Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
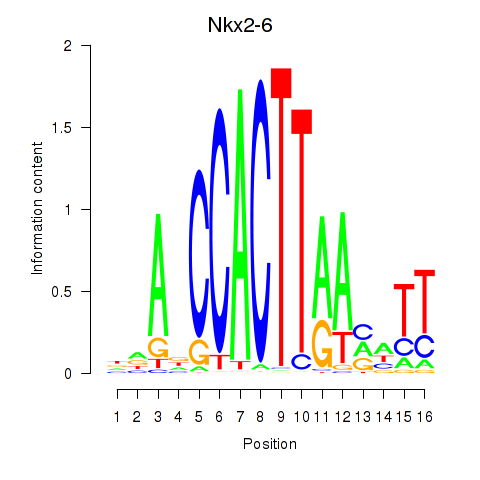
Results for Nkx2-6
Z-value: 0.72
Transcription factors associated with Nkx2-6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Nkx2-6
|
ENSMUSG00000044186.11 | NK2 homeobox 6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Nkx2-6 | mm39_v1_chr14_+_69409251_69409414 | -0.79 | 1.1e-08 | Click! |
Activity profile of Nkx2-6 motif
Sorted Z-values of Nkx2-6 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_-_41681273 | 18.30 |
ENSMUST00000031899.14
|
Kel
|
Kell blood group |
| chr4_+_46039202 | 9.15 |
ENSMUST00000156200.8
|
Tmod1
|
tropomodulin 1 |
| chr15_+_73594965 | 8.54 |
ENSMUST00000165541.8
ENSMUST00000167582.8 |
Ptp4a3
|
protein tyrosine phosphatase 4a3 |
| chr15_+_73595012 | 8.43 |
ENSMUST00000230044.2
|
Ptp4a3
|
protein tyrosine phosphatase 4a3 |
| chr7_-_115630282 | 3.74 |
ENSMUST00000206034.2
ENSMUST00000106612.8 |
Sox6
|
SRY (sex determining region Y)-box 6 |
| chr5_+_110478558 | 2.18 |
ENSMUST00000112481.2
|
Pole
|
polymerase (DNA directed), epsilon |
| chr4_-_109522502 | 1.68 |
ENSMUST00000063531.5
|
Cdkn2c
|
cyclin dependent kinase inhibitor 2C |
| chr15_-_78657640 | 1.65 |
ENSMUST00000018313.6
|
Mfng
|
MFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr5_+_145063568 | 1.34 |
ENSMUST00000138922.2
|
Arpc1b
|
actin related protein 2/3 complex, subunit 1B |
| chr10_+_4297762 | 1.20 |
ENSMUST00000215696.2
|
Akap12
|
A kinase (PRKA) anchor protein (gravin) 12 |
| chr2_+_3115250 | 1.06 |
ENSMUST00000072955.12
|
Fam171a1
|
family with sequence similarity 171, member A1 |
| chr19_+_8779903 | 0.97 |
ENSMUST00000172175.3
|
Zbtb3
|
zinc finger and BTB domain containing 3 |
| chr2_+_85715984 | 0.60 |
ENSMUST00000213441.3
|
Olfr1023
|
olfactory receptor 1023 |
| chr1_+_179788675 | 0.38 |
ENSMUST00000076687.12
ENSMUST00000097450.10 ENSMUST00000212756.2 |
Cdc42bpa
|
CDC42 binding protein kinase alpha |
| chr1_+_179788037 | 0.31 |
ENSMUST00000097453.9
ENSMUST00000111117.8 |
Cdc42bpa
|
CDC42 binding protein kinase alpha |
| chr12_-_114263874 | 0.15 |
ENSMUST00000103482.2
ENSMUST00000194159.2 |
Ighv9-4
|
immunoglobulin heavy variable 9-4 |
| chr4_+_62537750 | 0.15 |
ENSMUST00000084521.11
ENSMUST00000107424.2 |
Rgs3
|
regulator of G-protein signaling 3 |
| chr12_-_11485639 | 0.11 |
ENSMUST00000220506.2
|
Vsnl1
|
visinin-like 1 |
| chr11_+_103664976 | 0.09 |
ENSMUST00000000127.6
|
Wnt3
|
wingless-type MMTV integration site family, member 3 |
| chr7_-_115837034 | 0.08 |
ENSMUST00000182834.8
|
Plekha7
|
pleckstrin homology domain containing, family A member 7 |
| chr12_-_114073050 | 0.06 |
ENSMUST00000103472.4
|
Ighv9-2
|
immunoglobulin heavy variable V9-2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Nkx2-6
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 18.3 | GO:0031133 | regulation of axon diameter(GO:0031133) |
| 1.5 | 9.2 | GO:0030241 | skeletal muscle myosin thick filament assembly(GO:0030241) |
| 0.9 | 17.0 | GO:0043117 | positive regulation of vascular permeability(GO:0043117) |
| 0.7 | 2.2 | GO:0045004 | DNA replication proofreading(GO:0045004) |
| 0.3 | 3.7 | GO:2000741 | positive regulation of mesenchymal stem cell differentiation(GO:2000741) |
| 0.1 | 1.7 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.1 | 1.2 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.0 | 1.7 | GO:1904030 | negative regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045736) negative regulation of cyclin-dependent protein kinase activity(GO:1904030) |
| 0.0 | 0.1 | GO:0060061 | Spemann organizer formation(GO:0060061) canonical Wnt signaling pathway involved in midbrain dopaminergic neuron differentiation(GO:1904954) |
| 0.0 | 1.3 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 0.7 | GO:0040023 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
| 0.0 | 0.1 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.2 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.3 | 1.3 | GO:0036284 | tubulobulbar complex(GO:0036284) |
| 0.2 | 9.2 | GO:0005865 | striated muscle thin filament(GO:0005865) |
| 0.0 | 1.7 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 17.0 | GO:0005768 | endosome(GO:0005768) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.7 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.4 | 9.2 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.2 | 2.2 | GO:0008310 | single-stranded DNA 3'-5' exodeoxyribonuclease activity(GO:0008310) |
| 0.1 | 17.7 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 17.0 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.1 | 1.7 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.0 | 1.2 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.0 | 3.7 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 17.0 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.0 | 1.7 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 2.0 | PID CDC42 PATHWAY | CDC42 signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 9.2 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.1 | 2.2 | REACTOME REPAIR SYNTHESIS FOR GAP FILLING BY DNA POL IN TC NER | Genes involved in Repair synthesis for gap-filling by DNA polymerase in TC-NER |
| 0.1 | 1.7 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 1.7 | REACTOME G1 PHASE | Genes involved in G1 Phase |