Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
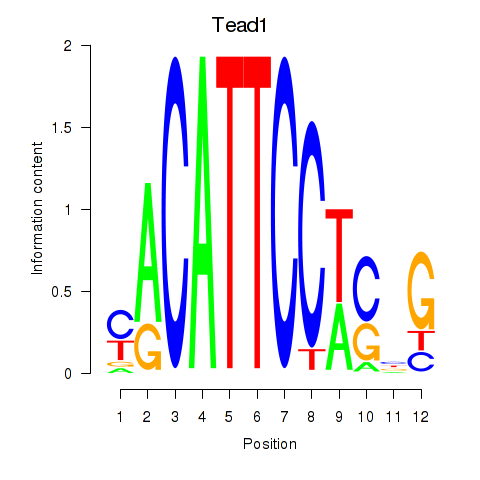
Results for Tead1
Z-value: 1.23
Transcription factors associated with Tead1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Tead1
|
ENSMUSG00000055320.19 | Tead1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Tead1 | mm39_v1_chr7_+_112278520_112278532 | 0.82 | 9.5e-10 | Click! |
Activity profile of Tead1 motif
Sorted Z-values of Tead1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Tead1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_43558386 | 7.14 |
ENSMUST00000130353.2
|
Tln1
|
talin 1 |
| chr7_-_33832640 | 5.11 |
ENSMUST00000038537.9
|
Wtip
|
WT1-interacting protein |
| chr12_+_85793313 | 4.13 |
ENSMUST00000040461.4
|
Flvcr2
|
feline leukemia virus subgroup C cellular receptor 2 |
| chr11_+_61967821 | 3.58 |
ENSMUST00000092415.9
ENSMUST00000201015.4 ENSMUST00000202744.4 ENSMUST00000201723.4 ENSMUST00000202179.2 |
Specc1
|
sperm antigen with calponin homology and coiled-coil domains 1 |
| chr8_+_57964921 | 3.28 |
ENSMUST00000067925.8
|
Hmgb2
|
high mobility group box 2 |
| chr7_-_143056252 | 3.23 |
ENSMUST00000010904.5
|
Phlda2
|
pleckstrin homology like domain, family A, member 2 |
| chr15_+_6416229 | 3.08 |
ENSMUST00000110664.9
ENSMUST00000110663.9 ENSMUST00000161812.8 ENSMUST00000160134.8 |
Dab2
|
disabled 2, mitogen-responsive phosphoprotein |
| chr18_+_50186349 | 2.86 |
ENSMUST00000148159.3
|
Tnfaip8
|
tumor necrosis factor, alpha-induced protein 8 |
| chr11_-_115903504 | 2.60 |
ENSMUST00000021114.5
|
Galk1
|
galactokinase 1 |
| chr11_-_100861713 | 2.51 |
ENSMUST00000060792.6
|
Cavin1
|
caveolae associated 1 |
| chr8_+_57964956 | 2.45 |
ENSMUST00000210871.2
|
Hmgb2
|
high mobility group box 2 |
| chr5_-_115438971 | 2.15 |
ENSMUST00000112090.2
|
Dynll1
|
dynein light chain LC8-type 1 |
| chr2_+_156617329 | 2.05 |
ENSMUST00000088552.7
|
Myl9
|
myosin, light polypeptide 9, regulatory |
| chr18_+_50164043 | 1.93 |
ENSMUST00000145726.2
ENSMUST00000128377.2 |
Tnfaip8
|
tumor necrosis factor, alpha-induced protein 8 |
| chr17_-_71575584 | 1.84 |
ENSMUST00000233148.2
|
Emilin2
|
elastin microfibril interfacer 2 |
| chr18_+_62086122 | 1.83 |
ENSMUST00000051720.6
ENSMUST00000235860.2 |
Sh3tc2
|
SH3 domain and tetratricopeptide repeats 2 |
| chr18_-_35795233 | 1.82 |
ENSMUST00000025209.12
ENSMUST00000096573.4 |
Spata24
|
spermatogenesis associated 24 |
| chr17_+_47904441 | 1.78 |
ENSMUST00000182874.3
|
Ccnd3
|
cyclin D3 |
| chr17_+_47904355 | 1.72 |
ENSMUST00000182209.8
|
Ccnd3
|
cyclin D3 |
| chr18_-_35795175 | 1.64 |
ENSMUST00000236574.2
ENSMUST00000236971.2 |
Spata24
|
spermatogenesis associated 24 |
| chr7_+_24603779 | 1.60 |
ENSMUST00000206906.2
ENSMUST00000206011.2 ENSMUST00000117419.8 ENSMUST00000205295.2 |
Arhgef1
|
Rho guanine nucleotide exchange factor (GEF) 1 |
| chr1_+_135764092 | 1.57 |
ENSMUST00000188028.7
ENSMUST00000178204.8 ENSMUST00000190451.7 ENSMUST00000189732.7 ENSMUST00000189355.7 |
Tnnt2
|
troponin T2, cardiac |
| chr2_-_60711706 | 1.43 |
ENSMUST00000164147.8
ENSMUST00000112509.2 |
Rbms1
|
RNA binding motif, single stranded interacting protein 1 |
| chr15_+_54274151 | 1.41 |
ENSMUST00000036737.4
|
Colec10
|
collectin sub-family member 10 |
| chr6_+_146789978 | 1.36 |
ENSMUST00000016631.14
ENSMUST00000203730.3 ENSMUST00000111623.9 |
Ppfibp1
|
PTPRF interacting protein, binding protein 1 (liprin beta 1) |
| chr1_+_51328265 | 1.33 |
ENSMUST00000051572.8
|
Cavin2
|
caveolae associated 2 |
| chr9_-_44624496 | 1.27 |
ENSMUST00000144251.8
ENSMUST00000156918.8 |
Phldb1
|
pleckstrin homology like domain, family B, member 1 |
| chr15_+_6416079 | 1.23 |
ENSMUST00000080880.12
|
Dab2
|
disabled 2, mitogen-responsive phosphoprotein |
| chr6_+_87405968 | 1.21 |
ENSMUST00000032125.7
|
Bmp10
|
bone morphogenetic protein 10 |
| chr6_-_42350188 | 1.14 |
ENSMUST00000073387.5
ENSMUST00000204357.2 |
Epha1
|
Eph receptor A1 |
| chr8_-_123278054 | 1.12 |
ENSMUST00000156333.9
ENSMUST00000067252.14 |
Piezo1
|
piezo-type mechanosensitive ion channel component 1 |
| chr14_-_54815000 | 1.10 |
ENSMUST00000054487.10
|
Ajuba
|
ajuba LIM protein |
| chr4_-_139695337 | 1.09 |
ENSMUST00000105031.4
|
Klhdc7a
|
kelch domain containing 7A |
| chr5_+_136996713 | 1.03 |
ENSMUST00000001790.6
|
Cldn15
|
claudin 15 |
| chr10_-_42152684 | 1.01 |
ENSMUST00000175881.8
ENSMUST00000056974.4 ENSMUST00000105502.8 ENSMUST00000105501.2 |
Foxo3
|
forkhead box O3 |
| chr7_+_3340013 | 0.96 |
ENSMUST00000204541.2
|
Myadm
|
myeloid-associated differentiation marker |
| chr17_-_71309815 | 0.95 |
ENSMUST00000123686.8
|
Myl12a
|
myosin, light chain 12A, regulatory, non-sarcomeric |
| chr14_-_70585874 | 0.95 |
ENSMUST00000152067.8
|
Slc39a14
|
solute carrier family 39 (zinc transporter), member 14 |
| chr5_+_117501557 | 0.94 |
ENSMUST00000111959.2
|
Wsb2
|
WD repeat and SOCS box-containing 2 |
| chr11_-_55310724 | 0.93 |
ENSMUST00000108858.8
ENSMUST00000141530.2 |
Sparc
|
secreted acidic cysteine rich glycoprotein |
| chr6_-_37419030 | 0.87 |
ENSMUST00000041093.6
|
Creb3l2
|
cAMP responsive element binding protein 3-like 2 |
| chr4_-_62278761 | 0.85 |
ENSMUST00000107461.2
ENSMUST00000084528.10 |
Fkbp15
|
FK506 binding protein 15 |
| chr17_+_69569184 | 0.85 |
ENSMUST00000224951.2
|
Epb41l3
|
erythrocyte membrane protein band 4.1 like 3 |
| chr6_+_17306334 | 0.81 |
ENSMUST00000007799.13
ENSMUST00000115456.6 |
Cav1
|
caveolin 1, caveolae protein |
| chr6_+_120341055 | 0.80 |
ENSMUST00000005108.10
|
Kdm5a
|
lysine (K)-specific demethylase 5A |
| chr4_+_15265798 | 0.77 |
ENSMUST00000062684.9
|
Tmem64
|
transmembrane protein 64 |
| chr7_-_109215754 | 0.72 |
ENSMUST00000084738.5
|
Denn2b
|
DENN domain containing 2B |
| chr3_+_31049896 | 0.72 |
ENSMUST00000108249.9
|
Prkci
|
protein kinase C, iota |
| chr11_-_5331734 | 0.66 |
ENSMUST00000172492.8
|
Znrf3
|
zinc and ring finger 3 |
| chr2_-_120234539 | 0.66 |
ENSMUST00000090042.12
ENSMUST00000110729.2 ENSMUST00000090046.12 |
Tmem87a
|
transmembrane protein 87A |
| chr9_-_8004586 | 0.65 |
ENSMUST00000086580.12
ENSMUST00000065353.13 |
Yap1
|
yes-associated protein 1 |
| chr19_-_57185928 | 0.61 |
ENSMUST00000111544.8
|
Ablim1
|
actin-binding LIM protein 1 |
| chr1_+_135693818 | 0.61 |
ENSMUST00000038945.6
|
Phlda3
|
pleckstrin homology like domain, family A, member 3 |
| chr4_-_129636073 | 0.61 |
ENSMUST00000066257.6
|
Khdrbs1
|
KH domain containing, RNA binding, signal transduction associated 1 |
| chr17_+_5991555 | 0.60 |
ENSMUST00000115791.10
ENSMUST00000080283.13 |
Synj2
|
synaptojanin 2 |
| chr16_-_38620688 | 0.59 |
ENSMUST00000057767.6
|
Upk1b
|
uroplakin 1B |
| chr7_-_28071919 | 0.59 |
ENSMUST00000119990.8
|
Plekhg2
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 2 |
| chr15_+_78726824 | 0.58 |
ENSMUST00000059619.3
|
Cdc42ep1
|
CDC42 effector protein (Rho GTPase binding) 1 |
| chr5_+_24631829 | 0.55 |
ENSMUST00000155598.8
|
Slc4a2
|
solute carrier family 4 (anion exchanger), member 2 |
| chr7_-_28071658 | 0.54 |
ENSMUST00000094644.11
|
Plekhg2
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 2 |
| chr14_+_5894220 | 0.53 |
ENSMUST00000063750.8
|
Rarb
|
retinoic acid receptor, beta |
| chr5_+_21850811 | 0.49 |
ENSMUST00000148873.8
ENSMUST00000072896.13 |
Armc10
|
armadillo repeat containing 10 |
| chr8_+_129450766 | 0.49 |
ENSMUST00000149116.2
|
Itgb1
|
integrin beta 1 (fibronectin receptor beta) |
| chr19_-_23425757 | 0.49 |
ENSMUST00000036069.8
|
Mamdc2
|
MAM domain containing 2 |
| chr8_+_122677231 | 0.48 |
ENSMUST00000093078.13
ENSMUST00000170857.8 ENSMUST00000026354.15 ENSMUST00000174753.8 ENSMUST00000172511.8 |
Banp
|
BTG3 associated nuclear protein |
| chr11_+_68961599 | 0.48 |
ENSMUST00000075980.12
ENSMUST00000094081.5 |
Tmem107
|
transmembrane protein 107 |
| chr18_+_60659257 | 0.47 |
ENSMUST00000223984.2
ENSMUST00000025505.7 ENSMUST00000223590.2 |
Dctn4
|
dynactin 4 |
| chr15_-_36283244 | 0.46 |
ENSMUST00000228358.2
ENSMUST00000022890.10 |
Rnf19a
|
ring finger protein 19A |
| chr1_-_155848917 | 0.46 |
ENSMUST00000138762.8
|
Cep350
|
centrosomal protein 350 |
| chr6_-_128101275 | 0.46 |
ENSMUST00000127105.8
|
Tspan9
|
tetraspanin 9 |
| chr4_-_62278673 | 0.42 |
ENSMUST00000084527.10
ENSMUST00000098033.10 |
Fkbp15
|
FK506 binding protein 15 |
| chr7_-_65020655 | 0.41 |
ENSMUST00000032729.8
|
Tjp1
|
tight junction protein 1 |
| chr5_+_103573367 | 0.41 |
ENSMUST00000048957.11
|
Ptpn13
|
protein tyrosine phosphatase, non-receptor type 13 |
| chr17_-_71309012 | 0.39 |
ENSMUST00000128179.2
ENSMUST00000150456.2 ENSMUST00000233357.2 ENSMUST00000233417.2 |
Myl12a
Gm49909
|
myosin, light chain 12A, regulatory, non-sarcomeric predicted gene, 49909 |
| chr12_-_98703664 | 0.39 |
ENSMUST00000170188.8
|
Ptpn21
|
protein tyrosine phosphatase, non-receptor type 21 |
| chr19_-_57185988 | 0.38 |
ENSMUST00000099294.9
|
Ablim1
|
actin-binding LIM protein 1 |
| chr11_-_85125889 | 0.36 |
ENSMUST00000018625.10
|
Appbp2
|
amyloid beta precursor protein (cytoplasmic tail) binding protein 2 |
| chr3_+_96011810 | 0.36 |
ENSMUST00000132980.8
ENSMUST00000138206.8 ENSMUST00000090785.9 ENSMUST00000035519.12 |
Otud7b
|
OTU domain containing 7B |
| chr1_+_189460461 | 0.36 |
ENSMUST00000097442.9
|
Ptpn14
|
protein tyrosine phosphatase, non-receptor type 14 |
| chr5_+_77099229 | 0.35 |
ENSMUST00000141687.2
|
Paics
|
phosphoribosylaminoimidazole carboxylase, phosphoribosylaminoribosylaminoimidazole, succinocarboxamide synthetase |
| chr19_-_57185808 | 0.35 |
ENSMUST00000111546.8
|
Ablim1
|
actin-binding LIM protein 1 |
| chr19_-_57185861 | 0.35 |
ENSMUST00000111550.8
|
Ablim1
|
actin-binding LIM protein 1 |
| chr11_+_115824108 | 0.34 |
ENSMUST00000140991.2
|
Sap30bp
|
SAP30 binding protein |
| chr18_+_44513547 | 0.34 |
ENSMUST00000202306.2
ENSMUST00000025350.10 |
Dcp2
|
decapping mRNA 2 |
| chr9_-_65298934 | 0.33 |
ENSMUST00000068307.4
|
Kbtbd13
|
kelch repeat and BTB (POZ) domain containing 13 |
| chr6_-_98319684 | 0.30 |
ENSMUST00000164491.3
|
Mdfic2
|
MyoD family inhibitor domain containing 2 |
| chr17_-_36012932 | 0.28 |
ENSMUST00000166980.9
ENSMUST00000145900.8 |
Ddr1
|
discoidin domain receptor family, member 1 |
| chr10_+_93476903 | 0.24 |
ENSMUST00000020204.5
|
Ntn4
|
netrin 4 |
| chr1_-_160134873 | 0.23 |
ENSMUST00000193185.6
|
Rabgap1l
|
RAB GTPase activating protein 1-like |
| chr2_-_25982160 | 0.18 |
ENSMUST00000114159.9
|
Nacc2
|
nucleus accumbens associated 2, BEN and BTB (POZ) domain containing |
| chr14_-_123151162 | 0.17 |
ENSMUST00000160401.8
|
Ggact
|
gamma-glutamylamine cyclotransferase |
| chr2_+_20524587 | 0.17 |
ENSMUST00000114604.9
ENSMUST00000066509.10 |
Etl4
|
enhancer trap locus 4 |
| chrX_-_103457431 | 0.17 |
ENSMUST00000033695.6
|
Abcb7
|
ATP-binding cassette, sub-family B (MDR/TAP), member 7 |
| chr6_+_34575435 | 0.17 |
ENSMUST00000079391.10
ENSMUST00000142512.8 ENSMUST00000115027.8 ENSMUST00000115026.8 |
Cald1
|
caldesmon 1 |
| chr16_+_26281885 | 0.13 |
ENSMUST00000161053.8
ENSMUST00000115302.2 |
Cldn16
|
claudin 16 |
| chr12_+_71184614 | 0.12 |
ENSMUST00000045907.16
|
2700049A03Rik
|
RIKEN cDNA 2700049A03 gene |
| chr11_+_62711057 | 0.12 |
ENSMUST00000055006.12
ENSMUST00000072639.10 |
Trim16
|
tripartite motif-containing 16 |
| chr3_+_122304778 | 0.11 |
ENSMUST00000198659.2
|
Bcar3
|
breast cancer anti-estrogen resistance 3 |
| chr17_-_48145466 | 0.11 |
ENSMUST00000066368.13
|
Mdfi
|
MyoD family inhibitor |
| chr2_+_91287846 | 0.07 |
ENSMUST00000028689.4
|
Lrp4
|
low density lipoprotein receptor-related protein 4 |
| chr18_-_9450097 | 0.06 |
ENSMUST00000053917.6
|
Ccny
|
cyclin Y |
| chr5_-_86616849 | 0.06 |
ENSMUST00000101073.3
|
Tmprss11a
|
transmembrane protease, serine 11a |
| chr5_-_135518098 | 0.06 |
ENSMUST00000201998.2
|
Hip1
|
huntingtin interacting protein 1 |
| chr6_-_119173348 | 0.04 |
ENSMUST00000187474.7
ENSMUST00000187940.7 |
Cacna1c
|
calcium channel, voltage-dependent, L type, alpha 1C subunit |
| chr7_+_140796096 | 0.02 |
ENSMUST00000153081.2
|
Rassf7
|
Ras association (RalGDS/AF-6) domain family (N-terminal) member 7 |
| chr11_+_115865535 | 0.02 |
ENSMUST00000021107.14
ENSMUST00000169928.8 ENSMUST00000106461.8 |
Itgb4
|
integrin beta 4 |
| chr6_-_128277701 | 0.00 |
ENSMUST00000143004.2
ENSMUST00000006311.13 ENSMUST00000112157.4 ENSMUST00000133118.2 |
Tead4
|
TEA domain family member 4 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 3.2 | GO:0060722 | spongiotrophoblast cell proliferation(GO:0060720) cell proliferation involved in embryonic placenta development(GO:0060722) |
| 0.9 | 2.6 | GO:0006059 | hexitol metabolic process(GO:0006059) glycolytic process from galactose(GO:0061623) |
| 0.7 | 4.3 | GO:0035026 | leading edge cell differentiation(GO:0035026) |
| 0.6 | 2.5 | GO:0006363 | termination of RNA polymerase I transcription(GO:0006363) |
| 0.6 | 5.7 | GO:0045654 | positive regulation of megakaryocyte differentiation(GO:0045654) |
| 0.6 | 8.0 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.6 | 5.6 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.4 | 1.6 | GO:0032972 | regulation of muscle filament sliding speed(GO:0032972) |
| 0.2 | 1.1 | GO:0033634 | positive regulation of cell-cell adhesion mediated by integrin(GO:0033634) |
| 0.2 | 0.7 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 0.2 | 0.6 | GO:0032241 | positive regulation of nucleobase-containing compound transport(GO:0032241) |
| 0.2 | 0.8 | GO:0031064 | negative regulation of histone deacetylation(GO:0031064) |
| 0.2 | 2.1 | GO:2000580 | regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000580) positive regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000582) |
| 0.2 | 0.8 | GO:0044339 | canonical Wnt signaling pathway involved in osteoblast differentiation(GO:0044339) |
| 0.2 | 0.5 | GO:0010710 | regulation of collagen catabolic process(GO:0010710) |
| 0.1 | 1.0 | GO:1901300 | positive regulation of hydrogen peroxide-mediated programmed cell death(GO:1901300) |
| 0.1 | 3.5 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.1 | 1.2 | GO:1903242 | regulation of cardiac muscle adaptation(GO:0010612) positive regulation of sarcomere organization(GO:0060298) regulation of cardiac muscle hypertrophy in response to stress(GO:1903242) |
| 0.1 | 1.3 | GO:1904261 | regulation of basement membrane assembly involved in embryonic body morphogenesis(GO:1904259) positive regulation of basement membrane assembly involved in embryonic body morphogenesis(GO:1904261) basement membrane assembly involved in embryonic body morphogenesis(GO:2001197) |
| 0.1 | 0.5 | GO:1904491 | protein localization to ciliary transition zone(GO:1904491) |
| 0.1 | 0.5 | GO:0040010 | positive regulation of growth rate(GO:0040010) |
| 0.1 | 0.7 | GO:0060242 | contact inhibition(GO:0060242) |
| 0.1 | 0.4 | GO:0071947 | protein deubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0071947) |
| 0.1 | 1.2 | GO:0032287 | peripheral nervous system myelin maintenance(GO:0032287) |
| 0.1 | 0.2 | GO:1900477 | negative regulation of G1/S transition of mitotic cell cycle by negative regulation of transcription from RNA polymerase II promoter(GO:1900477) |
| 0.1 | 4.8 | GO:0032611 | interleukin-1 beta production(GO:0032611) |
| 0.1 | 0.4 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.1 | 0.5 | GO:0003406 | retinal pigment epithelium development(GO:0003406) |
| 0.0 | 0.7 | GO:0035089 | establishment of apical/basal cell polarity(GO:0035089) |
| 0.0 | 0.5 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.0 | 0.6 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.0 | 1.3 | GO:0097320 | membrane tubulation(GO:0097320) |
| 0.0 | 0.2 | GO:0035426 | extracellular matrix-cell signaling(GO:0035426) |
| 0.0 | 0.6 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.0 | 0.3 | GO:0071044 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) histone mRNA catabolic process(GO:0071044) |
| 0.0 | 0.4 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.0 | 0.5 | GO:0034453 | microtubule anchoring(GO:0034453) |
| 0.0 | 0.1 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 0.0 | 2.0 | GO:0070527 | platelet aggregation(GO:0070527) |
| 0.0 | 1.7 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 1.1 | GO:0051496 | positive regulation of stress fiber assembly(GO:0051496) |
| 0.0 | 1.4 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.9 | GO:0006826 | iron ion transport(GO:0006826) |
| 0.0 | 0.4 | GO:0001946 | lymphangiogenesis(GO:0001946) |
| 0.0 | 0.5 | GO:0007097 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
| 0.0 | 0.1 | GO:0097105 | presynaptic membrane assembly(GO:0097105) |
| 0.0 | 0.1 | GO:2000588 | positive regulation of platelet-derived growth factor receptor-beta signaling pathway(GO:2000588) |
| 0.0 | 1.3 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.0 | 0.9 | GO:0010595 | positive regulation of endothelial cell migration(GO:0010595) |
| 0.0 | 0.9 | GO:0030968 | endoplasmic reticulum unfolded protein response(GO:0030968) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 4.3 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 0.2 | 0.9 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.2 | 1.6 | GO:1990584 | cardiac Troponin complex(GO:1990584) |
| 0.2 | 0.7 | GO:0071148 | TEAD-1-YAP complex(GO:0071148) TEAD-2-YAP complex(GO:0071149) |
| 0.2 | 0.5 | GO:0034665 | integrin alpha1-beta1 complex(GO:0034665) integrin alpha7-beta1 complex(GO:0034677) integrin alpha10-beta1 complex(GO:0034680) integrin alpha11-beta1 complex(GO:0034681) |
| 0.1 | 2.1 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.1 | 1.3 | GO:0045180 | basal cortex(GO:0045180) |
| 0.1 | 3.8 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 0.6 | GO:0031315 | extrinsic component of mitochondrial outer membrane(GO:0031315) |
| 0.0 | 6.5 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 0.5 | GO:0097197 | tetraspanin-enriched microdomain(GO:0097197) |
| 0.0 | 0.5 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.0 | 0.7 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
| 0.0 | 3.2 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 1.3 | GO:0099738 | cell cortex region(GO:0099738) |
| 0.0 | 0.9 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.0 | 7.2 | GO:0001726 | ruffle(GO:0001726) |
| 0.0 | 0.4 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 0.2 | GO:0030478 | actin cap(GO:0030478) |
| 0.0 | 5.7 | GO:0000793 | condensed chromosome(GO:0000793) |
| 0.0 | 1.4 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 2.2 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 1.0 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.0 | 0.5 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 0.0 | 0.1 | GO:0098890 | extrinsic component of postsynaptic membrane(GO:0098890) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 4.1 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.6 | 2.6 | GO:0004335 | galactokinase activity(GO:0004335) |
| 0.6 | 5.7 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.5 | 7.1 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.4 | 2.5 | GO:0042134 | rRNA primary transcript binding(GO:0042134) |
| 0.2 | 4.3 | GO:0098748 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.2 | 1.6 | GO:0030172 | troponin C binding(GO:0030172) troponin I binding(GO:0031013) |
| 0.2 | 0.8 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.2 | 1.1 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.1 | 2.1 | GO:0030235 | nitric-oxide synthase regulator activity(GO:0030235) |
| 0.1 | 1.2 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.1 | 1.1 | GO:0022833 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.1 | 4.8 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.1 | 0.5 | GO:0098639 | collagen binding involved in cell-matrix adhesion(GO:0098639) |
| 0.1 | 1.1 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.1 | 0.3 | GO:0004534 | 5'-3' exoribonuclease activity(GO:0004534) |
| 0.1 | 1.4 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.1 | 0.9 | GO:0015093 | ferrous iron transmembrane transporter activity(GO:0015093) |
| 0.1 | 3.5 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.1 | 1.4 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 0.6 | GO:0004439 | phosphatidylinositol-4,5-bisphosphate 5-phosphatase activity(GO:0004439) |
| 0.0 | 0.9 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 0.5 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.0 | 0.4 | GO:0071253 | connexin binding(GO:0071253) |
| 0.0 | 0.5 | GO:0003708 | retinoic acid receptor activity(GO:0003708) |
| 0.0 | 0.6 | GO:0010314 | phosphatidylinositol-5-phosphate binding(GO:0010314) |
| 0.0 | 0.2 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.0 | 0.2 | GO:0003839 | gamma-glutamylcyclotransferase activity(GO:0003839) |
| 0.0 | 0.4 | GO:1990380 | Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.0 | 1.3 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.0 | 0.6 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.0 | 0.6 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.0 | 1.3 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.0 | 0.6 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.0 | 0.7 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.0 | 0.4 | GO:0004697 | protein kinase C activity(GO:0004697) |
| 0.0 | 0.4 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.0 | 1.0 | GO:0001221 | transcription cofactor binding(GO:0001221) |
| 0.0 | 1.2 | GO:0035254 | glutamate receptor binding(GO:0035254) |
| 0.0 | 0.7 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 3.0 | GO:0003714 | transcription corepressor activity(GO:0003714) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 7.1 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.1 | 3.5 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.0 | 3.8 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 1.0 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.0 | 1.1 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.0 | 1.0 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.0 | 2.0 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 1.5 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.0 | 0.4 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.5 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.0 | 0.5 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 5.7 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.4 | 7.1 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.2 | 4.2 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.1 | 2.1 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.1 | 2.0 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 1.7 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.1 | 3.5 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.1 | 0.7 | REACTOME P75NTR RECRUITS SIGNALLING COMPLEXES | Genes involved in p75NTR recruits signalling complexes |
| 0.0 | 1.0 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 1.1 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.5 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.0 | 1.2 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.0 | 0.3 | REACTOME DESTABILIZATION OF MRNA BY BRF1 | Genes involved in Destabilization of mRNA by Butyrate Response Factor 1 (BRF1) |
| 0.0 | 0.6 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |