Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
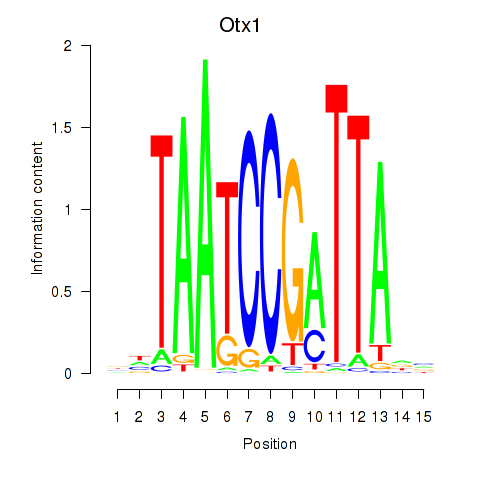
Results for Otx1
Z-value: 0.68
Transcription factors associated with Otx1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Otx1
|
ENSMUSG00000005917.16 | orthodenticle homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Otx1 | mm39_v1_chr11_-_21951605_21951631 | 0.28 | 1.0e-01 | Click! |
Activity profile of Otx1 motif
Sorted Z-values of Otx1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_+_3923086 | 4.61 |
ENSMUST00000117291.8
ENSMUST00000120585.8 ENSMUST00000043735.8 |
Mthfd1l
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1-like |
| chr8_+_117822593 | 3.95 |
ENSMUST00000034308.16
ENSMUST00000176860.2 |
Bco1
|
beta-carotene oxygenase 1 |
| chr14_-_87378641 | 3.77 |
ENSMUST00000168889.3
ENSMUST00000022599.14 |
Diaph3
|
diaphanous related formin 3 |
| chr11_+_94827050 | 3.72 |
ENSMUST00000001547.8
|
Col1a1
|
collagen, type I, alpha 1 |
| chr2_+_91376650 | 3.25 |
ENSMUST00000099716.11
ENSMUST00000046769.16 ENSMUST00000111337.3 |
Ckap5
|
cytoskeleton associated protein 5 |
| chr17_+_36172210 | 2.85 |
ENSMUST00000074259.15
ENSMUST00000174873.2 |
Nrm
|
nurim (nuclear envelope membrane protein) |
| chr11_+_78237492 | 2.48 |
ENSMUST00000100755.4
|
Unc119
|
unc-119 lipid binding chaperone |
| chr7_-_19410749 | 2.30 |
ENSMUST00000003074.16
|
Apoc2
|
apolipoprotein C-II |
| chr9_+_108356935 | 2.11 |
ENSMUST00000194147.2
ENSMUST00000065014.10 ENSMUST00000195483.6 ENSMUST00000195058.2 |
Lamb2
|
laminin, beta 2 |
| chr10_-_93725619 | 1.94 |
ENSMUST00000181091.8
ENSMUST00000181217.8 ENSMUST00000047910.15 ENSMUST00000180688.2 |
Metap2
|
methionine aminopeptidase 2 |
| chr13_-_61084358 | 1.85 |
ENSMUST00000225859.2
ENSMUST00000225167.2 ENSMUST00000021880.10 |
Gm49391
Ctla2a
|
predicted gene, 49391 cytotoxic T lymphocyte-associated protein 2 alpha |
| chr15_+_80017315 | 1.76 |
ENSMUST00000023050.9
|
Tab1
|
TGF-beta activated kinase 1/MAP3K7 binding protein 1 |
| chrX_-_165992145 | 1.67 |
ENSMUST00000112176.8
|
Tmsb4x
|
thymosin, beta 4, X chromosome |
| chr8_+_55024446 | 1.67 |
ENSMUST00000239166.2
ENSMUST00000239106.2 ENSMUST00000239152.2 |
Asb5
|
ankyrin repeat and SOCs box-containing 5 |
| chr17_+_36172235 | 1.55 |
ENSMUST00000172931.2
|
Nrm
|
nurim (nuclear envelope membrane protein) |
| chr16_+_49620883 | 1.50 |
ENSMUST00000229640.2
|
Cd47
|
CD47 antigen (Rh-related antigen, integrin-associated signal transducer) |
| chr1_-_165535617 | 1.46 |
ENSMUST00000040357.11
|
Rcsd1
|
RCSD domain containing 1 |
| chr11_-_106889291 | 1.45 |
ENSMUST00000124541.8
|
Kpna2
|
karyopherin (importin) alpha 2 |
| chr9_-_62895197 | 1.42 |
ENSMUST00000216209.2
|
Pias1
|
protein inhibitor of activated STAT 1 |
| chr5_-_104261556 | 1.40 |
ENSMUST00000031249.8
|
Sparcl1
|
SPARC-like 1 |
| chr4_-_11965691 | 1.39 |
ENSMUST00000108301.8
ENSMUST00000095144.10 ENSMUST00000108302.8 |
Pdp1
|
pyruvate dehyrogenase phosphatase catalytic subunit 1 |
| chr9_+_117888124 | 1.38 |
ENSMUST00000123690.2
|
Azi2
|
5-azacytidine induced gene 2 |
| chr5_-_104261285 | 1.33 |
ENSMUST00000199947.2
|
Sparcl1
|
SPARC-like 1 |
| chr1_-_85664246 | 1.31 |
ENSMUST00000064788.14
|
A630001G21Rik
|
RIKEN cDNA A630001G21 gene |
| chr5_-_65117375 | 1.26 |
ENSMUST00000062315.7
ENSMUST00000239485.2 ENSMUST00000201307.3 |
Tlr6
|
toll-like receptor 6 |
| chr12_-_26506422 | 1.17 |
ENSMUST00000020970.10
|
Rsad2
|
radical S-adenosyl methionine domain containing 2 |
| chr5_-_5799315 | 1.15 |
ENSMUST00000015796.9
|
Steap1
|
six transmembrane epithelial antigen of the prostate 1 |
| chr11_+_115455260 | 1.14 |
ENSMUST00000021085.11
|
Nup85
|
nucleoporin 85 |
| chr6_-_68609426 | 1.13 |
ENSMUST00000103328.3
|
Igkv10-96
|
immunoglobulin kappa variable 10-96 |
| chr9_-_53521585 | 1.12 |
ENSMUST00000034547.6
|
Acat1
|
acetyl-Coenzyme A acetyltransferase 1 |
| chr17_+_47922497 | 1.12 |
ENSMUST00000024778.3
|
Med20
|
mediator complex subunit 20 |
| chr10_-_111829393 | 1.10 |
ENSMUST00000161870.3
|
Glipr1
|
GLI pathogenesis-related 1 (glioma) |
| chr6_-_136805184 | 1.06 |
ENSMUST00000116514.4
|
Wbp11
|
WW domain binding protein 11 |
| chr18_+_24087725 | 1.00 |
ENSMUST00000225682.2
ENSMUST00000060762.6 |
Zfp397
|
zinc finger protein 397 |
| chr4_-_137512682 | 0.98 |
ENSMUST00000133473.2
|
Alpl
|
alkaline phosphatase, liver/bone/kidney |
| chrX_+_36390430 | 0.95 |
ENSMUST00000016553.5
|
Nkap
|
NFKB activating protein |
| chr3_+_101284391 | 0.95 |
ENSMUST00000043983.11
|
Igsf3
|
immunoglobulin superfamily, member 3 |
| chr12_-_114621406 | 0.94 |
ENSMUST00000192077.2
|
Ighv1-15
|
immunoglobulin heavy variable 1-15 |
| chr12_-_13299197 | 0.89 |
ENSMUST00000071103.10
|
Ddx1
|
DEAD box helicase 1 |
| chr2_-_153067297 | 0.89 |
ENSMUST00000099194.4
|
Tspyl3
|
TSPY-like 3 |
| chr18_-_68562385 | 0.88 |
ENSMUST00000052347.8
|
Mc2r
|
melanocortin 2 receptor |
| chr1_-_53391778 | 0.88 |
ENSMUST00000236737.2
ENSMUST00000027264.10 ENSMUST00000123519.9 |
Gm50478
Asnsd1
|
predicted gene, 50478 asparagine synthetase domain containing 1 |
| chr11_+_3913970 | 0.86 |
ENSMUST00000109985.8
ENSMUST00000020705.5 |
Pes1
|
pescadillo ribosomal biogenesis factor 1 |
| chr1_-_118239146 | 0.85 |
ENSMUST00000027623.9
|
Tsn
|
translin |
| chr2_-_12306722 | 0.82 |
ENSMUST00000028106.11
|
Itga8
|
integrin alpha 8 |
| chr13_+_21363602 | 0.77 |
ENSMUST00000222544.2
|
Trim27
|
tripartite motif-containing 27 |
| chr16_+_91444730 | 0.76 |
ENSMUST00000119368.8
ENSMUST00000114037.9 ENSMUST00000114036.9 ENSMUST00000122302.8 |
Son
|
Son DNA binding protein |
| chr15_+_102391614 | 0.75 |
ENSMUST00000229432.2
|
Pcbp2
|
poly(rC) binding protein 2 |
| chr1_+_118249558 | 0.74 |
ENSMUST00000027626.13
ENSMUST00000112688.10 |
Nifk
|
nucleolar protein interacting with the FHA domain of MKI67 |
| chr5_+_25427860 | 0.71 |
ENSMUST00000045737.14
|
Galnt11
|
polypeptide N-acetylgalactosaminyltransferase 11 |
| chr12_+_70499869 | 0.70 |
ENSMUST00000021471.13
|
Tmx1
|
thioredoxin-related transmembrane protein 1 |
| chr13_-_61045252 | 0.69 |
ENSMUST00000021884.10
|
Ctla2b
|
cytotoxic T lymphocyte-associated protein 2 beta |
| chr2_-_151822114 | 0.68 |
ENSMUST00000062047.6
|
Fam110a
|
family with sequence similarity 110, member A |
| chr19_-_10847121 | 0.67 |
ENSMUST00000120524.2
ENSMUST00000025645.14 |
Tmem132a
|
transmembrane protein 132A |
| chr17_-_47922374 | 0.66 |
ENSMUST00000024783.9
|
Bysl
|
bystin-like |
| chr19_+_5074070 | 0.63 |
ENSMUST00000025826.7
ENSMUST00000237371.2 ENSMUST00000235416.2 |
Slc29a2
|
solute carrier family 29 (nucleoside transporters), member 2 |
| chr9_-_20432562 | 0.61 |
ENSMUST00000215908.2
ENSMUST00000068296.8 ENSMUST00000174462.8 ENSMUST00000213418.2 |
Zfp266
|
zinc finger protein 266 |
| chr1_+_186947683 | 0.60 |
ENSMUST00000065573.14
ENSMUST00000110943.9 ENSMUST00000044812.12 |
Gpatch2
|
G patch domain containing 2 |
| chr8_-_46604742 | 0.57 |
ENSMUST00000041582.15
|
Snx25
|
sorting nexin 25 |
| chr1_+_131838294 | 0.56 |
ENSMUST00000062264.8
|
Nucks1
|
nuclear casein kinase and cyclin-dependent kinase substrate 1 |
| chr6_+_58810674 | 0.55 |
ENSMUST00000041401.11
|
Herc3
|
hect domain and RLD 3 |
| chr13_+_36142822 | 0.55 |
ENSMUST00000225537.2
|
Ppp1r3g
|
protein phosphatase 1, regulatory subunit 3G |
| chr10_-_39009844 | 0.55 |
ENSMUST00000134279.8
ENSMUST00000139743.8 ENSMUST00000149949.8 ENSMUST00000124941.8 ENSMUST00000125042.8 ENSMUST00000063204.9 |
Fam229b
|
family with sequence similarity 229, member B |
| chr13_-_63579497 | 0.53 |
ENSMUST00000160931.2
ENSMUST00000099444.10 ENSMUST00000220684.2 ENSMUST00000161977.8 ENSMUST00000163091.8 |
Fancc
|
Fanconi anemia, complementation group C |
| chr7_-_4066154 | 0.52 |
ENSMUST00000086401.10
ENSMUST00000068865.13 |
Lair1
|
leukocyte-associated Ig-like receptor 1 |
| chr11_-_59678462 | 0.51 |
ENSMUST00000125307.2
|
Pld6
|
phospholipase D family, member 6 |
| chr10_-_127047396 | 0.48 |
ENSMUST00000013970.9
|
Pip4k2c
|
phosphatidylinositol-5-phosphate 4-kinase, type II, gamma |
| chr13_+_21364069 | 0.48 |
ENSMUST00000021761.13
|
Trim27
|
tripartite motif-containing 27 |
| chr6_+_68657317 | 0.48 |
ENSMUST00000198735.2
|
Igkv10-95
|
immunoglobulin kappa variable 10-95 |
| chr2_+_154498917 | 0.47 |
ENSMUST00000044277.10
|
Chmp4b
|
charged multivesicular body protein 4B |
| chr13_-_61045212 | 0.47 |
ENSMUST00000171347.9
|
Ctla2b
|
cytotoxic T lymphocyte-associated protein 2 beta |
| chr8_-_94825556 | 0.46 |
ENSMUST00000034206.6
|
Bbs2
|
Bardet-Biedl syndrome 2 (human) |
| chr13_+_21364330 | 0.46 |
ENSMUST00000223065.2
|
Trim27
|
tripartite motif-containing 27 |
| chr19_+_12257218 | 0.45 |
ENSMUST00000207186.4
ENSMUST00000207915.2 |
Olfr1434
|
olfactory receptor 1434 |
| chr4_-_123644091 | 0.44 |
ENSMUST00000102636.4
|
Akirin1
|
akirin 1 |
| chr5_+_53713137 | 0.44 |
ENSMUST00000087360.9
|
Rbpj
|
recombination signal binding protein for immunoglobulin kappa J region |
| chr6_+_67838100 | 0.44 |
ENSMUST00000200586.2
ENSMUST00000103309.3 |
Igkv17-127
|
immunoglobulin kappa variable 17-127 |
| chr5_-_69748126 | 0.40 |
ENSMUST00000166298.8
|
Gnpda2
|
glucosamine-6-phosphate deaminase 2 |
| chr4_-_126094910 | 0.39 |
ENSMUST00000136157.8
|
Thrap3
|
thyroid hormone receptor associated protein 3 |
| chr2_+_112096154 | 0.39 |
ENSMUST00000110991.9
|
Slc12a6
|
solute carrier family 12, member 6 |
| chr16_-_63684477 | 0.39 |
ENSMUST00000232654.2
ENSMUST00000064405.8 |
Epha3
|
Eph receptor A3 |
| chr6_+_70332836 | 0.37 |
ENSMUST00000103390.3
|
Igkv8-18
|
immunoglobulin kappa variable 8-18 |
| chr5_+_31454787 | 0.36 |
ENSMUST00000201166.4
ENSMUST00000072228.9 |
Gckr
|
glucokinase regulatory protein |
| chr7_+_27537982 | 0.33 |
ENSMUST00000205701.2
|
Zfp59
|
zinc finger protein 59 |
| chr17_+_28075415 | 0.33 |
ENSMUST00000114849.3
|
Uhrf1bp1
|
UHRF1 (ICBP90) binding protein 1 |
| chr19_-_42768374 | 0.33 |
ENSMUST00000069298.13
ENSMUST00000160455.8 ENSMUST00000162004.8 |
Hps1
|
HPS1, biogenesis of lysosomal organelles complex 3 subunit 1 |
| chr16_-_18885809 | 0.33 |
ENSMUST00000200211.2
|
Iglj3
|
immunoglobulin lambda joining 3 |
| chr7_-_4066125 | 0.32 |
ENSMUST00000108600.9
ENSMUST00000205296.2 |
Lair1
|
leukocyte-associated Ig-like receptor 1 |
| chr12_-_98225676 | 0.31 |
ENSMUST00000021390.9
|
Galc
|
galactosylceramidase |
| chr17_+_21031817 | 0.31 |
ENSMUST00000232810.2
ENSMUST00000233712.2 ENSMUST00000232852.2 |
Vmn1r229
|
vomeronasal 1 receptor 229 |
| chr3_+_95018324 | 0.31 |
ENSMUST00000009102.9
|
Vps72
|
vacuolar protein sorting 72 |
| chr13_+_41169740 | 0.31 |
ENSMUST00000021790.7
|
Tmem14c
|
transmembrane protein 14C |
| chr7_+_103628383 | 0.30 |
ENSMUST00000098185.2
|
Olfr635
|
olfactory receptor 635 |
| chr7_-_4066194 | 0.30 |
ENSMUST00000086400.13
|
Lair1
|
leukocyte-associated Ig-like receptor 1 |
| chr4_+_74160705 | 0.29 |
ENSMUST00000077851.10
|
Kdm4c
|
lysine (K)-specific demethylase 4C |
| chr2_-_37312881 | 0.29 |
ENSMUST00000112936.4
ENSMUST00000112934.8 |
Rc3h2
|
ring finger and CCCH-type zinc finger domains 2 |
| chr7_-_7250327 | 0.29 |
ENSMUST00000170922.2
|
Vmn2r29
|
vomeronasal 2, receptor 29 |
| chr1_+_131838220 | 0.28 |
ENSMUST00000189946.7
|
Nucks1
|
nuclear casein kinase and cyclin-dependent kinase substrate 1 |
| chr2_-_156022054 | 0.28 |
ENSMUST00000126992.8
ENSMUST00000146288.8 ENSMUST00000029149.13 ENSMUST00000109587.9 ENSMUST00000109584.8 |
Rbm39
|
RNA binding motif protein 39 |
| chr7_-_37970734 | 0.28 |
ENSMUST00000032585.8
|
Pop4
|
processing of precursor 4, ribonuclease P/MRP family, (S. cerevisiae) |
| chr6_+_108771840 | 0.27 |
ENSMUST00000204483.2
|
Arl8b
|
ADP-ribosylation factor-like 8B |
| chr1_+_186947934 | 0.25 |
ENSMUST00000160471.8
|
Gpatch2
|
G patch domain containing 2 |
| chr1_-_186947651 | 0.25 |
ENSMUST00000183819.8
|
Spata17
|
spermatogenesis associated 17 |
| chr2_+_85715984 | 0.24 |
ENSMUST00000213441.3
|
Olfr1023
|
olfactory receptor 1023 |
| chr15_+_78819119 | 0.24 |
ENSMUST00000138880.9
ENSMUST00000041164.4 |
Nol12
|
nucleolar protein 12 |
| chr2_-_89273344 | 0.24 |
ENSMUST00000216123.2
|
Olfr1240
|
olfactory receptor 1240 |
| chr3_+_66892979 | 0.24 |
ENSMUST00000162362.8
ENSMUST00000065074.14 ENSMUST00000065047.13 |
Rsrc1
|
arginine/serine-rich coiled-coil 1 |
| chr13_-_21934675 | 0.24 |
ENSMUST00000102983.2
|
H4c12
|
H4 clustered histone 12 |
| chr12_-_114012399 | 0.22 |
ENSMUST00000103468.3
|
Ighv11-2
|
immunoglobulin heavy variable V11-2 |
| chr12_-_65120674 | 0.21 |
ENSMUST00000220983.2
ENSMUST00000220730.2 ENSMUST00000021332.10 |
Fkbp3
|
FK506 binding protein 3 |
| chr8_-_96615138 | 0.21 |
ENSMUST00000034097.8
|
Got2
|
glutamatic-oxaloacetic transaminase 2, mitochondrial |
| chr8_-_55171699 | 0.20 |
ENSMUST00000144711.9
|
Wdr17
|
WD repeat domain 17 |
| chr19_-_7017295 | 0.18 |
ENSMUST00000025918.9
|
Stip1
|
stress-induced phosphoprotein 1 |
| chr5_-_100521343 | 0.17 |
ENSMUST00000182433.8
|
Sec31a
|
Sec31 homolog A (S. cerevisiae) |
| chr18_-_70274639 | 0.16 |
ENSMUST00000121693.8
|
Rab27b
|
RAB27B, member RAS oncogene family |
| chr3_+_66893031 | 0.15 |
ENSMUST00000046542.13
ENSMUST00000162693.8 |
Rsrc1
|
arginine/serine-rich coiled-coil 1 |
| chr11_-_116165024 | 0.14 |
ENSMUST00000021133.16
|
Srp68
|
signal recognition particle 68 |
| chr2_+_112114906 | 0.11 |
ENSMUST00000053666.8
|
Slc12a6
|
solute carrier family 12, member 6 |
| chr19_-_41921676 | 0.11 |
ENSMUST00000075280.12
ENSMUST00000112123.4 |
Exosc1
|
exosome component 1 |
| chr7_+_86444235 | 0.11 |
ENSMUST00000233099.2
ENSMUST00000164996.2 |
Vmn2r77
|
vomeronasal 2, receptor 77 |
| chr1_-_186947618 | 0.10 |
ENSMUST00000110945.4
ENSMUST00000183931.8 ENSMUST00000027908.13 |
Spata17
|
spermatogenesis associated 17 |
| chr7_-_7483017 | 0.09 |
ENSMUST00000094866.7
|
Vmn2r32
|
vomeronasal 2, receptor 32 |
| chr10_-_7423341 | 0.09 |
ENSMUST00000169796.4
ENSMUST00000218087.2 |
Ulbp1
|
UL16 binding protein 1 |
| chr6_-_67919524 | 0.08 |
ENSMUST00000196768.2
|
Igkv9-124
|
immunoglobulin kappa chain variable 9-124 |
| chr11_-_116164928 | 0.08 |
ENSMUST00000106425.4
|
Srp68
|
signal recognition particle 68 |
| chr2_-_87881473 | 0.08 |
ENSMUST00000183862.3
|
Olfr1162
|
olfactory receptor 1162 |
| chr14_-_75185281 | 0.07 |
ENSMUST00000088970.7
ENSMUST00000228252.2 |
Lrch1
|
leucine-rich repeats and calponin homology (CH) domain containing 1 |
| chr13_+_18901459 | 0.05 |
ENSMUST00000072961.6
|
Vps41
|
VPS41 HOPS complex subunit |
| chr7_-_7822866 | 0.05 |
ENSMUST00000169683.2
|
Vmn2r35
|
vomeronasal 2, receptor 35 |
| chr7_+_135207505 | 0.04 |
ENSMUST00000210833.2
|
Ptpre
|
protein tyrosine phosphatase, receptor type, E |
| chr18_-_46730381 | 0.04 |
ENSMUST00000036030.14
|
Tmed7
|
transmembrane p24 trafficking protein 7 |
| chr7_-_9687512 | 0.03 |
ENSMUST00000165611.2
|
Vmn2r48
|
vomeronasal 2, receptor 48 |
| chr7_-_9839668 | 0.03 |
ENSMUST00000094863.6
|
Vmn2r51
|
vomeronasal 2, receptor 51 |
| chr15_-_57128522 | 0.03 |
ENSMUST00000137764.2
ENSMUST00000022995.13 |
Slc22a22
|
solute carrier family 22 (organic cation transporter), member 22 |
| chr9_+_106330437 | 0.02 |
ENSMUST00000185874.7
|
Pcbp4
|
poly(rC) binding protein 4 |
| chr7_+_48038274 | 0.01 |
ENSMUST00000056676.5
|
Mrgprb8
|
MAS-related GPR, member B8 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Otx1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 4.6 | GO:0009257 | 10-formyltetrahydrofolate biosynthetic process(GO:0009257) |
| 0.7 | 2.1 | GO:0072248 | metanephric glomerular visceral epithelial cell differentiation(GO:0072248) metanephric glomerular visceral epithelial cell development(GO:0072249) metanephric glomerular epithelial cell differentiation(GO:0072312) metanephric glomerular epithelial cell development(GO:0072313) |
| 0.6 | 2.5 | GO:2001287 | negative regulation of caveolin-mediated endocytosis(GO:2001287) |
| 0.6 | 2.3 | GO:0060697 | positive regulation of phospholipid catabolic process(GO:0060697) |
| 0.5 | 3.3 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.5 | 3.7 | GO:0036072 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.4 | 1.1 | GO:0006550 | isoleucine catabolic process(GO:0006550) |
| 0.3 | 1.3 | GO:0035666 | negative regulation of toll-like receptor 2 signaling pathway(GO:0034136) TRIF-dependent toll-like receptor signaling pathway(GO:0035666) detection of bacterial lipopeptide(GO:0070340) |
| 0.3 | 1.2 | GO:0034165 | positive regulation of toll-like receptor 9 signaling pathway(GO:0034165) |
| 0.3 | 1.7 | GO:1900041 | negative regulation of interleukin-2 secretion(GO:1900041) |
| 0.3 | 1.4 | GO:0044565 | dendritic cell proliferation(GO:0044565) |
| 0.3 | 0.8 | GO:2000721 | positive regulation of transcription from RNA polymerase II promoter involved in smooth muscle cell differentiation(GO:2000721) |
| 0.2 | 0.9 | GO:0006529 | asparagine biosynthetic process(GO:0006529) |
| 0.2 | 0.8 | GO:0060382 | release from viral latency(GO:0019046) regulation of DNA strand elongation(GO:0060382) |
| 0.2 | 4.0 | GO:0042574 | retinal metabolic process(GO:0042574) |
| 0.2 | 3.1 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.2 | 0.5 | GO:0030719 | oocyte construction(GO:0007308) oocyte axis specification(GO:0007309) oocyte anterior/posterior axis specification(GO:0007314) pole plasm assembly(GO:0007315) maternal determination of anterior/posterior axis, embryo(GO:0008358) P granule organization(GO:0030719) |
| 0.1 | 1.5 | GO:0071474 | cellular hyperosmotic response(GO:0071474) |
| 0.1 | 1.5 | GO:0008228 | opsonization(GO:0008228) |
| 0.1 | 1.9 | GO:0018206 | peptidyl-methionine modification(GO:0018206) |
| 0.1 | 0.4 | GO:0097156 | fasciculation of motor neuron axon(GO:0097156) |
| 0.1 | 1.8 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.1 | 0.5 | GO:0071477 | hypotonic salinity response(GO:0042539) cellular hypotonic salinity response(GO:0071477) |
| 0.1 | 0.5 | GO:0038108 | negative regulation of appetite by leptin-mediated signaling pathway(GO:0038108) |
| 0.1 | 1.1 | GO:0098706 | ferric iron import into cell(GO:0097461) ferric iron import across plasma membrane(GO:0098706) |
| 0.1 | 0.7 | GO:0018243 | protein O-linked glycosylation via threonine(GO:0018243) |
| 0.1 | 0.4 | GO:1905068 | positive regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901297) positive regulation of canonical Wnt signaling pathway involved in heart development(GO:1905068) |
| 0.1 | 1.4 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.1 | 0.9 | GO:1903608 | protein localization to cytoplasmic stress granule(GO:1903608) |
| 0.1 | 0.5 | GO:0090611 | ubiquitin-independent protein catabolic process via the multivesicular body sorting pathway(GO:0090611) |
| 0.1 | 0.2 | GO:0006533 | fumarate metabolic process(GO:0006106) aspartate catabolic process(GO:0006533) |
| 0.1 | 0.9 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) nucleolus organization(GO:0007000) |
| 0.1 | 0.9 | GO:0032808 | lacrimal gland development(GO:0032808) |
| 0.1 | 1.9 | GO:0045589 | regulation of regulatory T cell differentiation(GO:0045589) |
| 0.1 | 1.1 | GO:0045292 | mRNA cis splicing, via spliceosome(GO:0045292) |
| 0.1 | 0.5 | GO:2000467 | positive regulation of glycogen (starch) synthase activity(GO:2000467) |
| 0.1 | 0.4 | GO:0009750 | response to fructose(GO:0009750) |
| 0.0 | 1.4 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 0.3 | GO:0048069 | eye pigmentation(GO:0048069) melanosome assembly(GO:1903232) |
| 0.0 | 0.5 | GO:2000786 | positive regulation of autophagosome assembly(GO:2000786) |
| 0.0 | 0.8 | GO:0075522 | IRES-dependent viral translational initiation(GO:0075522) |
| 0.0 | 0.6 | GO:0060394 | negative regulation of pathway-restricted SMAD protein phosphorylation(GO:0060394) |
| 0.0 | 0.3 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.0 | 0.3 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.0 | 0.6 | GO:0015858 | nucleoside transport(GO:0015858) |
| 0.0 | 0.3 | GO:0061470 | T follicular helper cell differentiation(GO:0061470) |
| 0.0 | 0.3 | GO:1900113 | negative regulation of histone H3-K9 trimethylation(GO:1900113) |
| 0.0 | 1.0 | GO:0071425 | hematopoietic stem cell proliferation(GO:0071425) |
| 0.0 | 0.2 | GO:0071985 | multivesicular body sorting pathway(GO:0071985) |
| 0.0 | 1.4 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.0 | 0.4 | GO:0006044 | N-acetylglucosamine metabolic process(GO:0006044) |
| 0.0 | 0.5 | GO:0097150 | neuronal stem cell population maintenance(GO:0097150) |
| 0.0 | 1.1 | GO:0048246 | macrophage chemotaxis(GO:0048246) |
| 0.0 | 0.1 | GO:0042271 | susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.0 | 0.7 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.0 | 0.2 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.0 | 1.6 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.0 | 0.1 | GO:1902774 | late endosome to lysosome transport(GO:1902774) |
| 0.0 | 0.3 | GO:0043486 | histone exchange(GO:0043486) |
| 0.0 | 0.8 | GO:0000281 | mitotic cytokinesis(GO:0000281) |
| 0.0 | 0.4 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 3.7 | GO:0005584 | collagen type I trimer(GO:0005584) |
| 0.5 | 2.1 | GO:0005608 | laminin-3 complex(GO:0005608) |
| 0.2 | 0.9 | GO:0071920 | cleavage body(GO:0071920) |
| 0.2 | 2.3 | GO:0034363 | intermediate-density lipoprotein particle(GO:0034363) |
| 0.2 | 4.4 | GO:0005652 | nuclear lamina(GO:0005652) |
| 0.1 | 3.3 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.1 | 0.9 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.1 | 1.1 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.1 | 0.8 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.1 | 2.7 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.1 | 0.4 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.0 | 0.3 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.0 | 1.0 | GO:0065010 | extracellular membrane-bounded organelle(GO:0065010) |
| 0.0 | 0.5 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.0 | 1.7 | GO:0030904 | retromer complex(GO:0030904) |
| 0.0 | 0.5 | GO:0034464 | BBSome(GO:0034464) |
| 0.0 | 1.5 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.5 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.0 | 0.2 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) |
| 0.0 | 0.2 | GO:0032585 | multivesicular body membrane(GO:0032585) |
| 0.0 | 1.4 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.0 | 0.3 | GO:0031082 | BLOC complex(GO:0031082) |
| 0.0 | 5.4 | GO:0016607 | nuclear speck(GO:0016607) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 4.6 | GO:0004329 | formate-tetrahydrofolate ligase activity(GO:0004329) |
| 0.4 | 2.3 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 0.3 | 1.4 | GO:0004741 | [pyruvate dehydrogenase (lipoamide)] phosphatase activity(GO:0004741) |
| 0.3 | 1.3 | GO:0035663 | Toll-like receptor 2 binding(GO:0035663) |
| 0.3 | 1.1 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.2 | 0.9 | GO:0004066 | asparagine synthase (glutamine-hydrolyzing) activity(GO:0004066) |
| 0.2 | 3.7 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.2 | 1.4 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.2 | 0.9 | GO:0033677 | DNA/RNA helicase activity(GO:0033677) |
| 0.2 | 0.5 | GO:0016309 | 1-phosphatidylinositol-5-phosphate 4-kinase activity(GO:0016309) |
| 0.2 | 1.1 | GO:0016453 | acetyl-CoA C-acetyltransferase activity(GO:0003985) C-acetyltransferase activity(GO:0016453) |
| 0.1 | 0.9 | GO:0004977 | melanocortin receptor activity(GO:0004977) |
| 0.1 | 1.1 | GO:0052851 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.1 | 1.0 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.1 | 1.5 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 0.1 | 3.3 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 0.4 | GO:0004342 | glucosamine-6-phosphate deaminase activity(GO:0004342) |
| 0.1 | 0.5 | GO:0035827 | rubidium ion transmembrane transporter activity(GO:0035827) |
| 0.1 | 4.0 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.1 | 0.5 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.1 | 1.8 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.1 | 0.3 | GO:0000171 | ribonuclease MRP activity(GO:0000171) |
| 0.1 | 0.8 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.1 | 0.2 | GO:0080130 | L-phenylalanine:2-oxoglutarate aminotransferase activity(GO:0080130) |
| 0.1 | 0.8 | GO:1990829 | C-rich single-stranded DNA binding(GO:1990829) |
| 0.0 | 0.4 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) |
| 0.0 | 1.4 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 0.2 | GO:0005047 | signal recognition particle binding(GO:0005047) endoplasmic reticulum signal peptide binding(GO:0030942) |
| 0.0 | 0.4 | GO:0070095 | fructose-6-phosphate binding(GO:0070095) |
| 0.0 | 1.7 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 2.7 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.7 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 0.7 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.0 | 0.6 | GO:0034713 | type I transforming growth factor beta receptor binding(GO:0034713) |
| 0.0 | 3.8 | GO:0017048 | Rho GTPase binding(GO:0017048) |
| 0.0 | 1.2 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.0 | 0.6 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 0.0 | 1.1 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 1.9 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 0.3 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.0 | 1.9 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.2 | GO:0005527 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 1.2 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.0 | 1.5 | GO:0001104 | RNA polymerase II transcription cofactor activity(GO:0001104) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.1 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.1 | 1.4 | ST JAK STAT PATHWAY | Jak-STAT Pathway |
| 0.1 | 5.0 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.1 | 3.3 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 1.8 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 1.4 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.0 | 1.3 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 3.8 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.0 | 0.8 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.0 | 0.5 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.0 | 2.1 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.7 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.2 | 1.8 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.1 | 2.3 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.1 | 1.4 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.1 | 1.4 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.1 | 1.5 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.0 | 1.1 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 1.5 | REACTOME REGULATION OF GLUCOKINASE BY GLUCOKINASE REGULATORY PROTEIN | Genes involved in Regulation of Glucokinase by Glucokinase Regulatory Protein |
| 0.0 | 3.3 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 2.7 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 0.4 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.0 | 0.5 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 0.5 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.0 | 0.8 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 1.7 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 0.6 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 1.8 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.0 | 0.7 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |