Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
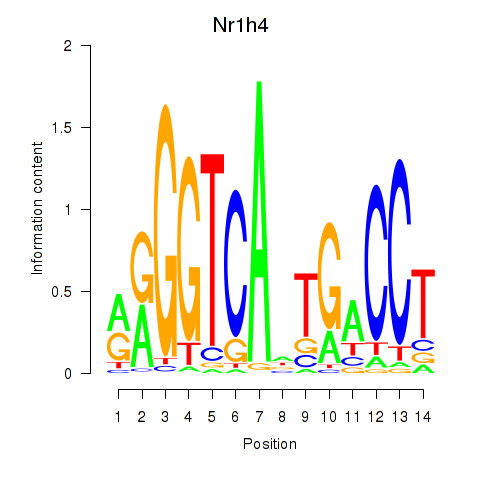
Results for Nr1h4
Z-value: 2.85
Transcription factors associated with Nr1h4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Nr1h4
|
ENSMUSG00000047638.16 | Nr1h4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Nr1h4 | mm39_v1_chr10_-_89369432_89369452 | 0.88 | 7.9e-13 | Click! |
Activity profile of Nr1h4 motif
Sorted Z-values of Nr1h4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Nr1h4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_-_93806593 | 26.20 |
ENSMUST00000109582.3
|
Ces1b
|
carboxylesterase 1B |
| chr7_-_105249308 | 19.15 |
ENSMUST00000210531.2
ENSMUST00000033185.10 |
Hpx
|
hemopexin |
| chr2_-_69172944 | 17.26 |
ENSMUST00000102709.8
ENSMUST00000102710.10 ENSMUST00000180142.2 |
Abcb11
|
ATP-binding cassette, sub-family B (MDR/TAP), member 11 |
| chr17_-_32639936 | 15.15 |
ENSMUST00000170392.9
ENSMUST00000237165.2 ENSMUST00000235892.2 ENSMUST00000114455.3 |
Pglyrp2
|
peptidoglycan recognition protein 2 |
| chr9_-_65330231 | 14.92 |
ENSMUST00000065894.7
|
Slc51b
|
solute carrier family 51, beta subunit |
| chr3_+_137983250 | 14.64 |
ENSMUST00000004232.10
|
Adh1
|
alcohol dehydrogenase 1 (class I) |
| chr4_+_133280680 | 13.98 |
ENSMUST00000042706.3
|
Nr0b2
|
nuclear receptor subfamily 0, group B, member 2 |
| chr7_-_99345016 | 13.85 |
ENSMUST00000107086.9
|
Slco2b1
|
solute carrier organic anion transporter family, member 2b1 |
| chr1_+_67162176 | 13.59 |
ENSMUST00000027144.8
|
Cps1
|
carbamoyl-phosphate synthetase 1 |
| chr6_+_141575226 | 13.14 |
ENSMUST00000042812.9
|
Slco1b2
|
solute carrier organic anion transporter family, member 1b2 |
| chr15_-_82648376 | 13.10 |
ENSMUST00000055721.6
|
Cyp2d40
|
cytochrome P450, family 2, subfamily d, polypeptide 40 |
| chr9_+_46179899 | 13.02 |
ENSMUST00000121598.8
|
Apoa5
|
apolipoprotein A-V |
| chr9_+_108539296 | 11.29 |
ENSMUST00000035222.6
|
Slc25a20
|
solute carrier family 25 (mitochondrial carnitine/acylcarnitine translocase), member 20 |
| chr11_-_116089595 | 11.11 |
ENSMUST00000072948.11
|
Acox1
|
acyl-Coenzyme A oxidase 1, palmitoyl |
| chr11_-_116089866 | 11.03 |
ENSMUST00000066587.12
|
Acox1
|
acyl-Coenzyme A oxidase 1, palmitoyl |
| chr19_+_38995463 | 10.76 |
ENSMUST00000025966.5
|
Cyp2c55
|
cytochrome P450, family 2, subfamily c, polypeptide 55 |
| chr11_+_72326337 | 9.97 |
ENSMUST00000076443.10
|
Ggt6
|
gamma-glutamyltransferase 6 |
| chr17_-_35081129 | 8.50 |
ENSMUST00000154526.8
|
Cfb
|
complement factor B |
| chr17_-_57535003 | 8.45 |
ENSMUST00000177046.2
ENSMUST00000024988.15 |
C3
|
complement component 3 |
| chr17_-_35081456 | 8.20 |
ENSMUST00000025229.11
ENSMUST00000176203.9 ENSMUST00000128767.8 |
Cfb
|
complement factor B |
| chr11_+_72326391 | 7.62 |
ENSMUST00000100903.3
|
Ggt6
|
gamma-glutamyltransferase 6 |
| chrX_-_51254129 | 7.41 |
ENSMUST00000033450.3
|
Gpc4
|
glypican 4 |
| chr11_+_72326358 | 6.91 |
ENSMUST00000108499.2
|
Ggt6
|
gamma-glutamyltransferase 6 |
| chr14_-_30645711 | 6.79 |
ENSMUST00000006697.17
|
Itih3
|
inter-alpha trypsin inhibitor, heavy chain 3 |
| chr9_-_21900724 | 6.61 |
ENSMUST00000045726.8
|
Rgl3
|
ral guanine nucleotide dissociation stimulator-like 3 |
| chr17_-_84154173 | 6.31 |
ENSMUST00000000687.9
|
Haao
|
3-hydroxyanthranilate 3,4-dioxygenase |
| chr7_-_44320244 | 6.29 |
ENSMUST00000048102.15
|
Myh14
|
myosin, heavy polypeptide 14 |
| chr9_+_107957621 | 6.26 |
ENSMUST00000035211.14
|
Mst1
|
macrophage stimulating 1 (hepatocyte growth factor-like) |
| chr1_+_163979384 | 6.21 |
ENSMUST00000086040.6
|
F5
|
coagulation factor V |
| chr9_+_107957640 | 6.12 |
ENSMUST00000162886.2
|
Mst1
|
macrophage stimulating 1 (hepatocyte growth factor-like) |
| chr7_-_30755007 | 6.10 |
ENSMUST00000206474.2
ENSMUST00000205807.2 ENSMUST00000039909.13 ENSMUST00000206305.2 ENSMUST00000205439.2 |
Fxyd1
|
FXYD domain-containing ion transport regulator 1 |
| chr11_+_97576619 | 5.95 |
ENSMUST00000107584.8
ENSMUST00000107585.9 |
Cisd3
|
CDGSH iron sulfur domain 3 |
| chr14_-_30645503 | 5.26 |
ENSMUST00000227995.2
|
Itih3
|
inter-alpha trypsin inhibitor, heavy chain 3 |
| chr16_-_22847808 | 5.17 |
ENSMUST00000115349.9
|
Kng2
|
kininogen 2 |
| chr11_+_97576724 | 5.12 |
ENSMUST00000107583.3
|
Cisd3
|
CDGSH iron sulfur domain 3 |
| chr16_-_22847760 | 5.10 |
ENSMUST00000039338.13
|
Kng2
|
kininogen 2 |
| chr13_+_25127127 | 5.08 |
ENSMUST00000021773.13
|
Gpld1
|
glycosylphosphatidylinositol specific phospholipase D1 |
| chr16_-_22847829 | 4.98 |
ENSMUST00000100046.9
|
Kng2
|
kininogen 2 |
| chr17_-_84154196 | 4.90 |
ENSMUST00000234214.2
|
Haao
|
3-hydroxyanthranilate 3,4-dioxygenase |
| chr8_+_13110921 | 4.79 |
ENSMUST00000211363.2
ENSMUST00000033822.4 |
Proz
|
protein Z, vitamin K-dependent plasma glycoprotein |
| chr4_-_141345549 | 4.76 |
ENSMUST00000053263.9
|
Tmem82
|
transmembrane protein 82 |
| chr13_+_91889626 | 4.73 |
ENSMUST00000022120.5
|
Acot12
|
acyl-CoA thioesterase 12 |
| chr7_-_126944578 | 4.69 |
ENSMUST00000060783.7
|
Zfp768
|
zinc finger protein 768 |
| chr17_-_79328157 | 4.57 |
ENSMUST00000168887.8
ENSMUST00000119284.8 |
Prkd3
|
protein kinase D3 |
| chr10_+_23770586 | 4.49 |
ENSMUST00000041416.8
|
Vnn1
|
vanin 1 |
| chr19_+_44980565 | 4.46 |
ENSMUST00000179305.2
|
Sema4g
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4G |
| chr5_+_137568982 | 4.41 |
ENSMUST00000196471.5
ENSMUST00000198783.5 |
Tfr2
|
transferrin receptor 2 |
| chr9_-_106353792 | 4.37 |
ENSMUST00000214682.2
ENSMUST00000112479.9 |
Parp3
|
poly (ADP-ribose) polymerase family, member 3 |
| chr10_-_80934708 | 4.35 |
ENSMUST00000117422.2
|
Creb3l3
|
cAMP responsive element binding protein 3-like 3 |
| chr15_-_82678490 | 4.25 |
ENSMUST00000006094.6
|
Cyp2d26
|
cytochrome P450, family 2, subfamily d, polypeptide 26 |
| chr19_-_38113056 | 4.17 |
ENSMUST00000236283.2
|
Rbp4
|
retinol binding protein 4, plasma |
| chr7_-_30754223 | 4.13 |
ENSMUST00000206012.2
ENSMUST00000108110.5 |
Fxyd1
|
FXYD domain-containing ion transport regulator 1 |
| chr11_+_101442961 | 4.07 |
ENSMUST00000103099.8
|
Nbr1
|
NBR1, autophagy cargo receptor |
| chr1_-_162687254 | 3.96 |
ENSMUST00000131058.8
|
Fmo1
|
flavin containing monooxygenase 1 |
| chr7_-_30754240 | 3.95 |
ENSMUST00000206860.2
ENSMUST00000071697.11 |
Fxyd1
|
FXYD domain-containing ion transport regulator 1 |
| chr17_-_87573294 | 3.91 |
ENSMUST00000145895.8
ENSMUST00000129616.8 ENSMUST00000155904.2 ENSMUST00000151155.8 ENSMUST00000144236.9 ENSMUST00000024963.11 |
Mcfd2
|
multiple coagulation factor deficiency 2 |
| chr7_-_30754193 | 3.87 |
ENSMUST00000205778.2
|
Fxyd1
|
FXYD domain-containing ion transport regulator 1 |
| chr10_+_116137277 | 3.85 |
ENSMUST00000092167.7
|
Ptprb
|
protein tyrosine phosphatase, receptor type, B |
| chr16_-_22848153 | 3.79 |
ENSMUST00000232459.2
|
Kng2
|
kininogen 2 |
| chr1_-_162687369 | 3.66 |
ENSMUST00000193078.6
|
Fmo1
|
flavin containing monooxygenase 1 |
| chr1_+_171052623 | 3.61 |
ENSMUST00000111321.8
ENSMUST00000005824.12 ENSMUST00000111320.8 ENSMUST00000111319.2 |
Apoa2
|
apolipoprotein A-II |
| chr10_-_78134026 | 3.60 |
ENSMUST00000105389.8
|
Agpat3
|
1-acylglycerol-3-phosphate O-acyltransferase 3 |
| chr3_+_121761471 | 3.31 |
ENSMUST00000196479.5
ENSMUST00000197155.5 |
Arhgap29
|
Rho GTPase activating protein 29 |
| chr11_-_100595019 | 3.29 |
ENSMUST00000017974.13
|
Dhx58
|
DEXH (Asp-Glu-X-His) box polypeptide 58 |
| chr17_+_25352353 | 3.26 |
ENSMUST00000162862.3
ENSMUST00000040729.9 |
Clcn7
|
chloride channel, voltage-sensitive 7 |
| chr17_-_25459086 | 3.19 |
ENSMUST00000038973.7
ENSMUST00000115154.11 |
Gnptg
|
N-acetylglucosamine-1-phosphotransferase, gamma subunit |
| chr2_-_174305856 | 3.10 |
ENSMUST00000016396.8
|
Atp5e
|
ATP synthase, H+ transporting, mitochondrial F1 complex, epsilon subunit |
| chr13_+_41013230 | 3.05 |
ENSMUST00000110191.10
|
Gcnt2
|
glucosaminyl (N-acetyl) transferase 2, I-branching enzyme |
| chr10_+_43777777 | 2.99 |
ENSMUST00000054418.12
|
Rtn4ip1
|
reticulon 4 interacting protein 1 |
| chr11_+_101443014 | 2.94 |
ENSMUST00000147239.8
|
Nbr1
|
NBR1, autophagy cargo receptor |
| chr11_+_66915969 | 2.83 |
ENSMUST00000079077.12
ENSMUST00000061786.6 |
Tmem220
|
transmembrane protein 220 |
| chr3_-_120965327 | 2.79 |
ENSMUST00000170781.2
ENSMUST00000039761.12 ENSMUST00000106467.8 ENSMUST00000106466.10 ENSMUST00000164925.9 |
Rwdd3
|
RWD domain containing 3 |
| chr9_+_44018583 | 2.74 |
ENSMUST00000152956.8
ENSMUST00000114815.3 ENSMUST00000206295.2 ENSMUST00000206769.2 ENSMUST00000205500.2 |
C1qtnf5
|
C1q and tumor necrosis factor related protein 5 |
| chr15_-_31367872 | 2.69 |
ENSMUST00000123325.9
|
Ankrd33b
|
ankyrin repeat domain 33B |
| chr13_+_41040657 | 2.68 |
ENSMUST00000069958.15
|
Gcnt2
|
glucosaminyl (N-acetyl) transferase 2, I-branching enzyme |
| chr9_+_21914083 | 2.66 |
ENSMUST00000216344.2
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr7_+_18725170 | 2.63 |
ENSMUST00000059331.9
ENSMUST00000131087.2 |
Mypop
|
Myb-related transcription factor, partner of profilin |
| chr15_-_31367668 | 2.59 |
ENSMUST00000110410.10
ENSMUST00000076942.5 |
Ankrd33b
|
ankyrin repeat domain 33B |
| chr17_-_74354844 | 2.56 |
ENSMUST00000043458.9
|
Srd5a2
|
steroid 5 alpha-reductase 2 |
| chr11_+_70410445 | 2.55 |
ENSMUST00000179000.2
|
Gltpd2
|
glycolipid transfer protein domain containing 2 |
| chr15_+_76264695 | 2.41 |
ENSMUST00000096385.11
ENSMUST00000160728.8 |
Mroh1
|
maestro heat-like repeat family member 1 |
| chr9_-_21913833 | 2.38 |
ENSMUST00000115336.10
|
Odad3
|
outer dynein arm docking complex subunit 3 |
| chr9_+_44018551 | 2.37 |
ENSMUST00000114821.9
ENSMUST00000114818.9 |
C1qtnf5
|
C1q and tumor necrosis factor related protein 5 |
| chr11_+_101443674 | 2.32 |
ENSMUST00000107213.8
ENSMUST00000107208.8 ENSMUST00000107212.8 ENSMUST00000127421.8 |
Nbr1
|
NBR1, autophagy cargo receptor |
| chr9_+_21279802 | 2.23 |
ENSMUST00000214474.2
|
Ilf3
|
interleukin enhancer binding factor 3 |
| chr15_+_102010632 | 2.23 |
ENSMUST00000229592.2
|
Tns2
|
tensin 2 |
| chr10_-_67384898 | 2.22 |
ENSMUST00000075686.7
|
Ado
|
2-aminoethanethiol (cysteamine) dioxygenase |
| chr9_-_59260713 | 2.20 |
ENSMUST00000026265.8
|
Bbs4
|
Bardet-Biedl syndrome 4 (human) |
| chr8_+_89247976 | 2.17 |
ENSMUST00000034086.13
|
Nkd1
|
naked cuticle 1 |
| chr9_+_21914334 | 2.17 |
ENSMUST00000115331.10
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr16_-_20549294 | 2.13 |
ENSMUST00000231826.2
ENSMUST00000076422.13 ENSMUST00000232217.2 |
Thpo
|
thrombopoietin |
| chr9_-_21913896 | 2.11 |
ENSMUST00000044926.6
|
Odad3
|
outer dynein arm docking complex subunit 3 |
| chr11_-_120715351 | 2.07 |
ENSMUST00000055655.9
|
Fasn
|
fatty acid synthase |
| chr2_-_130480014 | 2.05 |
ENSMUST00000089561.10
ENSMUST00000110260.8 |
Lzts3
|
leucine zipper, putative tumor suppressor family member 3 |
| chr3_+_154302311 | 2.00 |
ENSMUST00000192462.6
ENSMUST00000029850.15 |
Cryz
|
crystallin, zeta |
| chr1_-_162641495 | 1.98 |
ENSMUST00000144916.8
ENSMUST00000140274.2 |
Fmo4
|
flavin containing monooxygenase 4 |
| chr15_+_80556023 | 1.94 |
ENSMUST00000023044.7
|
Fam83f
|
family with sequence similarity 83, member F |
| chr9_+_21914296 | 1.91 |
ENSMUST00000003493.9
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr10_+_4561974 | 1.91 |
ENSMUST00000105590.8
ENSMUST00000067086.14 |
Esr1
|
estrogen receptor 1 (alpha) |
| chrM_+_8603 | 1.77 |
ENSMUST00000082409.1
|
mt-Co3
|
mitochondrially encoded cytochrome c oxidase III |
| chr9_+_56902172 | 1.65 |
ENSMUST00000034832.8
|
Ptpn9
|
protein tyrosine phosphatase, non-receptor type 9 |
| chr6_+_72391283 | 1.64 |
ENSMUST00000065906.9
|
Ggcx
|
gamma-glutamyl carboxylase |
| chr19_-_6919755 | 1.62 |
ENSMUST00000099782.10
|
Gpr137
|
G protein-coupled receptor 137 |
| chr5_-_147662798 | 1.57 |
ENSMUST00000110529.6
ENSMUST00000031653.12 |
Flt1
|
FMS-like tyrosine kinase 1 |
| chr9_-_63285080 | 1.56 |
ENSMUST00000034920.11
|
Map2k5
|
mitogen-activated protein kinase kinase 5 |
| chrX_-_137985960 | 1.48 |
ENSMUST00000033626.15
ENSMUST00000060824.4 |
Serpina7
|
serine (or cysteine) peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr12_+_76593799 | 1.45 |
ENSMUST00000218380.2
ENSMUST00000219751.2 |
Plekhg3
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 3 |
| chr9_-_106353303 | 1.39 |
ENSMUST00000156426.8
|
Parp3
|
poly (ADP-ribose) polymerase family, member 3 |
| chr9_+_44018520 | 1.39 |
ENSMUST00000114816.8
|
C1qtnf5
|
C1q and tumor necrosis factor related protein 5 |
| chr7_-_45362867 | 1.38 |
ENSMUST00000211340.2
|
Sphk2
|
sphingosine kinase 2 |
| chr2_-_90748331 | 1.35 |
ENSMUST00000111461.13
|
Ptpmt1
|
protein tyrosine phosphatase, mitochondrial 1 |
| chrX_+_20714782 | 1.33 |
ENSMUST00000001155.11
ENSMUST00000122312.8 ENSMUST00000120356.8 ENSMUST00000122850.2 |
Araf
|
Araf proto-oncogene, serine/threonine kinase |
| chr8_+_85807369 | 1.28 |
ENSMUST00000079764.14
|
Wdr83os
|
WD repeat domain 83 opposite strand |
| chr7_+_138968988 | 1.25 |
ENSMUST00000106098.8
ENSMUST00000026550.14 |
Inpp5a
|
inositol polyphosphate-5-phosphatase A |
| chr4_-_133360749 | 1.24 |
ENSMUST00000084238.5
|
Zdhhc18
|
zinc finger, DHHC domain containing 18 |
| chr1_-_43235914 | 1.22 |
ENSMUST00000187357.2
|
Fhl2
|
four and a half LIM domains 2 |
| chr4_-_135699885 | 1.21 |
ENSMUST00000067567.5
|
Lypla2
|
lysophospholipase 2 |
| chr10_-_89568106 | 1.19 |
ENSMUST00000020109.5
|
Actr6
|
ARP6 actin-related protein 6 |
| chrX_-_137985979 | 1.18 |
ENSMUST00000152457.2
|
Serpina7
|
serine (or cysteine) peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr9_+_21914513 | 1.15 |
ENSMUST00000215795.2
|
Prkcsh
|
protein kinase C substrate 80K-H |
| chr13_-_39144475 | 1.13 |
ENSMUST00000225331.2
ENSMUST00000224645.2 ENSMUST00000167513.3 ENSMUST00000225568.2 |
Slc35b3
|
solute carrier family 35, member B3 |
| chr6_-_29380423 | 1.10 |
ENSMUST00000147483.3
|
Opn1sw
|
opsin 1 (cone pigments), short-wave-sensitive (color blindness, tritan) |
| chr15_-_77811935 | 1.08 |
ENSMUST00000174529.2
ENSMUST00000173631.8 |
Txn2
|
thioredoxin 2 |
| chr9_-_20791012 | 1.07 |
ENSMUST00000043726.8
|
Angptl6
|
angiopoietin-like 6 |
| chr17_+_25459138 | 1.07 |
ENSMUST00000063574.7
|
Tsr3
|
TSR3 20S rRNA accumulation |
| chr9_-_50663648 | 1.06 |
ENSMUST00000217159.2
|
Hspb2
|
heat shock protein 2 |
| chr12_-_16660960 | 1.01 |
ENSMUST00000239165.2
ENSMUST00000111067.10 |
Lpin1
|
lipin 1 |
| chr14_-_31552335 | 1.01 |
ENSMUST00000228037.2
|
Ankrd28
|
ankyrin repeat domain 28 |
| chr12_-_103597663 | 0.97 |
ENSMUST00000121625.2
ENSMUST00000044231.12 |
Serpina10
|
serine (or cysteine) peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 10 |
| chr5_-_31359276 | 0.97 |
ENSMUST00000031562.11
|
Zfp513
|
zinc finger protein 513 |
| chrX_-_18327610 | 0.94 |
ENSMUST00000044188.5
|
Dipk2b
|
divergent protein kinase domain 2B |
| chr15_-_103231921 | 0.91 |
ENSMUST00000229551.2
|
Zfp385a
|
zinc finger protein 385A |
| chrX_-_138683102 | 0.89 |
ENSMUST00000101217.4
|
Ripply1
|
ripply transcriptional repressor 1 |
| chr6_-_86770504 | 0.88 |
ENSMUST00000204441.3
ENSMUST00000204398.2 ENSMUST00000001187.15 |
Anxa4
|
annexin A4 |
| chr7_+_18910340 | 0.87 |
ENSMUST00000117338.8
|
Eml2
|
echinoderm microtubule associated protein like 2 |
| chr1_-_171074652 | 0.85 |
ENSMUST00000013737.13
ENSMUST00000111318.8 |
Ndufs2
|
NADH:ubiquinone oxidoreductase core subunit S2 |
| chr3_-_107424637 | 0.83 |
ENSMUST00000166892.2
|
Slc6a17
|
solute carrier family 6 (neurotransmitter transporter), member 17 |
| chr11_-_63970270 | 0.82 |
ENSMUST00000049091.9
|
Cox10
|
heme A:farnesyltransferase cytochrome c oxidase assembly factor 10 |
| chr1_-_63215952 | 0.81 |
ENSMUST00000185412.7
ENSMUST00000027111.15 ENSMUST00000189664.2 |
Ndufs1
|
NADH:ubiquinone oxidoreductase core subunit S1 |
| chr7_-_4974167 | 0.80 |
ENSMUST00000133272.2
|
Sbk3
|
SH3 domain binding kinase family, member 3 |
| chr11_+_75404356 | 0.79 |
ENSMUST00000042808.13
ENSMUST00000118243.2 |
Scarf1
|
scavenger receptor class F, member 1 |
| chr7_+_16044336 | 0.78 |
ENSMUST00000136781.2
|
Bbc3
|
BCL2 binding component 3 |
| chr10_-_23226034 | 0.77 |
ENSMUST00000219315.2
|
Eya4
|
EYA transcriptional coactivator and phosphatase 4 |
| chr6_-_24528012 | 0.77 |
ENSMUST00000023851.9
|
Ndufa5
|
NADH:ubiquinone oxidoreductase subunit A5 |
| chr11_+_5905693 | 0.71 |
ENSMUST00000002818.9
|
Ykt6
|
YKT6 v-SNARE homolog (S. cerevisiae) |
| chr3_+_137329433 | 0.70 |
ENSMUST00000053855.8
|
Ddit4l
|
DNA-damage-inducible transcript 4-like |
| chr16_-_94171556 | 0.69 |
ENSMUST00000113906.9
|
Pigp
|
phosphatidylinositol glycan anchor biosynthesis, class P |
| chr2_-_120183575 | 0.68 |
ENSMUST00000028752.8
ENSMUST00000102501.10 |
Vps39
|
VPS39 HOPS complex subunit |
| chr13_-_40882417 | 0.65 |
ENSMUST00000225180.2
|
Tfap2a
|
transcription factor AP-2, alpha |
| chr5_-_129916283 | 0.63 |
ENSMUST00000094280.4
|
Chchd2
|
coiled-coil-helix-coiled-coil-helix domain containing 2 |
| chr7_+_44091822 | 0.63 |
ENSMUST00000058667.15
|
Lrrc4b
|
leucine rich repeat containing 4B |
| chr2_+_32511762 | 0.62 |
ENSMUST00000113278.9
|
Ak1
|
adenylate kinase 1 |
| chr2_+_83791847 | 0.61 |
ENSMUST00000178960.2
|
Gm13693
|
predicted gene 13693 |
| chr7_-_4973960 | 0.61 |
ENSMUST00000144863.8
|
Sbk3
|
SH3 domain binding kinase family, member 3 |
| chrX_+_76554608 | 0.60 |
ENSMUST00000088217.12
|
Tbl1x
|
transducin (beta)-like 1 X-linked |
| chrX_-_101687813 | 0.59 |
ENSMUST00000052012.14
ENSMUST00000043596.12 ENSMUST00000119229.8 ENSMUST00000122022.8 ENSMUST00000120270.8 ENSMUST00000113611.3 |
Phka1
|
phosphorylase kinase alpha 1 |
| chr17_+_24888641 | 0.59 |
ENSMUST00000234956.2
|
Zfp598
|
zinc finger protein 598 |
| chr2_+_128907854 | 0.58 |
ENSMUST00000035812.14
|
Ttl
|
tubulin tyrosine ligase |
| chr17_+_24888704 | 0.58 |
ENSMUST00000047179.7
|
Zfp598
|
zinc finger protein 598 |
| chr1_+_36550948 | 0.56 |
ENSMUST00000001166.14
ENSMUST00000097776.4 |
Cnnm3
|
cyclin M3 |
| chr15_+_101371353 | 0.53 |
ENSMUST00000088049.5
|
Krt86
|
keratin 86 |
| chr6_-_129600798 | 0.53 |
ENSMUST00000095412.10
|
Klrk1
|
killer cell lectin-like receptor subfamily K, member 1 |
| chr9_-_106533279 | 0.52 |
ENSMUST00000023959.13
ENSMUST00000201681.2 |
Grm2
|
glutamate receptor, metabotropic 2 |
| chr1_+_152830720 | 0.52 |
ENSMUST00000043313.15
ENSMUST00000186621.2 |
Nmnat2
|
nicotinamide nucleotide adenylyltransferase 2 |
| chr6_-_41752111 | 0.52 |
ENSMUST00000214976.3
|
Olfr459
|
olfactory receptor 459 |
| chr6_-_29380467 | 0.50 |
ENSMUST00000080428.13
|
Opn1sw
|
opsin 1 (cone pigments), short-wave-sensitive (color blindness, tritan) |
| chr5_-_31359559 | 0.50 |
ENSMUST00000202929.2
ENSMUST00000201231.2 ENSMUST00000114590.8 |
Zfp513
|
zinc finger protein 513 |
| chr4_+_42917228 | 0.48 |
ENSMUST00000107976.9
ENSMUST00000069184.9 |
Phf24
|
PHD finger protein 24 |
| chr15_-_90934021 | 0.48 |
ENSMUST00000109287.4
ENSMUST00000067205.16 ENSMUST00000088614.13 |
Kif21a
|
kinesin family member 21A |
| chr15_-_90934059 | 0.46 |
ENSMUST00000109288.9
ENSMUST00000100304.11 |
Kif21a
|
kinesin family member 21A |
| chr15_-_98465516 | 0.44 |
ENSMUST00000012104.7
|
Ccnt1
|
cyclin T1 |
| chr12_-_87346479 | 0.42 |
ENSMUST00000125733.8
|
Ism2
|
isthmin 2 |
| chr6_+_24528143 | 0.42 |
ENSMUST00000031696.10
|
Asb15
|
ankyrin repeat and SOCS box-containing 15 |
| chr4_-_135699205 | 0.41 |
ENSMUST00000105852.8
|
Lypla2
|
lysophospholipase 2 |
| chr1_+_92934050 | 0.41 |
ENSMUST00000059676.5
|
Aqp12
|
aquaporin 12 |
| chr1_-_63215812 | 0.40 |
ENSMUST00000185847.2
ENSMUST00000185732.7 ENSMUST00000188370.7 ENSMUST00000168099.9 |
Ndufs1
|
NADH:ubiquinone oxidoreductase core subunit S1 |
| chr3_+_137329791 | 0.40 |
ENSMUST00000165845.2
|
Ddit4l
|
DNA-damage-inducible transcript 4-like |
| chr9_+_50664207 | 0.39 |
ENSMUST00000034562.9
|
Cryab
|
crystallin, alpha B |
| chr17_-_5490504 | 0.39 |
ENSMUST00000005053.14
ENSMUST00000185896.2 ENSMUST00000188282.7 |
Tmem242
|
transmembrane protein 242 |
| chr9_+_50664288 | 0.39 |
ENSMUST00000214962.2
ENSMUST00000216755.2 |
Cryab
|
crystallin, alpha B |
| chr2_+_177404691 | 0.38 |
ENSMUST00000119838.9
|
Gm14322
|
predicted gene 14322 |
| chr7_+_128213084 | 0.37 |
ENSMUST00000043138.13
|
Inpp5f
|
inositol polyphosphate-5-phosphatase F |
| chr2_-_152793376 | 0.37 |
ENSMUST00000123121.9
|
Dusp15
|
dual specificity phosphatase-like 15 |
| chr8_+_93400916 | 0.35 |
ENSMUST00000034185.13
|
Irx6
|
Iroquois homeobox 6 |
| chr12_+_105672227 | 0.33 |
ENSMUST00000040876.7
|
Ak7
|
adenylate kinase 7 |
| chr4_-_133399909 | 0.33 |
ENSMUST00000062118.11
ENSMUST00000067902.13 |
Pigv
|
phosphatidylinositol glycan anchor biosynthesis, class V |
| chr1_-_153683908 | 0.33 |
ENSMUST00000141249.8
|
Rgsl1
|
regulator of G-protein signaling like 1 |
| chr19_-_24202344 | 0.32 |
ENSMUST00000099558.5
ENSMUST00000232956.2 |
Tjp2
|
tight junction protein 2 |
| chr5_-_5529119 | 0.32 |
ENSMUST00000115447.2
|
Pttg1ip2
|
PTTG1IP family member 2 |
| chr6_-_41751648 | 0.32 |
ENSMUST00000214752.2
|
Olfr459
|
olfactory receptor 459 |
| chr13_+_38041910 | 0.30 |
ENSMUST00000138110.8
|
Rreb1
|
ras responsive element binding protein 1 |
| chr5_-_37494179 | 0.29 |
ENSMUST00000114148.2
|
Evc
|
EvC ciliary complex subunit 1 |
| chr17_-_57348647 | 0.28 |
ENSMUST00000169012.2
ENSMUST00000058661.14 |
Slc25a41
|
solute carrier family 25, member 41 |
| chr14_+_53791444 | 0.25 |
ENSMUST00000198297.2
|
Trav14-1
|
T cell receptor alpha variable 14-1 |
| chr9_-_50663571 | 0.23 |
ENSMUST00000042790.5
|
Hspb2
|
heat shock protein 2 |
| chr4_-_15149755 | 0.22 |
ENSMUST00000108273.2
|
Necab1
|
N-terminal EF-hand calcium binding protein 1 |
| chr5_-_37494213 | 0.21 |
ENSMUST00000031005.11
|
Evc
|
EvC ciliary complex subunit 1 |
| chr10_+_80707669 | 0.20 |
ENSMUST00000219817.2
|
Tmprss9
|
transmembrane protease, serine 9 |
| chr6_-_129600812 | 0.20 |
ENSMUST00000168919.8
|
Klrk1
|
killer cell lectin-like receptor subfamily K, member 1 |
| chr4_+_53440389 | 0.19 |
ENSMUST00000107646.9
ENSMUST00000102911.10 |
Slc44a1
|
solute carrier family 44, member 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.2 | 16.6 | GO:0010901 | regulation of very-low-density lipoprotein particle remodeling(GO:0010901) |
| 3.7 | 22.1 | GO:0033540 | fatty acid beta-oxidation using acyl-CoA oxidase(GO:0033540) |
| 3.7 | 14.6 | GO:0006069 | ethanol oxidation(GO:0006069) |
| 2.8 | 13.9 | GO:0071718 | sodium-independent icosanoid transport(GO:0071718) |
| 2.7 | 19.2 | GO:0060332 | positive regulation of humoral immune response mediated by circulating immunoglobulin(GO:0002925) heme transport(GO:0015886) positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) |
| 2.5 | 15.1 | GO:0000270 | peptidoglycan metabolic process(GO:0000270) peptidoglycan catabolic process(GO:0009253) negative regulation of natural killer cell differentiation(GO:0032824) negative regulation of natural killer cell differentiation involved in immune response(GO:0032827) |
| 2.5 | 17.3 | GO:0046618 | drug export(GO:0046618) |
| 2.1 | 8.4 | GO:0001970 | positive regulation of activation of membrane attack complex(GO:0001970) |
| 1.9 | 13.6 | GO:0019856 | 'de novo' pyrimidine nucleobase biosynthetic process(GO:0006207) pyrimidine nucleobase biosynthetic process(GO:0019856) |
| 1.7 | 24.5 | GO:0006751 | glutathione catabolic process(GO:0006751) |
| 1.6 | 18.1 | GO:1903797 | positive regulation of sodium ion export(GO:1903275) positive regulation of sodium ion export from cell(GO:1903278) positive regulation of inorganic anion transmembrane transport(GO:1903797) |
| 1.4 | 12.4 | GO:0060763 | mammary duct terminal end bud growth(GO:0060763) |
| 1.3 | 5.1 | GO:0010982 | GPI anchor release(GO:0006507) regulation of high-density lipoprotein particle clearance(GO:0010982) |
| 1.3 | 11.3 | GO:0015879 | carnitine transport(GO:0015879) |
| 1.2 | 11.2 | GO:0046874 | quinolinate metabolic process(GO:0046874) |
| 1.1 | 14.9 | GO:0070863 | positive regulation of protein glycosylation(GO:0060050) positive regulation of protein exit from endoplasmic reticulum(GO:0070863) |
| 1.1 | 16.7 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 1.1 | 3.2 | GO:0016256 | N-glycan processing to lysosome(GO:0016256) |
| 1.0 | 5.8 | GO:1990166 | protein localization to site of double-strand break(GO:1990166) |
| 0.9 | 2.8 | GO:1900039 | positive regulation of cellular response to hypoxia(GO:1900039) regulation of hypoxia-inducible factor-1alpha signaling pathway(GO:1902071) |
| 0.9 | 10.8 | GO:0043651 | linoleic acid metabolic process(GO:0043651) |
| 0.9 | 2.7 | GO:0033189 | response to vitamin A(GO:0033189) |
| 0.8 | 4.2 | GO:0048807 | female genitalia morphogenesis(GO:0048807) |
| 0.8 | 13.1 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.8 | 5.7 | GO:0036438 | maintenance of lens transparency(GO:0036438) |
| 0.7 | 4.6 | GO:0089700 | protein kinase D signaling(GO:0089700) |
| 0.6 | 2.2 | GO:0038108 | negative regulation of appetite by leptin-mediated signaling pathway(GO:0038108) |
| 0.5 | 3.3 | GO:1900245 | negative regulation of MDA-5 signaling pathway(GO:0039534) positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.5 | 2.1 | GO:0038163 | thrombopoietin-mediated signaling pathway(GO:0038163) |
| 0.4 | 1.6 | GO:0017187 | peptidyl-glutamic acid carboxylation(GO:0017187) |
| 0.4 | 7.6 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.4 | 4.4 | GO:0097460 | ferrous iron import into cell(GO:0097460) ferrous iron import across plasma membrane(GO:0098707) |
| 0.4 | 1.6 | GO:0045415 | negative regulation of interleukin-8 biosynthetic process(GO:0045415) |
| 0.4 | 23.8 | GO:1900047 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.3 | 1.4 | GO:0006669 | sphinganine-1-phosphate biosynthetic process(GO:0006669) |
| 0.3 | 2.6 | GO:0061370 | testosterone biosynthetic process(GO:0061370) |
| 0.3 | 1.9 | GO:0060523 | prostate epithelial cord elongation(GO:0060523) |
| 0.3 | 7.9 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.3 | 1.2 | GO:0055014 | atrial cardiac muscle cell differentiation(GO:0055011) atrial cardiac muscle cell development(GO:0055014) |
| 0.3 | 4.4 | GO:1990440 | positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.3 | 12.0 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.3 | 2.0 | GO:0042737 | drug catabolic process(GO:0042737) |
| 0.3 | 1.6 | GO:0002084 | protein depalmitoylation(GO:0002084) macromolecule depalmitoylation(GO:0098734) |
| 0.3 | 17.4 | GO:0019369 | arachidonic acid metabolic process(GO:0019369) |
| 0.3 | 4.5 | GO:2000400 | positive regulation of T cell differentiation in thymus(GO:0033089) positive regulation of thymocyte aggregation(GO:2000400) |
| 0.3 | 2.1 | GO:0008611 | ether lipid biosynthetic process(GO:0008611) glycerol ether biosynthetic process(GO:0046504) ether biosynthetic process(GO:1901503) |
| 0.2 | 0.7 | GO:0030887 | positive regulation of myeloid dendritic cell activation(GO:0030887) |
| 0.2 | 2.2 | GO:0038031 | non-canonical Wnt signaling pathway via MAPK cascade(GO:0038030) non-canonical Wnt signaling pathway via JNK cascade(GO:0038031) |
| 0.2 | 0.9 | GO:0010609 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902164) |
| 0.2 | 1.6 | GO:0048050 | post-embryonic eye morphogenesis(GO:0048050) post-embryonic camera-type eye morphogenesis(GO:0048597) |
| 0.2 | 1.2 | GO:0072344 | rescue of stalled ribosome(GO:0072344) |
| 0.2 | 0.8 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.1 | 1.3 | GO:0032049 | cardiolipin biosynthetic process(GO:0032049) |
| 0.1 | 6.3 | GO:0071625 | vocalization behavior(GO:0071625) |
| 0.1 | 0.7 | GO:1902774 | late endosome to lysosome transport(GO:1902774) |
| 0.1 | 4.7 | GO:0006084 | acetyl-CoA metabolic process(GO:0006084) |
| 0.1 | 0.7 | GO:0003409 | optic cup structural organization(GO:0003409) |
| 0.1 | 14.0 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.1 | 0.9 | GO:0010968 | regulation of microtubule nucleation(GO:0010968) |
| 0.1 | 9.3 | GO:0045668 | negative regulation of osteoblast differentiation(GO:0045668) |
| 0.1 | 1.3 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.1 | 1.6 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.1 | 0.8 | GO:1900740 | positive regulation of thymocyte apoptotic process(GO:0070245) regulation of protein insertion into mitochondrial membrane involved in apoptotic signaling pathway(GO:1900739) positive regulation of protein insertion into mitochondrial membrane involved in apoptotic signaling pathway(GO:1900740) |
| 0.1 | 0.8 | GO:0015820 | branched-chain amino acid transport(GO:0015803) leucine transport(GO:0015820) |
| 0.1 | 0.6 | GO:0006172 | ADP biosynthetic process(GO:0006172) inosine biosynthetic process(GO:0046103) |
| 0.1 | 0.9 | GO:2000483 | negative regulation of interleukin-8 secretion(GO:2000483) |
| 0.1 | 1.1 | GO:0018904 | glycerol ether metabolic process(GO:0006662) ether metabolic process(GO:0018904) |
| 0.1 | 1.0 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 0.1 | 3.1 | GO:0015986 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.1 | 2.0 | GO:0042178 | xenobiotic catabolic process(GO:0042178) |
| 0.1 | 0.5 | GO:0034627 | nicotinamide nucleotide biosynthetic process from aspartate(GO:0019355) 'de novo' NAD biosynthetic process(GO:0034627) 'de novo' NAD biosynthetic process from aspartate(GO:0034628) |
| 0.1 | 4.5 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.1 | 0.8 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.1 | 0.8 | GO:0048680 | positive regulation of axon regeneration(GO:0048680) |
| 0.1 | 6.2 | GO:0070206 | protein trimerization(GO:0070206) |
| 0.1 | 0.9 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.1 | 1.4 | GO:0014850 | response to muscle activity(GO:0014850) |
| 0.1 | 1.2 | GO:0046855 | inositol phosphate dephosphorylation(GO:0046855) |
| 0.1 | 0.9 | GO:0032525 | somite rostral/caudal axis specification(GO:0032525) |
| 0.1 | 3.3 | GO:0009268 | response to pH(GO:0009268) |
| 0.1 | 0.4 | GO:0031161 | phosphatidylinositol catabolic process(GO:0031161) |
| 0.1 | 0.3 | GO:1903691 | positive regulation of wound healing, spreading of epidermal cells(GO:1903691) |
| 0.0 | 0.3 | GO:2001205 | negative regulation of osteoclast development(GO:2001205) |
| 0.0 | 0.6 | GO:1900037 | regulation of cellular response to hypoxia(GO:1900037) |
| 0.0 | 0.4 | GO:1901409 | positive regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901409) |
| 0.0 | 3.8 | GO:0022904 | respiratory electron transport chain(GO:0022904) |
| 0.0 | 1.2 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.0 | 2.2 | GO:0045071 | negative regulation of viral genome replication(GO:0045071) |
| 0.0 | 0.3 | GO:0060040 | retinal bipolar neuron differentiation(GO:0060040) |
| 0.0 | 0.2 | GO:0015871 | choline transport(GO:0015871) |
| 0.0 | 0.7 | GO:0006903 | vesicle targeting(GO:0006903) |
| 0.0 | 0.1 | GO:2001025 | positive regulation of response to drug(GO:2001025) |
| 0.0 | 1.0 | GO:0006505 | GPI anchor metabolic process(GO:0006505) |
| 0.0 | 0.2 | GO:0031639 | plasminogen activation(GO:0031639) |
| 0.0 | 1.1 | GO:0030490 | maturation of SSU-rRNA(GO:0030490) |
| 0.0 | 0.3 | GO:0003351 | epithelial cilium movement(GO:0003351) |
| 0.0 | 0.5 | GO:0003416 | endochondral bone growth(GO:0003416) bone growth(GO:0098868) |
| 0.0 | 0.5 | GO:0014047 | glutamate secretion(GO:0014047) |
| 0.0 | 1.3 | GO:0032006 | regulation of TOR signaling(GO:0032006) |
| 0.0 | 2.3 | GO:0007596 | blood coagulation(GO:0007596) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 7.9 | GO:0017177 | glucosidase II complex(GO:0017177) |
| 1.0 | 17.3 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.9 | 6.3 | GO:0097513 | myosin II filament(GO:0097513) |
| 0.7 | 16.6 | GO:0042627 | chylomicron(GO:0042627) |
| 0.3 | 3.1 | GO:0000275 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 0.3 | 4.4 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.3 | 1.9 | GO:0097550 | transcriptional preinitiation complex(GO:0097550) |
| 0.3 | 5.1 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.2 | 13.6 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.2 | 6.2 | GO:0031091 | platelet alpha granule(GO:0031091) |
| 0.2 | 22.1 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.2 | 61.3 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.2 | 9.3 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.2 | 14.5 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.2 | 4.7 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.2 | 2.2 | GO:0034464 | BBSome(GO:0034464) |
| 0.2 | 13.9 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 2.6 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.1 | 5.8 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.1 | 0.7 | GO:0097441 | basilar dendrite(GO:0097441) |
| 0.1 | 0.6 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.1 | 0.3 | GO:0031501 | mannosyltransferase complex(GO:0031501) |
| 0.1 | 2.2 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.1 | 4.2 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.1 | 2.8 | GO:0005747 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.1 | 0.7 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.0 | 0.4 | GO:0008024 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) |
| 0.0 | 0.6 | GO:0001520 | outer dense fiber(GO:0001520) |
| 0.0 | 10.0 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.6 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.0 | 12.1 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 0.6 | GO:0071006 | U2-type catalytic step 1 spliceosome(GO:0071006) |
| 0.0 | 0.8 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.0 | 0.7 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.0 | 1.2 | GO:0031430 | M band(GO:0031430) |
| 0.0 | 0.8 | GO:0070069 | cytochrome complex(GO:0070069) |
| 0.0 | 1.1 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 0.7 | GO:0031305 | integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 0.5 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 4.6 | GO:0005765 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.0 | 11.6 | GO:0005740 | mitochondrial envelope(GO:0005740) |
| 0.0 | 5.4 | GO:0043235 | receptor complex(GO:0043235) |
| 0.0 | 0.2 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.0 | 0.9 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 0.3 | GO:0005921 | gap junction(GO:0005921) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.9 | 14.6 | GO:0004024 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) |
| 4.5 | 13.6 | GO:0004088 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 3.2 | 19.2 | GO:0015232 | heme transporter activity(GO:0015232) |
| 3.2 | 22.1 | GO:0016401 | palmitoyl-CoA oxidase activity(GO:0016401) |
| 2.8 | 11.3 | GO:0015226 | amino-acid betaine transmembrane transporter activity(GO:0015199) carnitine transmembrane transporter activity(GO:0015226) |
| 2.7 | 10.8 | GO:0071614 | linoleic acid epoxygenase activity(GO:0071614) |
| 2.5 | 15.1 | GO:0008745 | N-acetylmuramoyl-L-alanine amidase activity(GO:0008745) peptidoglycan receptor activity(GO:0016019) |
| 2.2 | 13.0 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 2.0 | 24.5 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 2.0 | 44.2 | GO:0015125 | bile acid transmembrane transporter activity(GO:0015125) |
| 1.4 | 5.7 | GO:0008109 | N-acetyllactosaminide beta-1,6-N-acetylglucosaminyltransferase activity(GO:0008109) |
| 1.4 | 4.2 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 1.1 | 6.6 | GO:0008321 | Ral guanyl-nucleotide exchange factor activity(GO:0008321) |
| 0.9 | 4.7 | GO:0003986 | acetyl-CoA hydrolase activity(GO:0003986) |
| 0.9 | 3.6 | GO:0070653 | high-density lipoprotein particle receptor binding(GO:0070653) |
| 0.9 | 4.4 | GO:0004998 | transferrin receptor activity(GO:0004998) |
| 0.9 | 9.6 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.8 | 3.2 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 0.7 | 2.1 | GO:0016297 | [acyl-carrier-protein] S-malonyltransferase activity(GO:0004314) oleoyl-[acyl-carrier-protein] hydrolase activity(GO:0004320) myristoyl-[acyl-carrier-protein] hydrolase activity(GO:0016295) palmitoyl-[acyl-carrier-protein] hydrolase activity(GO:0016296) acyl-[acyl-carrier-protein] hydrolase activity(GO:0016297) S-acetyltransferase activity(GO:0016418) S-malonyltransferase activity(GO:0016419) malonyltransferase activity(GO:0016420) phosphopantetheine binding(GO:0031177) |
| 0.6 | 2.6 | GO:0047751 | 3-oxo-5-alpha-steroid 4-dehydrogenase activity(GO:0003865) cholestenone 5-alpha-reductase activity(GO:0047751) |
| 0.6 | 1.9 | GO:0038052 | type 1 metabotropic glutamate receptor binding(GO:0031798) RNA polymerase II transcription factor activity, estrogen-activated sequence-specific DNA binding(GO:0038052) |
| 0.6 | 5.8 | GO:0003909 | DNA ligase activity(GO:0003909) DNA ligase (ATP) activity(GO:0003910) |
| 0.6 | 5.1 | GO:0004630 | phospholipase D activity(GO:0004630) |
| 0.6 | 4.5 | GO:0034235 | GPI anchor binding(GO:0034235) |
| 0.5 | 2.0 | GO:0003960 | NADPH:quinone reductase activity(GO:0003960) |
| 0.5 | 15.0 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.4 | 11.2 | GO:0019825 | oxygen binding(GO:0019825) |
| 0.4 | 1.1 | GO:0046964 | 3'-phosphoadenosine 5'-phosphosulfate transmembrane transporter activity(GO:0046964) |
| 0.4 | 12.3 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.3 | 1.1 | GO:0008113 | peptide-methionine (S)-S-oxide reductase activity(GO:0008113) |
| 0.3 | 17.4 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.3 | 6.3 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.2 | 2.2 | GO:0034452 | dynactin binding(GO:0034452) |
| 0.2 | 0.7 | GO:0032394 | MHC class Ib receptor activity(GO:0032394) |
| 0.2 | 1.6 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.2 | 7.4 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.2 | 18.1 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.2 | 4.4 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.2 | 4.6 | GO:0004697 | protein kinase C activity(GO:0004697) |
| 0.2 | 1.2 | GO:0046030 | inositol-polyphosphate 5-phosphatase activity(GO:0004445) inositol trisphosphate phosphatase activity(GO:0046030) |
| 0.2 | 3.1 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.2 | 1.6 | GO:0098599 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.2 | 1.6 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.2 | 33.9 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.2 | 1.4 | GO:0008481 | sphinganine kinase activity(GO:0008481) D-erythro-sphingosine kinase activity(GO:0017050) |
| 0.2 | 3.6 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.1 | 3.3 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.1 | 25.7 | GO:0052689 | carboxylic ester hydrolase activity(GO:0052689) |
| 0.1 | 33.3 | GO:0004866 | endopeptidase inhibitor activity(GO:0004866) |
| 0.1 | 1.3 | GO:0004439 | phosphatidylinositol-4,5-bisphosphate 5-phosphatase activity(GO:0004439) |
| 0.1 | 0.8 | GO:0004311 | farnesyltranstransferase activity(GO:0004311) |
| 0.1 | 0.5 | GO:0000309 | nicotinamide-nucleotide adenylyltransferase activity(GO:0000309) nicotinate-nucleotide adenylyltransferase activity(GO:0004515) |
| 0.1 | 0.6 | GO:0004689 | phosphorylase kinase activity(GO:0004689) |
| 0.1 | 2.1 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 2.2 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.1 | 0.4 | GO:0052834 | inositol monophosphate 1-phosphatase activity(GO:0008934) inositol monophosphate 3-phosphatase activity(GO:0052832) inositol monophosphate 4-phosphatase activity(GO:0052833) inositol monophosphate phosphatase activity(GO:0052834) |
| 0.1 | 7.2 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.1 | 1.6 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.1 | 1.8 | GO:0015002 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.1 | 1.6 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 2.0 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.1 | 0.5 | GO:0001640 | adenylate cyclase inhibiting G-protein coupled glutamate receptor activity(GO:0001640) |
| 0.0 | 0.7 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.0 | 5.5 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.0 | 1.3 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.0 | 7.5 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 7.1 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 6.1 | GO:0043130 | ubiquitin binding(GO:0043130) |
| 0.0 | 0.2 | GO:0036313 | phosphatidylinositol 3-kinase catalytic subunit binding(GO:0036313) |
| 0.0 | 0.4 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
| 0.0 | 0.8 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.0 | 0.5 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.0 | 0.2 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.0 | 1.6 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 0.3 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.0 | 2.2 | GO:0004721 | phosphoprotein phosphatase activity(GO:0004721) |
| 0.0 | 0.9 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.0 | 0.6 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.0 | 1.2 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 0.2 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.0 | 0.3 | GO:0000030 | mannosyltransferase activity(GO:0000030) |
| 0.0 | 2.8 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 22.4 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.6 | 16.9 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.2 | 38.0 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.2 | 12.4 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.2 | 1.6 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.1 | 5.1 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.1 | 15.7 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 1.6 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.0 | 2.2 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.0 | 1.4 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.0 | 1.9 | PID ER NONGENOMIC PATHWAY | Plasma membrane estrogen receptor signaling |
| 0.0 | 2.1 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 1.2 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 1.3 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 25.1 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 1.4 | 15.0 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 1.2 | 13.9 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.9 | 22.1 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.8 | 11.2 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.6 | 16.6 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.5 | 6.2 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.5 | 6.4 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.5 | 7.9 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.5 | 11.3 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.4 | 7.4 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.2 | 6.3 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.2 | 2.6 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.2 | 1.6 | REACTOME OPSINS | Genes involved in Opsins |
| 0.2 | 1.6 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.2 | 15.9 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.1 | 3.1 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.1 | 7.6 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.1 | 2.1 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.1 | 3.6 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.1 | 3.9 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.1 | 1.0 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.0 | 12.4 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.0 | 0.8 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.0 | 2.8 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 1.0 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 1.4 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.6 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 0.5 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.0 | 0.6 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 2.3 | REACTOME RIG I MDA5 MEDIATED INDUCTION OF IFN ALPHA BETA PATHWAYS | Genes involved in RIG-I/MDA5 mediated induction of IFN-alpha/beta pathways |
| 0.0 | 0.6 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 0.3 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 0.4 | REACTOME ELONGATION ARREST AND RECOVERY | Genes involved in Elongation arrest and recovery |
| 0.0 | 1.3 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |