Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
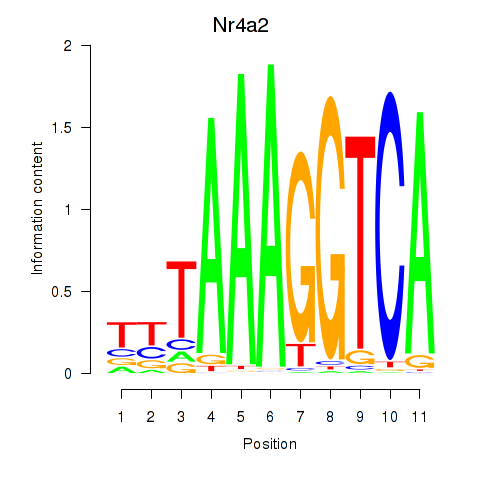
Results for Nr4a2
Z-value: 0.75
Transcription factors associated with Nr4a2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Nr4a2
|
ENSMUSG00000026826.14 | nuclear receptor subfamily 4, group A, member 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Nr4a2 | mm39_v1_chr2_-_57003064_57003084 | -0.28 | 1.0e-01 | Click! |
Activity profile of Nr4a2 motif
Sorted Z-values of Nr4a2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_+_76579960 | 3.14 |
ENSMUST00000229679.2
|
Gpt
|
glutamic pyruvic transaminase, soluble |
| chr7_-_14172434 | 2.50 |
ENSMUST00000210396.2
ENSMUST00000168252.9 |
Sult2a8
|
sulfotransferase family 2A, dehydroepiandrosterone (DHEA)-preferring, member 8 |
| chr9_-_21900724 | 1.99 |
ENSMUST00000045726.8
|
Rgl3
|
ral guanine nucleotide dissociation stimulator-like 3 |
| chr15_+_25940781 | 1.67 |
ENSMUST00000227275.2
|
Retreg1
|
reticulophagy regulator 1 |
| chr15_+_25940912 | 1.67 |
ENSMUST00000226438.2
|
Retreg1
|
reticulophagy regulator 1 |
| chr15_+_25940931 | 1.60 |
ENSMUST00000110438.3
|
Retreg1
|
reticulophagy regulator 1 |
| chr5_+_30437579 | 1.57 |
ENSMUST00000145167.9
|
Selenoi
|
selenoprotein I |
| chr7_+_51537645 | 1.54 |
ENSMUST00000208711.2
|
Gas2
|
growth arrest specific 2 |
| chr16_+_43960183 | 1.51 |
ENSMUST00000159514.8
ENSMUST00000161326.8 ENSMUST00000063520.15 ENSMUST00000063542.8 |
Naa50
|
N(alpha)-acetyltransferase 50, NatE catalytic subunit |
| chr16_-_43959993 | 1.38 |
ENSMUST00000137557.8
|
Atp6v1a
|
ATPase, H+ transporting, lysosomal V1 subunit A |
| chr2_-_73722932 | 1.29 |
ENSMUST00000154456.8
ENSMUST00000090802.11 ENSMUST00000055833.12 ENSMUST00000112007.8 ENSMUST00000112016.9 |
Atf2
|
activating transcription factor 2 |
| chr1_+_167445815 | 1.27 |
ENSMUST00000111380.2
|
Rxrg
|
retinoid X receptor gamma |
| chr2_+_61423469 | 1.21 |
ENSMUST00000112494.2
|
Tank
|
TRAF family member-associated Nf-kappa B activator |
| chr2_+_61423421 | 1.19 |
ENSMUST00000112495.8
ENSMUST00000112501.9 |
Tank
|
TRAF family member-associated Nf-kappa B activator |
| chr2_-_73722874 | 1.12 |
ENSMUST00000136958.8
ENSMUST00000112010.9 ENSMUST00000128531.8 ENSMUST00000112017.8 |
Atf2
|
activating transcription factor 2 |
| chr8_-_65186565 | 1.10 |
ENSMUST00000141021.2
|
Msmo1
|
methylsterol monoxygenase 1 |
| chr5_-_66308421 | 1.10 |
ENSMUST00000200775.4
ENSMUST00000094756.11 |
Rbm47
|
RNA binding motif protein 47 |
| chr9_-_21913833 | 1.05 |
ENSMUST00000115336.10
|
Odad3
|
outer dynein arm docking complex subunit 3 |
| chr1_-_63215952 | 1.05 |
ENSMUST00000185412.7
ENSMUST00000027111.15 ENSMUST00000189664.2 |
Ndufs1
|
NADH:ubiquinone oxidoreductase core subunit S1 |
| chr16_-_43959559 | 1.02 |
ENSMUST00000063661.13
ENSMUST00000114666.9 |
Atp6v1a
|
ATPase, H+ transporting, lysosomal V1 subunit A |
| chr8_+_105573693 | 1.01 |
ENSMUST00000055052.6
|
Ces2c
|
carboxylesterase 2C |
| chr9_-_21913896 | 0.99 |
ENSMUST00000044926.6
|
Odad3
|
outer dynein arm docking complex subunit 3 |
| chr16_-_43960045 | 0.97 |
ENSMUST00000147025.2
|
Atp6v1a
|
ATPase, H+ transporting, lysosomal V1 subunit A |
| chr8_-_71315902 | 0.94 |
ENSMUST00000212611.2
|
Kcnn1
|
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 1 |
| chr1_-_63215812 | 0.93 |
ENSMUST00000185847.2
ENSMUST00000185732.7 ENSMUST00000188370.7 ENSMUST00000168099.9 |
Ndufs1
|
NADH:ubiquinone oxidoreductase core subunit S1 |
| chr18_-_57108405 | 0.85 |
ENSMUST00000139243.9
ENSMUST00000025488.15 |
C330018D20Rik
|
RIKEN cDNA C330018D20 gene |
| chr11_+_4207557 | 0.84 |
ENSMUST00000066283.12
|
Lif
|
leukemia inhibitory factor |
| chrX_-_47602395 | 0.83 |
ENSMUST00000114945.9
ENSMUST00000037349.8 |
Aifm1
|
apoptosis-inducing factor, mitochondrion-associated 1 |
| chr10_-_42354276 | 0.83 |
ENSMUST00000151747.8
|
Afg1l
|
AFG1 like ATPase |
| chr19_+_3817396 | 0.77 |
ENSMUST00000052699.13
ENSMUST00000113974.11 ENSMUST00000113972.9 ENSMUST00000113973.8 ENSMUST00000113977.9 ENSMUST00000113968.9 |
Kmt5b
|
lysine methyltransferase 5B |
| chr9_-_21996693 | 0.76 |
ENSMUST00000179422.8
ENSMUST00000098937.10 ENSMUST00000177967.2 ENSMUST00000180180.8 |
Ecsit
|
ECSIT signalling integrator |
| chr5_-_66309244 | 0.73 |
ENSMUST00000167950.8
|
Rbm47
|
RNA binding motif protein 47 |
| chr10_-_42354482 | 0.72 |
ENSMUST00000041024.15
|
Afg1l
|
AFG1 like ATPase |
| chr4_+_102617495 | 0.71 |
ENSMUST00000072481.12
ENSMUST00000156596.8 ENSMUST00000080728.13 ENSMUST00000106882.9 |
Sgip1
|
SH3-domain GRB2-like (endophilin) interacting protein 1 |
| chr4_+_97665843 | 0.70 |
ENSMUST00000075448.13
ENSMUST00000092532.13 |
Nfia
|
nuclear factor I/A |
| chr13_+_93440265 | 0.63 |
ENSMUST00000109494.8
|
Homer1
|
homer scaffolding protein 1 |
| chr10_+_67021509 | 0.63 |
ENSMUST00000173689.8
|
Jmjd1c
|
jumonji domain containing 1C |
| chr9_+_108269992 | 0.59 |
ENSMUST00000192995.6
|
1700102P08Rik
|
RIKEN cDNA 1700102P08 gene |
| chrX_-_141173330 | 0.57 |
ENSMUST00000112907.8
|
Acsl4
|
acyl-CoA synthetase long-chain family member 4 |
| chr4_+_97665992 | 0.57 |
ENSMUST00000107062.9
ENSMUST00000052018.12 ENSMUST00000107057.8 |
Nfia
|
nuclear factor I/A |
| chr10_+_27950809 | 0.56 |
ENSMUST00000166468.2
ENSMUST00000218359.2 ENSMUST00000218276.2 |
Ptprk
|
protein tyrosine phosphatase, receptor type, K |
| chr1_-_123972900 | 0.55 |
ENSMUST00000112603.4
|
Dpp10
|
dipeptidylpeptidase 10 |
| chr5_-_66308666 | 0.55 |
ENSMUST00000201561.4
|
Rbm47
|
RNA binding motif protein 47 |
| chr5_-_88823049 | 0.55 |
ENSMUST00000133532.8
ENSMUST00000150438.2 |
Grsf1
|
G-rich RNA sequence binding factor 1 |
| chr5_-_121665249 | 0.55 |
ENSMUST00000152270.8
|
Mapkapk5
|
MAP kinase-activated protein kinase 5 |
| chr13_+_93440572 | 0.54 |
ENSMUST00000109493.9
|
Homer1
|
homer scaffolding protein 1 |
| chrX_-_139501246 | 0.54 |
ENSMUST00000112996.9
|
Tsc22d3
|
TSC22 domain family, member 3 |
| chr16_+_8647959 | 0.53 |
ENSMUST00000023150.7
|
1810013L24Rik
|
RIKEN cDNA 1810013L24 gene |
| chr6_+_15727798 | 0.53 |
ENSMUST00000128849.3
|
Mdfic
|
MyoD family inhibitor domain containing |
| chr10_+_98943999 | 0.51 |
ENSMUST00000161240.4
|
Galnt4
|
polypeptide N-acetylgalactosaminyltransferase 4 |
| chr2_+_127180559 | 0.51 |
ENSMUST00000059839.9
|
Astl
|
astacin-like metalloendopeptidase (M12 family) |
| chr4_+_102617332 | 0.50 |
ENSMUST00000066824.14
|
Sgip1
|
SH3-domain GRB2-like (endophilin) interacting protein 1 |
| chr9_-_114393406 | 0.50 |
ENSMUST00000111816.3
|
Trim71
|
tripartite motif-containing 71 |
| chr15_+_25940859 | 0.49 |
ENSMUST00000226750.2
|
Retreg1
|
reticulophagy regulator 1 |
| chr1_+_83094481 | 0.48 |
ENSMUST00000027351.13
ENSMUST00000113437.9 ENSMUST00000186832.2 |
Ccl20
|
chemokine (C-C motif) ligand 20 |
| chr19_+_3818112 | 0.48 |
ENSMUST00000005518.16
ENSMUST00000237440.2 ENSMUST00000152935.8 ENSMUST00000176262.8 ENSMUST00000176407.8 ENSMUST00000176926.8 ENSMUST00000176512.8 |
Kmt5b
|
lysine methyltransferase 5B |
| chr15_-_76113692 | 0.47 |
ENSMUST00000074834.12
|
Plec
|
plectin |
| chr9_+_108270020 | 0.47 |
ENSMUST00000035234.6
|
1700102P08Rik
|
RIKEN cDNA 1700102P08 gene |
| chr11_-_94568228 | 0.46 |
ENSMUST00000116349.9
|
Xylt2
|
xylosyltransferase II |
| chr19_+_36532061 | 0.45 |
ENSMUST00000169036.9
ENSMUST00000047247.12 |
Hectd2
|
HECT domain E3 ubiquitin protein ligase 2 |
| chr5_-_88823472 | 0.45 |
ENSMUST00000113234.8
ENSMUST00000153565.8 |
Grsf1
|
G-rich RNA sequence binding factor 1 |
| chr2_+_105499280 | 0.44 |
ENSMUST00000142772.8
|
Pax6
|
paired box 6 |
| chrX_-_16683578 | 0.43 |
ENSMUST00000040820.13
|
Maob
|
monoamine oxidase B |
| chr2_+_105499233 | 0.43 |
ENSMUST00000111086.11
ENSMUST00000111087.10 |
Pax6
|
paired box 6 |
| chr15_+_25774070 | 0.43 |
ENSMUST00000125667.3
|
Myo10
|
myosin X |
| chr1_+_131838220 | 0.43 |
ENSMUST00000189946.7
|
Nucks1
|
nuclear casein kinase and cyclin-dependent kinase substrate 1 |
| chr9_+_55056648 | 0.42 |
ENSMUST00000121677.8
|
Ube2q2
|
ubiquitin-conjugating enzyme E2Q family member 2 |
| chr2_+_4980986 | 0.41 |
ENSMUST00000027978.7
ENSMUST00000195688.2 |
Ucma
|
upper zone of growth plate and cartilage matrix associated |
| chr10_-_41179167 | 0.41 |
ENSMUST00000043814.5
|
Fig4
|
FIG4 phosphoinositide 5-phosphatase |
| chr17_-_36220518 | 0.41 |
ENSMUST00000141132.2
|
Atat1
|
alpha tubulin acetyltransferase 1 |
| chr9_-_42372710 | 0.38 |
ENSMUST00000066179.14
|
Tbcel
|
tubulin folding cofactor E-like |
| chr9_+_118335294 | 0.37 |
ENSMUST00000084820.6
|
Golga4
|
golgi autoantigen, golgin subfamily a, 4 |
| chr19_-_40371242 | 0.36 |
ENSMUST00000224583.2
|
Sorbs1
|
sorbin and SH3 domain containing 1 |
| chr7_+_24310171 | 0.36 |
ENSMUST00000206422.2
|
Phldb3
|
pleckstrin homology like domain, family B, member 3 |
| chr2_-_26127360 | 0.35 |
ENSMUST00000036187.9
|
Qsox2
|
quiescin Q6 sulfhydryl oxidase 2 |
| chr4_+_150999019 | 0.35 |
ENSMUST00000135169.8
|
Tnfrsf9
|
tumor necrosis factor receptor superfamily, member 9 |
| chr15_-_50753792 | 0.34 |
ENSMUST00000185183.2
|
Trps1
|
transcriptional repressor GATA binding 1 |
| chr2_+_4980923 | 0.34 |
ENSMUST00000167607.8
ENSMUST00000115010.9 |
Ucma
|
upper zone of growth plate and cartilage matrix associated |
| chr2_-_129541753 | 0.34 |
ENSMUST00000028883.12
|
Pdyn
|
prodynorphin |
| chr9_-_106533279 | 0.34 |
ENSMUST00000023959.13
ENSMUST00000201681.2 |
Grm2
|
glutamate receptor, metabotropic 2 |
| chr10_-_23226684 | 0.34 |
ENSMUST00000220299.2
|
Eya4
|
EYA transcriptional coactivator and phosphatase 4 |
| chr13_+_74269554 | 0.33 |
ENSMUST00000036208.7
ENSMUST00000225423.2 ENSMUST00000221703.2 |
Slc9a3
|
solute carrier family 9 (sodium/hydrogen exchanger), member 3 |
| chr2_+_79538124 | 0.33 |
ENSMUST00000090760.9
ENSMUST00000040863.11 ENSMUST00000111780.3 |
Ppp1r1c
|
protein phosphatase 1, regulatory inhibitor subunit 1C |
| chr12_+_56742413 | 0.33 |
ENSMUST00000001538.10
|
Pax9
|
paired box 9 |
| chr6_-_124791259 | 0.32 |
ENSMUST00000172132.10
ENSMUST00000239432.2 |
Tpi1
|
triosephosphate isomerase 1 |
| chr11_-_97242842 | 0.32 |
ENSMUST00000093942.5
|
Gpr179
|
G protein-coupled receptor 179 |
| chr2_+_30485048 | 0.32 |
ENSMUST00000102853.4
|
Cstad
|
CSA-conditional, T cell activation-dependent protein |
| chr17_-_36220924 | 0.31 |
ENSMUST00000141662.8
ENSMUST00000056034.13 ENSMUST00000077494.13 ENSMUST00000149277.8 ENSMUST00000061052.12 |
Atat1
|
alpha tubulin acetyltransferase 1 |
| chr10_-_12745109 | 0.30 |
ENSMUST00000218635.2
|
Utrn
|
utrophin |
| chr19_+_3817953 | 0.30 |
ENSMUST00000113970.8
|
Kmt5b
|
lysine methyltransferase 5B |
| chr5_+_63806451 | 0.30 |
ENSMUST00000159584.3
|
Nwd2
|
NACHT and WD repeat domain containing 2 |
| chr2_+_58991182 | 0.29 |
ENSMUST00000168631.8
ENSMUST00000102754.11 ENSMUST00000123908.8 |
Pkp4
|
plakophilin 4 |
| chr4_+_33081505 | 0.29 |
ENSMUST00000147889.2
|
Gabrr2
|
gamma-aminobutyric acid (GABA) C receptor, subunit rho 2 |
| chr4_-_150998857 | 0.29 |
ENSMUST00000105675.8
|
Park7
|
Parkinson disease (autosomal recessive, early onset) 7 |
| chr2_-_120394288 | 0.29 |
ENSMUST00000055241.13
ENSMUST00000135625.8 |
Zfp106
|
zinc finger protein 106 |
| chr3_-_87934772 | 0.28 |
ENSMUST00000005014.9
|
Hapln2
|
hyaluronan and proteoglycan link protein 2 |
| chr2_+_143874979 | 0.28 |
ENSMUST00000037722.9
ENSMUST00000110032.2 |
Banf2
|
BANF family member 2 |
| chr7_-_141009346 | 0.27 |
ENSMUST00000124444.2
|
Cend1
|
cell cycle exit and neuronal differentiation 1 |
| chr12_+_31488208 | 0.26 |
ENSMUST00000001254.6
|
Slc26a3
|
solute carrier family 26, member 3 |
| chr19_+_46140942 | 0.26 |
ENSMUST00000026254.14
|
Gbf1
|
golgi-specific brefeldin A-resistance factor 1 |
| chr7_-_134100659 | 0.26 |
ENSMUST00000238294.2
|
D7Ertd443e
|
DNA segment, Chr 7, ERATO Doi 443, expressed |
| chr4_-_141351110 | 0.25 |
ENSMUST00000038661.8
|
Slc25a34
|
solute carrier family 25, member 34 |
| chr7_-_133203838 | 0.25 |
ENSMUST00000033275.4
|
Tex36
|
testis expressed 36 |
| chr2_-_24809583 | 0.24 |
ENSMUST00000046227.12
ENSMUST00000114432.9 ENSMUST00000091348.11 ENSMUST00000102938.10 ENSMUST00000150379.2 ENSMUST00000152161.8 ENSMUST00000147147.8 |
Ehmt1
|
euchromatic histone methyltransferase 1 |
| chr4_-_42853888 | 0.24 |
ENSMUST00000107979.2
|
Fam205a1
|
family with sequence similarity 205, member A1 |
| chr3_+_84832783 | 0.24 |
ENSMUST00000107675.8
|
Fbxw7
|
F-box and WD-40 domain protein 7 |
| chr5_-_90487583 | 0.23 |
ENSMUST00000197021.2
|
Ankrd17
|
ankyrin repeat domain 17 |
| chr19_-_32038838 | 0.23 |
ENSMUST00000096119.5
|
Asah2
|
N-acylsphingosine amidohydrolase 2 |
| chr4_+_41966058 | 0.22 |
ENSMUST00000108026.3
|
Fam205a4
|
family with sequence similarity 205, member A4 |
| chr13_-_17979675 | 0.22 |
ENSMUST00000223490.2
|
Cdk13
|
cyclin-dependent kinase 13 |
| chr9_+_98746796 | 0.22 |
ENSMUST00000167951.3
|
Prr23a3
|
proline rich 23A, member 3 |
| chr2_-_58990967 | 0.21 |
ENSMUST00000226455.2
ENSMUST00000077687.6 |
Ccdc148
|
coiled-coil domain containing 148 |
| chr5_+_16758538 | 0.21 |
ENSMUST00000199581.5
|
Hgf
|
hepatocyte growth factor |
| chr6_+_3993774 | 0.21 |
ENSMUST00000031673.7
|
Gngt1
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 1 |
| chr1_+_87501721 | 0.21 |
ENSMUST00000166259.8
ENSMUST00000172222.8 ENSMUST00000163606.8 |
Neu2
|
neuraminidase 2 |
| chr6_-_99073156 | 0.21 |
ENSMUST00000175886.8
|
Foxp1
|
forkhead box P1 |
| chr7_-_141009264 | 0.21 |
ENSMUST00000164387.2
ENSMUST00000137488.2 ENSMUST00000084436.10 |
Cend1
|
cell cycle exit and neuronal differentiation 1 |
| chr9_-_18585826 | 0.20 |
ENSMUST00000208663.2
|
Muc16
|
mucin 16 |
| chr18_+_86729184 | 0.19 |
ENSMUST00000068423.10
|
Cbln2
|
cerebellin 2 precursor protein |
| chr7_+_4122555 | 0.18 |
ENSMUST00000079415.12
|
Ttyh1
|
tweety family member 1 |
| chr2_+_75662511 | 0.18 |
ENSMUST00000047232.14
ENSMUST00000111952.9 |
Agps
|
alkylglycerone phosphate synthase |
| chr2_-_155787197 | 0.18 |
ENSMUST00000040162.3
|
Gdf5
|
growth differentiation factor 5 |
| chr4_+_42318272 | 0.17 |
ENSMUST00000178192.3
|
Fam205a3
|
family with sequence similarity 205, member A3 |
| chr10_-_20600442 | 0.17 |
ENSMUST00000170265.8
|
Pde7b
|
phosphodiesterase 7B |
| chr6_+_34897904 | 0.17 |
ENSMUST00000185102.2
ENSMUST00000114997.3 |
Stra8
|
stimulated by retinoic acid gene 8 |
| chr17_+_69746321 | 0.17 |
ENSMUST00000169935.2
|
Akain1
|
A kinase (PRKA) anchor inhibitor 1 |
| chr6_+_113674011 | 0.17 |
ENSMUST00000089018.11
|
Tatdn2
|
TatD DNase domain containing 2 |
| chr19_+_11724913 | 0.16 |
ENSMUST00000025585.4
|
Cblif
|
cobalamin binding intrinsic factor |
| chr10_-_20600797 | 0.16 |
ENSMUST00000020165.14
|
Pde7b
|
phosphodiesterase 7B |
| chr3_-_92407800 | 0.15 |
ENSMUST00000062129.2
|
Sprr4
|
small proline-rich protein 4 |
| chr2_+_75662806 | 0.15 |
ENSMUST00000175646.2
|
Agps
|
alkylglycerone phosphate synthase |
| chr11_-_100713348 | 0.15 |
ENSMUST00000107358.9
|
Stat5b
|
signal transducer and activator of transcription 5B |
| chr14_-_78326435 | 0.15 |
ENSMUST00000118785.3
ENSMUST00000066437.5 |
Fam216b
|
family with sequence similarity 216, member B |
| chrX_-_166638057 | 0.15 |
ENSMUST00000238211.2
|
Frmpd4
|
FERM and PDZ domain containing 4 |
| chr6_-_3487369 | 0.14 |
ENSMUST00000201607.4
|
Hepacam2
|
HEPACAM family member 2 |
| chr12_-_75596441 | 0.14 |
ENSMUST00000218716.2
|
Ppp2r5e
|
protein phosphatase 2, regulatory subunit B', epsilon |
| chr10_+_93983844 | 0.14 |
ENSMUST00000105290.9
|
Nr2c1
|
nuclear receptor subfamily 2, group C, member 1 |
| chr10_+_110756031 | 0.14 |
ENSMUST00000220409.2
ENSMUST00000219502.2 |
Csrp2
|
cysteine and glycine-rich protein 2 |
| chr7_-_89176294 | 0.13 |
ENSMUST00000207932.2
|
Prss23
|
protease, serine 23 |
| chr2_-_101451383 | 0.12 |
ENSMUST00000090513.11
|
Iftap
|
intraflagellar transport associated protein |
| chr11_-_59029996 | 0.12 |
ENSMUST00000219084.3
|
Obscn
|
obscurin, cytoskeletal calmodulin and titin-interacting RhoGEF |
| chr12_-_76842263 | 0.12 |
ENSMUST00000082431.6
|
Gpx2
|
glutathione peroxidase 2 |
| chr7_+_120234399 | 0.12 |
ENSMUST00000033176.7
ENSMUST00000208400.2 |
Uqcrc2
|
ubiquinol cytochrome c reductase core protein 2 |
| chr6_+_34897874 | 0.12 |
ENSMUST00000114999.8
|
Stra8
|
stimulated by retinoic acid gene 8 |
| chr9_-_13358049 | 0.11 |
ENSMUST00000167906.3
|
Gm17571
|
predicted gene, 17571 |
| chr9_-_32255533 | 0.11 |
ENSMUST00000216033.2
|
Kcnj5
|
potassium inwardly-rectifying channel, subfamily J, member 5 |
| chr11_+_70548513 | 0.11 |
ENSMUST00000134087.8
|
Eno3
|
enolase 3, beta muscle |
| chr4_-_82939330 | 0.11 |
ENSMUST00000071708.12
|
Frem1
|
Fras1 related extracellular matrix protein 1 |
| chr10_+_42378193 | 0.10 |
ENSMUST00000105499.2
|
Snx3
|
sorting nexin 3 |
| chr10_+_90665639 | 0.10 |
ENSMUST00000179337.9
|
Anks1b
|
ankyrin repeat and sterile alpha motif domain containing 1B |
| chrX_+_48552803 | 0.10 |
ENSMUST00000130558.8
|
Arhgap36
|
Rho GTPase activating protein 36 |
| chr2_+_121125918 | 0.09 |
ENSMUST00000110639.8
|
Map1a
|
microtubule-associated protein 1 A |
| chr7_-_65177444 | 0.09 |
ENSMUST00000206228.2
|
Tjp1
|
tight junction protein 1 |
| chr16_+_21644692 | 0.09 |
ENSMUST00000232240.2
|
Map3k13
|
mitogen-activated protein kinase kinase kinase 13 |
| chrX_+_159551009 | 0.09 |
ENSMUST00000033650.14
|
Rs1
|
retinoschisis (X-linked, juvenile) 1 (human) |
| chr8_-_13544478 | 0.08 |
ENSMUST00000033828.7
|
Gas6
|
growth arrest specific 6 |
| chr17_+_70828649 | 0.07 |
ENSMUST00000233283.2
|
Dlgap1
|
DLG associated protein 1 |
| chr6_+_92068361 | 0.07 |
ENSMUST00000113460.8
|
Nr2c2
|
nuclear receptor subfamily 2, group C, member 2 |
| chr7_-_102566717 | 0.07 |
ENSMUST00000214160.2
ENSMUST00000215773.2 |
Olfr571
|
olfactory receptor 571 |
| chr18_+_57275854 | 0.06 |
ENSMUST00000139892.2
|
Megf10
|
multiple EGF-like-domains 10 |
| chr9_-_32255556 | 0.06 |
ENSMUST00000214223.2
|
Kcnj5
|
potassium inwardly-rectifying channel, subfamily J, member 5 |
| chrX_+_6327790 | 0.05 |
ENSMUST00000143641.4
|
Shroom4
|
shroom family member 4 |
| chr10_-_40178182 | 0.05 |
ENSMUST00000099945.6
ENSMUST00000238953.2 ENSMUST00000238969.2 |
Amd1
|
S-adenosylmethionine decarboxylase 1 |
| chr13_+_118851214 | 0.05 |
ENSMUST00000022246.9
|
Fgf10
|
fibroblast growth factor 10 |
| chrX_+_159551171 | 0.04 |
ENSMUST00000112368.3
|
Rs1
|
retinoschisis (X-linked, juvenile) 1 (human) |
| chrX_+_7445806 | 0.04 |
ENSMUST00000234363.2
ENSMUST00000235116.2 ENSMUST00000115739.9 ENSMUST00000234574.2 ENSMUST00000115740.9 |
Foxp3
|
forkhead box P3 |
| chr19_-_40371016 | 0.04 |
ENSMUST00000225766.3
|
Sorbs1
|
sorbin and SH3 domain containing 1 |
| chr5_+_16758777 | 0.04 |
ENSMUST00000030683.8
|
Hgf
|
hepatocyte growth factor |
| chr3_+_95041399 | 0.03 |
ENSMUST00000066386.6
|
Lysmd1
|
LysM, putative peptidoglycan-binding, domain containing 1 |
| chr7_+_4122523 | 0.03 |
ENSMUST00000119661.8
ENSMUST00000129423.8 |
Ttyh1
|
tweety family member 1 |
| chr19_+_43770619 | 0.03 |
ENSMUST00000026208.6
|
Abcc2
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 2 |
| chr14_+_27598021 | 0.01 |
ENSMUST00000211684.2
ENSMUST00000210924.2 |
Erc2
|
ELKS/RAB6-interacting/CAST family member 2 |
| chr2_+_128907854 | 0.01 |
ENSMUST00000035812.14
|
Ttl
|
tubulin tyrosine ligase |
| chr6_+_96090127 | 0.01 |
ENSMUST00000122120.8
|
Tafa1
|
TAFA chemokine like family member 1 |
| chr13_+_95012107 | 0.00 |
ENSMUST00000022195.13
|
Otp
|
orthopedia homeobox |
Network of associatons between targets according to the STRING database.
First level regulatory network of Nr4a2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 5.4 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.6 | 2.4 | GO:0032915 | positive regulation of transforming growth factor beta2 production(GO:0032915) |
| 0.6 | 2.4 | GO:2000158 | positive regulation of ubiquitin-specific protease activity(GO:2000158) |
| 0.3 | 1.5 | GO:0035694 | mitochondrial protein catabolic process(GO:0035694) |
| 0.2 | 0.7 | GO:0071929 | alpha-tubulin acetylation(GO:0071929) |
| 0.2 | 0.9 | GO:0021905 | pancreatic A cell development(GO:0003322) forebrain-midbrain boundary formation(GO:0021905) somatic motor neuron fate commitment(GO:0021917) regulation of transcription from RNA polymerase II promoter involved in somatic motor neuron fate commitment(GO:0021918) sensory neuron migration(GO:1904937) |
| 0.2 | 2.4 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.2 | 0.5 | GO:0045360 | regulation of interleukin-1 biosynthetic process(GO:0045360) positive regulation of interleukin-1 biosynthetic process(GO:0045362) |
| 0.2 | 1.6 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.1 | 0.8 | GO:1901674 | histone H3-K27 acetylation(GO:0043974) spongiotrophoblast differentiation(GO:0060708) regulation of histone H3-K27 acetylation(GO:1901674) |
| 0.1 | 3.4 | GO:0036295 | cellular response to increased oxygen levels(GO:0036295) |
| 0.1 | 1.6 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 0.1 | 0.5 | GO:0021933 | radial glia guided migration of cerebellar granule cell(GO:0021933) negative regulation of cerebellar granule cell precursor proliferation(GO:0021941) |
| 0.1 | 1.5 | GO:0070601 | centromeric sister chromatid cohesion(GO:0070601) |
| 0.1 | 0.6 | GO:0032304 | negative regulation of icosanoid secretion(GO:0032304) |
| 0.1 | 0.4 | GO:0019046 | release from viral latency(GO:0019046) regulation of DNA strand elongation(GO:0060382) |
| 0.1 | 0.3 | GO:0046166 | alditol catabolic process(GO:0019405) glyceraldehyde-3-phosphate biosynthetic process(GO:0046166) |
| 0.1 | 0.5 | GO:2000360 | negative regulation of binding of sperm to zona pellucida(GO:2000360) |
| 0.1 | 0.4 | GO:0014063 | negative regulation of serotonin secretion(GO:0014063) |
| 0.1 | 0.3 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.1 | 1.2 | GO:2000253 | positive regulation of feeding behavior(GO:2000253) |
| 0.1 | 0.3 | GO:1903198 | enzyme active site formation via L-cysteine sulfinic acid(GO:0018323) primary alcohol biosynthetic process(GO:0034309) cellular response to glyoxal(GO:0036471) glycolate biosynthetic process(GO:0046295) negative regulation of TRAIL-activated apoptotic signaling pathway(GO:1903122) regulation of pyrroline-5-carboxylate reductase activity(GO:1903167) positive regulation of pyrroline-5-carboxylate reductase activity(GO:1903168) regulation of tyrosine 3-monooxygenase activity(GO:1903176) positive regulation of tyrosine 3-monooxygenase activity(GO:1903178) L-dopa metabolic process(GO:1903184) L-dopa biosynthetic process(GO:1903185) glyoxal metabolic process(GO:1903189) regulation of L-dopa biosynthetic process(GO:1903195) positive regulation of L-dopa biosynthetic process(GO:1903197) regulation of L-dopa decarboxylase activity(GO:1903198) positive regulation of L-dopa decarboxylase activity(GO:1903200) positive regulation of cellular amino acid biosynthetic process(GO:2000284) |
| 0.1 | 0.3 | GO:0030210 | heparin biosynthetic process(GO:0030210) |
| 0.1 | 0.8 | GO:1902510 | regulation of apoptotic DNA fragmentation(GO:1902510) |
| 0.1 | 1.2 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.1 | 1.3 | GO:0072189 | ureter development(GO:0072189) |
| 0.1 | 0.6 | GO:0030718 | germ-line stem cell population maintenance(GO:0030718) |
| 0.1 | 0.4 | GO:2001180 | negative regulation of interleukin-10 secretion(GO:2001180) |
| 0.0 | 0.5 | GO:0090400 | stress-induced premature senescence(GO:0090400) |
| 0.0 | 0.5 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.0 | 0.2 | GO:1903378 | positive regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903378) |
| 0.0 | 0.4 | GO:0031642 | negative regulation of myelination(GO:0031642) |
| 0.0 | 0.3 | GO:0007527 | adult somatic muscle development(GO:0007527) |
| 0.0 | 0.2 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.0 | 0.3 | GO:0008065 | establishment of blood-nerve barrier(GO:0008065) |
| 0.0 | 0.6 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 0.0 | 0.2 | GO:0045409 | negative regulation of interleukin-6 biosynthetic process(GO:0045409) |
| 0.0 | 0.1 | GO:0070676 | intralumenal vesicle formation(GO:0070676) negative regulation of early endosome to late endosome transport(GO:2000642) |
| 0.0 | 0.1 | GO:0016062 | adaptation of rhodopsin mediated signaling(GO:0016062) light adaption(GO:0036367) |
| 0.0 | 0.3 | GO:0048133 | germ-line stem cell division(GO:0042078) male germ-line stem cell asymmetric division(GO:0048133) germline stem cell asymmetric division(GO:0098728) |
| 0.0 | 0.5 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.5 | GO:0045974 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.0 | 0.1 | GO:0009608 | response to symbiont(GO:0009608) response to symbiotic bacterium(GO:0009609) |
| 0.0 | 0.3 | GO:0098719 | sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.0 | 2.0 | GO:0042773 | ATP synthesis coupled electron transport(GO:0042773) |
| 0.0 | 1.3 | GO:1901522 | positive regulation of transcription from RNA polymerase II promoter involved in cellular response to chemical stimulus(GO:1901522) |
| 0.0 | 0.2 | GO:0006689 | ganglioside catabolic process(GO:0006689) oligosaccharide catabolic process(GO:0009313) |
| 0.0 | 0.0 | GO:0050787 | detoxification of mercury ion(GO:0050787) |
| 0.0 | 0.2 | GO:0060591 | chondroblast differentiation(GO:0060591) |
| 0.0 | 0.2 | GO:0072619 | interleukin-21 production(GO:0032625) interleukin-21 secretion(GO:0072619) |
| 0.0 | 0.3 | GO:0060665 | regulation of branching involved in salivary gland morphogenesis by mesenchymal-epithelial signaling(GO:0060665) |
| 0.0 | 0.3 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.0 | 0.1 | GO:0046544 | development of secondary male sexual characteristics(GO:0046544) |
| 0.0 | 0.1 | GO:0039663 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 0.0 | 0.2 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.0 | 0.3 | GO:0051454 | pH elevation(GO:0045852) intracellular pH elevation(GO:0051454) |
| 0.0 | 0.5 | GO:0050434 | positive regulation of viral transcription(GO:0050434) |
| 0.0 | 0.1 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.0 | 0.1 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.0 | 0.0 | GO:0002851 | positive regulation of tolerance induction dependent upon immune response(GO:0002654) regulation of peripheral tolerance induction(GO:0002658) positive regulation of peripheral tolerance induction(GO:0002660) regulation of peripheral T cell tolerance induction(GO:0002849) positive regulation of peripheral T cell tolerance induction(GO:0002851) |
| 0.0 | 1.1 | GO:0016126 | sterol biosynthetic process(GO:0016126) |
| 0.0 | 0.2 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.0 | 0.8 | GO:0045668 | negative regulation of osteoblast differentiation(GO:0045668) |
| 0.0 | 1.0 | GO:1903078 | positive regulation of protein localization to plasma membrane(GO:1903078) |
| 0.0 | 0.1 | GO:0048386 | positive regulation of retinoic acid receptor signaling pathway(GO:0048386) |
| 0.0 | 0.8 | GO:0001707 | mesoderm formation(GO:0001707) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 3.4 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.2 | 0.5 | GO:0032156 | septin cytoskeleton(GO:0032156) |
| 0.2 | 0.5 | GO:0060473 | cortical granule(GO:0060473) |
| 0.1 | 1.0 | GO:0031415 | NatA complex(GO:0031415) |
| 0.1 | 0.2 | GO:0002945 | cyclin K-CDK13 complex(GO:0002945) |
| 0.1 | 5.2 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.1 | 0.4 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.1 | 0.2 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.1 | 1.2 | GO:0030122 | AP-2 adaptor complex(GO:0030122) |
| 0.1 | 0.7 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.1 | 0.2 | GO:0030868 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.0 | 2.4 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 0.6 | GO:0044233 | ER-mitochondrion membrane contact site(GO:0044233) |
| 0.0 | 1.6 | GO:0000780 | condensed nuclear chromosome, centromeric region(GO:0000780) |
| 0.0 | 2.3 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.1 | GO:0005863 | striated muscle myosin thick filament(GO:0005863) |
| 0.0 | 1.2 | GO:0043034 | costamere(GO:0043034) |
| 0.0 | 1.0 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 0.5 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.0 | 0.8 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 0.3 | GO:0044292 | dendrite terminus(GO:0044292) |
| 0.0 | 0.3 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) contractile ring(GO:0070938) glycoprotein complex(GO:0090665) |
| 0.0 | 0.3 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.1 | GO:0005750 | mitochondrial respiratory chain complex III(GO:0005750) |
| 0.0 | 0.8 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 3.1 | GO:0047635 | L-alanine:2-oxoglutarate aminotransferase activity(GO:0004021) alanine-oxo-acid transaminase activity(GO:0047635) |
| 0.6 | 2.4 | GO:0035800 | deubiquitinase activator activity(GO:0035800) |
| 0.4 | 1.6 | GO:0004307 | ethanolaminephosphotransferase activity(GO:0004307) |
| 0.3 | 2.0 | GO:0008321 | Ral guanyl-nucleotide exchange factor activity(GO:0008321) |
| 0.2 | 2.4 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.2 | 1.6 | GO:0042799 | histone methyltransferase activity (H4-K20 specific)(GO:0042799) |
| 0.2 | 0.8 | GO:0005146 | leukemia inhibitory factor receptor binding(GO:0005146) |
| 0.2 | 0.5 | GO:0070002 | glutamic-type peptidase activity(GO:0070002) |
| 0.2 | 0.8 | GO:0004174 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 0.2 | 0.5 | GO:0030158 | protein xylosyltransferase activity(GO:0030158) |
| 0.1 | 1.3 | GO:0004886 | 9-cis retinoic acid receptor activity(GO:0004886) |
| 0.1 | 0.9 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.1 | 3.4 | GO:0046961 | proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.1 | 1.2 | GO:0031802 | type 5 metabotropic glutamate receptor binding(GO:0031802) |
| 0.1 | 0.4 | GO:0043812 | phosphatidylinositol-4-phosphate phosphatase activity(GO:0043812) |
| 0.1 | 0.6 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.1 | 0.3 | GO:0036478 | tyrosine 3-monooxygenase activator activity(GO:0036470) L-dopa decarboxylase activator activity(GO:0036478) cupric ion binding(GO:1903135) |
| 0.1 | 0.4 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.1 | 0.7 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.1 | 2.0 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 1.5 | GO:0010485 | H4 histone acetyltransferase activity(GO:0010485) |
| 0.1 | 0.5 | GO:0043426 | MRF binding(GO:0043426) |
| 0.1 | 0.2 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.0 | 0.4 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.0 | 0.2 | GO:0052795 | exo-alpha-sialidase activity(GO:0004308) alpha-sialidase activity(GO:0016997) exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.0 | 0.3 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.0 | 0.2 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.0 | 0.6 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.0 | 0.3 | GO:0001640 | adenylate cyclase inhibiting G-protein coupled glutamate receptor activity(GO:0001640) |
| 0.0 | 0.3 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.0 | 0.5 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.0 | 0.9 | GO:0070410 | AT DNA binding(GO:0003680) co-SMAD binding(GO:0070410) |
| 0.0 | 0.3 | GO:0016861 | intramolecular oxidoreductase activity, interconverting aldoses and ketoses(GO:0016861) |
| 0.0 | 0.2 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.0 | 0.6 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.0 | 0.6 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 0.2 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.0 | 0.5 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.0 | 0.5 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.0 | 0.5 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 1.1 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.0 | 0.6 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.0 | 0.2 | GO:0036122 | BMP binding(GO:0036122) |
| 0.0 | 0.1 | GO:0086089 | voltage-gated potassium channel activity involved in atrial cardiac muscle cell action potential repolarization(GO:0086089) |
| 0.0 | 0.3 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.0 | 0.6 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.0 | 0.5 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.0 | 0.1 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.0 | 0.3 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.4 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.0 | 1.3 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 1.3 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 0.8 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 0.1 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.0 | 1.5 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 0.8 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.0 | 0.6 | ST P38 MAPK PATHWAY | p38 MAPK Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.4 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.1 | 3.1 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.1 | 3.4 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.1 | 2.4 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.1 | 2.0 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.9 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.0 | 1.1 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.2 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.0 | 2.1 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.0 | 0.3 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.0 | 0.3 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.0 | 0.4 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 0.3 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.0 | 1.3 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |