Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
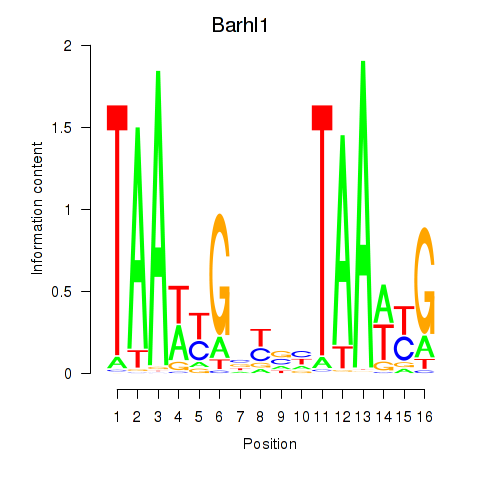
Results for Barhl1
Z-value: 0.62
Transcription factors associated with Barhl1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Barhl1
|
ENSMUSG00000026805.15 | BarH like homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Barhl1 | mm39_v1_chr2_-_28806639_28806680 | -0.36 | 3.0e-02 | Click! |
Activity profile of Barhl1 motif
Sorted Z-values of Barhl1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_90638580 | 9.76 |
ENSMUST00000042755.7
ENSMUST00000200693.2 |
Afp
|
alpha fetoprotein |
| chr16_-_16681839 | 4.26 |
ENSMUST00000100136.4
|
Igll1
|
immunoglobulin lambda-like polypeptide 1 |
| chr18_-_32692967 | 2.98 |
ENSMUST00000174000.2
ENSMUST00000174459.2 |
Gypc
|
glycophorin C |
| chr11_+_61967821 | 2.63 |
ENSMUST00000092415.9
ENSMUST00000201015.4 ENSMUST00000202744.4 ENSMUST00000201723.4 ENSMUST00000202179.2 |
Specc1
|
sperm antigen with calponin homology and coiled-coil domains 1 |
| chr15_-_100579450 | 2.08 |
ENSMUST00000230740.2
|
Cela1
|
chymotrypsin-like elastase family, member 1 |
| chr15_-_100579813 | 1.85 |
ENSMUST00000230572.2
|
Cela1
|
chymotrypsin-like elastase family, member 1 |
| chr11_+_116089678 | 1.63 |
ENSMUST00000021130.7
|
Ten1
|
TEN1 telomerase capping complex subunit |
| chr4_-_117013396 | 1.49 |
ENSMUST00000102696.5
|
Rps8
|
ribosomal protein S8 |
| chr5_+_149601688 | 1.38 |
ENSMUST00000100404.6
|
B3glct
|
beta-3-glucosyltransferase |
| chr8_-_85567256 | 1.32 |
ENSMUST00000003911.13
ENSMUST00000109761.9 ENSMUST00000128035.2 |
Rad23a
|
RAD23 homolog A, nucleotide excision repair protein |
| chr6_+_123100382 | 1.30 |
ENSMUST00000032248.8
|
Clec4a2
|
C-type lectin domain family 4, member a2 |
| chr17_+_7292967 | 1.22 |
ENSMUST00000097422.6
|
Gm1604b
|
predicted gene 1604b |
| chr3_-_75177378 | 1.14 |
ENSMUST00000039047.5
|
Serpini2
|
serine (or cysteine) peptidase inhibitor, clade I, member 2 |
| chrX_+_162694397 | 1.03 |
ENSMUST00000140845.2
|
Ap1s2
|
adaptor-related protein complex 1, sigma 2 subunit |
| chr3_-_88317601 | 0.84 |
ENSMUST00000193338.6
ENSMUST00000056370.13 |
Pmf1
|
polyamine-modulated factor 1 |
| chr7_+_97492124 | 0.82 |
ENSMUST00000033040.12
|
Pak1
|
p21 (RAC1) activated kinase 1 |
| chr18_-_42395207 | 0.74 |
ENSMUST00000097590.5
|
Lars
|
leucyl-tRNA synthetase |
| chr12_-_54842488 | 0.68 |
ENSMUST00000005798.9
|
Snx6
|
sorting nexin 6 |
| chr18_-_42395131 | 0.68 |
ENSMUST00000236102.2
|
Lars
|
leucyl-tRNA synthetase |
| chr10_+_58207229 | 0.67 |
ENSMUST00000238939.2
|
Lims1
|
LIM and senescent cell antigen-like domains 1 |
| chr6_-_129853617 | 0.65 |
ENSMUST00000014687.11
ENSMUST00000122219.2 |
Klra17
|
killer cell lectin-like receptor, subfamily A, member 17 |
| chr19_-_34143437 | 0.61 |
ENSMUST00000025686.9
|
Ankrd22
|
ankyrin repeat domain 22 |
| chr3_-_130503041 | 0.58 |
ENSMUST00000043937.9
|
Ostc
|
oligosaccharyltransferase complex subunit (non-catalytic) |
| chr15_-_58261093 | 0.57 |
ENSMUST00000227274.3
|
Anxa13
|
annexin A13 |
| chr5_-_23988696 | 0.52 |
ENSMUST00000119946.8
|
Pus7
|
pseudouridylate synthase 7 |
| chr11_+_78717398 | 0.52 |
ENSMUST00000147875.9
ENSMUST00000141321.2 |
Lyrm9
|
LYR motif containing 9 |
| chr2_-_104680104 | 0.51 |
ENSMUST00000028593.11
|
Prrg4
|
proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) |
| chr11_+_96209093 | 0.50 |
ENSMUST00000049241.9
|
Hoxb4
|
homeobox B4 |
| chr3_-_19319155 | 0.48 |
ENSMUST00000091314.11
|
Pde7a
|
phosphodiesterase 7A |
| chr10_+_128583734 | 0.48 |
ENSMUST00000163377.10
|
Pym1
|
PYM homolog 1, exon junction complex associated factor |
| chr11_+_101473062 | 0.45 |
ENSMUST00000039581.14
ENSMUST00000100403.9 ENSMUST00000107194.8 ENSMUST00000128614.2 |
Tmem106a
|
transmembrane protein 106A |
| chr5_-_23988551 | 0.44 |
ENSMUST00000148618.8
|
Pus7
|
pseudouridylate synthase 7 |
| chr17_-_25334879 | 0.44 |
ENSMUST00000024987.6
ENSMUST00000115181.9 |
Telo2
|
telomere maintenance 2 |
| chr6_+_113023419 | 0.44 |
ENSMUST00000138278.2
|
Thumpd3
|
THUMP domain containing 3 |
| chr10_-_126866682 | 0.43 |
ENSMUST00000040560.11
|
Tsfm
|
Ts translation elongation factor, mitochondrial |
| chr14_+_64097739 | 0.43 |
ENSMUST00000022528.6
|
Pinx1
|
PIN2/TERF1 interacting, telomerase inhibitor 1 |
| chr2_+_122256913 | 0.43 |
ENSMUST00000110525.8
|
Slc28a2
|
solute carrier family 28 (sodium-coupled nucleoside transporter), member 2 |
| chr2_-_73284262 | 0.42 |
ENSMUST00000102679.8
|
Wipf1
|
WAS/WASL interacting protein family, member 1 |
| chrX_+_41241049 | 0.42 |
ENSMUST00000128799.3
|
Stag2
|
stromal antigen 2 |
| chr6_+_88879233 | 0.41 |
ENSMUST00000055022.15
ENSMUST00000204765.3 ENSMUST00000153874.8 |
Tpra1
|
transmembrane protein, adipocyte asscociated 1 |
| chr3_-_146476331 | 0.37 |
ENSMUST00000106138.8
|
Prkacb
|
protein kinase, cAMP dependent, catalytic, beta |
| chr10_+_101994841 | 0.36 |
ENSMUST00000020039.13
|
Mgat4c
|
MGAT4 family, member C |
| chr6_-_41613322 | 0.33 |
ENSMUST00000031902.7
|
Trpv6
|
transient receptor potential cation channel, subfamily V, member 6 |
| chrX_+_108240356 | 0.33 |
ENSMUST00000139259.2
ENSMUST00000060013.4 |
Gm6377
|
predicted gene 6377 |
| chr2_-_86944911 | 0.28 |
ENSMUST00000216088.3
|
Olfr259
|
olfactory receptor 259 |
| chr14_-_26256025 | 0.28 |
ENSMUST00000139075.8
ENSMUST00000102956.8 |
Slmap
|
sarcolemma associated protein |
| chr15_+_25752963 | 0.28 |
ENSMUST00000022882.12
ENSMUST00000135173.8 |
Myo10
|
myosin X |
| chr6_+_42885812 | 0.28 |
ENSMUST00000216408.2
|
Olfr447
|
olfactory receptor 447 |
| chr11_+_54486925 | 0.27 |
ENSMUST00000218995.2
|
Rapgef6
|
Rap guanine nucleotide exchange factor (GEF) 6 |
| chr6_+_129568745 | 0.27 |
ENSMUST00000032268.14
ENSMUST00000112063.9 ENSMUST00000119520.8 |
Klrd1
|
killer cell lectin-like receptor, subfamily D, member 1 |
| chr3_-_146475974 | 0.26 |
ENSMUST00000106137.8
|
Prkacb
|
protein kinase, cAMP dependent, catalytic, beta |
| chr16_+_92089268 | 0.24 |
ENSMUST00000047383.10
|
Kcne2
|
potassium voltage-gated channel, Isk-related subfamily, gene 2 |
| chr1_-_75254989 | 0.23 |
ENSMUST00000039534.11
|
Resp18
|
regulated endocrine-specific protein 18 |
| chr11_-_79418500 | 0.22 |
ENSMUST00000154415.2
|
Evi2a
|
ecotropic viral integration site 2a |
| chr15_+_102240134 | 0.19 |
ENSMUST00000113682.9
ENSMUST00000001331.13 ENSMUST00000171244.2 |
Myg1
|
melanocyte proliferating gene 1 |
| chr2_-_111965322 | 0.19 |
ENSMUST00000213696.2
|
Olfr1316
|
olfactory receptor 1316 |
| chr2_-_85632888 | 0.18 |
ENSMUST00000217410.3
ENSMUST00000216425.3 |
Olfr1016
|
olfactory receptor 1016 |
| chr8_+_31579499 | 0.18 |
ENSMUST00000036631.14
|
Dusp26
|
dual specificity phosphatase 26 (putative) |
| chr2_-_45000389 | 0.18 |
ENSMUST00000201804.4
ENSMUST00000028229.13 ENSMUST00000202187.4 ENSMUST00000153561.6 ENSMUST00000201490.2 |
Zeb2
|
zinc finger E-box binding homeobox 2 |
| chr5_-_87630117 | 0.17 |
ENSMUST00000079811.13
ENSMUST00000144144.3 |
Ugt2a2
|
UDP glucuronosyltransferase 2 family, polypeptide A2 |
| chr9_+_19533374 | 0.17 |
ENSMUST00000213725.2
ENSMUST00000208694.2 |
Zfp317
|
zinc finger protein 317 |
| chr14_+_25980463 | 0.16 |
ENSMUST00000173155.8
|
Duxbl1
|
double homeobox B-like 1 |
| chr8_+_72635863 | 0.15 |
ENSMUST00000188685.7
|
Zfp709
|
zinc finger protein 709 |
| chr14_+_53599724 | 0.13 |
ENSMUST00000196105.2
|
Trav13n-4
|
T cell receptor alpha variable 13N-4 |
| chr17_+_37710293 | 0.12 |
ENSMUST00000216844.2
ENSMUST00000215974.2 ENSMUST00000215894.2 |
Olfr107
|
olfactory receptor 107 |
| chrX_-_74621828 | 0.11 |
ENSMUST00000033545.6
|
Rab39b
|
RAB39B, member RAS oncogene family |
| chr3_+_32490300 | 0.11 |
ENSMUST00000029201.14
|
Pik3ca
|
phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha |
| chr17_-_32491339 | 0.10 |
ENSMUST00000237008.2
|
Brd4
|
bromodomain containing 4 |
| chr8_+_31579633 | 0.10 |
ENSMUST00000170204.8
|
Dusp26
|
dual specificity phosphatase 26 (putative) |
| chr18_+_46730765 | 0.09 |
ENSMUST00000238168.2
ENSMUST00000078079.11 ENSMUST00000168382.2 ENSMUST00000235849.2 ENSMUST00000235973.2 ENSMUST00000235455.2 ENSMUST00000237478.2 |
Eif1a
|
eukaryotic translation initiation factor 1A |
| chr16_+_32002750 | 0.09 |
ENSMUST00000231512.2
|
Bex6
|
brain expressed family member 6 |
| chr10_+_69623262 | 0.09 |
ENSMUST00000183240.2
|
Ank3
|
ankyrin 3, epithelial |
| chr10_+_33913593 | 0.08 |
ENSMUST00000095758.3
|
Trappc3l
|
trafficking protein particle complex 3 like |
| chr4_+_112089442 | 0.07 |
ENSMUST00000038455.12
ENSMUST00000170945.2 |
Skint3
|
selection and upkeep of intraepithelial T cells 3 |
| chr14_-_9015639 | 0.06 |
ENSMUST00000112656.4
|
Synpr
|
synaptoporin |
| chr10_+_101994719 | 0.06 |
ENSMUST00000138522.8
ENSMUST00000163753.8 ENSMUST00000138016.8 |
Mgat4c
|
MGAT4 family, member C |
| chr15_+_28203872 | 0.05 |
ENSMUST00000067048.8
|
Dnah5
|
dynein, axonemal, heavy chain 5 |
| chr3_+_37402495 | 0.05 |
ENSMUST00000138563.9
|
Fgf2
|
fibroblast growth factor 2 |
| chr6_-_130044234 | 0.04 |
ENSMUST00000119096.2
|
Klra4
|
killer cell lectin-like receptor, subfamily A, member 4 |
| chr10_-_5019044 | 0.04 |
ENSMUST00000095899.5
|
Syne1
|
spectrin repeat containing, nuclear envelope 1 |
| chr6_+_90078412 | 0.03 |
ENSMUST00000089417.8
ENSMUST00000226577.2 |
Vmn1r50
|
vomeronasal 1 receptor 50 |
| chr3_+_37402795 | 0.03 |
ENSMUST00000200585.5
ENSMUST00000038885.10 |
Fgf2
|
fibroblast growth factor 2 |
| chr11_-_102076028 | 0.02 |
ENSMUST00000107156.9
ENSMUST00000021297.6 |
Lsm12
|
LSM12 homolog |
| chr19_-_11816583 | 0.02 |
ENSMUST00000214887.2
|
Olfr1417
|
olfactory receptor 1417 |
| chr2_-_79959178 | 0.00 |
ENSMUST00000102654.11
ENSMUST00000102655.10 |
Pde1a
|
phosphodiesterase 1A, calmodulin-dependent |
Network of associatons between targets according to the STRING database.
First level regulatory network of Barhl1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 9.8 | GO:0001542 | ovulation from ovarian follicle(GO:0001542) |
| 0.6 | 3.9 | GO:0060309 | elastin catabolic process(GO:0060309) |
| 0.5 | 1.4 | GO:0006429 | leucyl-tRNA aminoacylation(GO:0006429) |
| 0.2 | 0.8 | GO:0061534 | gamma-aminobutyric acid secretion, neurotransmission(GO:0061534) |
| 0.2 | 1.4 | GO:0006004 | fucose metabolic process(GO:0006004) |
| 0.1 | 0.4 | GO:1904742 | regulation of telomeric DNA binding(GO:1904742) |
| 0.1 | 0.4 | GO:1901642 | purine nucleoside transmembrane transport(GO:0015860) nucleoside transmembrane transport(GO:1901642) |
| 0.1 | 0.4 | GO:0070125 | mitochondrial translational elongation(GO:0070125) |
| 0.1 | 0.5 | GO:0048539 | bone marrow development(GO:0048539) |
| 0.1 | 1.6 | GO:0032211 | negative regulation of telomere maintenance via telomerase(GO:0032211) |
| 0.1 | 0.7 | GO:2000346 | negative regulation of hepatocyte proliferation(GO:2000346) |
| 0.1 | 0.3 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.1 | 0.3 | GO:1902310 | positive regulation of peptidyl-serine dephosphorylation(GO:1902310) |
| 0.0 | 0.4 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.0 | 0.6 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.0 | 0.4 | GO:0032876 | negative regulation of DNA endoreduplication(GO:0032876) |
| 0.0 | 1.0 | GO:0001522 | pseudouridine synthesis(GO:0001522) |
| 0.0 | 0.1 | GO:0044029 | DNA hypomethylation(GO:0044028) hypomethylation of CpG island(GO:0044029) |
| 0.0 | 0.4 | GO:0040016 | embryonic cleavage(GO:0040016) |
| 0.0 | 1.5 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.0 | 3.0 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 4.3 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 0.2 | GO:1901979 | regulation of inward rectifier potassium channel activity(GO:1901979) |
| 0.0 | 1.3 | GO:0045070 | positive regulation of viral genome replication(GO:0045070) |
| 0.0 | 0.1 | GO:0021762 | substantia nigra development(GO:0021762) |
| 0.0 | 0.3 | GO:1990035 | calcium ion import across plasma membrane(GO:0098703) calcium ion import into cell(GO:1990035) |
| 0.0 | 1.0 | GO:0060612 | adipose tissue development(GO:0060612) |
| 0.0 | 0.4 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.0 | 0.4 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 0.0 | 0.5 | GO:0000184 | nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:0000184) |
| 0.0 | 0.1 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.6 | GO:1990879 | CST complex(GO:1990879) |
| 0.2 | 0.8 | GO:0000444 | MIS12/MIND type complex(GO:0000444) |
| 0.1 | 1.4 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.1 | 0.7 | GO:0097422 | tubular endosome(GO:0097422) |
| 0.0 | 0.6 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.0 | 0.4 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.0 | 0.6 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.0 | 0.1 | GO:0005943 | phosphatidylinositol 3-kinase complex, class IA(GO:0005943) |
| 0.0 | 4.3 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 0.2 | GO:0048237 | rough endoplasmic reticulum lumen(GO:0048237) |
| 0.0 | 0.8 | GO:0071437 | invadopodium(GO:0071437) |
| 0.0 | 1.5 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 3.0 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 0.5 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.0 | 1.3 | GO:0000502 | proteasome complex(GO:0000502) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.4 | 9.8 | GO:0004566 | beta-glucuronidase activity(GO:0004566) |
| 0.5 | 1.4 | GO:0004819 | glutamine-tRNA ligase activity(GO:0004819) leucine-tRNA ligase activity(GO:0004823) |
| 0.1 | 0.4 | GO:0005415 | nucleoside:sodium symporter activity(GO:0005415) |
| 0.1 | 0.7 | GO:0034452 | dynactin binding(GO:0034452) |
| 0.1 | 0.3 | GO:0023029 | MHC class Ib protein binding(GO:0023029) |
| 0.1 | 1.3 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 0.1 | 0.6 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.1 | 0.4 | GO:0010521 | telomerase inhibitor activity(GO:0010521) |
| 0.1 | 1.0 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.1 | 0.6 | GO:1901611 | phosphatidylglycerol binding(GO:1901611) |
| 0.0 | 0.8 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.0 | 0.3 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.0 | 0.4 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.0 | 0.7 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.0 | 1.6 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.0 | 4.3 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 0.2 | GO:1902282 | voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization(GO:1902282) |
| 0.0 | 0.1 | GO:0034211 | GTP-dependent protein kinase activity(GO:0034211) |
| 0.0 | 0.4 | GO:0016423 | tRNA (guanine) methyltransferase activity(GO:0016423) |
| 0.0 | 3.9 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.4 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.0 | 0.3 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.0 | 0.1 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 9.8 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.9 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.0 | 4.9 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.6 | PID IL3 PATHWAY | IL3-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.0 | 0.8 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 1.4 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.0 | 1.5 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 0.7 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.6 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 0.4 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 1.3 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |