Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
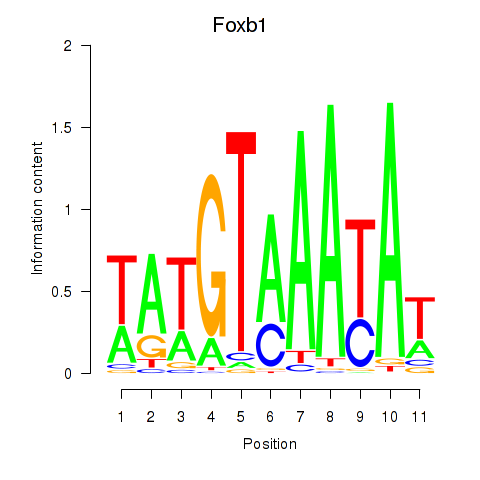
Results for Foxb1
Z-value: 0.55
Transcription factors associated with Foxb1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Foxb1
|
ENSMUSG00000059246.5 | forkhead box B1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Foxb1 | mm39_v1_chr9_-_69668204_69668222 | -0.15 | 4.0e-01 | Click! |
Activity profile of Foxb1 motif
Sorted Z-values of Foxb1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_20579322 | 5.58 |
ENSMUST00000087638.4
|
Aldh1a1
|
aldehyde dehydrogenase family 1, subfamily A1 |
| chr10_+_127734384 | 5.14 |
ENSMUST00000047134.8
|
Sdr9c7
|
4short chain dehydrogenase/reductase family 9C, member 7 |
| chr19_-_20704896 | 4.15 |
ENSMUST00000025656.4
|
Aldh1a7
|
aldehyde dehydrogenase family 1, subfamily A7 |
| chr7_+_119499322 | 1.55 |
ENSMUST00000106516.2
|
Lyrm1
|
LYR motif containing 1 |
| chr19_-_40175709 | 1.36 |
ENSMUST00000051846.13
|
Cyp2c70
|
cytochrome P450, family 2, subfamily c, polypeptide 70 |
| chr3_-_95811993 | 1.35 |
ENSMUST00000147962.3
ENSMUST00000036181.15 |
Car14
|
carbonic anhydrase 14 |
| chr3_-_146302343 | 1.33 |
ENSMUST00000029836.9
|
Dnase2b
|
deoxyribonuclease II beta |
| chr12_-_84497718 | 1.30 |
ENSMUST00000085192.7
ENSMUST00000220491.2 |
Aldh6a1
|
aldehyde dehydrogenase family 6, subfamily A1 |
| chr11_-_86884507 | 0.86 |
ENSMUST00000018571.5
|
Ypel2
|
yippee like 2 |
| chr7_+_27290969 | 0.79 |
ENSMUST00000108344.9
|
Akt2
|
thymoma viral proto-oncogene 2 |
| chr14_+_48358267 | 0.78 |
ENSMUST00000073150.6
|
Peli2
|
pellino 2 |
| chr3_+_96152813 | 0.74 |
ENSMUST00000078756.7
ENSMUST00000090779.4 |
H2ac18
Gm20634
|
H2A clustered histone 18 predicted gene 20634 |
| chr7_+_18962252 | 0.68 |
ENSMUST00000063976.9
|
Opa3
|
optic atrophy 3 |
| chr7_+_67305162 | 0.68 |
ENSMUST00000107470.2
|
Ttc23
|
tetratricopeptide repeat domain 23 |
| chr3_-_96147592 | 0.67 |
ENSMUST00000074976.8
|
H2ac19
|
H2A clustered histone 19 |
| chr7_+_27291126 | 0.64 |
ENSMUST00000167435.8
|
Akt2
|
thymoma viral proto-oncogene 2 |
| chr5_+_102629365 | 0.62 |
ENSMUST00000112854.8
|
Arhgap24
|
Rho GTPase activating protein 24 |
| chr1_-_170695328 | 0.62 |
ENSMUST00000027974.7
|
Atf6
|
activating transcription factor 6 |
| chr14_+_48358331 | 0.61 |
ENSMUST00000226513.2
|
Peli2
|
pellino 2 |
| chr4_-_135221810 | 0.60 |
ENSMUST00000105856.9
|
Nipal3
|
NIPA-like domain containing 3 |
| chr14_-_55880708 | 0.58 |
ENSMUST00000120041.8
ENSMUST00000121937.8 ENSMUST00000133707.2 ENSMUST00000002391.15 ENSMUST00000121791.8 |
Tm9sf1
|
transmembrane 9 superfamily member 1 |
| chr9_+_47441471 | 0.57 |
ENSMUST00000114548.8
ENSMUST00000152459.8 ENSMUST00000143026.9 ENSMUST00000085909.9 ENSMUST00000114547.8 ENSMUST00000239368.2 ENSMUST00000214542.2 ENSMUST00000034581.4 |
Cadm1
|
cell adhesion molecule 1 |
| chr7_+_18962301 | 0.57 |
ENSMUST00000161711.2
|
Opa3
|
optic atrophy 3 |
| chr12_-_32000534 | 0.56 |
ENSMUST00000172314.9
|
Hbp1
|
high mobility group box transcription factor 1 |
| chr5_+_3707171 | 0.54 |
ENSMUST00000198739.5
|
Tmbim7
|
transmembrane BAX inhibitor motif containing 7 |
| chr6_+_14901343 | 0.53 |
ENSMUST00000115477.8
|
Foxp2
|
forkhead box P2 |
| chr6_+_14901439 | 0.51 |
ENSMUST00000128567.8
|
Foxp2
|
forkhead box P2 |
| chr10_+_69761630 | 0.51 |
ENSMUST00000182029.8
|
Ank3
|
ankyrin 3, epithelial |
| chr16_-_44153288 | 0.51 |
ENSMUST00000136381.8
|
Sidt1
|
SID1 transmembrane family, member 1 |
| chr3_+_59939175 | 0.50 |
ENSMUST00000029325.5
|
Aadac
|
arylacetamide deacetylase |
| chr3_+_63148887 | 0.49 |
ENSMUST00000194324.6
|
Mme
|
membrane metallo endopeptidase |
| chr17_+_35188888 | 0.48 |
ENSMUST00000173680.2
|
Gm20481
|
predicted gene 20481 |
| chr2_-_147028309 | 0.47 |
ENSMUST00000067075.7
|
Nkx2-2
|
NK2 homeobox 2 |
| chr10_+_111342147 | 0.47 |
ENSMUST00000164773.2
|
Phlda1
|
pleckstrin homology like domain, family A, member 1 |
| chr5_+_102629240 | 0.47 |
ENSMUST00000073302.12
ENSMUST00000094559.9 |
Arhgap24
|
Rho GTPase activating protein 24 |
| chr11_+_78356523 | 0.47 |
ENSMUST00000001126.4
|
Slc46a1
|
solute carrier family 46, member 1 |
| chr4_-_135221926 | 0.47 |
ENSMUST00000102549.10
|
Nipal3
|
NIPA-like domain containing 3 |
| chr18_+_38809771 | 0.46 |
ENSMUST00000134388.2
ENSMUST00000148850.8 |
9630014M24Rik
Arhgap26
|
RIKEN cDNA 9630014M24 gene Rho GTPase activating protein 26 |
| chr2_-_45001141 | 0.43 |
ENSMUST00000201969.4
ENSMUST00000201623.4 |
Zeb2
|
zinc finger E-box binding homeobox 2 |
| chr12_-_32000209 | 0.42 |
ENSMUST00000176084.2
ENSMUST00000176103.8 ENSMUST00000167458.9 |
Hbp1
|
high mobility group box transcription factor 1 |
| chr12_+_112645237 | 0.41 |
ENSMUST00000174780.2
ENSMUST00000169593.2 ENSMUST00000173942.2 |
Zbtb42
|
zinc finger and BTB domain containing 42 |
| chr6_+_15184630 | 0.40 |
ENSMUST00000115470.3
|
Foxp2
|
forkhead box P2 |
| chr3_+_85946145 | 0.40 |
ENSMUST00000238331.2
|
Sh3d19
|
SH3 domain protein D19 |
| chr3_-_92922976 | 0.40 |
ENSMUST00000107301.2
ENSMUST00000029521.5 |
Crct1
|
cysteine-rich C-terminal 1 |
| chr6_+_8948608 | 0.40 |
ENSMUST00000160300.2
|
Nxph1
|
neurexophilin 1 |
| chr8_+_94763826 | 0.39 |
ENSMUST00000109556.9
ENSMUST00000093301.9 ENSMUST00000060632.8 |
Ogfod1
|
2-oxoglutarate and iron-dependent oxygenase domain containing 1 |
| chr16_-_44153498 | 0.39 |
ENSMUST00000047446.13
|
Sidt1
|
SID1 transmembrane family, member 1 |
| chrX_+_100419965 | 0.39 |
ENSMUST00000119080.8
|
Gjb1
|
gap junction protein, beta 1 |
| chr3_-_123483772 | 0.37 |
ENSMUST00000172537.3
|
Ndst3
|
N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 3 |
| chr6_+_15185202 | 0.36 |
ENSMUST00000154448.2
|
Foxp2
|
forkhead box P2 |
| chr6_-_41535322 | 0.33 |
ENSMUST00000193003.2
|
Trbv31
|
T cell receptor beta, variable 31 |
| chr1_-_39844467 | 0.33 |
ENSMUST00000171319.4
|
Gm3646
|
predicted gene 3646 |
| chr5_+_104318542 | 0.32 |
ENSMUST00000112771.2
|
Dspp
|
dentin sialophosphoprotein |
| chr12_+_56742413 | 0.31 |
ENSMUST00000001538.10
|
Pax9
|
paired box 9 |
| chr12_-_32000169 | 0.31 |
ENSMUST00000176520.8
|
Hbp1
|
high mobility group box transcription factor 1 |
| chr12_-_40249489 | 0.31 |
ENSMUST00000220951.2
|
Lsmem1
|
leucine-rich single-pass membrane protein 1 |
| chr4_+_11579648 | 0.31 |
ENSMUST00000180239.2
|
Fsbp
|
fibrinogen silencer binding protein |
| chr12_-_111946560 | 0.31 |
ENSMUST00000190680.2
|
Rd3l
|
retinal degeneration 3-like |
| chr6_-_41535292 | 0.30 |
ENSMUST00000103300.3
|
Trbv31
|
T cell receptor beta, variable 31 |
| chr11_-_98220466 | 0.30 |
ENSMUST00000041685.7
|
Neurod2
|
neurogenic differentiation 2 |
| chr15_+_102927366 | 0.29 |
ENSMUST00000165375.3
|
Hoxc4
|
homeobox C4 |
| chr3_-_129597679 | 0.29 |
ENSMUST00000185462.7
ENSMUST00000179187.2 |
Lrit3
|
leucine-rich repeat, immunoglobulin-like and transmembrane domains 3 |
| chr7_-_144761806 | 0.29 |
ENSMUST00000208788.2
|
Smim38
|
small integral membrane protein 38 |
| chr6_+_34840151 | 0.29 |
ENSMUST00000202010.2
|
Tmem140
|
transmembrane protein 140 |
| chr16_-_48232770 | 0.28 |
ENSMUST00000212197.2
|
Gm5485
|
predicted gene 5485 |
| chr4_-_139560831 | 0.28 |
ENSMUST00000030508.14
|
Pax7
|
paired box 7 |
| chr2_+_154042291 | 0.28 |
ENSMUST00000028987.7
|
Bpifb1
|
BPI fold containing family B, member 1 |
| chr5_-_51725059 | 0.28 |
ENSMUST00000127135.3
|
Ppargc1a
|
peroxisome proliferative activated receptor, gamma, coactivator 1 alpha |
| chr18_+_37085673 | 0.27 |
ENSMUST00000192512.6
ENSMUST00000192295.2 ENSMUST00000115661.5 |
Pcdha4
Gm42416
|
protocadherin alpha 4 predicted gene, 42416 |
| chr5_+_43673093 | 0.26 |
ENSMUST00000144558.3
|
C1qtnf7
|
C1q and tumor necrosis factor related protein 7 |
| chr17_+_70829050 | 0.25 |
ENSMUST00000133717.9
ENSMUST00000148486.8 |
Dlgap1
|
DLG associated protein 1 |
| chr10_+_69761597 | 0.24 |
ENSMUST00000182269.8
ENSMUST00000183261.8 ENSMUST00000183074.8 |
Ank3
|
ankyrin 3, epithelial |
| chr4_+_102287244 | 0.24 |
ENSMUST00000172616.2
|
Pde4b
|
phosphodiesterase 4B, cAMP specific |
| chr19_-_9065309 | 0.24 |
ENSMUST00000025554.3
|
Scgb1a1
|
secretoglobin, family 1A, member 1 (uteroglobin) |
| chr10_+_69761314 | 0.24 |
ENSMUST00000182692.8
ENSMUST00000092433.12 |
Ank3
|
ankyrin 3, epithelial |
| chr2_+_181409075 | 0.23 |
ENSMUST00000108757.9
|
Myt1
|
myelin transcription factor 1 |
| chr17_+_70829144 | 0.23 |
ENSMUST00000140728.8
|
Dlgap1
|
DLG associated protein 1 |
| chr17_-_35265702 | 0.22 |
ENSMUST00000097338.11
|
Msh5
|
mutS homolog 5 |
| chr12_-_40249314 | 0.22 |
ENSMUST00000095760.3
|
Lsmem1
|
leucine-rich single-pass membrane protein 1 |
| chr11_+_74252895 | 0.21 |
ENSMUST00000216362.2
|
Olfr412
|
olfactory receptor 412 |
| chr7_-_141009264 | 0.21 |
ENSMUST00000164387.2
ENSMUST00000137488.2 ENSMUST00000084436.10 |
Cend1
|
cell cycle exit and neuronal differentiation 1 |
| chr17_+_48037758 | 0.21 |
ENSMUST00000024782.12
ENSMUST00000144955.2 |
Pgc
|
progastricsin (pepsinogen C) |
| chrX_+_118836893 | 0.21 |
ENSMUST00000040961.3
ENSMUST00000113366.2 |
Pabpc5
|
poly(A) binding protein, cytoplasmic 5 |
| chr5_-_3707166 | 0.21 |
ENSMUST00000196304.2
|
Gatad1
|
GATA zinc finger domain containing 1 |
| chr4_+_127066667 | 0.20 |
ENSMUST00000106094.9
|
Dlgap3
|
DLG associated protein 3 |
| chrX_+_56257374 | 0.20 |
ENSMUST00000033466.2
|
Cd40lg
|
CD40 ligand |
| chr3_-_33136153 | 0.20 |
ENSMUST00000108225.10
|
Pex5l
|
peroxisomal biogenesis factor 5-like |
| chr14_-_52252318 | 0.20 |
ENSMUST00000228051.2
|
Zfp219
|
zinc finger protein 219 |
| chr7_-_141009346 | 0.20 |
ENSMUST00000124444.2
|
Cend1
|
cell cycle exit and neuronal differentiation 1 |
| chr18_-_25886750 | 0.19 |
ENSMUST00000224553.2
ENSMUST00000025117.14 |
Celf4
|
CUGBP, Elav-like family member 4 |
| chr4_-_102883905 | 0.19 |
ENSMUST00000084382.6
ENSMUST00000106869.3 |
Insl5
|
insulin-like 5 |
| chrX_+_108138965 | 0.19 |
ENSMUST00000033598.9
|
Sh3bgrl
|
SH3-binding domain glutamic acid-rich protein like |
| chr17_+_93506590 | 0.19 |
ENSMUST00000064775.8
|
Adcyap1
|
adenylate cyclase activating polypeptide 1 |
| chr15_+_102012782 | 0.18 |
ENSMUST00000230474.2
|
Tns2
|
tensin 2 |
| chr3_+_133942244 | 0.18 |
ENSMUST00000181904.3
|
Cxxc4
|
CXXC finger 4 |
| chr4_-_82970384 | 0.18 |
ENSMUST00000170248.9
|
Frem1
|
Fras1 related extracellular matrix protein 1 |
| chr14_+_33193765 | 0.16 |
ENSMUST00000208577.2
|
Frmpd2
|
FERM and PDZ domain containing 2 |
| chr8_-_65489791 | 0.16 |
ENSMUST00000124790.8
|
Apela
|
apelin receptor early endogenous ligand |
| chr17_+_93506435 | 0.15 |
ENSMUST00000234646.2
ENSMUST00000234081.2 |
Adcyap1
|
adenylate cyclase activating polypeptide 1 |
| chr2_+_181408833 | 0.15 |
ENSMUST00000108756.8
|
Myt1
|
myelin transcription factor 1 |
| chr7_-_119078330 | 0.15 |
ENSMUST00000207460.2
|
Umod
|
uromodulin |
| chr2_-_160701523 | 0.15 |
ENSMUST00000103112.8
|
Zhx3
|
zinc fingers and homeoboxes 3 |
| chr4_+_109092829 | 0.14 |
ENSMUST00000030285.8
|
Calr4
|
calreticulin 4 |
| chr1_+_179788037 | 0.14 |
ENSMUST00000097453.9
ENSMUST00000111117.8 |
Cdc42bpa
|
CDC42 binding protein kinase alpha |
| chr12_+_119291343 | 0.14 |
ENSMUST00000221917.2
|
Macc1
|
metastasis associated in colon cancer 1 |
| chr17_+_12597490 | 0.13 |
ENSMUST00000014578.7
|
Plg
|
plasminogen |
| chr2_+_109522781 | 0.12 |
ENSMUST00000111050.10
|
Bdnf
|
brain derived neurotrophic factor |
| chr12_+_113038376 | 0.11 |
ENSMUST00000109729.3
|
Tex22
|
testis expressed gene 22 |
| chr10_+_69761784 | 0.11 |
ENSMUST00000181974.8
ENSMUST00000182795.8 ENSMUST00000182437.8 |
Ank3
|
ankyrin 3, epithelial |
| chr10_-_49659355 | 0.10 |
ENSMUST00000105484.10
ENSMUST00000218598.2 ENSMUST00000079751.9 ENSMUST00000218441.2 |
Grik2
|
glutamate receptor, ionotropic, kainate 2 (beta 2) |
| chr18_+_69652751 | 0.10 |
ENSMUST00000200966.4
|
Tcf4
|
transcription factor 4 |
| chr1_+_187995096 | 0.10 |
ENSMUST00000060479.14
|
Ush2a
|
usherin |
| chr5_+_143803540 | 0.10 |
ENSMUST00000100487.6
|
Eif2ak1
|
eukaryotic translation initiation factor 2 alpha kinase 1 |
| chr2_-_60552980 | 0.09 |
ENSMUST00000028348.9
ENSMUST00000112517.8 |
Itgb6
|
integrin beta 6 |
| chr6_+_34840057 | 0.09 |
ENSMUST00000074949.4
|
Tmem140
|
transmembrane protein 140 |
| chr14_-_52252432 | 0.09 |
ENSMUST00000226527.2
|
Zfp219
|
zinc finger protein 219 |
| chr2_+_153742294 | 0.07 |
ENSMUST00000088955.12
ENSMUST00000135501.3 |
Bpifb6
|
BPI fold containing family B, member 6 |
| chr1_+_179788675 | 0.07 |
ENSMUST00000076687.12
ENSMUST00000097450.10 ENSMUST00000212756.2 |
Cdc42bpa
|
CDC42 binding protein kinase alpha |
| chr7_+_103593644 | 0.06 |
ENSMUST00000216006.2
|
Olfr633
|
olfactory receptor 633 |
| chr6_-_134768288 | 0.06 |
ENSMUST00000149776.3
|
Dusp16
|
dual specificity phosphatase 16 |
| chr3_-_102690026 | 0.06 |
ENSMUST00000172026.8
ENSMUST00000170856.6 |
Tshb
|
thyroid stimulating hormone, beta subunit |
| chr10_-_33662700 | 0.06 |
ENSMUST00000223295.2
|
Sult3a2
|
sulfotransferase family 3A, member 2 |
| chr19_-_11852453 | 0.06 |
ENSMUST00000213954.2
ENSMUST00000217617.2 |
Olfr1419
|
olfactory receptor 1419 |
| chr1_-_158642039 | 0.06 |
ENSMUST00000161589.3
|
Pappa2
|
pappalysin 2 |
| chr8_-_62576140 | 0.06 |
ENSMUST00000034052.14
ENSMUST00000034054.9 |
Anxa10
|
annexin A10 |
| chr1_+_70764874 | 0.05 |
ENSMUST00000053922.12
ENSMUST00000161937.2 ENSMUST00000162182.2 |
Vwc2l
|
von Willebrand factor C domain-containing protein 2-like |
| chr15_-_48655329 | 0.05 |
ENSMUST00000160658.8
ENSMUST00000100670.10 ENSMUST00000162830.8 |
Csmd3
|
CUB and Sushi multiple domains 3 |
| chr3_-_154760978 | 0.05 |
ENSMUST00000064076.6
|
Tnni3k
|
TNNI3 interacting kinase |
| chr7_-_103492361 | 0.04 |
ENSMUST00000063957.6
|
Hbb-bh1
|
hemoglobin Z, beta-like embryonic chain |
| chrX_-_42363663 | 0.03 |
ENSMUST00000016294.8
|
Tenm1
|
teneurin transmembrane protein 1 |
| chr6_-_3494587 | 0.03 |
ENSMUST00000049985.15
|
Hepacam2
|
HEPACAM family member 2 |
| chr5_-_87630117 | 0.03 |
ENSMUST00000079811.13
ENSMUST00000144144.3 |
Ugt2a2
|
UDP glucuronosyltransferase 2 family, polypeptide A2 |
| chr8_-_65489834 | 0.03 |
ENSMUST00000142822.4
|
Apela
|
apelin receptor early endogenous ligand |
| chrX_-_93166693 | 0.03 |
ENSMUST00000113925.8
|
Zfx
|
zinc finger protein X-linked |
| chr18_-_43820759 | 0.02 |
ENSMUST00000082254.8
|
Jakmip2
|
janus kinase and microtubule interacting protein 2 |
| chr15_+_43340609 | 0.02 |
ENSMUST00000022962.8
|
Emc2
|
ER membrane protein complex subunit 2 |
| chr9_-_101076198 | 0.02 |
ENSMUST00000066773.9
|
Ppp2r3a
|
protein phosphatase 2, regulatory subunit B'', alpha |
| chr4_+_109092610 | 0.02 |
ENSMUST00000106628.8
|
Calr4
|
calreticulin 4 |
| chr6_-_139478919 | 0.01 |
ENSMUST00000170650.3
|
Rergl
|
RERG/RAS-like |
| chr3_+_88762971 | 0.01 |
ENSMUST00000212694.2
|
Gon4l
|
gon-4-like (C.elegans) |
Network of associatons between targets according to the STRING database.
First level regulatory network of Foxb1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 5.6 | GO:0042904 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.3 | 1.3 | GO:0019859 | pyrimidine nucleobase catabolic process(GO:0006208) thymine catabolic process(GO:0006210) thymine metabolic process(GO:0019859) |
| 0.2 | 4.2 | GO:0042573 | retinoic acid metabolic process(GO:0042573) |
| 0.2 | 1.4 | GO:0071486 | cellular response to light intensity(GO:0071484) cellular response to high light intensity(GO:0071486) retinal rod cell apoptotic process(GO:0097473) |
| 0.2 | 0.5 | GO:0021529 | spinal cord oligodendrocyte cell differentiation(GO:0021529) spinal cord oligodendrocyte cell fate specification(GO:0021530) |
| 0.1 | 1.4 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.1 | 1.3 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 0.1 | 0.6 | GO:0042271 | susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.1 | 1.8 | GO:0060501 | positive regulation of epithelial cell proliferation involved in lung morphogenesis(GO:0060501) |
| 0.1 | 1.1 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.1 | 0.3 | GO:0071460 | cellular response to cell-matrix adhesion(GO:0071460) |
| 0.1 | 0.4 | GO:0021941 | radial glia guided migration of cerebellar granule cell(GO:0021933) negative regulation of cerebellar granule cell precursor proliferation(GO:0021941) |
| 0.1 | 0.3 | GO:1904633 | glomerular visceral epithelial cell apoptotic process(GO:1903210) regulation of glomerular visceral epithelial cell apoptotic process(GO:1904633) positive regulation of glomerular visceral epithelial cell apoptotic process(GO:1904635) positive regulation of progesterone biosynthetic process(GO:2000184) |
| 0.1 | 1.1 | GO:0010650 | positive regulation of cell communication by electrical coupling(GO:0010650) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 0.1 | 0.9 | GO:0033227 | dsRNA transport(GO:0033227) |
| 0.1 | 0.3 | GO:2000297 | negative regulation of synapse maturation(GO:2000297) |
| 0.1 | 0.5 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.1 | 0.2 | GO:0002225 | positive regulation of antimicrobial peptide production(GO:0002225) positive regulation of antibacterial peptide production(GO:0002803) |
| 0.1 | 0.3 | GO:0071651 | positive regulation of chemokine (C-C motif) ligand 5 production(GO:0071651) |
| 0.1 | 0.2 | GO:2001200 | positive regulation of dendritic cell differentiation(GO:2001200) |
| 0.1 | 0.5 | GO:0051958 | heme transport(GO:0015886) methotrexate transport(GO:0051958) |
| 0.1 | 1.3 | GO:0010745 | negative regulation of macrophage derived foam cell differentiation(GO:0010745) |
| 0.1 | 0.2 | GO:0032696 | negative regulation of interleukin-13 production(GO:0032696) |
| 0.1 | 1.3 | GO:0006309 | apoptotic DNA fragmentation(GO:0006309) |
| 0.1 | 1.1 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.0 | 0.3 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.0 | 0.4 | GO:0006449 | regulation of translational termination(GO:0006449) |
| 0.0 | 0.4 | GO:0015014 | heparan sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0015014) |
| 0.0 | 0.1 | GO:0061193 | taste bud development(GO:0061193) |
| 0.0 | 1.4 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.0 | 0.6 | GO:1990440 | positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.0 | 0.5 | GO:0070842 | aggresome assembly(GO:0070842) |
| 0.0 | 0.3 | GO:0009786 | regulation of asymmetric cell division(GO:0009786) |
| 0.0 | 0.5 | GO:0010898 | positive regulation of triglyceride catabolic process(GO:0010898) |
| 0.0 | 1.2 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.0 | 0.4 | GO:0015868 | purine ribonucleotide transport(GO:0015868) |
| 0.0 | 0.3 | GO:0014816 | skeletal muscle satellite cell differentiation(GO:0014816) |
| 0.0 | 0.3 | GO:0034144 | negative regulation of toll-like receptor 4 signaling pathway(GO:0034144) |
| 0.0 | 0.2 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) |
| 0.0 | 0.1 | GO:0046501 | protoporphyrinogen IX metabolic process(GO:0046501) |
| 0.0 | 0.2 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.0 | 0.1 | GO:0051918 | negative regulation of fibrinolysis(GO:0051918) |
| 0.0 | 0.1 | GO:0038044 | transforming growth factor-beta secretion(GO:0038044) |
| 0.0 | 0.2 | GO:1902866 | regulation of retina development in camera-type eye(GO:1902866) |
| 0.0 | 0.4 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.0 | 0.4 | GO:0060539 | diaphragm development(GO:0060539) |
| 0.0 | 0.2 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.0 | 0.1 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.0 | 0.2 | GO:2000253 | positive regulation of feeding behavior(GO:2000253) |
| 0.0 | 0.0 | GO:0072218 | ascending thin limb development(GO:0072021) thick ascending limb development(GO:0072023) metanephric ascending thin limb development(GO:0072218) metanephric thick ascending limb development(GO:0072233) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:1990844 | subsarcolemmal mitochondrion(GO:1990843) interfibrillar mitochondrion(GO:1990844) |
| 0.1 | 1.4 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.0 | 1.1 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.0 | 0.6 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.0 | 0.2 | GO:0017071 | intracellular cyclic nucleotide activated cation channel complex(GO:0017071) |
| 0.0 | 0.1 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.0 | 0.1 | GO:0044218 | other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 0.0 | 0.1 | GO:0034685 | integrin alphav-beta6 complex(GO:0034685) |
| 0.0 | 0.4 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 0.1 | GO:0032983 | kainate selective glutamate receptor complex(GO:0032983) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 9.7 | GO:0018479 | benzaldehyde dehydrogenase (NAD+) activity(GO:0018479) |
| 0.3 | 1.3 | GO:0004531 | deoxyribonuclease II activity(GO:0004531) |
| 0.2 | 5.1 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.1 | 0.9 | GO:0051033 | nucleic acid transmembrane transporter activity(GO:0051032) RNA transmembrane transporter activity(GO:0051033) |
| 0.1 | 0.4 | GO:0031544 | peptidyl-proline 3-dioxygenase activity(GO:0031544) |
| 0.1 | 0.5 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.1 | 1.3 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.1 | 0.2 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.1 | 1.1 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 0.4 | GO:0042328 | heparan sulfate N-acetylglucosaminyltransferase activity(GO:0042328) |
| 0.0 | 0.1 | GO:0005169 | neurotrophin TRKB receptor binding(GO:0005169) |
| 0.0 | 1.4 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.0 | 0.2 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.0 | 0.6 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 0.2 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.0 | 0.5 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.4 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.0 | 0.1 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.0 | 1.3 | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor(GO:0016620) |
| 0.0 | 0.2 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.0 | 0.1 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 0.0 | 1.1 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.0 | 1.1 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.0 | 0.3 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 2.8 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 0.3 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.4 | SA TRKA RECEPTOR | The TrkA receptor binds nerve growth factor to activate MAP kinase pathways and promote cell growth. |
| 0.0 | 1.3 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 0.9 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 5.6 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.1 | 1.4 | REACTOME IRAK1 RECRUITS IKK COMPLEX | Genes involved in IRAK1 recruits IKK complex |
| 0.1 | 1.4 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.1 | 0.6 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.1 | 1.3 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 1.1 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.5 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.4 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.0 | 0.6 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.1 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |