Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
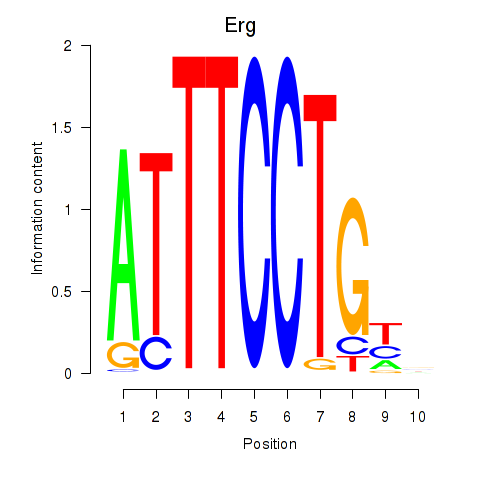
Results for Erg
Z-value: 2.95
Transcription factors associated with Erg
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Erg
|
ENSMUSG00000040732.21 | ETS transcription factor |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Erg | mm39_v1_chr16_-_95260104_95260243 | 0.77 | 3.7e-08 | Click! |
Activity profile of Erg motif
Sorted Z-values of Erg motif
Network of associatons between targets according to the STRING database.
First level regulatory network of Erg
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 26.8 | 80.4 | GO:0002148 | hypochlorous acid metabolic process(GO:0002148) hypochlorous acid biosynthetic process(GO:0002149) |
| 13.1 | 39.4 | GO:0035606 | peptidyl-cysteine S-trans-nitrosylation(GO:0035606) |
| 4.8 | 14.3 | GO:0001807 | regulation of type IV hypersensitivity(GO:0001807) |
| 4.1 | 12.3 | GO:0002625 | regulation of T cell antigen processing and presentation(GO:0002625) |
| 3.8 | 11.4 | GO:0071460 | cellular response to cell-matrix adhesion(GO:0071460) |
| 3.1 | 9.3 | GO:0042986 | positive regulation of amyloid precursor protein biosynthetic process(GO:0042986) |
| 3.0 | 15.0 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 2.5 | 15.1 | GO:0035585 | calcium-mediated signaling using extracellular calcium source(GO:0035585) |
| 2.5 | 7.4 | GO:0002880 | chronic inflammatory response to non-antigenic stimulus(GO:0002545) regulation of chronic inflammatory response to non-antigenic stimulus(GO:0002880) |
| 2.3 | 9.3 | GO:0033367 | protein localization to secretory granule(GO:0033366) protein localization to mast cell secretory granule(GO:0033367) protease localization to mast cell secretory granule(GO:0033368) maintenance of protein location in mast cell secretory granule(GO:0033370) T cell secretory granule organization(GO:0033371) maintenance of protease location in mast cell secretory granule(GO:0033373) protein localization to T cell secretory granule(GO:0033374) protease localization to T cell secretory granule(GO:0033375) maintenance of protein location in T cell secretory granule(GO:0033377) maintenance of protease location in T cell secretory granule(GO:0033379) granzyme B localization to T cell secretory granule(GO:0033380) maintenance of granzyme B location in T cell secretory granule(GO:0033382) |
| 2.2 | 8.8 | GO:0045659 | negative regulation of neutrophil differentiation(GO:0045659) |
| 2.1 | 12.8 | GO:0006742 | NADP catabolic process(GO:0006742) |
| 2.1 | 8.4 | GO:0090116 | C-5 methylation of cytosine(GO:0090116) positive regulation of methylation-dependent chromatin silencing(GO:0090309) |
| 2.1 | 8.3 | GO:0010286 | heat acclimation(GO:0010286) |
| 2.0 | 13.8 | GO:1903527 | positive regulation of membrane tubulation(GO:1903527) |
| 1.8 | 7.1 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 1.7 | 15.5 | GO:1901164 | negative regulation of trophoblast cell migration(GO:1901164) |
| 1.7 | 12.0 | GO:0010936 | negative regulation of macrophage cytokine production(GO:0010936) |
| 1.7 | 8.5 | GO:0038032 | termination of G-protein coupled receptor signaling pathway(GO:0038032) |
| 1.7 | 28.7 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 1.7 | 11.7 | GO:0003376 | sphingosine-1-phosphate signaling pathway(GO:0003376) |
| 1.7 | 8.3 | GO:1901355 | response to rapamycin(GO:1901355) |
| 1.7 | 9.9 | GO:0032439 | endosome localization(GO:0032439) negative regulation of vacuolar transport(GO:1903336) |
| 1.6 | 40.7 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 1.5 | 5.9 | GO:0014022 | neural plate elongation(GO:0014022) convergent extension involved in neural plate elongation(GO:0022007) |
| 1.4 | 15.2 | GO:0034723 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 1.4 | 12.4 | GO:0002291 | T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:0002291) |
| 1.3 | 3.8 | GO:2000097 | regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 1.3 | 12.7 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 1.3 | 3.8 | GO:2000566 | positive regulation of CD8-positive, alpha-beta T cell proliferation(GO:2000566) |
| 1.2 | 9.9 | GO:0071802 | negative regulation of podosome assembly(GO:0071802) |
| 1.2 | 16.0 | GO:0043313 | regulation of neutrophil degranulation(GO:0043313) |
| 1.2 | 5.9 | GO:0031509 | telomeric heterochromatin assembly(GO:0031509) negative regulation of chromosome condensation(GO:1902340) |
| 1.2 | 9.3 | GO:0097411 | hypoxia-inducible factor-1alpha signaling pathway(GO:0097411) |
| 1.1 | 3.4 | GO:0030472 | mitotic spindle organization in nucleus(GO:0030472) |
| 1.1 | 3.3 | GO:0046035 | CMP salvage(GO:0006238) CMP biosynthetic process(GO:0009224) pyrimidine ribonucleotide salvage(GO:0010138) pyrimidine nucleotide salvage(GO:0032262) UMP salvage(GO:0044206) CMP metabolic process(GO:0046035) |
| 1.1 | 3.2 | GO:0090673 | endothelial cell-matrix adhesion(GO:0090673) |
| 1.0 | 12.5 | GO:0046645 | positive regulation of gamma-delta T cell differentiation(GO:0045588) positive regulation of gamma-delta T cell activation(GO:0046645) |
| 1.0 | 3.1 | GO:1904959 | regulation of cytochrome-c oxidase activity(GO:1904959) |
| 1.0 | 4.0 | GO:0061534 | gamma-aminobutyric acid secretion, neurotransmission(GO:0061534) |
| 1.0 | 3.0 | GO:0000389 | mRNA 3'-splice site recognition(GO:0000389) |
| 1.0 | 7.7 | GO:0045046 | peroxisomal membrane transport(GO:0015919) protein import into peroxisome membrane(GO:0045046) |
| 1.0 | 5.8 | GO:0002528 | regulation of vascular permeability involved in acute inflammatory response(GO:0002528) |
| 0.9 | 17.1 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.9 | 13.2 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.9 | 4.7 | GO:0045347 | negative regulation of MHC class II biosynthetic process(GO:0045347) |
| 0.9 | 7.3 | GO:0072378 | blood coagulation, fibrin clot formation(GO:0072378) |
| 0.9 | 2.6 | GO:0006427 | histidyl-tRNA aminoacylation(GO:0006427) |
| 0.9 | 3.5 | GO:0021993 | initiation of neural tube closure(GO:0021993) |
| 0.9 | 4.3 | GO:0002309 | T cell proliferation involved in immune response(GO:0002309) |
| 0.8 | 5.0 | GO:1901228 | positive regulation of transcription from RNA polymerase II promoter involved in heart development(GO:1901228) |
| 0.8 | 2.5 | GO:0060715 | syncytiotrophoblast cell differentiation involved in labyrinthine layer development(GO:0060715) |
| 0.8 | 3.3 | GO:0071544 | diphosphoinositol polyphosphate catabolic process(GO:0071544) |
| 0.8 | 12.1 | GO:0045651 | positive regulation of macrophage differentiation(GO:0045651) |
| 0.8 | 3.2 | GO:2000349 | negative regulation of CD40 signaling pathway(GO:2000349) |
| 0.8 | 4.8 | GO:0051140 | regulation of NK T cell proliferation(GO:0051140) positive regulation of NK T cell proliferation(GO:0051142) |
| 0.8 | 3.2 | GO:0060800 | regulation of cell differentiation involved in embryonic placenta development(GO:0060800) |
| 0.8 | 5.5 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 0.8 | 3.9 | GO:1904975 | response to bleomycin(GO:1904975) cellular response to bleomycin(GO:1904976) |
| 0.7 | 7.5 | GO:0010724 | regulation of definitive erythrocyte differentiation(GO:0010724) |
| 0.7 | 6.6 | GO:0051534 | negative regulation of NFAT protein import into nucleus(GO:0051534) |
| 0.7 | 2.2 | GO:0038091 | VEGF-activated platelet-derived growth factor receptor signaling pathway(GO:0038086) positive regulation of cell proliferation by VEGF-activated platelet derived growth factor receptor signaling pathway(GO:0038091) |
| 0.7 | 14.5 | GO:0050930 | induction of positive chemotaxis(GO:0050930) |
| 0.7 | 6.6 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.6 | 5.0 | GO:0006868 | glutamine transport(GO:0006868) |
| 0.6 | 15.6 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.6 | 4.4 | GO:0002322 | B cell proliferation involved in immune response(GO:0002322) |
| 0.6 | 4.3 | GO:1903142 | positive regulation of endothelial cell development(GO:1901552) positive regulation of establishment of endothelial barrier(GO:1903142) |
| 0.6 | 6.0 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.6 | 3.6 | GO:2001170 | negative regulation of ATP biosynthetic process(GO:2001170) |
| 0.6 | 1.7 | GO:0051892 | negative regulation of cardioblast differentiation(GO:0051892) |
| 0.6 | 10.1 | GO:0002726 | positive regulation of T cell cytokine production(GO:0002726) |
| 0.6 | 3.9 | GO:0007190 | activation of adenylate cyclase activity(GO:0007190) |
| 0.6 | 2.2 | GO:0046084 | adenine salvage(GO:0006168) adenine metabolic process(GO:0046083) adenine biosynthetic process(GO:0046084) |
| 0.6 | 5.0 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.5 | 1.6 | GO:2000670 | positive regulation of dendritic cell apoptotic process(GO:2000670) |
| 0.5 | 2.2 | GO:0071586 | CAAX-box protein processing(GO:0071586) CAAX-box protein maturation(GO:0080120) |
| 0.5 | 1.6 | GO:0060671 | epithelial cell differentiation involved in embryonic placenta development(GO:0060671) epithelial cell morphogenesis involved in placental branching(GO:0060672) |
| 0.5 | 1.6 | GO:0032685 | negative regulation of granulocyte macrophage colony-stimulating factor production(GO:0032685) |
| 0.5 | 3.1 | GO:0051387 | negative regulation of neurotrophin TRK receptor signaling pathway(GO:0051387) |
| 0.5 | 1.5 | GO:1902626 | assembly of large subunit precursor of preribosome(GO:1902626) |
| 0.5 | 1.5 | GO:0030538 | embryonic genitalia morphogenesis(GO:0030538) |
| 0.5 | 6.5 | GO:0002002 | regulation of angiotensin levels in blood(GO:0002002) |
| 0.5 | 21.4 | GO:0015804 | neutral amino acid transport(GO:0015804) |
| 0.5 | 3.0 | GO:2000434 | regulation of protein neddylation(GO:2000434) negative regulation of protein neddylation(GO:2000435) |
| 0.5 | 1.4 | GO:0019389 | glucuronoside metabolic process(GO:0019389) |
| 0.5 | 7.5 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.5 | 0.9 | GO:0032053 | ciliary basal body organization(GO:0032053) |
| 0.5 | 1.4 | GO:2000847 | negative regulation of corticosteroid hormone secretion(GO:2000847) negative regulation of glucocorticoid secretion(GO:2000850) |
| 0.5 | 3.7 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.5 | 4.1 | GO:0090232 | positive regulation of spindle checkpoint(GO:0090232) |
| 0.5 | 1.4 | GO:0090076 | maintenance of mitochondrion location(GO:0051659) relaxation of skeletal muscle(GO:0090076) |
| 0.4 | 1.8 | GO:0071348 | cellular response to interleukin-11(GO:0071348) |
| 0.4 | 3.5 | GO:0043654 | recognition of apoptotic cell(GO:0043654) |
| 0.4 | 1.7 | GO:0001928 | regulation of exocyst assembly(GO:0001928) |
| 0.4 | 1.7 | GO:1901165 | positive regulation of trophoblast cell migration(GO:1901165) |
| 0.4 | 3.3 | GO:1900748 | positive regulation of vascular endothelial growth factor signaling pathway(GO:1900748) |
| 0.4 | 6.6 | GO:0046007 | negative regulation of activated T cell proliferation(GO:0046007) |
| 0.4 | 4.9 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.4 | 3.7 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.4 | 2.8 | GO:0010756 | positive regulation of plasminogen activation(GO:0010756) |
| 0.4 | 9.0 | GO:0038092 | nodal signaling pathway(GO:0038092) |
| 0.4 | 7.0 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.4 | 2.6 | GO:0010897 | negative regulation of triglyceride catabolic process(GO:0010897) |
| 0.4 | 11.5 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.4 | 2.2 | GO:0040010 | positive regulation of growth rate(GO:0040010) |
| 0.4 | 5.1 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 0.4 | 1.4 | GO:0010726 | positive regulation of hydrogen peroxide metabolic process(GO:0010726) |
| 0.3 | 18.1 | GO:0035411 | catenin import into nucleus(GO:0035411) |
| 0.3 | 2.8 | GO:0035385 | Roundabout signaling pathway(GO:0035385) |
| 0.3 | 5.4 | GO:0048194 | Golgi vesicle budding(GO:0048194) |
| 0.3 | 12.4 | GO:0002467 | germinal center formation(GO:0002467) |
| 0.3 | 2.0 | GO:1990481 | mRNA pseudouridine synthesis(GO:1990481) |
| 0.3 | 1.0 | GO:1903904 | negative regulation of establishment of T cell polarity(GO:1903904) negative regulation of T cell migration(GO:2000405) |
| 0.3 | 2.5 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.3 | 3.7 | GO:0010669 | epithelial structure maintenance(GO:0010669) |
| 0.3 | 6.5 | GO:0010759 | positive regulation of macrophage chemotaxis(GO:0010759) |
| 0.3 | 1.8 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 0.3 | 1.5 | GO:0038026 | reelin-mediated signaling pathway(GO:0038026) |
| 0.3 | 0.9 | GO:0044878 | mitotic cytokinesis checkpoint(GO:0044878) |
| 0.3 | 0.3 | GO:0032752 | interleukin-3 production(GO:0032632) regulation of interleukin-3 production(GO:0032672) positive regulation of interleukin-3 production(GO:0032752) interleukin-3 biosynthetic process(GO:0042223) regulation of interleukin-3 biosynthetic process(GO:0045399) positive regulation of interleukin-3 biosynthetic process(GO:0045401) positive regulation of granulocyte macrophage colony-stimulating factor biosynthetic process(GO:0045425) |
| 0.3 | 3.2 | GO:0098789 | pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.3 | 12.8 | GO:0006779 | porphyrin-containing compound biosynthetic process(GO:0006779) |
| 0.3 | 1.4 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 0.3 | 1.7 | GO:0050916 | sensory perception of sweet taste(GO:0050916) sensory perception of umami taste(GO:0050917) |
| 0.3 | 1.4 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.3 | 0.8 | GO:0070358 | actin polymerization-dependent cell motility(GO:0070358) |
| 0.3 | 4.4 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.3 | 5.5 | GO:0009950 | dorsal/ventral axis specification(GO:0009950) |
| 0.3 | 7.2 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.3 | 1.9 | GO:0000447 | endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000447) |
| 0.3 | 2.7 | GO:0090037 | positive regulation of protein kinase C signaling(GO:0090037) |
| 0.3 | 7.4 | GO:0007130 | synaptonemal complex assembly(GO:0007130) |
| 0.3 | 3.7 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.3 | 1.0 | GO:0071883 | activation of MAPK activity by adrenergic receptor signaling pathway(GO:0071883) |
| 0.3 | 3.1 | GO:0023041 | neuronal signal transduction(GO:0023041) |
| 0.3 | 29.2 | GO:0030168 | platelet activation(GO:0030168) |
| 0.2 | 6.2 | GO:0006379 | mRNA cleavage(GO:0006379) |
| 0.2 | 3.9 | GO:0071285 | cellular response to lithium ion(GO:0071285) |
| 0.2 | 19.2 | GO:0007189 | adenylate cyclase-activating G-protein coupled receptor signaling pathway(GO:0007189) |
| 0.2 | 4.1 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.2 | 2.9 | GO:0051418 | interphase microtubule nucleation by interphase microtubule organizing center(GO:0051415) microtubule nucleation by microtubule organizing center(GO:0051418) |
| 0.2 | 2.4 | GO:2000675 | negative regulation of type B pancreatic cell apoptotic process(GO:2000675) |
| 0.2 | 3.1 | GO:0061635 | regulation of protein complex stability(GO:0061635) |
| 0.2 | 4.3 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.2 | 1.4 | GO:2000504 | positive regulation of blood vessel remodeling(GO:2000504) |
| 0.2 | 21.5 | GO:2000649 | regulation of sodium ion transmembrane transporter activity(GO:2000649) |
| 0.2 | 2.1 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.2 | 0.8 | GO:0046882 | negative regulation of follicle-stimulating hormone secretion(GO:0046882) |
| 0.2 | 0.8 | GO:0036228 | protein targeting to nuclear inner membrane(GO:0036228) |
| 0.2 | 0.8 | GO:0007185 | transmembrane receptor protein tyrosine phosphatase signaling pathway(GO:0007185) |
| 0.2 | 1.6 | GO:0032596 | protein transport into membrane raft(GO:0032596) |
| 0.2 | 5.3 | GO:2000401 | regulation of lymphocyte migration(GO:2000401) |
| 0.2 | 3.7 | GO:0031954 | positive regulation of protein autophosphorylation(GO:0031954) |
| 0.2 | 2.4 | GO:0071073 | positive regulation of phospholipid biosynthetic process(GO:0071073) |
| 0.2 | 4.6 | GO:0036344 | platelet formation(GO:0030220) platelet morphogenesis(GO:0036344) |
| 0.2 | 0.8 | GO:1902774 | late endosome to lysosome transport(GO:1902774) |
| 0.2 | 0.8 | GO:0086053 | AV node cell to bundle of His cell communication by electrical coupling(GO:0086053) |
| 0.2 | 2.4 | GO:0097242 | beta-amyloid clearance(GO:0097242) |
| 0.2 | 3.4 | GO:0050913 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) sensory perception of bitter taste(GO:0050913) |
| 0.2 | 0.6 | GO:0070317 | negative regulation of G0 to G1 transition(GO:0070317) |
| 0.2 | 2.0 | GO:0090091 | positive regulation of extracellular matrix disassembly(GO:0090091) negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 0.2 | 1.1 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.2 | 0.6 | GO:0002946 | tRNA C5-cytosine methylation(GO:0002946) |
| 0.2 | 1.1 | GO:0043402 | glucocorticoid mediated signaling pathway(GO:0043402) |
| 0.2 | 5.8 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.2 | 20.4 | GO:0007229 | integrin-mediated signaling pathway(GO:0007229) |
| 0.2 | 1.6 | GO:0045759 | negative regulation of action potential(GO:0045759) |
| 0.2 | 7.4 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.2 | 3.4 | GO:0046085 | adenosine metabolic process(GO:0046085) |
| 0.2 | 16.5 | GO:0042100 | B cell proliferation(GO:0042100) |
| 0.2 | 3.8 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 0.2 | 1.5 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.2 | 4.5 | GO:0007095 | mitotic G2 DNA damage checkpoint(GO:0007095) |
| 0.2 | 2.7 | GO:0007096 | regulation of exit from mitosis(GO:0007096) |
| 0.2 | 1.8 | GO:1990440 | positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.2 | 0.5 | GO:0039663 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 0.1 | 1.2 | GO:0032792 | negative regulation of CREB transcription factor activity(GO:0032792) |
| 0.1 | 0.7 | GO:0097531 | mast cell chemotaxis(GO:0002551) mast cell migration(GO:0097531) |
| 0.1 | 5.6 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.1 | 0.8 | GO:0006543 | glutamine catabolic process(GO:0006543) |
| 0.1 | 7.5 | GO:0006730 | one-carbon metabolic process(GO:0006730) |
| 0.1 | 0.8 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
| 0.1 | 6.1 | GO:0033119 | negative regulation of RNA splicing(GO:0033119) |
| 0.1 | 0.8 | GO:0000479 | endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000479) |
| 0.1 | 2.7 | GO:0071157 | negative regulation of cell cycle arrest(GO:0071157) |
| 0.1 | 0.9 | GO:0099590 | neurotransmitter receptor internalization(GO:0099590) |
| 0.1 | 1.4 | GO:0051895 | negative regulation of focal adhesion assembly(GO:0051895) negative regulation of adherens junction organization(GO:1903392) |
| 0.1 | 5.1 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.1 | 3.3 | GO:0010447 | response to acidic pH(GO:0010447) |
| 0.1 | 2.4 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.1 | 5.1 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.1 | 3.5 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.1 | 0.3 | GO:1904093 | regulation of autophagic cell death(GO:1904092) negative regulation of autophagic cell death(GO:1904093) |
| 0.1 | 5.2 | GO:0002026 | regulation of the force of heart contraction(GO:0002026) |
| 0.1 | 1.6 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.1 | 2.7 | GO:0030225 | macrophage differentiation(GO:0030225) |
| 0.1 | 1.6 | GO:0002756 | MyD88-independent toll-like receptor signaling pathway(GO:0002756) |
| 0.1 | 6.5 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.1 | 1.4 | GO:0046825 | regulation of protein export from nucleus(GO:0046825) |
| 0.1 | 7.4 | GO:0006661 | phosphatidylinositol biosynthetic process(GO:0006661) |
| 0.1 | 1.8 | GO:0071545 | inositol phosphate catabolic process(GO:0071545) |
| 0.1 | 0.5 | GO:0048050 | post-embryonic eye morphogenesis(GO:0048050) post-embryonic camera-type eye morphogenesis(GO:0048597) |
| 0.1 | 0.4 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.1 | 0.5 | GO:2000255 | negative regulation of male germ cell proliferation(GO:2000255) |
| 0.1 | 1.2 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.1 | 0.9 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.1 | 1.7 | GO:0033234 | negative regulation of protein sumoylation(GO:0033234) |
| 0.1 | 1.5 | GO:1990001 | inhibition of cysteine-type endopeptidase activity(GO:0097340) zymogen inhibition(GO:0097341) inhibition of cysteine-type endopeptidase activity involved in apoptotic process(GO:1990001) |
| 0.1 | 0.5 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.1 | 7.1 | GO:0007596 | blood coagulation(GO:0007596) |
| 0.1 | 1.1 | GO:1904659 | glucose transmembrane transport(GO:1904659) |
| 0.1 | 1.3 | GO:0060218 | hematopoietic stem cell differentiation(GO:0060218) |
| 0.1 | 0.4 | GO:0002903 | negative regulation of B cell apoptotic process(GO:0002903) |
| 0.1 | 1.4 | GO:0006656 | phosphatidylcholine biosynthetic process(GO:0006656) |
| 0.1 | 0.4 | GO:0007144 | female meiosis I(GO:0007144) |
| 0.1 | 3.2 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.1 | 5.5 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.1 | 0.6 | GO:0070836 | caveola assembly(GO:0070836) |
| 0.1 | 15.1 | GO:0000398 | RNA splicing, via transesterification reactions with bulged adenosine as nucleophile(GO:0000377) mRNA splicing, via spliceosome(GO:0000398) |
| 0.1 | 1.0 | GO:0021694 | cerebellar Purkinje cell layer formation(GO:0021694) |
| 0.1 | 0.2 | GO:0016340 | calcium-dependent cell-matrix adhesion(GO:0016340) |
| 0.1 | 2.6 | GO:0007257 | activation of JUN kinase activity(GO:0007257) |
| 0.1 | 6.9 | GO:0031532 | actin cytoskeleton reorganization(GO:0031532) |
| 0.1 | 0.4 | GO:0048680 | positive regulation of axon regeneration(GO:0048680) |
| 0.1 | 1.4 | GO:0035455 | response to interferon-alpha(GO:0035455) |
| 0.1 | 0.8 | GO:0021670 | lateral ventricle development(GO:0021670) |
| 0.1 | 5.4 | GO:0016525 | negative regulation of angiogenesis(GO:0016525) |
| 0.1 | 7.6 | GO:0007160 | cell-matrix adhesion(GO:0007160) |
| 0.1 | 0.2 | GO:0042539 | hypotonic salinity response(GO:0042539) cellular hypotonic salinity response(GO:0071477) |
| 0.1 | 1.3 | GO:1900745 | positive regulation of p38MAPK cascade(GO:1900745) |
| 0.1 | 2.0 | GO:0002292 | T cell differentiation involved in immune response(GO:0002292) |
| 0.1 | 1.7 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.0 | 3.6 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.0 | 0.6 | GO:0045779 | negative regulation of bone resorption(GO:0045779) negative regulation of bone remodeling(GO:0046851) |
| 0.0 | 3.1 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.0 | 6.6 | GO:0000910 | cytokinesis(GO:0000910) |
| 0.0 | 0.3 | GO:0002051 | osteoblast fate commitment(GO:0002051) |
| 0.0 | 0.2 | GO:0071847 | TNFSF11-mediated signaling pathway(GO:0071847) negative regulation of osteoclast development(GO:2001205) |
| 0.0 | 1.6 | GO:0090162 | establishment of epithelial cell polarity(GO:0090162) |
| 0.0 | 0.6 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.0 | 2.9 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.0 | 1.9 | GO:0035458 | cellular response to interferon-beta(GO:0035458) |
| 0.0 | 2.0 | GO:0006360 | transcription from RNA polymerase I promoter(GO:0006360) |
| 0.0 | 1.3 | GO:0048266 | behavioral response to pain(GO:0048266) |
| 0.0 | 0.1 | GO:0032960 | regulation of inositol trisphosphate biosynthetic process(GO:0032960) positive regulation of inositol trisphosphate biosynthetic process(GO:0032962) |
| 0.0 | 0.3 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.0 | 0.3 | GO:0038180 | nerve growth factor signaling pathway(GO:0038180) |
| 0.0 | 1.8 | GO:0001541 | ovarian follicle development(GO:0001541) |
| 0.0 | 1.3 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.0 | 2.3 | GO:0006367 | transcription initiation from RNA polymerase II promoter(GO:0006367) |
| 0.0 | 2.2 | GO:0007188 | adenylate cyclase-modulating G-protein coupled receptor signaling pathway(GO:0007188) |
| 0.0 | 1.9 | GO:0030183 | B cell differentiation(GO:0030183) |
| 0.0 | 2.9 | GO:0051028 | mRNA transport(GO:0051028) |
| 0.0 | 0.9 | GO:0007099 | centriole replication(GO:0007099) |
| 0.0 | 0.7 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 0.5 | GO:0051457 | maintenance of protein location in nucleus(GO:0051457) |
| 0.0 | 2.1 | GO:0045637 | regulation of myeloid cell differentiation(GO:0045637) |
| 0.0 | 1.3 | GO:0008156 | negative regulation of DNA replication(GO:0008156) |
| 0.0 | 0.2 | GO:0097369 | sodium ion import(GO:0097369) sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.0 | 0.4 | GO:0048490 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.0 | 1.4 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.3 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.0 | 0.7 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.0 | 1.8 | GO:0007030 | Golgi organization(GO:0007030) |
| 0.0 | 4.9 | GO:0006936 | muscle contraction(GO:0006936) |
| 0.0 | 0.2 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.0 | 0.1 | GO:1990743 | protein sialylation(GO:1990743) |
| 0.0 | 0.5 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.0 | 0.6 | GO:0051567 | histone H3-K9 methylation(GO:0051567) |
| 0.0 | 2.2 | GO:0042113 | B cell activation(GO:0042113) |
| 0.0 | 2.6 | GO:0042254 | ribosome biogenesis(GO:0042254) |
| 0.0 | 0.5 | GO:1903078 | positive regulation of protein localization to plasma membrane(GO:1903078) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.6 | 78.7 | GO:0005766 | primary lysosome(GO:0005766) azurophil granule(GO:0042582) |
| 2.5 | 12.4 | GO:0034687 | integrin alphaL-beta2 complex(GO:0034687) |
| 2.4 | 7.1 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 2.2 | 15.1 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 2.0 | 11.9 | GO:0005826 | actomyosin contractile ring(GO:0005826) |
| 1.7 | 22.4 | GO:0031091 | platelet alpha granule(GO:0031091) |
| 1.7 | 5.0 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 1.5 | 25.7 | GO:0001931 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 1.5 | 14.9 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 1.4 | 4.1 | GO:0030905 | retromer, tubulation complex(GO:0030905) |
| 1.3 | 19.9 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 1.3 | 6.6 | GO:0071953 | elastic fiber(GO:0071953) |
| 1.2 | 6.0 | GO:0000799 | nuclear condensin complex(GO:0000799) |
| 1.1 | 15.0 | GO:0000778 | condensed nuclear chromosome kinetochore(GO:0000778) |
| 0.8 | 9.3 | GO:0042629 | mast cell granule(GO:0042629) |
| 0.8 | 3.4 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.8 | 5.6 | GO:0097443 | sorting endosome(GO:0097443) |
| 0.7 | 10.9 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.7 | 2.9 | GO:0008275 | gamma-tubulin small complex(GO:0008275) |
| 0.7 | 4.2 | GO:0005683 | U7 snRNP(GO:0005683) |
| 0.7 | 1.4 | GO:0031673 | H zone(GO:0031673) |
| 0.7 | 3.3 | GO:0045160 | myosin I complex(GO:0045160) |
| 0.6 | 3.8 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.6 | 3.7 | GO:0000120 | RNA polymerase I transcription factor complex(GO:0000120) |
| 0.6 | 2.5 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.6 | 6.2 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.6 | 7.4 | GO:0000801 | central element(GO:0000801) |
| 0.6 | 3.7 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.6 | 3.7 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.6 | 13.3 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.6 | 16.1 | GO:0031362 | anchored component of external side of plasma membrane(GO:0031362) |
| 0.6 | 41.3 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.5 | 5.9 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.5 | 2.6 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 0.5 | 3.7 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.5 | 3.7 | GO:0036396 | MIS complex(GO:0036396) |
| 0.5 | 7.3 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.5 | 15.3 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.4 | 1.7 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 0.4 | 14.1 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.4 | 23.0 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.4 | 3.7 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.4 | 3.2 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.4 | 3.2 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 0.4 | 7.7 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.3 | 7.6 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.3 | 16.9 | GO:0002102 | podosome(GO:0002102) |
| 0.3 | 1.0 | GO:0071001 | U4/U6 snRNP(GO:0071001) |
| 0.3 | 6.6 | GO:0071004 | U2-type prespliceosome(GO:0071004) |
| 0.3 | 8.3 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.3 | 5.3 | GO:0036038 | MKS complex(GO:0036038) |
| 0.3 | 5.0 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.3 | 0.8 | GO:0043512 | inhibin-betaglycan-ActRII complex(GO:0034673) inhibin A complex(GO:0043512) |
| 0.3 | 4.5 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.3 | 1.0 | GO:1990696 | stereocilia ankle link complex(GO:0002142) USH2 complex(GO:1990696) |
| 0.2 | 2.0 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.2 | 1.0 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.2 | 18.9 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.2 | 13.1 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.2 | 9.6 | GO:0008305 | integrin complex(GO:0008305) |
| 0.2 | 1.3 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.2 | 25.2 | GO:0001725 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.2 | 2.4 | GO:0016589 | NURF complex(GO:0016589) |
| 0.2 | 0.6 | GO:0098843 | postsynaptic endocytic zone(GO:0098843) |
| 0.2 | 2.0 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.2 | 1.4 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.2 | 2.6 | GO:0071437 | invadopodium(GO:0071437) |
| 0.2 | 8.5 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.2 | 3.9 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.2 | 9.3 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.2 | 3.2 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.1 | 3.1 | GO:0033276 | transcription factor TFTC complex(GO:0033276) |
| 0.1 | 0.9 | GO:0098536 | deuterosome(GO:0098536) |
| 0.1 | 0.3 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 0.1 | 3.9 | GO:0043196 | varicosity(GO:0043196) |
| 0.1 | 3.6 | GO:0005605 | basal lamina(GO:0005605) |
| 0.1 | 0.6 | GO:0060053 | neurofilament cytoskeleton(GO:0060053) |
| 0.1 | 4.7 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.1 | 1.4 | GO:0042587 | glycogen granule(GO:0042587) |
| 0.1 | 6.3 | GO:0032809 | neuronal cell body membrane(GO:0032809) |
| 0.1 | 8.8 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.1 | 2.4 | GO:0051286 | cell tip(GO:0051286) |
| 0.1 | 0.8 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.1 | 15.1 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.1 | 1.7 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.1 | 1.2 | GO:0030123 | AP-3 adaptor complex(GO:0030123) |
| 0.1 | 1.3 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.1 | 0.5 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.1 | 0.9 | GO:0090543 | Flemming body(GO:0090543) |
| 0.1 | 2.3 | GO:0031527 | filopodium membrane(GO:0031527) |
| 0.1 | 14.0 | GO:0005884 | actin filament(GO:0005884) |
| 0.1 | 1.7 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.1 | 2.2 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.1 | 12.4 | GO:0000922 | spindle pole(GO:0000922) |
| 0.1 | 1.5 | GO:0031414 | N-terminal protein acetyltransferase complex(GO:0031414) |
| 0.1 | 5.2 | GO:0031672 | A band(GO:0031672) |
| 0.1 | 2.8 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.1 | 1.2 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.1 | 3.5 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.1 | 13.7 | GO:0001726 | ruffle(GO:0001726) |
| 0.1 | 3.1 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.1 | 5.3 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.1 | 2.1 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.1 | 5.2 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.1 | 1.9 | GO:0030673 | axolemma(GO:0030673) |
| 0.1 | 1.1 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.1 | 0.7 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.1 | 2.0 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.1 | 1.5 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.0 | 0.3 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.0 | 0.7 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.0 | 0.1 | GO:1904602 | serotonin-activated cation-selective channel complex(GO:1904602) |
| 0.0 | 0.3 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.0 | 0.2 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.0 | 0.4 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.0 | 0.5 | GO:0016342 | catenin complex(GO:0016342) |
| 0.0 | 3.3 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 0.7 | GO:0035267 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 4.9 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 1.0 | GO:0044452 | nucleolar part(GO:0044452) |
| 0.0 | 0.1 | GO:0012507 | ER to Golgi transport vesicle membrane(GO:0012507) |
| 0.0 | 2.4 | GO:0000502 | proteasome complex(GO:0000502) |
| 0.0 | 0.9 | GO:0097546 | ciliary base(GO:0097546) |
| 0.0 | 3.1 | GO:0005923 | bicellular tight junction(GO:0005923) |
| 0.0 | 2.1 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 7.4 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 27.1 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.0 | 0.3 | GO:0000145 | exocyst(GO:0000145) |
| 0.0 | 0.5 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 1.5 | GO:0009898 | cytoplasmic side of plasma membrane(GO:0009898) |
| 0.0 | 0.3 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 0.0 | 2.5 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.0 | 0.2 | GO:0002080 | acrosomal membrane(GO:0002080) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.0 | 50.4 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 2.6 | 12.8 | GO:0042610 | CD8 receptor binding(GO:0042610) |
| 2.5 | 12.4 | GO:0030369 | ICAM-3 receptor activity(GO:0030369) |
| 2.4 | 14.4 | GO:0046624 | sphingolipid transporter activity(GO:0046624) |
| 2.0 | 9.8 | GO:0070051 | fibrinogen binding(GO:0070051) |
| 1.8 | 5.3 | GO:0051916 | granulocyte colony-stimulating factor binding(GO:0051916) |
| 1.7 | 15.5 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 1.5 | 19.9 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 1.3 | 13.3 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 1.2 | 9.9 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 1.2 | 4.7 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 1.1 | 9.1 | GO:0016312 | inositol bisphosphate phosphatase activity(GO:0016312) |
| 1.1 | 16.9 | GO:0030274 | LIM domain binding(GO:0030274) |
| 1.1 | 79.3 | GO:0004601 | peroxidase activity(GO:0004601) |
| 1.1 | 16.4 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 1.1 | 11.8 | GO:0004908 | interleukin-1 receptor activity(GO:0004908) |
| 1.1 | 3.2 | GO:0008281 | sulfonylurea receptor activity(GO:0008281) |
| 1.1 | 10.5 | GO:0004064 | arylesterase activity(GO:0004064) |
| 1.0 | 4.2 | GO:0071208 | histone pre-mRNA DCP binding(GO:0071208) |
| 1.0 | 4.1 | GO:1990460 | leptin receptor binding(GO:1990460) |
| 0.9 | 11.3 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.9 | 3.6 | GO:0051435 | BH4 domain binding(GO:0051435) |
| 0.9 | 2.6 | GO:0004821 | histidine-tRNA ligase activity(GO:0004821) |
| 0.9 | 7.7 | GO:0016308 | 1-phosphatidylinositol-4-phosphate 5-kinase activity(GO:0016308) |
| 0.8 | 3.4 | GO:0008160 | protein tyrosine phosphatase activator activity(GO:0008160) |
| 0.8 | 6.6 | GO:0019960 | C-X3-C chemokine binding(GO:0019960) |
| 0.8 | 7.7 | GO:0030911 | TPR domain binding(GO:0030911) |
| 0.8 | 2.3 | GO:0052593 | tryptamine:oxygen oxidoreductase (deaminating) activity(GO:0052593) aminoacetone:oxygen oxidoreductase(deaminating) activity(GO:0052594) aliphatic-amine oxidase activity(GO:0052595) phenethylamine:oxygen oxidoreductase (deaminating) activity(GO:0052596) |
| 0.8 | 10.5 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.7 | 2.2 | GO:0002055 | adenine binding(GO:0002055) adenine phosphoribosyltransferase activity(GO:0003999) |
| 0.7 | 6.7 | GO:0043208 | glycosphingolipid binding(GO:0043208) |
| 0.7 | 2.2 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 0.7 | 7.3 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.7 | 5.0 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.7 | 41.4 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.7 | 8.4 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.7 | 5.3 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.6 | 9.3 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.6 | 5.5 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.6 | 8.3 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.5 | 4.7 | GO:0015181 | arginine transmembrane transporter activity(GO:0015181) |
| 0.5 | 6.1 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.5 | 1.5 | GO:0034189 | very-low-density lipoprotein particle binding(GO:0034189) |
| 0.5 | 15.0 | GO:0043274 | phospholipase binding(GO:0043274) |
| 0.5 | 3.3 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.5 | 3.3 | GO:0034431 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.5 | 6.9 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.5 | 2.8 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.5 | 4.6 | GO:0043184 | vascular endothelial growth factor receptor 2 binding(GO:0043184) |
| 0.4 | 2.7 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.4 | 11.4 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.4 | 1.7 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 0.4 | 2.5 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 0.4 | 56.0 | GO:0004540 | ribonuclease activity(GO:0004540) |
| 0.4 | 1.6 | GO:0004949 | cannabinoid receptor activity(GO:0004949) |
| 0.4 | 3.6 | GO:0043262 | adenosine-diphosphatase activity(GO:0043262) |
| 0.4 | 28.7 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.4 | 27.6 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.4 | 1.9 | GO:0008853 | exodeoxyribonuclease III activity(GO:0008853) |
| 0.4 | 19.8 | GO:0015175 | neutral amino acid transmembrane transporter activity(GO:0015175) |
| 0.4 | 2.8 | GO:0043141 | ATP-dependent 5'-3' DNA helicase activity(GO:0043141) |
| 0.3 | 3.1 | GO:1903136 | cuprous ion binding(GO:1903136) |
| 0.3 | 2.7 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.3 | 2.4 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 0.3 | 3.7 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.3 | 5.7 | GO:0001163 | RNA polymerase I regulatory region sequence-specific DNA binding(GO:0001163) RNA polymerase I CORE element sequence-specific DNA binding(GO:0001164) |
| 0.3 | 3.0 | GO:0055104 | ligase inhibitor activity(GO:0055104) ubiquitin ligase inhibitor activity(GO:1990948) |
| 0.3 | 3.2 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.3 | 11.0 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.3 | 1.9 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.3 | 4.0 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.3 | 5.5 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.3 | 5.1 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.3 | 3.9 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.3 | 2.0 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.3 | 9.9 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.3 | 0.8 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.3 | 21.5 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.3 | 12.3 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.3 | 13.5 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.2 | 3.9 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.2 | 1.4 | GO:0019961 | interferon binding(GO:0019961) |
| 0.2 | 1.4 | GO:0030229 | very-low-density lipoprotein particle receptor activity(GO:0030229) |
| 0.2 | 1.4 | GO:0008970 | phosphatidylcholine 1-acylhydrolase activity(GO:0008970) |
| 0.2 | 1.8 | GO:0042731 | PH domain binding(GO:0042731) |
| 0.2 | 12.1 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.2 | 10.8 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.2 | 3.7 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.2 | 3.5 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.2 | 2.6 | GO:0046934 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol bisphosphate kinase activity(GO:0052813) |
| 0.2 | 1.4 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) |
| 0.2 | 1.0 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.2 | 0.8 | GO:0086077 | gap junction channel activity involved in AV node cell-bundle of His cell electrical coupling(GO:0086077) |
| 0.2 | 1.5 | GO:0001727 | lipid kinase activity(GO:0001727) |
| 0.2 | 7.6 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.2 | 5.6 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.2 | 4.7 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.2 | 3.6 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.2 | 14.9 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.2 | 0.5 | GO:0004958 | prostaglandin F receptor activity(GO:0004958) |
| 0.2 | 3.5 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.2 | 3.3 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.2 | 5.7 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.2 | 15.6 | GO:0005518 | collagen binding(GO:0005518) |
| 0.2 | 36.2 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.2 | 5.3 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.2 | 4.3 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.2 | 2.0 | GO:0031697 | beta-1 adrenergic receptor binding(GO:0031697) |
| 0.2 | 1.0 | GO:0070815 | peptidyl-lysine 5-dioxygenase activity(GO:0070815) |
| 0.2 | 1.1 | GO:0043423 | 3-phosphoinositide-dependent protein kinase binding(GO:0043423) |
| 0.2 | 0.6 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.2 | 5.4 | GO:0043548 | phosphatidylinositol 3-kinase binding(GO:0043548) |
| 0.1 | 8.5 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.1 | 1.2 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.1 | 3.6 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.1 | 3.4 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.1 | 0.8 | GO:0004359 | glutaminase activity(GO:0004359) |
| 0.1 | 0.9 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.1 | 4.0 | GO:0005112 | Notch binding(GO:0005112) |
| 0.1 | 1.2 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.1 | 9.0 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.1 | 10.1 | GO:0004004 | RNA helicase activity(GO:0003724) ATP-dependent RNA helicase activity(GO:0004004) |
| 0.1 | 1.5 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.1 | 1.7 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.1 | 0.5 | GO:0033906 | hyaluronoglucuronidase activity(GO:0033906) |
| 0.1 | 0.6 | GO:0070012 | oligopeptidase activity(GO:0070012) |
| 0.1 | 1.7 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.1 | 2.0 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.1 | 0.6 | GO:0034211 | GTP-dependent protein kinase activity(GO:0034211) |
| 0.1 | 5.6 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.1 | 4.6 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 3.3 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.1 | 1.1 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.1 | 1.4 | GO:0005338 | nucleotide-sugar transmembrane transporter activity(GO:0005338) |
| 0.1 | 1.0 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.1 | 0.8 | GO:0034711 | inhibin binding(GO:0034711) |
| 0.1 | 31.8 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.1 | 1.1 | GO:0005412 | glucose:sodium symporter activity(GO:0005412) |
| 0.1 | 4.1 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.1 | 0.4 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.1 | 2.6 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.1 | 0.6 | GO:0016428 | tRNA (cytosine-5-)-methyltransferase activity(GO:0016428) |
| 0.1 | 2.6 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.1 | 14.4 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 1.5 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.1 | 5.0 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.1 | 3.6 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.1 | 0.8 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.1 | 4.4 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.1 | 15.5 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.1 | 0.5 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.1 | 0.6 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.1 | 0.3 | GO:0019767 | IgE receptor activity(GO:0019767) |
| 0.1 | 0.4 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.1 | 4.8 | GO:0017048 | Rho GTPase binding(GO:0017048) |
| 0.1 | 0.3 | GO:0042296 | ISG15 transferase activity(GO:0042296) |
| 0.0 | 0.4 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.0 | 0.7 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.0 | 3.1 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 4.6 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 0.2 | GO:0035827 | rubidium ion transmembrane transporter activity(GO:0035827) |
| 0.0 | 2.0 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.0 | 2.3 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.0 | 6.2 | GO:0019903 | protein phosphatase binding(GO:0019903) |
| 0.0 | 0.3 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.0 | 1.0 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.0 | 0.8 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.6 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.0 | 0.2 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.0 | 0.9 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 0.1 | GO:0022850 | serotonin-gated cation channel activity(GO:0022850) |
| 0.0 | 1.0 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.0 | 1.6 | GO:0031072 | heat shock protein binding(GO:0031072) |
| 0.0 | 0.4 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.0 | 0.2 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.0 | 0.1 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.0 | 0.0 | GO:0045131 | pre-mRNA branch point binding(GO:0045131) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 5.5 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 1.3 | 79.1 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 1.0 | 14.3 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 1.0 | 13.3 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.9 | 9.2 | ST JAK STAT PATHWAY | Jak-STAT Pathway |
| 0.9 | 31.7 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.9 | 15.4 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.8 | 18.5 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.8 | 47.9 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.7 | 39.9 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.7 | 13.2 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.7 | 8.5 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.6 | 63.9 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.6 | 11.4 | PID AVB3 OPN PATHWAY | Osteopontin-mediated events |
| 0.6 | 22.2 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.6 | 44.7 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.6 | 14.4 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.6 | 7.3 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.5 | 31.2 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.5 | 20.0 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.3 | 11.0 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.3 | 17.5 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.3 | 5.7 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.2 | 2.2 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.2 | 11.2 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.2 | 5.7 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.2 | 3.9 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.2 | 7.9 | PID INTEGRIN1 PATHWAY | Beta1 integrin cell surface interactions |
| 0.2 | 4.3 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.2 | 2.7 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.2 | 3.9 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.2 | 9.6 | ST INTEGRIN SIGNALING PATHWAY | Integrin Signaling Pathway |
| 0.2 | 31.6 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.2 | 9.8 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.1 | 14.0 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.1 | 1.2 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.1 | 4.4 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.1 | 2.7 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.1 | 2.4 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.1 | 3.4 | PID FOXO PATHWAY | FoxO family signaling |
| 0.1 | 3.3 | PID MYC PATHWAY | C-MYC pathway |
| 0.1 | 3.3 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.1 | 5.6 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.1 | 2.4 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 2.2 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.1 | 0.7 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
| 0.1 | 2.8 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.1 | 12.6 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.1 | 4.5 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.1 | 1.5 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.1 | 1.4 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.1 | 4.4 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 6.6 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.0 | 0.7 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.8 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.0 | 0.8 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.0 | 1.4 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 0.6 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 1.0 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 0.2 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 29.3 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 1.1 | 22.9 | REACTOME P130CAS LINKAGE TO MAPK SIGNALING FOR INTEGRINS | Genes involved in p130Cas linkage to MAPK signaling for integrins |
| 1.0 | 16.9 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.7 | 10.7 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.6 | 7.8 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.6 | 40.7 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.6 | 15.5 | REACTOME INHIBITION OF THE PROTEOLYTIC ACTIVITY OF APC C REQUIRED FOR THE ONSET OF ANAPHASE BY MITOTIC SPINDLE CHECKPOINT COMPONENTS | Genes involved in Inhibition of the proteolytic activity of APC/C required for the onset of anaphase by mitotic spindle checkpoint components |
| 0.6 | 3.7 | REACTOME INHIBITION OF REPLICATION INITIATION OF DAMAGED DNA BY RB1 E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |
| 0.6 | 8.3 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.6 | 12.8 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.6 | 14.3 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.6 | 28.3 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.5 | 7.3 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.5 | 31.5 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.4 | 9.6 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.4 | 3.2 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.4 | 8.4 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.4 | 10.5 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.4 | 5.9 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.4 | 23.0 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.3 | 5.4 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.3 | 28.7 | REACTOME ANTIGEN PROCESSING CROSS PRESENTATION | Genes involved in Antigen processing-Cross presentation |
| 0.3 | 11.8 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.3 | 4.2 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.3 | 12.2 | REACTOME G BETA GAMMA SIGNALLING THROUGH PLC BETA | Genes involved in G beta:gamma signalling through PLC beta |
| 0.3 | 6.2 | REACTOME PROCESSING OF CAPPED INTRONLESS PRE MRNA | Genes involved in Processing of Capped Intronless Pre-mRNA |
| 0.3 | 7.3 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.3 | 20.0 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.3 | 6.0 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.2 | 16.1 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.2 | 7.6 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.2 | 3.3 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.2 | 3.9 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.2 | 10.4 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.2 | 4.0 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.2 | 7.6 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.2 | 45.9 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.2 | 3.7 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |
| 0.2 | 1.7 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |
| 0.2 | 2.9 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.2 | 16.7 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.2 | 10.8 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.1 | 5.7 | REACTOME TRANSPORT OF RIBONUCLEOPROTEINS INTO THE HOST NUCLEUS | Genes involved in Transport of Ribonucleoproteins into the Host Nucleus |
| 0.1 | 4.7 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.1 | 4.7 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 4.6 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.1 | 23.8 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.1 | 3.4 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.1 | 3.4 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 1.7 | REACTOME G ALPHA1213 SIGNALLING EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.1 | 0.5 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.1 | 1.4 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.1 | 3.3 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.1 | 2.0 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.1 | 0.8 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.1 | 1.4 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.1 | 4.1 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.1 | 3.2 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.1 | 1.2 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.1 | 1.3 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.1 | 1.6 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 1.1 | REACTOME ACTIVATION OF NF KAPPAB IN B CELLS | Genes involved in Activation of NF-kappaB in B Cells |
| 0.1 | 2.7 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.1 | 6.6 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 0.8 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |
| 0.0 | 0.6 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 0.5 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.0 | 1.2 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 0.9 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.0 | 1.5 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 0.5 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.0 | 0.2 | REACTOME PROCESSING OF CAPPED INTRON CONTAINING PRE MRNA | Genes involved in Processing of Capped Intron-Containing Pre-mRNA |
| 0.0 | 0.3 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 1.0 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 1.1 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 0.4 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |