Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
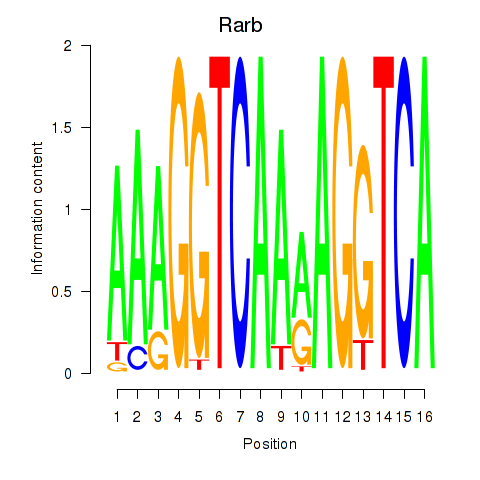
Results for Rarb
Z-value: 0.46
Transcription factors associated with Rarb
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Rarb
|
ENSMUSG00000017491.10 | retinoic acid receptor, beta |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Rarb | mm39_v1_chr14_+_5894220_5894254 | 0.18 | 2.8e-01 | Click! |
Activity profile of Rarb motif
Sorted Z-values of Rarb motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_-_85797946 | 2.77 |
ENSMUST00000032074.5
|
Nat8f5
|
N-acetyltransferase 8 (GCN5-related) family member 5 |
| chr5_-_87074380 | 1.68 |
ENSMUST00000031183.3
|
Ugt2b1
|
UDP glucuronosyltransferase 2 family, polypeptide B1 |
| chr12_+_8027767 | 1.43 |
ENSMUST00000037520.14
|
Apob
|
apolipoprotein B |
| chr12_+_8027640 | 1.39 |
ENSMUST00000171271.8
ENSMUST00000037811.13 |
Apob
|
apolipoprotein B |
| chr5_-_87288177 | 1.24 |
ENSMUST00000067790.7
|
Ugt2b5
|
UDP glucuronosyltransferase 2 family, polypeptide B5 |
| chr5_-_87054796 | 1.02 |
ENSMUST00000031181.16
ENSMUST00000113333.2 |
Ugt2b34
|
UDP glucuronosyltransferase 2 family, polypeptide B34 |
| chr7_-_115630282 | 0.80 |
ENSMUST00000206034.2
ENSMUST00000106612.8 |
Sox6
|
SRY (sex determining region Y)-box 6 |
| chr1_+_4878046 | 0.78 |
ENSMUST00000027036.11
ENSMUST00000150971.8 ENSMUST00000119612.9 ENSMUST00000137887.8 ENSMUST00000115529.8 |
Lypla1
|
lysophospholipase 1 |
| chr5_-_33375509 | 0.61 |
ENSMUST00000201475.2
|
Spon2
|
spondin 2, extracellular matrix protein |
| chr19_+_28967875 | 0.56 |
ENSMUST00000224599.2
ENSMUST00000050148.5 |
Cdc37l1
|
cell division cycle 37-like 1 |
| chr1_+_191553556 | 0.49 |
ENSMUST00000027931.8
|
Nek2
|
NIMA (never in mitosis gene a)-related expressed kinase 2 |
| chr6_+_108771840 | 0.45 |
ENSMUST00000204483.2
|
Arl8b
|
ADP-ribosylation factor-like 8B |
| chr10_-_92558219 | 0.45 |
ENSMUST00000020163.7
|
Nedd1
|
neural precursor cell expressed, developmentally down-regulated gene 1 |
| chr17_+_12803019 | 0.40 |
ENSMUST00000046959.9
ENSMUST00000233066.2 |
Slc22a2
|
solute carrier family 22 (organic cation transporter), member 2 |
| chr10_+_67021509 | 0.39 |
ENSMUST00000173689.8
|
Jmjd1c
|
jumonji domain containing 1C |
| chr2_+_3771709 | 0.35 |
ENSMUST00000177037.2
|
Fam107b
|
family with sequence similarity 107, member B |
| chr6_-_85797917 | 0.35 |
ENSMUST00000174143.2
|
Nat8f6
|
N-acetyltransferase 8 (GCN5-related) family member 6 |
| chr7_+_35148461 | 0.23 |
ENSMUST00000118969.8
|
Slc7a9
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 9 |
| chr2_+_30171055 | 0.23 |
ENSMUST00000143119.3
|
Gm28038
|
predicted gene, 28038 |
| chr10_-_126737185 | 0.22 |
ENSMUST00000168520.3
ENSMUST00000026504.13 |
Atp23
|
ATP23 metallopeptidase and ATP synthase assembly factor homolog |
| chr7_+_35148579 | 0.19 |
ENSMUST00000032703.10
|
Slc7a9
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 9 |
| chr7_+_35148188 | 0.19 |
ENSMUST00000118383.8
|
Slc7a9
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 9 |
| chr11_+_105069591 | 0.13 |
ENSMUST00000106939.9
|
Tlk2
|
tousled-like kinase 2 (Arabidopsis) |
| chr19_+_44980565 | 0.12 |
ENSMUST00000179305.2
|
Sema4g
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4G |
| chr7_+_45267024 | 0.09 |
ENSMUST00000008605.6
ENSMUST00000209379.2 |
Fut1
|
fucosyltransferase 1 |
| chr15_-_98560739 | 0.08 |
ENSMUST00000162384.2
ENSMUST00000003450.15 |
Ddx23
|
DEAD box helicase 23 |
| chr7_+_127503251 | 0.05 |
ENSMUST00000071056.14
|
Bckdk
|
branched chain ketoacid dehydrogenase kinase |
| chr2_-_163259012 | 0.04 |
ENSMUST00000127038.2
|
Oser1
|
oxidative stress responsive serine rich 1 |
| chr4_-_118314647 | 0.03 |
ENSMUST00000106375.2
ENSMUST00000168404.9 ENSMUST00000006556.11 |
Mpl
|
myeloproliferative leukemia virus oncogene |
| chr19_-_8763609 | 0.02 |
ENSMUST00000177216.8
ENSMUST00000176610.9 ENSMUST00000177056.8 |
Taf6l
|
TATA-box binding protein associated factor 6 like |
| chr11_+_76795346 | 0.02 |
ENSMUST00000072633.4
|
Tmigd1
|
transmembrane and immunoglobulin domain containing 1 |
| chr11_-_71092124 | 0.02 |
ENSMUST00000108514.10
|
Nlrp1b
|
NLR family, pyrin domain containing 1B |
| chr19_-_8763771 | 0.01 |
ENSMUST00000176496.8
|
Taf6l
|
TATA-box binding protein associated factor 6 like |
Network of associatons between targets according to the STRING database.
First level regulatory network of Rarb
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.7 | GO:0018879 | biphenyl metabolic process(GO:0018879) |
| 0.2 | 2.8 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 0.2 | 0.6 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.2 | 0.6 | GO:0015811 | L-cystine transport(GO:0015811) |
| 0.1 | 0.8 | GO:0002084 | protein depalmitoylation(GO:0002084) macromolecule depalmitoylation(GO:0098734) |
| 0.1 | 0.5 | GO:0030997 | regulation of centriole-centriole cohesion(GO:0030997) |
| 0.1 | 0.8 | GO:2000741 | positive regulation of mesenchymal stem cell differentiation(GO:2000741) |
| 0.1 | 2.8 | GO:0001702 | gastrulation with mouth forming second(GO:0001702) |
| 0.0 | 0.4 | GO:0030718 | germ-line stem cell population maintenance(GO:0030718) |
| 0.0 | 0.4 | GO:0015697 | quaternary ammonium group transport(GO:0015697) |
| 0.0 | 0.5 | GO:0071539 | protein localization to centrosome(GO:0071539) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 2.8 | GO:0034359 | mature chylomicron(GO:0034359) |
| 0.1 | 0.5 | GO:0000931 | gamma-tubulin large complex(GO:0000931) gamma-tubulin ring complex(GO:0008274) |
| 0.0 | 0.2 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.8 | GO:0035473 | lipase binding(GO:0035473) |
| 0.2 | 0.6 | GO:0015184 | L-cystine transmembrane transporter activity(GO:0015184) |
| 0.1 | 0.4 | GO:0005277 | acetylcholine transmembrane transporter activity(GO:0005277) norepinephrine transmembrane transporter activity(GO:0005333) acetate ester transmembrane transporter activity(GO:1901375) |
| 0.1 | 3.9 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.1 | 0.8 | GO:0098599 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.0 | 0.1 | GO:0031127 | galactoside 2-alpha-L-fucosyltransferase activity(GO:0008107) alpha-(1,2)-fucosyltransferase activity(GO:0031127) |
| 0.0 | 2.8 | GO:0008080 | N-acetyltransferase activity(GO:0008080) |
| 0.0 | 0.6 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.0 | 0.4 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.8 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 0.6 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.8 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.4 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 0.5 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.0 | 0.8 | REACTOME ENOS ACTIVATION AND REGULATION | Genes involved in eNOS activation and regulation |
| 0.0 | 0.5 | REACTOME APC CDC20 MEDIATED DEGRADATION OF NEK2A | Genes involved in APC-Cdc20 mediated degradation of Nek2A |
| 0.0 | 0.6 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |