Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
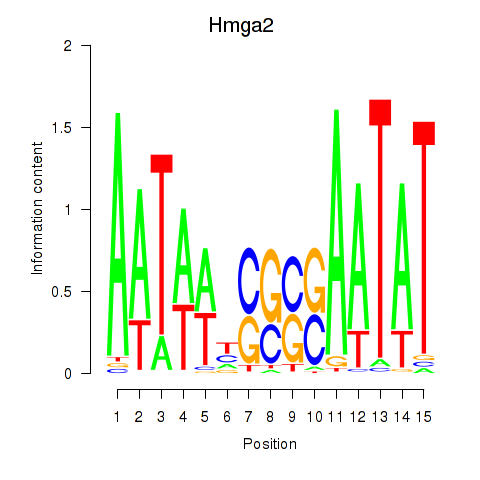
Results for Hmga2
Z-value: 0.68
Transcription factors associated with Hmga2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hmga2
|
ENSMUSG00000056758.15 | high mobility group AT-hook 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Hmga2 | mm39_v1_chr10_-_120312374_120312388 | -0.25 | 1.5e-01 | Click! |
Activity profile of Hmga2 motif
Sorted Z-values of Hmga2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_+_41498716 | 6.44 |
ENSMUST00000070380.5
|
Prss2
|
protease, serine 2 |
| chr6_-_41012435 | 4.91 |
ENSMUST00000031931.6
|
2210010C04Rik
|
RIKEN cDNA 2210010C04 gene |
| chr15_+_4756684 | 3.41 |
ENSMUST00000161997.8
ENSMUST00000022788.15 |
C6
|
complement component 6 |
| chr15_+_4756657 | 3.18 |
ENSMUST00000162585.8
|
C6
|
complement component 6 |
| chr6_+_41369290 | 3.10 |
ENSMUST00000049079.9
|
Gm5771
|
predicted gene 5771 |
| chr5_-_87288177 | 2.87 |
ENSMUST00000067790.7
|
Ugt2b5
|
UDP glucuronosyltransferase 2 family, polypeptide B5 |
| chr19_+_40078132 | 2.63 |
ENSMUST00000068094.13
ENSMUST00000080171.3 |
Cyp2c50
|
cytochrome P450, family 2, subfamily c, polypeptide 50 |
| chr5_-_87240405 | 2.51 |
ENSMUST00000132667.2
ENSMUST00000145617.8 ENSMUST00000094649.11 |
Ugt2b36
|
UDP glucuronosyltransferase 2 family, polypeptide B36 |
| chr5_-_87572060 | 2.38 |
ENSMUST00000072818.6
|
Ugt2b38
|
UDP glucuronosyltransferase 2 family, polypeptide B38 |
| chr19_-_39637489 | 1.77 |
ENSMUST00000067328.7
|
Cyp2c67
|
cytochrome P450, family 2, subfamily c, polypeptide 67 |
| chr16_-_18232202 | 1.60 |
ENSMUST00000165430.8
ENSMUST00000147720.3 |
Comt
|
catechol-O-methyltransferase |
| chr5_-_87402659 | 1.50 |
ENSMUST00000075858.4
|
Ugt2b37
|
UDP glucuronosyltransferase 2 family, polypeptide B37 |
| chr6_+_41435846 | 1.22 |
ENSMUST00000031910.8
|
Prss1
|
protease, serine 1 (trypsin 1) |
| chr2_-_93988229 | 1.17 |
ENSMUST00000028619.5
|
Hsd17b12
|
hydroxysteroid (17-beta) dehydrogenase 12 |
| chr13_+_4283729 | 0.97 |
ENSMUST00000081326.7
|
Akr1c19
|
aldo-keto reductase family 1, member C19 |
| chr7_+_143376871 | 0.97 |
ENSMUST00000128454.8
ENSMUST00000073878.12 |
Dhcr7
|
7-dehydrocholesterol reductase |
| chr6_+_42222841 | 0.95 |
ENSMUST00000031897.8
|
Gstk1
|
glutathione S-transferase kappa 1 |
| chr5_+_35740371 | 0.79 |
ENSMUST00000068947.14
ENSMUST00000114237.8 ENSMUST00000156125.8 ENSMUST00000202266.4 ENSMUST00000068563.12 |
Acox3
|
acyl-Coenzyme A oxidase 3, pristanoyl |
| chr5_+_87148697 | 0.74 |
ENSMUST00000031186.9
|
Ugt2b35
|
UDP glucuronosyltransferase 2 family, polypeptide B35 |
| chr11_-_115518774 | 0.67 |
ENSMUST00000154623.2
ENSMUST00000106503.10 ENSMUST00000141614.3 |
Slc25a19
|
solute carrier family 25 (mitochondrial thiamine pyrophosphate carrier), member 19 |
| chr15_-_82291372 | 0.66 |
ENSMUST00000230198.2
ENSMUST00000230248.2 ENSMUST00000072776.5 ENSMUST00000229911.2 |
Cyp2d10
|
cytochrome P450, family 2, subfamily d, polypeptide 10 |
| chrX_+_41149264 | 0.65 |
ENSMUST00000224454.2
|
Xiap
|
X-linked inhibitor of apoptosis |
| chr6_-_40976413 | 0.61 |
ENSMUST00000166306.3
|
Gm2663
|
predicted gene 2663 |
| chr7_-_14226851 | 0.58 |
ENSMUST00000108524.4
ENSMUST00000211740.2 ENSMUST00000209744.2 |
Sult2a7
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 7 |
| chr8_-_85620537 | 0.56 |
ENSMUST00000003907.14
ENSMUST00000109745.8 ENSMUST00000142748.2 |
Gcdh
|
glutaryl-Coenzyme A dehydrogenase |
| chr3_-_40801251 | 0.55 |
ENSMUST00000204017.2
ENSMUST00000026859.11 |
Mfsd8
|
major facilitator superfamily domain containing 8 |
| chr3_+_144283355 | 0.51 |
ENSMUST00000151086.3
|
Selenof
|
selenoprotein F |
| chr1_+_178146689 | 0.47 |
ENSMUST00000027781.7
|
Cox20
|
cytochrome c oxidase assembly protein 20 |
| chr3_+_40848580 | 0.46 |
ENSMUST00000159774.7
ENSMUST00000203472.3 ENSMUST00000203650.3 ENSMUST00000108077.10 ENSMUST00000203892.2 |
Abhd18
|
abhydrolase domain containing 18 |
| chrX_-_163763337 | 0.45 |
ENSMUST00000112248.9
|
Mospd2
|
motile sperm domain containing 2 |
| chr9_-_103165423 | 0.45 |
ENSMUST00000123530.8
|
1300017J02Rik
|
RIKEN cDNA 1300017J02 gene |
| chr6_+_40619913 | 0.43 |
ENSMUST00000238599.2
|
Mgam
|
maltase-glucoamylase |
| chr7_+_3620356 | 0.40 |
ENSMUST00000076657.11
ENSMUST00000108644.8 |
Ndufa3
|
NADH:ubiquinone oxidoreductase subunit A3 |
| chr17_+_73414977 | 0.40 |
ENSMUST00000130574.4
ENSMUST00000149064.9 ENSMUST00000067545.8 |
Lclat1
|
lysocardiolipin acyltransferase 1 |
| chr12_+_8258166 | 0.40 |
ENSMUST00000220274.2
|
Ldah
|
lipid droplet associated hydrolase |
| chr3_+_62245765 | 0.39 |
ENSMUST00000079300.13
|
Arhgef26
|
Rho guanine nucleotide exchange factor (GEF) 26 |
| chr1_+_172525613 | 0.39 |
ENSMUST00000038495.5
|
Crp
|
C-reactive protein, pentraxin-related |
| chr12_-_72964646 | 0.39 |
ENSMUST00000044000.12
|
4930447C04Rik
|
RIKEN cDNA 4930447C04 gene |
| chr7_+_129193581 | 0.37 |
ENSMUST00000084519.7
|
Wdr11
|
WD repeat domain 11 |
| chr13_+_74554509 | 0.37 |
ENSMUST00000222435.2
|
Ftl1-ps1
|
ferritin light polypeptide 1, pseudogene 1 |
| chr17_+_44389704 | 0.37 |
ENSMUST00000154166.8
ENSMUST00000024756.5 |
Enpp5
|
ectonucleotide pyrophosphatase/phosphodiesterase 5 |
| chr11_-_83193412 | 0.35 |
ENSMUST00000176374.2
|
Pex12
|
peroxisomal biogenesis factor 12 |
| chr7_-_24423715 | 0.35 |
ENSMUST00000081657.6
|
Lypd11
|
Ly6/PLAUR domain containing 11 |
| chr19_-_4384029 | 0.34 |
ENSMUST00000176653.2
|
Kdm2a
|
lysine (K)-specific demethylase 2A |
| chr10_+_125802084 | 0.34 |
ENSMUST00000074807.8
|
Lrig3
|
leucine-rich repeats and immunoglobulin-like domains 3 |
| chrX_+_100683662 | 0.34 |
ENSMUST00000119299.8
ENSMUST00000044475.5 |
Ogt
|
O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase) |
| chr12_+_8258107 | 0.33 |
ENSMUST00000037383.13
ENSMUST00000218883.2 ENSMUST00000218086.2 ENSMUST00000169104.3 ENSMUST00000217999.2 |
Ldah
|
lipid droplet associated hydrolase |
| chrM_+_9870 | 0.32 |
ENSMUST00000084013.1
|
mt-Nd4l
|
mitochondrially encoded NADH dehydrogenase 4L |
| chr8_+_71849497 | 0.31 |
ENSMUST00000002473.10
|
Babam1
|
BRISC and BRCA1 A complex member 1 |
| chr12_+_84332006 | 0.30 |
ENSMUST00000123614.8
ENSMUST00000147363.8 ENSMUST00000135001.8 ENSMUST00000146377.8 |
Ptgr2
|
prostaglandin reductase 2 |
| chr11_+_22921991 | 0.29 |
ENSMUST00000049506.8
|
Zrsr1
|
zinc finger (CCCH type), RNA binding motif and serine/arginine rich 1 |
| chr3_+_63148887 | 0.29 |
ENSMUST00000194324.6
|
Mme
|
membrane metallo endopeptidase |
| chrX_+_102400061 | 0.29 |
ENSMUST00000116547.3
|
Chic1
|
cysteine-rich hydrophobic domain 1 |
| chrM_+_10167 | 0.29 |
ENSMUST00000082414.1
|
mt-Nd4
|
mitochondrially encoded NADH dehydrogenase 4 |
| chr4_+_130001349 | 0.29 |
ENSMUST00000030563.6
|
Pef1
|
penta-EF hand domain containing 1 |
| chr14_+_55842002 | 0.28 |
ENSMUST00000138037.2
|
Irf9
|
interferon regulatory factor 9 |
| chr3_+_40801405 | 0.28 |
ENSMUST00000108078.9
|
Abhd18
|
abhydrolase domain containing 18 |
| chr10_-_93375832 | 0.28 |
ENSMUST00000016034.3
|
Amdhd1
|
amidohydrolase domain containing 1 |
| chr16_-_17019352 | 0.26 |
ENSMUST00000090192.12
|
Ube2l3
|
ubiquitin-conjugating enzyme E2L 3 |
| chr17_+_28075495 | 0.26 |
ENSMUST00000233809.2
|
Uhrf1bp1
|
UHRF1 (ICBP90) binding protein 1 |
| chr5_-_96312006 | 0.25 |
ENSMUST00000137207.2
|
Cnot6l
|
CCR4-NOT transcription complex, subunit 6-like |
| chr16_+_29398165 | 0.25 |
ENSMUST00000161186.8
ENSMUST00000038867.13 |
Opa1
|
OPA1, mitochondrial dynamin like GTPase |
| chr11_-_116197994 | 0.25 |
ENSMUST00000124281.2
|
Exoc7
|
exocyst complex component 7 |
| chr12_+_84147571 | 0.24 |
ENSMUST00000222921.2
|
Acot6
|
acyl-CoA thioesterase 6 |
| chr10_+_78864575 | 0.24 |
ENSMUST00000203906.3
|
Olfr57
|
olfactory receptor 57 |
| chr4_+_102971909 | 0.23 |
ENSMUST00000143417.8
|
Mier1
|
MEIR1 treanscription regulator |
| chr13_+_21938258 | 0.23 |
ENSMUST00000091709.3
|
H2bc15
|
H2B clustered histone 15 |
| chr13_+_19362068 | 0.23 |
ENSMUST00000103553.3
|
Trgv7
|
T cell receptor gamma, variable 7 |
| chr3_+_85481416 | 0.23 |
ENSMUST00000107672.8
ENSMUST00000127348.8 ENSMUST00000107674.2 |
Gatb
|
glutamyl-tRNA(Gln) amidotransferase, subunit B |
| chr19_-_5560473 | 0.22 |
ENSMUST00000237111.2
ENSMUST00000070172.6 |
Snx32
|
sorting nexin 32 |
| chr1_+_87983099 | 0.22 |
ENSMUST00000138182.8
ENSMUST00000113142.10 |
Ugt1a10
|
UDP glycosyltransferase 1 family, polypeptide A10 |
| chr11_+_62842019 | 0.22 |
ENSMUST00000035854.4
|
Cdrt4
|
CMT1A duplicated region transcript 4 |
| chr1_+_131936022 | 0.22 |
ENSMUST00000146432.2
|
Elk4
|
ELK4, member of ETS oncogene family |
| chr8_+_46984016 | 0.22 |
ENSMUST00000152423.2
|
Acsl1
|
acyl-CoA synthetase long-chain family member 1 |
| chr12_-_115825934 | 0.21 |
ENSMUST00000198777.2
|
Ighv1-77
|
immunoglobulin heavy variable 1-77 |
| chr4_+_137589548 | 0.21 |
ENSMUST00000102518.10
|
Ece1
|
endothelin converting enzyme 1 |
| chr16_-_95883714 | 0.21 |
ENSMUST00000113827.4
|
Brwd1
|
bromodomain and WD repeat domain containing 1 |
| chr5_+_14564932 | 0.21 |
ENSMUST00000182407.8
ENSMUST00000030691.17 |
Pclo
|
piccolo (presynaptic cytomatrix protein) |
| chr16_+_96162854 | 0.20 |
ENSMUST00000113794.8
|
Igsf5
|
immunoglobulin superfamily, member 5 |
| chrX_-_95499889 | 0.19 |
ENSMUST00000164693.8
ENSMUST00000119035.9 |
Hsf3
|
heat shock transcription factor 3 |
| chr8_-_69848167 | 0.19 |
ENSMUST00000072427.7
ENSMUST00000213012.2 ENSMUST00000239456.2 |
Gm10033
|
predicted gene 10033 |
| chr13_-_94422337 | 0.19 |
ENSMUST00000022197.15
ENSMUST00000152555.8 |
Scamp1
|
secretory carrier membrane protein 1 |
| chr14_+_53743184 | 0.18 |
ENSMUST00000103583.5
|
Trav10
|
T cell receptor alpha variable 10 |
| chr1_+_87983189 | 0.18 |
ENSMUST00000173325.2
|
Ugt1a10
|
UDP glycosyltransferase 1 family, polypeptide A10 |
| chr16_-_17019321 | 0.18 |
ENSMUST00000232668.2
ENSMUST00000231643.2 ENSMUST00000232035.2 |
Ube2l3
|
ubiquitin-conjugating enzyme E2L 3 |
| chrX_-_5995457 | 0.18 |
ENSMUST00000101698.4
ENSMUST00000117544.2 |
Ezhip
|
EZH inhibitory protein |
| chr5_-_74692327 | 0.18 |
ENSMUST00000072857.13
ENSMUST00000113542.9 ENSMUST00000151474.3 |
Scfd2
|
Sec1 family domain containing 2 |
| chr1_+_173877941 | 0.18 |
ENSMUST00000062665.4
|
Olfr432
|
olfactory receptor 432 |
| chr3_-_89152320 | 0.17 |
ENSMUST00000107464.8
ENSMUST00000090924.13 |
Trim46
|
tripartite motif-containing 46 |
| chr14_-_36641470 | 0.17 |
ENSMUST00000182042.2
|
Ccser2
|
coiled-coil serine rich 2 |
| chrM_+_9459 | 0.17 |
ENSMUST00000082411.1
|
mt-Nd3
|
mitochondrially encoded NADH dehydrogenase 3 |
| chr4_-_150087587 | 0.17 |
ENSMUST00000084117.13
|
H6pd
|
hexose-6-phosphate dehydrogenase (glucose 1-dehydrogenase) |
| chr11_+_83193495 | 0.17 |
ENSMUST00000176430.8
ENSMUST00000065692.14 ENSMUST00000142680.2 |
Ap2b1
|
adaptor-related protein complex 2, beta 1 subunit |
| chr6_+_116627567 | 0.16 |
ENSMUST00000067354.10
ENSMUST00000178241.4 |
Depp1
|
DEPP1 autophagy regulator |
| chr9_-_8134295 | 0.16 |
ENSMUST00000037397.8
|
Cep126
|
centrosomal protein 126 |
| chr9_-_20553576 | 0.16 |
ENSMUST00000155301.8
|
Fbxl12
|
F-box and leucine-rich repeat protein 12 |
| chr5_-_107073704 | 0.16 |
ENSMUST00000200249.2
ENSMUST00000112690.10 |
Hfm1
|
HFM1, ATP-dependent DNA helicase homolog |
| chr14_-_54949596 | 0.16 |
ENSMUST00000064290.8
|
Cebpe
|
CCAAT/enhancer binding protein (C/EBP), epsilon |
| chr13_+_41403317 | 0.16 |
ENSMUST00000165561.4
|
Smim13
|
small integral membrane protein 13 |
| chr19_+_37674029 | 0.16 |
ENSMUST00000073391.5
|
Cyp26c1
|
cytochrome P450, family 26, subfamily c, polypeptide 1 |
| chr14_-_54520382 | 0.15 |
ENSMUST00000059996.7
|
Olfr49
|
olfactory receptor 49 |
| chr6_+_116627635 | 0.15 |
ENSMUST00000204555.2
|
Depp1
|
DEPP1 autophagy regulator |
| chr14_-_36641270 | 0.15 |
ENSMUST00000182797.8
|
Ccser2
|
coiled-coil serine rich 2 |
| chr3_+_93349637 | 0.15 |
ENSMUST00000064257.6
|
Tchh
|
trichohyalin |
| chr11_+_77353218 | 0.14 |
ENSMUST00000102493.8
|
Coro6
|
coronin 6 |
| chr12_+_85871404 | 0.14 |
ENSMUST00000177188.8
ENSMUST00000095536.10 ENSMUST00000110220.9 ENSMUST00000040179.14 |
Ttll5
|
tubulin tyrosine ligase-like family, member 5 |
| chr1_+_46105898 | 0.14 |
ENSMUST00000069293.10
|
Dnah7b
|
dynein, axonemal, heavy chain 7B |
| chrX_+_73352694 | 0.14 |
ENSMUST00000130581.2
|
Gdi1
|
guanosine diphosphate (GDP) dissociation inhibitor 1 |
| chr2_-_84509172 | 0.14 |
ENSMUST00000111665.8
|
Tmx2
|
thioredoxin-related transmembrane protein 2 |
| chr11_+_100225233 | 0.14 |
ENSMUST00000017309.2
|
Gast
|
gastrin |
| chr2_-_127498129 | 0.14 |
ENSMUST00000028853.7
|
Mal
|
myelin and lymphocyte protein, T cell differentiation protein |
| chr17_+_31514780 | 0.13 |
ENSMUST00000237460.2
|
Slc37a1
|
solute carrier family 37 (glycerol-3-phosphate transporter), member 1 |
| chr6_-_106777014 | 0.13 |
ENSMUST00000013882.10
ENSMUST00000113239.10 |
Crbn
|
cereblon |
| chr16_+_32219324 | 0.13 |
ENSMUST00000115149.3
|
Tm4sf19
|
transmembrane 4 L six family member 19 |
| chr2_-_52225146 | 0.13 |
ENSMUST00000075301.10
|
Neb
|
nebulin |
| chr7_+_27207226 | 0.13 |
ENSMUST00000125990.2
ENSMUST00000065487.7 |
Prx
|
periaxin |
| chr6_-_69658959 | 0.13 |
ENSMUST00000103345.4
|
Igkv4-51
|
immunoglobulin kappa chain variable 4-51 |
| chr2_-_132787790 | 0.13 |
ENSMUST00000038280.5
|
Fermt1
|
fermitin family member 1 |
| chr6_-_128558560 | 0.13 |
ENSMUST00000060574.9
|
A2ml1
|
alpha-2-macroglobulin like 1 |
| chr4_+_109091682 | 0.13 |
ENSMUST00000106629.8
|
Calr4
|
calreticulin 4 |
| chr5_+_90608751 | 0.13 |
ENSMUST00000031314.10
|
Alb
|
albumin |
| chr2_-_155534295 | 0.13 |
ENSMUST00000041059.12
|
Trpc4ap
|
transient receptor potential cation channel, subfamily C, member 4 associated protein |
| chr4_+_44756608 | 0.13 |
ENSMUST00000143385.2
|
Zcchc7
|
zinc finger, CCHC domain containing 7 |
| chr2_-_18053595 | 0.13 |
ENSMUST00000142856.2
|
Skida1
|
SKI/DACH domain containing 1 |
| chr7_+_23451695 | 0.13 |
ENSMUST00000228331.2
|
Vmn1r174
|
vomeronasal 1 receptor 174 |
| chr5_+_7354130 | 0.12 |
ENSMUST00000160634.2
ENSMUST00000159546.2 |
Tex47
|
testis expressed 47 |
| chr4_+_101353742 | 0.12 |
ENSMUST00000154120.9
ENSMUST00000106930.8 |
Dnajc6
|
DnaJ heat shock protein family (Hsp40) member C6 |
| chrX_-_16683578 | 0.12 |
ENSMUST00000040820.13
|
Maob
|
monoamine oxidase B |
| chr16_+_96162949 | 0.12 |
ENSMUST00000000163.13
ENSMUST00000081093.10 ENSMUST00000113795.8 |
Igsf5
Gm49948
|
immunoglobulin superfamily, member 5 predicted gene, 49948 |
| chrX_-_138963461 | 0.12 |
ENSMUST00000133780.8
|
Nup62cl
|
nucleoporin 62 C-terminal like |
| chr19_-_57185861 | 0.12 |
ENSMUST00000111550.8
|
Ablim1
|
actin-binding LIM protein 1 |
| chr2_+_179684288 | 0.12 |
ENSMUST00000041126.9
|
Ss18l1
|
SS18, nBAF chromatin remodeling complex subunit like 1 |
| chr16_-_18407558 | 0.12 |
ENSMUST00000232589.2
|
Tbx1
|
T-box 1 |
| chr4_+_111830119 | 0.12 |
ENSMUST00000106568.8
ENSMUST00000055014.11 ENSMUST00000163281.2 |
Skint7
|
selection and upkeep of intraepithelial T cells 7 |
| chr19_+_6234392 | 0.11 |
ENSMUST00000025699.9
ENSMUST00000113528.2 |
Majin
|
membrane anchored junction protein |
| chr14_-_52616625 | 0.11 |
ENSMUST00000214980.2
|
Olfr1512
|
olfactory receptor 1512 |
| chr2_-_162929732 | 0.11 |
ENSMUST00000094653.6
|
Gtsf1l
|
gametocyte specific factor 1-like |
| chr7_-_8203319 | 0.11 |
ENSMUST00000086282.13
ENSMUST00000146278.9 ENSMUST00000142934.3 |
Vmn2r42
|
vomeronasal 2, receptor 42 |
| chr7_-_9575286 | 0.11 |
ENSMUST00000174433.2
|
Gm10302
|
predicted gene 10302 |
| chr12_-_36303394 | 0.11 |
ENSMUST00000221155.2
|
Lrrc72
|
leucine rich repeat containing 72 |
| chr7_-_101517874 | 0.11 |
ENSMUST00000150184.2
|
Folr1
|
folate receptor 1 (adult) |
| chr2_+_163389068 | 0.11 |
ENSMUST00000109411.8
ENSMUST00000018094.13 |
Hnf4a
|
hepatic nuclear factor 4, alpha |
| chr12_-_115019136 | 0.11 |
ENSMUST00000103519.2
ENSMUST00000192724.2 |
Ighv1-49
|
immunoglobulin heavy variable 1-49 |
| chr1_+_46106006 | 0.11 |
ENSMUST00000238212.2
|
Dnah7b
|
dynein, axonemal, heavy chain 7B |
| chr1_+_74627506 | 0.11 |
ENSMUST00000113732.2
|
Bcs1l
|
BCS1-like (yeast) |
| chr10_+_128540049 | 0.10 |
ENSMUST00000217836.2
|
Pmel
|
premelanosome protein |
| chr14_+_53728572 | 0.10 |
ENSMUST00000181210.3
ENSMUST00000183488.2 |
Trav6-5
|
T cell receptor alpha variable 6-5 |
| chr7_-_7340608 | 0.10 |
ENSMUST00000174368.9
ENSMUST00000072475.9 |
Vmn2r30
|
vomeronasal 2, receptor 30 |
| chr18_+_37125424 | 0.10 |
ENSMUST00000194038.2
|
Pcdha8
|
protocadherin alpha 8 |
| chr2_+_67935015 | 0.10 |
ENSMUST00000042456.4
|
B3galt1
|
UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 1 |
| chr7_-_8492074 | 0.10 |
ENSMUST00000164845.4
|
Vmn2r45
|
vomeronasal 2, receptor 45 |
| chr6_-_123395075 | 0.10 |
ENSMUST00000172199.3
|
Vmn2r20
|
vomeronasal 2, receptor 20 |
| chr4_+_44756553 | 0.10 |
ENSMUST00000107824.9
|
Zcchc7
|
zinc finger, CCHC domain containing 7 |
| chr16_+_29398149 | 0.10 |
ENSMUST00000160597.8
|
Opa1
|
OPA1, mitochondrial dynamin like GTPase |
| chr17_+_21446349 | 0.10 |
ENSMUST00000235895.2
|
Vmn1r234
|
vomeronasal 1 receptor 234 |
| chr7_-_101518217 | 0.10 |
ENSMUST00000123321.8
|
Folr1
|
folate receptor 1 (adult) |
| chr19_+_45549009 | 0.10 |
ENSMUST00000047057.9
|
Gm17018
|
predicted gene 17018 |
| chr4_-_32923505 | 0.10 |
ENSMUST00000084749.8
|
Ankrd6
|
ankyrin repeat domain 6 |
| chr4_-_32923389 | 0.09 |
ENSMUST00000035719.11
|
Ankrd6
|
ankyrin repeat domain 6 |
| chr9_+_3013140 | 0.09 |
ENSMUST00000143083.3
|
Gm10721
|
predicted gene 10721 |
| chr5_-_151510389 | 0.09 |
ENSMUST00000165928.4
|
Vmn2r18
|
vomeronasal 2, receptor 18 |
| chr7_-_10900086 | 0.09 |
ENSMUST00000067210.12
|
Zscan4d
|
zinc finger and SCAN domain containing 4D |
| chr8_-_3928495 | 0.09 |
ENSMUST00000209176.2
ENSMUST00000011445.8 |
Cd209d
|
CD209d antigen |
| chrX_+_139808351 | 0.09 |
ENSMUST00000033806.5
|
Vsig1
|
V-set and immunoglobulin domain containing 1 |
| chr7_-_42291971 | 0.09 |
ENSMUST00000098503.9
|
Zfp976
|
zinc finger protein 976 |
| chr18_+_66005891 | 0.09 |
ENSMUST00000173985.10
|
Grp
|
gastrin releasing peptide |
| chr2_+_87854404 | 0.08 |
ENSMUST00000217006.2
|
Olfr1161
|
olfactory receptor 1161 |
| chr11_+_95925711 | 0.08 |
ENSMUST00000006217.10
ENSMUST00000107700.4 |
Snf8
|
SNF8, ESCRT-II complex subunit, homolog (S. cerevisiae) |
| chr11_-_116197478 | 0.08 |
ENSMUST00000126731.8
|
Exoc7
|
exocyst complex component 7 |
| chr11_-_116197523 | 0.08 |
ENSMUST00000133468.2
ENSMUST00000106411.10 ENSMUST00000106413.10 ENSMUST00000021147.14 |
Exoc7
|
exocyst complex component 7 |
| chr9_-_14682300 | 0.08 |
ENSMUST00000191047.7
ENSMUST00000060330.5 |
1700012B09Rik
|
RIKEN cDNA 1700012B09 gene |
| chr7_-_12156516 | 0.08 |
ENSMUST00000120220.3
|
Zfp551
|
zinc fingr protein 551 |
| chr4_-_86476502 | 0.08 |
ENSMUST00000030216.6
|
Saxo1
|
stabilizer of axonemal microtubules 1 |
| chrX_+_138963623 | 0.08 |
ENSMUST00000044806.3
|
Dnaaf6b
|
dynein axonemal assembly factor 6B |
| chr9_+_3004457 | 0.08 |
ENSMUST00000178348.2
|
Gm11168
|
predicted gene 11168 |
| chr1_+_134422366 | 0.08 |
ENSMUST00000047978.9
|
Rabif
|
RAB interacting factor |
| chr2_+_150940165 | 0.08 |
ENSMUST00000109888.2
|
Gm14147
|
predicted gene 14147 |
| chr7_-_103420801 | 0.08 |
ENSMUST00000106878.3
|
Olfr69
|
olfactory receptor 69 |
| chr9_+_88721217 | 0.07 |
ENSMUST00000163255.9
ENSMUST00000186363.2 |
Trim43c
|
tripartite motif-containing 43C |
| chr7_+_6233178 | 0.07 |
ENSMUST00000165445.3
|
Zscan5b
|
zinc finger and SCAN domain containing 5B |
| chr4_+_52964547 | 0.07 |
ENSMUST00000215010.2
ENSMUST00000215127.2 |
Olfr270
|
olfactory receptor 270 |
| chr14_+_65835995 | 0.07 |
ENSMUST00000150897.8
|
Nuggc
|
nuclear GTPase, germinal center associated |
| chr4_+_57637817 | 0.07 |
ENSMUST00000150412.4
|
Pakap
|
paralemmin A kinase anchor protein |
| chr4_-_144513121 | 0.07 |
ENSMUST00000105746.3
|
Aadacl4fm5
|
AADACL4 family member 5 |
| chr9_+_92191415 | 0.07 |
ENSMUST00000150594.8
ENSMUST00000098477.8 |
1700057G04Rik
|
RIKEN cDNA 1700057G04 gene |
| chr2_-_30249202 | 0.07 |
ENSMUST00000100215.11
ENSMUST00000113620.10 ENSMUST00000163668.3 ENSMUST00000028214.15 ENSMUST00000113621.10 |
Sh3glb2
|
SH3-domain GRB2-like endophilin B2 |
| chr9_-_55917834 | 0.07 |
ENSMUST00000217105.2
|
4930563M21Rik
|
RIKEN cDNA 4930563M21 gene |
| chr7_-_7692512 | 0.07 |
ENSMUST00000173459.3
|
Vmn2r34
|
vomeronasal 2, receptor 34 |
| chr2_-_86208737 | 0.07 |
ENSMUST00000217435.2
|
Olfr1057
|
olfactory receptor 1057 |
| chr1_+_118104272 | 0.07 |
ENSMUST00000186264.2
|
Gm29106
|
predicted gene 29106 |
| chr11_+_108573428 | 0.07 |
ENSMUST00000106718.10
ENSMUST00000106715.8 ENSMUST00000106724.10 |
Cep112
|
centrosomal protein 112 |
| chr11_+_58062467 | 0.07 |
ENSMUST00000020820.2
|
Mrpl22
|
mitochondrial ribosomal protein L22 |
| chr2_+_111130980 | 0.07 |
ENSMUST00000216697.2
ENSMUST00000213823.2 |
Olfr1279
|
olfactory receptor 1279 |
| chr15_-_37458768 | 0.06 |
ENSMUST00000116445.9
|
Ncald
|
neurocalcin delta |
| chr6_-_70021662 | 0.06 |
ENSMUST00000196959.2
|
Igkv8-34
|
immunoglobulin kappa variable 8-34 |
| chr13_+_22534534 | 0.06 |
ENSMUST00000226909.2
ENSMUST00000227167.2 ENSMUST00000226786.2 |
Vmn1r198
|
vomeronasal 1 receptor 198 |
| chr11_-_69706124 | 0.06 |
ENSMUST00000210714.2
|
Gm39566
|
predicted gene, 39566 |
| chr9_+_3025417 | 0.06 |
ENSMUST00000075573.7
|
Gm10717
|
predicted gene 10717 |
| chr9_+_56858162 | 0.06 |
ENSMUST00000068856.5
|
Snupn
|
snurportin 1 |
| chr2_+_5956251 | 0.06 |
ENSMUST00000060092.13
|
Upf2
|
UPF2 regulator of nonsense transcripts homolog (yeast) |
Network of associatons between targets according to the STRING database.
First level regulatory network of Hmga2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 6.6 | GO:0001970 | positive regulation of activation of membrane attack complex(GO:0001970) |
| 0.4 | 1.6 | GO:0045963 | negative regulation of catecholamine metabolic process(GO:0045914) negative regulation of dopamine metabolic process(GO:0045963) |
| 0.3 | 1.0 | GO:0016131 | brassinosteroid metabolic process(GO:0016131) brassinosteroid biosynthetic process(GO:0016132) |
| 0.3 | 6.0 | GO:0031000 | response to caffeine(GO:0031000) |
| 0.3 | 2.6 | GO:0043651 | linoleic acid metabolic process(GO:0043651) |
| 0.2 | 0.6 | GO:0046949 | fatty-acyl-CoA biosynthetic process(GO:0046949) |
| 0.2 | 0.7 | GO:0030974 | thiamine pyrophosphate transport(GO:0030974) |
| 0.2 | 0.6 | GO:1902528 | regulation of protein linear polyubiquitination(GO:1902528) positive regulation of protein linear polyubiquitination(GO:1902530) |
| 0.1 | 0.8 | GO:0033540 | fatty acid beta-oxidation using acyl-CoA oxidase(GO:0033540) |
| 0.1 | 0.4 | GO:0010705 | meiotic DNA double-strand break processing involved in reciprocal meiotic recombination(GO:0010705) |
| 0.1 | 0.3 | GO:0032474 | otolith morphogenesis(GO:0032474) |
| 0.1 | 0.3 | GO:1902527 | positive regulation of protein monoubiquitination(GO:1902527) |
| 0.1 | 0.2 | GO:0090526 | regulation of gluconeogenesis involved in cellular glucose homeostasis(GO:0090526) |
| 0.1 | 0.2 | GO:0070681 | glutaminyl-tRNAGln biosynthesis via transamidation(GO:0070681) |
| 0.1 | 0.2 | GO:0010814 | substance P catabolic process(GO:0010814) calcitonin catabolic process(GO:0010816) endothelin maturation(GO:0034959) |
| 0.1 | 0.3 | GO:0010636 | positive regulation of mitochondrial fusion(GO:0010636) |
| 0.1 | 0.2 | GO:0099526 | presynaptic signal transduction(GO:0098928) presynapse to nucleus signaling pathway(GO:0099526) |
| 0.1 | 0.3 | GO:0006548 | histidine catabolic process(GO:0006548) |
| 0.0 | 1.4 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.0 | 0.4 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.0 | 0.1 | GO:0051659 | maintenance of mitochondrion location(GO:0051659) |
| 0.0 | 0.3 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.0 | 0.1 | GO:0021644 | vagus nerve morphogenesis(GO:0021644) |
| 0.0 | 0.3 | GO:0033184 | positive regulation of histone ubiquitination(GO:0033184) |
| 0.0 | 0.5 | GO:0035092 | sperm chromatin condensation(GO:0035092) |
| 0.0 | 0.1 | GO:1902569 | regulation of activation of JAK2 kinase activity(GO:0010534) activation of JAK2 kinase activity(GO:0042977) negative regulation of activation of JAK2 kinase activity(GO:1902569) |
| 0.0 | 0.4 | GO:0052696 | flavonoid glucuronidation(GO:0052696) xenobiotic glucuronidation(GO:0052697) |
| 0.0 | 0.1 | GO:2001201 | regulation of transforming growth factor-beta secretion(GO:2001201) |
| 0.0 | 0.6 | GO:0033617 | mitochondrial respiratory chain complex IV assembly(GO:0033617) |
| 0.0 | 0.1 | GO:0014063 | negative regulation of serotonin secretion(GO:0014063) |
| 0.0 | 0.4 | GO:0035635 | entry of bacterium into host cell(GO:0035635) regulation of entry of bacterium into host cell(GO:2000535) |
| 0.0 | 0.2 | GO:0099590 | neurotransmitter receptor internalization(GO:0099590) |
| 0.0 | 0.2 | GO:0071231 | neural crest cell migration involved in heart formation(GO:0003147) anterior neural tube closure(GO:0061713) cellular response to folic acid(GO:0071231) |
| 0.0 | 0.2 | GO:0016103 | diterpenoid catabolic process(GO:0016103) retinoic acid catabolic process(GO:0034653) |
| 0.0 | 3.9 | GO:0007586 | digestion(GO:0007586) |
| 0.0 | 0.1 | GO:0032290 | peripheral nervous system myelin formation(GO:0032290) |
| 0.0 | 0.3 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.0 | 0.1 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.0 | 0.1 | GO:0072318 | clathrin coat disassembly(GO:0072318) |
| 0.0 | 0.1 | GO:0051030 | RNA import into nucleus(GO:0006404) snRNA transport(GO:0051030) |
| 0.0 | 0.2 | GO:0044539 | long-chain fatty acid import(GO:0044539) |
| 0.0 | 0.0 | GO:1900135 | positive regulation of renin secretion into blood stream(GO:1900135) |
| 0.0 | 0.7 | GO:0090077 | foam cell differentiation(GO:0090077) |
| 0.0 | 0.4 | GO:0016558 | protein import into peroxisome matrix(GO:0016558) |
| 0.0 | 0.7 | GO:0019369 | arachidonic acid metabolic process(GO:0019369) |
| 0.0 | 0.1 | GO:0036343 | psychomotor behavior(GO:0036343) positive regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900738) |
| 0.0 | 0.2 | GO:0031937 | positive regulation of chromatin silencing(GO:0031937) |
| 0.0 | 0.2 | GO:0036159 | inner dynein arm assembly(GO:0036159) |
| 0.0 | 0.4 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.0 | 0.1 | GO:0097240 | meiotic telomere tethering at nuclear periphery(GO:0044821) meiotic attachment of telomere to nuclear envelope(GO:0070197) chromosome attachment to the nuclear envelope(GO:0097240) |
| 0.0 | 0.1 | GO:0001766 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 0.0 | 0.1 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.0 | 0.2 | GO:0070932 | histone H3 deacetylation(GO:0070932) |
| 0.0 | 0.3 | GO:0006693 | prostanoid metabolic process(GO:0006692) prostaglandin metabolic process(GO:0006693) |
| 0.0 | 0.4 | GO:0051443 | positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
| 0.0 | 0.1 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.0 | 0.1 | GO:1990592 | protein polyufmylation(GO:1990564) protein K69-linked ufmylation(GO:1990592) |
| 0.0 | 1.0 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.0 | 0.1 | GO:0070417 | cellular response to cold(GO:0070417) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 6.6 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.1 | 0.4 | GO:1990429 | Pex17p-Pex14p docking complex(GO:1990415) peroxisomal importomer complex(GO:1990429) |
| 0.1 | 0.2 | GO:0030956 | glutamyl-tRNA(Gln) amidotransferase complex(GO:0030956) |
| 0.1 | 0.3 | GO:0070469 | respiratory chain(GO:0070469) |
| 0.1 | 0.2 | GO:0044317 | rod spherule(GO:0044317) |
| 0.1 | 0.3 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.0 | 0.2 | GO:1990769 | proximal neuron projection(GO:1990769) |
| 0.0 | 1.0 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.0 | 0.3 | GO:0089701 | U2AF(GO:0089701) |
| 0.0 | 0.4 | GO:0000801 | central element(GO:0000801) |
| 0.0 | 0.4 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.0 | 0.3 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.0 | 0.1 | GO:0000814 | ESCRT II complex(GO:0000814) |
| 0.0 | 0.8 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 0.0 | GO:0001405 | presequence translocase-associated import motor(GO:0001405) |
| 0.0 | 0.3 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.0 | 0.2 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.0 | 0.1 | GO:0005879 | axonemal microtubule(GO:0005879) |
| 0.0 | 0.2 | GO:0036156 | inner dynein arm(GO:0036156) |
| 0.0 | 0.1 | GO:0032585 | multivesicular body membrane(GO:0032585) |
| 0.0 | 0.7 | GO:0031304 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 0.9 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.3 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 0.5 | GO:0042588 | zymogen granule(GO:0042588) |
| 0.0 | 0.1 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.0 | 6.0 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.6 | GO:0071614 | linoleic acid epoxygenase activity(GO:0071614) |
| 0.5 | 1.6 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 0.4 | 1.2 | GO:0016509 | long-chain-3-hydroxyacyl-CoA dehydrogenase activity(GO:0016509) |
| 0.3 | 1.0 | GO:0047598 | sterol delta7 reductase activity(GO:0009918) 7-dehydrocholesterol reductase activity(GO:0047598) |
| 0.3 | 10.4 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.1 | 0.3 | GO:0016262 | protein N-acetylglucosaminyltransferase activity(GO:0016262) |
| 0.1 | 0.7 | GO:0015234 | thiamine transmembrane transporter activity(GO:0015234) |
| 0.1 | 0.2 | GO:0050567 | glutaminyl-tRNA synthase (glutamine-hydrolyzing) activity(GO:0050567) |
| 0.1 | 16.3 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 0.4 | GO:0032450 | maltose alpha-glucosidase activity(GO:0032450) |
| 0.1 | 0.8 | GO:0003997 | acyl-CoA oxidase activity(GO:0003997) |
| 0.1 | 0.3 | GO:0045322 | unmethylated CpG binding(GO:0045322) |
| 0.1 | 0.2 | GO:0047936 | glucose 1-dehydrogenase [NAD(P)] activity(GO:0047936) |
| 0.0 | 0.4 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.0 | 1.4 | GO:0008392 | arachidonic acid epoxygenase activity(GO:0008392) |
| 0.0 | 0.2 | GO:0043758 | acetate-CoA ligase (ADP-forming) activity(GO:0043758) |
| 0.0 | 1.0 | GO:0015035 | protein disulfide oxidoreductase activity(GO:0015035) |
| 0.0 | 0.6 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.0 | 0.4 | GO:0097027 | ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.0 | 0.1 | GO:0070540 | stearic acid binding(GO:0070540) |
| 0.0 | 0.1 | GO:0005093 | Rab GDP-dissociation inhibitor activity(GO:0005093) |
| 0.0 | 0.3 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 0.0 | 0.5 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.0 | 0.7 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) aromatase activity(GO:0070330) |
| 0.0 | 0.3 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 0.0 | 0.1 | GO:0016784 | 3-mercaptopyruvate sulfurtransferase activity(GO:0016784) |
| 0.0 | 0.2 | GO:0008401 | retinoic acid 4-hydroxylase activity(GO:0008401) |
| 0.0 | 0.1 | GO:0047275 | glucosaminylgalactosylglucosylceramide beta-galactosyltransferase activity(GO:0047275) |
| 0.0 | 0.8 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.3 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.0 | 0.1 | GO:0015119 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.0 | 0.2 | GO:0051870 | methotrexate binding(GO:0051870) |
| 0.0 | 0.4 | GO:0004551 | nucleotide diphosphatase activity(GO:0004551) |
| 0.0 | 0.4 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.0 | 0.1 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.0 | 0.2 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 0.7 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.0 | 0.3 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 7.2 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.3 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.0 | 0.6 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 0.1 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.0 | 0.1 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 6.4 | REACTOME DEFENSINS | Genes involved in Defensins |
| 0.2 | 7.0 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.1 | 0.4 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.0 | 1.4 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 1.2 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 1.0 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.8 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.0 | 1.6 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.0 | 0.1 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.6 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |