.1 | 0.5 | GO:0071883 | activation of MAPK activity by adrenergic receptor signaling pathway(GO:0071883) |
| 0.1 | 4.1 | GO:0005978 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.1 | 0.3 | GO:0001550 | ovarian cumulus expansion(GO:0001550) fused antrum stage(GO:0048165) |
| 0.0 | 7.4 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 0.3 | GO:0019348 | dolichol metabolic process(GO:0019348) |
| 0.0 | 1.2 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.0 | 0.4 | GO:0070245 | positive regulation of thymocyte apoptotic process(GO:0070245) |
| 0.0 | 0.4 | GO:0032876 | negative regulation of DNA endoreduplication(GO:0032876) |
| 0.0 | 0.2 | GO:0010916 | regulation of very-low-density lipoprotein particle clearance(GO:0010915) negative regulation of very-low-density lipoprotein particle clearance(GO:0010916) |
| 0.0 | 3.6 | GO:0016126 | sterol biosynthetic process(GO:0016126) |
| 0.0 | 2.2 | GO:0006414 | translational elongation(GO:0006414) |
| 0.0 | 2.6 | GO:0016239 | positive regulation of macroautophagy(GO:0016239) |
| 0.0 | 0.4 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.0 | 0.9 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.0 | 0.5 | GO:0086069 | bundle of His cell to Purkinje myocyte communication(GO:0086069) |
| 0.0 | 0.2 | GO:0046013 | regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.0 | 0.2 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.0 | 0.1 | GO:0015825 | L-serine transport(GO:0015825) |
| 0.0 | 0.7 | GO:2000785 | regulation of autophagosome assembly(GO:2000785) |
| 0.0 | 0.2 | GO:0002155 | thyroid hormone mediated signaling pathway(GO:0002154) regulation of thyroid hormone mediated signaling pathway(GO:0002155) |
| 0.0 | 1.6 | GO:0001541 | ovarian follicle development(GO:0001541) |
| 0.0 | 0.5 | GO:0033141 | positive regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033141) |
| 0.0 | 0.7 | GO:0001921 | positive regulation of receptor recycling(GO:0001921) |
| 0.0 | 1.5 | GO:0006367 | transcription initiation from RNA polymerase II promoter(GO:0006367) |
| 0.0 | 0.2 | GO:1900037 | regulation of cellular response to hypoxia(GO:1900037) |
| 0.0 | 1.5 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.9 | 11.5 | GO:0031021 | interphase microtubule organizing center(GO:0031021) |
| 1.3 | 7.7 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 0.8 | 2.4 | GO:0070992 | translation initiation complex(GO:0070992) |
| 0.3 | 2.2 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.2 | 2.4 | GO:0034363 | intermediate-density lipoprotein particle(GO:0034363) |
| 0.2 | 2.1 | GO:0042629 | mast cell granule(GO:0042629) |
| 0.2 | 2.6 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.2 | 3.6 | GO:0005639 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.1 | 0.7 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.1 | 0.8 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.1 | 1.1 | GO:0033162 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.1 | 0.6 | GO:0097058 | CRLF-CLCF1 complex(GO:0097058) |
| 0.1 | 1.5 | GO:0098643 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.1 | 0.3 | GO:1904602 | serotonin-activated cation-selective channel complex(GO:1904602) |
| 0.1 | 1.2 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.1 | 1.6 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.1 | 5.0 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 1.4 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.1 | 0.7 | GO:0071141 | SMAD protein complex(GO:0071141) |
| 0.1 | 0.9 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.1 | 1.4 | GO:0033276 | transcription factor TFTC complex(GO:0033276) |
| 0.1 | 7.4 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.1 | 1.0 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.0 | 0.0 | GO:0044301 | climbing fiber(GO:0044301) |
| 0.0 | 2.9 | GO:0060170 | ciliary membrane(GO:0060170) |
| 0.0 | 1.5 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.0 | 4.1 | GO:0016234 | inclusion body(GO:0016234) |
| 0.0 | 0.5 | GO:0001527 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.0 | 0.9 | GO:0044216 | other organism(GO:0044215) other organism cell(GO:0044216) other organism part(GO:0044217) |
| 0.0 | 3.9 | GO:0036126 | sperm flagellum(GO:0036126) |
| 0.0 | 2.9 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 3.2 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.0 | 39.9 | GO:0031410 | cytoplasmic vesicle(GO:0031410) |
| 0.0 | 2.7 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 1.2 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 6.7 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 1.9 | GO:0005923 | bicellular tight junction(GO:0005923) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 4.1 | GO:0004373 | glycogen (starch) synthase activity(GO:0004373) |
| 1.3 | 7.7 | GO:0004748 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 1.1 | 7.9 | GO:0052798 | beta-galactoside alpha-2,3-sialyltransferase activity(GO:0052798) |
| 1.0 | 3.1 | GO:0017084 | glutamate 5-kinase activity(GO:0004349) glutamate-5-semialdehyde dehydrogenase activity(GO:0004350) delta1-pyrroline-5-carboxylate synthetase activity(GO:0017084) amino acid kinase activity(GO:0019202) |
| 0.7 | 2.1 | GO:0019959 | interleukin-8 binding(GO:0019959) |
| 0.6 | 10.1 | GO:0019841 | retinol binding(GO:0019841) |
| 0.5 | 4.2 | GO:0008865 | fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.4 | 2.4 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 0.4 | 1.1 | GO:0036461 | BLOC-2 complex binding(GO:0036461) |
| 0.3 | 3.6 | GO:0070087 | chromo shadow domain binding(GO:0070087) |
| 0.3 | 2.2 | GO:0047179 | platelet-activating factor acetyltransferase activity(GO:0047179) |
| 0.3 | 2.9 | GO:0016861 | intramolecular oxidoreductase activity, interconverting aldoses and ketoses(GO:0016861) |
| 0.3 | 14.5 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.2 | 0.7 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.2 | 0.7 | GO:0035500 | MH2 domain binding(GO:0035500) |
| 0.2 | 1.4 | GO:0031812 | G-protein coupled nucleotide receptor binding(GO:0031811) P2Y1 nucleotide receptor binding(GO:0031812) |
| 0.2 | 2.1 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.2 | 2.1 | GO:0031802 | type 5 metabotropic glutamate receptor binding(GO:0031802) |
| 0.1 | 8.6 | GO:0070888 | bHLH transcription factor binding(GO:0043425) E-box binding(GO:0070888) |
| 0.1 | 13.3 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.1 | 1.4 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.1 | 2.2 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.1 | 0.3 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 0.1 | 0.8 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.1 | 0.2 | GO:0031531 | thyrotropin-releasing hormone receptor binding(GO:0031531) |
| 0.1 | 0.9 | GO:0017151 | DEAD/H-box RNA helicase binding(GO:0017151) |
| 0.1 | 0.5 | GO:0031013 | troponin I binding(GO:0031013) |
| 0.1 | 1.9 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 0.9 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 0.1 | 1.6 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.1 | 1.2 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.1 | 0.3 | GO:0022850 | serotonin-gated cation channel activity(GO:0022850) |
| 0.1 | 7.4 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.1 | 0.6 | GO:0005127 | ciliary neurotrophic factor receptor binding(GO:0005127) |
| 0.1 | 0.4 | GO:0048019 | receptor antagonist activity(GO:0048019) |
| 0.0 | 0.5 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.0 | 3.3 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.0 | 0.7 | GO:0043008 | ATP-dependent protein binding(GO:0043008) |
| 0.0 | 0.2 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.0 | 1.7 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.0 | 1.5 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 4.6 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 0.9 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.0 | 0.4 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 0.0 | 1.9 | GO:0008374 | O-acyltransferase activity(GO:0008374) |
| 0.0 | 0.5 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.0 | 0.2 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.0 | 0.2 | GO:1990380 | Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.0 | 0.7 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.0 | 2.4 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 1.5 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.5 | GO:0008198 | ferrous iron binding(GO:0008198) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 11.5 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 1.9 | ST JAK STAT PATHWAY | Jak-STAT Pathway |
| 0.1 | 4.1 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 7.7 | PID E2F PATHWAY | E2F transcription factor network |
| 0.1 | 9.3 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.1 | 2.3 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.1 | 3.3 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 2.1 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.0 | 1.6 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.0 | 1.4 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.0 | 1.5 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.0 | 1.5 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 0.4 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.0 | 0.3 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 0.9 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 0.7 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 7.7 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.3 | 7.9 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.2 | 9.3 | REACTOME METABOLISM OF STEROID HORMONES AND VITAMINS A AND D | Genes involved in Metabolism of steroid hormones and vitamins A and D |
| 0.1 | 1.7 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.1 | 2.4 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.1 | 3.6 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 3.1 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.1 | 11.9 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.1 | 2.1 | REACTOME PYRUVATE METABOLISM | Genes involved in Pyruvate metabolism |
| 0.1 | 3.2 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.1 | 5.0 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.1 | 1.6 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 1.5 | REACTOME SIGNALING BY CONSTITUTIVELY ACTIVE EGFR | Genes involved in Signaling by constitutively active EGFR |
| 0.0 | 2.1 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.9 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 2.4 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 0.7 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.0 | 2.2 | REACTOME TRANSLATION | Genes involved in Translation |
| 0.0 | 1.4 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.0 | 2.6 | REACTOME GLUCOSE METABOLISM | Genes involved in Glucose metabolism |
| 0.0 | 1.5 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.5 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.0 | 0.4 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |