Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
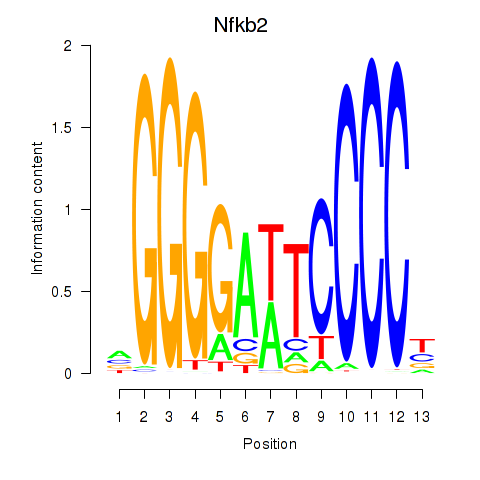
Results for Nfkb2
Z-value: 0.42
Transcription factors associated with Nfkb2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Nfkb2
|
ENSMUSG00000025225.16 | nuclear factor of kappa light polypeptide gene enhancer in B cells 2, p49/p100 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Nfkb2 | mm39_v1_chr19_+_46292759_46292845 | 0.49 | 2.6e-03 | Click! |
Activity profile of Nfkb2 motif
Sorted Z-values of Nfkb2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr14_+_80237691 | 3.84 |
ENSMUST00000228749.2
ENSMUST00000088735.4 |
Olfm4
|
olfactomedin 4 |
| chr17_+_56259617 | 2.10 |
ENSMUST00000003274.8
|
Ebi3
|
Epstein-Barr virus induced gene 3 |
| chr7_-_44181477 | 2.00 |
ENSMUST00000098483.9
ENSMUST00000035323.6 |
Spib
|
Spi-B transcription factor (Spi-1/PU.1 related) |
| chrX_-_7537580 | 1.99 |
ENSMUST00000033486.6
|
Plp2
|
proteolipid protein 2 |
| chr7_+_44866635 | 1.96 |
ENSMUST00000097216.5
ENSMUST00000209343.2 ENSMUST00000209678.2 |
Tead2
|
TEA domain family member 2 |
| chr1_+_153616090 | 1.44 |
ENSMUST00000027748.8
|
Rgs16
|
regulator of G-protein signaling 16 |
| chr8_+_79755194 | 1.29 |
ENSMUST00000119254.8
ENSMUST00000238669.2 |
Zfp827
|
zinc finger protein 827 |
| chr19_-_46327024 | 1.22 |
ENSMUST00000236046.2
ENSMUST00000236980.2 ENSMUST00000235485.2 ENSMUST00000236061.2 ENSMUST00000236236.2 ENSMUST00000236768.2 ENSMUST00000236651.2 |
Cuedc2
|
CUE domain containing 2 |
| chr19_-_46327071 | 1.16 |
ENSMUST00000235977.2
ENSMUST00000167861.8 ENSMUST00000051234.9 ENSMUST00000236066.2 |
Cuedc2
|
CUE domain containing 2 |
| chr11_+_70548022 | 1.15 |
ENSMUST00000157027.8
ENSMUST00000072841.12 ENSMUST00000108548.8 ENSMUST00000126241.8 |
Eno3
|
enolase 3, beta muscle |
| chr8_+_79754980 | 1.04 |
ENSMUST00000087927.11
ENSMUST00000098614.9 |
Zfp827
|
zinc finger protein 827 |
| chr19_+_46292759 | 0.83 |
ENSMUST00000237330.2
|
Nfkb2
|
nuclear factor of kappa light polypeptide gene enhancer in B cells 2, p49/p100 |
| chr8_+_120729459 | 0.80 |
ENSMUST00000108972.4
|
Crispld2
|
cysteine-rich secretory protein LCCL domain containing 2 |
| chr11_+_70548622 | 0.79 |
ENSMUST00000170716.8
|
Eno3
|
enolase 3, beta muscle |
| chr14_-_30348153 | 0.77 |
ENSMUST00000112211.9
ENSMUST00000112210.11 |
Prkcd
|
protein kinase C, delta |
| chr19_+_6391148 | 0.75 |
ENSMUST00000025897.13
ENSMUST00000130382.8 |
Map4k2
|
mitogen-activated protein kinase kinase kinase kinase 2 |
| chr2_+_102380357 | 0.71 |
ENSMUST00000028612.8
|
Pamr1
|
peptidase domain containing associated with muscle regeneration 1 |
| chr7_+_3339059 | 0.69 |
ENSMUST00000096744.8
|
Myadm
|
myeloid-associated differentiation marker |
| chr11_-_54853729 | 0.68 |
ENSMUST00000108885.8
ENSMUST00000102730.9 ENSMUST00000018482.13 ENSMUST00000108886.8 ENSMUST00000102731.8 |
Tnip1
|
TNFAIP3 interacting protein 1 |
| chr19_+_46293210 | 0.65 |
ENSMUST00000236591.2
|
Nfkb2
|
nuclear factor of kappa light polypeptide gene enhancer in B cells 2, p49/p100 |
| chr7_-_44702269 | 0.62 |
ENSMUST00000057293.8
|
Prr12
|
proline rich 12 |
| chr14_-_30329765 | 0.60 |
ENSMUST00000112207.8
ENSMUST00000112206.8 ENSMUST00000112202.8 ENSMUST00000112203.2 |
Prkcd
|
protein kinase C, delta |
| chr5_-_137529251 | 0.58 |
ENSMUST00000132525.8
|
Gnb2
|
guanine nucleotide binding protein (G protein), beta 2 |
| chr19_+_46293160 | 0.55 |
ENSMUST00000073116.13
|
Nfkb2
|
nuclear factor of kappa light polypeptide gene enhancer in B cells 2, p49/p100 |
| chr5_-_137529465 | 0.54 |
ENSMUST00000150063.9
|
Gnb2
|
guanine nucleotide binding protein (G protein), beta 2 |
| chr11_+_70548513 | 0.50 |
ENSMUST00000134087.8
|
Eno3
|
enolase 3, beta muscle |
| chr13_+_43938251 | 0.42 |
ENSMUST00000015540.4
|
Cd83
|
CD83 antigen |
| chr7_-_125799644 | 0.39 |
ENSMUST00000168189.8
|
Xpo6
|
exportin 6 |
| chr15_-_102154874 | 0.37 |
ENSMUST00000063339.14
|
Rarg
|
retinoic acid receptor, gamma |
| chr2_-_93292257 | 0.35 |
ENSMUST00000150508.8
|
Cd82
|
CD82 antigen |
| chr3_+_97565528 | 0.35 |
ENSMUST00000045743.13
|
Prkab2
|
protein kinase, AMP-activated, beta 2 non-catalytic subunit |
| chr7_-_19363280 | 0.34 |
ENSMUST00000094762.10
ENSMUST00000049912.15 ENSMUST00000098754.5 |
Relb
|
avian reticuloendotheliosis viral (v-rel) oncogene related B |
| chr7_-_125799570 | 0.34 |
ENSMUST00000009344.16
|
Xpo6
|
exportin 6 |
| chr9_+_5308828 | 0.33 |
ENSMUST00000162846.8
ENSMUST00000027012.14 |
Casp4
|
caspase 4, apoptosis-related cysteine peptidase |
| chr7_+_28988724 | 0.31 |
ENSMUST00000207714.2
ENSMUST00000048187.6 |
Ppp1r14a
|
protein phosphatase 1, regulatory inhibitor subunit 14A |
| chr4_+_129878890 | 0.31 |
ENSMUST00000106017.8
ENSMUST00000121049.8 |
Adgrb2
|
adhesion G protein-coupled receptor B2 |
| chr2_-_37312881 | 0.30 |
ENSMUST00000112936.4
ENSMUST00000112934.8 |
Rc3h2
|
ring finger and CCCH-type zinc finger domains 2 |
| chr19_+_53517528 | 0.30 |
ENSMUST00000038287.7
|
Dusp5
|
dual specificity phosphatase 5 |
| chr17_-_36291087 | 0.28 |
ENSMUST00000055454.14
|
Prr3
|
proline-rich polypeptide 3 |
| chr9_-_106762818 | 0.28 |
ENSMUST00000185707.2
|
Rbm15b
|
RNA binding motif protein 15B |
| chr3_-_107667499 | 0.28 |
ENSMUST00000153114.2
ENSMUST00000118593.8 ENSMUST00000120243.8 |
Csf1
|
colony stimulating factor 1 (macrophage) |
| chr7_+_30121776 | 0.26 |
ENSMUST00000108175.8
|
Nfkbid
|
nuclear factor of kappa light polypeptide gene enhancer in B cells inhibitor, delta |
| chr4_+_124880223 | 0.25 |
ENSMUST00000030687.8
|
Rspo1
|
R-spondin 1 |
| chr8_+_124138163 | 0.21 |
ENSMUST00000071134.4
ENSMUST00000212743.2 |
Tubb3
|
tubulin, beta 3 class III |
| chr4_-_125021610 | 0.20 |
ENSMUST00000036188.8
|
Zc3h12a
|
zinc finger CCCH type containing 12A |
| chr17_+_36290743 | 0.19 |
ENSMUST00000087200.4
|
Gnl1
|
guanine nucleotide binding protein-like 1 |
| chr11_+_101066867 | 0.18 |
ENSMUST00000103109.4
|
Cntnap1
|
contactin associated protein-like 1 |
| chr3_+_31149201 | 0.16 |
ENSMUST00000118470.8
ENSMUST00000029194.12 ENSMUST00000123532.2 |
Skil
|
SKI-like |
| chr8_-_105357941 | 0.13 |
ENSMUST00000034351.8
|
Rrad
|
Ras-related associated with diabetes |
| chr11_-_71928300 | 0.12 |
ENSMUST00000048207.10
|
Aipl1
|
aryl hydrocarbon receptor-interacting protein-like 1 |
| chr1_+_149975782 | 0.11 |
ENSMUST00000035065.9
|
Ptgs2
|
prostaglandin-endoperoxide synthase 2 |
| chr9_-_44646487 | 0.11 |
ENSMUST00000034611.15
|
Phldb1
|
pleckstrin homology like domain, family B, member 1 |
| chr9_-_107486381 | 0.09 |
ENSMUST00000102531.7
ENSMUST00000102530.8 ENSMUST00000195057.2 ENSMUST00000102532.10 |
Sema3b
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3B |
| chr1_-_63153675 | 0.06 |
ENSMUST00000097718.9
|
Ino80d
|
INO80 complex subunit D |
| chr14_+_55829165 | 0.06 |
ENSMUST00000019443.15
|
Rnf31
|
ring finger protein 31 |
| chr3_+_31149247 | 0.06 |
ENSMUST00000117728.8
|
Skil
|
SKI-like |
| chr11_+_77576981 | 0.05 |
ENSMUST00000100802.11
ENSMUST00000181023.2 |
Nufip2
|
nuclear fragile X mental retardation protein interacting protein 2 |
| chr5_+_90907207 | 0.04 |
ENSMUST00000031318.6
|
Cxcl5
|
chemokine (C-X-C motif) ligand 5 |
| chr17_-_88372315 | 0.03 |
ENSMUST00000235056.2
ENSMUST00000130379.9 |
Fbxo11
|
F-box protein 11 |
| chr7_-_125799585 | 0.02 |
ENSMUST00000163959.2
|
Xpo6
|
exportin 6 |
| chr7_+_18910340 | 0.02 |
ENSMUST00000117338.8
|
Eml2
|
echinoderm microtubule associated protein like 2 |
| chr4_+_129878627 | 0.02 |
ENSMUST00000120204.8
|
Adgrb2
|
adhesion G protein-coupled receptor B2 |
| chr11_+_69920542 | 0.01 |
ENSMUST00000232266.2
ENSMUST00000132597.5 |
Dlg4
|
discs large MAGUK scaffold protein 4 |
| chr6_+_68547717 | 0.00 |
ENSMUST00000196839.5
ENSMUST00000103327.3 |
Igkv12-98
|
immunoglobulin kappa variable 12-98 |
| chr7_+_16043502 | 0.00 |
ENSMUST00000002152.13
|
Bbc3
|
BCL2 binding component 3 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Nfkb2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.0 | GO:0002268 | follicular dendritic cell activation(GO:0002266) follicular dendritic cell differentiation(GO:0002268) |
| 0.3 | 1.4 | GO:0060697 | positive regulation of phospholipid catabolic process(GO:0060697) |
| 0.3 | 2.4 | GO:0010936 | negative regulation of macrophage cytokine production(GO:0010936) |
| 0.1 | 0.9 | GO:1903003 | positive regulation of protein deubiquitination(GO:1903003) |
| 0.1 | 0.3 | GO:1903969 | regulation of macrophage colony-stimulating factor signaling pathway(GO:1902226) regulation of response to macrophage colony-stimulating factor(GO:1903969) regulation of cellular response to macrophage colony-stimulating factor stimulus(GO:1903972) |
| 0.1 | 0.7 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.1 | 2.0 | GO:0048368 | lateral mesoderm development(GO:0048368) |
| 0.1 | 0.4 | GO:0003430 | growth plate cartilage chondrocyte growth(GO:0003430) |
| 0.1 | 3.8 | GO:1900026 | positive regulation of substrate adhesion-dependent cell spreading(GO:1900026) |
| 0.0 | 0.3 | GO:1903265 | positive regulation of tumor necrosis factor-mediated signaling pathway(GO:1903265) |
| 0.0 | 0.4 | GO:0032713 | negative regulation of interleukin-4 production(GO:0032713) |
| 0.0 | 2.0 | GO:0030225 | macrophage differentiation(GO:0030225) |
| 0.0 | 0.1 | GO:0090362 | positive regulation of platelet-derived growth factor production(GO:0090362) |
| 0.0 | 0.3 | GO:0061470 | T follicular helper cell differentiation(GO:0061470) |
| 0.0 | 0.2 | GO:0033085 | negative regulation of T cell differentiation in thymus(GO:0033085) |
| 0.0 | 0.1 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.0 | 0.3 | GO:2000254 | regulation of male germ cell proliferation(GO:2000254) |
| 0.0 | 0.7 | GO:0006903 | vesicle targeting(GO:0006903) |
| 0.0 | 0.3 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.0 | 2.4 | GO:0006096 | glycolytic process(GO:0006096) |
| 0.0 | 0.1 | GO:1901842 | negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.0 | 0.2 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.0 | 0.3 | GO:0043011 | myeloid dendritic cell differentiation(GO:0043011) |
| 0.0 | 0.2 | GO:1902231 | positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902231) |
| 0.0 | 0.8 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.0 | 0.1 | GO:1904259 | regulation of basement membrane assembly involved in embryonic body morphogenesis(GO:1904259) positive regulation of basement membrane assembly involved in embryonic body morphogenesis(GO:1904261) basement membrane assembly involved in embryonic body morphogenesis(GO:2001197) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.0 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 0.4 | 2.0 | GO:0033257 | Bcl3/NF-kappaB2 complex(GO:0033257) |
| 0.2 | 2.4 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.2 | 3.8 | GO:0042581 | specific granule(GO:0042581) |
| 0.1 | 0.3 | GO:0097169 | AIM2 inflammasome complex(GO:0097169) |
| 0.1 | 0.3 | GO:1990682 | CSF1-CSF1R complex(GO:1990682) |
| 0.0 | 0.2 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.0 | 0.3 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.0 | 1.1 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.1 | GO:0071797 | LUBAC complex(GO:0071797) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.4 | GO:0070976 | calcium-independent protein kinase C activity(GO:0004699) TIR domain binding(GO:0070976) |
| 0.2 | 2.4 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.1 | 0.3 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 0.1 | 2.0 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 0.1 | 2.0 | GO:0019956 | chemokine binding(GO:0019956) |
| 0.0 | 0.7 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.0 | 0.3 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.0 | 1.6 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.0 | 1.1 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 0.3 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.0 | 0.4 | GO:0003708 | retinoic acid receptor activity(GO:0003708) |
| 0.0 | 2.1 | GO:0004896 | cytokine receptor activity(GO:0004896) |
| 0.0 | 0.3 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.0 | 0.8 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.0 | 0.2 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.0 | 0.7 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.1 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.0 | 2.0 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.0 | 1.4 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 0.3 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.0 | 0.7 | PID TNF PATHWAY | TNF receptor signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.0 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.0 | 2.4 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 2.0 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 1.4 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 1.1 | REACTOME G BETA GAMMA SIGNALLING THROUGH PLC BETA | Genes involved in G beta:gamma signalling through PLC beta |
| 0.0 | 0.3 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.0 | 0.2 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |