Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
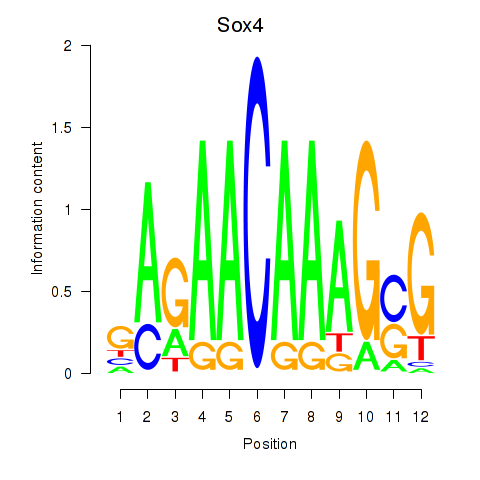
Results for Sox4
Z-value: 2.03
Transcription factors associated with Sox4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Sox4
|
ENSMUSG00000076431.5 | Sox4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Sox4 | mm39_v1_chr13_-_29137673_29137696 | 0.53 | 9.3e-04 | Click! |
Activity profile of Sox4 motif
Sorted Z-values of Sox4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Sox4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_+_114914880 | 13.53 |
ENSMUST00000161601.8
|
Tal1
|
T cell acute lymphocytic leukemia 1 |
| chrX_+_72683020 | 13.31 |
ENSMUST00000019701.9
|
Dusp9
|
dual specificity phosphatase 9 |
| chr7_-_103502404 | 12.51 |
ENSMUST00000033229.5
|
Hbb-y
|
hemoglobin Y, beta-like embryonic chain |
| chr17_+_12338161 | 8.18 |
ENSMUST00000024594.9
|
Agpat4
|
1-acylglycerol-3-phosphate O-acyltransferase 4 (lysophosphatidic acid acyltransferase, delta) |
| chr7_-_100164007 | 7.58 |
ENSMUST00000207405.2
|
Dnajb13
|
DnaJ heat shock protein family (Hsp40) member B13 |
| chr8_+_85696695 | 6.60 |
ENSMUST00000164807.2
|
Prdx2
|
peroxiredoxin 2 |
| chr13_+_117356808 | 6.54 |
ENSMUST00000022242.9
|
Emb
|
embigin |
| chr12_+_24758724 | 5.96 |
ENSMUST00000153058.8
|
Rrm2
|
ribonucleotide reductase M2 |
| chr8_+_85696453 | 5.87 |
ENSMUST00000125893.8
|
Prdx2
|
peroxiredoxin 2 |
| chr8_+_85696396 | 5.84 |
ENSMUST00000109733.8
|
Prdx2
|
peroxiredoxin 2 |
| chr9_-_103357564 | 5.59 |
ENSMUST00000124310.5
|
Bfsp2
|
beaded filament structural protein 2, phakinin |
| chr17_-_79662514 | 5.40 |
ENSMUST00000068958.9
|
Cdc42ep3
|
CDC42 effector protein (Rho GTPase binding) 3 |
| chr8_+_95720864 | 5.30 |
ENSMUST00000212141.2
|
Adgrg1
|
adhesion G protein-coupled receptor G1 |
| chr12_+_24758968 | 5.23 |
ENSMUST00000154588.2
|
Rrm2
|
ribonucleotide reductase M2 |
| chr16_+_16964801 | 5.18 |
ENSMUST00000232479.2
ENSMUST00000232344.2 ENSMUST00000069064.7 |
Ydjc
|
YdjC homolog (bacterial) |
| chr8_+_95721019 | 5.12 |
ENSMUST00000212976.2
ENSMUST00000212995.2 |
Adgrg1
|
adhesion G protein-coupled receptor G1 |
| chr17_+_29712008 | 5.03 |
ENSMUST00000234665.2
|
Pim1
|
proviral integration site 1 |
| chr8_+_95721378 | 4.82 |
ENSMUST00000212956.2
ENSMUST00000212531.2 |
Adgrg1
|
adhesion G protein-coupled receptor G1 |
| chr5_+_110987839 | 4.53 |
ENSMUST00000200172.2
ENSMUST00000066160.3 |
Chek2
|
checkpoint kinase 2 |
| chr7_-_143102792 | 4.33 |
ENSMUST00000072727.7
ENSMUST00000207948.2 ENSMUST00000208190.2 |
Nap1l4
|
nucleosome assembly protein 1-like 4 |
| chr15_+_76131020 | 4.30 |
ENSMUST00000229380.2
|
Grina
|
glutamate receptor, ionotropic, N-methyl D-aspartate-associated protein 1 (glutamate binding) |
| chr15_-_79169671 | 4.24 |
ENSMUST00000170955.2
ENSMUST00000165408.8 |
Baiap2l2
|
BAI1-associated protein 2-like 2 |
| chr9_-_110886306 | 4.05 |
ENSMUST00000195968.2
ENSMUST00000111888.3 |
Ccrl2
|
chemokine (C-C motif) receptor-like 2 |
| chr11_+_69737437 | 3.70 |
ENSMUST00000152566.8
ENSMUST00000108633.9 |
Plscr3
|
phospholipid scramblase 3 |
| chr11_+_69737491 | 3.63 |
ENSMUST00000019605.4
|
Plscr3
|
phospholipid scramblase 3 |
| chr11_+_69737200 | 3.38 |
ENSMUST00000108632.8
|
Plscr3
|
phospholipid scramblase 3 |
| chrX_-_144288071 | 3.37 |
ENSMUST00000112835.8
ENSMUST00000143610.3 |
Amot
|
angiomotin |
| chr19_-_7218512 | 3.37 |
ENSMUST00000025675.11
|
Naa40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit |
| chr6_-_47571901 | 3.25 |
ENSMUST00000081721.13
ENSMUST00000114618.8 ENSMUST00000114616.8 |
Ezh2
|
enhancer of zeste 2 polycomb repressive complex 2 subunit |
| chr12_+_24881582 | 3.07 |
ENSMUST00000221952.2
ENSMUST00000078902.8 ENSMUST00000110942.11 |
Mboat2
|
membrane bound O-acyltransferase domain containing 2 |
| chr8_-_4309257 | 3.05 |
ENSMUST00000053252.9
|
Ctxn1
|
cortexin 1 |
| chr11_-_106050927 | 3.03 |
ENSMUST00000045923.10
|
Limd2
|
LIM domain containing 2 |
| chr11_+_101333238 | 2.99 |
ENSMUST00000107249.8
|
Rpl27
|
ribosomal protein L27 |
| chr4_+_129407374 | 2.90 |
ENSMUST00000062356.7
|
Marcksl1
|
MARCKS-like 1 |
| chr11_+_121128042 | 2.88 |
ENSMUST00000103015.4
|
Narf
|
nuclear prelamin A recognition factor |
| chrX_+_70600481 | 2.85 |
ENSMUST00000123100.2
|
Hmgb3
|
high mobility group box 3 |
| chr6_+_71684846 | 2.75 |
ENSMUST00000212792.2
|
Reep1
|
receptor accessory protein 1 |
| chr19_-_46315543 | 2.70 |
ENSMUST00000223917.2
ENSMUST00000224447.2 ENSMUST00000041391.5 ENSMUST00000096029.12 |
Psd
|
pleckstrin and Sec7 domain containing |
| chr11_+_45946800 | 2.69 |
ENSMUST00000011400.8
|
Adam19
|
a disintegrin and metallopeptidase domain 19 (meltrin beta) |
| chr1_-_79838897 | 2.63 |
ENSMUST00000190724.2
|
Serpine2
|
serine (or cysteine) peptidase inhibitor, clade E, member 2 |
| chr11_+_101333115 | 2.57 |
ENSMUST00000077856.13
|
Rpl27
|
ribosomal protein L27 |
| chrX_+_162873183 | 2.57 |
ENSMUST00000015545.10
|
Cltrn
|
collectrin, amino acid transport regulator |
| chr13_+_21365308 | 2.56 |
ENSMUST00000221464.2
|
Trim27
|
tripartite motif-containing 27 |
| chr12_-_75678092 | 2.54 |
ENSMUST00000238938.2
|
Rplp2-ps1
|
ribosomal protein, large P2, pseudogene 1 |
| chr7_-_143102524 | 2.50 |
ENSMUST00000208093.2
ENSMUST00000209098.2 |
Nap1l4
|
nucleosome assembly protein 1-like 4 |
| chr15_+_76130947 | 2.42 |
ENSMUST00000229772.2
ENSMUST00000230347.2 ENSMUST00000023225.8 |
Grina
|
glutamate receptor, ionotropic, N-methyl D-aspartate-associated protein 1 (glutamate binding) |
| chr10_-_116899664 | 2.39 |
ENSMUST00000218719.2
ENSMUST00000219573.2 ENSMUST00000047672.9 |
Cct2
|
chaperonin containing Tcp1, subunit 2 (beta) |
| chrX_+_158623460 | 2.37 |
ENSMUST00000112451.8
ENSMUST00000112453.9 |
Sh3kbp1
|
SH3-domain kinase binding protein 1 |
| chr6_-_16898440 | 2.31 |
ENSMUST00000031533.11
|
Tfec
|
transcription factor EC |
| chr19_+_7394951 | 2.31 |
ENSMUST00000159348.3
|
2700081O15Rik
|
RIKEN cDNA 2700081O15 gene |
| chr1_-_55066629 | 2.27 |
ENSMUST00000027127.14
|
Sf3b1
|
splicing factor 3b, subunit 1 |
| chr1_-_153363354 | 2.24 |
ENSMUST00000186380.7
ENSMUST00000188345.2 ENSMUST00000042141.12 |
Dhx9
|
DEAH (Asp-Glu-Ala-His) box polypeptide 9 |
| chr3_-_95041246 | 2.21 |
ENSMUST00000172572.9
ENSMUST00000173462.3 |
Scnm1
|
sodium channel modifier 1 |
| chr16_-_90841360 | 2.21 |
ENSMUST00000118522.8
|
Paxbp1
|
PAX3 and PAX7 binding protein 1 |
| chr7_-_126831803 | 2.11 |
ENSMUST00000133913.8
|
Septin1
|
septin 1 |
| chr3_-_143910926 | 2.10 |
ENSMUST00000120539.8
ENSMUST00000196264.5 |
Lmo4
|
LIM domain only 4 |
| chr5_+_25452342 | 2.05 |
ENSMUST00000114950.2
|
Galnt11
|
polypeptide N-acetylgalactosaminyltransferase 11 |
| chr7_+_44866095 | 1.88 |
ENSMUST00000209437.2
|
Tead2
|
TEA domain family member 2 |
| chr12_-_110662723 | 1.86 |
ENSMUST00000021698.13
|
Hsp90aa1
|
heat shock protein 90, alpha (cytosolic), class A member 1 |
| chr3_+_89072096 | 1.83 |
ENSMUST00000121212.9
ENSMUST00000152205.5 ENSMUST00000090927.12 ENSMUST00000148265.8 ENSMUST00000121931.8 |
Clk2
|
CDC-like kinase 2 |
| chr12_-_110662677 | 1.77 |
ENSMUST00000124156.8
|
Hsp90aa1
|
heat shock protein 90, alpha (cytosolic), class A member 1 |
| chr7_+_121818692 | 1.76 |
ENSMUST00000033152.5
|
Chp2
|
calcineurin-like EF hand protein 2 |
| chr7_-_110681402 | 1.60 |
ENSMUST00000159305.2
|
Eif4g2
|
eukaryotic translation initiation factor 4, gamma 2 |
| chr12_-_111638722 | 1.58 |
ENSMUST00000001304.9
|
Ckb
|
creatine kinase, brain |
| chr17_+_75312520 | 1.58 |
ENSMUST00000234490.2
ENSMUST00000001927.12 |
Ltbp1
|
latent transforming growth factor beta binding protein 1 |
| chr1_-_36982747 | 1.53 |
ENSMUST00000185964.3
|
Tmem131
|
transmembrane protein 131 |
| chr6_+_71684872 | 1.49 |
ENSMUST00000212631.2
|
Reep1
|
receptor accessory protein 1 |
| chr16_-_18885809 | 1.49 |
ENSMUST00000200211.2
|
Iglj3
|
immunoglobulin lambda joining 3 |
| chr13_-_59930059 | 1.43 |
ENSMUST00000225581.2
|
Gm49354
|
predicted gene, 49354 |
| chr2_-_165740864 | 1.38 |
ENSMUST00000136842.2
|
Zmynd8
|
zinc finger, MYND-type containing 8 |
| chr1_-_131127825 | 1.34 |
ENSMUST00000068564.15
|
Rassf5
|
Ras association (RalGDS/AF-6) domain family member 5 |
| chr14_-_61283911 | 1.25 |
ENSMUST00000111234.10
ENSMUST00000224371.2 |
Tnfrsf19
|
tumor necrosis factor receptor superfamily, member 19 |
| chr5_+_122534900 | 1.25 |
ENSMUST00000196969.5
|
Arpc3
|
actin related protein 2/3 complex, subunit 3 |
| chr8_-_85696040 | 1.20 |
ENSMUST00000214133.2
ENSMUST00000147812.8 |
Gm49661
Rnaseh2a
|
predicted gene, 49661 ribonuclease H2, large subunit |
| chr18_+_82572595 | 1.16 |
ENSMUST00000152071.9
ENSMUST00000142850.9 ENSMUST00000080658.12 ENSMUST00000133193.9 ENSMUST00000123251.9 ENSMUST00000153478.9 ENSMUST00000075372.13 ENSMUST00000102812.12 ENSMUST00000062446.15 ENSMUST00000114674.11 ENSMUST00000132369.3 |
Mbp
|
myelin basic protein |
| chr12_-_81827893 | 1.15 |
ENSMUST00000035987.9
|
Map3k9
|
mitogen-activated protein kinase kinase kinase 9 |
| chr12_-_79343040 | 1.12 |
ENSMUST00000218377.2
ENSMUST00000021547.8 |
Zfyve26
|
zinc finger, FYVE domain containing 26 |
| chr1_+_31215482 | 1.02 |
ENSMUST00000062560.14
|
Lgsn
|
lengsin, lens protein with glutamine synthetase domain |
| chr11_-_115704447 | 0.93 |
ENSMUST00000041684.11
ENSMUST00000156812.2 |
Caskin2
|
CASK-interacting protein 2 |
| chr1_-_36596598 | 0.93 |
ENSMUST00000191642.6
ENSMUST00000193382.6 ENSMUST00000195620.6 |
Sema4c
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4C |
| chr3_+_146205562 | 0.93 |
ENSMUST00000090031.12
ENSMUST00000118280.2 |
Gng5
|
guanine nucleotide binding protein (G protein), gamma 5 |
| chr15_-_44291699 | 0.88 |
ENSMUST00000038719.8
|
Nudcd1
|
NudC domain containing 1 |
| chr2_-_26854283 | 0.84 |
ENSMUST00000136710.2
ENSMUST00000064244.11 ENSMUST00000114020.10 |
Rexo4
|
REX4, 3'-5' exonuclease |
| chr5_-_136227760 | 0.81 |
ENSMUST00000149151.2
ENSMUST00000151786.8 |
Prkrip1
|
Prkr interacting protein 1 (IL11 inducible) |
| chr9_+_59614877 | 0.78 |
ENSMUST00000128944.8
ENSMUST00000098661.10 |
Gramd2
|
GRAM domain containing 2 |
| chr2_+_32460868 | 0.75 |
ENSMUST00000140592.8
ENSMUST00000028151.7 |
Dpm2
|
dolichol-phosphate (beta-D) mannosyltransferase 2 |
| chrX_+_132751729 | 0.69 |
ENSMUST00000033602.9
|
Tnmd
|
tenomodulin |
| chr7_+_126832399 | 0.68 |
ENSMUST00000056232.7
|
Zfp553
|
zinc finger protein 553 |
| chr3_-_127631068 | 0.68 |
ENSMUST00000051737.8
ENSMUST00000200409.5 |
Ap1ar
|
adaptor-related protein complex 1 associated regulatory protein |
| chr14_-_52495160 | 0.67 |
ENSMUST00000200169.6
|
Chd8
|
chromodomain helicase DNA binding protein 8 |
| chr8_-_85696369 | 0.66 |
ENSMUST00000109736.9
ENSMUST00000140561.8 |
Rnaseh2a
|
ribonuclease H2, large subunit |
| chr3_+_146205864 | 0.65 |
ENSMUST00000119130.2
|
Gng5
|
guanine nucleotide binding protein (G protein), gamma 5 |
| chr16_-_19334384 | 0.65 |
ENSMUST00000054606.2
|
Olfr167
|
olfactory receptor 167 |
| chr15_+_44291470 | 0.62 |
ENSMUST00000226827.2
ENSMUST00000060652.5 |
Eny2
|
ENY2 transcription and export complex 2 subunit |
| chr4_+_83335947 | 0.62 |
ENSMUST00000030206.10
ENSMUST00000071544.11 |
Snapc3
|
small nuclear RNA activating complex, polypeptide 3 |
| chr3_-_143908111 | 0.60 |
ENSMUST00000121796.8
|
Lmo4
|
LIM domain only 4 |
| chr9_+_13677266 | 0.59 |
ENSMUST00000152532.8
|
Mtmr2
|
myotubularin related protein 2 |
| chr3_+_8574420 | 0.58 |
ENSMUST00000029002.9
|
Stmn2
|
stathmin-like 2 |
| chr19_+_34899778 | 0.57 |
ENSMUST00000223907.2
|
Kif20b
|
kinesin family member 20B |
| chr11_+_54205722 | 0.52 |
ENSMUST00000072178.11
ENSMUST00000101211.9 ENSMUST00000101213.9 ENSMUST00000064690.10 ENSMUST00000108899.8 |
Acsl6
|
acyl-CoA synthetase long-chain family member 6 |
| chr11_+_22462088 | 0.50 |
ENSMUST00000059319.8
|
Tmem17
|
transmembrane protein 17 |
| chr11_+_94858417 | 0.49 |
ENSMUST00000038928.7
|
H1f9
|
H1.9 linker histone |
| chr3_-_143908060 | 0.48 |
ENSMUST00000121112.6
|
Lmo4
|
LIM domain only 4 |
| chr2_+_78699360 | 0.44 |
ENSMUST00000028398.14
|
Ube2e3
|
ubiquitin-conjugating enzyme E2E 3 |
| chr7_-_30614249 | 0.44 |
ENSMUST00000190950.7
ENSMUST00000187137.7 ENSMUST00000190638.7 |
Mag
|
myelin-associated glycoprotein |
| chr19_+_34899752 | 0.44 |
ENSMUST00000087341.7
|
Kif20b
|
kinesin family member 20B |
| chr3_-_50398027 | 0.43 |
ENSMUST00000029297.6
ENSMUST00000194462.6 |
Slc7a11
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 11 |
| chr16_+_84631789 | 0.42 |
ENSMUST00000114184.8
|
Gabpa
|
GA repeat binding protein, alpha |
| chr4_-_141327146 | 0.40 |
ENSMUST00000141518.8
ENSMUST00000127455.8 ENSMUST00000105784.8 |
Fblim1
|
filamin binding LIM protein 1 |
| chr7_+_126832215 | 0.39 |
ENSMUST00000106312.4
|
Zfp553
|
zinc finger protein 553 |
| chr10_-_17823736 | 0.38 |
ENSMUST00000037879.8
|
Heca
|
hdc homolog, cell cycle regulator |
| chr17_-_36015484 | 0.38 |
ENSMUST00000117301.8
|
Ddr1
|
discoidin domain receptor family, member 1 |
| chr1_+_39940189 | 0.33 |
ENSMUST00000191761.6
ENSMUST00000193682.6 ENSMUST00000195860.6 ENSMUST00000195259.6 ENSMUST00000195636.6 ENSMUST00000192509.6 |
Map4k4
|
mitogen-activated protein kinase kinase kinase kinase 4 |
| chr15_+_30457772 | 0.32 |
ENSMUST00000228282.2
|
Ctnnd2
|
catenin (cadherin associated protein), delta 2 |
| chr19_+_5790918 | 0.31 |
ENSMUST00000081496.6
|
Ltbp3
|
latent transforming growth factor beta binding protein 3 |
| chr1_-_131441962 | 0.27 |
ENSMUST00000185445.3
|
Srgap2
|
SLIT-ROBO Rho GTPase activating protein 2 |
| chr11_+_67128843 | 0.26 |
ENSMUST00000018632.11
|
Myh4
|
myosin, heavy polypeptide 4, skeletal muscle |
| chr14_+_102077937 | 0.23 |
ENSMUST00000159026.8
|
Lmo7
|
LIM domain only 7 |
| chr12_-_72283465 | 0.19 |
ENSMUST00000021497.16
ENSMUST00000137990.2 |
Rtn1
|
reticulon 1 |
| chr12_-_72455708 | 0.19 |
ENSMUST00000078505.14
|
Rtn1
|
reticulon 1 |
| chr5_+_20112704 | 0.17 |
ENSMUST00000115267.7
|
Magi2
|
membrane associated guanylate kinase, WW and PDZ domain containing 2 |
| chrX_-_9983836 | 0.17 |
ENSMUST00000115543.3
ENSMUST00000044789.10 ENSMUST00000115544.9 |
Srpx
|
sushi-repeat-containing protein |
| chr6_+_48578935 | 0.17 |
ENSMUST00000204095.3
|
Zfp775
|
zinc finger protein 775 |
| chr18_+_37622518 | 0.16 |
ENSMUST00000055949.4
|
Pcdhb18
|
protocadherin beta 18 |
| chr11_-_49077864 | 0.15 |
ENSMUST00000153999.3
ENSMUST00000066531.13 |
Btnl9
|
butyrophilin-like 9 |
| chr11_-_49077986 | 0.15 |
ENSMUST00000046522.13
|
Btnl9
|
butyrophilin-like 9 |
| chr17_-_36014892 | 0.14 |
ENSMUST00000097333.10
ENSMUST00000003628.13 |
Ddr1
|
discoidin domain receptor family, member 1 |
| chr5_+_20112500 | 0.12 |
ENSMUST00000101558.10
|
Magi2
|
membrane associated guanylate kinase, WW and PDZ domain containing 2 |
| chr12_+_86999366 | 0.11 |
ENSMUST00000191463.2
|
Cipc
|
CLOCK interacting protein, circadian |
| chr3_-_96833336 | 0.10 |
ENSMUST00000062944.7
|
Gja8
|
gap junction protein, alpha 8 |
| chr5_-_24650164 | 0.09 |
ENSMUST00000115043.8
ENSMUST00000115041.2 ENSMUST00000030800.13 |
Fastk
|
Fas-activated serine/threonine kinase |
| chr16_-_31094095 | 0.09 |
ENSMUST00000060188.14
|
Ppp1r2
|
protein phosphatase 1, regulatory inhibitor subunit 2 |
| chr17_-_36008863 | 0.07 |
ENSMUST00000146472.8
|
Ddr1
|
discoidin domain receptor family, member 1 |
| chr4_+_136013372 | 0.00 |
ENSMUST00000069195.5
ENSMUST00000130658.2 |
Zfp46
|
zinc finger protein 46 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.5 | 13.5 | GO:0060217 | hemangioblast cell differentiation(GO:0060217) |
| 1.9 | 7.6 | GO:1904158 | axonemal central apparatus assembly(GO:1904158) |
| 1.5 | 18.3 | GO:0002536 | respiratory burst involved in inflammatory response(GO:0002536) |
| 1.4 | 12.5 | GO:0015671 | oxygen transport(GO:0015671) |
| 1.4 | 4.2 | GO:0051325 | interphase(GO:0051325) mitotic interphase(GO:0051329) |
| 1.1 | 3.4 | GO:0003365 | establishment of cell polarity involved in ameboidal cell migration(GO:0003365) |
| 1.1 | 3.2 | GO:0071707 | immunoglobulin heavy chain V-D-J recombination(GO:0071707) |
| 0.9 | 2.6 | GO:0044704 | mating plug formation(GO:0042628) single-organism reproductive behavior(GO:0044704) post-mating behavior(GO:0045297) seminal vesicle development(GO:0061107) |
| 0.6 | 15.2 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.6 | 13.3 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.5 | 6.5 | GO:0035879 | lactate transport(GO:0015727) lactate transmembrane transport(GO:0035873) plasma membrane lactate transport(GO:0035879) |
| 0.5 | 3.6 | GO:0045583 | regulation of cytotoxic T cell differentiation(GO:0045583) positive regulation of cytotoxic T cell differentiation(GO:0045585) |
| 0.5 | 11.2 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 0.5 | 2.9 | GO:0045578 | negative regulation of B cell differentiation(GO:0045578) |
| 0.4 | 2.2 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.4 | 2.6 | GO:1900041 | negative regulation of interleukin-2 secretion(GO:1900041) |
| 0.4 | 5.0 | GO:1902033 | vitamin D receptor signaling pathway(GO:0070561) regulation of hematopoietic stem cell proliferation(GO:1902033) |
| 0.3 | 1.0 | GO:0006542 | glutamine biosynthetic process(GO:0006542) |
| 0.3 | 5.4 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.3 | 1.9 | GO:0043137 | DNA replication, removal of RNA primer(GO:0043137) |
| 0.3 | 2.4 | GO:0090666 | scaRNA localization to Cajal body(GO:0090666) |
| 0.3 | 1.2 | GO:1904209 | chemokine (C-C motif) ligand 2 secretion(GO:0035926) regulation of chemokine (C-C motif) ligand 2 secretion(GO:1904207) positive regulation of chemokine (C-C motif) ligand 2 secretion(GO:1904209) |
| 0.3 | 5.1 | GO:0070307 | lens fiber cell development(GO:0070307) |
| 0.3 | 2.1 | GO:0018243 | protein O-linked glycosylation via threonine(GO:0018243) |
| 0.2 | 1.4 | GO:1903756 | regulation of transcription from RNA polymerase II promoter by histone modification(GO:1903756) negative regulation of transcription from RNA polymerase II promoter by histone modification(GO:1903758) |
| 0.2 | 6.7 | GO:1902236 | negative regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signaling pathway(GO:1902236) |
| 0.2 | 3.2 | GO:0021527 | spinal cord association neuron differentiation(GO:0021527) |
| 0.2 | 4.2 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.1 | 0.6 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 0.1 | 1.6 | GO:0072513 | positive regulation of secondary heart field cardioblast proliferation(GO:0072513) |
| 0.1 | 2.2 | GO:0014842 | regulation of skeletal muscle satellite cell proliferation(GO:0014842) |
| 0.1 | 4.2 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.1 | 3.4 | GO:0043968 | N-terminal protein amino acid acetylation(GO:0006474) histone H2A acetylation(GO:0043968) |
| 0.1 | 0.7 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) |
| 0.1 | 0.7 | GO:0019348 | dolichol metabolic process(GO:0019348) |
| 0.1 | 0.5 | GO:0010747 | positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) |
| 0.1 | 0.3 | GO:1902460 | transforming growth factor beta activation(GO:0036363) regulation of mesenchymal stem cell proliferation(GO:1902460) positive regulation of mesenchymal stem cell proliferation(GO:1902462) |
| 0.1 | 1.8 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.1 | 10.7 | GO:0055092 | cholesterol homeostasis(GO:0042632) sterol homeostasis(GO:0055092) |
| 0.1 | 1.6 | GO:0030644 | cellular chloride ion homeostasis(GO:0030644) |
| 0.1 | 0.5 | GO:0032954 | regulation of cytokinetic process(GO:0032954) regulation of mitotic cytokinetic process(GO:1903436) positive regulation of mitotic cytokinetic process(GO:1903438) positive regulation of mitotic cytokinesis(GO:1903490) |
| 0.1 | 6.8 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.1 | 2.7 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 1.8 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.1 | 1.9 | GO:0030903 | notochord development(GO:0030903) |
| 0.1 | 0.3 | GO:0031269 | pseudopodium assembly(GO:0031269) |
| 0.1 | 0.6 | GO:2000645 | regulation of receptor catabolic process(GO:2000644) negative regulation of receptor catabolic process(GO:2000645) |
| 0.1 | 0.7 | GO:2000270 | negative regulation of fibroblast apoptotic process(GO:2000270) |
| 0.1 | 0.3 | GO:0045653 | negative regulation of megakaryocyte differentiation(GO:0045653) |
| 0.1 | 0.9 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.1 | 1.6 | GO:0032967 | positive regulation of collagen biosynthetic process(GO:0032967) |
| 0.1 | 0.6 | GO:0009301 | snRNA transcription(GO:0009301) |
| 0.0 | 10.7 | GO:0008654 | phospholipid biosynthetic process(GO:0008654) |
| 0.0 | 2.7 | GO:0006509 | membrane protein ectodomain proteolysis(GO:0006509) |
| 0.0 | 0.4 | GO:0089711 | L-glutamate transmembrane transport(GO:0089711) |
| 0.0 | 0.4 | GO:0048711 | positive regulation of astrocyte differentiation(GO:0048711) |
| 0.0 | 2.3 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.0 | 2.3 | GO:0001825 | blastocyst formation(GO:0001825) |
| 0.0 | 0.2 | GO:0060244 | negative regulation of cell proliferation involved in contact inhibition(GO:0060244) |
| 0.0 | 1.2 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 0.3 | GO:0097118 | neuroligin clustering involved in postsynaptic membrane assembly(GO:0097118) |
| 0.0 | 0.9 | GO:0021535 | cell migration in hindbrain(GO:0021535) |
| 0.0 | 0.7 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.0 | 5.1 | GO:0006364 | rRNA processing(GO:0006364) |
| 0.0 | 0.3 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.0 | 1.6 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.0 | 1.2 | GO:0007257 | activation of JUN kinase activity(GO:0007257) |
| 0.0 | 0.4 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.5 | 13.5 | GO:0033193 | Lsd1/2 complex(GO:0033193) |
| 1.9 | 11.2 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 1.6 | 12.5 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 1.4 | 15.2 | GO:0097451 | glial limiting end-foot(GO:0097451) |
| 0.6 | 3.6 | GO:0097226 | sperm mitochondrial sheath(GO:0097226) |
| 0.5 | 1.6 | GO:0038045 | large latent transforming growth factor-beta complex(GO:0038045) |
| 0.5 | 1.9 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 0.4 | 2.9 | GO:0005638 | lamin filament(GO:0005638) |
| 0.4 | 1.6 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.4 | 2.2 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) |
| 0.3 | 4.2 | GO:0071439 | clathrin complex(GO:0071439) |
| 0.3 | 1.9 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.2 | 2.6 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.2 | 0.7 | GO:0033185 | dolichol-phosphate-mannose synthase complex(GO:0033185) |
| 0.1 | 1.2 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.1 | 2.4 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.1 | 4.2 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) |
| 0.1 | 1.6 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 3.2 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.1 | 2.3 | GO:0071004 | U2-type prespliceosome(GO:0071004) |
| 0.1 | 5.6 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.1 | 1.2 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.1 | 2.6 | GO:0030904 | retromer complex(GO:0030904) |
| 0.1 | 3.4 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.1 | 15.4 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 2.7 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 5.1 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 5.6 | GO:0036126 | sperm flagellum(GO:0036126) |
| 0.0 | 0.6 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.0 | 6.4 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 0.5 | GO:0036038 | MKS complex(GO:0036038) |
| 0.0 | 0.5 | GO:1990023 | mitotic spindle midzone(GO:1990023) |
| 0.0 | 3.3 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 0.3 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.0 | 0.7 | GO:0071339 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.0 | 0.3 | GO:0036057 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.0 | 0.3 | GO:0032982 | myosin filament(GO:0032982) |
| 0.0 | 2.4 | GO:0030139 | endocytic vesicle(GO:0030139) |
| 0.0 | 0.9 | GO:0030672 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.1 | 12.5 | GO:0031721 | hemoglobin alpha binding(GO:0031721) |
| 1.9 | 11.2 | GO:0004748 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 1.2 | 18.3 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.8 | 3.4 | GO:0043532 | angiostatin binding(GO:0043532) |
| 0.8 | 13.3 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.7 | 2.2 | GO:0033680 | ATP-dependent DNA/RNA helicase activity(GO:0033680) triplex DNA binding(GO:0045142) |
| 0.6 | 4.2 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.5 | 3.6 | GO:0002135 | CTP binding(GO:0002135) |
| 0.4 | 3.1 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.4 | 1.6 | GO:0050436 | microfibril binding(GO:0050436) |
| 0.4 | 5.0 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.4 | 3.4 | GO:1990189 | peptide-serine-N-acetyltransferase activity(GO:1990189) |
| 0.4 | 10.7 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.4 | 8.2 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.3 | 1.0 | GO:0016211 | glutamate-ammonia ligase activity(GO:0004356) ammonia ligase activity(GO:0016211) |
| 0.3 | 3.2 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.3 | 1.9 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.3 | 1.9 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.2 | 5.4 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.2 | 4.1 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.2 | 1.2 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.2 | 0.7 | GO:0004582 | dolichyl-phosphate beta-D-mannosyltransferase activity(GO:0004582) |
| 0.2 | 4.8 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.2 | 15.2 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.2 | 0.7 | GO:0035650 | AP-1 adaptor complex binding(GO:0035650) |
| 0.1 | 10.1 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 2.9 | GO:0005521 | lamin binding(GO:0005521) |
| 0.1 | 2.1 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.1 | 6.8 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.1 | 10.0 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.1 | 1.9 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 0.1 | 0.6 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.1 | 2.7 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.1 | 0.4 | GO:0000099 | sulfur amino acid transmembrane transporter activity(GO:0000099) |
| 0.0 | 1.4 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.3 | GO:0070699 | type II activin receptor binding(GO:0070699) |
| 0.0 | 0.4 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.0 | 0.7 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 0.5 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 5.2 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.3 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.0 | 1.8 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.0 | 1.6 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 2.7 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 3.0 | GO:0001158 | enhancer sequence-specific DNA binding(GO:0001158) |
| 0.0 | 1.1 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.0 | 0.5 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.0 | 4.3 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.0 | 2.0 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 13.3 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.2 | 5.0 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.1 | 3.6 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.1 | 11.2 | PID E2F PATHWAY | E2F transcription factor network |
| 0.1 | 4.2 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 3.2 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.0 | 2.4 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.0 | 1.2 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.0 | 1.3 | PID CD8 TCR PATHWAY | TCR signaling in naïve CD8+ T cells |
| 0.0 | 4.7 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.2 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 1.1 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 0.3 | PID EPHB FWD PATHWAY | EPHB forward signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 11.2 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.3 | 4.2 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.3 | 3.1 | REACTOME ACYL CHAIN REMODELLING OF PC | Genes involved in Acyl chain remodelling of PC |
| 0.2 | 8.2 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.2 | 3.6 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.1 | 3.4 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 2.4 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.1 | 2.4 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.0 | 1.9 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 5.6 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 2.3 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 1.6 | REACTOME G BETA GAMMA SIGNALLING THROUGH PLC BETA | Genes involved in G beta:gamma signalling through PLC beta |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.0 | 0.6 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.0 | 2.1 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.6 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 3 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 3 Promoter |
| 0.0 | 2.2 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 1.6 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |