Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
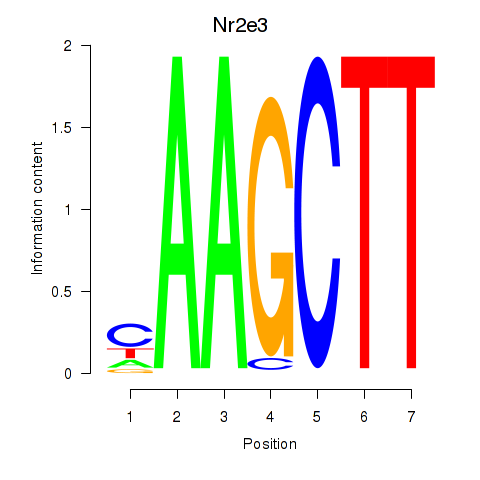
Results for Nr2e3
Z-value: 1.80
Transcription factors associated with Nr2e3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Nr2e3
|
ENSMUSG00000032292.9 | Nr2e3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Nr2e3 | mm39_v1_chr9_-_59857355_59857407 | 0.30 | 7.1e-02 | Click! |
Activity profile of Nr2e3 motif
Sorted Z-values of Nr2e3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Nr2e3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_-_90603013 | 21.51 |
ENSMUST00000069960.12
ENSMUST00000117167.2 |
S100a9
|
S100 calcium binding protein A9 (calgranulin B) |
| chr7_+_100143250 | 9.03 |
ENSMUST00000153287.8
|
Ucp2
|
uncoupling protein 2 (mitochondrial, proton carrier) |
| chr7_-_110462446 | 6.46 |
ENSMUST00000033050.5
|
Lyve1
|
lymphatic vessel endothelial hyaluronan receptor 1 |
| chr19_+_38121248 | 6.38 |
ENSMUST00000025956.13
|
Pde6c
|
phosphodiesterase 6C, cGMP specific, cone, alpha prime |
| chr19_+_38121214 | 6.22 |
ENSMUST00000112329.3
|
Pde6c
|
phosphodiesterase 6C, cGMP specific, cone, alpha prime |
| chr11_-_55075855 | 6.19 |
ENSMUST00000039305.6
|
Slc36a2
|
solute carrier family 36 (proton/amino acid symporter), member 2 |
| chr4_+_123176570 | 5.85 |
ENSMUST00000106243.8
ENSMUST00000106241.8 ENSMUST00000080178.13 |
Pabpc4
|
poly(A) binding protein, cytoplasmic 4 |
| chr7_-_103492361 | 5.64 |
ENSMUST00000063957.6
|
Hbb-bh1
|
hemoglobin Z, beta-like embryonic chain |
| chr6_+_142244145 | 4.72 |
ENSMUST00000041993.3
|
Iapp
|
islet amyloid polypeptide |
| chr2_-_164198427 | 4.49 |
ENSMUST00000109367.10
|
Slpi
|
secretory leukocyte peptidase inhibitor |
| chr2_-_164197987 | 4.37 |
ENSMUST00000165980.2
|
Slpi
|
secretory leukocyte peptidase inhibitor |
| chr3_-_10273628 | 4.28 |
ENSMUST00000029041.6
|
Fabp4
|
fatty acid binding protein 4, adipocyte |
| chr3_-_137687284 | 4.20 |
ENSMUST00000136613.4
ENSMUST00000029806.13 |
Dapp1
|
dual adaptor for phosphotyrosine and 3-phosphoinositides 1 |
| chr8_+_84335176 | 3.91 |
ENSMUST00000212300.2
|
Dnajb1
|
DnaJ heat shock protein family (Hsp40) member B1 |
| chr3_-_88410495 | 3.79 |
ENSMUST00000120377.8
ENSMUST00000029699.13 |
Lmna
|
lamin A |
| chr7_-_19005721 | 3.36 |
ENSMUST00000032561.9
|
Vasp
|
vasodilator-stimulated phosphoprotein |
| chr9_+_106080307 | 3.05 |
ENSMUST00000024047.12
ENSMUST00000216348.2 |
Twf2
|
twinfilin actin binding protein 2 |
| chr13_-_29137673 | 3.02 |
ENSMUST00000067230.6
|
Sox4
|
SRY (sex determining region Y)-box 4 |
| chr9_+_65494469 | 2.85 |
ENSMUST00000239405.2
ENSMUST00000047099.13 ENSMUST00000131483.3 ENSMUST00000141046.3 |
Pif1
|
PIF1 5'-to-3' DNA helicase |
| chr16_+_17051423 | 2.79 |
ENSMUST00000090190.14
ENSMUST00000232082.2 ENSMUST00000232426.2 |
Hic2
Gm49573
|
hypermethylated in cancer 2 predicted gene, 49573 |
| chr8_+_89015705 | 2.65 |
ENSMUST00000171456.9
|
Adcy7
|
adenylate cyclase 7 |
| chr11_+_69804714 | 2.63 |
ENSMUST00000072581.9
ENSMUST00000116358.8 |
Gps2
|
G protein pathway suppressor 2 |
| chr4_+_119052476 | 2.59 |
ENSMUST00000030395.9
|
Svbp
|
small vasohibin binding protein |
| chr14_+_76741918 | 2.46 |
ENSMUST00000022587.10
ENSMUST00000134109.2 |
Tsc22d1
|
TSC22 domain family, member 1 |
| chr10_-_45346297 | 2.40 |
ENSMUST00000079390.7
|
Lin28b
|
lin-28 homolog B (C. elegans) |
| chr11_+_67167950 | 2.39 |
ENSMUST00000019625.12
|
Myh8
|
myosin, heavy polypeptide 8, skeletal muscle, perinatal |
| chr9_+_113641615 | 2.39 |
ENSMUST00000111838.10
ENSMUST00000166734.10 ENSMUST00000214522.2 ENSMUST00000163895.3 |
Clasp2
|
CLIP associating protein 2 |
| chr5_-_22755274 | 2.32 |
ENSMUST00000030872.12
|
Orc5
|
origin recognition complex, subunit 5 |
| chr14_+_76741625 | 2.30 |
ENSMUST00000177207.2
|
Tsc22d1
|
TSC22 domain family, member 1 |
| chr4_+_119052693 | 2.25 |
ENSMUST00000097908.4
|
Svbp
|
small vasohibin binding protein |
| chrX_+_20483742 | 2.20 |
ENSMUST00000115375.8
ENSMUST00000115374.8 ENSMUST00000084383.10 |
Rbm10
|
RNA binding motif protein 10 |
| chrX_-_93585668 | 2.17 |
ENSMUST00000026142.8
|
Maged1
|
MAGE family member D1 |
| chr6_-_36787096 | 2.13 |
ENSMUST00000201321.2
ENSMUST00000101534.5 |
Ptn
|
pleiotrophin |
| chr11_-_100986192 | 2.10 |
ENSMUST00000019447.15
|
Psmc3ip
|
proteasome (prosome, macropain) 26S subunit, ATPase 3, interacting protein |
| chr10_-_119948890 | 2.07 |
ENSMUST00000020449.12
|
Helb
|
helicase (DNA) B |
| chr17_+_35460722 | 2.03 |
ENSMUST00000068056.12
ENSMUST00000174757.8 ENSMUST00000173731.8 |
Ddx39b
|
DEAD box helicase 39b |
| chr13_+_112425322 | 2.00 |
ENSMUST00000022275.14
|
Ankrd55
|
ankyrin repeat domain 55 |
| chr7_+_30450896 | 1.99 |
ENSMUST00000182229.8
ENSMUST00000080518.14 ENSMUST00000182227.8 ENSMUST00000182721.8 |
Sbsn
|
suprabasin |
| chr7_+_4918199 | 1.93 |
ENSMUST00000116354.4
|
Zfp628
|
zinc finger protein 628 |
| chr8_+_79755194 | 1.91 |
ENSMUST00000119254.8
ENSMUST00000238669.2 |
Zfp827
|
zinc finger protein 827 |
| chr7_+_101714943 | 1.91 |
ENSMUST00000094130.4
ENSMUST00000084843.10 |
Xndc1
Xntrpc
|
Xrcc1 N-terminal domain containing 1 Xndc1-transient receptor potential cation channel, subfamily C, member 2 readthrough |
| chr7_-_110673269 | 1.90 |
ENSMUST00000163014.2
|
Eif4g2
|
eukaryotic translation initiation factor 4, gamma 2 |
| chr5_+_76331727 | 1.87 |
ENSMUST00000031144.14
|
Tmem165
|
transmembrane protein 165 |
| chr19_-_7318798 | 1.87 |
ENSMUST00000165965.8
ENSMUST00000051711.16 ENSMUST00000169541.8 ENSMUST00000165989.2 |
Mark2
|
MAP/microtubule affinity regulating kinase 2 |
| chr4_+_119052548 | 1.86 |
ENSMUST00000106345.3
|
Svbp
|
small vasohibin binding protein |
| chrX_+_168662592 | 1.85 |
ENSMUST00000112105.8
ENSMUST00000078947.12 |
Mid1
|
midline 1 |
| chr6_+_136509922 | 1.81 |
ENSMUST00000187429.4
|
Atf7ip
|
activating transcription factor 7 interacting protein |
| chr5_-_65117375 | 1.79 |
ENSMUST00000062315.7
ENSMUST00000239485.2 ENSMUST00000201307.3 |
Tlr6
|
toll-like receptor 6 |
| chr2_+_124452194 | 1.79 |
ENSMUST00000051419.15
ENSMUST00000076335.12 ENSMUST00000078621.12 ENSMUST00000077847.12 |
Sema6d
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D |
| chr12_-_54842488 | 1.78 |
ENSMUST00000005798.9
|
Snx6
|
sorting nexin 6 |
| chr15_+_6329263 | 1.76 |
ENSMUST00000078019.13
|
Dab2
|
disabled 2, mitogen-responsive phosphoprotein |
| chr1_+_90131702 | 1.74 |
ENSMUST00000065587.5
ENSMUST00000159654.2 |
Ackr3
|
atypical chemokine receptor 3 |
| chr2_-_45002902 | 1.74 |
ENSMUST00000076836.13
ENSMUST00000176732.8 ENSMUST00000200844.4 |
Zeb2
|
zinc finger E-box binding homeobox 2 |
| chr19_-_40365318 | 1.64 |
ENSMUST00000239304.2
|
Sorbs1
|
sorbin and SH3 domain containing 1 |
| chr16_-_25924527 | 1.58 |
ENSMUST00000039990.6
|
P3h2
|
prolyl 3-hydroxylase 2 |
| chr12_-_110669076 | 1.58 |
ENSMUST00000155242.8
|
Hsp90aa1
|
heat shock protein 90, alpha (cytosolic), class A member 1 |
| chr13_+_35925296 | 1.56 |
ENSMUST00000163595.3
|
Cdyl
|
chromodomain protein, Y chromosome-like |
| chr17_+_75485791 | 1.54 |
ENSMUST00000135447.8
ENSMUST00000112516.8 |
Ltbp1
|
latent transforming growth factor beta binding protein 1 |
| chr15_+_6329278 | 1.47 |
ENSMUST00000159046.2
ENSMUST00000161040.8 |
Dab2
|
disabled 2, mitogen-responsive phosphoprotein |
| chr8_+_105267431 | 1.47 |
ENSMUST00000056051.11
|
Car7
|
carbonic anhydrase 7 |
| chr2_+_24839758 | 1.45 |
ENSMUST00000028350.9
|
Zmynd19
|
zinc finger, MYND domain containing 19 |
| chr8_+_79754980 | 1.42 |
ENSMUST00000087927.11
ENSMUST00000098614.9 |
Zfp827
|
zinc finger protein 827 |
| chr7_-_118091135 | 1.41 |
ENSMUST00000178344.3
|
Itpripl2
|
inositol 1,4,5-triphosphate receptor interacting protein-like 2 |
| chr5_-_65492940 | 1.39 |
ENSMUST00000203471.3
ENSMUST00000172732.8 ENSMUST00000204965.3 |
Rfc1
|
replication factor C (activator 1) 1 |
| chr3_-_79053182 | 1.37 |
ENSMUST00000118340.7
|
Rapgef2
|
Rap guanine nucleotide exchange factor (GEF) 2 |
| chrX_-_7054952 | 1.34 |
ENSMUST00000004428.14
|
Clcn5
|
chloride channel, voltage-sensitive 5 |
| chr7_+_92426772 | 1.32 |
ENSMUST00000208945.2
|
Rab30
|
RAB30, member RAS oncogene family |
| chr7_+_101714692 | 1.32 |
ENSMUST00000106950.8
ENSMUST00000146450.8 |
Xndc1
|
Xrcc1 N-terminal domain containing 1 |
| chrX_+_41238193 | 1.29 |
ENSMUST00000115073.9
ENSMUST00000115072.8 |
Stag2
|
stromal antigen 2 |
| chr8_+_84334805 | 1.29 |
ENSMUST00000005620.10
|
Dnajb1
|
DnaJ heat shock protein family (Hsp40) member B1 |
| chr13_-_74882374 | 1.29 |
ENSMUST00000220738.2
|
Cast
|
calpastatin |
| chr5_+_65505657 | 1.26 |
ENSMUST00000031096.11
|
Klb
|
klotho beta |
| chr2_+_124452493 | 1.26 |
ENSMUST00000103239.10
ENSMUST00000103240.9 |
Sema6d
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D |
| chr2_-_126517458 | 1.25 |
ENSMUST00000103227.8
ENSMUST00000110425.9 ENSMUST00000089745.11 |
Gabpb1
|
GA repeat binding protein, beta 1 |
| chr5_+_35156454 | 1.25 |
ENSMUST00000114283.8
|
Rgs12
|
regulator of G-protein signaling 12 |
| chr6_-_129599645 | 1.23 |
ENSMUST00000032252.8
|
Klrk1
|
killer cell lectin-like receptor subfamily K, member 1 |
| chr4_-_129534403 | 1.23 |
ENSMUST00000084264.12
|
Txlna
|
taxilin alpha |
| chr19_+_39102342 | 1.23 |
ENSMUST00000087234.3
|
Cyp2c66
|
cytochrome P450, family 2, subfamily c, polypeptide 66 |
| chr2_-_126517383 | 1.20 |
ENSMUST00000103226.10
ENSMUST00000110424.9 |
Gabpb1
|
GA repeat binding protein, beta 1 |
| chr4_-_109522502 | 1.18 |
ENSMUST00000063531.5
|
Cdkn2c
|
cyclin dependent kinase inhibitor 2C |
| chr16_+_31241085 | 1.17 |
ENSMUST00000089759.9
|
Bdh1
|
3-hydroxybutyrate dehydrogenase, type 1 |
| chr2_+_18059982 | 1.16 |
ENSMUST00000028076.15
|
Mllt10
|
myeloid/lymphoid or mixed-lineage leukemia; translocated to, 10 |
| chr18_+_35347983 | 1.12 |
ENSMUST00000235449.2
ENSMUST00000235269.2 |
Ctnna1
|
catenin (cadherin associated protein), alpha 1 |
| chr13_-_74882328 | 1.07 |
ENSMUST00000223309.2
|
Cast
|
calpastatin |
| chr6_-_122317484 | 1.07 |
ENSMUST00000112600.9
|
Phc1
|
polyhomeotic 1 |
| chr8_+_107237483 | 1.05 |
ENSMUST00000080797.8
|
Cdh3
|
cadherin 3 |
| chrX_+_7656225 | 1.04 |
ENSMUST00000136930.8
ENSMUST00000115675.9 ENSMUST00000101694.10 |
Gripap1
|
GRIP1 associated protein 1 |
| chr10_+_119655294 | 1.03 |
ENSMUST00000105262.9
ENSMUST00000147454.8 ENSMUST00000138410.8 ENSMUST00000144825.8 ENSMUST00000148954.8 ENSMUST00000144959.8 |
Grip1
|
glutamate receptor interacting protein 1 |
| chr8_-_58106057 | 1.01 |
ENSMUST00000034021.12
|
Galnt7
|
polypeptide N-acetylgalactosaminyltransferase 7 |
| chr8_-_61407760 | 1.00 |
ENSMUST00000110302.8
|
Clcn3
|
chloride channel, voltage-sensitive 3 |
| chr2_+_127750978 | 0.99 |
ENSMUST00000110344.2
|
Acoxl
|
acyl-Coenzyme A oxidase-like |
| chr11_+_100210705 | 0.98 |
ENSMUST00000049385.14
|
Eif1
|
eukaryotic translation initiation factor 1 |
| chr11_+_67090878 | 0.98 |
ENSMUST00000124516.8
ENSMUST00000018637.15 ENSMUST00000129018.8 |
Myh1
|
myosin, heavy polypeptide 1, skeletal muscle, adult |
| chr17_+_75485906 | 0.97 |
ENSMUST00000112514.2
|
Ltbp1
|
latent transforming growth factor beta binding protein 1 |
| chr15_-_80989200 | 0.97 |
ENSMUST00000109579.9
|
Mrtfa
|
myocardin related transcription factor A |
| chr9_+_65494429 | 0.96 |
ENSMUST00000134538.9
|
Pif1
|
PIF1 5'-to-3' DNA helicase |
| chr13_-_92931317 | 0.88 |
ENSMUST00000022213.8
|
Thbs4
|
thrombospondin 4 |
| chr8_-_58106027 | 0.87 |
ENSMUST00000110316.3
|
Galnt7
|
polypeptide N-acetylgalactosaminyltransferase 7 |
| chr5_-_69699965 | 0.85 |
ENSMUST00000031045.10
|
Yipf7
|
Yip1 domain family, member 7 |
| chr4_-_49521036 | 0.82 |
ENSMUST00000057829.4
|
Mrpl50
|
mitochondrial ribosomal protein L50 |
| chr15_-_76906832 | 0.82 |
ENSMUST00000019037.10
ENSMUST00000169226.9 |
Mb
|
myoglobin |
| chr1_-_97904958 | 0.82 |
ENSMUST00000161567.8
|
Pam
|
peptidylglycine alpha-amidating monooxygenase |
| chr14_+_20724366 | 0.81 |
ENSMUST00000048657.10
|
Sec24c
|
Sec24 related gene family, member C (S. cerevisiae) |
| chr11_-_65053710 | 0.76 |
ENSMUST00000093002.12
ENSMUST00000047463.15 |
Arhgap44
|
Rho GTPase activating protein 44 |
| chr5_-_69699932 | 0.76 |
ENSMUST00000202423.2
|
Yipf7
|
Yip1 domain family, member 7 |
| chrX_-_58211440 | 0.73 |
ENSMUST00000119306.2
|
Fgf13
|
fibroblast growth factor 13 |
| chr9_-_111086528 | 0.71 |
ENSMUST00000199404.2
|
Mlh1
|
mutL homolog 1 |
| chr3_-_110158280 | 0.69 |
ENSMUST00000190378.2
ENSMUST00000106567.2 |
Prmt6
|
protein arginine N-methyltransferase 6 |
| chr6_-_68887957 | 0.69 |
ENSMUST00000200454.2
|
Igkv4-86
|
immunoglobulin kappa variable 4-86 |
| chrX_+_41241049 | 0.68 |
ENSMUST00000128799.3
|
Stag2
|
stromal antigen 2 |
| chr17_+_87270504 | 0.66 |
ENSMUST00000024956.15
|
Rhoq
|
ras homolog family member Q |
| chrX_+_7656251 | 0.65 |
ENSMUST00000140540.2
|
Gripap1
|
GRIP1 associated protein 1 |
| chr14_+_20724378 | 0.63 |
ENSMUST00000224492.2
ENSMUST00000223751.2 ENSMUST00000225108.2 ENSMUST00000224754.2 |
Sec24c
|
Sec24 related gene family, member C (S. cerevisiae) |
| chr2_+_65451100 | 0.62 |
ENSMUST00000144254.6
ENSMUST00000028377.14 |
Scn2a
|
sodium channel, voltage-gated, type II, alpha |
| chr19_-_45619559 | 0.61 |
ENSMUST00000160718.9
|
Fbxw4
|
F-box and WD-40 domain protein 4 |
| chr1_-_37575313 | 0.58 |
ENSMUST00000042161.15
|
Mgat4a
|
mannoside acetylglucosaminyltransferase 4, isoenzyme A |
| chr9_-_77255069 | 0.58 |
ENSMUST00000184848.8
ENSMUST00000184415.8 |
Mlip
|
muscular LMNA-interacting protein |
| chr5_-_65492907 | 0.57 |
ENSMUST00000203581.3
|
Rfc1
|
replication factor C (activator 1) 1 |
| chr7_-_105218472 | 0.57 |
ENSMUST00000187683.7
ENSMUST00000210079.2 ENSMUST00000187051.7 ENSMUST00000189265.7 ENSMUST00000190369.7 |
Apbb1
|
amyloid beta (A4) precursor protein-binding, family B, member 1 |
| chr18_+_35695485 | 0.56 |
ENSMUST00000235199.2
ENSMUST00000237744.2 ENSMUST00000236276.2 |
Matr3
|
matrin 3 |
| chr1_-_179572765 | 0.55 |
ENSMUST00000211943.3
ENSMUST00000131716.4 ENSMUST00000221136.2 |
Kif28
|
kinesin family member 28 |
| chr15_-_81756076 | 0.55 |
ENSMUST00000023117.10
|
Phf5a
|
PHD finger protein 5A |
| chr9_-_20085353 | 0.53 |
ENSMUST00000215984.3
|
Olfr870
|
olfactory receptor 870 |
| chr3_-_143908111 | 0.52 |
ENSMUST00000121796.8
|
Lmo4
|
LIM domain only 4 |
| chr2_+_104961228 | 0.51 |
ENSMUST00000111098.8
ENSMUST00000111099.2 |
Wt1
|
Wilms tumor 1 homolog |
| chr8_-_24928953 | 0.49 |
ENSMUST00000052622.6
|
Tcim
|
transcriptional and immune response regulator |
| chr5_-_5315968 | 0.48 |
ENSMUST00000115451.8
ENSMUST00000115452.8 ENSMUST00000131392.8 |
Cdk14
|
cyclin-dependent kinase 14 |
| chr4_-_136776006 | 0.44 |
ENSMUST00000049583.8
|
Zbtb40
|
zinc finger and BTB domain containing 40 |
| chr1_-_160040286 | 0.44 |
ENSMUST00000195654.2
ENSMUST00000014370.11 |
Cacybp
|
calcyclin binding protein |
| chr9_-_77255099 | 0.43 |
ENSMUST00000184138.8
ENSMUST00000184006.8 ENSMUST00000185144.8 ENSMUST00000034910.16 |
Mlip
|
muscular LMNA-interacting protein |
| chr3_-_143908060 | 0.41 |
ENSMUST00000121112.6
|
Lmo4
|
LIM domain only 4 |
| chr13_+_49697919 | 0.39 |
ENSMUST00000177948.2
ENSMUST00000021820.14 |
Aspn
|
asporin |
| chr8_-_55171699 | 0.37 |
ENSMUST00000144711.9
|
Wdr17
|
WD repeat domain 17 |
| chr1_-_58463200 | 0.35 |
ENSMUST00000034868.14
|
Clk1
|
CDC-like kinase 1 |
| chr7_+_120634834 | 0.33 |
ENSMUST00000207351.2
|
Mettl9
|
methyltransferase like 9 |
| chr3_+_103739877 | 0.32 |
ENSMUST00000062945.12
|
Bcl2l15
|
BCLl2-like 15 |
| chr10_-_129524028 | 0.31 |
ENSMUST00000203785.3
ENSMUST00000217576.2 |
Olfr802
|
olfactory receptor 802 |
| chr8_+_22717338 | 0.31 |
ENSMUST00000160585.2
ENSMUST00000162447.2 |
Thsd1
|
thrombospondin, type I, domain 1 |
| chr16_-_16377982 | 0.30 |
ENSMUST00000161861.8
|
Fgd4
|
FYVE, RhoGEF and PH domain containing 4 |
| chr4_-_154721288 | 0.30 |
ENSMUST00000030902.13
ENSMUST00000105637.8 ENSMUST00000070313.14 ENSMUST00000105636.8 ENSMUST00000105638.9 ENSMUST00000097759.9 ENSMUST00000124771.2 |
Prdm16
|
PR domain containing 16 |
| chr2_-_25471703 | 0.29 |
ENSMUST00000114217.3
ENSMUST00000191602.2 |
Ajm1
|
apical junction component 1 |
| chr16_-_22258469 | 0.28 |
ENSMUST00000079601.13
|
Etv5
|
ets variant 5 |
| chr7_-_105217851 | 0.28 |
ENSMUST00000188368.7
ENSMUST00000187057.7 |
Apbb1
|
amyloid beta (A4) precursor protein-binding, family B, member 1 |
| chr18_+_61688378 | 0.27 |
ENSMUST00000165721.8
ENSMUST00000115246.9 ENSMUST00000166990.8 ENSMUST00000163205.8 ENSMUST00000170862.8 |
Csnk1a1
|
casein kinase 1, alpha 1 |
| chrX_-_133012600 | 0.27 |
ENSMUST00000033610.13
|
Nox1
|
NADPH oxidase 1 |
| chr1_+_177273226 | 0.26 |
ENSMUST00000077225.8
|
Zbtb18
|
zinc finger and BTB domain containing 18 |
| chr16_-_63684477 | 0.26 |
ENSMUST00000232654.2
ENSMUST00000064405.8 |
Epha3
|
Eph receptor A3 |
| chr6_+_34840057 | 0.25 |
ENSMUST00000074949.4
|
Tmem140
|
transmembrane protein 140 |
| chr1_+_97697881 | 0.25 |
ENSMUST00000112844.10
ENSMUST00000112842.8 ENSMUST00000027571.13 |
Gin1
|
gypsy retrotransposon integrase 1 |
| chr19_+_4264470 | 0.23 |
ENSMUST00000237171.2
|
Gm45928
|
predicted gene, 45928 |
| chr11_+_19874354 | 0.23 |
ENSMUST00000093299.13
|
Spred2
|
sprouty-related EVH1 domain containing 2 |
| chr11_+_93934940 | 0.21 |
ENSMUST00000132079.8
|
Spag9
|
sperm associated antigen 9 |
| chr14_-_78866714 | 0.20 |
ENSMUST00000228362.2
ENSMUST00000227767.2 |
Dgkh
|
diacylglycerol kinase, eta |
| chr1_+_151631088 | 0.19 |
ENSMUST00000188145.7
ENSMUST00000059498.12 |
Edem3
|
ER degradation enhancer, mannosidase alpha-like 3 |
| chr12_-_115884332 | 0.18 |
ENSMUST00000103548.3
|
Ighv1-81
|
immunoglobulin heavy variable 1-81 |
| chr2_-_140513382 | 0.17 |
ENSMUST00000110057.3
|
Flrt3
|
fibronectin leucine rich transmembrane protein 3 |
| chr19_+_4264292 | 0.17 |
ENSMUST00000046506.7
|
Clcf1
|
cardiotrophin-like cytokine factor 1 |
| chr17_+_87270707 | 0.15 |
ENSMUST00000139344.2
|
Rhoq
|
ras homolog family member Q |
| chr1_+_178015287 | 0.15 |
ENSMUST00000159284.2
|
Desi2
|
desumoylating isopeptidase 2 |
| chr2_-_140513320 | 0.13 |
ENSMUST00000056760.4
|
Flrt3
|
fibronectin leucine rich transmembrane protein 3 |
| chrX_-_133012457 | 0.13 |
ENSMUST00000159259.3
ENSMUST00000113275.10 |
Nox1
|
NADPH oxidase 1 |
| chr6_-_122317457 | 0.12 |
ENSMUST00000160843.8
|
Phc1
|
polyhomeotic 1 |
| chr11_+_34264757 | 0.10 |
ENSMUST00000165963.9
ENSMUST00000093192.4 |
Insyn2b
|
inhibitory synaptic factor family member 2B |
| chr11_+_96173355 | 0.08 |
ENSMUST00000125410.2
|
Hoxb8
|
homeobox B8 |
| chr8_+_22717308 | 0.08 |
ENSMUST00000069828.10
|
Thsd1
|
thrombospondin, type I, domain 1 |
| chr1_-_75293447 | 0.08 |
ENSMUST00000189551.7
|
Dnpep
|
aspartyl aminopeptidase |
| chr12_+_95662124 | 0.07 |
ENSMUST00000110117.2
|
Flrt2
|
fibronectin leucine rich transmembrane protein 2 |
| chr14_-_62998561 | 0.07 |
ENSMUST00000053959.7
ENSMUST00000223585.2 |
Ints6
|
integrator complex subunit 6 |
| chr6_+_122530758 | 0.06 |
ENSMUST00000043301.14
|
Aicda
|
activation-induced cytidine deaminase |
| chr4_-_120808820 | 0.05 |
ENSMUST00000106280.8
|
Zfp69
|
zinc finger protein 69 |
| chr14_-_9015639 | 0.05 |
ENSMUST00000112656.4
|
Synpr
|
synaptoporin |
| chr3_+_68375495 | 0.05 |
ENSMUST00000182532.8
|
Schip1
|
schwannomin interacting protein 1 |
| chr5_+_20112704 | 0.05 |
ENSMUST00000115267.7
|
Magi2
|
membrane associated guanylate kinase, WW and PDZ domain containing 2 |
| chr18_+_65054188 | 0.05 |
ENSMUST00000236898.2
ENSMUST00000237410.2 |
Nedd4l
|
neural precursor cell expressed, developmentally down-regulated gene 4-like |
| chr11_+_19874403 | 0.04 |
ENSMUST00000093298.12
|
Spred2
|
sprouty-related EVH1 domain containing 2 |
| chr14_+_102077937 | 0.03 |
ENSMUST00000159026.8
|
Lmo7
|
LIM domain only 7 |
| chr2_+_67004178 | 0.03 |
ENSMUST00000239009.2
ENSMUST00000238912.2 |
Xirp2
|
xin actin-binding repeat containing 2 |
| chr3_+_69129745 | 0.02 |
ENSMUST00000183126.2
|
Arl14
|
ADP-ribosylation factor-like 14 |
| chr11_+_96173475 | 0.02 |
ENSMUST00000168043.2
|
Hoxb8
|
homeobox B8 |
| chr14_+_102078038 | 0.01 |
ENSMUST00000159314.8
|
Lmo7
|
LIM domain only 7 |
| chr9_-_77255171 | 0.01 |
ENSMUST00000185039.8
|
Mlip
|
muscular LMNA-interacting protein |
| chr2_+_83554741 | 0.00 |
ENSMUST00000028499.11
|
Itgav
|
integrin alpha V |
| chr16_-_63684425 | 0.00 |
ENSMUST00000232049.2
|
Epha3
|
Eph receptor A3 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.2 | 21.5 | GO:0035606 | peptidyl-cysteine S-trans-nitrosylation(GO:0035606) |
| 1.5 | 6.2 | GO:0035524 | proline transmembrane transport(GO:0035524) glycine import(GO:0036233) |
| 0.8 | 12.6 | GO:0046549 | retinal cone cell development(GO:0046549) |
| 0.7 | 6.5 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.6 | 3.8 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.6 | 3.0 | GO:0035910 | ascending aorta development(GO:0035905) ascending aorta morphogenesis(GO:0035910) |
| 0.6 | 1.8 | GO:0034136 | negative regulation of toll-like receptor 2 signaling pathway(GO:0034136) |
| 0.6 | 4.7 | GO:0097647 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.5 | 3.2 | GO:0035026 | leading edge cell differentiation(GO:0035026) |
| 0.5 | 2.1 | GO:1904395 | positive regulation of skeletal muscle acetylcholine-gated channel clustering(GO:1904395) |
| 0.5 | 2.1 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.5 | 1.4 | GO:2000670 | positive regulation of dendritic cell apoptotic process(GO:2000670) |
| 0.5 | 9.0 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.4 | 1.2 | GO:0030887 | positive regulation of myeloid dendritic cell activation(GO:0030887) |
| 0.4 | 2.4 | GO:0010587 | miRNA catabolic process(GO:0010587) |
| 0.4 | 3.8 | GO:0044806 | G-quadruplex DNA unwinding(GO:0044806) |
| 0.4 | 1.9 | GO:0032472 | Golgi calcium ion transport(GO:0032472) |
| 0.4 | 5.2 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 0.4 | 1.5 | GO:0032849 | positive regulation of cellular pH reduction(GO:0032849) |
| 0.4 | 1.1 | GO:0051794 | regulation of catagen(GO:0051794) |
| 0.3 | 1.0 | GO:0097401 | synaptic vesicle lumen acidification(GO:0097401) |
| 0.3 | 6.5 | GO:0006027 | glycosaminoglycan catabolic process(GO:0006027) |
| 0.3 | 3.1 | GO:0032532 | regulation of microvillus length(GO:0032532) |
| 0.3 | 0.9 | GO:0071603 | endothelial cell-cell adhesion(GO:0071603) |
| 0.3 | 2.0 | GO:2000002 | negative regulation of DNA damage checkpoint(GO:2000002) |
| 0.3 | 8.9 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.2 | 1.2 | GO:0038032 | termination of G-protein coupled receptor signaling pathway(GO:0038032) |
| 0.2 | 2.5 | GO:0098887 | neurotransmitter receptor transport, endosome to postsynaptic membrane(GO:0098887) |
| 0.2 | 2.4 | GO:1903690 | negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 0.2 | 0.7 | GO:0034970 | histone H3-R2 methylation(GO:0034970) |
| 0.2 | 1.6 | GO:0045583 | regulation of cytotoxic T cell differentiation(GO:0045583) positive regulation of cytotoxic T cell differentiation(GO:0045585) |
| 0.2 | 1.5 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.2 | 3.7 | GO:0071285 | cellular response to lithium ion(GO:0071285) |
| 0.2 | 1.1 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.2 | 0.7 | GO:0043060 | meiotic metaphase I plate congression(GO:0043060) meiotic metaphase plate congression(GO:0051311) |
| 0.2 | 0.5 | GO:2001076 | negative regulation of metanephric glomerulus development(GO:0072299) negative regulation of metanephric glomerular mesangial cell proliferation(GO:0072302) regulation of metanephric ureteric bud development(GO:2001074) positive regulation of metanephric ureteric bud development(GO:2001076) |
| 0.2 | 2.2 | GO:0034393 | positive regulation of smooth muscle cell apoptotic process(GO:0034393) |
| 0.2 | 1.3 | GO:0090080 | positive regulation of MAPKKK cascade by fibroblast growth factor receptor signaling pathway(GO:0090080) |
| 0.1 | 0.8 | GO:0050757 | thymidylate synthase biosynthetic process(GO:0050757) regulation of thymidylate synthase biosynthetic process(GO:0050758) negative regulation of thymidylate synthase biosynthetic process(GO:0050760) |
| 0.1 | 0.4 | GO:0045726 | positive regulation of integrin biosynthetic process(GO:0045726) |
| 0.1 | 1.9 | GO:0035372 | protein localization to microtubule(GO:0035372) |
| 0.1 | 1.6 | GO:0019511 | peptidyl-proline hydroxylation(GO:0019511) |
| 0.1 | 2.4 | GO:0007343 | egg activation(GO:0007343) |
| 0.1 | 1.2 | GO:0032876 | negative regulation of DNA endoreduplication(GO:0032876) |
| 0.1 | 2.4 | GO:0030049 | muscle filament sliding(GO:0030049) |
| 0.1 | 2.6 | GO:0007190 | activation of adenylate cyclase activity(GO:0007190) |
| 0.1 | 6.7 | GO:0010596 | negative regulation of endothelial cell migration(GO:0010596) |
| 0.1 | 0.7 | GO:0045196 | establishment or maintenance of neuroblast polarity(GO:0045196) establishment of neuroblast polarity(GO:0045200) |
| 0.1 | 0.8 | GO:0031179 | peptide modification(GO:0031179) |
| 0.1 | 1.0 | GO:0009048 | dosage compensation(GO:0007549) dosage compensation by inactivation of X chromosome(GO:0009048) |
| 0.1 | 0.3 | GO:0097156 | fasciculation of motor neuron axon(GO:0097156) |
| 0.1 | 1.4 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.1 | 0.4 | GO:0070171 | negative regulation of tooth mineralization(GO:0070171) |
| 0.1 | 2.2 | GO:0090190 | positive regulation of branching involved in ureteric bud morphogenesis(GO:0090190) |
| 0.1 | 2.5 | GO:0032967 | positive regulation of collagen biosynthetic process(GO:0032967) |
| 0.1 | 2.9 | GO:0014912 | negative regulation of smooth muscle cell migration(GO:0014912) |
| 0.1 | 1.8 | GO:0045898 | regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045898) |
| 0.1 | 0.6 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.1 | 0.9 | GO:0021527 | spinal cord association neuron differentiation(GO:0021527) |
| 0.1 | 2.3 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.0 | 5.9 | GO:0043488 | regulation of mRNA stability(GO:0043488) |
| 0.0 | 1.7 | GO:1902230 | negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902230) |
| 0.0 | 1.2 | GO:0045736 | negative regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045736) negative regulation of cyclin-dependent protein kinase activity(GO:1904030) |
| 0.0 | 1.0 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.0 | 2.1 | GO:0007131 | reciprocal meiotic recombination(GO:0007131) reciprocal DNA recombination(GO:0035825) |
| 0.0 | 1.6 | GO:0045725 | positive regulation of glycogen biosynthetic process(GO:0045725) |
| 0.0 | 0.3 | GO:0030035 | microspike assembly(GO:0030035) |
| 0.0 | 0.3 | GO:0003344 | pericardium morphogenesis(GO:0003344) |
| 0.0 | 2.0 | GO:0046329 | negative regulation of JNK cascade(GO:0046329) |
| 0.0 | 0.3 | GO:0090336 | positive regulation of brown fat cell differentiation(GO:0090336) |
| 0.0 | 1.8 | GO:0045197 | establishment or maintenance of epithelial cell apical/basal polarity(GO:0045197) |
| 0.0 | 0.1 | GO:0090310 | negative regulation of methylation-dependent chromatin silencing(GO:0090310) |
| 0.0 | 0.8 | GO:1904376 | negative regulation of protein localization to plasma membrane(GO:1903077) negative regulation of protein localization to cell periphery(GO:1904376) |
| 0.0 | 1.7 | GO:0006493 | protein O-linked glycosylation(GO:0006493) |
| 0.0 | 0.2 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 0.0 | 3.4 | GO:0030838 | positive regulation of actin filament polymerization(GO:0030838) |
| 0.0 | 1.0 | GO:0010614 | negative regulation of cardiac muscle hypertrophy(GO:0010614) |
| 0.0 | 0.2 | GO:0015074 | DNA integration(GO:0015074) |
| 0.0 | 1.8 | GO:0042147 | retrograde transport, endosome to Golgi(GO:0042147) |
| 0.0 | 0.5 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.0 | 1.9 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.0 | 0.3 | GO:0043517 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) |
| 0.0 | 1.2 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.0 | 0.6 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.0 | 0.1 | GO:0061343 | cell adhesion involved in heart morphogenesis(GO:0061343) |
| 0.0 | 0.4 | GO:0060416 | response to growth hormone(GO:0060416) |
| 0.0 | 0.1 | GO:0021516 | dorsal spinal cord development(GO:0021516) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 5.2 | GO:0099522 | region of cytosol(GO:0099522) postsynaptic cytosol(GO:0099524) |
| 0.8 | 2.5 | GO:0038045 | large latent transforming growth factor-beta complex(GO:0038045) |
| 0.8 | 2.4 | GO:1904511 | cortical microtubule plus-end(GO:1903754) cytoplasmic microtubule plus-end(GO:1904511) |
| 0.7 | 5.6 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.6 | 3.2 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 0.5 | 3.8 | GO:0005638 | lamin filament(GO:0005638) |
| 0.3 | 2.0 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 0.3 | 2.1 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.3 | 1.6 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.3 | 1.6 | GO:0097226 | sperm mitochondrial sheath(GO:0097226) |
| 0.2 | 0.7 | GO:0005712 | chiasma(GO:0005712) late recombination nodule(GO:0005715) |
| 0.2 | 1.8 | GO:0097422 | tubular endosome(GO:0097422) |
| 0.2 | 1.7 | GO:0098837 | postsynaptic recycling endosome(GO:0098837) |
| 0.2 | 2.3 | GO:0000808 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.2 | 2.0 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.1 | 1.9 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 1.9 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.1 | 3.4 | GO:0032982 | myosin filament(GO:0032982) |
| 0.1 | 2.6 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.1 | 1.2 | GO:0001739 | sex chromatin(GO:0001739) |
| 0.1 | 0.5 | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex(GO:0000308) |
| 0.1 | 1.4 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.1 | 0.8 | GO:1990812 | growth cone lamellipodium(GO:1990761) growth cone filopodium(GO:1990812) |
| 0.1 | 1.1 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.1 | 5.0 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.1 | 3.8 | GO:0005657 | replication fork(GO:0005657) |
| 0.1 | 0.4 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.0 | 34.4 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 1.9 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.0 | 6.0 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 1.0 | GO:0042581 | specific granule(GO:0042581) |
| 0.0 | 0.5 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 0.4 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.0 | 0.6 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.0 | 2.9 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 1.3 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 1.3 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 0.8 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 6.9 | GO:0005774 | vacuolar membrane(GO:0005774) |
| 0.0 | 6.9 | GO:0045177 | apical part of cell(GO:0045177) |
| 0.0 | 0.8 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.0 | 1.4 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 1.1 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.7 | 21.5 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 2.1 | 6.2 | GO:0005302 | L-tyrosine transmembrane transporter activity(GO:0005302) |
| 1.5 | 9.0 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 1.4 | 5.6 | GO:0031721 | hemoglobin alpha binding(GO:0031721) |
| 1.3 | 3.8 | GO:0033680 | ATP-dependent DNA/RNA helicase activity(GO:0033680) |
| 0.7 | 2.1 | GO:0017116 | single-stranded DNA-dependent ATP-dependent DNA helicase activity(GO:0017116) |
| 0.6 | 2.5 | GO:0050436 | microfibril binding(GO:0050436) |
| 0.6 | 1.8 | GO:0035663 | Toll-like receptor 2 binding(GO:0035663) |
| 0.5 | 2.1 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 0.5 | 1.6 | GO:0019797 | procollagen-proline 3-dioxygenase activity(GO:0019797) |
| 0.5 | 12.6 | GO:0030553 | cGMP binding(GO:0030553) |
| 0.4 | 1.2 | GO:0032394 | MHC class Ib receptor activity(GO:0032394) |
| 0.4 | 2.0 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.4 | 1.2 | GO:0003858 | 3-hydroxybutyrate dehydrogenase activity(GO:0003858) |
| 0.3 | 2.4 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 0.3 | 1.0 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.2 | 1.7 | GO:0019958 | C-X-C chemokine binding(GO:0019958) |
| 0.2 | 4.2 | GO:0005522 | profilin binding(GO:0005522) |
| 0.2 | 1.2 | GO:0008390 | testosterone 16-alpha-hydroxylase activity(GO:0008390) |
| 0.2 | 6.5 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.2 | 1.6 | GO:0002135 | CTP binding(GO:0002135) |
| 0.2 | 0.8 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.2 | 5.9 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.2 | 2.6 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.2 | 2.0 | GO:0033170 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.2 | 0.7 | GO:0032137 | guanine/thymine mispair binding(GO:0032137) |
| 0.2 | 3.2 | GO:0035615 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.2 | 1.2 | GO:0034452 | dynactin binding(GO:0034452) |
| 0.2 | 3.1 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.2 | 2.1 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.1 | 2.3 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.1 | 0.8 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.1 | 0.7 | GO:0035241 | protein-arginine omega-N monomethyltransferase activity(GO:0035241) histone methyltransferase activity (H4-R3 specific)(GO:0044020) |
| 0.1 | 5.2 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.1 | 2.4 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.1 | 1.4 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.1 | 1.9 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.1 | 4.2 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.1 | 1.9 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.1 | 1.0 | GO:0003997 | acyl-CoA oxidase activity(GO:0003997) |
| 0.1 | 1.5 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.1 | 2.4 | GO:0000146 | microfilament motor activity(GO:0000146) |
| 0.1 | 0.5 | GO:0010385 | double-stranded methylated DNA binding(GO:0010385) hemi-methylated DNA-binding(GO:0044729) |
| 0.1 | 3.1 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.1 | 0.3 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.1 | 4.3 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.1 | 1.3 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.1 | 0.4 | GO:0030171 | voltage-gated proton channel activity(GO:0030171) |
| 0.1 | 3.1 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 1.2 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.1 | 0.6 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.1 | 8.4 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 0.3 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.0 | 0.6 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.0 | 3.0 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.0 | 1.1 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.0 | 1.6 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.0 | 1.5 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 0.8 | GO:0048156 | tau protein binding(GO:0048156) |
| 0.0 | 4.7 | GO:0005179 | hormone activity(GO:0005179) |
| 0.0 | 1.6 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.0 | 1.0 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.0 | 0.3 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) |
| 0.0 | 1.2 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 1.6 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.0 | 1.0 | GO:0001191 | transcriptional repressor activity, RNA polymerase II transcription factor binding(GO:0001191) |
| 0.0 | 0.2 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.0 | 4.0 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 0.7 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.0 | 0.2 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 1.8 | GO:0051219 | phosphoprotein binding(GO:0051219) |
| 0.0 | 0.5 | GO:0030332 | cyclin binding(GO:0030332) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 23.3 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.4 | 12.6 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.1 | 10.2 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 2.1 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.1 | 2.6 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.1 | 4.5 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.1 | 4.2 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.1 | 1.6 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.0 | 4.3 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 0.8 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 2.2 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 1.6 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 4.3 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 3.8 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 2.0 | PID FAS PATHWAY | FAS (CD95) signaling pathway |
| 0.0 | 9.3 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.9 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 2.4 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 2.4 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 1.2 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 1.8 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 3.3 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 6.5 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.2 | 2.3 | REACTOME E2F ENABLED INHIBITION OF PRE REPLICATION COMPLEX FORMATION | Genes involved in E2F-enabled inhibition of pre-replication complex formation |
| 0.2 | 3.2 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.2 | 4.7 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.2 | 2.6 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.1 | 4.3 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.1 | 2.2 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.1 | 3.1 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.1 | 3.4 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 6.2 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.1 | 5.0 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.1 | 1.4 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.1 | 1.6 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.1 | 7.1 | REACTOME RESPIRATORY ELECTRON TRANSPORT ATP SYNTHESIS BY CHEMIOSMOTIC COUPLING AND HEAT PRODUCTION BY UNCOUPLING PROTEINS | Genes involved in Respiratory electron transport, ATP synthesis by chemiosmotic coupling, and heat production by uncoupling proteins. |
| 0.1 | 0.9 | REACTOME SIGNALING BY PDGF | Genes involved in Signaling by PDGF |
| 0.1 | 2.0 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 4.0 | REACTOME MUSCLE CONTRACTION | Genes involved in Muscle contraction |
| 0.0 | 1.3 | REACTOME FGFR LIGAND BINDING AND ACTIVATION | Genes involved in FGFR ligand binding and activation |
| 0.0 | 1.9 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.6 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 1.2 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 0.7 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.0 | 1.2 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.0 | 1.8 | REACTOME MYD88 MAL CASCADE INITIATED ON PLASMA MEMBRANE | Genes involved in MyD88:Mal cascade initiated on plasma membrane |
| 0.0 | 0.6 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.6 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |