Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
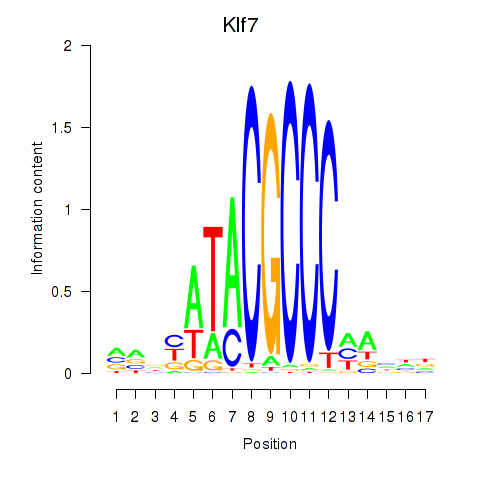
Results for Klf7
Z-value: 0.89
Transcription factors associated with Klf7
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Klf7
|
ENSMUSG00000025959.14 | Kruppel-like factor 7 (ubiquitous) |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Klf7 | mm39_v1_chr1_-_64160557_64160650 | 0.38 | 2.4e-02 | Click! |
Activity profile of Klf7 motif
Sorted Z-values of Klf7 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chrX_+_135171002 | 10.18 |
ENSMUST00000178632.8
ENSMUST00000053540.11 |
Bex3
|
brain expressed X-linked 3 |
| chrX_+_135171049 | 9.28 |
ENSMUST00000113112.2
|
Bex3
|
brain expressed X-linked 3 |
| chr8_+_85628557 | 8.10 |
ENSMUST00000067060.10
ENSMUST00000239392.2 |
Klf1
|
Kruppel-like factor 1 (erythroid) |
| chr5_+_33815466 | 6.03 |
ENSMUST00000074849.13
ENSMUST00000079534.11 ENSMUST00000201633.2 |
Tacc3
|
transforming, acidic coiled-coil containing protein 3 |
| chr4_+_140737955 | 5.98 |
ENSMUST00000071977.9
|
Mfap2
|
microfibrillar-associated protein 2 |
| chr11_-_101998648 | 4.51 |
ENSMUST00000177304.8
ENSMUST00000017455.15 |
Pyy
|
peptide YY |
| chr5_-_136164840 | 3.94 |
ENSMUST00000150406.2
ENSMUST00000006301.11 |
Lrwd1
|
leucine-rich repeats and WD repeat domain containing 1 |
| chr3_+_51131868 | 3.76 |
ENSMUST00000023849.15
ENSMUST00000167780.2 |
Noct
|
nocturnin |
| chr7_-_127593003 | 3.49 |
ENSMUST00000033056.5
|
Pycard
|
PYD and CARD domain containing |
| chr19_-_4109446 | 3.49 |
ENSMUST00000189808.7
|
Gstp3
|
glutathione S-transferase pi 3 |
| chr11_-_61384998 | 3.47 |
ENSMUST00000101085.9
ENSMUST00000079080.13 ENSMUST00000108714.2 |
Mapk7
|
mitogen-activated protein kinase 7 |
| chr15_+_98972850 | 3.45 |
ENSMUST00000039665.8
|
Troap
|
trophinin associated protein |
| chr7_-_132378584 | 3.44 |
ENSMUST00000106168.2
|
Fam53b
|
family with sequence similarity 53, member B |
| chr17_-_24470356 | 3.07 |
ENSMUST00000115390.5
|
Ccnf
|
cyclin F |
| chr15_-_98851423 | 3.03 |
ENSMUST00000134214.3
|
Gm49450
|
predicted gene, 49450 |
| chr1_+_63216281 | 2.67 |
ENSMUST00000188524.2
|
Eef1b2
|
eukaryotic translation elongation factor 1 beta 2 |
| chr11_+_63023893 | 2.62 |
ENSMUST00000108700.2
|
Pmp22
|
peripheral myelin protein 22 |
| chr15_-_98851566 | 2.60 |
ENSMUST00000097014.7
|
Tuba1a
|
tubulin, alpha 1A |
| chr3_-_103698391 | 2.58 |
ENSMUST00000106845.9
ENSMUST00000029438.15 |
Hipk1
|
homeodomain interacting protein kinase 1 |
| chr1_+_63215976 | 2.39 |
ENSMUST00000129339.8
|
Eef1b2
|
eukaryotic translation elongation factor 1 beta 2 |
| chr9_-_45847344 | 2.18 |
ENSMUST00000034590.4
|
Tagln
|
transgelin |
| chr4_+_63477018 | 2.08 |
ENSMUST00000077709.11
|
Tmem268
|
transmembrane protein 268 |
| chr10_+_127512933 | 2.06 |
ENSMUST00000118612.8
ENSMUST00000048099.5 |
Nemp1
|
nuclear envelope integral membrane protein 1 |
| chr8_-_85413707 | 1.96 |
ENSMUST00000238301.2
|
Nacc1
|
nucleus accumbens associated 1, BEN and BTB (POZ) domain containing |
| chr17_+_46513666 | 1.85 |
ENSMUST00000087031.7
|
Xpo5
|
exportin 5 |
| chr11_-_40624200 | 1.76 |
ENSMUST00000020579.9
|
Hmmr
|
hyaluronan mediated motility receptor (RHAMM) |
| chr17_-_46513499 | 1.73 |
ENSMUST00000024749.9
|
Polh
|
polymerase (DNA directed), eta (RAD 30 related) |
| chr2_-_151815307 | 1.67 |
ENSMUST00000109863.2
|
Fam110a
|
family with sequence similarity 110, member A |
| chr5_+_115769960 | 1.65 |
ENSMUST00000031492.15
|
Rab35
|
RAB35, member RAS oncogene family |
| chr15_-_34443054 | 1.62 |
ENSMUST00000142643.2
|
Rpl30
|
ribosomal protein L30 |
| chr7_-_44578834 | 1.62 |
ENSMUST00000107857.11
ENSMUST00000167930.8 ENSMUST00000085399.13 ENSMUST00000166972.9 |
Ap2a1
|
adaptor-related protein complex 2, alpha 1 subunit |
| chr5_+_136164990 | 1.57 |
ENSMUST00000136634.8
ENSMUST00000041100.4 |
Alkbh4
|
alkB homolog 4, lysine demethylase |
| chr8_-_3671270 | 1.43 |
ENSMUST00000159548.2
ENSMUST00000019614.13 |
Xab2
|
XPA binding protein 2 |
| chr1_+_92759324 | 1.43 |
ENSMUST00000045970.8
|
Gpc1
|
glypican 1 |
| chr7_+_25380195 | 1.39 |
ENSMUST00000205658.2
|
B9d2
|
B9 protein domain 2 |
| chr12_-_79343040 | 1.34 |
ENSMUST00000218377.2
ENSMUST00000021547.8 |
Zfyve26
|
zinc finger, FYVE domain containing 26 |
| chr14_+_30201569 | 1.33 |
ENSMUST00000022535.9
ENSMUST00000223658.2 |
Dcp1a
|
decapping mRNA 1A |
| chr15_-_90563510 | 1.29 |
ENSMUST00000014777.9
ENSMUST00000064391.12 |
Cpne8
|
copine VIII |
| chr8_-_85692634 | 1.20 |
ENSMUST00000109738.10
ENSMUST00000065049.15 ENSMUST00000128972.9 |
Rnaseh2a
|
ribonuclease H2, large subunit |
| chr4_-_156077061 | 1.14 |
ENSMUST00000052185.5
|
B3galt6
|
UDP-Gal:betaGal beta 1,3-galactosyltransferase, polypeptide 6 |
| chr7_+_25380263 | 1.14 |
ENSMUST00000205325.2
ENSMUST00000206913.2 |
B9d2
|
B9 protein domain 2 |
| chr8_+_106587268 | 1.08 |
ENSMUST00000212610.2
ENSMUST00000212484.2 ENSMUST00000212200.2 |
Nutf2
|
nuclear transport factor 2 |
| chr8_+_106587212 | 1.04 |
ENSMUST00000008594.9
|
Nutf2
|
nuclear transport factor 2 |
| chr17_+_36227869 | 0.94 |
ENSMUST00000151664.8
|
Ppp1r10
|
protein phosphatase 1, regulatory subunit 10 |
| chr6_+_83996344 | 0.94 |
ENSMUST00000204987.3
ENSMUST00000203695.3 ENSMUST00000204354.3 ENSMUST00000089595.12 ENSMUST00000113818.8 ENSMUST00000081904.7 |
Dysf
|
dysferlin |
| chr5_-_124325213 | 0.91 |
ENSMUST00000161938.8
|
Pitpnm2
|
phosphatidylinositol transfer protein, membrane-associated 2 |
| chr9_-_32255604 | 0.89 |
ENSMUST00000034533.7
|
Kcnj5
|
potassium inwardly-rectifying channel, subfamily J, member 5 |
| chr16_+_4756963 | 0.85 |
ENSMUST00000229961.2
ENSMUST00000050881.10 |
Nudt16l1
|
nudix (nucleoside diphosphate linked moiety X)-type motif 16-like 1 |
| chr9_-_36678868 | 0.81 |
ENSMUST00000217599.2
ENSMUST00000120381.9 |
Stt3a
|
STT3, subunit of the oligosaccharyltransferase complex, homolog A (S. cerevisiae) |
| chr11_-_101193106 | 0.78 |
ENSMUST00000130916.8
|
Becn1
|
beclin 1, autophagy related |
| chr15_-_10485976 | 0.76 |
ENSMUST00000169050.8
ENSMUST00000022855.12 |
Brix1
|
BRX1, biogenesis of ribosomes |
| chr7_-_68398917 | 0.75 |
ENSMUST00000118110.3
|
Arrdc4
|
arrestin domain containing 4 |
| chr9_-_32255556 | 0.73 |
ENSMUST00000214223.2
|
Kcnj5
|
potassium inwardly-rectifying channel, subfamily J, member 5 |
| chr11_-_78067446 | 0.72 |
ENSMUST00000148154.3
ENSMUST00000017549.13 |
Nek8
|
NIMA (never in mitosis gene a)-related expressed kinase 8 |
| chr2_-_156154667 | 0.68 |
ENSMUST00000079125.8
|
Scand1
|
SCAN domain-containing 1 |
| chr11_+_80274105 | 0.67 |
ENSMUST00000165565.8
ENSMUST00000188489.7 ENSMUST00000017567.14 ENSMUST00000108216.8 ENSMUST00000053740.15 |
Zfp207
|
zinc finger protein 207 |
| chr9_-_32255533 | 0.66 |
ENSMUST00000216033.2
|
Kcnj5
|
potassium inwardly-rectifying channel, subfamily J, member 5 |
| chr11_-_115977755 | 0.65 |
ENSMUST00000074628.13
ENSMUST00000106444.4 |
Wbp2
|
WW domain binding protein 2 |
| chr11_-_101193032 | 0.61 |
ENSMUST00000140706.8
ENSMUST00000170502.2 ENSMUST00000172233.8 |
Becn1
|
beclin 1, autophagy related |
| chr18_+_35252470 | 0.54 |
ENSMUST00000237154.2
|
Ctnna1
|
catenin (cadherin associated protein), alpha 1 |
| chr18_+_11790409 | 0.51 |
ENSMUST00000047322.8
|
Rbbp8
|
retinoblastoma binding protein 8, endonuclease |
| chr11_+_65698001 | 0.51 |
ENSMUST00000071465.9
ENSMUST00000018491.8 |
Zkscan6
|
zinc finger with KRAB and SCAN domains 6 |
| chr2_+_181138958 | 0.49 |
ENSMUST00000149163.8
ENSMUST00000000844.15 ENSMUST00000184849.8 ENSMUST00000108800.8 ENSMUST00000069712.9 |
Tpd52l2
|
tumor protein D52-like 2 |
| chr2_+_181139016 | 0.48 |
ENSMUST00000108799.10
|
Tpd52l2
|
tumor protein D52-like 2 |
| chr15_-_102630496 | 0.43 |
ENSMUST00000171838.2
|
Calcoco1
|
calcium binding and coiled coil domain 1 |
| chr4_-_132056725 | 0.42 |
ENSMUST00000127402.8
ENSMUST00000105962.10 ENSMUST00000030730.14 ENSMUST00000105960.3 |
Trnau1ap
|
tRNA selenocysteine 1 associated protein 1 |
| chr1_+_120048890 | 0.41 |
ENSMUST00000027637.13
ENSMUST00000112644.9 ENSMUST00000056038.15 |
3110009E18Rik
|
RIKEN cDNA 3110009E18 gene |
| chr15_+_98532866 | 0.40 |
ENSMUST00000230490.2
|
Cacnb3
|
calcium channel, voltage-dependent, beta 3 subunit |
| chr9_-_71393175 | 0.38 |
ENSMUST00000233263.2
ENSMUST00000034720.12 |
Polr2m
|
polymerase (RNA) II (DNA directed) polypeptide M |
| chr18_+_37818263 | 0.36 |
ENSMUST00000194418.2
|
Pcdhga4
|
protocadherin gamma subfamily A, 4 |
| chr7_-_100661181 | 0.36 |
ENSMUST00000178340.3
ENSMUST00000037540.5 |
P2ry2
|
purinergic receptor P2Y, G-protein coupled 2 |
| chr18_+_88989914 | 0.36 |
ENSMUST00000023828.9
|
Rttn
|
rotatin |
| chr15_-_102630589 | 0.35 |
ENSMUST00000023818.11
|
Calcoco1
|
calcium binding and coiled coil domain 1 |
| chr7_-_68398989 | 0.34 |
ENSMUST00000048068.15
|
Arrdc4
|
arrestin domain containing 4 |
| chr7_-_100661220 | 0.31 |
ENSMUST00000207916.2
|
P2ry2
|
purinergic receptor P2Y, G-protein coupled 2 |
| chr17_+_36227433 | 0.30 |
ENSMUST00000087211.9
|
Ppp1r10
|
protein phosphatase 1, regulatory subunit 10 |
| chr15_+_85620308 | 0.29 |
ENSMUST00000057979.6
|
Ppara
|
peroxisome proliferator activated receptor alpha |
| chr1_-_58484528 | 0.27 |
ENSMUST00000114345.9
ENSMUST00000117069.8 ENSMUST00000114348.8 ENSMUST00000190048.7 ENSMUST00000185990.2 ENSMUST00000081677.12 |
Ppil3
|
peptidylprolyl isomerase (cyclophilin)-like 3 |
| chr6_-_121107953 | 0.25 |
ENSMUST00000204248.2
|
Mical3
|
microtubule associated monooxygenase, calponin and LIM domain containing 3 |
| chr5_+_136165055 | 0.25 |
ENSMUST00000143229.2
|
Alkbh4
|
alkB homolog 4, lysine demethylase |
| chr11_-_60618022 | 0.24 |
ENSMUST00000002889.5
|
Flii
|
flightless I actin binding protein |
| chr9_-_114762879 | 0.23 |
ENSMUST00000084853.4
|
Gpd1l
|
glycerol-3-phosphate dehydrogenase 1-like |
| chr11_+_68322945 | 0.21 |
ENSMUST00000021283.8
|
Pik3r5
|
phosphoinositide-3-kinase regulatory subunit 5 |
| chr11_-_78427061 | 0.21 |
ENSMUST00000017759.9
ENSMUST00000108277.3 |
Tnfaip1
|
tumor necrosis factor, alpha-induced protein 1 (endothelial) |
| chr17_+_64203017 | 0.18 |
ENSMUST00000000129.14
|
Fer
|
fer (fms/fps related) protein kinase |
| chr5_+_45677571 | 0.14 |
ENSMUST00000156481.8
ENSMUST00000119579.3 ENSMUST00000118833.3 |
Med28
|
mediator complex subunit 28 |
| chr7_-_115797722 | 0.09 |
ENSMUST00000181981.8
|
Plekha7
|
pleckstrin homology domain containing, family A member 7 |
| chr14_+_79689230 | 0.09 |
ENSMUST00000100359.3
ENSMUST00000226192.2 |
Kbtbd6
|
kelch repeat and BTB (POZ) domain containing 6 |
| chr7_-_109215960 | 0.08 |
ENSMUST00000077909.9
|
Denn2b
|
DENN domain containing 2B |
| chr8_+_70605404 | 0.03 |
ENSMUST00000110146.9
ENSMUST00000110143.8 ENSMUST00000110141.9 ENSMUST00000110140.2 |
Mef2b
|
myocyte enhancer factor 2B |
| chr1_+_171156568 | 0.01 |
ENSMUST00000111300.8
|
Dedd
|
death effector domain-containing |
Network of associatons between targets according to the STRING database.
First level regulatory network of Klf7
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 3.5 | GO:0002277 | myeloid dendritic cell activation involved in immune response(GO:0002277) positive regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002582) positive regulation of antigen processing and presentation of peptide antigen via MHC class II(GO:0002588) |
| 1.2 | 3.5 | GO:0070376 | regulation of ERK5 cascade(GO:0070376) |
| 0.6 | 1.8 | GO:0036090 | cleavage furrow ingression(GO:0036090) |
| 0.5 | 1.6 | GO:1900126 | negative regulation of hyaluronan biosynthetic process(GO:1900126) |
| 0.5 | 2.1 | GO:1904046 | negative regulation of vascular endothelial growth factor production(GO:1904046) |
| 0.5 | 1.4 | GO:0000349 | generation of catalytic spliceosome for first transesterification step(GO:0000349) |
| 0.5 | 1.4 | GO:0030200 | heparan sulfate proteoglycan catabolic process(GO:0030200) |
| 0.5 | 1.8 | GO:1900368 | regulation of RNA interference(GO:1900368) |
| 0.5 | 3.2 | GO:0071169 | establishment of protein localization to chromatin(GO:0071169) |
| 0.4 | 3.1 | GO:0000320 | re-entry into mitotic cell cycle(GO:0000320) |
| 0.4 | 8.1 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.4 | 5.1 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.3 | 0.8 | GO:0043686 | co-translational protein modification(GO:0043686) |
| 0.3 | 6.0 | GO:0022027 | interkinetic nuclear migration(GO:0022027) astral microtubule organization(GO:0030953) |
| 0.2 | 4.5 | GO:0032096 | negative regulation of response to food(GO:0032096) |
| 0.2 | 2.3 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.2 | 1.7 | GO:0006290 | pyrimidine dimer repair(GO:0006290) |
| 0.2 | 1.7 | GO:0048227 | plasma membrane to endosome transport(GO:0048227) |
| 0.2 | 1.2 | GO:0043137 | DNA replication, removal of RNA primer(GO:0043137) |
| 0.2 | 2.6 | GO:0061072 | iris morphogenesis(GO:0061072) |
| 0.1 | 1.4 | GO:0043652 | engulfment of apoptotic cell(GO:0043652) |
| 0.1 | 18.9 | GO:0008625 | extrinsic apoptotic signaling pathway via death domain receptors(GO:0008625) |
| 0.1 | 6.0 | GO:0030220 | platelet formation(GO:0030220) |
| 0.1 | 0.8 | GO:2001032 | regulation of double-strand break repair via nonhomologous end joining(GO:2001032) |
| 0.1 | 2.6 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.1 | 5.4 | GO:0006414 | translational elongation(GO:0006414) |
| 0.1 | 0.7 | GO:0071442 | positive regulation of histone H3-K14 acetylation(GO:0071442) |
| 0.1 | 0.2 | GO:0046168 | glycerol-3-phosphate catabolic process(GO:0046168) |
| 0.1 | 0.4 | GO:0051643 | endoplasmic reticulum localization(GO:0051643) |
| 0.1 | 3.4 | GO:0035411 | catenin import into nucleus(GO:0035411) |
| 0.1 | 0.7 | GO:0060406 | positive regulation of penile erection(GO:0060406) |
| 0.1 | 0.5 | GO:0010792 | DNA double-strand break processing involved in repair via single-strand annealing(GO:0010792) |
| 0.1 | 0.9 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.1 | 0.7 | GO:0035330 | regulation of hippo signaling(GO:0035330) |
| 0.0 | 1.6 | GO:0031640 | killing of cells of other organism(GO:0031640) disruption of cells of other organism(GO:0044364) |
| 0.0 | 1.1 | GO:0051443 | positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
| 0.0 | 0.2 | GO:0036215 | response to stem cell factor(GO:0036215) cellular response to stem cell factor stimulus(GO:0036216) Fc-epsilon receptor signaling pathway(GO:0038095) Kit signaling pathway(GO:0038109) |
| 0.0 | 0.4 | GO:0061577 | calcium ion transmembrane transport via high voltage-gated calcium channel(GO:0061577) regulation of membrane repolarization during action potential(GO:0098903) |
| 0.0 | 0.8 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 1.1 | GO:0006024 | glycosaminoglycan biosynthetic process(GO:0006024) |
| 0.0 | 0.7 | GO:0007094 | mitotic spindle assembly checkpoint(GO:0007094) |
| 0.0 | 0.2 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.0 | 0.1 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 3.5 | GO:0097169 | AIM2 inflammasome complex(GO:0097169) |
| 0.7 | 5.1 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.6 | 1.8 | GO:0031074 | nucleocytoplasmic shuttling complex(GO:0031074) nuclear RNA export factor complex(GO:0042272) |
| 0.4 | 3.9 | GO:0031933 | telomeric heterochromatin(GO:0031933) |
| 0.4 | 6.0 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.3 | 1.4 | GO:0034271 | phosphatidylinositol 3-kinase complex, class III, type I(GO:0034271) phosphatidylinositol 3-kinase complex, class III, type II(GO:0034272) |
| 0.2 | 1.2 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.2 | 2.1 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.1 | 2.5 | GO:0036038 | MKS complex(GO:0036038) |
| 0.1 | 1.2 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.1 | 1.4 | GO:0071014 | post-mRNA release spliceosomal complex(GO:0071014) |
| 0.1 | 0.8 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.1 | 1.8 | GO:0070938 | contractile ring(GO:0070938) |
| 0.1 | 1.6 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.0 | 0.2 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.0 | 6.0 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 0.2 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 0.0 | 1.1 | GO:0005797 | Golgi medial cisterna(GO:0005797) |
| 0.0 | 1.7 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.0 | 5.1 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 3.1 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 2.6 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 0.4 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 3.2 | GO:0030315 | T-tubule(GO:0030315) |
| 0.0 | 1.6 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 0.6 | GO:0043218 | compact myelin(GO:0043218) |
| 0.0 | 8.3 | GO:0000790 | nuclear chromatin(GO:0000790) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 6.0 | GO:0070051 | fibrinogen binding(GO:0070051) |
| 0.6 | 19.5 | GO:0005123 | death receptor binding(GO:0005123) |
| 0.5 | 1.4 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.3 | 3.5 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.3 | 2.3 | GO:0086089 | voltage-gated potassium channel activity involved in atrial cardiac muscle cell action potential repolarization(GO:0086089) |
| 0.3 | 1.8 | GO:0070883 | pre-miRNA binding(GO:0070883) |
| 0.2 | 0.7 | GO:0045030 | UTP-activated nucleotide receptor activity(GO:0045030) |
| 0.2 | 5.1 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.2 | 1.2 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.2 | 3.9 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.2 | 3.8 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.1 | 1.6 | GO:0035368 | selenocysteine insertion sequence binding(GO:0035368) |
| 0.1 | 4.5 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.1 | 3.5 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.1 | 1.1 | GO:0008499 | UDP-galactose:beta-N-acetylglucosamine beta-1,3-galactosyltransferase activity(GO:0008499) |
| 0.1 | 0.8 | GO:0004576 | oligosaccharyl transferase activity(GO:0004576) dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.1 | 1.8 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 2.1 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.1 | 1.7 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 0.2 | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity(GO:0004367) |
| 0.0 | 0.8 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
| 0.0 | 2.5 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.8 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 5.6 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.0 | 0.5 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) |
| 0.0 | 1.4 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 1.3 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.0 | 0.2 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 1.4 | GO:0043548 | phosphatidylinositol 3-kinase binding(GO:0043548) |
| 0.0 | 1.7 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 2.6 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 1.3 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 1.2 | GO:0016706 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, 2-oxoglutarate as one donor, and incorporation of one atom each of oxygen into both donors(GO:0016706) |
| 0.0 | 0.4 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.0 | 0.9 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 1.6 | GO:0008565 | protein transporter activity(GO:0008565) |
| 0.0 | 0.7 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 6.0 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 3.5 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.1 | 2.3 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.1 | 1.6 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.1 | 6.5 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 2.6 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.0 | 2.6 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.0 | 0.8 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 1.4 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.0 | 2.8 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.0 | 2.4 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.5 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.2 | 3.5 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.1 | 1.8 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.1 | 2.6 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.1 | 1.6 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.1 | 1.4 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.1 | 1.4 | REACTOME FORMATION OF TRANSCRIPTION COUPLED NER TC NER REPAIR COMPLEX | Genes involved in Formation of transcription-coupled NER (TC-NER) repair complex |
| 0.1 | 1.8 | REACTOME MICRORNA MIRNA BIOGENESIS | Genes involved in MicroRNA (miRNA) Biogenesis |
| 0.1 | 1.3 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.0 | 2.3 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.0 | 0.7 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 1.1 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.0 | 6.7 | REACTOME TRANSLATION | Genes involved in Translation |
| 0.0 | 4.5 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 2.5 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 0.4 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.0 | 0.5 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.0 | 1.7 | REACTOME DNA REPAIR | Genes involved in DNA Repair |
| 0.0 | 0.2 | REACTOME G BETA GAMMA SIGNALLING THROUGH PI3KGAMMA | Genes involved in G beta:gamma signalling through PI3Kgamma |