Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
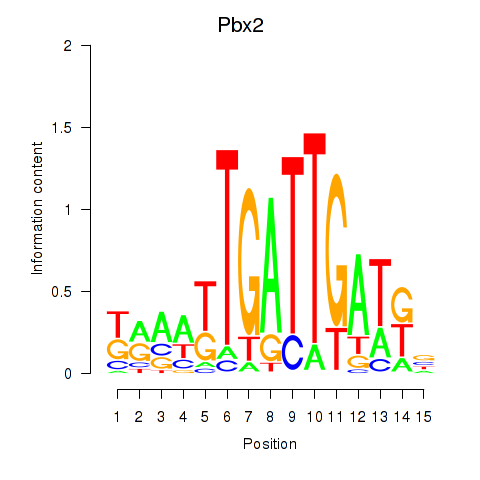
Results for Pbx2
Z-value: 1.01
Transcription factors associated with Pbx2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Pbx2
|
ENSMUSG00000034673.15 | pre B cell leukemia homeobox 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Pbx2 | mm39_v1_chr17_+_34811217_34811303 | -0.81 | 1.5e-09 | Click! |
Activity profile of Pbx2 motif
Sorted Z-values of Pbx2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_61972348 | 24.16 |
ENSMUST00000074018.4
|
Mup20
|
major urinary protein 20 |
| chr13_-_24098981 | 9.57 |
ENSMUST00000110407.4
|
Slc17a4
|
solute carrier family 17 (sodium phosphate), member 4 |
| chr13_-_24098951 | 8.91 |
ENSMUST00000021769.16
|
Slc17a4
|
solute carrier family 17 (sodium phosphate), member 4 |
| chr7_-_46392403 | 8.83 |
ENSMUST00000128088.4
|
Saa1
|
serum amyloid A 1 |
| chr7_+_46401214 | 7.10 |
ENSMUST00000210769.2
ENSMUST00000210272.2 ENSMUST00000075982.4 |
Saa2
|
serum amyloid A 2 |
| chr6_-_3988900 | 5.42 |
ENSMUST00000183682.3
|
Tfpi2
|
tissue factor pathway inhibitor 2 |
| chr4_-_96552349 | 5.17 |
ENSMUST00000030299.8
|
Cyp2j5
|
cytochrome P450, family 2, subfamily j, polypeptide 5 |
| chr3_-_107851021 | 5.15 |
ENSMUST00000106684.8
ENSMUST00000106685.9 |
Gstm6
|
glutathione S-transferase, mu 6 |
| chrX_-_8059597 | 5.10 |
ENSMUST00000143223.2
ENSMUST00000033509.15 |
Ebp
|
phenylalkylamine Ca2+ antagonist (emopamil) binding protein |
| chr3_-_107850707 | 4.89 |
ENSMUST00000106681.3
|
Gstm6
|
glutathione S-transferase, mu 6 |
| chr5_-_87572060 | 4.78 |
ENSMUST00000072818.6
|
Ugt2b38
|
UDP glucuronosyltransferase 2 family, polypeptide B38 |
| chr5_-_87288177 | 4.40 |
ENSMUST00000067790.7
|
Ugt2b5
|
UDP glucuronosyltransferase 2 family, polypeptide B5 |
| chr2_-_62242562 | 4.40 |
ENSMUST00000047812.8
|
Dpp4
|
dipeptidylpeptidase 4 |
| chr6_-_3988835 | 4.30 |
ENSMUST00000203257.2
|
Tfpi2
|
tissue factor pathway inhibitor 2 |
| chr3_+_130411097 | 3.99 |
ENSMUST00000166187.8
ENSMUST00000072271.13 |
Etnppl
|
ethanolamine phosphate phospholyase |
| chr7_+_51537645 | 3.66 |
ENSMUST00000208711.2
|
Gas2
|
growth arrest specific 2 |
| chr8_-_95422851 | 3.22 |
ENSMUST00000034227.6
|
Pllp
|
plasma membrane proteolipid |
| chr19_+_39980868 | 3.19 |
ENSMUST00000049178.3
|
Cyp2c37
|
cytochrome P450, family 2. subfamily c, polypeptide 37 |
| chr13_+_4099001 | 3.08 |
ENSMUST00000118717.10
|
Akr1c14
|
aldo-keto reductase family 1, member C14 |
| chr4_+_140970161 | 2.97 |
ENSMUST00000138096.8
ENSMUST00000006618.9 ENSMUST00000125392.8 |
Arhgef19
|
Rho guanine nucleotide exchange factor (GEF) 19 |
| chr2_-_34951443 | 2.92 |
ENSMUST00000028233.7
|
Hc
|
hemolytic complement |
| chr3_+_122688721 | 2.77 |
ENSMUST00000023820.6
|
Fabp2
|
fatty acid binding protein 2, intestinal |
| chrM_+_11735 | 2.76 |
ENSMUST00000082418.1
|
mt-Nd5
|
mitochondrially encoded NADH dehydrogenase 5 |
| chr6_-_127086480 | 2.64 |
ENSMUST00000039913.9
|
Tigar
|
Trp53 induced glycolysis regulatory phosphatase |
| chr11_+_70104736 | 2.58 |
ENSMUST00000171032.8
|
Slc16a11
|
solute carrier family 16 (monocarboxylic acid transporters), member 11 |
| chr11_-_109986763 | 2.39 |
ENSMUST00000046223.14
ENSMUST00000106664.10 ENSMUST00000106662.2 |
Abca8a
|
ATP-binding cassette, sub-family A (ABC1), member 8a |
| chr11_+_97576619 | 2.35 |
ENSMUST00000107584.8
ENSMUST00000107585.9 |
Cisd3
|
CDGSH iron sulfur domain 3 |
| chr12_+_76302064 | 2.25 |
ENSMUST00000021443.7
|
Mthfd1
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent), methenyltetrahydrofolate cyclohydrolase, formyltetrahydrofolate synthase |
| chr1_+_155911136 | 2.19 |
ENSMUST00000111757.10
|
Tor1aip2
|
torsin A interacting protein 2 |
| chr12_+_76302145 | 2.19 |
ENSMUST00000220046.2
|
Mthfd1
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent), methenyltetrahydrofolate cyclohydrolase, formyltetrahydrofolate synthase |
| chr1_+_88066086 | 2.16 |
ENSMUST00000014263.6
|
Ugt1a6a
|
UDP glucuronosyltransferase 1 family, polypeptide A6A |
| chr6_+_8259405 | 2.02 |
ENSMUST00000160705.8
ENSMUST00000159433.8 |
Umad1
|
UMAP1-MVP12 associated (UMA) domain containing 1 |
| chr11_-_115078653 | 1.99 |
ENSMUST00000103041.8
|
Nat9
|
N-acetyltransferase 9 (GCN5-related, putative) |
| chr6_+_8259379 | 1.97 |
ENSMUST00000162034.8
|
Umad1
|
UMAP1-MVP12 associated (UMA) domain containing 1 |
| chr6_+_124281607 | 1.93 |
ENSMUST00000032234.5
ENSMUST00000112541.8 |
Cd163
|
CD163 antigen |
| chr6_+_8259288 | 1.91 |
ENSMUST00000159335.8
|
Umad1
|
UMAP1-MVP12 associated (UMA) domain containing 1 |
| chr19_-_3625698 | 1.90 |
ENSMUST00000172362.3
ENSMUST00000025846.16 ENSMUST00000226109.2 ENSMUST00000113997.9 |
Ppp6r3
|
protein phosphatase 6, regulatory subunit 3 |
| chr2_-_75768752 | 1.83 |
ENSMUST00000099996.5
|
Ttc30b
|
tetratricopeptide repeat domain 30B |
| chrX_-_36137764 | 1.80 |
ENSMUST00000047486.6
|
C330007P06Rik
|
RIKEN cDNA C330007P06 gene |
| chr11_+_97576724 | 1.72 |
ENSMUST00000107583.3
|
Cisd3
|
CDGSH iron sulfur domain 3 |
| chr3_-_65300000 | 1.69 |
ENSMUST00000029414.12
|
Ssr3
|
signal sequence receptor, gamma |
| chr15_+_92495007 | 1.65 |
ENSMUST00000035399.10
|
Pdzrn4
|
PDZ domain containing RING finger 4 |
| chr13_-_86194889 | 1.60 |
ENSMUST00000131011.2
|
Cox7c
|
cytochrome c oxidase subunit 7C |
| chr10_-_62486948 | 1.57 |
ENSMUST00000020270.6
|
Ddx50
|
DExD box helicase 50 |
| chr1_-_80191649 | 1.54 |
ENSMUST00000058748.2
|
Fam124b
|
family with sequence similarity 124, member B |
| chr12_+_44268134 | 1.53 |
ENSMUST00000122902.8
|
Pnpla8
|
patatin-like phospholipase domain containing 8 |
| chr11_-_60243695 | 1.53 |
ENSMUST00000095254.12
ENSMUST00000102683.11 ENSMUST00000093048.13 ENSMUST00000093046.13 ENSMUST00000064019.15 ENSMUST00000102682.5 |
Tom1l2
|
target of myb1-like 2 (chicken) |
| chr19_+_8897732 | 1.41 |
ENSMUST00000096243.7
|
B3gat3
|
beta-1,3-glucuronyltransferase 3 (glucuronosyltransferase I) |
| chr9_+_119170360 | 1.40 |
ENSMUST00000039784.12
|
Acaa1a
|
acetyl-Coenzyme A acyltransferase 1A |
| chr11_-_96720738 | 1.40 |
ENSMUST00000107657.8
ENSMUST00000081775.12 |
Nfe2l1
|
nuclear factor, erythroid derived 2,-like 1 |
| chr11_-_31621727 | 1.38 |
ENSMUST00000109415.2
|
Bod1
|
biorientation of chromosomes in cell division 1 |
| chr17_+_85265420 | 1.38 |
ENSMUST00000080217.14
ENSMUST00000112304.10 |
Ppm1b
|
protein phosphatase 1B, magnesium dependent, beta isoform |
| chr5_-_25047577 | 1.32 |
ENSMUST00000030787.9
|
Rheb
|
Ras homolog enriched in brain |
| chr8_-_41507808 | 1.30 |
ENSMUST00000093534.11
|
Mtus1
|
mitochondrial tumor suppressor 1 |
| chr6_+_8259328 | 1.28 |
ENSMUST00000159378.8
|
Umad1
|
UMAP1-MVP12 associated (UMA) domain containing 1 |
| chr1_-_169796709 | 1.26 |
ENSMUST00000027989.13
ENSMUST00000111353.4 |
Hsd17b7
|
hydroxysteroid (17-beta) dehydrogenase 7 |
| chr5_+_144192033 | 1.23 |
ENSMUST00000056578.7
|
Bri3
|
brain protein I3 |
| chr18_+_3383230 | 1.20 |
ENSMUST00000162301.8
ENSMUST00000161317.2 |
Cul2
|
cullin 2 |
| chrX_+_37689503 | 1.19 |
ENSMUST00000000365.3
|
Mcts1
|
malignant T cell amplified sequence 1 |
| chr7_-_3298243 | 1.17 |
ENSMUST00000108653.4
|
Nlrp12
|
NLR family, pyrin domain containing 12 |
| chr2_-_177567397 | 1.17 |
ENSMUST00000108934.9
ENSMUST00000081529.11 |
Zfp972
|
zinc finger protein 972 |
| chr1_+_136552639 | 1.14 |
ENSMUST00000047734.15
ENSMUST00000112046.2 |
Zfp281
|
zinc finger protein 281 |
| chr3_+_20011201 | 1.13 |
ENSMUST00000091309.12
ENSMUST00000108329.8 ENSMUST00000003714.13 |
Cp
|
ceruloplasmin |
| chr3_-_65299967 | 1.05 |
ENSMUST00000119896.2
|
Ssr3
|
signal sequence receptor, gamma |
| chr3_+_20011251 | 1.05 |
ENSMUST00000108328.8
|
Cp
|
ceruloplasmin |
| chr7_+_29607917 | 1.02 |
ENSMUST00000186475.2
|
Zfp383
|
zinc finger protein 383 |
| chr9_-_108140925 | 0.98 |
ENSMUST00000171412.7
ENSMUST00000195429.6 ENSMUST00000080435.9 |
Dag1
|
dystroglycan 1 |
| chr1_+_44158111 | 0.96 |
ENSMUST00000155917.8
|
Bivm
|
basic, immunoglobulin-like variable motif containing |
| chr7_-_12829100 | 0.88 |
ENSMUST00000209822.3
ENSMUST00000235753.2 |
Vmn1r85
|
vomeronasal 1 receptor 85 |
| chr7_+_133239414 | 0.87 |
ENSMUST00000128901.9
|
Edrf1
|
erythroid differentiation regulatory factor 1 |
| chr12_-_83968507 | 0.87 |
ENSMUST00000222439.2
ENSMUST00000135962.8 ENSMUST00000155112.8 ENSMUST00000136848.8 ENSMUST00000126943.2 ENSMUST00000117217.8 |
Numb
|
NUMB endocytic adaptor protein |
| chr4_-_150087587 | 0.87 |
ENSMUST00000084117.13
|
H6pd
|
hexose-6-phosphate dehydrogenase (glucose 1-dehydrogenase) |
| chr3_-_88679881 | 0.86 |
ENSMUST00000090945.5
|
Syt11
|
synaptotagmin XI |
| chr15_-_100945261 | 0.83 |
ENSMUST00000222611.2
|
Tmdd1
|
transmembrane and death domain 1 |
| chr5_+_104318542 | 0.79 |
ENSMUST00000112771.2
|
Dspp
|
dentin sialophosphoprotein |
| chr11_-_109986804 | 0.79 |
ENSMUST00000100287.9
|
Abca8a
|
ATP-binding cassette, sub-family A (ABC1), member 8a |
| chr1_-_85239791 | 0.75 |
ENSMUST00000162421.2
|
C130026I21Rik
|
RIKEN cDNA C130026I21 gene |
| chr5_-_3697806 | 0.75 |
ENSMUST00000119783.2
ENSMUST00000007559.15 |
Gatad1
|
GATA zinc finger domain containing 1 |
| chr7_+_43256593 | 0.74 |
ENSMUST00000120935.2
ENSMUST00000127765.8 ENSMUST00000032661.14 |
Zfp819
|
zinc finger protein 819 |
| chr3_+_20111958 | 0.74 |
ENSMUST00000002502.12
|
Hltf
|
helicase-like transcription factor |
| chr17_+_75742881 | 0.71 |
ENSMUST00000164192.9
|
Rasgrp3
|
RAS, guanyl releasing protein 3 |
| chr3_+_29136449 | 0.71 |
ENSMUST00000146943.8
|
Egfem1
|
EGF-like and EMI domain containing 1 |
| chr13_+_60749735 | 0.70 |
ENSMUST00000226059.2
ENSMUST00000077453.13 |
Dapk1
|
death associated protein kinase 1 |
| chr14_+_53194239 | 0.69 |
ENSMUST00000199800.2
|
Trav15d-2-dv6d-2
|
T cell receptor alpha variable 15D-2-DV6D-2 |
| chr17_-_35265702 | 0.68 |
ENSMUST00000097338.11
|
Msh5
|
mutS homolog 5 |
| chr9_+_95441652 | 0.67 |
ENSMUST00000079597.7
|
Paqr9
|
progestin and adipoQ receptor family member IX |
| chr15_-_5137951 | 0.66 |
ENSMUST00000141020.2
|
Card6
|
caspase recruitment domain family, member 6 |
| chr4_+_102843540 | 0.65 |
ENSMUST00000030248.12
ENSMUST00000125417.9 ENSMUST00000169211.3 |
Dynlt5
|
dynein light chain Tctex-type 5 |
| chr1_-_181992051 | 0.65 |
ENSMUST00000227629.2
ENSMUST00000227586.2 |
Vmn1r1
|
vomeronasal 1 receptor 1 |
| chr8_+_91555449 | 0.65 |
ENSMUST00000109614.9
|
Chd9
|
chromodomain helicase DNA binding protein 9 |
| chr12_+_119278005 | 0.64 |
ENSMUST00000222784.2
|
Macc1
|
metastasis associated in colon cancer 1 |
| chr11_-_58504307 | 0.63 |
ENSMUST00000048801.8
|
Lypd8l
|
LY6/PLAUR domain containing 8 like |
| chr1_+_60948149 | 0.63 |
ENSMUST00000027164.9
|
Ctla4
|
cytotoxic T-lymphocyte-associated protein 4 |
| chr9_-_20371123 | 0.61 |
ENSMUST00000162438.2
|
Zfp26
|
zinc finger protein 26 |
| chr18_-_23171713 | 0.60 |
ENSMUST00000081423.13
|
Nol4
|
nucleolar protein 4 |
| chrX_-_43879055 | 0.60 |
ENSMUST00000060481.9
|
Dcaf12l1
|
DDB1 and CUL4 associated factor 12-like 1 |
| chr6_-_13677928 | 0.59 |
ENSMUST00000203078.2
ENSMUST00000045235.8 |
Bmt2
|
base methyltransferase of 25S rRNA 2 |
| chr1_-_155910546 | 0.58 |
ENSMUST00000169241.8
|
Tor1aip1
|
torsin A interacting protein 1 |
| chr5_-_72893941 | 0.56 |
ENSMUST00000169534.6
|
Txk
|
TXK tyrosine kinase |
| chr2_-_101451383 | 0.56 |
ENSMUST00000090513.11
|
Iftap
|
intraflagellar transport associated protein |
| chr11_-_99241924 | 0.55 |
ENSMUST00000017732.3
|
Krt27
|
keratin 27 |
| chr15_-_5137975 | 0.53 |
ENSMUST00000118365.3
|
Card6
|
caspase recruitment domain family, member 6 |
| chr15_-_101471324 | 0.53 |
ENSMUST00000023714.5
|
Krt90
|
keratin 90 |
| chr4_-_118667141 | 0.52 |
ENSMUST00000084313.5
ENSMUST00000219094.2 |
Olfr1335
|
olfactory receptor 1335 |
| chr14_+_123897383 | 0.52 |
ENSMUST00000049681.14
|
Itgbl1
|
integrin, beta-like 1 |
| chr1_+_134487928 | 0.52 |
ENSMUST00000112198.3
|
Kdm5b
|
lysine (K)-specific demethylase 5B |
| chr1_+_169756217 | 0.51 |
ENSMUST00000094348.4
|
Ccdc190
|
coiled-coil domain containing 190 |
| chr9_-_58277734 | 0.51 |
ENSMUST00000040217.6
|
Tbc1d21
|
TBC1 domain family, member 21 |
| chr7_+_119289249 | 0.51 |
ENSMUST00000047045.10
|
Acsm4
|
acyl-CoA synthetase medium-chain family member 4 |
| chr13_-_99653045 | 0.50 |
ENSMUST00000064762.6
|
Map1b
|
microtubule-associated protein 1B |
| chr4_+_147576874 | 0.50 |
ENSMUST00000105721.9
|
Zfp982
|
zinc finger protein 982 |
| chr2_-_136727599 | 0.50 |
ENSMUST00000231720.2
|
ENSMUSG00000116563.2
|
novel protein |
| chrX_+_36920563 | 0.49 |
ENSMUST00000096454.2
|
Rhox7a
|
reproductive homeobox 7A |
| chr2_-_151510453 | 0.48 |
ENSMUST00000180195.8
ENSMUST00000096439.4 |
Rad21l
|
RAD21-like (S. pombe) |
| chr2_-_88590348 | 0.48 |
ENSMUST00000131038.4
ENSMUST00000213138.3 |
Olfr1199
|
olfactory receptor 1199 |
| chr3_+_145823205 | 0.47 |
ENSMUST00000140214.3
|
Mcoln3
|
mucolipin 3 |
| chr8_-_84963653 | 0.46 |
ENSMUST00000039480.7
|
Zswim4
|
zinc finger SWIM-type containing 4 |
| chr14_+_63284438 | 0.45 |
ENSMUST00000067990.8
ENSMUST00000111203.2 |
Defb42
|
defensin beta 42 |
| chr1_+_134487893 | 0.45 |
ENSMUST00000047714.14
|
Kdm5b
|
lysine (K)-specific demethylase 5B |
| chr17_-_52140305 | 0.44 |
ENSMUST00000133574.8
|
Satb1
|
special AT-rich sequence binding protein 1 |
| chr19_-_5168251 | 0.44 |
ENSMUST00000113728.8
ENSMUST00000113727.8 ENSMUST00000025798.13 |
Klc2
|
kinesin light chain 2 |
| chr8_-_55177510 | 0.44 |
ENSMUST00000175915.8
|
Wdr17
|
WD repeat domain 17 |
| chr19_-_11833365 | 0.44 |
ENSMUST00000079875.4
|
Olfr1418
|
olfactory receptor 1418 |
| chrX_-_36986109 | 0.41 |
ENSMUST00000115156.2
|
Rhox7b
|
reproductive homeobox 7B |
| chr1_+_157353696 | 0.41 |
ENSMUST00000111700.8
|
Sec16b
|
SEC16 homolog B (S. cerevisiae) |
| chr3_-_152687877 | 0.39 |
ENSMUST00000044278.6
|
St6galnac5
|
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 5 |
| chr13_-_43324937 | 0.39 |
ENSMUST00000222160.2
ENSMUST00000179852.9 ENSMUST00000021797.9 |
Tbc1d7
|
TBC1 domain family, member 7 |
| chr11_-_99313078 | 0.39 |
ENSMUST00000017741.4
|
Krt12
|
keratin 12 |
| chr1_-_138770167 | 0.39 |
ENSMUST00000112026.4
ENSMUST00000019374.14 |
Lhx9
|
LIM homeobox protein 9 |
| chr7_+_106737534 | 0.39 |
ENSMUST00000213367.3
ENSMUST00000214819.3 ENSMUST00000216871.3 ENSMUST00000215284.3 ENSMUST00000209942.2 |
Olfr716
|
olfactory receptor 716 |
| chr17_-_28584182 | 0.38 |
ENSMUST00000041819.14
|
Tulp1
|
tubby like protein 1 |
| chr3_+_152052102 | 0.38 |
ENSMUST00000117492.9
ENSMUST00000026507.13 ENSMUST00000197748.5 |
Usp33
|
ubiquitin specific peptidase 33 |
| chr11_+_88964667 | 0.37 |
ENSMUST00000100619.11
|
Gm525
|
predicted gene 525 |
| chr6_+_40468167 | 0.37 |
ENSMUST00000064932.6
|
Tas2r137
|
taste receptor, type 2, member 137 |
| chr4_+_146033882 | 0.37 |
ENSMUST00000105730.2
ENSMUST00000091878.6 |
Zfp987
|
zinc finger protein 987 |
| chr14_+_53574579 | 0.37 |
ENSMUST00000179580.3
|
Trav13n-3
|
T cell receptor alpha variable 13N-3 |
| chr6_-_67743756 | 0.36 |
ENSMUST00000103306.2
|
Igkv1-131
|
immunoglobulin kappa variable 1-131 |
| chr19_-_32080496 | 0.36 |
ENSMUST00000235213.2
ENSMUST00000236504.2 |
Asah2
|
N-acylsphingosine amidohydrolase 2 |
| chr7_-_121700958 | 0.35 |
ENSMUST00000139456.2
ENSMUST00000106471.9 ENSMUST00000123296.8 ENSMUST00000033157.10 |
Ndufab1
|
NADH:ubiquinone oxidoreductase subunit AB1 |
| chr7_+_99384352 | 0.34 |
ENSMUST00000098264.2
|
Olfr520
|
olfactory receptor 520 |
| chr7_-_27749453 | 0.34 |
ENSMUST00000140053.3
ENSMUST00000032824.10 |
Psmc4
|
proteasome (prosome, macropain) 26S subunit, ATPase, 4 |
| chr16_-_37205302 | 0.33 |
ENSMUST00000114781.8
ENSMUST00000114780.8 |
Stxbp5l
|
syntaxin binding protein 5-like |
| chr6_-_129303659 | 0.33 |
ENSMUST00000203159.2
|
Clec2m
|
C-type lectin domain family 2, member m |
| chr4_-_59138983 | 0.32 |
ENSMUST00000107547.2
|
Shoc1
|
shortage in chiasmata 1 |
| chr13_-_21798192 | 0.32 |
ENSMUST00000051874.6
|
Olfr1362
|
olfactory receptor 1362 |
| chr7_+_18451998 | 0.32 |
ENSMUST00000051973.9
ENSMUST00000208221.2 ENSMUST00000108481.8 |
Psg22
|
pregnancy-specific glycoprotein 22 |
| chr7_-_18165959 | 0.30 |
ENSMUST00000019291.7
|
Psg28
|
pregnancy-specific glycoprotein 28 |
| chr1_+_60948307 | 0.29 |
ENSMUST00000097720.4
|
Ctla4
|
cytotoxic T-lymphocyte-associated protein 4 |
| chr9_-_6269846 | 0.29 |
ENSMUST00000051706.6
|
Ddi1
|
DNA-damage inducible 1 |
| chr3_-_98417351 | 0.29 |
ENSMUST00000179429.6
|
Hsd3b4
|
hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 4 |
| chrX_-_20955370 | 0.28 |
ENSMUST00000040667.13
|
Zfp300
|
zinc finger protein 300 |
| chr9_-_80347482 | 0.27 |
ENSMUST00000085289.12
ENSMUST00000185068.2 ENSMUST00000113250.10 |
Impg1
|
interphotoreceptor matrix proteoglycan 1 |
| chr18_+_46983105 | 0.26 |
ENSMUST00000025358.4
ENSMUST00000234519.2 |
Lvrn
|
laeverin |
| chr1_+_164103127 | 0.26 |
ENSMUST00000027867.7
|
Ccdc181
|
coiled-coil domain containing 181 |
| chr2_-_175017664 | 0.24 |
ENSMUST00000109054.3
|
Gm14443
|
predicted gene 14443 |
| chr17_+_9068805 | 0.24 |
ENSMUST00000115720.8
|
Pde10a
|
phosphodiesterase 10A |
| chr2_-_89555356 | 0.24 |
ENSMUST00000216203.2
ENSMUST00000213196.2 |
Olfr1252
|
olfactory receptor 1252 |
| chr4_+_146599377 | 0.23 |
ENSMUST00000140089.7
ENSMUST00000179175.3 |
Zfp981
|
zinc finger protein 981 |
| chr4_-_146993984 | 0.23 |
ENSMUST00000238583.2
ENSMUST00000049821.4 |
Gm21411
|
predicted gene, 21411 |
| chr15_-_98118858 | 0.23 |
ENSMUST00000142443.8
ENSMUST00000170618.8 |
Gm44579
Olfr287
|
predicted gene 44579 olfactory receptor 287 |
| chrX_-_121307036 | 0.23 |
ENSMUST00000079490.6
|
Nap1l3
|
nucleosome assembly protein 1-like 3 |
| chr7_-_103291882 | 0.23 |
ENSMUST00000213536.2
ENSMUST00000216570.2 |
Olfr622
|
olfactory receptor 622 |
| chr9_-_70841881 | 0.22 |
ENSMUST00000214995.2
|
Lipc
|
lipase, hepatic |
| chr7_+_6048160 | 0.22 |
ENSMUST00000037728.13
ENSMUST00000121583.2 |
Nlrp4c
|
NLR family, pyrin domain containing 4C |
| chr11_+_58311921 | 0.21 |
ENSMUST00000013797.3
|
1810065E05Rik
|
RIKEN cDNA 1810065E05 gene |
| chr16_-_37205277 | 0.21 |
ENSMUST00000114787.8
ENSMUST00000114782.8 ENSMUST00000114775.8 |
Stxbp5l
|
syntaxin binding protein 5-like |
| chr4_-_147787010 | 0.20 |
ENSMUST00000117638.2
|
Zfp534
|
zinc finger protein 534 |
| chr4_+_145397238 | 0.20 |
ENSMUST00000105738.9
|
Zfp980
|
zinc finger protein 980 |
| chr13_-_25454058 | 0.20 |
ENSMUST00000057866.13
|
Nrsn1
|
neurensin 1 |
| chr9_+_92191415 | 0.20 |
ENSMUST00000150594.8
ENSMUST00000098477.8 |
1700057G04Rik
|
RIKEN cDNA 1700057G04 gene |
| chr4_+_147056433 | 0.19 |
ENSMUST00000146688.3
|
Zfp989
|
zinc finger protein 989 |
| chr9_+_38686470 | 0.18 |
ENSMUST00000071681.4
|
Olfr921
|
olfactory receptor 921 |
| chrX_-_91643178 | 0.18 |
ENSMUST00000113955.2
|
Mageb18
|
MAGE family member B18 |
| chr2_-_57942844 | 0.18 |
ENSMUST00000090940.6
|
Ermn
|
ermin, ERM-like protein |
| chr2_+_29014106 | 0.18 |
ENSMUST00000129544.8
|
Setx
|
senataxin |
| chr12_+_103547657 | 0.18 |
ENSMUST00000190664.2
|
Ppp4r4
|
protein phosphatase 4, regulatory subunit 4 |
| chr6_+_78347636 | 0.17 |
ENSMUST00000204873.3
|
Reg3b
|
regenerating islet-derived 3 beta |
| chr6_+_41112064 | 0.17 |
ENSMUST00000103272.4
|
Trbv14
|
T cell receptor beta, variable 14 |
| chrX_+_72108393 | 0.17 |
ENSMUST00000060418.8
|
Pnma3
|
paraneoplastic antigen MA3 |
| chr4_+_146093394 | 0.17 |
ENSMUST00000168483.9
|
Zfp600
|
zinc finger protein 600 |
| chr3_-_151455514 | 0.17 |
ENSMUST00000029671.9
|
Ifi44
|
interferon-induced protein 44 |
| chr4_+_12089373 | 0.16 |
ENSMUST00000095143.9
ENSMUST00000063839.6 |
Rbm12b2
|
RNA binding motif protein 12 B2 |
| chr5_-_122492223 | 0.16 |
ENSMUST00000117263.8
ENSMUST00000049009.7 |
Rad9b
|
RAD9 checkpoint clamp component B |
| chr6_-_69920632 | 0.16 |
ENSMUST00000198880.5
ENSMUST00000103371.3 |
Igkv12-38
|
immunoglobulin kappa chain variable 12-38 |
| chr15_+_82230155 | 0.16 |
ENSMUST00000023086.15
|
Smdt1
|
single-pass membrane protein with aspartate rich tail 1 |
| chrX_+_111150171 | 0.16 |
ENSMUST00000164272.3
ENSMUST00000132037.2 |
4933403O08Rik
|
RIKEN cDNA 4933403O08 gene |
| chr2_-_113588983 | 0.15 |
ENSMUST00000099575.4
|
Grem1
|
gremlin 1, DAN family BMP antagonist |
| chr15_+_77392466 | 0.15 |
ENSMUST00000172191.3
|
Apol11a
|
apolipoprotein L 11a |
| chr12_-_55349760 | 0.15 |
ENSMUST00000021410.10
|
Ppp2r3c
|
protein phosphatase 2, regulatory subunit B'', gamma |
| chr2_+_128809268 | 0.15 |
ENSMUST00000110320.9
ENSMUST00000110319.3 |
Zc3h6
|
zinc finger CCCH type containing 6 |
| chr4_+_147106307 | 0.15 |
ENSMUST00000075775.6
|
Rex2
|
reduced expression 2 |
| chr1_+_15875846 | 0.13 |
ENSMUST00000188371.7
ENSMUST00000027057.8 |
Terf1
|
telomeric repeat binding factor 1 |
| chr2_-_4656870 | 0.13 |
ENSMUST00000192470.2
ENSMUST00000035721.14 ENSMUST00000152362.7 |
Prpf18
|
pre-mRNA processing factor 18 |
| chr16_-_50151350 | 0.13 |
ENSMUST00000114488.8
|
Bbx
|
bobby sox HMG box containing |
| chr16_-_93400691 | 0.13 |
ENSMUST00000023669.14
ENSMUST00000233931.2 ENSMUST00000154355.3 |
Setd4
|
SET domain containing 4 |
| chr10_+_81442137 | 0.12 |
ENSMUST00000119492.2
|
BC025920
|
cDNA sequence BC025920 |
| chr19_-_32038838 | 0.12 |
ENSMUST00000096119.5
|
Asah2
|
N-acylsphingosine amidohydrolase 2 |
| chr5_+_8710059 | 0.12 |
ENSMUST00000047753.5
|
Abcb1a
|
ATP-binding cassette, sub-family B (MDR/TAP), member 1A |
| chr5_-_151510389 | 0.12 |
ENSMUST00000165928.4
|
Vmn2r18
|
vomeronasal 2, receptor 18 |
| chr17_-_35334404 | 0.11 |
ENSMUST00000172854.2
ENSMUST00000062657.5 |
Ly6g5b
|
lymphocyte antigen 6 complex, locus G5B |
Network of associatons between targets according to the STRING database.
First level regulatory network of Pbx2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.0 | 24.2 | GO:0008355 | olfactory learning(GO:0008355) |
| 1.7 | 5.2 | GO:2000863 | positive regulation of estrogen secretion(GO:2000863) |
| 1.1 | 4.4 | GO:0000105 | histidine biosynthetic process(GO:0000105) 10-formyltetrahydrofolate biosynthetic process(GO:0009257) |
| 0.9 | 2.6 | GO:1902689 | negative regulation of NAD metabolic process(GO:1902689) negative regulation of glucose catabolic process to lactate via pyruvate(GO:1904024) |
| 0.9 | 4.4 | GO:0036343 | psychomotor behavior(GO:0036343) |
| 0.7 | 2.9 | GO:0010760 | negative regulation of macrophage chemotaxis(GO:0010760) |
| 0.5 | 1.5 | GO:0046338 | phosphatidylethanolamine catabolic process(GO:0046338) |
| 0.5 | 1.0 | GO:2000861 | estrogen secretion(GO:0035937) regulation of estrogen secretion(GO:2000861) |
| 0.4 | 1.2 | GO:0008588 | release of cytoplasmic sequestered NF-kappaB(GO:0008588) |
| 0.3 | 1.0 | GO:0021682 | nerve maturation(GO:0021682) |
| 0.3 | 3.2 | GO:0031580 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 0.3 | 0.9 | GO:0030860 | regulation of polarized epithelial cell differentiation(GO:0030860) |
| 0.3 | 18.0 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.3 | 1.4 | GO:0050651 | dermatan sulfate proteoglycan biosynthetic process(GO:0050651) |
| 0.3 | 0.8 | GO:0071460 | cellular response to cell-matrix adhesion(GO:0071460) |
| 0.2 | 0.9 | GO:1905154 | negative regulation of membrane invagination(GO:1905154) calcium ion regulated lysosome exocytosis(GO:1990927) |
| 0.2 | 1.2 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.2 | 1.4 | GO:0006499 | N-terminal protein myristoylation(GO:0006499) |
| 0.2 | 2.2 | GO:0052697 | flavonoid glucuronidation(GO:0052696) xenobiotic glucuronidation(GO:0052697) |
| 0.2 | 2.7 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.2 | 0.9 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 0.1 | 2.2 | GO:0090435 | protein localization to nuclear envelope(GO:0090435) |
| 0.1 | 1.3 | GO:2000074 | regulation of type B pancreatic cell development(GO:2000074) |
| 0.1 | 17.9 | GO:0035725 | sodium ion transmembrane transport(GO:0035725) |
| 0.1 | 0.3 | GO:2001201 | regulation of transforming growth factor-beta secretion(GO:2001201) |
| 0.1 | 1.6 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.1 | 3.2 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.1 | 0.6 | GO:0060335 | positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) |
| 0.1 | 6.4 | GO:0006695 | cholesterol biosynthetic process(GO:0006695) |
| 0.1 | 2.2 | GO:0046688 | response to copper ion(GO:0046688) |
| 0.1 | 0.5 | GO:0097240 | meiotic telomere tethering at nuclear periphery(GO:0044821) meiotic attachment of telomere to nuclear envelope(GO:0070197) chromosome attachment to the nuclear envelope(GO:0097240) |
| 0.1 | 0.2 | GO:0031554 | regulation of DNA-templated transcription, termination(GO:0031554) |
| 0.1 | 0.4 | GO:0035262 | gonad morphogenesis(GO:0035262) |
| 0.1 | 0.6 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.1 | 0.2 | GO:1900158 | negative regulation of monocyte chemotaxis(GO:0090027) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.0 | 0.9 | GO:0006098 | pentose-phosphate shunt(GO:0006098) |
| 0.0 | 0.1 | GO:0000350 | generation of catalytic spliceosome for second transesterification step(GO:0000350) |
| 0.0 | 0.2 | GO:0080163 | regulation of protein serine/threonine phosphatase activity(GO:0080163) |
| 0.0 | 2.1 | GO:0042773 | ATP synthesis coupled electron transport(GO:0042773) |
| 0.0 | 0.7 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.0 | 0.7 | GO:0071447 | cellular response to hydroperoxide(GO:0071447) |
| 0.0 | 0.4 | GO:0048207 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.0 | 0.4 | GO:0009249 | protein lipoylation(GO:0009249) |
| 0.0 | 1.3 | GO:0010758 | regulation of macrophage chemotaxis(GO:0010758) |
| 0.0 | 1.4 | GO:0008206 | bile acid metabolic process(GO:0008206) |
| 0.0 | 9.5 | GO:0010466 | negative regulation of peptidase activity(GO:0010466) |
| 0.0 | 0.4 | GO:0061303 | cornea development in camera-type eye(GO:0061303) |
| 0.0 | 0.1 | GO:2001025 | positive regulation of response to drug(GO:2001025) |
| 0.0 | 0.1 | GO:0002759 | regulation of antimicrobial humoral response(GO:0002759) |
| 0.0 | 1.9 | GO:0043666 | regulation of phosphoprotein phosphatase activity(GO:0043666) |
| 0.0 | 0.1 | GO:0097694 | establishment of RNA localization to telomere(GO:0097694) |
| 0.0 | 0.5 | GO:0061162 | establishment of monopolar cell polarity(GO:0061162) |
| 0.0 | 0.7 | GO:0006301 | postreplication repair(GO:0006301) |
| 0.0 | 0.4 | GO:1902018 | negative regulation of cilium assembly(GO:1902018) |
| 0.0 | 1.1 | GO:0050892 | intestinal absorption(GO:0050892) |
| 0.0 | 0.6 | GO:0010172 | embryonic body morphogenesis(GO:0010172) |
| 0.0 | 0.4 | GO:1903546 | protein localization to photoreceptor outer segment(GO:1903546) |
| 0.0 | 0.4 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.0 | 1.5 | GO:0045839 | negative regulation of mitotic nuclear division(GO:0045839) |
| 0.0 | 0.3 | GO:0045899 | positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.0 | 1.2 | GO:0042073 | intraciliary transport(GO:0042073) |
| 0.0 | 0.4 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.0 | 0.4 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.0 | 3.6 | GO:0007050 | cell cycle arrest(GO:0007050) |
| 0.0 | 0.2 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.0 | 0.5 | GO:0031069 | hair follicle morphogenesis(GO:0031069) |
| 0.0 | 2.6 | GO:0015718 | monocarboxylic acid transport(GO:0015718) |
| 0.0 | 0.0 | GO:0030845 | regulation of inositol-polyphosphate 5-phosphatase activity(GO:0010924) positive regulation of inositol-polyphosphate 5-phosphatase activity(GO:0010925) phospholipase C-inhibiting G-protein coupled receptor signaling pathway(GO:0030845) regulation of cell diameter(GO:0060305) |
| 0.0 | 0.3 | GO:0035634 | response to stilbenoid(GO:0035634) |
| 0.0 | 3.0 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.0 | 3.3 | GO:0006869 | lipid transport(GO:0006869) |
| 0.0 | 0.2 | GO:0031573 | DNA replication checkpoint(GO:0000076) intra-S DNA damage checkpoint(GO:0031573) |
| 0.0 | 0.4 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.1 | GO:0044829 | positive regulation by host of viral genome replication(GO:0044829) |
| 0.0 | 0.1 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.0 | 0.6 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 4.4 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.4 | 16.0 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.3 | 2.9 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.3 | 2.8 | GO:0045179 | apical cortex(GO:0045179) |
| 0.1 | 1.2 | GO:0030891 | VCB complex(GO:0030891) |
| 0.1 | 3.2 | GO:0043218 | compact myelin(GO:0043218) |
| 0.1 | 0.2 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.1 | 1.0 | GO:0016011 | dystroglycan complex(GO:0016011) |
| 0.1 | 0.5 | GO:0034991 | nuclear meiotic cohesin complex(GO:0034991) |
| 0.1 | 0.9 | GO:0032009 | early phagosome(GO:0032009) |
| 0.0 | 1.8 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.0 | 1.4 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 0.7 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.0 | 0.3 | GO:0031595 | nuclear proteasome complex(GO:0031595) |
| 0.0 | 0.2 | GO:0030896 | checkpoint clamp complex(GO:0030896) |
| 0.0 | 2.2 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 0.3 | GO:0002177 | manchette(GO:0002177) |
| 0.0 | 0.2 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.0 | 0.9 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.0 | 0.5 | GO:0043196 | varicosity(GO:0043196) |
| 0.0 | 0.4 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.0 | 1.5 | GO:0031903 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 8.9 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.0 | 0.1 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.0 | 2.6 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 9.1 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 0.1 | GO:0070187 | telosome(GO:0070187) |
| 0.0 | 28.2 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.0 | 0.9 | GO:0030136 | clathrin-coated vesicle(GO:0030136) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.8 | 24.2 | GO:0005186 | pheromone activity(GO:0005186) |
| 1.3 | 4.0 | GO:0016838 | carbon-oxygen lyase activity, acting on phosphates(GO:0016838) |
| 1.2 | 18.5 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 1.1 | 4.4 | GO:0004329 | formate-tetrahydrofolate ligase activity(GO:0004329) |
| 0.7 | 5.1 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 0.5 | 3.1 | GO:0047023 | androsterone dehydrogenase activity(GO:0047023) |
| 0.5 | 1.4 | GO:0008775 | acetate CoA-transferase activity(GO:0008775) |
| 0.4 | 15.9 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.3 | 12.7 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.3 | 2.8 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.3 | 0.9 | GO:0047936 | glucose 1-dehydrogenase [NAD(P)] activity(GO:0047936) |
| 0.3 | 2.6 | GO:0004331 | fructose-2,6-bisphosphate 2-phosphatase activity(GO:0004331) |
| 0.2 | 1.0 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.2 | 8.4 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.2 | 4.4 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.2 | 3.2 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.2 | 1.3 | GO:0000253 | 3-keto sterol reductase activity(GO:0000253) |
| 0.2 | 2.2 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.2 | 10.0 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.2 | 1.5 | GO:0047499 | calcium-independent phospholipase A2 activity(GO:0047499) |
| 0.1 | 0.4 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.1 | 1.0 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.1 | 4.1 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.1 | 0.4 | GO:0031177 | phosphopantetheine binding(GO:0031177) |
| 0.1 | 2.8 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 0.6 | GO:0016433 | rRNA (adenine) methyltransferase activity(GO:0016433) |
| 0.1 | 9.8 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 0.5 | GO:0003996 | acyl-CoA ligase activity(GO:0003996) |
| 0.1 | 2.8 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.1 | 0.5 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.1 | 0.2 | GO:0001160 | transcription termination site sequence-specific DNA binding(GO:0001147) transcription termination site DNA binding(GO:0001160) |
| 0.1 | 0.9 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.1 | 0.7 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.0 | 0.3 | GO:0036402 | proteasome-activating ATPase activity(GO:0036402) |
| 0.0 | 0.1 | GO:0000386 | second spliceosomal transesterification activity(GO:0000386) |
| 0.0 | 0.2 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) |
| 0.0 | 0.1 | GO:0090555 | phosphatidylethanolamine-translocating ATPase activity(GO:0090555) |
| 0.0 | 0.4 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
| 0.0 | 2.6 | GO:0008028 | monocarboxylic acid transmembrane transporter activity(GO:0008028) |
| 0.0 | 0.2 | GO:0030297 | transmembrane receptor protein tyrosine kinase activator activity(GO:0030297) |
| 0.0 | 0.3 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 0.0 | 0.3 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.0 | 0.3 | GO:0003854 | 3-beta-hydroxy-delta5-steroid dehydrogenase activity(GO:0003854) |
| 0.0 | 1.5 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 2.7 | GO:0004866 | endopeptidase inhibitor activity(GO:0004866) |
| 0.0 | 0.7 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 3.0 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.0 | 0.1 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.0 | 1.4 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.0 | 1.3 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 3.0 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.0 | 0.1 | GO:0034597 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) phosphatidylinositol-4,5-bisphosphate 4-phosphatase activity(GO:0034597) |
| 0.0 | 1.3 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.0 | 2.0 | GO:0008080 | N-acetyltransferase activity(GO:0008080) |
| 0.0 | 1.4 | GO:0019905 | syntaxin binding(GO:0019905) |
| 0.0 | 0.4 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.0 | 0.3 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.0 | 3.7 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.0 | 2.1 | GO:0004386 | helicase activity(GO:0004386) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 8.3 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.1 | 1.0 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.0 | 1.2 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.0 | 4.1 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 2.2 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 1.2 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 0.7 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 0.9 | PID MET PATHWAY | Signaling events mediated by Hepatocyte Growth Factor Receptor (c-Met) |
| 0.0 | 1.3 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.0 | 1.2 | SIG PIP3 SIGNALING IN CARDIAC MYOCTES | Genes related to PIP3 signaling in cardiac myocytes |
| 0.0 | 0.9 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 8.8 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.2 | 6.4 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.2 | 4.4 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.1 | 3.7 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.1 | 1.3 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.1 | 1.5 | REACTOME ACYL CHAIN REMODELLING OF PE | Genes involved in Acyl chain remodelling of PE |
| 0.1 | 4.4 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 2.2 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.1 | 0.7 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.0 | 1.2 | REACTOME OXYGEN DEPENDENT PROLINE HYDROXYLATION OF HYPOXIA INDUCIBLE FACTOR ALPHA | Genes involved in Oxygen-dependent Proline Hydroxylation of Hypoxia-inducible Factor Alpha |
| 0.0 | 1.4 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.0 | 0.7 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.0 | 2.0 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.9 | REACTOME CTLA4 INHIBITORY SIGNALING | Genes involved in CTLA4 inhibitory signaling |
| 0.0 | 0.9 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 0.1 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.0 | 1.4 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.0 | 0.4 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 0.5 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |