Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
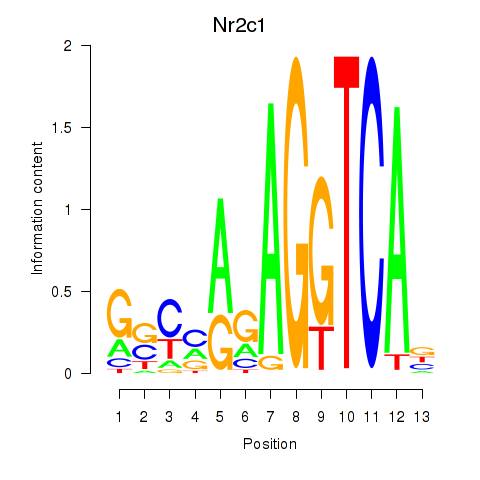
Results for Nr2c1
Z-value: 1.17
Transcription factors associated with Nr2c1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Nr2c1
|
ENSMUSG00000005897.15 | Nr2c1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Nr2c1 | mm39_v1_chr10_+_93983844_93983885 | -0.13 | 4.4e-01 | Click! |
Activity profile of Nr2c1 motif
Sorted Z-values of Nr2c1 motif
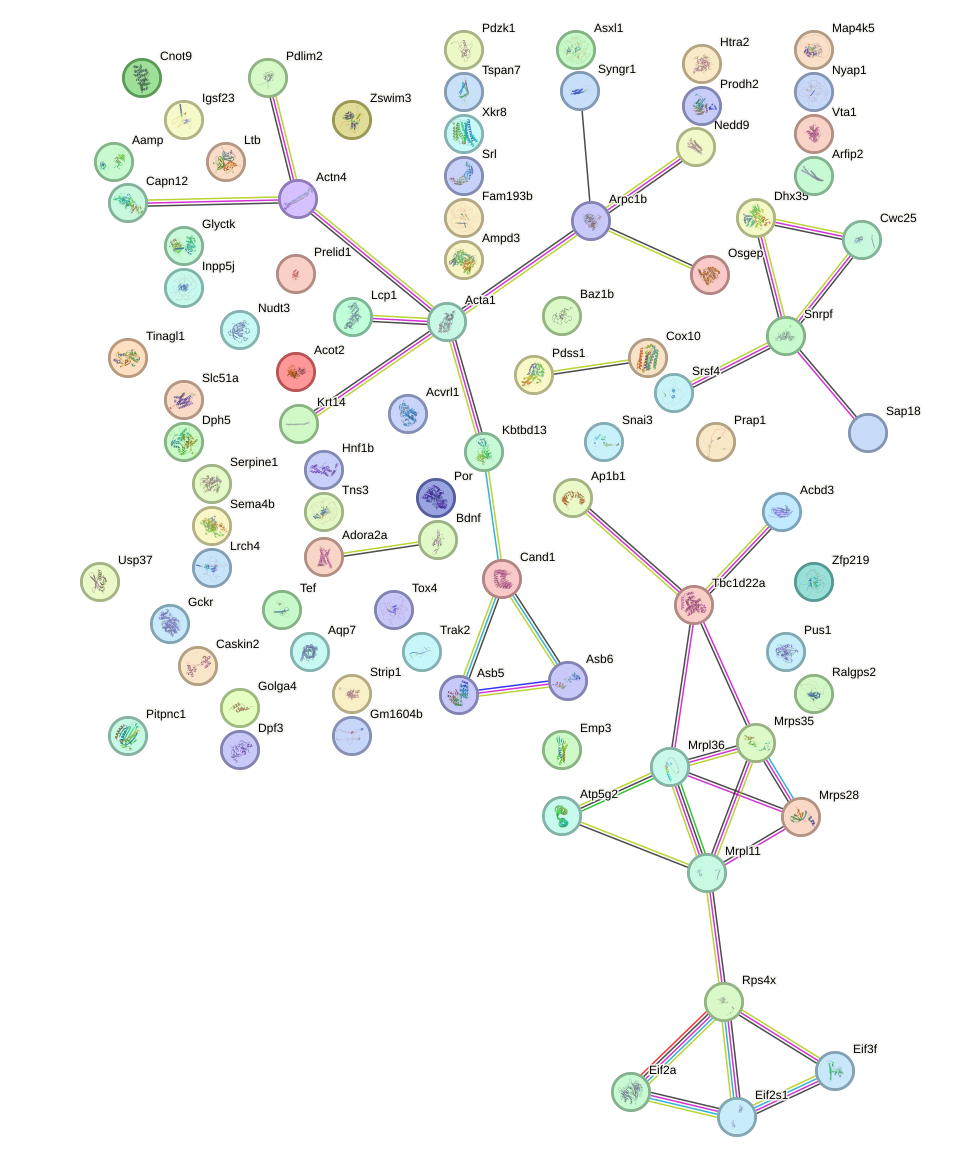
Network of associatons between targets according to the STRING database.
First level regulatory network of Nr2c1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_+_79975520 | 3.91 |
ENSMUST00000009728.13
ENSMUST00000009727.12 |
Syngr1
|
synaptogyrin 1 |
| chr7_+_30193047 | 2.93 |
ENSMUST00000058280.13
ENSMUST00000133318.8 ENSMUST00000142575.8 ENSMUST00000131040.2 |
Prodh2
|
proline dehydrogenase (oxidase) 2 |
| chr8_-_124621483 | 2.84 |
ENSMUST00000034453.6
ENSMUST00000212584.2 |
Acta1
|
actin, alpha 1, skeletal muscle |
| chr8_-_123187406 | 2.49 |
ENSMUST00000006762.7
|
Snai3
|
snail family zinc finger 3 |
| chrX_+_10351360 | 2.44 |
ENSMUST00000076354.13
ENSMUST00000115526.2 |
Tspan7
|
tetraspanin 7 |
| chr1_-_59012579 | 2.29 |
ENSMUST00000173590.2
ENSMUST00000027186.12 |
Trak2
|
trafficking protein, kinesin binding 2 |
| chr11_+_4936824 | 2.21 |
ENSMUST00000109897.8
ENSMUST00000009234.16 |
Ap1b1
|
adaptor protein complex AP-1, beta 1 subunit |
| chr7_+_110367375 | 2.04 |
ENSMUST00000170374.8
|
Ampd3
|
adenosine monophosphate deaminase 3 |
| chr7_-_45570828 | 1.90 |
ENSMUST00000038876.13
|
Emp3
|
epithelial membrane protein 3 |
| chr5_+_145060489 | 1.86 |
ENSMUST00000136074.2
|
Arpc1b
|
actin related protein 2/3 complex, subunit 1B |
| chr7_+_108533613 | 1.77 |
ENSMUST00000033342.7
|
Eif3f
|
eukaryotic translation initiation factor 3, subunit F |
| chr6_-_83030759 | 1.75 |
ENSMUST00000134606.8
|
Htra2
|
HtrA serine peptidase 2 |
| chr7_-_45570254 | 1.73 |
ENSMUST00000164119.3
|
Emp3
|
epithelial membrane protein 3 |
| chr10_-_93425553 | 1.72 |
ENSMUST00000020203.7
|
Snrpf
|
small nuclear ribonucleoprotein polypeptide F |
| chr7_-_45570538 | 1.70 |
ENSMUST00000210297.2
|
Emp3
|
epithelial membrane protein 3 |
| chr2_-_30720345 | 1.68 |
ENSMUST00000041726.4
|
Asb6
|
ankyrin repeat and SOCS box-containing 6 |
| chr7_-_45570674 | 1.53 |
ENSMUST00000210939.2
|
Emp3
|
epithelial membrane protein 3 |
| chr14_-_70414236 | 1.53 |
ENSMUST00000153735.8
|
Pdlim2
|
PDZ and LIM domain 2 |
| chr2_+_164647002 | 1.43 |
ENSMUST00000052107.5
|
Zswim3
|
zinc finger SWIM-type containing 3 |
| chr11_-_3454766 | 1.37 |
ENSMUST00000044507.12
|
Inpp5j
|
inositol polyphosphate 5-phosphatase J |
| chr17_+_35413415 | 1.35 |
ENSMUST00000025262.6
ENSMUST00000173600.2 |
Ltb
|
lymphotoxin B |
| chr7_+_79836581 | 1.25 |
ENSMUST00000032754.9
|
Sema4b
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4B |
| chr6_+_125122172 | 1.25 |
ENSMUST00000119527.8
ENSMUST00000117675.8 |
Iffo1
|
intermediate filament family orphan 1 |
| chr3_+_96737385 | 1.22 |
ENSMUST00000058865.14
|
Pdzk1
|
PDZ domain containing 1 |
| chr2_+_22785534 | 1.21 |
ENSMUST00000053729.14
|
Pdss1
|
prenyl (solanesyl) diphosphate synthase, subunit 1 |
| chr1_+_74545203 | 1.13 |
ENSMUST00000087215.7
|
Cnot9
|
CCR4-NOT transcription complex, subunit 9 |
| chr9_-_106035332 | 1.10 |
ENSMUST00000112543.9
|
Glyctk
|
glycerate kinase |
| chr6_-_83031358 | 1.10 |
ENSMUST00000113962.8
ENSMUST00000089645.13 ENSMUST00000113963.8 |
Htra2
|
HtrA serine peptidase 2 |
| chr17_-_27816151 | 1.08 |
ENSMUST00000231742.2
|
Nudt3
|
nudix (nucleotide diphosphate linked moiety X)-type motif 3 |
| chr13_+_73476629 | 1.07 |
ENSMUST00000221730.2
|
Mrpl36
|
mitochondrial ribosomal protein L36 |
| chrX_-_101232978 | 1.07 |
ENSMUST00000033683.8
|
Rps4x
|
ribosomal protein S4, X-linked |
| chr9_-_106035308 | 1.05 |
ENSMUST00000159809.2
ENSMUST00000162562.2 ENSMUST00000036382.13 |
Glyctk
|
glycerate kinase |
| chr4_-_41048124 | 1.05 |
ENSMUST00000030136.13
|
Aqp7
|
aquaporin 7 |
| chr5_+_31454787 | 1.05 |
ENSMUST00000201166.4
ENSMUST00000072228.9 |
Gckr
|
glucokinase regulatory protein |
| chr17_-_13211374 | 1.04 |
ENSMUST00000159551.8
ENSMUST00000160781.8 |
Wtap
|
Wilms tumour 1-associating protein |
| chr7_-_19684654 | 1.02 |
ENSMUST00000043440.8
|
Igsf23
|
immunoglobulin superfamily, member 23 |
| chr12_-_83534482 | 1.02 |
ENSMUST00000177959.8
ENSMUST00000178756.8 |
Dpf3
|
D4, zinc and double PHD fingers, family 3 |
| chr4_+_131600918 | 1.02 |
ENSMUST00000053819.6
|
Srsf4
|
serine and arginine-rich splicing factor 4 |
| chr5_+_135216090 | 1.01 |
ENSMUST00000002825.6
|
Baz1b
|
bromodomain adjacent to zinc finger domain, 1B |
| chr14_-_51162083 | 0.92 |
ENSMUST00000160375.8
ENSMUST00000162177.8 |
Osgep
|
O-sialoglycoprotein endopeptidase |
| chr10_+_75152705 | 0.88 |
ENSMUST00000105420.3
|
Adora2a
|
adenosine A2a receptor |
| chr17_-_13211075 | 0.88 |
ENSMUST00000159986.8
ENSMUST00000007007.14 |
Wtap
|
Wilms tumour 1-associating protein |
| chr5_+_137628377 | 0.83 |
ENSMUST00000175968.8
|
Lrch4
|
leucine-rich repeats and calponin homology (CH) domain containing 4 |
| chr15_+_79025523 | 0.81 |
ENSMUST00000040077.8
|
Polr2f
|
polymerase (RNA) II (DNA directed) polypeptide F |
| chr13_-_41513215 | 0.80 |
ENSMUST00000224803.2
|
Nedd9
|
neural precursor cell expressed, developmentally down-regulated gene 9 |
| chr7_-_28297565 | 0.77 |
ENSMUST00000040531.9
ENSMUST00000108283.8 |
Samd4b
Pak4
|
sterile alpha motif domain containing 4B p21 (RAC1) activated kinase 4 |
| chr1_+_133237516 | 0.76 |
ENSMUST00000094557.7
ENSMUST00000192465.2 ENSMUST00000193888.6 ENSMUST00000194044.6 ENSMUST00000184603.8 |
Golt1a
Gm28040
Gm28040
|
golgi transport 1A predicted gene, 28040 predicted gene, 28040 |
| chr5_+_135717890 | 0.75 |
ENSMUST00000005651.13
ENSMUST00000122113.8 |
Por
|
P450 (cytochrome) oxidoreductase |
| chr14_-_51162346 | 0.73 |
ENSMUST00000159292.8
|
Osgep
|
O-sialoglycoprotein endopeptidase |
| chr4_-_130068484 | 0.73 |
ENSMUST00000132545.3
ENSMUST00000175992.8 ENSMUST00000105999.9 |
Tinagl1
|
tubulointerstitial nephritis antigen-like 1 |
| chr2_+_22785593 | 0.73 |
ENSMUST00000152170.8
|
Pdss1
|
prenyl (solanesyl) diphosphate synthase, subunit 1 |
| chr15_-_102579463 | 0.70 |
ENSMUST00000185641.7
|
Atp5g2
|
ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C2 (subunit 9) |
| chr8_+_96551974 | 0.69 |
ENSMUST00000074053.6
|
Sap18b
|
Sin3-associated polypeptide 18B |
| chr13_+_55468313 | 0.69 |
ENSMUST00000021942.8
|
Prelid1
|
PRELI domain containing 1 |
| chr11_+_83741689 | 0.69 |
ENSMUST00000108114.9
|
Hnf1b
|
HNF1 homeobox B |
| chr15_+_101026803 | 0.67 |
ENSMUST00000119063.8
ENSMUST00000121718.8 |
Acvrl1
|
activin A receptor, type II-like 1 |
| chr11_+_83741657 | 0.65 |
ENSMUST00000021016.10
|
Hnf1b
|
HNF1 homeobox B |
| chr17_+_7292967 | 0.64 |
ENSMUST00000097422.6
|
Gm1604b
|
predicted gene 1604b |
| chr15_-_79025387 | 0.63 |
ENSMUST00000187550.7
ENSMUST00000188562.7 ENSMUST00000190509.7 ENSMUST00000190730.7 ENSMUST00000190959.7 ENSMUST00000169604.8 ENSMUST00000186053.7 |
1700088E04Rik
|
RIKEN cDNA 1700088E04 gene |
| chr7_+_139673300 | 0.63 |
ENSMUST00000026540.9
|
Prap1
|
proline-rich acidic protein 1 |
| chr1_-_156767123 | 0.63 |
ENSMUST00000189316.7
ENSMUST00000190648.7 ENSMUST00000172057.8 ENSMUST00000191605.7 |
Ralgps2
|
Ral GEF with PH domain and SH3 binding motif 2 |
| chr3_-_8988854 | 0.62 |
ENSMUST00000042148.6
|
Mrps28
|
mitochondrial ribosomal protein S28 |
| chr8_+_55024446 | 0.59 |
ENSMUST00000239166.2
ENSMUST00000239106.2 ENSMUST00000239152.2 |
Asb5
|
ankyrin repeat and SOCs box-containing 5 |
| chr19_-_45986919 | 0.56 |
ENSMUST00000045396.9
|
Armh3
|
armadillo-like helical domain containing 3 |
| chr3_+_58433236 | 0.56 |
ENSMUST00000029387.15
|
Eif2a
|
eukaryotic translation initiation factor 2A |
| chr11_-_63970270 | 0.55 |
ENSMUST00000049091.9
|
Cox10
|
heme A:farnesyltransferase cytochrome c oxidase assembly factor 10 |
| chr2_+_153187729 | 0.53 |
ENSMUST00000227428.2
ENSMUST00000109790.2 |
Asxl1
|
ASXL transcriptional regulator 1 |
| chr10_-_14581203 | 0.53 |
ENSMUST00000149485.2
ENSMUST00000154132.8 |
Vta1
|
vesicle (multivesicular body) trafficking 1 |
| chr1_-_156766957 | 0.52 |
ENSMUST00000171292.8
ENSMUST00000063199.13 ENSMUST00000027886.14 |
Ralgps2
|
Ral GEF with PH domain and SH3 binding motif 2 |
| chr15_+_101026454 | 0.52 |
ENSMUST00000117984.8
|
Acvrl1
|
activin A receptor, type II-like 1 |
| chr10_-_119075910 | 0.50 |
ENSMUST00000020315.13
|
Cand1
|
cullin associated and neddylation disassociated 1 |
| chr5_-_137101108 | 0.50 |
ENSMUST00000077523.4
ENSMUST00000041388.11 |
Serpine1
|
serine (or cysteine) peptidase inhibitor, clade E, member 1 |
| chr14_+_75373766 | 0.49 |
ENSMUST00000145303.8
|
Lcp1
|
lymphocyte cytosolic protein 1 |
| chr11_-_115698894 | 0.48 |
ENSMUST00000132780.8
|
Caskin2
|
CASK-interacting protein 2 |
| chr13_-_55718899 | 0.48 |
ENSMUST00000225240.3
ENSMUST00000021957.8 |
Fam193b
|
family with sequence similarity 193, member B |
| chr3_-_107538996 | 0.47 |
ENSMUST00000064759.7
|
Strip1
|
striatin interacting protein 1 |
| chr16_-_4340920 | 0.46 |
ENSMUST00000090500.10
ENSMUST00000023161.8 |
Srl
|
sarcalumenin |
| chr19_+_5012362 | 0.45 |
ENSMUST00000236917.2
|
Mrpl11
|
mitochondrial ribosomal protein L11 |
| chr7_-_28597523 | 0.44 |
ENSMUST00000216863.2
|
Actn4
|
actinin alpha 4 |
| chr14_+_58036071 | 0.44 |
ENSMUST00000111268.8
|
Sap18
|
Sin3-associated polypeptide 18 |
| chr2_+_109522781 | 0.44 |
ENSMUST00000111050.10
|
Bdnf
|
brain derived neurotrophic factor |
| chr9_+_118335294 | 0.43 |
ENSMUST00000084820.6
|
Golga4
|
golgi autoantigen, golgin subfamily a, 4 |
| chr2_+_158636727 | 0.43 |
ENSMUST00000029186.14
ENSMUST00000109478.9 ENSMUST00000156893.2 |
Dhx35
|
DEAH (Asp-Glu-Ala-His) box polypeptide 35 |
| chr15_+_81695615 | 0.42 |
ENSMUST00000023024.8
|
Tef
|
thyrotroph embryonic factor |
| chr19_+_5012396 | 0.39 |
ENSMUST00000235229.2
|
Mrpl11
|
mitochondrial ribosomal protein L11 |
| chr3_-_94693780 | 0.38 |
ENSMUST00000107273.9
ENSMUST00000238849.2 |
Cgn
|
cingulin |
| chr14_+_58035640 | 0.36 |
ENSMUST00000111269.2
|
Sap18
|
Sin3-associated polypeptide 18 |
| chr3_+_115681486 | 0.35 |
ENSMUST00000189799.7
ENSMUST00000200258.2 |
Dph5
|
diphthamide biosynthesis 5 |
| chr12_+_84034628 | 0.35 |
ENSMUST00000021649.8
|
Acot2
|
acyl-CoA thioesterase 2 |
| chr15_+_86098660 | 0.35 |
ENSMUST00000063414.9
|
Tbc1d22a
|
TBC1 domain family, member 22a |
| chr17_+_75772475 | 0.35 |
ENSMUST00000095204.6
|
Rasgrp3
|
RAS, guanyl releasing protein 3 |
| chr11_-_107228382 | 0.35 |
ENSMUST00000040380.13
|
Pitpnc1
|
phosphatidylinositol transfer protein, cytoplasmic 1 |
| chr14_+_58036093 | 0.34 |
ENSMUST00000111267.2
|
Sap18
|
Sin3-associated polypeptide 18 |
| chr5_-_110927803 | 0.33 |
ENSMUST00000112426.8
|
Pus1
|
pseudouridine synthase 1 |
| chr1_-_74323795 | 0.33 |
ENSMUST00000178235.8
ENSMUST00000006462.14 |
Aamp
|
angio-associated migratory protein |
| chr7_+_28580847 | 0.33 |
ENSMUST00000066880.6
|
Capn12
|
calpain 12 |
| chr14_-_52252003 | 0.31 |
ENSMUST00000226522.2
|
Zfp219
|
zinc finger protein 219 |
| chr1_+_180553569 | 0.30 |
ENSMUST00000027780.6
|
Acbd3
|
acyl-Coenzyme A binding domain containing 3 |
| chr17_+_57369231 | 0.29 |
ENSMUST00000097299.10
ENSMUST00000169543.8 ENSMUST00000163763.2 |
Crb3
|
crumbs family member 3 |
| chr11_-_100098333 | 0.29 |
ENSMUST00000007272.8
|
Krt14
|
keratin 14 |
| chr12_-_115083839 | 0.29 |
ENSMUST00000103521.3
|
Ighv1-50
|
immunoglobulin heavy variable 1-50 |
| chr9_-_65298934 | 0.28 |
ENSMUST00000068307.4
|
Kbtbd13
|
kelch repeat and BTB (POZ) domain containing 13 |
| chr19_+_5012336 | 0.28 |
ENSMUST00000237974.2
|
Mrpl11
|
mitochondrial ribosomal protein L11 |
| chr13_+_104424359 | 0.26 |
ENSMUST00000065766.7
|
Adamts6
|
a disintegrin-like and metallopeptidase (reprolysin type) with thrombospondin type 1 motif, 6 |
| chr11_-_97657327 | 0.26 |
ENSMUST00000107579.2
ENSMUST00000018685.9 |
Cwc25
|
CWC25 spliceosome-associated protein |
| chr11_-_8574887 | 0.26 |
ENSMUST00000239091.2
|
Tns3
|
tensin 3 |
| chr1_-_74544946 | 0.25 |
ENSMUST00000044260.11
ENSMUST00000186282.7 |
Usp37
|
ubiquitin specific peptidase 37 |
| chr16_-_89940652 | 0.25 |
ENSMUST00000114124.9
|
Tiam1
|
T cell lymphoma invasion and metastasis 1 |
| chr5_-_110928436 | 0.24 |
ENSMUST00000149208.2
ENSMUST00000031483.15 ENSMUST00000086643.12 ENSMUST00000170468.8 ENSMUST00000031481.13 |
Pus1
|
pseudouridine synthase 1 |
| chr16_-_32306683 | 0.23 |
ENSMUST00000042042.9
|
Slc51a
|
solute carrier family 51, alpha subunit |
| chr4_-_132459762 | 0.21 |
ENSMUST00000045550.5
|
Xkr8
|
X-linked Kx blood group related 8 |
| chr1_-_74583434 | 0.19 |
ENSMUST00000189257.7
|
Usp37
|
ubiquitin specific peptidase 37 |
| chr5_-_137739863 | 0.19 |
ENSMUST00000061789.14
|
Nyap1
|
neuronal tyrosine-phosphorylated phosphoinositide 3-kinase adaptor 1 |
| chr5_-_121974325 | 0.18 |
ENSMUST00000137682.2
|
Sh2b3
|
SH2B adaptor protein 3 |
| chr1_-_156767196 | 0.18 |
ENSMUST00000185198.7
|
Ralgps2
|
Ral GEF with PH domain and SH3 binding motif 2 |
| chr16_-_55659194 | 0.17 |
ENSMUST00000096026.9
ENSMUST00000036273.13 ENSMUST00000114457.8 |
Nfkbiz
|
nuclear factor of kappa light polypeptide gene enhancer in B cells inhibitor, zeta |
| chr10_+_75152908 | 0.16 |
ENSMUST00000219044.2
|
Adora2a
|
adenosine A2a receptor |
| chr2_+_172994841 | 0.16 |
ENSMUST00000029017.6
|
Pck1
|
phosphoenolpyruvate carboxykinase 1, cytosolic |
| chr4_+_156300325 | 0.14 |
ENSMUST00000105572.3
|
Perm1
|
PPARGC1 and ESRR induced regulator, muscle 1 |
| chr14_+_75373915 | 0.14 |
ENSMUST00000122840.8
|
Lcp1
|
lymphocyte cytosolic protein 1 |
| chr11_+_98274637 | 0.14 |
ENSMUST00000008021.3
|
Tcap
|
titin-cap |
| chr4_-_11076159 | 0.13 |
ENSMUST00000058183.9
|
Ndufaf6
|
NADH:ubiquinone oxidoreductase complex assembly factor 6 |
| chr2_+_110427643 | 0.12 |
ENSMUST00000045972.13
ENSMUST00000111026.3 |
Slc5a12
|
solute carrier family 5 (sodium/glucose cotransporter), member 12 |
| chr10_+_4382467 | 0.12 |
ENSMUST00000095893.11
ENSMUST00000118544.8 ENSMUST00000117489.8 |
Armt1
|
acidic residue methyltransferase 1 |
| chr4_+_110254907 | 0.11 |
ENSMUST00000097920.9
ENSMUST00000080744.13 |
Agbl4
|
ATP/GTP binding protein-like 4 |
| chr14_+_51162425 | 0.10 |
ENSMUST00000049411.12
ENSMUST00000136753.8 ENSMUST00000154288.3 |
Apex1
|
apurinic/apyrimidinic endonuclease 1 |
| chr9_-_60594742 | 0.10 |
ENSMUST00000114032.8
ENSMUST00000166168.8 ENSMUST00000132366.2 |
Lrrc49
|
leucine rich repeat containing 49 |
| chr1_+_59296065 | 0.10 |
ENSMUST00000160662.8
ENSMUST00000114248.3 |
Cdk15
|
cyclin-dependent kinase 15 |
| chr7_+_82297803 | 0.09 |
ENSMUST00000141726.8
ENSMUST00000179489.8 ENSMUST00000039881.4 |
Efl1
|
elongation factor like GTPase 1 |
| chr17_+_38110779 | 0.09 |
ENSMUST00000215168.2
ENSMUST00000216478.2 |
Olfr124
|
olfactory receptor 124 |
| chr13_-_93328719 | 0.08 |
ENSMUST00000048702.7
|
Tent2
|
terminal nucleotidyltransferase 2 |
| chr17_+_75772567 | 0.08 |
ENSMUST00000234660.2
|
Rasgrp3
|
RAS, guanyl releasing protein 3 |
| chr2_-_62404195 | 0.08 |
ENSMUST00000174234.8
ENSMUST00000000402.16 ENSMUST00000174448.8 ENSMUST00000102732.10 |
Fap
|
fibroblast activation protein |
| chr17_-_26014613 | 0.08 |
ENSMUST00000235889.2
|
Gm50367
|
predicted gene, 50367 |
| chr3_+_68479578 | 0.08 |
ENSMUST00000170788.9
|
Schip1
|
schwannomin interacting protein 1 |
| chr4_+_117109204 | 0.08 |
ENSMUST00000125943.8
ENSMUST00000106434.8 |
Tmem53
|
transmembrane protein 53 |
| chr4_+_110254858 | 0.07 |
ENSMUST00000106589.9
ENSMUST00000106587.9 ENSMUST00000106591.8 ENSMUST00000106592.8 |
Agbl4
|
ATP/GTP binding protein-like 4 |
| chr19_+_10914517 | 0.06 |
ENSMUST00000037261.4
|
Ptgdr2
|
prostaglandin D2 receptor 2 |
| chr2_+_106525938 | 0.06 |
ENSMUST00000016530.14
|
Mpped2
|
metallophosphoesterase domain containing 2 |
| chr13_-_93328883 | 0.06 |
ENSMUST00000225868.2
|
Tent2
|
terminal nucleotidyltransferase 2 |
| chr1_-_74323536 | 0.05 |
ENSMUST00000190008.2
|
Aamp
|
angio-associated migratory protein |
| chr9_-_110818679 | 0.05 |
ENSMUST00000084922.6
ENSMUST00000199891.2 |
Rtp3
|
receptor transporter protein 3 |
| chr4_-_120966396 | 0.04 |
ENSMUST00000106268.4
|
Tmco2
|
transmembrane and coiled-coil domains 2 |
| chr3_+_103716836 | 0.04 |
ENSMUST00000076599.8
ENSMUST00000106824.8 ENSMUST00000106823.8 ENSMUST00000047285.7 |
Ap4b1
|
adaptor-related protein complex AP-4, beta 1 |
| chr15_-_76396151 | 0.03 |
ENSMUST00000023214.11
|
Dgat1
|
diacylglycerol O-acyltransferase 1 |
| chr4_-_82939330 | 0.03 |
ENSMUST00000071708.12
|
Frem1
|
Fras1 related extracellular matrix protein 1 |
| chr14_+_58035614 | 0.03 |
ENSMUST00000128764.9
|
Sap18
|
Sin3-associated polypeptide 18 |
| chr19_+_5687503 | 0.03 |
ENSMUST00000025867.6
|
Rela
|
v-rel reticuloendotheliosis viral oncogene homolog A (avian) |
| chr17_-_52117894 | 0.03 |
ENSMUST00000176669.8
|
Satb1
|
special AT-rich sequence binding protein 1 |
| chr1_-_164285914 | 0.02 |
ENSMUST00000027863.13
|
Atp1b1
|
ATPase, Na+/K+ transporting, beta 1 polypeptide |
| chr3_+_86131970 | 0.01 |
ENSMUST00000192145.6
ENSMUST00000194759.6 ENSMUST00000107635.7 |
Lrba
|
LPS-responsive beige-like anchor |
| chr3_-_79535966 | 0.01 |
ENSMUST00000120992.8
|
Etfdh
|
electron transferring flavoprotein, dehydrogenase |
| chr14_-_52252432 | 0.00 |
ENSMUST00000226527.2
|
Zfp219
|
zinc finger protein 219 |
| chr7_-_126483851 | 0.00 |
ENSMUST00000071268.11
ENSMUST00000117394.2 |
Taok2
|
TAO kinase 2 |
| chr17_+_35069347 | 0.00 |
ENSMUST00000097343.11
ENSMUST00000173357.8 ENSMUST00000173065.8 ENSMUST00000165953.3 |
Nelfe
|
negative elongation factor complex member E, Rdbp |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 2.9 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 0.9 | 2.8 | GO:1904924 | negative regulation of mitophagy in response to mitochondrial depolarization(GO:1904924) |
| 0.4 | 1.7 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.3 | 1.3 | GO:0061235 | regulation of pronephros size(GO:0035565) mesonephros morphogenesis(GO:0061206) mesonephric nephron development(GO:0061215) mesonephric nephron morphogenesis(GO:0061228) mesenchymal stem cell maintenance involved in mesonephric nephron morphogenesis(GO:0061235) regulation of mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:0061295) negative regulation of mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:0061296) mesenchymal cell apoptotic process involved in mesonephric nephron morphogenesis(GO:1901146) |
| 0.3 | 3.0 | GO:0030240 | skeletal muscle thin filament assembly(GO:0030240) |
| 0.3 | 1.1 | GO:0071544 | diphosphoinositol polyphosphate catabolic process(GO:0071544) |
| 0.3 | 1.0 | GO:0014057 | positive regulation of acetylcholine secretion, neurotransmission(GO:0014057) |
| 0.3 | 0.8 | GO:0098749 | cerebellar neuron development(GO:0098749) |
| 0.2 | 2.3 | GO:0098937 | dendritic transport(GO:0098935) anterograde dendritic transport(GO:0098937) |
| 0.2 | 6.9 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.2 | 1.1 | GO:0070295 | glycerol transport(GO:0015793) renal water absorption(GO:0070295) |
| 0.2 | 2.0 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.2 | 0.8 | GO:0043602 | nitrate catabolic process(GO:0043602) nitric oxide catabolic process(GO:0046210) cellular organohalogen metabolic process(GO:0090345) cellular organofluorine metabolic process(GO:0090346) |
| 0.2 | 1.1 | GO:2000327 | regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000325) positive regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000327) |
| 0.2 | 4.3 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.2 | 0.5 | GO:2000097 | regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 0.2 | 1.4 | GO:0045084 | positive regulation of interleukin-12 biosynthetic process(GO:0045084) |
| 0.1 | 1.0 | GO:0009750 | response to fructose(GO:0009750) |
| 0.1 | 1.9 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.1 | 1.2 | GO:0015879 | carnitine transport(GO:0015879) |
| 0.1 | 0.5 | GO:0035521 | monoubiquitinated histone deubiquitination(GO:0035521) monoubiquitinated histone H2A deubiquitination(GO:0035522) |
| 0.1 | 1.3 | GO:0032485 | regulation of Ral protein signal transduction(GO:0032485) |
| 0.1 | 0.5 | GO:0014873 | response to muscle activity involved in regulation of muscle adaptation(GO:0014873) |
| 0.1 | 0.6 | GO:1990928 | response to amino acid starvation(GO:1990928) |
| 0.1 | 0.6 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.1 | 0.4 | GO:1902396 | protein localization to bicellular tight junction(GO:1902396) |
| 0.1 | 1.2 | GO:0090500 | lymphatic endothelial cell differentiation(GO:0060836) endocardial cushion to mesenchymal transition(GO:0090500) |
| 0.1 | 0.6 | GO:1990481 | mRNA pseudouridine synthesis(GO:1990481) |
| 0.1 | 0.3 | GO:1904268 | regulation of Schwann cell chemotaxis(GO:1904266) positive regulation of Schwann cell chemotaxis(GO:1904268) Schwann cell chemotaxis(GO:1990751) |
| 0.1 | 1.9 | GO:0006744 | ubiquinone biosynthetic process(GO:0006744) |
| 0.1 | 1.8 | GO:0075522 | IRES-dependent viral translational initiation(GO:0075522) |
| 0.1 | 0.5 | GO:0071985 | multivesicular body sorting pathway(GO:0071985) |
| 0.1 | 0.2 | GO:0070782 | phosphatidylserine exposure on apoptotic cell surface(GO:0070782) |
| 0.1 | 0.4 | GO:0017182 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.1 | 2.2 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.1 | 1.4 | GO:0031115 | negative regulation of microtubule polymerization(GO:0031115) |
| 0.0 | 0.7 | GO:1901857 | negative regulation of mitochondrial membrane potential(GO:0010917) positive regulation of cellular respiration(GO:1901857) |
| 0.0 | 0.8 | GO:0045199 | maintenance of epithelial cell apical/basal polarity(GO:0045199) |
| 0.0 | 1.0 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.0 | 0.3 | GO:0045110 | intermediate filament bundle assembly(GO:0045110) |
| 0.0 | 0.2 | GO:0006114 | glycerol biosynthetic process(GO:0006114) positive regulation of transcription from RNA polymerase II promoter in response to acidic pH(GO:0061402) |
| 0.0 | 1.7 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 1.9 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 0.2 | GO:0035609 | C-terminal protein deglutamylation(GO:0035609) |
| 0.0 | 0.4 | GO:0042760 | very long-chain fatty acid catabolic process(GO:0042760) |
| 0.0 | 1.1 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 0.4 | GO:0035871 | protein K11-linked deubiquitination(GO:0035871) |
| 0.0 | 0.6 | GO:0071803 | positive regulation of podosome assembly(GO:0071803) |
| 0.0 | 0.1 | GO:0010710 | regulation of collagen catabolic process(GO:0010710) melanocyte proliferation(GO:0097325) |
| 0.0 | 1.0 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.0 | 0.5 | GO:0045899 | positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.0 | 0.7 | GO:0015986 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.0 | 1.1 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.0 | 0.1 | GO:0042256 | mature ribosome assembly(GO:0042256) |
| 0.0 | 0.0 | GO:1901738 | regulation of vitamin A metabolic process(GO:1901738) |
| 0.0 | 0.1 | GO:2000255 | negative regulation of male germ cell proliferation(GO:2000255) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.9 | GO:0036284 | tubulobulbar complex(GO:0036284) |
| 0.3 | 1.7 | GO:0005683 | U7 snRNP(GO:0005683) |
| 0.3 | 1.9 | GO:0036396 | MIS complex(GO:0036396) |
| 0.2 | 1.9 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.2 | 2.8 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.2 | 1.7 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
| 0.2 | 1.8 | GO:0071541 | eukaryotic translation initiation factor 3 complex, eIF3m(GO:0071541) |
| 0.1 | 1.1 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.1 | 0.5 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.1 | 0.6 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.1 | 0.8 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.0 | 2.8 | GO:0005865 | striated muscle thin filament(GO:0005865) |
| 0.0 | 1.0 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 0.0 | 2.2 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.0 | 0.7 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 1.2 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.0 | 3.3 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 1.5 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.0 | 1.0 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.0 | 2.2 | GO:0005762 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 1.0 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.0 | 0.4 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.0 | 0.4 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.0 | 0.4 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.0 | 0.3 | GO:0060091 | kinocilium(GO:0060091) |
| 0.0 | 0.6 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 0.6 | GO:0005763 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 1.1 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 1.5 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 1.4 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 1.5 | GO:0097517 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 0.4 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 0.1 | GO:0071438 | invadopodium membrane(GO:0071438) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.7 | GO:0061711 | N(6)-L-threonylcarbamoyladenine synthase(GO:0061711) |
| 0.5 | 1.9 | GO:0050347 | trans-hexaprenyltranstransferase activity(GO:0000010) trans-octaprenyltranstransferase activity(GO:0050347) |
| 0.5 | 1.4 | GO:0052658 | inositol-1,4,5-trisphosphate 5-phosphatase activity(GO:0052658) |
| 0.2 | 1.1 | GO:0015254 | glycerol channel activity(GO:0015254) |
| 0.2 | 2.0 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.2 | 0.8 | GO:0008941 | nitric oxide dioxygenase activity(GO:0008941) oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of two atoms of oxygen into one donor(GO:0016708) iron-cytochrome-c reductase activity(GO:0047726) |
| 0.2 | 1.2 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.2 | 1.1 | GO:0000298 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.2 | 1.2 | GO:0005124 | scavenger receptor binding(GO:0005124) |
| 0.1 | 0.4 | GO:0005169 | neurotrophin TRKB receptor binding(GO:0005169) |
| 0.1 | 0.7 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.1 | 1.0 | GO:0070095 | fructose-6-phosphate binding(GO:0070095) |
| 0.1 | 1.0 | GO:0001609 | G-protein coupled adenosine receptor activity(GO:0001609) |
| 0.1 | 1.5 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.1 | 2.9 | GO:0071949 | FAD binding(GO:0071949) |
| 0.1 | 2.0 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.1 | 1.0 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.1 | 0.8 | GO:0001055 | RNA polymerase I activity(GO:0001054) RNA polymerase II activity(GO:0001055) |
| 0.1 | 0.6 | GO:0004311 | farnesyltranstransferase activity(GO:0004311) |
| 0.1 | 0.2 | GO:0004611 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 0.0 | 0.3 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.0 | 0.7 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 1.0 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.1 | GO:0016890 | site-specific endodeoxyribonuclease activity, specific for altered base(GO:0016890) |
| 0.0 | 2.3 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.6 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.0 | 0.8 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.0 | 0.3 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.0 | 0.1 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.0 | 1.4 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 2.2 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.0 | 0.7 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.0 | 0.1 | GO:0004958 | prostaglandin F receptor activity(GO:0004958) |
| 0.0 | 1.1 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 0.5 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 2.2 | GO:0008565 | protein transporter activity(GO:0008565) |
| 0.0 | 0.5 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.0 | 0.1 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 2.6 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 1.7 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 0.1 | GO:0010340 | carboxyl-O-methyltransferase activity(GO:0010340) protein carboxyl O-methyltransferase activity(GO:0051998) |
| 0.0 | 0.3 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 0.2 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.8 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 1.8 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 1.6 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 1.2 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.0 | 1.9 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 1.5 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.8 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 1.2 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 0.3 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.2 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.1 | 1.7 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.1 | 2.0 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 1.1 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.0 | 0.8 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 2.8 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 1.0 | REACTOME NUCLEOTIDE LIKE PURINERGIC RECEPTORS | Genes involved in Nucleotide-like (purinergic) receptors |
| 0.0 | 0.8 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.0 | 1.0 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.0 | 1.0 | REACTOME REGULATION OF GLUCOKINASE BY GLUCOKINASE REGULATORY PROTEIN | Genes involved in Regulation of Glucokinase by Glucokinase Regulatory Protein |
| 0.0 | 1.3 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.0 | 1.4 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.0 | 0.4 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 0.4 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.0 | 0.5 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.0 | 0.5 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.0 | 0.2 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |