Project
avrg: GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
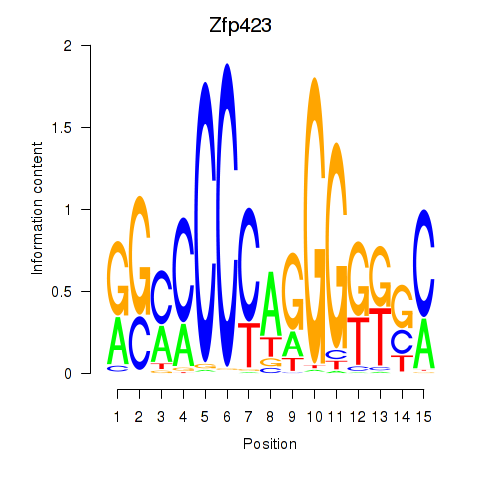
Results for Zfp423
Z-value: 1.00
Transcription factors associated with Zfp423
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Zfp423
|
ENSMUSG00000045333.16 | Zfp423 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Zfp423 | mm39_v1_chr8_-_88531018_88531067 | 0.45 | 5.5e-03 | Click! |
Activity profile of Zfp423 motif
Sorted Z-values of Zfp423 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Zfp423
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_+_86605146 | 6.21 |
ENSMUST00000043400.9
|
Asprv1
|
aspartic peptidase, retroviral-like 1 |
| chr12_+_109425769 | 5.29 |
ENSMUST00000173812.2
|
Dlk1
|
delta like non-canonical Notch ligand 1 |
| chr10_+_60185093 | 3.28 |
ENSMUST00000105459.2
|
Vsir
|
V-set immunoregulatory receptor |
| chr9_-_37464200 | 3.23 |
ENSMUST00000065668.12
|
Nrgn
|
neurogranin |
| chrX_-_74174450 | 3.22 |
ENSMUST00000114092.8
ENSMUST00000132501.8 ENSMUST00000153318.8 ENSMUST00000155742.2 |
Mpp1
|
membrane protein, palmitoylated |
| chrX_-_74174608 | 3.17 |
ENSMUST00000033775.9
|
Mpp1
|
membrane protein, palmitoylated |
| chr11_+_54793569 | 3.02 |
ENSMUST00000082430.11
|
Gpx3
|
glutathione peroxidase 3 |
| chr15_+_78128990 | 2.99 |
ENSMUST00000096357.12
|
Ncf4
|
neutrophil cytosolic factor 4 |
| chr19_+_10019023 | 2.94 |
ENSMUST00000237672.2
|
Fads3
|
fatty acid desaturase 3 |
| chr2_+_154633265 | 2.84 |
ENSMUST00000140713.3
ENSMUST00000137333.8 |
Raly
a
|
hnRNP-associated with lethal yellow nonagouti |
| chrX_-_74174524 | 2.83 |
ENSMUST00000114091.8
|
Mpp1
|
membrane protein, palmitoylated |
| chr11_+_54793743 | 2.47 |
ENSMUST00000149324.3
|
Gpx3
|
glutathione peroxidase 3 |
| chr4_-_106321363 | 2.23 |
ENSMUST00000049507.6
|
Pcsk9
|
proprotein convertase subtilisin/kexin type 9 |
| chr15_+_78129040 | 2.12 |
ENSMUST00000133618.3
|
Ncf4
|
neutrophil cytosolic factor 4 |
| chr17_-_23864237 | 1.97 |
ENSMUST00000024696.9
|
Mmp25
|
matrix metallopeptidase 25 |
| chr4_+_126915104 | 1.92 |
ENSMUST00000030623.8
|
Sfpq
|
splicing factor proline/glutamine rich (polypyrimidine tract binding protein associated) |
| chr19_+_7245591 | 1.86 |
ENSMUST00000066646.12
|
Rcor2
|
REST corepressor 2 |
| chr8_-_84425724 | 1.85 |
ENSMUST00000005616.16
|
Pkn1
|
protein kinase N1 |
| chr17_+_69463786 | 1.82 |
ENSMUST00000112680.8
ENSMUST00000080208.7 ENSMUST00000225977.2 |
Epb41l3
|
erythrocyte membrane protein band 4.1 like 3 |
| chr19_+_10018914 | 1.62 |
ENSMUST00000115995.4
|
Fads3
|
fatty acid desaturase 3 |
| chr16_+_49675969 | 1.59 |
ENSMUST00000229101.2
ENSMUST00000230836.2 ENSMUST00000229206.2 ENSMUST00000084838.14 ENSMUST00000230281.2 |
Cd47
|
CD47 antigen (Rh-related antigen, integrin-associated signal transducer) |
| chr5_-_22549688 | 1.57 |
ENSMUST00000062372.14
ENSMUST00000161356.8 |
Reln
|
reelin |
| chr7_-_141023199 | 1.52 |
ENSMUST00000106005.9
|
Pidd1
|
p53 induced death domain protein 1 |
| chr4_+_129407374 | 1.50 |
ENSMUST00000062356.7
|
Marcksl1
|
MARCKS-like 1 |
| chr16_+_17307503 | 1.43 |
ENSMUST00000023448.15
|
Aifm3
|
apoptosis-inducing factor, mitochondrion-associated 3 |
| chrX_+_100298134 | 1.28 |
ENSMUST00000062000.6
|
Foxo4
|
forkhead box O4 |
| chr19_+_6451667 | 1.28 |
ENSMUST00000113471.3
ENSMUST00000113469.3 |
Rasgrp2
|
RAS, guanyl releasing protein 2 |
| chr7_-_5017642 | 1.14 |
ENSMUST00000207412.2
ENSMUST00000077385.15 ENSMUST00000165320.3 |
Fiz1
|
Flt3 interacting zinc finger protein 1 |
| chr7_-_126625617 | 1.13 |
ENSMUST00000032916.6
|
Maz
|
MYC-associated zinc finger protein (purine-binding transcription factor) |
| chr4_-_116982804 | 1.07 |
ENSMUST00000183310.2
|
Btbd19
|
BTB (POZ) domain containing 19 |
| chr2_+_38229270 | 1.06 |
ENSMUST00000143783.9
|
Lhx2
|
LIM homeobox protein 2 |
| chr7_+_127084283 | 0.88 |
ENSMUST00000048896.8
|
Fbrs
|
fibrosin |
| chr19_-_10847121 | 0.87 |
ENSMUST00000120524.2
ENSMUST00000025645.14 |
Tmem132a
|
transmembrane protein 132A |
| chr15_-_103242697 | 0.86 |
ENSMUST00000229373.2
|
Zfp385a
|
zinc finger protein 385A |
| chr7_+_18737944 | 0.73 |
ENSMUST00000053713.5
|
Irf2bp1
|
interferon regulatory factor 2 binding protein 1 |
| chr2_+_164627743 | 0.62 |
ENSMUST00000174070.8
ENSMUST00000172577.8 ENSMUST00000056181.7 |
Snx21
|
sorting nexin family member 21 |
| chr9_+_120321557 | 0.60 |
ENSMUST00000007139.6
|
Eif1b
|
eukaryotic translation initiation factor 1B |
| chr3_+_114697710 | 0.60 |
ENSMUST00000081752.13
|
Olfm3
|
olfactomedin 3 |
| chr17_-_32385826 | 0.53 |
ENSMUST00000087723.5
|
Notch3
|
notch 3 |
| chr2_+_164627913 | 0.50 |
ENSMUST00000152471.2
|
Snx21
|
sorting nexin family member 21 |
| chr8_+_27750349 | 0.49 |
ENSMUST00000033880.7
|
Eif4ebp1
|
eukaryotic translation initiation factor 4E binding protein 1 |
| chr4_+_154321982 | 0.47 |
ENSMUST00000152159.8
|
Megf6
|
multiple EGF-like-domains 6 |
| chr17_-_37178079 | 0.47 |
ENSMUST00000025329.13
ENSMUST00000174195.8 |
Trim15
|
tripartite motif-containing 15 |
| chr17_+_69463846 | 0.47 |
ENSMUST00000238898.2
|
Epb41l3
|
erythrocyte membrane protein band 4.1 like 3 |
| chr10_-_14581203 | 0.47 |
ENSMUST00000149485.2
ENSMUST00000154132.8 |
Vta1
|
vesicle (multivesicular body) trafficking 1 |
| chr19_+_41818409 | 0.44 |
ENSMUST00000087155.5
|
Frat1
|
frequently rearranged in advanced T cell lymphomas |
| chr6_+_7844759 | 0.43 |
ENSMUST00000040159.6
|
C1galt1
|
core 1 synthase, glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase, 1 |
| chr7_-_19192672 | 0.43 |
ENSMUST00000239292.2
ENSMUST00000085715.7 |
Mark4
|
MAP/microtubule affinity regulating kinase 4 |
| chr17_-_23939700 | 0.39 |
ENSMUST00000201734.4
|
1520401A03Rik
|
RIKEN cDNA 1520401A03 gene |
| chr4_-_148711211 | 0.36 |
ENSMUST00000186947.7
ENSMUST00000105699.8 |
Tardbp
|
TAR DNA binding protein |
| chr2_+_106523532 | 0.30 |
ENSMUST00000111063.8
|
Mpped2
|
metallophosphoesterase domain containing 2 |
| chr19_-_5781503 | 0.29 |
ENSMUST00000162976.2
|
Znrd2
|
zinc ribbon domain containing 2 |
| chr15_+_101191077 | 0.28 |
ENSMUST00000071328.7
|
Smim41
|
small integral membrane protein 41 |
| chr15_+_101191099 | 0.25 |
ENSMUST00000191426.2
|
Smim41
|
small integral membrane protein 41 |
| chr7_-_79443536 | 0.24 |
ENSMUST00000032760.6
|
Mesp1
|
mesoderm posterior 1 |
| chr16_-_45474413 | 0.24 |
ENSMUST00000036732.9
|
BC016579
|
cDNA sequence, BC016579 |
| chr16_-_20972750 | 0.23 |
ENSMUST00000170665.3
|
Teddm3
|
transmembrane epididymal family member 3 |
| chr5_-_136594286 | 0.21 |
ENSMUST00000176172.8
|
Cux1
|
cut-like homeobox 1 |
| chr10_-_80223475 | 0.20 |
ENSMUST00000105350.3
|
Mex3d
|
mex3 RNA binding family member D |
| chr18_+_34892599 | 0.20 |
ENSMUST00000097622.4
|
Fam53c
|
family with sequence similarity 53, member C |
| chr2_+_93017916 | 0.18 |
ENSMUST00000090554.11
|
Trp53i11
|
transformation related protein 53 inducible protein 11 |
| chr2_+_93017887 | 0.14 |
ENSMUST00000150462.8
ENSMUST00000111266.8 |
Trp53i11
|
transformation related protein 53 inducible protein 11 |
| chr12_-_69230760 | 0.14 |
ENSMUST00000110620.2
ENSMUST00000110619.2 ENSMUST00000054544.7 |
Rpl36al
|
ribosomal protein L36A-like |
| chr1_+_156386414 | 0.14 |
ENSMUST00000166172.9
ENSMUST00000027888.13 |
Abl2
|
v-abl Abelson murine leukemia viral oncogene 2 (arg, Abelson-related gene) |
| chr17_-_25652750 | 0.14 |
ENSMUST00000159610.8
ENSMUST00000159048.8 ENSMUST00000078496.12 |
Cacna1h
|
calcium channel, voltage-dependent, T type, alpha 1H subunit |
| chr7_+_63803583 | 0.13 |
ENSMUST00000085222.12
ENSMUST00000206277.2 |
Trpm1
|
transient receptor potential cation channel, subfamily M, member 1 |
| chr11_+_81992662 | 0.10 |
ENSMUST00000000194.4
|
Ccl12
|
chemokine (C-C motif) ligand 12 |
| chr10_+_80972089 | 0.09 |
ENSMUST00000048128.15
|
Zbtb7a
|
zinc finger and BTB domain containing 7a |
| chr7_+_63803663 | 0.02 |
ENSMUST00000206314.2
|
Trpm1
|
transient receptor potential cation channel, subfamily M, member 1 |
| chr7_-_45519853 | 0.01 |
ENSMUST00000211713.2
|
Grin2d
|
glutamate receptor, ionotropic, NMDA2D (epsilon 4) |
| chr19_+_55730242 | 0.00 |
ENSMUST00000111662.11
ENSMUST00000041717.14 |
Tcf7l2
|
transcription factor 7 like 2, T cell specific, HMG box |
| chr10_+_80972368 | 0.00 |
ENSMUST00000119606.8
ENSMUST00000146895.2 ENSMUST00000121840.8 |
Zbtb7a
|
zinc finger and BTB domain containing 7a |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.8 | GO:0040030 | regulation of molecular function, epigenetic(GO:0040030) |
| 0.6 | 2.2 | GO:0032805 | positive regulation of low-density lipoprotein particle receptor catabolic process(GO:0032805) |
| 0.5 | 3.3 | GO:2000562 | negative regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000562) |
| 0.4 | 1.6 | GO:0097477 | lateral motor column neuron migration(GO:0097477) |
| 0.2 | 2.0 | GO:0060022 | hard palate development(GO:0060022) |
| 0.2 | 9.2 | GO:0090022 | regulation of neutrophil chemotaxis(GO:0090022) |
| 0.2 | 0.9 | GO:1902164 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902164) |
| 0.2 | 2.3 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.2 | 1.8 | GO:0035405 | histone-threonine phosphorylation(GO:0035405) |
| 0.2 | 5.1 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.2 | 5.5 | GO:0042744 | hydrogen peroxide catabolic process(GO:0042744) |
| 0.2 | 1.3 | GO:0070317 | negative regulation of G0 to G1 transition(GO:0070317) |
| 0.2 | 1.9 | GO:0045876 | positive regulation of sister chromatid cohesion(GO:0045876) |
| 0.1 | 5.3 | GO:0045780 | positive regulation of bone resorption(GO:0045780) positive regulation of bone remodeling(GO:0046852) |
| 0.1 | 1.6 | GO:0008228 | opsonization(GO:0008228) |
| 0.1 | 1.1 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.1 | 0.5 | GO:1990928 | response to amino acid starvation(GO:1990928) |
| 0.1 | 0.5 | GO:1901252 | regulation of intracellular transport of viral material(GO:1901252) |
| 0.1 | 1.5 | GO:0006977 | DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest(GO:0006977) |
| 0.1 | 0.2 | GO:0090081 | regulation of heart induction by regulation of canonical Wnt signaling pathway(GO:0090081) |
| 0.1 | 3.2 | GO:1900273 | positive regulation of long-term synaptic potentiation(GO:1900273) |
| 0.1 | 0.5 | GO:0072103 | glomerulus vasculature morphogenesis(GO:0072103) glomerular capillary formation(GO:0072104) |
| 0.1 | 0.5 | GO:0071985 | multivesicular body sorting pathway(GO:0071985) |
| 0.1 | 0.4 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.1 | 1.1 | GO:0045199 | maintenance of epithelial cell apical/basal polarity(GO:0045199) |
| 0.0 | 0.1 | GO:2000501 | natural killer cell chemotaxis(GO:0035747) negative regulation of lymphocyte migration(GO:2000402) regulation of natural killer cell chemotaxis(GO:2000501) |
| 0.0 | 1.4 | GO:0097194 | execution phase of apoptosis(GO:0097194) |
| 0.0 | 6.1 | GO:0016485 | protein processing(GO:0016485) |
| 0.0 | 1.3 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.0 | 0.7 | GO:0042462 | eye photoreceptor cell development(GO:0042462) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.2 | GO:1990666 | PCSK9-LDLR complex(GO:1990666) PCSK9-AnxA2 complex(GO:1990667) |
| 0.3 | 5.1 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.2 | 3.2 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.2 | 1.9 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.1 | 2.3 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.0 | 9.2 | GO:0030863 | cortical cytoskeleton(GO:0030863) |
| 0.0 | 1.8 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.1 | GO:0035841 | new growing cell tip(GO:0035841) |
| 0.0 | 2.8 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 1.1 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 0.4 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.0 | 7.8 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 1.9 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.2 | GO:0034189 | very-low-density lipoprotein particle binding(GO:0034189) |
| 0.5 | 2.8 | GO:0031781 | melanocortin receptor binding(GO:0031779) type 3 melanocortin receptor binding(GO:0031781) type 4 melanocortin receptor binding(GO:0031782) |
| 0.5 | 1.8 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 0.4 | 5.1 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.3 | 5.5 | GO:0008430 | selenium binding(GO:0008430) |
| 0.3 | 1.6 | GO:0070326 | very-low-density lipoprotein particle receptor binding(GO:0070326) |
| 0.2 | 6.2 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.1 | 1.6 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 0.1 | 3.2 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.0 | 1.4 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 0.4 | GO:0048531 | beta-1,3-galactosyltransferase activity(GO:0048531) |
| 0.0 | 2.0 | GO:0016504 | peptidase activator activity(GO:0016504) |
| 0.0 | 0.5 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.0 | 1.3 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 0.1 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.0 | 1.1 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.0 | 2.3 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.8 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 5.8 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 1.2 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 1.6 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 1.3 | PID RAS PATHWAY | Regulation of Ras family activation |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 0.5 | REACTOME SIGNALING BY NOTCH3 | Genes involved in Signaling by NOTCH3 |
| 0.1 | 5.3 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.1 | 5.1 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.1 | 2.0 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 1.6 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.0 | 1.3 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 1.3 | REACTOME PIP3 ACTIVATES AKT SIGNALING | Genes involved in PIP3 activates AKT signaling |
| 0.0 | 0.5 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 0.4 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.0 | 0.5 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |