Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
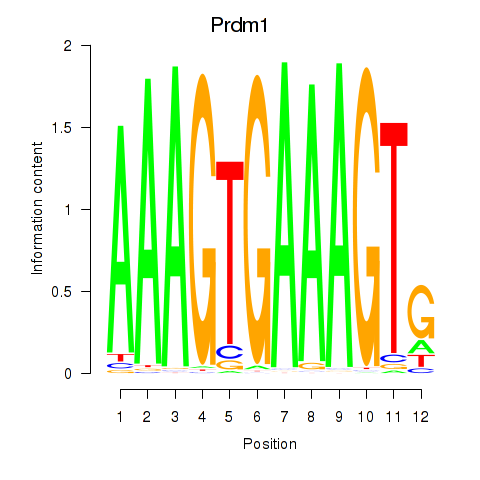
Results for Prdm1
Z-value: 0.86
Transcription factors associated with Prdm1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Prdm1
|
ENSMUSG00000038151.14 | PR domain containing 1, with ZNF domain |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Prdm1 | mm39_v1_chr10_-_44334711_44334747 | -0.23 | 1.9e-01 | Click! |
Activity profile of Prdm1 motif
Sorted Z-values of Prdm1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_40078132 | 8.55 |
ENSMUST00000068094.13
ENSMUST00000080171.3 |
Cyp2c50
|
cytochrome P450, family 2, subfamily c, polypeptide 50 |
| chr6_-_55152002 | 7.99 |
ENSMUST00000003569.6
|
Inmt
|
indolethylamine N-methyltransferase |
| chr15_+_77613239 | 7.99 |
ENSMUST00000230979.2
ENSMUST00000109775.4 |
Apol9b
|
apolipoprotein L 9b |
| chr15_+_77613348 | 5.85 |
ENSMUST00000230742.2
|
Apol9b
|
apolipoprotein L 9b |
| chr5_+_31078775 | 4.92 |
ENSMUST00000201621.4
|
Khk
|
ketohexokinase |
| chr8_-_45786226 | 4.45 |
ENSMUST00000095328.6
|
Cyp4v3
|
cytochrome P450, family 4, subfamily v, polypeptide 3 |
| chr15_+_33083358 | 4.27 |
ENSMUST00000228916.2
ENSMUST00000226483.2 ENSMUST00000228737.2 |
Cpq
|
carboxypeptidase Q |
| chr6_-_48549594 | 4.11 |
ENSMUST00000009425.7
ENSMUST00000204267.3 ENSMUST00000204930.3 ENSMUST00000204182.2 |
Rarres2
|
retinoic acid receptor responder (tazarotene induced) 2 |
| chr5_+_31078911 | 3.66 |
ENSMUST00000201571.4
|
Khk
|
ketohexokinase |
| chr3_-_113368407 | 3.58 |
ENSMUST00000106540.8
|
Amy1
|
amylase 1, salivary |
| chr14_+_40827317 | 3.48 |
ENSMUST00000047286.7
|
Mat1a
|
methionine adenosyltransferase I, alpha |
| chr11_-_5900019 | 3.26 |
ENSMUST00000102920.4
|
Gck
|
glucokinase |
| chr8_+_70536103 | 3.19 |
ENSMUST00000007738.11
|
Hapln4
|
hyaluronan and proteoglycan link protein 4 |
| chr3_+_129630380 | 3.17 |
ENSMUST00000077918.7
|
Cfi
|
complement component factor i |
| chr14_+_40826970 | 3.14 |
ENSMUST00000225720.2
|
Mat1a
|
methionine adenosyltransferase I, alpha |
| chr14_+_40827108 | 3.02 |
ENSMUST00000224514.2
|
Mat1a
|
methionine adenosyltransferase I, alpha |
| chr8_-_93956143 | 2.90 |
ENSMUST00000176282.2
ENSMUST00000034173.14 |
Ces1e
|
carboxylesterase 1E |
| chr15_+_25843225 | 2.88 |
ENSMUST00000022881.15
|
Retreg1
|
reticulophagy regulator 1 |
| chr8_-_85526653 | 2.84 |
ENSMUST00000126806.2
ENSMUST00000076715.13 |
Nfix
|
nuclear factor I/X |
| chr9_-_106353792 | 2.78 |
ENSMUST00000214682.2
ENSMUST00000112479.9 |
Parp3
|
poly (ADP-ribose) polymerase family, member 3 |
| chr4_-_107928567 | 2.72 |
ENSMUST00000106701.2
|
Scp2
|
sterol carrier protein 2, liver |
| chr15_+_100202642 | 2.70 |
ENSMUST00000067752.5
ENSMUST00000229588.2 |
Mettl7a1
|
methyltransferase like 7A1 |
| chr6_+_117145496 | 2.50 |
ENSMUST00000112866.8
ENSMUST00000112871.8 ENSMUST00000073043.5 |
Cxcl12
|
chemokine (C-X-C motif) ligand 12 |
| chr10_+_116137277 | 2.49 |
ENSMUST00000092167.7
|
Ptprb
|
protein tyrosine phosphatase, receptor type, B |
| chr9_-_106353571 | 2.45 |
ENSMUST00000123555.8
ENSMUST00000125850.2 |
Parp3
|
poly (ADP-ribose) polymerase family, member 3 |
| chr4_-_42756489 | 2.45 |
ENSMUST00000140546.3
ENSMUST00000102957.6 |
Ccl19
|
chemokine (C-C motif) ligand 19 |
| chr11_-_73215442 | 2.45 |
ENSMUST00000021119.9
|
Aspa
|
aspartoacylase |
| chr3_+_81839908 | 2.29 |
ENSMUST00000029649.3
|
Ctso
|
cathepsin O |
| chr9_-_107546166 | 2.28 |
ENSMUST00000177567.8
|
Slc38a3
|
solute carrier family 38, member 3 |
| chr4_-_42773987 | 2.25 |
ENSMUST00000095114.5
|
Ccl21a
|
chemokine (C-C motif) ligand 21A (serine) |
| chr8_-_85526972 | 2.24 |
ENSMUST00000099070.10
|
Nfix
|
nuclear factor I/X |
| chr2_+_43445333 | 2.22 |
ENSMUST00000028223.9
ENSMUST00000112826.8 |
Kynu
|
kynureninase |
| chr6_-_23839136 | 2.12 |
ENSMUST00000166458.9
ENSMUST00000142913.9 ENSMUST00000069074.14 ENSMUST00000115361.9 ENSMUST00000018122.14 ENSMUST00000115356.3 |
Cadps2
|
Ca2+-dependent activator protein for secretion 2 |
| chr11_+_120421496 | 2.08 |
ENSMUST00000026119.8
|
Gcgr
|
glucagon receptor |
| chr11_-_120618052 | 2.06 |
ENSMUST00000106148.10
ENSMUST00000026144.5 |
Dcxr
|
dicarbonyl L-xylulose reductase |
| chr3_+_122688721 | 1.99 |
ENSMUST00000023820.6
|
Fabp2
|
fatty acid binding protein 2, intestinal |
| chr8_+_127790772 | 1.95 |
ENSMUST00000079777.12
ENSMUST00000160272.8 ENSMUST00000162907.8 ENSMUST00000162536.8 ENSMUST00000026921.13 ENSMUST00000162665.8 ENSMUST00000162602.8 ENSMUST00000160581.8 ENSMUST00000161355.8 ENSMUST00000162531.8 ENSMUST00000160766.8 ENSMUST00000159537.8 |
Pard3
|
par-3 family cell polarity regulator |
| chr13_+_74787952 | 1.90 |
ENSMUST00000221822.2
ENSMUST00000221526.2 |
Erap1
|
endoplasmic reticulum aminopeptidase 1 |
| chr17_-_32643067 | 1.87 |
ENSMUST00000237130.2
|
Pglyrp2
|
peptidoglycan recognition protein 2 |
| chr12_+_37292029 | 1.78 |
ENSMUST00000160390.2
|
Agmo
|
alkylglycerol monooxygenase |
| chr2_+_43445359 | 1.77 |
ENSMUST00000050511.7
|
Kynu
|
kynureninase |
| chr15_+_32920869 | 1.73 |
ENSMUST00000022871.7
|
Sdc2
|
syndecan 2 |
| chr9_-_44714263 | 1.71 |
ENSMUST00000044694.8
|
Ttc36
|
tetratricopeptide repeat domain 36 |
| chr3_-_107952146 | 1.66 |
ENSMUST00000178808.8
ENSMUST00000106670.2 ENSMUST00000029489.15 |
Gstm4
|
glutathione S-transferase, mu 4 |
| chr14_-_73622638 | 1.66 |
ENSMUST00000228637.2
ENSMUST00000022704.9 |
Itm2b
|
integral membrane protein 2B |
| chr11_+_102993865 | 1.62 |
ENSMUST00000152971.2
|
Acbd4
|
acyl-Coenzyme A binding domain containing 4 |
| chr14_-_45715308 | 1.61 |
ENSMUST00000141424.2
|
Fermt2
|
fermitin family member 2 |
| chr9_-_107546195 | 1.60 |
ENSMUST00000192990.6
|
Slc38a3
|
solute carrier family 38, member 3 |
| chr10_+_128158413 | 1.58 |
ENSMUST00000219836.2
|
Cnpy2
|
canopy FGF signaling regulator 2 |
| chr1_+_139429430 | 1.55 |
ENSMUST00000027615.7
|
F13b
|
coagulation factor XIII, beta subunit |
| chr3_+_145464413 | 1.53 |
ENSMUST00000029845.15
|
Ddah1
|
dimethylarginine dimethylaminohydrolase 1 |
| chr12_+_37291632 | 1.51 |
ENSMUST00000049874.14
|
Agmo
|
alkylglycerol monooxygenase |
| chr3_+_142406827 | 1.44 |
ENSMUST00000044392.11
ENSMUST00000199519.5 |
Kyat3
|
kynurenine aminotransferase 3 |
| chr13_-_12479804 | 1.43 |
ENSMUST00000124888.8
|
Lgals8
|
lectin, galactose binding, soluble 8 |
| chr18_-_13013030 | 1.40 |
ENSMUST00000119512.8
|
Osbpl1a
|
oxysterol binding protein-like 1A |
| chr11_-_77784922 | 1.38 |
ENSMUST00000017597.5
|
Pipox
|
pipecolic acid oxidase |
| chr10_-_95159933 | 1.37 |
ENSMUST00000053594.7
|
Cradd
|
CASP2 and RIPK1 domain containing adaptor with death domain |
| chr10_+_128158328 | 1.36 |
ENSMUST00000219037.2
ENSMUST00000026446.4 |
Cnpy2
|
canopy FGF signaling regulator 2 |
| chr1_-_184578057 | 1.36 |
ENSMUST00000068725.10
|
Mtarc2
|
mitochondrial amidoxime reducing component 2 |
| chr17_+_34138611 | 1.34 |
ENSMUST00000234247.2
|
Tapbp
|
TAP binding protein |
| chr2_+_68966125 | 1.34 |
ENSMUST00000041865.8
|
Nostrin
|
nitric oxide synthase trafficker |
| chr16_-_30086317 | 1.34 |
ENSMUST00000064856.9
|
Cpn2
|
carboxypeptidase N, polypeptide 2 |
| chr17_-_34406193 | 1.33 |
ENSMUST00000173831.3
|
Psmb9
|
proteasome (prosome, macropain) subunit, beta type 9 (large multifunctional peptidase 2) |
| chr3_-_101511971 | 1.33 |
ENSMUST00000036493.8
|
Atp1a1
|
ATPase, Na+/K+ transporting, alpha 1 polypeptide |
| chr14_+_64890008 | 1.32 |
ENSMUST00000224503.2
|
Kif13b
|
kinesin family member 13B |
| chr13_+_33188511 | 1.32 |
ENSMUST00000006391.5
|
Serpinb9
|
serine (or cysteine) peptidase inhibitor, clade B, member 9 |
| chr3_+_142406787 | 1.32 |
ENSMUST00000106218.8
|
Kyat3
|
kynurenine aminotransferase 3 |
| chr12_+_37291728 | 1.31 |
ENSMUST00000160768.8
|
Agmo
|
alkylglycerol monooxygenase |
| chr15_-_89263448 | 1.26 |
ENSMUST00000049968.9
|
Odf3b
|
outer dense fiber of sperm tails 3B |
| chr4_+_42114817 | 1.24 |
ENSMUST00000098123.4
|
Gm13304
|
predicted gene 13304 |
| chr14_-_25927674 | 1.24 |
ENSMUST00000100811.6
|
Tmem254a
|
transmembrane protein 254a |
| chr15_-_89263790 | 1.23 |
ENSMUST00000238996.2
|
Odf3b
|
outer dense fiber of sperm tails 3B |
| chr5_-_105387395 | 1.23 |
ENSMUST00000065588.7
|
Gbp10
|
guanylate-binding protein 10 |
| chr13_+_24023428 | 1.22 |
ENSMUST00000091698.12
ENSMUST00000110422.3 ENSMUST00000166467.9 |
Slc17a3
|
solute carrier family 17 (sodium phosphate), member 3 |
| chr1_-_106687457 | 1.22 |
ENSMUST00000010049.6
|
Kdsr
|
3-ketodihydrosphingosine reductase |
| chr4_-_25281750 | 1.20 |
ENSMUST00000038705.8
ENSMUST00000102994.10 |
Ufl1
|
UFM1 specific ligase 1 |
| chr17_+_37504783 | 1.20 |
ENSMUST00000038844.7
|
Ubd
|
ubiquitin D |
| chr5_+_92285748 | 1.19 |
ENSMUST00000031355.10
ENSMUST00000202155.2 |
Uso1
|
USO1 vesicle docking factor |
| chr4_-_96479793 | 1.17 |
ENSMUST00000055693.9
|
Cyp2j9
|
cytochrome P450, family 2, subfamily j, polypeptide 9 |
| chr4_+_41903610 | 1.17 |
ENSMUST00000098128.4
|
Ccl21d
|
chemokine (C-C motif) ligand 21D |
| chr4_-_148236516 | 1.15 |
ENSMUST00000056965.12
ENSMUST00000168503.8 ENSMUST00000152098.8 |
Fbxo6
|
F-box protein 6 |
| chr14_+_52122299 | 1.15 |
ENSMUST00000047899.13
ENSMUST00000164902.8 |
Mettl17
|
methyltransferase like 17 |
| chr4_+_42255693 | 1.15 |
ENSMUST00000178864.3
|
Ccl21b
|
chemokine (C-C motif) ligand 21B (leucine) |
| chr14_+_64889973 | 1.13 |
ENSMUST00000100473.5
|
Kif13b
|
kinesin family member 13B |
| chr17_+_34138699 | 1.12 |
ENSMUST00000234320.2
|
Tapbp
|
TAP binding protein |
| chr2_+_85850977 | 1.11 |
ENSMUST00000213774.2
ENSMUST00000214546.2 ENSMUST00000215682.2 |
Olfr1033
|
olfactory receptor 1033 |
| chr3_+_106629209 | 1.08 |
ENSMUST00000106736.3
ENSMUST00000154973.2 ENSMUST00000098750.5 ENSMUST00000150513.2 |
Lrif1
|
ligand dependent nuclear receptor interacting factor 1 |
| chr14_-_101878106 | 1.07 |
ENSMUST00000100339.9
|
Commd6
|
COMM domain containing 6 |
| chr16_-_35691914 | 1.07 |
ENSMUST00000042665.9
|
Parp14
|
poly (ADP-ribose) polymerase family, member 14 |
| chr1_+_177269845 | 1.07 |
ENSMUST00000195002.2
|
Zbtb18
|
zinc finger and BTB domain containing 18 |
| chr14_-_55909527 | 1.05 |
ENSMUST00000010520.10
|
Nedd8
|
neural precursor cell expressed, developmentally down-regulated gene 8 |
| chr2_+_11710101 | 1.04 |
ENSMUST00000138349.8
ENSMUST00000135341.8 ENSMUST00000128156.9 |
Il15ra
|
interleukin 15 receptor, alpha chain |
| chr17_-_13135232 | 1.03 |
ENSMUST00000079121.4
|
Mrpl18
|
mitochondrial ribosomal protein L18 |
| chr2_-_155661324 | 1.02 |
ENSMUST00000124586.2
|
BC029722
|
cDNA sequence BC029722 |
| chr8_+_89423645 | 1.02 |
ENSMUST00000043526.15
ENSMUST00000211554.2 ENSMUST00000209532.2 ENSMUST00000209559.2 |
Cyld
|
CYLD lysine 63 deubiquitinase |
| chr5_+_115061293 | 1.02 |
ENSMUST00000031540.11
ENSMUST00000112143.4 |
Oasl1
|
2'-5' oligoadenylate synthetase-like 1 |
| chr1_+_91226058 | 0.98 |
ENSMUST00000027532.13
|
Scly
|
selenocysteine lyase |
| chr3_-_65300000 | 0.97 |
ENSMUST00000029414.12
|
Ssr3
|
signal sequence receptor, gamma |
| chr13_+_24023386 | 0.95 |
ENSMUST00000039721.14
|
Slc17a3
|
solute carrier family 17 (sodium phosphate), member 3 |
| chr8_+_95161006 | 0.93 |
ENSMUST00000211816.2
|
Nlrc5
|
NLR family, CARD domain containing 5 |
| chr16_-_38533597 | 0.92 |
ENSMUST00000023487.5
|
Arhgap31
|
Rho GTPase activating protein 31 |
| chr11_-_120464062 | 0.91 |
ENSMUST00000026122.11
|
P4hb
|
prolyl 4-hydroxylase, beta polypeptide |
| chr13_-_92620507 | 0.90 |
ENSMUST00000040106.9
|
Fam151b
|
family with sequence similarity 151, member B |
| chr1_-_183766195 | 0.89 |
ENSMUST00000050306.8
|
1700056E22Rik
|
RIKEN cDNA 1700056E22 gene |
| chr4_+_42612121 | 0.86 |
ENSMUST00000178168.3
|
Gm10591
|
predicted gene 10591 |
| chr14_-_101877870 | 0.85 |
ENSMUST00000168587.3
|
Commd6
|
COMM domain containing 6 |
| chr8_-_106665060 | 0.84 |
ENSMUST00000034369.10
|
Psmb10
|
proteasome (prosome, macropain) subunit, beta type 10 |
| chr17_+_34043536 | 0.83 |
ENSMUST00000048249.8
|
Ndufa7
|
NADH:ubiquinone oxidoreductase subunit A7 |
| chr2_+_11710523 | 0.83 |
ENSMUST00000138856.2
ENSMUST00000078834.12 ENSMUST00000114834.10 ENSMUST00000114833.10 ENSMUST00000114831.9 |
Il15ra
|
interleukin 15 receptor, alpha chain |
| chr4_-_45489794 | 0.83 |
ENSMUST00000146236.8
|
Shb
|
src homology 2 domain-containing transforming protein B |
| chr6_+_40619913 | 0.83 |
ENSMUST00000238599.2
|
Mgam
|
maltase-glucoamylase |
| chr1_-_172418058 | 0.82 |
ENSMUST00000065679.8
|
Slamf8
|
SLAM family member 8 |
| chr11_-_120463667 | 0.80 |
ENSMUST00000168360.2
|
P4hb
|
prolyl 4-hydroxylase, beta polypeptide |
| chr13_+_104953679 | 0.80 |
ENSMUST00000022230.15
|
Srek1ip1
|
splicing regulatory glutamine/lysine-rich protein 1interacting protein 1 |
| chr6_-_39095144 | 0.79 |
ENSMUST00000038398.7
|
Parp12
|
poly (ADP-ribose) polymerase family, member 12 |
| chr11_-_53750016 | 0.79 |
ENSMUST00000117316.8
ENSMUST00000120776.8 ENSMUST00000121435.2 |
Gm12216
|
predicted gene 12216 |
| chr2_-_51862941 | 0.78 |
ENSMUST00000145481.8
ENSMUST00000112705.9 |
Nmi
|
N-myc (and STAT) interactor |
| chr2_+_11710246 | 0.78 |
ENSMUST00000148748.8
|
Il15ra
|
interleukin 15 receptor, alpha chain |
| chr7_+_100355798 | 0.78 |
ENSMUST00000107042.9
ENSMUST00000207564.2 ENSMUST00000049053.9 |
Fam168a
|
family with sequence similarity 168, member A |
| chr17_-_30845845 | 0.77 |
ENSMUST00000235547.2
ENSMUST00000188852.2 |
Glo1
1700097N02Rik
|
glyoxalase 1 RIKEN cDNA 1700097N02 gene |
| chr10_-_75633362 | 0.77 |
ENSMUST00000120177.8
|
Gstt1
|
glutathione S-transferase, theta 1 |
| chr7_-_19297001 | 0.76 |
ENSMUST00000058444.10
|
Ppp1r37
|
protein phosphatase 1, regulatory subunit 37 |
| chr17_+_25114090 | 0.76 |
ENSMUST00000043907.14
|
Mrps34
|
mitochondrial ribosomal protein S34 |
| chr9_-_106353303 | 0.75 |
ENSMUST00000156426.8
|
Parp3
|
poly (ADP-ribose) polymerase family, member 3 |
| chr17_+_85264134 | 0.74 |
ENSMUST00000112305.10
|
Ppm1b
|
protein phosphatase 1B, magnesium dependent, beta isoform |
| chr1_+_183766572 | 0.73 |
ENSMUST00000048655.8
|
Dusp10
|
dual specificity phosphatase 10 |
| chr11_-_70350783 | 0.73 |
ENSMUST00000019064.9
|
Cxcl16
|
chemokine (C-X-C motif) ligand 16 |
| chr3_+_106389732 | 0.72 |
ENSMUST00000029508.11
|
Dennd2d
|
DENN/MADD domain containing 2D |
| chr3_+_106628987 | 0.72 |
ENSMUST00000130105.2
|
Lrif1
|
ligand dependent nuclear receptor interacting factor 1 |
| chrX_-_151820545 | 0.71 |
ENSMUST00000051484.5
|
Mageh1
|
MAGE family member H1 |
| chr10_-_75633563 | 0.71 |
ENSMUST00000139724.3
|
Gstt1
|
glutathione S-transferase, theta 1 |
| chr14_-_51134930 | 0.70 |
ENSMUST00000227271.2
|
Klhl33
|
kelch-like 33 |
| chr9_+_54858066 | 0.70 |
ENSMUST00000034848.14
|
Psma4
|
proteasome subunit alpha 4 |
| chr11_+_48967411 | 0.68 |
ENSMUST00000109202.3
|
Ifi47
|
interferon gamma inducible protein 47 |
| chr9_+_38629560 | 0.67 |
ENSMUST00000001544.12
ENSMUST00000118144.8 |
Vwa5a
|
von Willebrand factor A domain containing 5A |
| chr2_-_101451383 | 0.67 |
ENSMUST00000090513.11
|
Iftap
|
intraflagellar transport associated protein |
| chr10_-_116732813 | 0.65 |
ENSMUST00000048229.9
|
Myrfl
|
myelin regulatory factor-like |
| chr7_+_130972867 | 0.64 |
ENSMUST00000075610.13
|
Pstk
|
phosphoseryl-tRNA kinase |
| chr5_-_93354348 | 0.63 |
ENSMUST00000058550.15
|
Ccni
|
cyclin I |
| chr15_-_77480311 | 0.63 |
ENSMUST00000089465.6
|
Apol10b
|
apolipoprotein L 10B |
| chr8_+_93553901 | 0.63 |
ENSMUST00000034187.9
|
Mmp2
|
matrix metallopeptidase 2 |
| chr16_+_35892437 | 0.62 |
ENSMUST00000163352.9
ENSMUST00000231468.2 |
Ccdc58
|
coiled-coil domain containing 58 |
| chr3_+_142326363 | 0.61 |
ENSMUST00000165774.8
|
Gbp2
|
guanylate binding protein 2 |
| chr15_+_66763169 | 0.61 |
ENSMUST00000005255.9
|
Ccn4
|
cellular communication network factor 4 |
| chrX_+_160500051 | 0.60 |
ENSMUST00000112338.2
|
Rai2
|
retinoic acid induced 2 |
| chr2_+_31462659 | 0.60 |
ENSMUST00000113482.8
|
Fubp3
|
far upstream element (FUSE) binding protein 3 |
| chr17_-_36206812 | 0.59 |
ENSMUST00000059740.15
|
2310061I04Rik
|
RIKEN cDNA 2310061I04 gene |
| chr15_+_90108480 | 0.59 |
ENSMUST00000100309.3
ENSMUST00000231200.2 |
Alg10b
|
asparagine-linked glycosylation 10B (alpha-1,2-glucosyltransferase) |
| chr19_-_6134903 | 0.59 |
ENSMUST00000160977.8
ENSMUST00000159859.2 ENSMUST00000025707.9 ENSMUST00000160712.8 ENSMUST00000237738.2 |
Zfpl1
|
zinc finger like protein 1 |
| chr18_+_77031788 | 0.59 |
ENSMUST00000097522.11
ENSMUST00000097521.12 ENSMUST00000145634.9 |
Hdhd2
|
haloacid dehalogenase-like hydrolase domain containing 2 |
| chr5_-_92582969 | 0.59 |
ENSMUST00000135112.8
|
Nup54
|
nucleoporin 54 |
| chr9_-_123461593 | 0.58 |
ENSMUST00000026273.11
|
Slc6a20b
|
solute carrier family 6 (neurotransmitter transporter), member 20B |
| chr9_+_54858092 | 0.58 |
ENSMUST00000172407.8
|
Psma4
|
proteasome subunit alpha 4 |
| chr14_-_51134906 | 0.58 |
ENSMUST00000170855.2
|
Klhl33
|
kelch-like 33 |
| chr7_-_18852282 | 0.58 |
ENSMUST00000141380.3
|
Meiosin
|
meiosis initiator |
| chr14_-_55909314 | 0.57 |
ENSMUST00000163750.8
|
Nedd8
|
neural precursor cell expressed, developmentally down-regulated gene 8 |
| chr1_+_85177316 | 0.57 |
ENSMUST00000161424.5
ENSMUST00000113402.4 |
Gm7609
|
predicted pseudogene 7609 |
| chr2_+_65499097 | 0.57 |
ENSMUST00000200829.4
|
Scn2a
|
sodium channel, voltage-gated, type II, alpha |
| chr15_-_89263466 | 0.56 |
ENSMUST00000228111.2
|
Odf3b
|
outer dense fiber of sperm tails 3B |
| chr3_-_88332401 | 0.56 |
ENSMUST00000168755.7
ENSMUST00000193433.6 ENSMUST00000195657.6 ENSMUST00000057935.9 |
Slc25a44
|
solute carrier family 25, member 44 |
| chr11_+_119158713 | 0.56 |
ENSMUST00000106259.9
|
Gaa
|
glucosidase, alpha, acid |
| chr2_+_11710633 | 0.56 |
ENSMUST00000114832.3
|
Il15ra
|
interleukin 15 receptor, alpha chain |
| chr2_-_51863203 | 0.55 |
ENSMUST00000028314.9
|
Nmi
|
N-myc (and STAT) interactor |
| chr8_-_11528615 | 0.54 |
ENSMUST00000033900.7
|
Rab20
|
RAB20, member RAS oncogene family |
| chr8_+_47192767 | 0.54 |
ENSMUST00000034041.9
ENSMUST00000208507.2 ENSMUST00000207105.2 |
Irf2
|
interferon regulatory factor 2 |
| chr16_+_43184191 | 0.54 |
ENSMUST00000156367.8
|
Zbtb20
|
zinc finger and BTB domain containing 20 |
| chr17_+_31514780 | 0.54 |
ENSMUST00000237460.2
|
Slc37a1
|
solute carrier family 37 (glycerol-3-phosphate transporter), member 1 |
| chr16_+_10307589 | 0.53 |
ENSMUST00000230892.2
ENSMUST00000230146.2 ENSMUST00000230392.2 |
Ciita
|
class II transactivator |
| chr17_-_26004507 | 0.53 |
ENSMUST00000140738.8
ENSMUST00000145053.2 ENSMUST00000138759.8 ENSMUST00000133071.8 ENSMUST00000077938.10 |
Haghl
|
hydroxyacylglutathione hydrolase-like |
| chr2_-_90735171 | 0.53 |
ENSMUST00000005647.4
|
Ndufs3
|
NADH:ubiquinone oxidoreductase core subunit S3 |
| chr6_-_125208738 | 0.51 |
ENSMUST00000043422.8
|
Tapbpl
|
TAP binding protein-like |
| chr5_+_28370687 | 0.50 |
ENSMUST00000036177.9
|
En2
|
engrailed 2 |
| chr14_+_55909692 | 0.50 |
ENSMUST00000002397.7
|
Gmpr2
|
guanosine monophosphate reductase 2 |
| chr16_-_38253507 | 0.50 |
ENSMUST00000002926.8
|
Pla1a
|
phospholipase A1 member A |
| chr5_-_93354287 | 0.50 |
ENSMUST00000144514.3
|
Ccni
|
cyclin I |
| chr3_-_115800989 | 0.49 |
ENSMUST00000067485.4
|
Slc30a7
|
solute carrier family 30 (zinc transporter), member 7 |
| chr9_+_44394080 | 0.49 |
ENSMUST00000220303.2
|
Bcl9l
|
B cell CLL/lymphoma 9-like |
| chr14_-_30850881 | 0.49 |
ENSMUST00000203261.3
|
Smim4
|
small integral membrane protein 4 |
| chr17_-_37334091 | 0.49 |
ENSMUST00000167275.3
|
Mog
|
myelin oligodendrocyte glycoprotein |
| chr7_+_29883569 | 0.49 |
ENSMUST00000098594.4
|
Cox7a1
|
cytochrome c oxidase subunit 7A1 |
| chr8_-_84059048 | 0.48 |
ENSMUST00000177594.8
ENSMUST00000053902.4 |
Elmod2
|
ELMO/CED-12 domain containing 2 |
| chr11_+_119158829 | 0.47 |
ENSMUST00000026666.13
ENSMUST00000106258.2 |
Gaa
|
glucosidase, alpha, acid |
| chr10_-_29411857 | 0.47 |
ENSMUST00000092623.5
|
Rspo3
|
R-spondin 3 |
| chr11_-_98478018 | 0.46 |
ENSMUST00000052919.8
|
Ormdl3
|
ORM1-like 3 (S. cerevisiae) |
| chr13_+_22017906 | 0.46 |
ENSMUST00000180288.2
|
H2bc24
|
H2B clustered histone 24 |
| chr17_-_32253467 | 0.46 |
ENSMUST00000002145.12
ENSMUST00000133308.3 |
Hsf2bp
|
heat shock transcription factor 2 binding protein |
| chr5_-_127709961 | 0.46 |
ENSMUST00000155321.2
|
Slc15a4
|
solute carrier family 15, member 4 |
| chr8_-_45863572 | 0.46 |
ENSMUST00000209651.2
ENSMUST00000211370.2 ENSMUST00000034056.12 ENSMUST00000167106.3 |
Tlr3
|
toll-like receptor 3 |
| chr12_+_8258107 | 0.45 |
ENSMUST00000037383.13
ENSMUST00000218883.2 ENSMUST00000218086.2 ENSMUST00000169104.3 ENSMUST00000217999.2 |
Ldah
|
lipid droplet associated hydrolase |
| chr16_+_97337275 | 0.45 |
ENSMUST00000024112.8
|
Mx2
|
MX dynamin-like GTPase 2 |
| chr7_-_7212997 | 0.45 |
ENSMUST00000074455.9
|
Zfp772
|
zinc finger protein 772 |
| chr2_+_97298002 | 0.44 |
ENSMUST00000059049.8
|
Lrrc4c
|
leucine rich repeat containing 4C |
| chr4_+_123077515 | 0.43 |
ENSMUST00000152194.2
|
Hpcal4
|
hippocalcin-like 4 |
| chr1_+_180978491 | 0.43 |
ENSMUST00000134115.8
ENSMUST00000111059.2 |
Cnih4
|
cornichon family AMPA receptor auxiliary protein 4 |
| chr12_+_71021395 | 0.43 |
ENSMUST00000160027.8
ENSMUST00000160864.8 |
Psma3
|
proteasome subunit alpha 3 |
| chr1_+_85454323 | 0.43 |
ENSMUST00000239236.2
|
Gm7592
|
predicted gene 7592 |
| chr16_-_18882012 | 0.42 |
ENSMUST00000199490.2
|
Iglj1
|
immunoglobulin lambda joining 1 |
| chr7_+_29883611 | 0.42 |
ENSMUST00000208441.2
|
Cox7a1
|
cytochrome c oxidase subunit 7A1 |
| chr6_-_8180174 | 0.42 |
ENSMUST00000213284.2
|
Col28a1
|
collagen, type XXVIII, alpha 1 |
| chr18_-_66155651 | 0.41 |
ENSMUST00000143990.2
|
Lman1
|
lectin, mannose-binding, 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Prdm1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.2 | 9.6 | GO:0009087 | methionine catabolic process(GO:0009087) |
| 2.1 | 8.6 | GO:0034285 | response to sucrose(GO:0009744) response to disaccharide(GO:0034285) |
| 1.6 | 4.7 | GO:2000547 | regulation of dendritic cell dendrite assembly(GO:2000547) |
| 1.3 | 4.0 | GO:0019442 | tryptophan catabolic process to acetyl-CoA(GO:0019442) |
| 1.3 | 3.9 | GO:2000487 | asparagine transport(GO:0006867) positive regulation of glutamine transport(GO:2000487) |
| 1.0 | 6.0 | GO:1990166 | protein localization to site of double-strand break(GO:1990166) |
| 0.7 | 8.6 | GO:0043651 | linoleic acid metabolic process(GO:0043651) |
| 0.7 | 2.1 | GO:0042732 | D-xylose metabolic process(GO:0042732) |
| 0.7 | 4.1 | GO:0061760 | antifungal innate immune response(GO:0061760) |
| 0.7 | 2.7 | GO:0032382 | positive regulation of intracellular lipid transport(GO:0032379) positive regulation of intracellular sterol transport(GO:0032382) positive regulation of intracellular cholesterol transport(GO:0032385) lipid hydroperoxide transport(GO:1901373) |
| 0.7 | 3.3 | GO:0009732 | detection of carbohydrate stimulus(GO:0009730) detection of hexose stimulus(GO:0009732) detection of monosaccharide stimulus(GO:0034287) detection of glucose(GO:0051594) |
| 0.6 | 2.5 | GO:1903237 | negative regulation of leukocyte tethering or rolling(GO:1903237) |
| 0.6 | 2.5 | GO:0050823 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-dependent(GO:0002479) peptide stabilization(GO:0050822) peptide antigen stabilization(GO:0050823) |
| 0.5 | 0.5 | GO:0002590 | regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002589) negative regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002590) |
| 0.5 | 1.5 | GO:0018900 | dichloromethane metabolic process(GO:0018900) |
| 0.5 | 1.4 | GO:0006553 | lysine metabolic process(GO:0006553) |
| 0.4 | 1.3 | GO:1902524 | positive regulation of protein K48-linked ubiquitination(GO:1902524) |
| 0.4 | 2.9 | GO:0045716 | positive regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045716) |
| 0.4 | 4.6 | GO:0046485 | ether lipid metabolic process(GO:0046485) |
| 0.3 | 2.4 | GO:0006083 | acetate metabolic process(GO:0006083) |
| 0.3 | 1.0 | GO:0035928 | rRNA import into mitochondrion(GO:0035928) |
| 0.3 | 1.0 | GO:0043181 | vacuolar sequestering(GO:0043181) |
| 0.3 | 1.7 | GO:0018916 | nitrobenzene metabolic process(GO:0018916) |
| 0.3 | 2.9 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.3 | 1.9 | GO:0000270 | peptidoglycan metabolic process(GO:0000270) peptidoglycan catabolic process(GO:0009253) negative regulation of natural killer cell differentiation(GO:0032824) negative regulation of natural killer cell differentiation involved in immune response(GO:0032827) |
| 0.3 | 1.9 | GO:0003383 | apical constriction(GO:0003383) |
| 0.3 | 0.8 | GO:1902623 | negative regulation of neutrophil migration(GO:1902623) |
| 0.3 | 1.3 | GO:1903416 | response to glycoside(GO:1903416) |
| 0.3 | 2.1 | GO:1990504 | dense core granule exocytosis(GO:1990504) |
| 0.3 | 1.3 | GO:0042270 | protection from natural killer cell mediated cytotoxicity(GO:0042270) |
| 0.3 | 1.0 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 0.3 | 2.1 | GO:0033762 | response to glucagon(GO:0033762) |
| 0.3 | 4.3 | GO:0006590 | thyroid hormone generation(GO:0006590) |
| 0.2 | 5.1 | GO:0021684 | cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.2 | 1.2 | GO:1990592 | protein polyufmylation(GO:1990564) protein K69-linked ufmylation(GO:1990592) |
| 0.2 | 0.9 | GO:0036228 | protein targeting to nuclear inner membrane(GO:0036228) |
| 0.2 | 2.8 | GO:0070189 | kynurenine metabolic process(GO:0070189) |
| 0.2 | 1.4 | GO:0002317 | plasma cell differentiation(GO:0002317) |
| 0.2 | 1.0 | GO:0097343 | ripoptosome assembly(GO:0097343) ripoptosome assembly involved in necroptotic process(GO:1901026) |
| 0.2 | 0.8 | GO:1905053 | regulation of base-excision repair(GO:1905051) positive regulation of base-excision repair(GO:1905053) |
| 0.2 | 1.7 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.2 | 1.9 | GO:0019885 | antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.2 | 0.5 | GO:0046038 | GMP catabolic process(GO:0046038) |
| 0.2 | 0.9 | GO:0060335 | positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) |
| 0.2 | 0.5 | GO:0015817 | histidine transport(GO:0015817) |
| 0.1 | 2.2 | GO:0015747 | urate transport(GO:0015747) |
| 0.1 | 0.8 | GO:0045354 | interferon-alpha biosynthetic process(GO:0045349) regulation of interferon-alpha biosynthetic process(GO:0045354) |
| 0.1 | 1.0 | GO:0016259 | selenocysteine metabolic process(GO:0016259) |
| 0.1 | 1.3 | GO:0019243 | methylglyoxal catabolic process to D-lactate via S-lactoyl-glutathione(GO:0019243) methylglyoxal catabolic process(GO:0051596) methylglyoxal catabolic process to lactate(GO:0061727) |
| 0.1 | 0.6 | GO:0014012 | peripheral nervous system axon regeneration(GO:0014012) |
| 0.1 | 0.7 | GO:0060266 | negative regulation of respiratory burst involved in inflammatory response(GO:0060266) |
| 0.1 | 1.5 | GO:0006527 | arginine catabolic process(GO:0006527) |
| 0.1 | 0.3 | GO:0090071 | negative regulation of ribosome biogenesis(GO:0090071) |
| 0.1 | 0.5 | GO:1900060 | negative regulation of ceramide biosynthetic process(GO:1900060) |
| 0.1 | 0.3 | GO:0042138 | meiotic DNA double-strand break formation(GO:0042138) |
| 0.1 | 2.8 | GO:0032825 | positive regulation of natural killer cell differentiation(GO:0032825) |
| 0.1 | 0.7 | GO:0006499 | N-terminal protein myristoylation(GO:0006499) |
| 0.1 | 0.4 | GO:0000966 | RNA 5'-end processing(GO:0000966) mitochondrial tRNA processing(GO:0090646) |
| 0.1 | 1.2 | GO:0070842 | aggresome assembly(GO:0070842) |
| 0.1 | 0.4 | GO:0090156 | lysosomal microautophagy(GO:0016237) piecemeal microautophagy of nucleus(GO:0034727) suppression by virus of host autophagy(GO:0039521) amino acid homeostasis(GO:0080144) negative regulation of sphingolipid biosynthetic process(GO:0090155) cellular sphingolipid homeostasis(GO:0090156) negative regulation of sphingolipid biosynthesis involved in cellular sphingolipid homeostasis(GO:0090157) |
| 0.1 | 0.5 | GO:0090383 | phagosome acidification(GO:0090383) |
| 0.1 | 1.2 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.1 | 0.8 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.1 | 1.4 | GO:0006977 | DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest(GO:0006977) |
| 0.1 | 0.8 | GO:1900194 | negative regulation of oocyte maturation(GO:1900194) |
| 0.1 | 0.4 | GO:0051581 | negative regulation of neurotransmitter uptake(GO:0051581) negative regulation of serotonin uptake(GO:0051612) |
| 0.1 | 0.3 | GO:2000836 | positive regulation of androgen secretion(GO:2000836) |
| 0.1 | 0.6 | GO:0001514 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.1 | 0.3 | GO:0002121 | inter-male aggressive behavior(GO:0002121) |
| 0.1 | 0.3 | GO:0010636 | positive regulation of mitochondrial fusion(GO:0010636) |
| 0.1 | 1.0 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.1 | 0.5 | GO:0045343 | MHC class I biosynthetic process(GO:0045341) regulation of MHC class I biosynthetic process(GO:0045343) positive regulation of MHC class I biosynthetic process(GO:0045345) |
| 0.1 | 0.4 | GO:0042985 | negative regulation of amyloid precursor protein biosynthetic process(GO:0042985) |
| 0.1 | 1.5 | GO:0006516 | glycoprotein catabolic process(GO:0006516) |
| 0.1 | 0.2 | GO:1904430 | negative regulation of t-circle formation(GO:1904430) |
| 0.1 | 0.2 | GO:0002182 | cytoplasmic translational elongation(GO:0002182) regulation of cytoplasmic translational elongation(GO:1900247) negative regulation of cytoplasmic translational elongation(GO:1900248) |
| 0.1 | 1.6 | GO:0045116 | protein neddylation(GO:0045116) |
| 0.1 | 0.6 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.1 | 1.2 | GO:0046519 | sphingoid metabolic process(GO:0046519) |
| 0.0 | 2.0 | GO:0050892 | intestinal absorption(GO:0050892) |
| 0.0 | 0.3 | GO:0019364 | pyridine nucleotide catabolic process(GO:0019364) |
| 0.0 | 0.5 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.0 | 0.3 | GO:0007341 | penetration of zona pellucida(GO:0007341) |
| 0.0 | 0.4 | GO:0010700 | negative regulation of norepinephrine secretion(GO:0010700) |
| 0.0 | 0.8 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 3.0 | GO:0071346 | cellular response to interferon-gamma(GO:0071346) |
| 0.0 | 8.0 | GO:0009308 | amine metabolic process(GO:0009308) |
| 0.0 | 0.4 | GO:0006561 | proline biosynthetic process(GO:0006561) |
| 0.0 | 0.3 | GO:0035694 | mitochondrial protein catabolic process(GO:0035694) |
| 0.0 | 0.3 | GO:0010701 | positive regulation of norepinephrine secretion(GO:0010701) |
| 0.0 | 0.4 | GO:0072602 | interleukin-4 secretion(GO:0072602) |
| 0.0 | 0.2 | GO:1904784 | NLRP1 inflammasome complex assembly(GO:1904784) |
| 0.0 | 0.3 | GO:0007039 | protein catabolic process in the vacuole(GO:0007039) |
| 0.0 | 0.6 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.0 | 0.3 | GO:0072619 | interleukin-21 production(GO:0032625) interleukin-21 secretion(GO:0072619) |
| 0.0 | 4.4 | GO:0019395 | fatty acid oxidation(GO:0019395) |
| 0.0 | 0.5 | GO:0061088 | sequestering of zinc ion(GO:0032119) regulation of sequestering of zinc ion(GO:0061088) |
| 0.0 | 0.3 | GO:0045198 | establishment of epithelial cell apical/basal polarity(GO:0045198) |
| 0.0 | 0.3 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.0 | 0.6 | GO:1990403 | embryonic brain development(GO:1990403) |
| 0.0 | 0.2 | GO:0070268 | cornification(GO:0070268) |
| 0.0 | 0.7 | GO:0010818 | T cell chemotaxis(GO:0010818) |
| 0.0 | 0.2 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.0 | 0.2 | GO:0051771 | negative regulation of nitric-oxide synthase biosynthetic process(GO:0051771) |
| 0.0 | 0.6 | GO:0002360 | T cell lineage commitment(GO:0002360) |
| 0.0 | 0.4 | GO:0031573 | intra-S DNA damage checkpoint(GO:0031573) |
| 0.0 | 0.5 | GO:0060670 | branching involved in labyrinthine layer morphogenesis(GO:0060670) |
| 0.0 | 3.6 | GO:0016052 | carbohydrate catabolic process(GO:0016052) |
| 0.0 | 0.3 | GO:0070102 | interleukin-6-mediated signaling pathway(GO:0070102) |
| 0.0 | 0.2 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.0 | 0.2 | GO:0035457 | cellular response to interferon-alpha(GO:0035457) |
| 0.0 | 0.7 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.0 | 0.1 | GO:0015755 | fructose transport(GO:0015755) |
| 0.0 | 0.1 | GO:0015722 | canalicular bile acid transport(GO:0015722) |
| 0.0 | 0.1 | GO:0002838 | T cell mediated immune response to tumor cell(GO:0002424) negative regulation of response to tumor cell(GO:0002835) negative regulation of immune response to tumor cell(GO:0002838) regulation of T cell mediated immune response to tumor cell(GO:0002840) |
| 0.0 | 0.1 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.0 | 0.1 | GO:0036337 | Fas signaling pathway(GO:0036337) |
| 0.0 | 0.2 | GO:0061303 | cornea development in camera-type eye(GO:0061303) |
| 0.0 | 0.3 | GO:0043383 | negative T cell selection(GO:0043383) |
| 0.0 | 0.5 | GO:0090077 | foam cell differentiation(GO:0090077) |
| 0.0 | 0.3 | GO:0035634 | response to stilbenoid(GO:0035634) |
| 0.0 | 0.1 | GO:0051490 | negative regulation of filopodium assembly(GO:0051490) |
| 0.0 | 1.6 | GO:0048041 | cell-substrate adherens junction assembly(GO:0007045) focal adhesion assembly(GO:0048041) |
| 0.0 | 0.1 | GO:0045065 | cytotoxic T cell differentiation(GO:0045065) |
| 0.0 | 1.0 | GO:0021766 | hippocampus development(GO:0021766) |
| 0.0 | 0.2 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.0 | 0.8 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 1.2 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.0 | 1.7 | GO:0048814 | regulation of dendrite morphogenesis(GO:0048814) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.2 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.3 | 1.7 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 0.3 | 2.5 | GO:0042825 | TAP complex(GO:0042825) |
| 0.2 | 1.9 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.2 | 2.0 | GO:0045179 | apical cortex(GO:0045179) |
| 0.2 | 2.2 | GO:0031315 | extrinsic component of mitochondrial outer membrane(GO:0031315) |
| 0.2 | 3.3 | GO:0045180 | basal cortex(GO:0045180) |
| 0.2 | 0.5 | GO:1902560 | GMP reductase complex(GO:1902560) |
| 0.2 | 1.7 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 0.1 | 1.3 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.1 | 6.0 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.1 | 1.5 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.1 | 0.5 | GO:0035339 | SPOTS complex(GO:0035339) |
| 0.1 | 1.0 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.1 | 0.9 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.1 | 3.0 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.1 | 0.4 | GO:0048476 | Holliday junction resolvase complex(GO:0048476) |
| 0.1 | 0.2 | GO:0043564 | Ku70:Ku80 complex(GO:0043564) |
| 0.1 | 1.0 | GO:0033655 | host cell cytoplasm(GO:0030430) host cell cytoplasm part(GO:0033655) |
| 0.0 | 1.2 | GO:0012507 | ER to Golgi transport vesicle membrane(GO:0012507) |
| 0.0 | 0.6 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.0 | 2.7 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 1.4 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.0 | 5.9 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 2.9 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 1.7 | GO:0030660 | Golgi-associated vesicle membrane(GO:0030660) |
| 0.0 | 0.1 | GO:0046691 | intracellular canaliculus(GO:0046691) |
| 0.0 | 0.3 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.0 | 2.1 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 0.2 | GO:0072558 | NLRP1 inflammasome complex(GO:0072558) |
| 0.0 | 1.4 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.9 | GO:0005746 | mitochondrial respiratory chain(GO:0005746) |
| 0.0 | 1.6 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.0 | 0.7 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 0.0 | 0.3 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.0 | 2.1 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 0.9 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 1.1 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 1.1 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.0 | 1.3 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 0.4 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 0.1 | GO:0030134 | ER to Golgi transport vesicle(GO:0030134) |
| 0.0 | 0.4 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.0 | 23.0 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.9 | 8.6 | GO:0004454 | ketohexokinase activity(GO:0004454) |
| 2.4 | 9.6 | GO:0004478 | methionine adenosyltransferase activity(GO:0004478) |
| 2.1 | 8.6 | GO:0071614 | linoleic acid epoxygenase activity(GO:0071614) |
| 1.5 | 4.6 | GO:0050479 | glyceryl-ether monooxygenase activity(GO:0050479) |
| 1.0 | 8.0 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 1.0 | 4.0 | GO:0016823 | hydrolase activity, acting on acid carbon-carbon bonds(GO:0016822) hydrolase activity, acting on acid carbon-carbon bonds, in ketonic substances(GO:0016823) |
| 1.0 | 3.9 | GO:0015182 | L-asparagine transmembrane transporter activity(GO:0015182) |
| 0.8 | 2.4 | GO:0019807 | aspartoacylase activity(GO:0019807) |
| 0.7 | 2.1 | GO:0004967 | glucagon receptor activity(GO:0004967) |
| 0.7 | 2.7 | GO:0050632 | propanoyl-CoA C-acyltransferase activity(GO:0033814) propionyl-CoA C2-trimethyltridecanoyltransferase activity(GO:0050632) phosphatidylethanolamine transporter activity(GO:1904121) |
| 0.6 | 2.4 | GO:0031735 | CCR10 chemokine receptor binding(GO:0031735) |
| 0.6 | 4.3 | GO:0070573 | metallodipeptidase activity(GO:0070573) |
| 0.6 | 6.0 | GO:0003910 | DNA ligase activity(GO:0003909) DNA ligase (ATP) activity(GO:0003910) |
| 0.6 | 2.8 | GO:0047804 | cysteine-S-conjugate beta-lyase activity(GO:0047804) |
| 0.5 | 3.6 | GO:0004556 | alpha-amylase activity(GO:0004556) |
| 0.5 | 1.5 | GO:0019120 | hydrolase activity, acting on acid halide bonds(GO:0016824) hydrolase activity, acting on acid halide bonds, in C-halide compounds(GO:0019120) alkylhalidase activity(GO:0047651) |
| 0.4 | 0.8 | GO:0016160 | amylase activity(GO:0016160) |
| 0.4 | 2.5 | GO:0046979 | TAP1 binding(GO:0046978) TAP2 binding(GO:0046979) |
| 0.3 | 3.3 | GO:0004340 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) |
| 0.3 | 1.0 | GO:0070279 | vitamin B6 binding(GO:0070279) |
| 0.3 | 1.9 | GO:0008745 | N-acetylmuramoyl-L-alanine amidase activity(GO:0008745) peptidoglycan receptor activity(GO:0016019) |
| 0.3 | 1.7 | GO:0004656 | procollagen-proline 4-dioxygenase activity(GO:0004656) |
| 0.3 | 2.2 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 0.3 | 1.0 | GO:0004574 | oligo-1,6-glucosidase activity(GO:0004574) maltose alpha-glucosidase activity(GO:0032450) |
| 0.2 | 1.4 | GO:0016647 | oxidoreductase activity, acting on the CH-NH group of donors, oxygen as acceptor(GO:0016647) |
| 0.2 | 0.5 | GO:0005290 | L-histidine transmembrane transporter activity(GO:0005290) |
| 0.2 | 2.0 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.2 | 2.5 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.2 | 1.9 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.2 | 0.5 | GO:0016657 | GMP reductase activity(GO:0003920) oxidoreductase activity, acting on NAD(P)H, nitrogenous group as acceptor(GO:0016657) |
| 0.1 | 3.9 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.1 | 4.0 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.1 | 1.5 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.1 | 2.7 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.1 | 0.4 | GO:0016426 | tRNA (adenine) methyltransferase activity(GO:0016426) |
| 0.1 | 1.0 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.1 | 0.8 | GO:0015315 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.1 | 1.3 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) steroid hormone binding(GO:1990239) |
| 0.1 | 2.8 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 0.4 | GO:0019779 | Atg12 activating enzyme activity(GO:0019778) Atg8 activating enzyme activity(GO:0019779) |
| 0.1 | 0.8 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.1 | 1.0 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.1 | 1.5 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.1 | 0.5 | GO:0008970 | phosphatidylcholine 1-acylhydrolase activity(GO:0008970) |
| 0.1 | 1.9 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.1 | 3.4 | GO:0016655 | oxidoreductase activity, acting on NAD(P)H, quinone or similar compound as acceptor(GO:0016655) |
| 0.1 | 0.4 | GO:0004735 | pyrroline-5-carboxylate reductase activity(GO:0004735) |
| 0.1 | 1.0 | GO:0008097 | 5S rRNA binding(GO:0008097) |
| 0.1 | 3.6 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 0.5 | GO:0004416 | hydroxyacylglutathione hydrolase activity(GO:0004416) |
| 0.1 | 1.4 | GO:0070513 | death domain binding(GO:0070513) |
| 0.1 | 0.3 | GO:0003829 | beta-1,3-galactosyl-O-glycosyl-glycoprotein beta-1,6-N-acetylglucosaminyltransferase activity(GO:0003829) |
| 0.1 | 0.3 | GO:0030284 | estrogen receptor activity(GO:0030284) |
| 0.1 | 2.5 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.1 | 1.2 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.0 | 3.7 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.0 | 0.6 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.0 | 0.7 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.0 | 0.4 | GO:0008821 | crossover junction endodeoxyribonuclease activity(GO:0008821) |
| 0.0 | 0.3 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.0 | 0.4 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 0.0 | 0.6 | GO:0046527 | glucosyltransferase activity(GO:0046527) |
| 0.0 | 0.1 | GO:0047936 | glucose 1-dehydrogenase [NAD(P)] activity(GO:0047936) |
| 0.0 | 1.7 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 0.6 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.0 | 0.9 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.4 | GO:0001609 | G-protein coupled adenosine receptor activity(GO:0001609) |
| 0.0 | 0.2 | GO:0004661 | protein geranylgeranyltransferase activity(GO:0004661) |
| 0.0 | 0.3 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.0 | 4.5 | GO:0020037 | heme binding(GO:0020037) |
| 0.0 | 0.3 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.0 | 0.9 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.2 | GO:0048019 | receptor antagonist activity(GO:0048019) |
| 0.0 | 1.1 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.2 | GO:0051575 | 5'-deoxyribose-5-phosphate lyase activity(GO:0051575) |
| 0.0 | 1.3 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.0 | 0.1 | GO:0008147 | structural constituent of bone(GO:0008147) |
| 0.0 | 0.2 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.0 | 1.8 | GO:0042974 | retinoic acid receptor binding(GO:0042974) |
| 0.0 | 2.5 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 1.7 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 0.4 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.0 | 0.3 | GO:0003854 | 3-beta-hydroxy-delta5-steroid dehydrogenase activity(GO:0003854) |
| 0.0 | 1.4 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 0.8 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.0 | 0.6 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.0 | 0.4 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 2.5 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 0.3 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.0 | 0.1 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.0 | 0.8 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.2 | GO:0046703 | natural killer cell lectin-like receptor binding(GO:0046703) |
| 0.0 | 0.8 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 0.1 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.0 | 0.2 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.0 | 0.8 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.0 | 0.7 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.0 | 0.5 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 0.4 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.3 | GO:0046961 | proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 3.2 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.1 | 4.0 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.1 | 2.4 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.1 | 2.5 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 1.2 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 0.8 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.8 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.0 | 1.0 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 6.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.1 | PID HIV NEF PATHWAY | HIV-1 Nef: Negative effector of Fas and TNF-alpha |
| 0.0 | 1.9 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 1.7 | ST INTEGRIN SIGNALING PATHWAY | Integrin Signaling Pathway |
| 0.0 | 0.1 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.0 | 0.4 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 1.1 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.0 | 1.0 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 0.7 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 4.0 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.3 | 9.6 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.2 | 3.2 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.2 | 0.8 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.1 | 2.7 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 7ALPHA HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 7alpha-hydroxycholesterol |
| 0.1 | 1.5 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.1 | 5.7 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 1.9 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.1 | 3.3 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 1.7 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.1 | 2.0 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.1 | 1.9 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.1 | 1.7 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.1 | 0.8 | REACTOME TRAF6 MEDIATED INDUCTION OF TAK1 COMPLEX | Genes involved in TRAF6 mediated induction of TAK1 complex |
| 0.1 | 3.9 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.1 | 1.6 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 1.3 | REACTOME ENOS ACTIVATION AND REGULATION | Genes involved in eNOS activation and regulation |
| 0.0 | 8.8 | REACTOME METABOLISM OF CARBOHYDRATES | Genes involved in Metabolism of carbohydrates |
| 0.0 | 0.6 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 1.6 | REACTOME CDK MEDIATED PHOSPHORYLATION AND REMOVAL OF CDC6 | Genes involved in CDK-mediated phosphorylation and removal of Cdc6 |
| 0.0 | 1.3 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 1.2 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.6 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 1.3 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.8 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.0 | 0.7 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.0 | 1.0 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 0.5 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.0 | 0.4 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 1.2 | REACTOME GLUCAGON SIGNALING IN METABOLIC REGULATION | Genes involved in Glucagon signaling in metabolic regulation |
| 0.0 | 0.5 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.0 | 0.5 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 1.4 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.6 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 0.2 | REACTOME INTEGRATION OF PROVIRUS | Genes involved in Integration of provirus |
| 0.0 | 0.6 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.0 | 0.8 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.0 | 0.6 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.3 | REACTOME REGULATION OF INSULIN SECRETION BY GLUCAGON LIKE PEPTIDE1 | Genes involved in Regulation of Insulin Secretion by Glucagon-like Peptide-1 |