Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
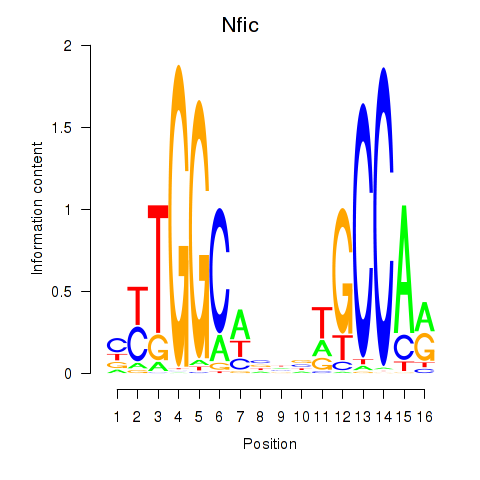
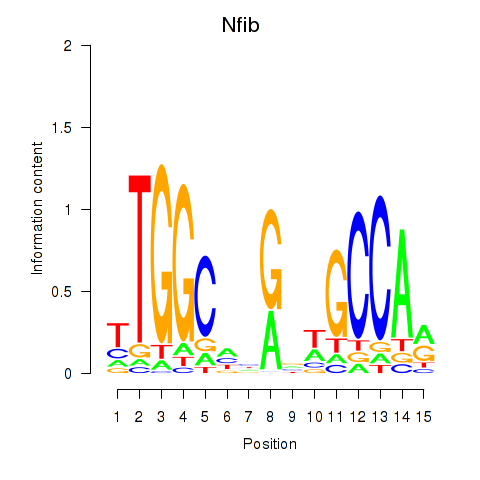
Results for Nfic_Nfib
Z-value: 1.52
Transcription factors associated with Nfic_Nfib
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Nfic
|
ENSMUSG00000055053.19 | nuclear factor I/C |
|
Nfib
|
ENSMUSG00000008575.18 | nuclear factor I/B |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Nfib | mm39_v1_chr4_-_82623972_82623997 | 0.39 | 1.8e-02 | Click! |
| Nfic | mm39_v1_chr10_-_81243475_81243515 | 0.28 | 9.7e-02 | Click! |
Activity profile of Nfic_Nfib motif
Sorted Z-values of Nfic_Nfib motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_-_109678685 | 11.01 |
ENSMUST00000112022.5
|
Camp
|
cathelicidin antimicrobial peptide |
| chrX_+_135171002 | 9.98 |
ENSMUST00000178632.8
ENSMUST00000053540.11 |
Bex3
|
brain expressed X-linked 3 |
| chr5_-_145742672 | 9.41 |
ENSMUST00000067479.6
|
Cyp3a44
|
cytochrome P450, family 3, subfamily a, polypeptide 44 |
| chrX_+_135171049 | 8.62 |
ENSMUST00000113112.2
|
Bex3
|
brain expressed X-linked 3 |
| chr5_-_145521533 | 8.38 |
ENSMUST00000075837.8
|
Cyp3a41b
|
cytochrome P450, family 3, subfamily a, polypeptide 41B |
| chr8_+_95710977 | 8.33 |
ENSMUST00000093271.8
|
Adgrg1
|
adhesion G protein-coupled receptor G1 |
| chr3_-_107129038 | 7.25 |
ENSMUST00000029504.9
|
Cym
|
chymosin |
| chr11_+_61376257 | 6.84 |
ENSMUST00000064783.10
ENSMUST00000040522.7 |
Mfap4
|
microfibrillar-associated protein 4 |
| chr7_+_46700349 | 6.78 |
ENSMUST00000010451.8
|
Tmem86a
|
transmembrane protein 86A |
| chr3_+_90576285 | 6.59 |
ENSMUST00000069927.10
|
S100a8
|
S100 calcium binding protein A8 (calgranulin A) |
| chrX_-_134968985 | 6.41 |
ENSMUST00000049130.8
|
Bex2
|
brain expressed X-linked 2 |
| chr19_+_4644425 | 5.84 |
ENSMUST00000238089.2
|
Pcx
|
pyruvate carboxylase |
| chr2_+_13579092 | 5.46 |
ENSMUST00000193675.2
|
Vim
|
vimentin |
| chr2_+_13578738 | 5.43 |
ENSMUST00000141365.3
ENSMUST00000028062.8 |
Vim
|
vimentin |
| chr17_+_32755525 | 5.30 |
ENSMUST00000169591.8
ENSMUST00000003416.15 |
Cyp4f16
|
cytochrome P450, family 4, subfamily f, polypeptide 16 |
| chrX_-_51702790 | 4.83 |
ENSMUST00000069360.14
|
Gpc3
|
glypican 3 |
| chr1_+_172328768 | 4.64 |
ENSMUST00000111228.2
|
Tagln2
|
transgelin 2 |
| chr11_+_115353290 | 4.62 |
ENSMUST00000106532.4
ENSMUST00000092445.12 ENSMUST00000153466.2 |
Slc16a5
|
solute carrier family 16 (monocarboxylic acid transporters), member 5 |
| chr14_+_30856687 | 4.46 |
ENSMUST00000090212.5
|
Nt5dc2
|
5'-nucleotidase domain containing 2 |
| chr17_-_28736483 | 4.40 |
ENSMUST00000114792.8
ENSMUST00000177939.8 |
Fkbp5
|
FK506 binding protein 5 |
| chr3_-_104725853 | 4.28 |
ENSMUST00000106775.8
ENSMUST00000166979.8 |
Mov10
|
Mov10 RISC complex RNA helicase |
| chr5_-_35683035 | 4.27 |
ENSMUST00000038676.7
|
Cpz
|
carboxypeptidase Z |
| chr6_+_78402956 | 4.26 |
ENSMUST00000079926.6
|
Reg1
|
regenerating islet-derived 1 |
| chr3_-_104725581 | 4.07 |
ENSMUST00000168015.8
|
Mov10
|
Mov10 RISC complex RNA helicase |
| chr19_+_4644365 | 4.06 |
ENSMUST00000113825.4
|
Pcx
|
pyruvate carboxylase |
| chr11_+_109376432 | 3.98 |
ENSMUST00000106697.8
|
Arsg
|
arylsulfatase G |
| chr3_-_104725535 | 3.97 |
ENSMUST00000002297.12
|
Mov10
|
Mov10 RISC complex RNA helicase |
| chrX_-_51702813 | 3.95 |
ENSMUST00000114857.2
|
Gpc3
|
glypican 3 |
| chr5_-_134776101 | 3.84 |
ENSMUST00000015138.13
|
Eln
|
elastin |
| chr15_-_103275190 | 3.80 |
ENSMUST00000023128.8
|
Itga5
|
integrin alpha 5 (fibronectin receptor alpha) |
| chr8_+_95703728 | 3.77 |
ENSMUST00000179619.9
|
Adgrg1
|
adhesion G protein-coupled receptor G1 |
| chr8_-_124621483 | 3.76 |
ENSMUST00000034453.6
ENSMUST00000212584.2 |
Acta1
|
actin, alpha 1, skeletal muscle |
| chr2_+_164611812 | 3.75 |
ENSMUST00000088248.13
ENSMUST00000001439.7 |
Ube2c
|
ubiquitin-conjugating enzyme E2C |
| chr11_+_3939924 | 3.74 |
ENSMUST00000109981.2
|
Gal3st1
|
galactose-3-O-sulfotransferase 1 |
| chr18_+_37939442 | 3.71 |
ENSMUST00000076807.7
|
Pcdhgc3
|
protocadherin gamma subfamily C, 3 |
| chr5_+_130398261 | 3.63 |
ENSMUST00000086029.10
|
Caln1
|
calneuron 1 |
| chr2_+_172994841 | 3.62 |
ENSMUST00000029017.6
|
Pck1
|
phosphoenolpyruvate carboxykinase 1, cytosolic |
| chr17_+_43671314 | 3.41 |
ENSMUST00000226087.2
|
Adgrf5
|
adhesion G protein-coupled receptor F5 |
| chr6_+_135339929 | 3.41 |
ENSMUST00000032330.16
|
Emp1
|
epithelial membrane protein 1 |
| chr6_+_135339543 | 3.35 |
ENSMUST00000205156.3
|
Emp1
|
epithelial membrane protein 1 |
| chr7_-_110581652 | 3.15 |
ENSMUST00000005751.13
|
Irag1
|
inositol 1,4,5-triphosphate receptor associated 1 |
| chr9_+_44045859 | 3.12 |
ENSMUST00000034650.15
ENSMUST00000098852.3 ENSMUST00000216002.2 |
Mcam
|
melanoma cell adhesion molecule |
| chr1_-_88205185 | 3.09 |
ENSMUST00000147393.2
|
Hjurp
|
Holliday junction recognition protein |
| chr5_+_115682138 | 3.00 |
ENSMUST00000202564.2
|
Pxn
|
paxillin |
| chrX_+_149829131 | 2.93 |
ENSMUST00000112685.8
|
Fgd1
|
FYVE, RhoGEF and PH domain containing 1 |
| chr1_+_40305738 | 2.92 |
ENSMUST00000114795.3
|
Il1r1
|
interleukin 1 receptor, type I |
| chr5_-_122188165 | 2.88 |
ENSMUST00000154139.8
|
Cux2
|
cut-like homeobox 2 |
| chr1_+_74414354 | 2.86 |
ENSMUST00000187516.7
ENSMUST00000027368.6 |
Slc11a1
|
solute carrier family 11 (proton-coupled divalent metal ion transporters), member 1 |
| chrX_-_106446928 | 2.84 |
ENSMUST00000033591.6
|
Itm2a
|
integral membrane protein 2A |
| chr1_-_88205233 | 2.81 |
ENSMUST00000065420.12
ENSMUST00000054674.15 |
Hjurp
|
Holliday junction recognition protein |
| chrX_-_149595873 | 2.74 |
ENSMUST00000131241.2
ENSMUST00000147152.3 ENSMUST00000143843.8 |
Maged2
|
MAGE family member D2 |
| chrX_-_149596680 | 2.67 |
ENSMUST00000112700.8
|
Maged2
|
MAGE family member D2 |
| chr9_+_59563872 | 2.64 |
ENSMUST00000215623.2
ENSMUST00000215660.2 ENSMUST00000217353.2 |
Pkm
|
pyruvate kinase, muscle |
| chr15_+_81686622 | 2.63 |
ENSMUST00000109553.10
|
Tef
|
thyrotroph embryonic factor |
| chr11_-_120930193 | 2.59 |
ENSMUST00000026159.6
|
Cd7
|
CD7 antigen |
| chr6_-_136918495 | 2.54 |
ENSMUST00000111892.8
|
Arhgdib
|
Rho, GDP dissociation inhibitor (GDI) beta |
| chr9_-_71070506 | 2.53 |
ENSMUST00000074465.9
|
Aqp9
|
aquaporin 9 |
| chr9_+_59563838 | 2.49 |
ENSMUST00000163694.4
|
Pkm
|
pyruvate kinase, muscle |
| chr11_+_96941420 | 2.49 |
ENSMUST00000090020.13
|
Osbpl7
|
oxysterol binding protein-like 7 |
| chr9_-_53521585 | 2.47 |
ENSMUST00000034547.6
|
Acat1
|
acetyl-Coenzyme A acetyltransferase 1 |
| chr9_+_69361348 | 2.44 |
ENSMUST00000134907.8
|
Anxa2
|
annexin A2 |
| chr9_+_69360902 | 2.38 |
ENSMUST00000034756.15
ENSMUST00000123470.8 |
Anxa2
|
annexin A2 |
| chr4_+_129941633 | 2.38 |
ENSMUST00000044565.15
ENSMUST00000132251.2 |
Col16a1
|
collagen, type XVI, alpha 1 |
| chr1_+_92900834 | 2.37 |
ENSMUST00000186298.7
ENSMUST00000027489.9 |
Gpr35
|
G protein-coupled receptor 35 |
| chr7_+_19144950 | 2.33 |
ENSMUST00000208710.2
ENSMUST00000003643.3 |
Ckm
|
creatine kinase, muscle |
| chr3_+_88523440 | 2.32 |
ENSMUST00000177498.8
ENSMUST00000176500.8 |
Arhgef2
|
rho/rac guanine nucleotide exchange factor (GEF) 2 |
| chr6_-_136918671 | 2.31 |
ENSMUST00000032344.12
|
Arhgdib
|
Rho, GDP dissociation inhibitor (GDI) beta |
| chr11_+_96941637 | 2.29 |
ENSMUST00000168565.2
|
Osbpl7
|
oxysterol binding protein-like 7 |
| chr14_+_71179308 | 2.29 |
ENSMUST00000227633.2
|
Gfra2
|
glial cell line derived neurotrophic factor family receptor alpha 2 |
| chr11_+_58808716 | 2.27 |
ENSMUST00000069941.13
|
Btnl10
|
butyrophilin-like 10 |
| chr3_-_115923098 | 2.23 |
ENSMUST00000196449.5
|
Vcam1
|
vascular cell adhesion molecule 1 |
| chr18_-_33597060 | 2.23 |
ENSMUST00000168890.2
|
Nrep
|
neuronal regeneration related protein |
| chr11_+_67167950 | 2.23 |
ENSMUST00000019625.12
|
Myh8
|
myosin, heavy polypeptide 8, skeletal muscle, perinatal |
| chr18_-_33596890 | 2.22 |
ENSMUST00000237066.2
|
Nrep
|
neuronal regeneration related protein |
| chr10_-_105410280 | 2.17 |
ENSMUST00000061506.9
|
Tmtc2
|
transmembrane and tetratricopeptide repeat containing 2 |
| chr15_+_34306812 | 2.17 |
ENSMUST00000226766.2
ENSMUST00000163455.9 ENSMUST00000022947.7 ENSMUST00000228570.2 ENSMUST00000227759.2 |
Matn2
|
matrilin 2 |
| chr6_+_17306334 | 2.16 |
ENSMUST00000007799.13
ENSMUST00000115456.6 |
Cav1
|
caveolin 1, caveolae protein |
| chr11_+_115044966 | 2.15 |
ENSMUST00000021076.6
|
Rab37
|
RAB37, member RAS oncogene family |
| chr19_-_42741148 | 2.03 |
ENSMUST00000236659.2
ENSMUST00000076505.4 |
Pyroxd2
|
pyridine nucleotide-disulphide oxidoreductase domain 2 |
| chr2_+_38229270 | 2.01 |
ENSMUST00000143783.9
|
Lhx2
|
LIM homeobox protein 2 |
| chr6_-_83633064 | 2.00 |
ENSMUST00000014686.3
|
Clec4f
|
C-type lectin domain family 4, member f |
| chr1_-_162687369 | 1.99 |
ENSMUST00000193078.6
|
Fmo1
|
flavin containing monooxygenase 1 |
| chr16_+_34605282 | 1.97 |
ENSMUST00000023538.9
|
Mylk
|
myosin, light polypeptide kinase |
| chr7_+_100143250 | 1.96 |
ENSMUST00000153287.8
|
Ucp2
|
uncoupling protein 2 (mitochondrial, proton carrier) |
| chr18_-_33596468 | 1.96 |
ENSMUST00000171533.9
|
Nrep
|
neuronal regeneration related protein |
| chr11_+_104441489 | 1.95 |
ENSMUST00000018800.9
|
Myl4
|
myosin, light polypeptide 4 |
| chr4_-_63321591 | 1.94 |
ENSMUST00000035724.5
|
Akna
|
AT-hook transcription factor |
| chr19_-_5974825 | 1.93 |
ENSMUST00000055458.6
|
Cdc42ep2
|
CDC42 effector protein (Rho GTPase binding) 2 |
| chr8_+_95722289 | 1.93 |
ENSMUST00000211984.2
|
Adgrg1
|
adhesion G protein-coupled receptor G1 |
| chr4_+_137408975 | 1.93 |
ENSMUST00000047243.12
|
Rap1gap
|
Rap1 GTPase-activating protein |
| chr9_-_56068282 | 1.92 |
ENSMUST00000034876.10
|
Tspan3
|
tetraspanin 3 |
| chr17_+_47908025 | 1.89 |
ENSMUST00000183206.2
|
Ccnd3
|
cyclin D3 |
| chr18_+_35252470 | 1.89 |
ENSMUST00000237154.2
|
Ctnna1
|
catenin (cadherin associated protein), alpha 1 |
| chr10_+_81012465 | 1.88 |
ENSMUST00000047864.11
|
Eef2
|
eukaryotic translation elongation factor 2 |
| chr1_-_162687254 | 1.87 |
ENSMUST00000131058.8
|
Fmo1
|
flavin containing monooxygenase 1 |
| chr7_+_100142977 | 1.85 |
ENSMUST00000129324.8
|
Ucp2
|
uncoupling protein 2 (mitochondrial, proton carrier) |
| chr3_-_131065658 | 1.82 |
ENSMUST00000029610.9
|
Hadh
|
hydroxyacyl-Coenzyme A dehydrogenase |
| chr5_-_135107078 | 1.81 |
ENSMUST00000067935.11
ENSMUST00000076203.3 |
Vps37d
|
vacuolar protein sorting 37D |
| chr7_+_44866095 | 1.81 |
ENSMUST00000209437.2
|
Tead2
|
TEA domain family member 2 |
| chr4_-_132260799 | 1.79 |
ENSMUST00000152993.8
ENSMUST00000067496.7 |
Atpif1
|
ATPase inhibitory factor 1 |
| chr1_-_90771674 | 1.78 |
ENSMUST00000097653.11
ENSMUST00000188587.7 ENSMUST00000056925.16 ENSMUST00000187753.2 |
Col6a3
|
collagen, type VI, alpha 3 |
| chr19_+_6135013 | 1.76 |
ENSMUST00000025704.3
|
Cdca5
|
cell division cycle associated 5 |
| chr11_-_3489228 | 1.76 |
ENSMUST00000075118.10
ENSMUST00000136243.2 ENSMUST00000020721.15 ENSMUST00000170588.8 |
Smtn
|
smoothelin |
| chr18_-_78166595 | 1.76 |
ENSMUST00000091813.12
|
Slc14a1
|
solute carrier family 14 (urea transporter), member 1 |
| chr7_-_4755971 | 1.72 |
ENSMUST00000183971.8
ENSMUST00000182173.2 ENSMUST00000182738.8 ENSMUST00000182111.8 ENSMUST00000184143.8 ENSMUST00000182048.2 ENSMUST00000063324.14 |
Cox6b2
|
cytochrome c oxidase subunit 6B2 |
| chr12_-_79030250 | 1.72 |
ENSMUST00000070174.14
|
Tmem229b
|
transmembrane protein 229B |
| chr18_-_36648850 | 1.70 |
ENSMUST00000025363.7
|
Hbegf
|
heparin-binding EGF-like growth factor |
| chr18_-_33596792 | 1.69 |
ENSMUST00000051087.16
|
Nrep
|
neuronal regeneration related protein |
| chr13_+_43938251 | 1.69 |
ENSMUST00000015540.4
|
Cd83
|
CD83 antigen |
| chr6_+_72074718 | 1.68 |
ENSMUST00000187007.3
|
St3gal5
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 5 |
| chr1_-_87501548 | 1.68 |
ENSMUST00000068681.12
|
Ngef
|
neuronal guanine nucleotide exchange factor |
| chr1_-_72914036 | 1.68 |
ENSMUST00000027377.9
|
Igfbp5
|
insulin-like growth factor binding protein 5 |
| chr18_-_78166539 | 1.65 |
ENSMUST00000160292.8
|
Slc14a1
|
solute carrier family 14 (urea transporter), member 1 |
| chr11_-_116001037 | 1.64 |
ENSMUST00000106441.8
ENSMUST00000021120.6 |
Trim47
|
tripartite motif-containing 47 |
| chr17_+_29538157 | 1.63 |
ENSMUST00000114699.9
ENSMUST00000155348.3 ENSMUST00000234618.2 |
Pi16
|
peptidase inhibitor 16 |
| chr3_-_94566107 | 1.62 |
ENSMUST00000196655.5
ENSMUST00000200407.2 ENSMUST00000006123.11 ENSMUST00000196733.5 |
Tuft1
|
tuftelin 1 |
| chr16_-_17394495 | 1.58 |
ENSMUST00000231645.2
ENSMUST00000232226.2 ENSMUST00000232336.2 ENSMUST00000232385.2 ENSMUST00000231615.2 ENSMUST00000231283.2 ENSMUST00000172164.10 |
Slc7a4
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 4 |
| chr9_+_46151994 | 1.57 |
ENSMUST00000034585.7
|
Apoa4
|
apolipoprotein A-IV |
| chr9_-_56151334 | 1.56 |
ENSMUST00000188142.7
|
Peak1
|
pseudopodium-enriched atypical kinase 1 |
| chr12_+_3941728 | 1.56 |
ENSMUST00000172689.8
ENSMUST00000111186.8 |
Dnmt3a
|
DNA methyltransferase 3A |
| chr3_-_10273628 | 1.56 |
ENSMUST00000029041.6
|
Fabp4
|
fatty acid binding protein 4, adipocyte |
| chr2_+_36120438 | 1.54 |
ENSMUST00000062069.6
|
Ptgs1
|
prostaglandin-endoperoxide synthase 1 |
| chr11_+_115768323 | 1.54 |
ENSMUST00000222123.2
|
Myo15b
|
myosin XVB |
| chr3_-_121325887 | 1.53 |
ENSMUST00000039197.9
|
Slc44a3
|
solute carrier family 44, member 3 |
| chr8_-_65146079 | 1.52 |
ENSMUST00000048967.9
|
Cpe
|
carboxypeptidase E |
| chr13_+_55517545 | 1.52 |
ENSMUST00000063771.14
|
Rgs14
|
regulator of G-protein signaling 14 |
| chr6_+_72074545 | 1.52 |
ENSMUST00000069994.11
ENSMUST00000114112.4 |
St3gal5
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 5 |
| chr7_+_130633776 | 1.48 |
ENSMUST00000084509.7
ENSMUST00000213064.3 ENSMUST00000208311.4 |
Dmbt1
|
deleted in malignant brain tumors 1 |
| chr7_-_126275529 | 1.46 |
ENSMUST00000106372.11
ENSMUST00000155419.3 ENSMUST00000106373.9 |
Sult1a1
|
sulfotransferase family 1A, phenol-preferring, member 1 |
| chr8_+_95703506 | 1.44 |
ENSMUST00000212581.2
|
Adgrg1
|
adhesion G protein-coupled receptor G1 |
| chr3_-_144466602 | 1.44 |
ENSMUST00000059091.6
|
Clca3a1
|
chloride channel accessory 3A1 |
| chr7_-_118594365 | 1.43 |
ENSMUST00000008878.10
|
Gprc5b
|
G protein-coupled receptor, family C, group 5, member B |
| chr2_+_84891281 | 1.40 |
ENSMUST00000238769.2
|
Tnks1bp1
|
tankyrase 1 binding protein 1 |
| chr11_+_58808830 | 1.39 |
ENSMUST00000020792.12
ENSMUST00000108818.4 |
Btnl10
|
butyrophilin-like 10 |
| chr4_+_43957677 | 1.37 |
ENSMUST00000107855.2
|
Glipr2
|
GLI pathogenesis-related 2 |
| chr11_-_76359222 | 1.35 |
ENSMUST00000065028.14
|
Abr
|
active BCR-related gene |
| chr1_+_40264211 | 1.34 |
ENSMUST00000027241.11
|
Il1r1
|
interleukin 1 receptor, type I |
| chr5_+_135754568 | 1.34 |
ENSMUST00000127096.2
|
Por
|
P450 (cytochrome) oxidoreductase |
| chr7_+_78563964 | 1.34 |
ENSMUST00000120331.4
|
Isg20
|
interferon-stimulated protein |
| chr2_-_32314017 | 1.33 |
ENSMUST00000113307.9
|
Slc25a25
|
solute carrier family 25 (mitochondrial carrier, phosphate carrier), member 25 |
| chr8_-_95602952 | 1.33 |
ENSMUST00000046461.9
|
Dok4
|
docking protein 4 |
| chr9_+_88209250 | 1.33 |
ENSMUST00000034992.8
|
Nt5e
|
5' nucleotidase, ecto |
| chr7_-_105131407 | 1.31 |
ENSMUST00000047040.4
|
Cavin3
|
caveolae associated 3 |
| chr15_-_98505508 | 1.29 |
ENSMUST00000096224.6
|
Adcy6
|
adenylate cyclase 6 |
| chr15_+_54609011 | 1.28 |
ENSMUST00000050027.9
|
Ccn3
|
cellular communication network factor 3 |
| chr11_-_109986763 | 1.25 |
ENSMUST00000046223.14
ENSMUST00000106664.10 ENSMUST00000106662.2 |
Abca8a
|
ATP-binding cassette, sub-family A (ABC1), member 8a |
| chr1_-_162687488 | 1.24 |
ENSMUST00000134098.8
ENSMUST00000111518.3 |
Fmo1
|
flavin containing monooxygenase 1 |
| chr4_-_93223746 | 1.23 |
ENSMUST00000066774.6
|
Tusc1
|
tumor suppressor candidate 1 |
| chr10_-_75757393 | 1.22 |
ENSMUST00000121304.2
ENSMUST00000000925.10 |
Smarcb1
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1 |
| chr10_-_62163444 | 1.21 |
ENSMUST00000139228.8
|
Hk1
|
hexokinase 1 |
| chr1_+_51328265 | 1.20 |
ENSMUST00000051572.8
|
Cavin2
|
caveolae associated 2 |
| chr12_-_26506422 | 1.17 |
ENSMUST00000020970.10
|
Rsad2
|
radical S-adenosyl methionine domain containing 2 |
| chr7_-_44174065 | 1.16 |
ENSMUST00000165208.4
|
Mybpc2
|
myosin binding protein C, fast-type |
| chr10_-_76949510 | 1.16 |
ENSMUST00000105409.8
|
Col18a1
|
collagen, type XVIII, alpha 1 |
| chr11_-_69556888 | 1.16 |
ENSMUST00000108654.3
|
Cd68
|
CD68 antigen |
| chr11_-_69556904 | 1.16 |
ENSMUST00000018918.12
|
Cd68
|
CD68 antigen |
| chr7_+_126811831 | 1.15 |
ENSMUST00000127710.3
|
Mylpf
|
myosin light chain, phosphorylatable, fast skeletal muscle |
| chr10_-_76949762 | 1.15 |
ENSMUST00000072755.12
|
Col18a1
|
collagen, type XVIII, alpha 1 |
| chr4_+_94627513 | 1.13 |
ENSMUST00000073939.13
ENSMUST00000102798.8 |
Tek
|
TEK receptor tyrosine kinase |
| chr19_+_6214416 | 1.13 |
ENSMUST00000045042.8
ENSMUST00000237511.2 |
Batf2
|
basic leucine zipper transcription factor, ATF-like 2 |
| chr2_-_93680024 | 1.13 |
ENSMUST00000068513.11
ENSMUST00000041593.15 ENSMUST00000130077.2 |
Accs
|
1-aminocyclopropane-1-carboxylate synthase (non-functional) |
| chr5_-_38637474 | 1.13 |
ENSMUST00000143758.8
ENSMUST00000156272.8 |
Slc2a9
|
solute carrier family 2 (facilitated glucose transporter), member 9 |
| chr2_-_32341408 | 1.11 |
ENSMUST00000028160.15
ENSMUST00000113310.9 |
Slc25a25
|
solute carrier family 25 (mitochondrial carrier, phosphate carrier), member 25 |
| chr17_+_31515163 | 1.11 |
ENSMUST00000235972.2
ENSMUST00000165149.3 ENSMUST00000236251.2 |
Slc37a1
|
solute carrier family 37 (glycerol-3-phosphate transporter), member 1 |
| chr3_+_95136248 | 1.10 |
ENSMUST00000107201.9
|
Cdc42se1
|
CDC42 small effector 1 |
| chr11_-_109986804 | 1.08 |
ENSMUST00000100287.9
|
Abca8a
|
ATP-binding cassette, sub-family A (ABC1), member 8a |
| chr10_-_75768302 | 1.08 |
ENSMUST00000120281.8
ENSMUST00000000924.13 |
Mmp11
|
matrix metallopeptidase 11 |
| chr10_+_75407356 | 1.07 |
ENSMUST00000143226.8
ENSMUST00000124259.8 |
Ggt1
|
gamma-glutamyltransferase 1 |
| chr8_-_106223502 | 1.05 |
ENSMUST00000212303.2
|
Zdhhc1
|
zinc finger, DHHC domain containing 1 |
| chr12_-_112640378 | 1.05 |
ENSMUST00000130342.2
|
Akt1
|
thymoma viral proto-oncogene 1 |
| chr2_-_179867605 | 1.04 |
ENSMUST00000015791.6
|
Lama5
|
laminin, alpha 5 |
| chr5_+_137743992 | 1.01 |
ENSMUST00000100540.10
|
Tsc22d4
|
TSC22 domain family, member 4 |
| chr7_+_79836581 | 1.00 |
ENSMUST00000032754.9
|
Sema4b
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4B |
| chr3_+_79536378 | 1.00 |
ENSMUST00000029388.10
|
4930579G24Rik
|
RIKEN cDNA 4930579G24 gene |
| chr15_-_66703471 | 1.00 |
ENSMUST00000164163.8
|
Sla
|
src-like adaptor |
| chr9_-_21710029 | 0.99 |
ENSMUST00000034717.7
|
Kank2
|
KN motif and ankyrin repeat domains 2 |
| chr11_+_50492899 | 0.98 |
ENSMUST00000142118.3
ENSMUST00000040523.9 |
Adamts2
|
a disintegrin-like and metallopeptidase (reprolysin type) with thrombospondin type 1 motif, 2 |
| chr5_+_137756407 | 0.98 |
ENSMUST00000141733.8
ENSMUST00000110985.2 |
Tsc22d4
|
TSC22 domain family, member 4 |
| chr14_+_62569517 | 0.98 |
ENSMUST00000022499.13
|
Rnaseh2b
|
ribonuclease H2, subunit B |
| chr17_+_29537798 | 0.97 |
ENSMUST00000114701.10
|
Pi16
|
peptidase inhibitor 16 |
| chr1_+_136059101 | 0.96 |
ENSMUST00000075164.11
|
Kif21b
|
kinesin family member 21B |
| chr19_-_46950737 | 0.96 |
ENSMUST00000168536.9
ENSMUST00000236783.2 ENSMUST00000238106.2 |
Nt5c2
|
5'-nucleotidase, cytosolic II |
| chr4_-_57916283 | 0.96 |
ENSMUST00000063816.6
|
D630039A03Rik
|
RIKEN cDNA D630039A03 gene |
| chr2_-_166902307 | 0.96 |
ENSMUST00000155281.8
|
Znfx1
|
zinc finger, NFX1-type containing 1 |
| chr9_+_50663171 | 0.94 |
ENSMUST00000214609.2
|
Cryab
|
crystallin, alpha B |
| chr11_+_94520567 | 0.94 |
ENSMUST00000021239.7
|
Lrrc59
|
leucine rich repeat containing 59 |
| chr1_-_156501860 | 0.92 |
ENSMUST00000188964.7
ENSMUST00000190607.2 ENSMUST00000079625.11 |
Tor3a
|
torsin family 3, member A |
| chr5_-_137530214 | 0.92 |
ENSMUST00000140139.2
|
Gnb2
|
guanine nucleotide binding protein (G protein), beta 2 |
| chrX_+_20529137 | 0.92 |
ENSMUST00000001989.9
|
Uba1
|
ubiquitin-like modifier activating enzyme 1 |
| chr7_+_78564062 | 0.91 |
ENSMUST00000205981.2
|
Isg20
|
interferon-stimulated protein |
| chr19_-_46033353 | 0.90 |
ENSMUST00000026252.14
ENSMUST00000156585.9 ENSMUST00000185355.7 ENSMUST00000152946.8 |
Ldb1
|
LIM domain binding 1 |
| chr5_+_137744228 | 0.87 |
ENSMUST00000100539.10
|
Tsc22d4
|
TSC22 domain family, member 4 |
| chr2_+_32536594 | 0.86 |
ENSMUST00000113272.8
ENSMUST00000009705.14 ENSMUST00000167841.8 |
Eng
|
endoglin |
| chr19_-_46917661 | 0.86 |
ENSMUST00000236727.2
|
Nt5c2
|
5'-nucleotidase, cytosolic II |
| chr2_+_152907875 | 0.86 |
ENSMUST00000238488.2
ENSMUST00000129377.8 ENSMUST00000109800.2 |
Ccm2l
|
cerebral cavernous malformation 2-like |
| chr2_+_11710523 | 0.85 |
ENSMUST00000138856.2
ENSMUST00000078834.12 ENSMUST00000114834.10 ENSMUST00000114833.10 ENSMUST00000114831.9 |
Il15ra
|
interleukin 15 receptor, alpha chain |
| chr11_+_115331365 | 0.85 |
ENSMUST00000093914.5
|
Trim80
|
tripartite motif-containing 80 |
| chr6_+_29433247 | 0.85 |
ENSMUST00000101617.9
ENSMUST00000065090.8 |
Flnc
|
filamin C, gamma |
| chr2_+_84880776 | 0.85 |
ENSMUST00000111605.9
|
Tnks1bp1
|
tankyrase 1 binding protein 1 |
| chr2_+_84966569 | 0.85 |
ENSMUST00000057019.9
|
Aplnr
|
apelin receptor |
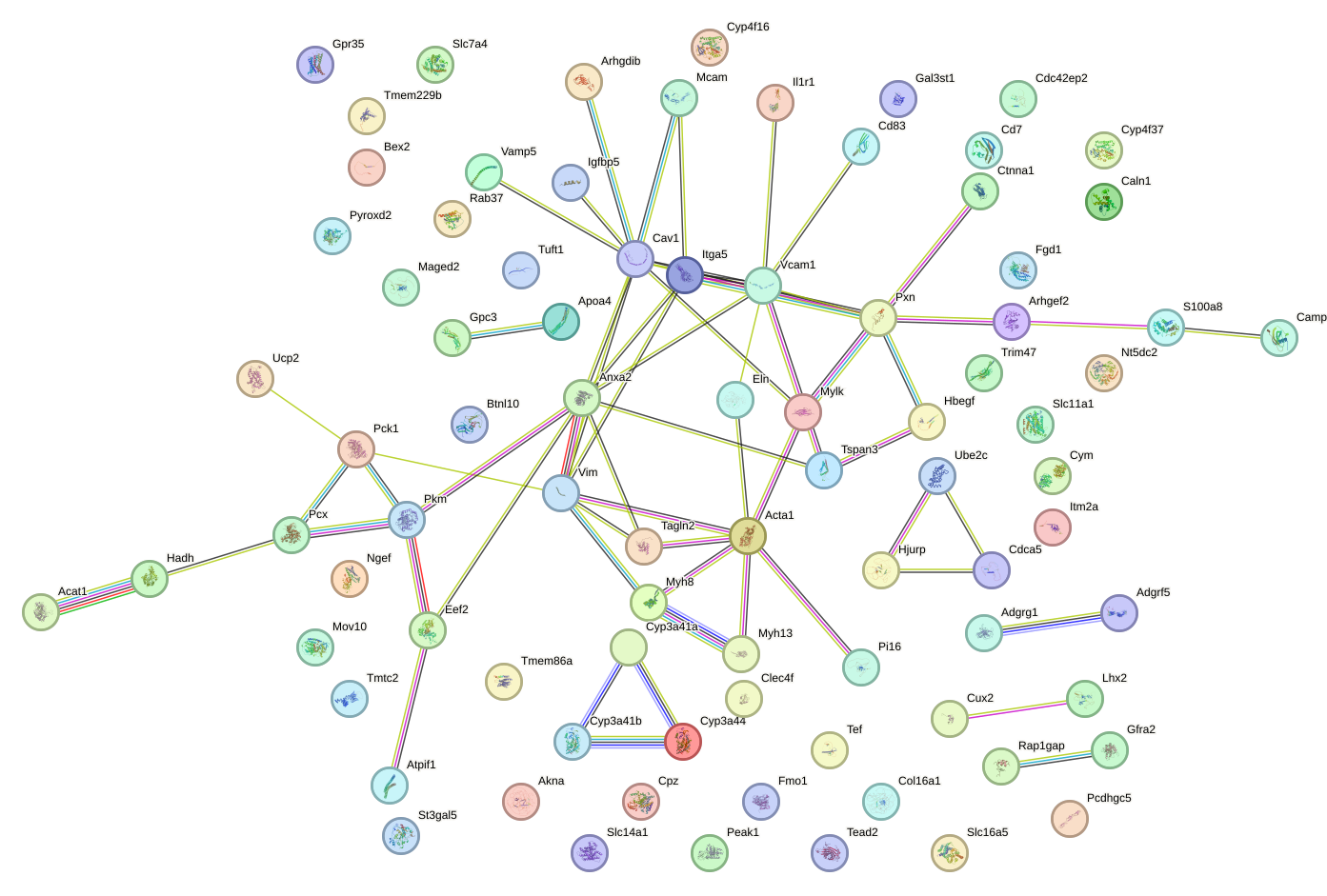
Network of associatons between targets according to the STRING database.
First level regulatory network of Nfic_Nfib
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.1 | 12.3 | GO:0035279 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing(GO:0098795) |
| 2.0 | 9.9 | GO:0019074 | viral genome packaging(GO:0019072) viral RNA genome packaging(GO:0019074) |
| 1.8 | 8.8 | GO:0072138 | mesenchymal cell proliferation involved in ureteric bud development(GO:0072138) |
| 1.7 | 1.7 | GO:0051542 | elastin biosynthetic process(GO:0051542) |
| 1.6 | 6.6 | GO:0070488 | neutrophil aggregation(GO:0070488) |
| 1.4 | 10.9 | GO:1900147 | Schwann cell migration(GO:0036135) regulation of Schwann cell migration(GO:1900147) |
| 1.1 | 4.3 | GO:0010286 | heat acclimation(GO:0010286) |
| 1.0 | 5.9 | GO:0034080 | CENP-A containing nucleosome assembly(GO:0034080) CENP-A containing chromatin organization(GO:0061641) |
| 1.0 | 2.9 | GO:0070839 | divalent metal ion export(GO:0070839) |
| 0.9 | 3.6 | GO:0061402 | positive regulation of transcription from RNA polymerase II promoter in response to acidic pH(GO:0061402) |
| 0.9 | 0.9 | GO:1905072 | cardiac jelly development(GO:1905072) |
| 0.8 | 12.6 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 0.8 | 2.5 | GO:0006550 | isoleucine catabolic process(GO:0006550) |
| 0.8 | 2.4 | GO:1904456 | negative regulation of neuronal action potential(GO:1904456) |
| 0.8 | 5.5 | GO:0031536 | positive regulation of exit from mitosis(GO:0031536) |
| 0.8 | 4.5 | GO:0032804 | negative regulation of low-density lipoprotein particle receptor catabolic process(GO:0032804) |
| 0.7 | 2.2 | GO:0031104 | dendrite regeneration(GO:0031104) |
| 0.6 | 2.5 | GO:0015851 | nucleobase transport(GO:0015851) pyrimidine nucleobase transport(GO:0015855) |
| 0.6 | 5.0 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.6 | 15.5 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.6 | 6.8 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.5 | 4.8 | GO:1901164 | negative regulation of trophoblast cell migration(GO:1901164) |
| 0.5 | 3.2 | GO:0071918 | urea transmembrane transport(GO:0071918) |
| 0.5 | 5.1 | GO:0042866 | pyruvate biosynthetic process(GO:0042866) |
| 0.5 | 1.5 | GO:0030070 | insulin processing(GO:0030070) |
| 0.5 | 1.9 | GO:1904425 | negative regulation of GTP binding(GO:1904425) |
| 0.5 | 3.8 | GO:0030240 | skeletal muscle thin filament assembly(GO:0030240) |
| 0.5 | 1.9 | GO:0060709 | glycogen cell differentiation involved in embryonic placenta development(GO:0060709) |
| 0.4 | 1.3 | GO:1904116 | response to vasopressin(GO:1904116) cellular response to vasopressin(GO:1904117) |
| 0.4 | 1.7 | GO:0014734 | skeletal muscle hypertrophy(GO:0014734) |
| 0.4 | 3.7 | GO:0019375 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.4 | 2.8 | GO:0002317 | plasma cell differentiation(GO:0002317) |
| 0.4 | 2.0 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 0.4 | 3.1 | GO:0071802 | negative regulation of podosome assembly(GO:0071802) |
| 0.4 | 2.2 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.4 | 0.7 | GO:0060279 | positive regulation of ovulation(GO:0060279) |
| 0.4 | 3.3 | GO:0071073 | positive regulation of phospholipid biosynthetic process(GO:0071073) |
| 0.3 | 1.3 | GO:0046210 | nitrate catabolic process(GO:0043602) nitric oxide catabolic process(GO:0046210) cellular organohalogen metabolic process(GO:0090345) cellular organofluorine metabolic process(GO:0090346) |
| 0.3 | 1.3 | GO:0046086 | adenosine biosynthetic process(GO:0046086) |
| 0.3 | 1.3 | GO:1901003 | negative regulation of fermentation(GO:1901003) |
| 0.3 | 2.2 | GO:2000767 | positive regulation of cytoplasmic translation(GO:2000767) |
| 0.3 | 1.9 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.3 | 4.7 | GO:0071397 | cellular response to cholesterol(GO:0071397) |
| 0.3 | 1.2 | GO:1900110 | negative regulation of histone H3-K9 dimethylation(GO:1900110) |
| 0.3 | 0.9 | GO:0043973 | histone H3-K4 acetylation(GO:0043973) |
| 0.3 | 1.8 | GO:1901526 | positive regulation of macromitophagy(GO:1901526) positive regulation of mitophagy in response to mitochondrial depolarization(GO:1904925) |
| 0.3 | 1.2 | GO:0034165 | positive regulation of toll-like receptor 9 signaling pathway(GO:0034165) |
| 0.3 | 0.8 | GO:0014739 | positive regulation of muscle hyperplasia(GO:0014739) |
| 0.3 | 3.8 | GO:0033631 | cell-cell adhesion mediated by integrin(GO:0033631) |
| 0.2 | 3.7 | GO:0061469 | regulation of type B pancreatic cell proliferation(GO:0061469) |
| 0.2 | 1.2 | GO:0072656 | maintenance of protein location in mitochondrion(GO:0072656) |
| 0.2 | 2.5 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.2 | 0.7 | GO:0030200 | heparan sulfate proteoglycan catabolic process(GO:0030200) |
| 0.2 | 2.0 | GO:0021978 | telencephalon regionalization(GO:0021978) |
| 0.2 | 1.1 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.2 | 1.5 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.2 | 4.1 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.2 | 0.8 | GO:0002030 | inhibitory G-protein coupled receptor phosphorylation(GO:0002030) |
| 0.2 | 1.7 | GO:0032713 | negative regulation of interleukin-4 production(GO:0032713) |
| 0.2 | 5.6 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.2 | 0.6 | GO:1903977 | regulation of macrophage colony-stimulating factor signaling pathway(GO:1902226) regulation of response to macrophage colony-stimulating factor(GO:1903969) regulation of cellular response to macrophage colony-stimulating factor stimulus(GO:1903972) positive regulation of glial cell migration(GO:1903977) |
| 0.2 | 0.8 | GO:0051933 | amino acid neurotransmitter reuptake(GO:0051933) glutamate reuptake(GO:0051935) |
| 0.2 | 2.2 | GO:0060710 | chorio-allantoic fusion(GO:0060710) |
| 0.2 | 0.9 | GO:1904715 | negative regulation of chaperone-mediated autophagy(GO:1904715) |
| 0.2 | 0.5 | GO:2001178 | mediator complex assembly(GO:0036034) regulation of mediator complex assembly(GO:2001176) positive regulation of mediator complex assembly(GO:2001178) |
| 0.2 | 0.9 | GO:0090271 | positive regulation of fibroblast growth factor production(GO:0090271) |
| 0.2 | 3.3 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.2 | 0.5 | GO:0019417 | sulfur oxidation(GO:0019417) |
| 0.2 | 0.8 | GO:0048050 | post-embryonic eye morphogenesis(GO:0048050) post-embryonic camera-type eye morphogenesis(GO:0048597) |
| 0.2 | 0.6 | GO:0055048 | regulation of spindle elongation(GO:0032887) regulation of mitotic spindle elongation(GO:0032888) anastral spindle assembly(GO:0055048) protein localization to spindle pole body(GO:0071988) regulation of protein localization to spindle pole body(GO:1902363) positive regulation of protein localization to spindle pole body(GO:1902365) positive regulation of mitotic spindle elongation(GO:1902846) |
| 0.2 | 0.5 | GO:2000820 | negative regulation of transcription from RNA polymerase II promoter involved in smooth muscle cell differentiation(GO:2000820) |
| 0.1 | 4.4 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.1 | 2.6 | GO:0061052 | negative regulation of cell growth involved in cardiac muscle cell development(GO:0061052) |
| 0.1 | 1.1 | GO:0060313 | negative regulation of neutrophil degranulation(GO:0043314) negative regulation of blood vessel remodeling(GO:0060313) |
| 0.1 | 0.7 | GO:0006566 | threonine metabolic process(GO:0006566) |
| 0.1 | 1.6 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.1 | 1.6 | GO:0044027 | hypermethylation of CpG island(GO:0044027) |
| 0.1 | 0.6 | GO:2000588 | positive regulation of platelet-derived growth factor receptor-beta signaling pathway(GO:2000588) |
| 0.1 | 0.6 | GO:0044339 | canonical Wnt signaling pathway involved in osteoblast differentiation(GO:0044339) |
| 0.1 | 18.4 | GO:0008625 | extrinsic apoptotic signaling pathway via death domain receptors(GO:0008625) |
| 0.1 | 2.3 | GO:2000353 | positive regulation of endothelial cell apoptotic process(GO:2000353) |
| 0.1 | 1.1 | GO:1901748 | peptide modification(GO:0031179) leukotriene D4 metabolic process(GO:1901748) leukotriene D4 biosynthetic process(GO:1901750) |
| 0.1 | 0.4 | GO:0015783 | GDP-fucose transport(GO:0015783) purine nucleotide-sugar transport(GO:0036079) pyrimidine nucleotide-sugar transmembrane transport(GO:0090481) |
| 0.1 | 0.5 | GO:0042245 | tRNA wobble cytosine modification(GO:0002101) RNA repair(GO:0042245) |
| 0.1 | 1.9 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.1 | 0.5 | GO:0060376 | positive regulation of mast cell differentiation(GO:0060376) |
| 0.1 | 2.5 | GO:0030049 | muscle filament sliding(GO:0030049) |
| 0.1 | 0.4 | GO:0031269 | pseudopodium assembly(GO:0031269) |
| 0.1 | 0.4 | GO:1900369 | negative regulation of RNA interference(GO:1900369) |
| 0.1 | 2.5 | GO:0046415 | urate metabolic process(GO:0046415) |
| 0.1 | 2.0 | GO:0051132 | NK T cell activation(GO:0051132) |
| 0.1 | 1.4 | GO:0060907 | positive regulation of macrophage cytokine production(GO:0060907) |
| 0.1 | 2.8 | GO:0071378 | growth hormone receptor signaling pathway(GO:0060396) cellular response to growth hormone stimulus(GO:0071378) |
| 0.1 | 1.5 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 0.1 | 1.2 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 0.1 | 1.6 | GO:0010226 | response to lithium ion(GO:0010226) cellular response to lithium ion(GO:0071285) |
| 0.1 | 0.3 | GO:1904020 | regulation of G-protein coupled receptor internalization(GO:1904020) |
| 0.1 | 2.4 | GO:0031953 | negative regulation of protein autophosphorylation(GO:0031953) |
| 0.1 | 2.6 | GO:0007614 | short-term memory(GO:0007614) |
| 0.1 | 3.9 | GO:0014823 | response to activity(GO:0014823) |
| 0.1 | 0.3 | GO:0002476 | antigen processing and presentation of endogenous peptide antigen via MHC class Ib(GO:0002476) antigen processing and presentation of exogenous peptide antigen via MHC class Ib(GO:0002477) antigen processing and presentation of exogenous protein antigen via MHC class Ib, TAP-dependent(GO:0002481) cytosol to ER transport(GO:0046967) |
| 0.1 | 0.6 | GO:0034154 | toll-like receptor 7 signaling pathway(GO:0034154) |
| 0.1 | 0.1 | GO:0061341 | non-canonical Wnt signaling pathway involved in heart development(GO:0061341) planar cell polarity pathway involved in heart morphogenesis(GO:0061346) |
| 0.1 | 0.3 | GO:0070563 | negative regulation of vitamin D receptor signaling pathway(GO:0070563) |
| 0.1 | 1.4 | GO:0061002 | negative regulation of dendritic spine morphogenesis(GO:0061002) |
| 0.1 | 0.8 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.1 | 1.4 | GO:0046085 | adenosine metabolic process(GO:0046085) |
| 0.1 | 1.1 | GO:0034116 | positive regulation of heterotypic cell-cell adhesion(GO:0034116) |
| 0.1 | 0.5 | GO:0002023 | reduction of food intake in response to dietary excess(GO:0002023) positive regulation of somatostatin secretion(GO:0090274) |
| 0.1 | 2.8 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.1 | 0.2 | GO:0002426 | immunoglobulin production in mucosal tissue(GO:0002426) |
| 0.1 | 2.1 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.1 | 1.8 | GO:0048368 | lateral mesoderm development(GO:0048368) |
| 0.1 | 0.3 | GO:1904706 | negative regulation of vascular smooth muscle cell proliferation(GO:1904706) |
| 0.1 | 0.7 | GO:0006072 | glycerol-3-phosphate metabolic process(GO:0006072) |
| 0.1 | 0.3 | GO:0046351 | disaccharide biosynthetic process(GO:0046351) |
| 0.1 | 0.6 | GO:0036506 | maintenance of unfolded protein(GO:0036506) maintenance of unfolded protein involved in ERAD pathway(GO:1904378) |
| 0.1 | 0.3 | GO:0072566 | chemokine (C-X-C motif) ligand 1 production(GO:0072566) regulation of chemokine (C-X-C motif) ligand 1 production(GO:2000338) |
| 0.1 | 0.4 | GO:0097309 | cap1 mRNA methylation(GO:0097309) |
| 0.1 | 2.2 | GO:0031954 | positive regulation of protein autophosphorylation(GO:0031954) |
| 0.1 | 0.2 | GO:0051799 | negative regulation of hair follicle development(GO:0051799) |
| 0.1 | 0.2 | GO:0098749 | cerebellar neuron development(GO:0098749) |
| 0.1 | 0.6 | GO:1905146 | lysosomal protein catabolic process(GO:1905146) |
| 0.1 | 2.2 | GO:0051894 | positive regulation of focal adhesion assembly(GO:0051894) |
| 0.1 | 1.5 | GO:0051923 | sulfation(GO:0051923) |
| 0.1 | 0.2 | GO:0003245 | cardiac muscle tissue growth involved in heart morphogenesis(GO:0003245) |
| 0.1 | 1.4 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.1 | 1.3 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 7.7 | GO:0017015 | regulation of transforming growth factor beta receptor signaling pathway(GO:0017015) |
| 0.1 | 0.2 | GO:0072365 | regulation of cellular ketone metabolic process by negative regulation of transcription from RNA polymerase II promoter(GO:0072365) |
| 0.1 | 3.6 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.1 | 1.5 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.1 | 0.9 | GO:0032825 | positive regulation of natural killer cell differentiation(GO:0032825) |
| 0.1 | 0.3 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.1 | 0.1 | GO:0061198 | fungiform papilla formation(GO:0061198) |
| 0.1 | 0.2 | GO:0007308 | oocyte construction(GO:0007308) oocyte axis specification(GO:0007309) oocyte anterior/posterior axis specification(GO:0007314) pole plasm assembly(GO:0007315) maternal determination of anterior/posterior axis, embryo(GO:0008358) P granule organization(GO:0030719) |
| 0.0 | 2.1 | GO:0048873 | homeostasis of number of cells within a tissue(GO:0048873) |
| 0.0 | 0.8 | GO:0097066 | response to thyroid hormone(GO:0097066) |
| 0.0 | 1.1 | GO:0043011 | myeloid dendritic cell differentiation(GO:0043011) |
| 0.0 | 0.4 | GO:1903003 | positive regulation of protein deubiquitination(GO:1903003) |
| 0.0 | 0.3 | GO:1900454 | positive regulation of long term synaptic depression(GO:1900454) |
| 0.0 | 3.3 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.1 | GO:0044704 | mating plug formation(GO:0042628) single-organism reproductive behavior(GO:0044704) post-mating behavior(GO:0045297) seminal vesicle development(GO:0061107) |
| 0.0 | 0.2 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.0 | 1.5 | GO:0010718 | positive regulation of epithelial to mesenchymal transition(GO:0010718) |
| 0.0 | 1.8 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.0 | 0.7 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.0 | 0.7 | GO:0046185 | aldehyde catabolic process(GO:0046185) |
| 0.0 | 2.0 | GO:0032781 | positive regulation of ATPase activity(GO:0032781) |
| 0.0 | 0.4 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 0.0 | 0.2 | GO:0070458 | detoxification of nitrogen compound(GO:0051410) cellular detoxification of nitrogen compound(GO:0070458) |
| 0.0 | 0.9 | GO:0060445 | branching involved in salivary gland morphogenesis(GO:0060445) |
| 0.0 | 0.3 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.0 | 3.4 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.0 | 0.7 | GO:0016242 | negative regulation of macroautophagy(GO:0016242) |
| 0.0 | 3.5 | GO:0030038 | contractile actin filament bundle assembly(GO:0030038) stress fiber assembly(GO:0043149) |
| 0.0 | 0.4 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.0 | 1.0 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.0 | 0.1 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.0 | 1.4 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.0 | 0.3 | GO:2001199 | regulation of dendritic cell differentiation(GO:2001198) negative regulation of dendritic cell differentiation(GO:2001199) |
| 0.0 | 0.5 | GO:0032211 | negative regulation of telomere maintenance via telomerase(GO:0032211) |
| 0.0 | 0.2 | GO:0097475 | motor neuron migration(GO:0097475) |
| 0.0 | 0.2 | GO:0001574 | ganglioside biosynthetic process(GO:0001574) |
| 0.0 | 0.4 | GO:0051451 | myoblast migration(GO:0051451) |
| 0.0 | 0.5 | GO:0000338 | protein deneddylation(GO:0000338) |
| 0.0 | 0.8 | GO:0097352 | autophagosome maturation(GO:0097352) |
| 0.0 | 0.4 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.0 | 1.1 | GO:0046329 | negative regulation of JNK cascade(GO:0046329) |
| 0.0 | 0.1 | GO:0021648 | vestibulocochlear nerve morphogenesis(GO:0021648) vestibulocochlear nerve formation(GO:0021650) |
| 0.0 | 0.1 | GO:1904714 | regulation of chaperone-mediated autophagy(GO:1904714) |
| 0.0 | 1.1 | GO:0051438 | regulation of ubiquitin-protein transferase activity(GO:0051438) |
| 0.0 | 0.3 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.0 | 1.2 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 0.5 | GO:0010591 | regulation of lamellipodium assembly(GO:0010591) |
| 0.0 | 0.1 | GO:0003195 | mitral valve formation(GO:0003192) tricuspid valve formation(GO:0003195) right ventricular cardiac muscle tissue morphogenesis(GO:0003221) |
| 0.0 | 0.2 | GO:0002483 | antigen processing and presentation of endogenous peptide antigen(GO:0002483) antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.0 | 0.6 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.0 | 0.2 | GO:0032099 | negative regulation of appetite(GO:0032099) |
| 0.0 | 0.2 | GO:0019321 | pentose metabolic process(GO:0019321) |
| 0.0 | 0.8 | GO:0048747 | muscle fiber development(GO:0048747) |
| 0.0 | 0.1 | GO:0044351 | macropinocytosis(GO:0044351) |
| 0.0 | 0.2 | GO:0071243 | cellular response to arsenic-containing substance(GO:0071243) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 10.7 | GO:0071953 | elastic fiber(GO:0071953) |
| 1.7 | 5.1 | GO:1902912 | pyruvate kinase complex(GO:1902912) |
| 1.6 | 4.8 | GO:1990667 | PCSK9-AnxA2 complex(GO:1990667) |
| 1.4 | 15.5 | GO:0097451 | glial limiting end-foot(GO:0097451) |
| 1.0 | 10.9 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.7 | 2.9 | GO:0030312 | external encapsulating structure(GO:0030312) |
| 0.5 | 1.8 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 0.4 | 10.6 | GO:0042581 | specific granule(GO:0042581) |
| 0.3 | 1.0 | GO:0043259 | laminin-10 complex(GO:0043259) |
| 0.2 | 1.3 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.2 | 1.5 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.2 | 0.5 | GO:1990879 | CST complex(GO:1990879) |
| 0.2 | 1.7 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.2 | 1.8 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.2 | 2.2 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.2 | 5.9 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.1 | 0.6 | GO:1990682 | CSF1-CSF1R complex(GO:1990682) |
| 0.1 | 9.8 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.1 | 0.7 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.1 | 3.7 | GO:0032982 | myosin filament(GO:0032982) |
| 0.1 | 1.8 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.1 | 1.9 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.1 | 0.6 | GO:0061673 | mitotic spindle astral microtubule(GO:0061673) |
| 0.1 | 4.3 | GO:0042588 | zymogen granule(GO:0042588) |
| 0.1 | 3.8 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.1 | 0.8 | GO:0097443 | sorting endosome(GO:0097443) |
| 0.1 | 0.6 | GO:0098890 | extrinsic component of postsynaptic membrane(GO:0098890) |
| 0.1 | 0.3 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.1 | 1.7 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.1 | 1.9 | GO:0016342 | catenin complex(GO:0016342) |
| 0.1 | 11.6 | GO:0097517 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.1 | 1.6 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.1 | 5.4 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 3.6 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.1 | 1.1 | GO:0031462 | Cul2-RING ubiquitin ligase complex(GO:0031462) |
| 0.1 | 11.2 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.1 | 3.5 | GO:0008305 | integrin complex(GO:0008305) |
| 0.1 | 1.1 | GO:1990023 | mitotic spindle midzone(GO:1990023) |
| 0.1 | 2.8 | GO:0001741 | XY body(GO:0001741) |
| 0.1 | 1.6 | GO:0042627 | chylomicron(GO:0042627) |
| 0.1 | 0.6 | GO:0072379 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.1 | 1.7 | GO:0005753 | mitochondrial proton-transporting ATP synthase complex(GO:0005753) |
| 0.1 | 0.8 | GO:0097449 | astrocyte projection(GO:0097449) |
| 0.1 | 0.6 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.1 | 0.7 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.1 | 0.4 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.1 | 6.5 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 0.6 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.1 | 1.3 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.0 | 1.6 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 1.0 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.0 | 0.6 | GO:0031045 | dense core granule(GO:0031045) |
| 0.0 | 0.6 | GO:0032009 | early phagosome(GO:0032009) |
| 0.0 | 2.2 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 0.3 | GO:0042825 | TAP complex(GO:0042825) |
| 0.0 | 0.9 | GO:0030867 | rough endoplasmic reticulum membrane(GO:0030867) |
| 0.0 | 0.8 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 0.2 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.0 | 2.6 | GO:0016459 | myosin complex(GO:0016459) |
| 0.0 | 17.3 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.0 | 5.8 | GO:0045111 | intermediate filament cytoskeleton(GO:0045111) |
| 0.0 | 2.2 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 7.6 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 16.3 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.0 | 1.9 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 0.4 | GO:0090543 | Flemming body(GO:0090543) |
| 0.0 | 0.5 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 3.0 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 0.3 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 0.1 | GO:0030891 | VCB complex(GO:0030891) |
| 0.0 | 0.2 | GO:0031933 | telomeric heterochromatin(GO:0031933) |
| 0.0 | 0.5 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 1.7 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 0.4 | GO:0000145 | exocyst(GO:0000145) |
| 0.0 | 1.1 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 0.2 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.0 | 0.3 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.0 | 0.6 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.0 | 0.1 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.0 | 1.9 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 5.8 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 0.4 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.0 | 1.8 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 0.1 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.0 | 1.6 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.0 | 0.5 | GO:0008180 | COP9 signalosome(GO:0008180) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.5 | 9.9 | GO:0004736 | pyruvate carboxylase activity(GO:0004736) |
| 2.2 | 17.8 | GO:0050649 | testosterone 6-beta-hydroxylase activity(GO:0050649) |
| 1.8 | 10.9 | GO:1990254 | keratin filament binding(GO:1990254) |
| 1.2 | 3.6 | GO:0004611 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 1.1 | 3.2 | GO:0047291 | lactosylceramide alpha-2,3-sialyltransferase activity(GO:0047291) |
| 1.1 | 4.3 | GO:0004909 | interleukin-1, Type I, activating receptor activity(GO:0004909) |
| 1.0 | 5.1 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.8 | 5.9 | GO:0015265 | urea channel activity(GO:0015265) |
| 0.8 | 6.6 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.8 | 3.0 | GO:0051435 | BH4 domain binding(GO:0051435) |
| 0.7 | 2.8 | GO:0001733 | galactosylceramide sulfotransferase activity(GO:0001733) |
| 0.7 | 18.0 | GO:0005123 | death receptor binding(GO:0005123) |
| 0.6 | 3.8 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 0.6 | 6.7 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.5 | 4.8 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 0.5 | 4.8 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.5 | 5.1 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.5 | 2.3 | GO:0016167 | glial cell-derived neurotrophic factor receptor activity(GO:0016167) |
| 0.4 | 2.2 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 0.4 | 1.8 | GO:0043532 | angiostatin binding(GO:0043532) |
| 0.4 | 2.9 | GO:0016453 | acetyl-CoA C-acetyltransferase activity(GO:0003985) C-acetyltransferase activity(GO:0016453) |
| 0.4 | 1.5 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.4 | 2.9 | GO:0005534 | galactose binding(GO:0005534) |
| 0.3 | 1.3 | GO:0008941 | nitric oxide dioxygenase activity(GO:0008941) oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of two atoms of oxygen into one donor(GO:0016708) iron-cytochrome-c reductase activity(GO:0047726) |
| 0.3 | 2.3 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.3 | 2.2 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.3 | 7.2 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.3 | 8.8 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.2 | 2.2 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.2 | 7.3 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) |
| 0.2 | 1.9 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.2 | 18.1 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.2 | 0.7 | GO:0004368 | glycerol-3-phosphate dehydrogenase activity(GO:0004368) oxidoreductase activity, acting on the CH-OH group of donors, quinone or similar compound as acceptor(GO:0016901) |
| 0.2 | 2.4 | GO:0015217 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.2 | 4.0 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.2 | 4.4 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.2 | 1.8 | GO:0003857 | 3-hydroxyacyl-CoA dehydrogenase activity(GO:0003857) |
| 0.2 | 2.4 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.2 | 1.6 | GO:0060228 | phosphatidylcholine-sterol O-acyltransferase activator activity(GO:0060228) |
| 0.2 | 0.6 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 0.2 | 1.3 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.2 | 1.3 | GO:0008294 | calcium- and calmodulin-responsive adenylate cyclase activity(GO:0008294) |
| 0.2 | 2.9 | GO:0015299 | solute:proton antiporter activity(GO:0015299) |
| 0.2 | 0.7 | GO:0008732 | threonine aldolase activity(GO:0004793) L-allo-threonine aldolase activity(GO:0008732) |
| 0.2 | 1.7 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.2 | 0.5 | GO:0035939 | microsatellite binding(GO:0035939) |
| 0.2 | 1.4 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.2 | 1.2 | GO:0019158 | fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.1 | 0.8 | GO:0038085 | vascular endothelial growth factor binding(GO:0038085) |
| 0.1 | 1.1 | GO:0034235 | GPI anchor binding(GO:0034235) |
| 0.1 | 0.5 | GO:0004165 | dodecenoyl-CoA delta-isomerase activity(GO:0004165) |
| 0.1 | 1.6 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.1 | 0.4 | GO:0005457 | GDP-fucose transmembrane transporter activity(GO:0005457) purine nucleotide-sugar transmembrane transporter activity(GO:0036080) |
| 0.1 | 1.9 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.1 | 0.8 | GO:0015315 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.1 | 0.6 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.1 | 0.4 | GO:0003692 | left-handed Z-DNA binding(GO:0003692) |
| 0.1 | 2.0 | GO:0015143 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.1 | 0.3 | GO:0015433 | peptide antigen-transporting ATPase activity(GO:0015433) tapasin binding(GO:0046980) |
| 0.1 | 7.3 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.1 | 0.2 | GO:0032129 | histone deacetylase activity (H3-K9 specific)(GO:0032129) NAD-dependent histone deacetylase activity (H3-K9 specific)(GO:0046969) |
| 0.1 | 2.7 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.1 | 3.1 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.1 | 0.3 | GO:0004478 | methionine adenosyltransferase activity(GO:0004478) |
| 0.1 | 0.7 | GO:0004030 | aldehyde dehydrogenase [NAD(P)+] activity(GO:0004030) |
| 0.1 | 2.5 | GO:0000146 | microfilament motor activity(GO:0000146) |
| 0.1 | 0.7 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.1 | 0.5 | GO:0004920 | interleukin-10 receptor activity(GO:0004920) |
| 0.1 | 1.9 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.1 | 1.2 | GO:0001164 | RNA polymerase I regulatory region sequence-specific DNA binding(GO:0001163) RNA polymerase I CORE element sequence-specific DNA binding(GO:0001164) |
| 0.1 | 0.5 | GO:0043734 | DNA-N1-methyladenine dioxygenase activity(GO:0043734) methylcytosine dioxygenase activity(GO:0070579) |
| 0.1 | 0.3 | GO:0070976 | TIR domain binding(GO:0070976) |
| 0.1 | 0.2 | GO:0019150 | D-ribulokinase activity(GO:0019150) |
| 0.1 | 1.9 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.1 | 4.4 | GO:0008028 | monocarboxylic acid transmembrane transporter activity(GO:0008028) |
| 0.1 | 0.5 | GO:0004046 | aminoacylase activity(GO:0004046) |
| 0.1 | 3.5 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.1 | 3.3 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.1 | 0.4 | GO:0004483 | mRNA (nucleoside-2'-O-)-methyltransferase activity(GO:0004483) |
| 0.1 | 4.2 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.1 | 0.4 | GO:0047288 | monosialoganglioside sialyltransferase activity(GO:0047288) |
| 0.1 | 11.2 | GO:0004386 | helicase activity(GO:0004386) |
| 0.1 | 1.3 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 0.1 | 6.6 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.1 | 0.5 | GO:0016308 | 1-phosphatidylinositol-4-phosphate 5-kinase activity(GO:0016308) |
| 0.0 | 0.6 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 0.0 | 0.1 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 0.0 | 1.4 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 0.0 | 0.4 | GO:0015272 | ATP-activated inward rectifier potassium channel activity(GO:0015272) |
| 0.0 | 0.3 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.0 | 0.2 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.0 | 2.0 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 0.5 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.0 | 0.1 | GO:0018169 | ribosomal S6-glutamic acid ligase activity(GO:0018169) |
| 0.0 | 0.8 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.0 | 0.6 | GO:0035325 | Toll-like receptor binding(GO:0035325) |
| 0.0 | 1.1 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.6 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.0 | 1.8 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.0 | 0.2 | GO:0047179 | platelet-activating factor acetyltransferase activity(GO:0047179) |
| 0.0 | 0.2 | GO:0016888 | endodeoxyribonuclease activity, producing 5'-phosphomonoesters(GO:0016888) |
| 0.0 | 6.3 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 2.3 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 0.3 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.0 | 3.8 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.0 | 0.2 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.2 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.0 | 1.1 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 1.3 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.0 | 0.4 | GO:0004862 | cAMP-dependent protein kinase inhibitor activity(GO:0004862) |
| 0.0 | 0.2 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.0 | 0.5 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.0 | 1.6 | GO:0030295 | protein kinase activator activity(GO:0030295) |
| 0.0 | 1.3 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.3 | GO:0050308 | sugar-phosphatase activity(GO:0050308) |
| 0.0 | 0.1 | GO:0035851 | Krueppel-associated box domain binding(GO:0035851) |
| 0.0 | 0.8 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.9 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 0.1 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 0.0 | 0.2 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.0 | 0.4 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.3 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.0 | 0.3 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.1 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.0 | 0.5 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.0 | 1.6 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.3 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.0 | 1.3 | GO:0051213 | dioxygenase activity(GO:0051213) |
| 0.0 | 5.2 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 1.2 | GO:0043621 | protein self-association(GO:0043621) |
| 0.0 | 0.1 | GO:0070320 | inward rectifier potassium channel inhibitor activity(GO:0070320) |
| 0.0 | 0.3 | GO:0016675 | oxidoreductase activity, acting on a heme group of donors(GO:0016675) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 11.0 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.2 | 12.9 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.2 | 6.9 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.1 | 11.1 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 5.2 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.1 | 8.9 | PID INTEGRIN1 PATHWAY | Beta1 integrin cell surface interactions |
| 0.1 | 0.8 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.1 | 11.2 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.1 | 4.3 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.1 | 2.1 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.1 | 4.2 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.1 | 1.3 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 3.4 | PID ARF6 TRAFFICKING PATHWAY | Arf6 trafficking events |
| 0.1 | 2.9 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 2.6 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.1 | 11.5 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 4.7 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.0 | 0.9 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 1.2 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 0.6 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.0 | 3.6 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.0 | 2.7 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 0.8 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.0 | 3.1 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 1.5 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.0 | 1.3 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.0 | 4.7 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.7 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 1.8 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.0 | 1.8 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
| 0.0 | 1.8 | PID ERBB1 DOWNSTREAM PATHWAY | ErbB1 downstream signaling |
| 0.0 | 0.2 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.0 | 0.2 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 1.0 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 1.0 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.0 | 0.2 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.0 | 0.2 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 0.6 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 4.0 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.4 | 9.5 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.3 | 12.3 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.3 | 4.8 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.3 | 10.5 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.3 | 14.0 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.3 | 5.9 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.2 | 3.8 | REACTOME PHOSPHORYLATION OF THE APC C | Genes involved in Phosphorylation of the APC/C |
| 0.2 | 2.5 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.1 | 1.8 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.1 | 4.7 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.1 | 1.7 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.1 | 1.8 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 1.5 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.1 | 1.7 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.1 | 2.5 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.1 | 9.3 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.1 | 5.3 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.1 | 3.0 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 0.8 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.1 | 1.7 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.1 | 4.4 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.1 | 4.5 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 4.2 | REACTOME SIGNAL TRANSDUCTION BY L1 | Genes involved in Signal transduction by L1 |
| 0.1 | 0.9 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.1 | 1.9 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.1 | 1.5 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.1 | 2.1 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 2.2 | REACTOME MUSCLE CONTRACTION | Genes involved in Muscle contraction |
| 0.1 | 1.7 | REACTOME SHC1 EVENTS IN ERBB4 SIGNALING | Genes involved in SHC1 events in ERBB4 signaling |
| 0.1 | 0.9 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.1 | 1.5 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.1 | 1.8 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 2.5 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 0.8 | REACTOME INITIAL TRIGGERING OF COMPLEMENT | Genes involved in Initial triggering of complement |
| 0.0 | 0.6 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.0 | 1.5 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.0 | 0.6 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.0 | 1.5 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 1.8 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.5 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.0 | 2.6 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.8 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.7 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 3.2 | REACTOME RESPIRATORY ELECTRON TRANSPORT ATP SYNTHESIS BY CHEMIOSMOTIC COUPLING AND HEAT PRODUCTION BY UNCOUPLING PROTEINS | Genes involved in Respiratory electron transport, ATP synthesis by chemiosmotic coupling, and heat production by uncoupling proteins. |
| 0.0 | 1.5 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.0 | 0.8 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 1.5 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.0 | 0.6 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.5 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.0 | 1.2 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 2.9 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.5 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.0 | 0.4 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.0 | 1.9 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 0.2 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.0 | 0.7 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |